Abstract
Many populations of elasmobranchs (sharks and rays) are experiencing severe declines due to the high demand for shark fins in Asia, the activities of unregulated fisheries, and increases in shark and ray catches. Recently, the effects of the decline in the populations of marine fish species on genetic diversity have drawn increasing attention; however, only a few studies have addressed the genetic diversity of shark and ray populations. Here, we report the results of a quantitative analysis of the genetic diversity of shark and ray species over the past 20 years and discuss the importance and utility of this genetic information for fisheries management and conservation policies. Furthermore, we suggest future actions important for minimizing the gaps in our current knowledge of the genetic diversity of shark and ray species and to minimize the information gap between genetic scientists and policymakers. We suggest that shark and ray fisheries management and conservation policies consider genetic diversity information, such as the management unit, effective population size (Ne), haplotype and nucleotide diversity, observed heterozygosity, and allelic richness, because the long-term survival of a species is strongly dependent on the levels of genetic diversity within and between populations. In addition, sharks and rays are a group of particular interest for genetic conservation due to their remarkable life histories.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic data have aided conservation research and management by facilitating the detection of genetically distinct populations, the measurement of genetic connectivity and the identification of the risks associated with demographic change and inbreeding (Allendorf et al. 2013). A good example for which genetic information has been considered in fisheries management is the Pacific salmon (Oncorhynchus spp.), for which genetic data have influenced conservation efforts associated with population restoration (Waples 1995). However, the effective application of genetic data to the management plans for several marine species, including sharks and rays, remains a challenge (Kenchington et al. 2003).
The effects of population-level declines are of major concern in conservation biology because small populations suffer from inbreeding and genetic drift. These effects lead to loss of genetic diversity, which has several potential consequences, such as compromising the ability of a population to evolve in order to cope with environmental changes and reducing its chances of long-term persistence (Frankham et al. 2002). Therefore, councils of evolutionary biologists and fisheries scientists are interested in elucidating the genetic patterns and demographic connectivity of different groups of individuals or populations as well as the distributions of genetic variation within and between populations (Waples and Gaggiotti 2006; Grant and Cheng 2012; Ovenden 2013).
Sharks and rays (including stingrays and skates) are groups of interest to conservationists due to their ecological importance in the marine environment and their current high levels of overexploitation (Dulvy et al. 2014). Currently, there are more than 1160 validly named species of elasmobranchs in the world (Weigmann 2016), representing a significant number of apex and mesopredators that occupy top positions in the food chain (Heithaus et al. 2008; Ferretti et al. 2010). However, despite their ecological importance, elasmobranchs are one of the most imperiled groups of marine species worldwide (Cortés 2002; Bräutigam et al. 2015) due to their life history characteristics, including late sexual maturity, lengthy pregnancy, low fertility, slow growth and long life span, making them particularly susceptible to anthropogenic pressures such as overfishing, environmental changes, and pollution (Seitz and Poulakis 2006; Dulvy et al. 2014). Indeed, these anthropogenic pressures can cause changes in genetic diversity through population reduction, thus compromising these species’ ability to evolve (DiBattista 2008).
Currently, massive population-level declines and extinction risks due to overfishing over recent decades present significant threats to sharks and rays in all oceans (Ferretti et al. 2010; Worm et al. 2013; Dulvy et al. 2014). The main issues that jeopardize shark and ray species include the high demand for shark fins and gill plates in Asia, unregulated fisheries, bycatching, and increased shark fishing due to the collapse of other fisheries (Musick et al. 2000; Clarke et al. 2006; Herndon et al. 2010; Dulvy et al. 2014; McClenachan et al. 2016). According to a study by Worm et al. (2013), the global catch of sharks from reported and unreported landings, discards, and shark finning was estimated as approximately 100 million tons in 2010. Such fishing pressures are more challenging to elasmobranchs because of their high susceptibility relative to most teleosts and because sharks and rays require several decades to recover from overfishing (Stevens et al. 2000).
In general, fisheries management of shark and ray relies on a series of studies on the basic biology, life history, and population ecology of elasmobranchs (Simpfendorfer et al. 2011). However, the population genetic diversity of sharks and rays is generally neglected in fisheries management, and the possibility of change appears distant, as many international conservation efforts currently fail to acknowledge genetic variation (Laikre 2010; Ovenden et al. 2013). Therefore, the expansion of global population genetics studies describing the genetic diversity of shark and ray species worldwide is urgently needed in order to identify genetically distinct populations and to preserve genetic diversity. It is imperative to address the severe factors that jeopardize shark and ray populations.
