Abstract
Future barley cultivars will have to produce under the constraints of higher temperature in combination with increased concentrations of atmospheric carbon dioxide and ozone as a consequence of climate change. A diverse set of 167 spring barley genotypes was cultivated under elevated levels of temperature (+5 °C) and [CO2] (700 ppm) as single factors and in combination as well as under elevated [O3] (100–150 ppb) as single factor. The setting in general resembled changes projected by IPCC (AR5) to take place at the end of this century. A genome-wide association study (GWAS) was performed to identify markers for increased primary production under climate change conditions and reveal possible genes of interest. Phenotyped traits included grain yield, number of grains, number of ears per plant, aboveground vegetative biomass, harvest index and stability of the production parameters over the five applied treatments. The GWAS encompassed 7864 SNP markers (Illumina iselect), a compressed mixed linear model with the GAPIT package, and conservative validation of markers. A total of 60 marker-trait associations [−log10(P value) 2.97–5.58] were identified, e.g. grain yield under elevated temperature on barley chromosome 2H, static stability of grain yield on 7H, sites for exploitation of elevated [CO2] on 4H and 7H and associations under the two-factor treatment. Marker-trait associations identified from single-factor treatments were not retrieved, when elevated [CO2] and temperature were combined emphasizing the need for multifactor experiments. This GWA study identified markers and chromosome regions to be targeted in breeding for development of climate resilient cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rapid changes in growth environments induced by altered climate conditions urge the need to breed climate resilient crop cultivars. Traditionally, characterization and subsequent introduction of genes into elite germplasm have increased tolerance to pest and pathogens. However, in a future climate of increasing temperatures as well as elevated atmospheric concentration of carbon dioxide ([CO2]) and ozone ([O3]), abiotic stresses demand a more extensive focus in crop improvement. Therefore, molecular targeting of genes providing tolerance to abiotic stresses is an important tool in the development of climate resilient cultivars.
Barley (Hordeum vulgare L.) is an economically important crop plant widely used as feed for livestock and as ingredient in beverage and food products for human consumption (e.g. Newton et al. 2011). Studies have reported high phenotypic plasticity in wild barley, which may explain its large geographical distribution and wide adaptation to diverse environmental conditions (Nevo et al. 2012). As a diploid, inbreeding, temperate crop, barley has traditionally been considered a model for plant genetic research. Large collections of germplasm containing geographically diverse elite varieties, landraces and wild accessions are readily available and possibly contain alleles that could counteract or even exploit the projected future increases in temperature, [CO2] and [O3] (The International Barley Genome Sequencing Consortium 2012).
Marker-assisted selection, where markers for desired agricultural traits are applied to verify loci related to a phenotype, is a method to accelerate plant breeding. However, robust phenotyping and reliable markers are needed. In recent years, the cost of genotyping has decreased considerably (Wetterstrand 2014) and accelerated the identification of markers associated with agricultural traits encoded by quantitative trait loci (QTL) in segregating progenies, as well as in diverse sets of accessions through genome-wide association study (GWAS). The emergence of high-throughput SNP genotyping platforms has enabled the implementation of GWAS in barley (Close et al. 2009; Waugh et al. 2009; Comadran et al. 2012). The advantage of GWAS to traditional bi-parental mapping is that alleles present within diverse sets of accessions can be identified, and not only alleles present in the parents of segregating crosses (Zhu et al. 2008). To avoid an inflated rate of false positives, the frequency of an allele within the set of accessions must reach an adequate level (minor allele frequency) to be revealed. Hence, the minor allele frequency limits the use of GWAS to find rare alleles (Tabangin et al. 2009). As GWAS analyses the linkage disequilibrium (LD) between marker loci, the genetic structure (relatedness) within the set of accessions must be accounted for.
LD is found to be extensive in barley (Caldwell et al. 2006; Comadran et al. 2011), and therefore, barley is well fitted for identification of marker-trait associations by GWAS with a reasonable number of markers. Numerous studies have reported QTL and marker-trait associations for agricultural traits in barley (e.g. Kraakman et al. 2004; Schweizer and Stein 2011; Varshney et al. 2012; Tondelli et al. 2013); other studies led to a better understanding of plant tolerance to stresses, e.g. salt and aluminium (Cai et al. 2013; Long et al. 2013). Exploring QTL and marker-trait associations under projected climate change conditions could improve the understanding of genes and processes operating under such conditions. However, phenotypic responses to elevated temperature, [CO2] and/or [O3] have not yet been included in a GWAS.
Already, climate changes have been found to cause a 5.5 % decrease in global wheat yield in the period from 1980 to 2008 (Lobell et al. 2011). According to IPCC (Intergovernmental Panel on Climate Change), the [CO2] may increase to about 1,000 ppm (mean value) at the end of the twenty-first century from approximately 400 ppm of today (Collins et al. 2013). In the same period, the [O3] is expected to increase with 8 ppb in average per year from the present levels at 32–62 ppb but with large local and seasonal variation (Collins et al. 2013; Ellermann et al. 2013). The effect of the elevated greenhouse gasses is that average temperature will increase up to +5 °C though with large regional differences; also for the occurrence of extreme climate events, local variation will occur (Collins et al. 2013).
Even though plants have a long evolutionary history of adapting to changing environmental conditions, the present climatic changes might be progressing with a pace that exceeds natural adaptation (Rosenzweig and Parry 1994; Svenning and Sandel 2013). In recent years, barley production has stagnated in Europe (FAOSTAT 2014) due to climatic, agronomic and/or socio-economic reasons. Hence, to maintain and preferable increase crop productivity to meet the future need from higher living standards and population growth (UN 2012; IPCC 2014), the cultivars of the future should be designed to possess stability, e.g. towards extreme climate shifts during the growing season, and to exploit the elevated [CO2].
In the search for markers associated with production traits, e.g. grain yield and its components as well as their stability under climate change conditions, we applied 7864 SNP markers to a set of spring barley accessions consisting of landraces, old and new cultivars as well as breeder lines. The climate treatments represented elevated levels of temperature (+5 °C), [CO2] (700 ppm) and [O3] (100–150 ppb) as single factors, and the elevated levels of temperature and [CO2] were combined in a two-factor treatment.
Materials and methods
Plant material
A total of 127 predominantly Nordic spring barley accessions of both 2- and 6-row types were included in the analysis (Table 1). Our set of accessions covered 38 landraces, 31 old cultivars (1883–1974), 53 modern cultivars (1975–2013) and 5 breeder lines. The separation into new and old cultivars was based on the introduction of exotic gene pools as disease resistance in the period after 1975 (Backes et al. 2003). Further details of the material are shown in Online Resources 1.
Phenotyping
The accessions were grown to maturity in the RERAF phytotron (http://www.eco.kt.dtu.dk/Research/Research_Facilitites/RERAF) under four climate treatments and a control. In the climate treatments temperature, [CO2] and [O3] were applied at levels projected by IPCC at the end of the twenty-first century (Collins et al. 2013). In the control treatment, ambient temperature was 19/12 °C (day/night) and [CO2] at 385 ppm. The climate treatments were (1) [CO2] at 700 ppm; (2) temperature at 24/17 °C (day/night); (3) the combination of elevated temperature (24/17 °C) and [CO2] (700 ppm); and (4) O3 added at 100–150 ppb. For each accession, eight plants were grown in 11 L pots (plant density 151 plant m−2). The pots were placed at identical positions in all treatments and rotated within and between chambers once a week during the growing period to eliminate potential chamber-specific effects. Light exposure, humidity level, water and fertilizer supply were identical between all treatments. The light regime averaged approximately 400 mol photons m−2 s−1 at canopy height (ca. 1 m) and was applied in a 16/8 h day/night cycle. The relative humidity was constant 55/70 % (day/night). Water was supplied by a dripping system at 4.4 L m−2 day−1 in the morning, and 10 g NPK fertilizer (21-3-10, Yara) was supplemented to the sphagnum substrate (Pindstrup Substrate No. 6, Pindstrup Mosebrug A/S, Denmark) at the time of sowing. Watering was reduced stepwise when 2/3 of the accessions in a treatment had reached Zadoks growth stage (ZGS) 90 and ended at ZGS 99 (Zadoks et al. 1974). Plants were harvested and threshed individually.
The production parameters measured after harvest were grain yield (GY, g plant−1), aboveground vegetative biomass (BM, g plant−1), total number of ears (ET, no. plant−1) and number of ears with grains (EG, no. plant−1). From the measured values, the following parameters were calculated: number of grains (GN, no. plant−1, GY/weight of 40 grains), harvest index (HI, %), ∆ (delta) values and static and dynamic stability. Harvest index was calculated as grain yield relative to aboveground vegetative biomass, and ∆ values were defined as the measure of a production parameter of a given accession in a climate treatment relative to the measure of this production parameter in the ambient treatment. The response to the overall production parameters in the climate treatments was calculated in % to the ambient treatment and significance according to T test; 0.001:***, 0.01:** and 0.05:* in Microsoft excel version 2010. Static stability was found according to environmental variance (S 2; Roemer 1917), and dynamic stability according to Wricke’s ecovalence (W 2; Wricke 1962). The stability measures were calculated over all five treatments for BM, ET, EG, GN and HI. S 2 is defined as:
where R ij is the observed yield of the accession i in the treatment j, m i is mean yield of the accession across treatments and e the number of environments, in the present study treatments. W 2 is defined as:
where R ij is the observed yield of the accession i in the treatment j, m i is mean yield of the accession across treatments, m j is mean yield across treatment j of all accession and m is the grand mean. Further details on experimental set-up, phenotyping of traits and details on RERAF can be found in Ingvordsen et al. (2015).
Genotyping
Genomic DNA was extracted using the CTAB procedure (Cetyl Trimethyl Ammonium Bromide; Rogers and Bendich 1985). Plant material was collected at seedling stage and freeze dried. For cultivars and breeder lines, DNA was extracted from one individual. For some of the landraces (25 out of 38), normally two but up to six individuals were included and treated as separate genotypes, however, linked to the same phenotype. Diversity found within the landrace accessions is given in Online Resources 2. In total, 192 samples were genotyped by TraitGenetics (TraitGenetics GmbH, Gatersleben, http://www.traitgenetics.com/en/) with a total of 7864 SNP (single nucleotide polymorphism) markers from the Illumina iselect array (Comadran et al. 2012).
In filtering SNP markers, maximally 50 % missing data were accepted, and an allele was considered rare when present in only 7 % of the genotypes, and the marker in question was excluded.