Against this background, we conduct a critical review and discuss the importance of including genetic diversity data in shark and ray fisheries management plans and, consequently, in conservation policies. Specifically, we discuss the importance of sharks and rays within a conservation genetics context, presenting the possible effects of fishing on their genetic diversity, and we address the current limitations and the need for an increase in genetic studies of this taxonomic group in order to assess genetic diversity across geographical ranges. In addition, we suggest future actions important to minimize the knowledge gap between shark and ray geneticists and the authors of conservation policies.
What makes sharks and rays particularly interesting to conservation genetics?
In addition to their ecological importance, elasmobranchs are a group of particular interest to conservation geneticists—researchers who use genetic/genomic techniques to solve problems in conservation biology—due to the remarkable features of their life histories. These features include (i) the evolutionary uniqueness of elasmobranchs, (ii) their reproductive strategy, (iii) the effects of overfishing on evolution, (iv) their broad geographic distribution, and (v) the limited number of studies describing their genetic diversity.
Evolutionary uniqueness
Sharks and rays compose a major lineage of evolutionarily unique vertebrates consisting of approximately 1160 living species; these species represent a small fraction (< 3.0%) of modern fish fauna (Nelson et al. 2016; Weigmann 2016). Compared with marine teleosts, sharks and rays present a low species richness (1160 shark and ray species versus 30,000 teleost species). In particular, some shark and ray orders contain only one family and few genera and species, such as Echinorhiniformes (1 genus, 2 species), Pristiophoriformes (2 genera, 7 species), and Heterodontiformes (9 species), and there are even several monotypic families, such as the shark families Mitsukurinidae, Cetorhinidae, Pseudocarchariidae, and Leptochariidae and the ray families Hypnidae, Hexatrygonidae, and Plesiobatidae (Ebert et al. 2013; Last et al. 2016). Furthermore, intrinsic factors, such as diversity of form and function as a means of successful evolutionary resilience, contribute to a lower historic extinction rate and a higher evolutionary adaptability for shark and ray species, allowing them to inhabit several marine and freshwater ecosystems (Ferretti et al. 2010; Ebert et al. 2013; Richards et al. 2013). In addition, over the past 455 million years, sharks have been able to survive mass extinctions that have left ocean waters with far fewer fish (Grogan et al. 2012). Such resilience suggests that sharks have unique genetic properties that support their adaptability and evolutionary success; therefore, their genetic properties must be preserved.
Reproductive strategy
Shark and ray species exhibit a wide diversity of reproductive strategies, including multiple paternity, parthenogenesis, sperm storage, and philopatry, and these strategies can have considerable effects on genetic diversity (Chapman et al. 2004; Daly-Engel et al. 2010; Conrath and Musick 2012; Bernal et al. 2015). For example, multiple paternity has been documented in many shark and ray species (e.g., Chevolot et al. 2007; Daly-Engel et al. 2010; Byrne and Avise 2012), and whether multiple paternity assists in maintaining genetic diversity is a subject of debate (Zeh and Zeh 2003; Karl 2008). Theoretical studies argue that under natural conditions, an increase in multiple paternity will reduce effective population size (Ne) and consequently the genetic diversity (Ramakrishnan et al. 2004). On the other hand, multiple matings and sperm storage events could increase the Ne after a bottleneck (Karl 2008). In addition, Byrne and Avise (2012) posited the “sperm storage” theory, in which females mating with multiple males promotes competition among the sperm, which might lead either to improved fertilization success or to better genes for their zygotes.
Parthenogenesis, or “virgin birth” (the production of offspring without fertilization by a male), has been documented in sharks and rays (e.g., Chapman et al. 2007; Portnoy et al. 2014a, b; Fields et al. 2015). Although it is difficult to estimate the possible effects on wild populations, this reproductive strategy can be advantageous because of its adaptive significance (Booth and Schuett 2011). In particular, at low population densities, when females undergo fertilization failure because of the difficulty in finding males, facultative parthenogenesis could have adaptive significance (Fields et al. 2015). On the other hand, due to elevated homozygosity, parthenogenesis is believed to increase inbreeding, reduce fitness, increase the likelihood of the fixation of deleterious alleles, and consequently increase the probability of extinction (Watts et al. 2006; Chapman et al. 2007; Booth and Schuett 2011). The first report of facultative parthenogenesis was just recently documented in wild populations of smalltooth sawfish (Pristis pectinata, Pristidae), with five individuals reportedly close to or in complete homozygosity (Fields et al. 2015).