Genetic structure and association analysis
Genetic relatedness between accessions was analysed with the dataset including all markers using a distance matrix based on ‘Simple Matching’ and a subsequent principal coordinate analysis (PCoA). The PCoA was computed using the macro ‘Diversity’ in Microsoft Excel (2010) (programmed by G. Backes, co-author). The bayesian cluster analysis using the software ‘STRUCTURE’ (Pritchard et al. 2000) was also applied on the dataset. For the STRUCTURE analysis, a panmixis model was applied with a length of burn-in period and number of MCM repeats after burin of 10,000 in the 20 first rounds to determine the number of groups and 100,000, respectively, in the second rounds determining the Q-values. For the choice of the optimal number of groups, the method of Evanno et al. (2005) was applied. Linkage disequilibrium for a random sample of SNP marker pairs with a distance smaller than 50 cM (10 % of all linked pairs) was calculated in Microsoft Excel (2010) by the macro ‘Assoc’ (programmed by G. Backes). The calculated values were, together with a locally weighted scatterplot smoothing curve, visualized using R version 2.15.3 (R Core Team 2014).
Associated markers were identified in a two-step analysis by the use of the macro ‘Assoc.’, which calls the R-package GAPIT (Genomic Association and Prediction Integrated Tool; Lipka et al. 2012) in R version 2.15.3 (R Core Team 2014). In step one, the optimal model was identified, and in step two, the association analysis was performed with the identified model. In the optimization step, the models included two different distance matrices, one calculated by efficient mixed-model association (EMMA; Kang et al. 2008) and the other a simple-match distance matrix calculated on all polymorphic SNP markers. Each of the two models was tested including matrices with and without the vector of Q-values calculated from STRUCTURE. To choose the optimal model, a compressed mixed linear model was applied (Zhang et al. 2010) on each phenotype. Maximum likelihood and heritability of the model were calculated, as well as goodness of fit of the predicted versus the observed P values. In addition and for reason of comparison, a non-stratified naive model was applied. The optimization step was performed for each phenotyped trait with two partly overlapping datasets. One dataset included 7864 SNP markers (AllM), whereas the second dataset included the 3967 SNP markers with established position (MwP); hence, MwP was a subset of the AllM dataset. Subsequently, association analyses were performed with each of the two datasets with the optimized model including genetic structure and with the naive model.
For each phenotypic trait, the critical P values for assessing the significance of associations in the naive model were corrected for multiple comparisons using a false discovery rate (FDR = 0.05) (Benjamini and Hochberg 1995). Considering the stringency of the model used for accounting for population structure, most of the false positives were inherently controlled and no further corrections were applied. To further minimize false positives, the cut-off for accepting associations was arbitrary set to −log10(P value) > 2.95 and had to be meet by both datasets.
Linkage disequilibrium and bioinformatics of associated markers
Linkage disequilibrium (LD) between SNP markers positioned on the same chromosome was determined by TASSEL version 4.0 (Bradbury et al. 2007). When LD was found, the respective markers were treated as a LD block. The position of the LD blocks and associated markers was determined in cM and as bin according to the BinMap 2005 with its unique segregation patterns separating the bins by single recombinant events. In practice GrainGenes 2.0 (http://wheat.pw.usda.gov) was utilized, and the barley maps OPA123-2008, OPA 2011, OWB, Stein 2006 and/or OWB OPA2008 when necessary. Known genes that co-localized with SNP markers were determined through marker position from HarvEst Barley version 2.02 (http://harvest.ucr.edu; Alpert et al. 2011), IPK barley BLAST server, Gatersleben (http://webblast.ipk-gatersleben.de/barley/) and/or MIPS barley genome database (http://mips.helmholtz-muenchen.de/plant/barley/index.jsp).
Results
Effects of applied treatments to production parameters and stability
The production parameters were found to be strongly affected by the climate treatments as displayed in Table 2, and differences between cultivar responses were prominent as reported by Ingvordsen et al. (2015). Elevated temperature decreased overall grain yield (55.8 %), number of grains and also aboveground vegetative biomass and HI. Elevated [CO2] increased all production parameters apart from HI (grain yield increased 14.5 %). In the two-factor treatment, grain yield decreased (29.6 %) together with number of grains, whereas number of grain-bearing ears and total number of ears increased; the aboveground vegetative biomass was overall not influenced, and as a result, HI decreased. Elevated [O3] resulted in reduced grain yield (15.8 %), but had a positive effect on number of ears produced; aboveground vegetative biomass was unaffected and HI decreased (Table 2). The stability measures calculated over all treatments for each trait varied among the accessions as seen in Table 2. For GY, the environmental variance, S 2, ranged from 0.58 to 16.79, and the dynamic stability, W 2, ranged from 0.27 to 33.60.
Marker analysis
Genotyping succeeded for 6985 of the SNP markers, and after filtering, the AllM dataset contained 5978 polymorphic SNP markers and the MwP dataset 3679 in 167 genotypes. Heterozygotes were excluded from analysis. All phenotypic traits, except for ∆ET under elevated [O3], passed the optimization step. The LD decay of all SNP markers was found to decrease considerable after 40 cM and sufficient to perform GWAS (Online Resources 3). Only nine out of the 25 landraces with more than one individual showed above 10 % polymorphism in alleles (10.2–41.8 %, Online Resources 2).
Genetic structure and optimization
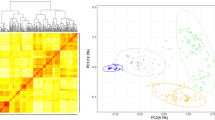
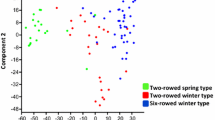
Three groups within the 167 genotypes were formed by the principal coordinate analysis (PCoA) and the Bayesian clustering (Fig. 1, Online Resources 4). The groups corresponded reasonably well with row type and time of registration at the official variety list. One group was comprised primarily by 6-rowed accessions with different year of registration (landraces, old and modern cultivars) (Fig. 1a, III), whereas the 2-rowed accessions were divided into two groups. One group of 2-rowed accessions included modern cultivars (Fig. 1a, I), and the other group included primarily old cultivars and landraces (Fig. 1a, II). No grouping could be identified by country of origin.
Principal coordinate analysis (a) and Bayesian cluster analysis (b) of 167 spring barley genotypes based on 6624 SNP markers. a I, II and III indicate potential groups. Open circle 6-rowed landrace, open triangle 6-rowed cultivar from before 1975, open square 6-rowed cultivar from after 1975, grey circle 2-rowed landrace, grey triangle 2-rowed cultivar from before 1975, grey square 2-rowed cultivar from after 1975 and filled diamond 2- or 6-rowed segregating landrace. b Bar plot sorted by Q-values from STRUCTURE at K = 3 showing three major groups. Q-values available in Online Resources 4
In the optimization step, the compressed mixed linear model (Zhang et al. 2010) with EMMA as correction for genetic structure was found to be the best fit for the majority of the phenotypes, and the model was applied in the association analyses for all phenotypes.
Marker associations and LD
Sixty associations with −log10(P value) values from 2.97 to 5.58 were found between phenotypic traits and 44 of the applied SNP markers (Table 3; Fig. 2, Online Resources 5). The marker-trait associations were predominantly identified in the single-factor treatments of elevated [CO2] and temperature for the phenotypes GY, ∆GY, ∆GN, EG and HI. Few markers were found associated with traits under the two-factor treatment or with S 2 for either of the phenotyped traits (Table 3).
In total, 12 LD blocks together with 15 single (not located to a LD block) marker-trait associations were identified (Table 3, Fig. 2, Online Resources 6). From the analysis of LD between markers on the same chromosome, 10 LD blocks included two to five marker-trait associations and two LD blocks constituted one marker each that associated with more than one phenotype. All marker-trait associations are reported in Online Resources 5. Marker-trait associations for thousand grain weight and grain per ear were identified but not reported since they generally were found to have low −log10(P values).
Generally, the LD blocks included markers associated with similar phenotypic traits under one climate treatment, e.g. LD block 2H-2 comprised markers for GY ∆GY under elevated temperature (Table 3). LD block 2H-6 and 6H-10 deviated by including markers associated with different phenotypic traits in different climatic treatments, i.e. ∆GY under elevated temperature together with ∆ET in the two-factor treatment (Table 3). Only LD block 2H-6 included markers associated with phenotypes in the two-factor treatment, as well as one of the single-factor treatments; however, the markers were not found associated with the same trait.
The LD blocks 2H-2 and 2H-3 hold markers that associated with GY, ∆GY, BM or ∆BM under elevated temperature (Table 3) indicating an area important for temperature tolerance. Also, LD block 3H-7 indicated an area related to elevated temperature with associations for ∆GN, HI and ∆GY under elevated temperature. Associations for GY, ΔGY and ΔGN under elevated [CO2] were found with six markers in LD block 4H-9 and 7H-11 pointing towards chromosome areas for exploitation of elevated [CO2]. For the two-factor treatment, associated markers were found in LD block 2H-6. Also, two single markers, SCRI_RS_144841 and SCRI_RS_213333, were found associated under the two-factor treatment with HI and ΔGN, respectively (Table 3).
Associations with climate stability across the five applied treatments for either of the production parameters were found in four of the LD blocks and with four single markers. Two of the markers were both found in LD block 7H-12 associating with S 2 of HI and GY (Table 3). One marker associated with W 2 of GY was found in LD block 7H-11 together with marker-trait associations for ∆GN, ∆GY and GY under elevated [CO2] suggesting exploitation of elevated [CO2] involved in dynamic stability of GY.
Discussion
Overall response to the applied treatments
The identified responses of the production to the climate treatments support the projected decrease in future primary production (IPCC 2014). In agreement with previous studies (e.g. Morison and Lawlor 1999; Fuhrer 2003; Bokszczanin and Fragkostefanakis 2013), elevated temperature and [O3] were found to cause decrease, and elevated [CO2] to cause increase in the grain yield; however, variation in response was identified between the accessions as described by Ingvordsen et al. (2015). Under the two-factor treatment, the most realistic future climate scenario, grain yield was decreased by 30 %, whereas quantity of aboveground vegetative biomass was generally maintained. Identification of the genomic regions involved in these responses may serve in developing cultivars that can combat effects of the future climate changes and increase our knowledge on the underlying mechanisms for a high and stable production.
Genetic structure and relatedness between accessions
In this study, a separation of the 167 barley genotypes tested was observed according to year of registration and row type (2- and 6-rowed). The same grouping has also been reported in earlier studies on European and Nordic spring barley accessions (Backes et al. 2003; Brantestam et al. 2007). Even though the agroenvironmental conditions for cultivation of spring barley differ widely in the Northern European region, we did not observe any genetic separation according to country of origin. Likewise, no separation according to country of origin was observed in the study by Brantestam et al. (2007). One reason could be that local adaptation or breeding for regional conditions could be present, obscuring the national origin. However, the lack of grouping correlating to ‘country of origin’ within the Nordic countries Denmark, Finland, Norway and Sweden could also suggest that the cultivars are widely adapted to Nordic conditions or that they represent common germplasm independent of national breeding programs.