Another important reproductive strategy that can affect genetic diversity is natal philopatry, which is defined by the return of a far-ranging individual to its exact birthplace (Chapman et al. 2015). For instance, sex-biased dispersion, such as male-biased dispersal and female philopatry to a coastal nursery has been documented for the great white shark (Carcharodon carcharias, Lamnidae; Pardini et al. 2001) and bonnethead (Sphyrna tiburo, Sphyrnidae; Portnoy et al. 2015). According to Portnoy et al. (2015), sex-biased dispersion can facilitate sorting of locally adaptive variation, with the dispersion of one sex facilitating the movement of potentially adaptive variation among locations and environments.
Effects of overfishing on the evolutionary process
Overfishing may impact evolutionary processes mainly by changing body size and by promoting early sexual maturity; additionally, overfishing affects bioeconomics and macroecological patterns (Belgrano and Fowler 2013; Heino et al. 2015). For sharks, only a few studies based exclusively on phenotypic traits have shown direct evidence of the influence of fisheries on evolution. Walker et al. (1998) reported changes in the growth rate of gummy sharks (Mustelus antarcticus, Triakidae) caused by length-selective fishing mortality. Furthermore, Clarke et al. (2013) reported that the median lengths of silky sharks (Carcharhinus falciformis, Carcharhinidae) and oceanic whitetip sharks (Carcharhinus longimanus, Carcharhinidae) decreased significantly in the Pacific Ocean between 1995 and 2010. Though these phenotypic changes may indicate an evolutionary response to overfishing, the possible genetic consequences are unknown. Recently, Gallagher et al. (2014) suggested that the ecological, behavioral, and physiological adaptations of hammerhead sharks (Sphyrnidae) that once promoted evolutionary success are now maladaptive under current levels and modes of exploitation. For example, the high agility that supports their prey capture strategy of burst swimming behavior also results in a high rate (60–80%) of at-vessel and post-release mortality (Gallagher et al. 2014). However, though no studies of direct fisheries-induced evolution of shark and ray species exist, the examples cited above suggest that evolutionary traits and unique adaptations can be affected by overexploitation.
Broad geographic distribution
Many sharks and rays are widely distributed and highly mobile (e.g., the shortfin mako Isurus oxyrinchus, Lamnidae, and the pelagic stingray Pteroplatytrygon violacea, Dasyatidae), features that make it difficult to sample enough individuals from different locations to allow for the identification of discrete populations over the entire distribution of the species. For example, at least 150 shark species regularly migrate across national boundaries, and ¼ of threatened shark species have ranges that include at least 18 countries (Dulvy et al. 2014).
Unlike bony fishes and other marine organisms, shark and ray species do not have a planktonic larval stage with dispersal via ocean currents. Instead, their dispersal is mediated entirely by the active movement of adult individuals. In general, large migratory and oceanic species such as the blue shark (Prionace glauca, Carcharhinidae) tend to present more homogeneous populations (Taguchi et al. 2015), whereas smaller, more coastal species, such as the spot-tail shark (Carcharhinus sorrah, Carcharhinidae), commonly exist in isolated populations (Giles et al. 2014). These coastal sharks and rays tend to experience more obstacles, such as marine barriers and oceanographic heterogeneity. Therefore, assessing the extant genetic diversity throughout their distribution is imperative to avoid local gene pool erosion. For example, the blue shark is likely the most wide-ranging species of shark and the most heavily fished shark species in the world, but only a few range-limited studies (e.g., Ovenden et al. 2009; King et al. 2015; Taguchi et al. 2015; Li et al. 2016) have attempted to describe the genetic structure and diversity of this species. These studies did not indicate any genetic differentiation in the Pacific and Indo-Pacific regions, a result that was attributed mainly to the blue shark’s high agility. Nevertheless, the authors indicated the need for cooperative fisheries management among different countries, even though management programs do not have any genetic data available along the broad distribution of the blue shark. Consequently, there is a considerable gap to obtaining a holistic view of the population structure of this species. Moreover, it will not be possible to detect negative changes and reductions in genetic diversity unless the major distribution points of sharks and rays are studied.
Although there are currently no global genetic population studies of the blue shark, a few widely distributed shark species, such as the scalloped hammerhead (Sphyrna lewini, Sphyrnidae), the whale shark (Rhincodon typus, Rhincodontidae), the sand tiger shark (Carcharias taurus, Odontaspididae), the sandbar shark (Carcharhinus plumbeus, Carcharhinidae), the dusky shark (Carcharhinus obscurus, Carcharhinidae), the copper shark (Carcharhinus brachyurus, Carcharhinidae) and the silky shark (Carcharhinus falciformis, Carcharhinidae), have been studied globally. Although these studies are incipient and some of them report only limited genetic markers, they demonstrate that different populations of single shark species are genetically discrete entities worldwide that may have different levels of genetic diversity (Duncan et al. 2006; Castro et al. 2007; Ahonen et al. 2009; Portnoy et al. 2010; Benavides et al. 2011b; Clarke et al. 2015). Therefore, each discrete shark population should be managed separately to reduce the risk of depleting their genetic resources.