Conservative validation of marker-trait associations
The outcome from a GWAS is the results of the applied statistical model and dependent on the set of accessions studied. Additionally, type of molecular markers, phenotypes and phenotyping method influence the output. Several studies have debated the statistical challenges and the inputs (Jannink 2007; Stich et al. 2008; Matthies et al. 2012). In the present study, we applied a conservative approach, where markers were only accepted when found significant [−log10(P value) > 2.95] in four analyses (both AllM and MwP tested with the optimized and naive model). Taking such conservative approach to avoid false positives consequently caused a higher rate of rejected marker-trait associations. However, the conservative procedure of validation was applied to achieve marker-trait associations with known chromosome position sufficiently solid to be exploited in marker-assisted selection.
Markers for breeding of cultivars for the future climate
Since 1975, [CO2] has increased from 330 to 400 ppm of today (Collins et al. 2013). However, a correlating positive response in cereal grain yield to this change in [CO2] is found absent or weak, despite that modern cultivars have been developed along with the gradually increasing [CO2] (Manderscheid and Weigel 1997; Ziska et al. 2004; Franzaring et al. 2013; Ingvordsen et al. 2015). The markers that associated with increased grain yield and grain number under elevated [CO2] identified in LD block 4H-9 and 7H-11 are possibly of value, when aiming to improve response of grain yield to elevated [CO2]. Further, a putative sucrose synthase, a key enzyme in the sucrose metabolism forming carbohydrates (Barrero-Sicilia et al. 2011), is co-localizing with one of the markers in LD block 7H-11 (BOPA2_12_30880; Alpert et al. 2011). In that same region, a sucrose synthase was previously found, decisive for malting quality (Matthies et al. 2011). The role of sucrose synthase in the energy storage metabolism should be further explored under conditions of elevated [CO2], as this enzyme could potentially improve grain yield in a changed climate.
The identification of markers associated with either grain yield or vegetative aboveground biomass under elevated temperature (LD block 2H-2 and 2H-3) calls for further investigations of genes in that region. The presence of loci for grain yield has previously been reported by Varshney et al. (2012) and Hayes et al. (1993) in this chromosome region. Interestingly, the pseudo-response regulator gene Ppd-H1, found to provide adaptation to different environments through photoperiod responses, is positioned in this region on 2H (Turner et al. 2005; Wang et al. 2010). Within the QTL reported by Wang et al. (2010), the strongest candidate gene for Ppd-H1 was found located at chromosome 2H bin 3, where we here identified four markers associated with grain yield or grain yield relative to ambient under elevated temperature (Table 3, LD block 2H-2) pointing to a possible link between photoperiod response and grain yield. However, the bins span large genomic areas and connection remains therefore speculative. In favour of the connection between photoperiod and increased grain yield is the previously suggested photoperiod control of the growth period of the spike, where prolongation of this period leads to increased number of grains (Miralles and Slafer 2007). Further, in barley, genetic variability has been identified for this developmental phase, where stems and spikes grow actively (Whitechurch et al. 2007). The markers are found associated under elevated temperature, where number of grains decreased significantly and ears with grains also decreased. Therefore, another physiological explanation could be the ability to maintain floret fertility under elevated temperature, which has been found to abort florets (Ugarte et al. 2007; Rajala et al. 2011). In order to explain the underlying mechanism of the found association or perhaps connect photoperiod to temperature tolerance, further studies are necessary.
In LD block 2H-4, three markers associated either with number of ears with grains or with total number of ears under all applied treatments except for the two-factor treatment. The gene product co-localized with the marker positioned at 66.3 cM (BOPA1_4037-916, Online Resource 5) shows homology to a PsbP family protein, which plays a role in the oxygen evolving complex of photosystem II that is essential for normal photosynthetic activity (Roose et al. 2007; HarvEST 2014; Alpert et al. 2011). The marker at 66.9 cM (SCRI_RS_6727, Table 3), also within LD block 2H-4, that associated with total number of ears both under elevated temperature and elevated [CO2] co-localizes with a putative peroxidase 18-like protein. Several peroxidases have been found involved in abiotic and biotic stress responses. Elevated temperature as an abiotic stressor has previously been found to decrease ear production (Köszegi et al. 2005). Here, we found the marker SCRI_RS_6727 in LD block 2H-4 to be associated with total number of ears under elevated temperature and therefore potentially beneficial in marker-assisted selection for maintaining the production of ears under elevated temperature. However, further studies are necessary to verify whether the maintained ear production is also accompanied by grain yield.
Climate stability of yield and aboveground vegetative biomass was estimated by two stability measures (S 2 and W 2); however, few marker associations with stability were identified. This might suggest that climate stability is highly complex with interaction of numerous genes. An accession with a low value of S 2 was interpreted as environmentally stable, a precious trait under future climate variability. The two markers associated with either S 2 of grain yield or HI in LD block 7H-12 co-localize with the ubiquitin-conjugating enzyme 18 (Alpert et al. 2011). With ubiquitin being involved in protein degradation, the found co-localization with this ubiquitin enzyme indicates the involvement of protein turnover in stability to climate variation. However, the complexity of the processes that govern yield will make further interpretation highly speculative; this complexity might also be the reason for the scarcity of published results on QTL for stability in barley (Kraakman et al. 2004; Lacaze et al. 2009). However, not least due to the threat of climate change, this area of research is developing (reviewed by Korte and Farlow 2013) valorizing markers-trait associations for stability.
Marker-trait associations in the two-factor treatment differ from those in the constituent single-factor treatments
It could be assumed that a marker or LD block associated with a trait under either elevated temperature or [CO2] would also be associated with this trait, when both of these single factors were elevated simultaneously. However, only three markers were found to be associated with phenotypes in the two-factor treatment, while 31 marker-trait associations were identified under the corresponding single-factor treatments. This difference in number of marker-trait associations suggests the responses in the combined treatment to involve the interaction of multiple genes, and therefore, associations cannot easily be detected by GWAS. Interestingly, only in one case, LD block 2H-6, two linked markers were found for both the two-factor treatment and one of the component single-factor treatments viz. temperature, however, not with the same production parameter (ΔGY, temperature and ΔET, two-factor; Table 3). The absent overlap of marker traits from single-factor and their two-factor treatments may be attributable to the opposite effects of elevated temperature and [CO2] on plant production (Table 2). The apparent absence of similarity in genetic regulation of effects in the combined and the single-factor treatments emphasizes the need for multifactor studies, when the need is to identify marker-phenotype association for multifaceted future climate conditions.
Preferably, association studies should be performed under field conditions with elevated climatic factors. However, multifactor field studies including temperature are challenging in set-up. Despite several attempts and developments of technologies, it is difficult to increase temperature more than 1–2 °C under field conditions without several redundant experimental effects (Kimball et al. 2007; Bruhn et al. 2013). For GWAS, an appropriate—and large—quantity of accessions must be included and that defines the size of the facility needed to manipulate the climate and further excludes several enclosure facilities due to size.
In conclusion, our results have revealed potential genome sites to be exploited in the breeding of climate resilient cultivars. We have identified SNP candidates for grain yield and other production parameters under conditions of elevated temperature, [CO2] and [O3] for further use in marker-assisted selection.
Abbreviations
- AllM:
-
Dataset of all markers
- BM:
-
Aboveground vegetative biomass
- EG:
-
Number of ears with grains
- ET:
-
Number of ears
- GN:
-
Number of grains
- GWAS:
-
Genome-wide association study
- GY:
-
Grain yield
- HI:
-
Harvest index
- LD:
-
Linkage disequilibrium
- MwP:
-
Markers from the AllM dataset with identified position
- QTL:
-
Quantitative trait loci
- SNP:
-
Single nucleotide polymorphism
References
Alpert M, Wanamaker S, Duma D et al (2011) A genome sequence resource for barley. Barley Genet Newsl 41:10–11
Backes G, Hatz B, Jahoor A, Fischbeck G (2003) RFLP diversity within and between major groups of barley in Europe. Plant Breed 122:291–299. doi:10.1046/j.1439-0523.2003.00810.x
Barrero-Sicilia C, Hernando-Amado S, González-Melendi P, Carbonero P (2011) Structure, expression profile and subcellular localisation of four different sucrose synthase genes from barley. Planta 234:391–403. doi:10.1007/s00425-011-1408-x
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 57:289–300
Bokszczanin KL, Fragkostefanakis S (2013) Perspectives on deciphering mechanisms underlying plant heat stress response and thermotolerance. Front Plant Sci 4:315. doi:10.3389/fpls.2013.00315
Bradbury PJ, Zhang Z, Kroon DE et al (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. doi:10.1093/bioinformatics/btm308
Brantestam AK, Bothmer R, Dayteg C et al (2007) Genetic diversity changes and relationships in spring barley (Hordeum vulgare L.) germplasm of Nordic and Baltic areas as shown by SSR markers. Genet Resour Crop Evol 54:749–758. doi:10.1007/s10722-006-9159-4
Bruhn D, Larsen KS, de Dato GD et al (2013) Improving the performance of infrared reflective night curtains for warming field plots. Agric For Meteorol 173:53–62. doi:10.1016/j.agrformet.2013.01.004
Cai S, Wu D, Jabeen Z et al (2013) Genome-wide association analysis of aluminum tolerance in cultivated and Tibetan wild barley. PLoS ONE 8:e69776. doi:10.1371/journal.pone.0069776
Caldwell KS, Russell J, Langridge P, Powell W (2006) Extreme population-dependent linkage disequilibrium detected in an inbreeding plant species, Hordeum vulgare. Genetics 172:557–567. doi:10.1534/genetics.104.038489
Close TJ, Bhat PR, Lonardi S et al (2009) Development and implementation of high-throughput SNP genotyping in barley. BMC Genom 10:582. doi:10.1186/1471-2164-10-582
Collins M, Knutti R, Arblaster J, Dufresne J-L et al (2013) Long-term climate change: projections, commitments and irreversibility. In: Stocker TF, Qin D, Plattner G-K, Tignor M, Allen SK, Boschung J, Nauels A, Xia Y, Bex V, Midgley PM (eds.) Climate change 2013: the physical science basis. Contribution of working group I to the fifth assessment report of the Intergovernmental Panel on Climate Change. Cambridge University Press, Cambridge
Comadran J, Ramsay L, MacKenzie K et al (2011) Patterns of polymorphism and linkage disequilibrium in cultivated barley. Theor Appl Genet 122:523–531. doi:10.1007/s00122-010-1466-7
Comadran J, Kilian B, Russell J et al (2012) Natural variation in a homolog of Antirrhinum CENTRORADIALIS contributed to spring growth habit and environmental adaptation in cultivated barley. Nat Genet 44:1388–1392. doi:10.1038/ng.2447
Ellermann T, Nøjgaard JK, Nordstrøm C, et al (2013) The Danish air quality monitoring programme. Annual summary for 2012. 59 pp
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. doi:10.1111/j.1365-294X.2005.02553.x
FAOSTAT (2014) Food supply, crops primary equivalent. http://faostat3.fao.org. Accessed Feb 2014
Franzaring J, Holz I, Fangmeier A (2013) Responses of old and modern cereals to CO2-fertilisation. 64:943–956. doi:10.1071/CP13311
Fuhrer J (2003) Agroecosystem responses to combinations of elevated CO2, ozone, and global climate change. Agric Ecosyst Environ 97:1–20. doi:10.1016/S0167-8809(03)00125-7
Hayes PM, Liu BH, Knapp SJ et al (1993) Quantitative trait locus effects and environmental interaction in a sample of North American barley germ plasm. Theor Appl Genet 87:392–401. doi:10.1007/BF01184929
Ingvordsen IPCC (2014) Summary for policymakers; climate change 2014: impacts, adaptation, and vulnerability. Contribution of working group II to the fifth assessment report of the Intergovernmental Panel on Climate Change 2014, pp 1–44
Ingvordsen CH, Backes G, Lyngkjær MF et al (2015) Significant decrease in yield under future climate conditions: stability and production of 138 spring barley accessions. Eur J Agron 63:105–113. doi:10.1016/j.eja.2014.12.003
Jannink J-L (2007) Identifying quantitative trait locus by genetic background interactions in association studies. Genetics 176:553–561. doi:10.1534/genetics.106.062992
Kang HM, Zaitlen NA, Wade CM et al (2008) Efficient control of population structure in model organism association mapping. Genetics 178:1709–1723. doi:10.1534/genetics.107.080101
Kimball B, Conley MM, Wang S et al (2007) Infrared heater arrays for warming ecosystem field plots. Glob Change Biol 14:309–320. doi:10.1111/j.1365-2486.2007.01486.x
Korte A, Farlow A (2013) The advantages and limitations of trait analysis with GWAS: a review. Plant Methods 9:29. doi:10.1186/1746-4811-9-29
Köszegi B, Karsai I, Mészáros K, Kovács G (2005) Study of the early growth and tillering capacity of barley (Hordeum vulgare L.) under changing temperature and light conditions. Acta Agron Hung 53:371–376
Kraakman ATW, Niks RE, Van den Berg PM et al (2004) Linkage disequilibrium mapping of yield and yield stability in modern spring barley cultivars. Genetics 168:435–446. doi:10.1534/genetics.104.026831
Lacaze X, Hayes PM, Korol A (2009) Genetics of phenotypic plasticity: QTL analysis in barley, Hordeum vulgare. Heredity 102:163–173. doi:10.1038/hdy.2008.76
Lipka AE, Tian F, Wang Q et al (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28:2397–2399. doi:10.1093/bioinformatics/bts444
Lobell DB, Schlenker W, Costa-Roberts J (2011) Climate trends and global crop production since 1980. Science 333:616–620. doi:10.1126/science.1204531
Long NV, Dolstra O, Malosetti M et al (2013) Association mapping of salt tolerance in barley (Hordeum vulgare L.). Theor Appl Genet Theor Angew Genet 126:2335–2351. doi:10.1007/s00122-013-2139-0
Manderscheid R, Weigel HJ (1997) Photosynthetic and growth responses of old and modern spring wheat cultivars to atmospheric CO2 enrichment. Agric Ecosyst Environ 64:65–73. doi:10.1016/S0167-8809(97)00020-0
Matthies IE, Sharma S, Weise S, Röder MS (2011) Sequence variation in the barley genes encoding sucrose synthase I and sucrose phosphate synthase II, and its association with variation in grain traits and malting quality. Euphytica 184:73–83. doi:10.1007/s10681-011-0563-2
Matthies IE, Hintum T, Weise S, Röder MS (2012) Population structure revealed by different marker types (SSR or DArT) has an impact on the results of genome-wide association mapping in European barley cultivars. Mol Breed 30:951–966. doi:10.1007/s11032-011-9678-3
Miralles DJ, Slafer GA (2007) Sink limitations to yield in wheat: How could it be reduced? J Agric Sci 145:139–149. doi:10.1017/S0021859607006752
Morison JIL, Lawlor DW (1999) Interactions between increasing CO2 concentration and temperature on plant growth. Plant Cell Environ 22:659–682
Nevo E, Fu Y-B, Pavlicek T et al (2012) Evolution of wild cereals during 28 years of global warming in Israel. Proc Natl Acad Sci USA 109:3412–3415. doi:10.1073/pnas.1121411109
Newton AC, Flavell AJ, George TS et al (2011) Crops that feed the world 4. Barley: a resilient crop? Strengths and weaknesses in the context of food security. Food Secur 3:141–178. doi:10.1007/s12571-011-0126-3
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rajala A, Hakala K, Mäkelä P, Peltonen-Sainio P (2011) Drought effect on grain number and grain weight at spike and spikelet level in six-row spring barley. J Agron Crop Sci 197:103–112. doi:10.1111/j.1439-037X.2010.00449.x
Roemer T (1917) Sind die ertragsreichen Sorten ertragssicherer? Mitteilung Deutsche Landwirtschafts-Gesellschaft 32:87–89
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5:69–76. doi:10.1007/BF00020088
Roose JL, Wegener KM, Pakrasi HB (2007) The extrinsic proteins of photosystem II. Photosynth Res 92:369–387. doi:10.1007/s11120-006-9117-1
Rosenzweig C, Parry ML (1994) Potential impact of climate change on world food supply. Nature 367:133–138
Schweizer P, Stein N (2011) Large-scale data integration reveals colocalization of gene functional groups with meta-QTL for multiple disease resistance in barley. Mol Plant Microbe Interact 24:1492–1501. doi:10.1094/MPMI-05-11-0107
Stich B, Möhring J, Piepho H-P et al (2008) Comparison of mixed-model approaches for association mapping. Genetics 178:1745–1754. doi:10.1534/genetics.107.079707
Svenning J-C, Sandel B (2013) Disequilibrium vegetation dynamics under future climate change. Am J Bot 100:1266–1286. doi:10.3732/ajb.1200469
Tabangin ME, Woo JG, Martin LJ (2009) The effect of minor allele frequency on the likelihood of obtaining false positives. BMC Proc 3(Suppl 7):S41. doi:10.1186/1753-6561-3-S7-S41
R Core Team (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
The International Barley Genome Sequencing Consortium (2012) A physical, genetic and functional sequence assembly of the barley genome. Nature 491:711–718. doi:10.1038/nature11543
Tondelli A, Xu X, Moragues M et al (2013) Structural and temporal variation in genetic diversity of european spring two-row barley cultivars and association mapping of quantitative traits. Plant Genome 6:1–14. doi:10.3835/plantgenome2013.03.0007
Turner A, Beales J, Faure S et al (2005) The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310:1031–1034. doi:10.1126/science.1117619
Ugarte C, Calderini DF, Slafer GA (2007) Grain weight and grain number responsiveness to pre-anthesis temperature in wheat, barley and triticale. Fields Crop Res 100:240–248. doi:10.1016/j.fcr.2006.07.010
UN (2012) World population prospects. United Nations, Department of Economic and Social Affairs Population Division. http://esa.un.org/wpp. Accessed 17 Jan 2014
Varshney RK, Paulo MJ, Grando S et al (2012) Genome wide association analyses for drought tolerance related traits in barley (Hordeum vulgare L.). Fields Crop Res 126:171–180. doi:10.1016/j.fcr.2011.10.008
Wang G, Schmalenbach I, von Korff M et al (2010) Association of barley photoperiod and vernalization genes with QTLs for flowering time and agronomic traits in a BC2DH population and a set of wild barley introgression lines. Theor Appl Genet 120:1559–1574. doi:10.1007/s00122-010-1276-y
Waugh R, Jannink J-L, Muehlbauer GJ, Ramsay L (2009) The emergence of whole genome association scans in barley. Curr Opin Plant Biol 12:218–222. doi:10.1016/j.pbi.2008.12.007
Wetterstrand K (2014) DNA sequencing costs: The NHGRI Genome Sequencing Program (GSP). http://www.genome.gov/sequencingcosts. Accessed 23 Jan 2014
Whitechurch EM, Slafer GA, Miralles DJ (2007) Variability in the duration of stem elongation in wheat and barley genotypes. J Agron Crop Sci 193:138–145. doi:10.1111/j.1439-037X.2007.00260.x
Wricke G (1962) Über eine Methode zur Erfassung der ökologischen Streubreite in Feldversuchen. Z Pflanzenzuchtg 47:92–96
Zadoks JC, Chang TT, Konzak CF (1974) A decimal code for the growth stages of cereals. Weed Res 14:415–421
Zhang Z, Ersoz E, Lai C-Q et al (2010) Mixed linear model approach adapted for genome-wide association studies. Nat Genet 42:355–360. doi:10.1038/ng.546
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome J 1:5. doi:10.3835/plantgenome2008.02.0089
Ziska LH, Morris CF, Goins EW (2004) Quantitative and qualitative evaluation of selected wheat varieties released since 1903 to increasing atmospheric carbon dioxide: Can yield sensitivity to carbon dioxide be a factor in wheat performance? Glob Change Biol 10:1810–1819. doi:10.1111/j.1365-2486.2004.00840.x
Acknowledgments
We thank all involved in the cultivation and harvest procedures and Allan Murphy and Esben Højrup for technical assistance in RERAF during the experiment. The accessions were provided by NordGen (the Nordic Genetic Resource Center; http://www.nordgen.org/), breeders of the Nordic network ‘Sustainable primary production in a changing climate’ (NordForsk) and a few came from the BAR-OF project (ICROFS, Denmark). The CO2 used in this study was generously supplied by Air Liquide Danmark A/S. The Nordic Council of Ministers supported the major part of this research through the network, ‘Sustainable primary production in a changing climate’. Other funders were The Danish Council for Independent Research: Technology and Production Sciences (FTP) via the project ‘Climate Change Effects on Plant Health’ (project 0602-00589B), and COBRA (Core Organic II).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ingvordsen, C.H., Backes, G., Lyngkjær, M.F. et al. Genome-wide association study of production and stability traits in barley cultivated under future climate scenarios. Mol Breeding 35, 84 (2015). https://doi.org/10.1007/s11032-015-0283-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-015-0283-8