Few studies describe the genetic diversity of sharks and rays
Despite an increase in the number of genetics studies in the last decade (Fig. 1), currently, only ~ 10% of shark and ray species have been investigated in terms of their population genetic structure, genetic diversity and demographic history. For example, no population genetics study performed to date has aimed at describing the genetic diversity and identifying the discrete populations along the distribution range of the pelagic stingray, a cosmopolitan species frequently caught as bycatch in pelagic longline fisheries around the world (Forselledo et al. 2008). The same is true for species with narrow geographic distributions and species that are critically endangered, such as the daggernose shark (Isogomphodon oxyrhynchus, Carcharhinidae) (Lessa et al. 2016). Furthermore, the majority of studies represent the first genetic examination of a particular species (Dudgeon et al. 2012), although there are a few exceptions, such as studies in which some species, including the white shark and the scalloped hammerhead, are re-examined. The absence of genetic evaluations of many shark species complicates the transition from the current overexploitation and short- and long-term conservation.
The effects of fishing on the genetic diversity of shark and ray populations
Despite the negative relationship between overexploited populations and genetic diversity, the use of genetics in the management plans of threatened species remains a challenge (Laikre 2010). The effects of population declines on marine fishes in terms of decreased genetic diversity have recently drawn attention. Many authors claim the need for an application of genetic diversity metrics in fisheries management plans because of the harmful consequences of inbreeding, the loss of genetic diversity, the loss of evolutionary potential and changes in population structures of species (Kenchington et al. 2003; Allendorf et al. 2008; Frankham 2010; Laikre et al. 2010b; Hoban et al. 2013a, b).
Studies have used historical and contemporary samples to address the question of how fisheries affect the genetic diversity (mtDNA and microsatellites) of fish and marine mammal stocks (Hauser et al. 2002; Pichler and Baker 2000). For example, the heterozygosity (microsatellites), number of alleles per locus, and Ne of the New Zealand snapper (Pagrus auratus, Sparidae) declined between 1950 and 1988 after the creation of a fishery for this population (Hauser et al. 2002). McCusker and Bentzen (2010) found a positive relationship between genetic diversity (mtDNA and microsatellites) and fish stock abundance. Meanwhile, Pinsky and Palumbi (2014) compiled data for 140 species of marine fishes across 11,049 loci and clearly showed a reduction in allelic richness in 9 overfished stocks among 12 genera and families. According to Allendorf et al. (2008), uncontrolled harvesting may lead to genetic impacts, such as the alteration of population subdivision, loss of genetic variation, and selective genetic changes. To date, no study has assessed the direct relationship between abundance and genetic diversity metrics (i.e., nucleotide and haplotype diversity, observed heterozygosity and allelic richness) in shark and ray species. Nevertheless, many species that are under intense fishing pressure have shown low values of genetic diversity, as indicated mainly by nucleotide diversity and observed heterozygosity (Table 1; Fig. 3). Although the low genetic diversity of shark and ray species is probably more associated with bottlenecks and the slow rate of molecular evolution, regardless of whether the cause is historical or cotemporary, the current levels of genetic diversity should be taken into account for conservation policies (Martin et al. 1992; Hoelzel et al. 2006; O’Brien et al. 2013; Allendorf et al. 2013).
Traditional management plans seek to increase the number of individuals in different populations (Hauser and Carvalho 2008). However, even large populations (census population size—Nc) may face a substantial loss of genetic variation because the Ne, which determines the strength of genetic drift in a population, is often much smaller than Nc in overexploited marine fish species (Ryman et al. 1995; Allendorf et al. 2008; Hare et al. 2011). For example, millions of individuals may be equivalent to a Ne of only hundreds or thousands (Ryman et al. 1995; Hauser et al. 2002). Therefore, as Ne decreases, genetic drift erodes genetic variation, increasing the probability of fixation of deleterious alleles and reducing the resilience of overfished species (Hare et al. 2011). Although Ne is one of the most important genetic parameters of wildlife populations, estimations of contemporary Ne are highly limited for shark and ray species (Table 2). Several studies have shown Ne values lower than 500 for some shark and ray species, such as the zebra shark (Stegostoma fasciatum, Stegostomidae) (Dudgeon and Ovenden 2015) and the smalltooth sawfish (Chapman et al. 2011) (Table 2). Recent studies suggest that at a minimum, an Ne of ≥ 100 individuals is suggested to prevent short-term genetic erosion and a 10% loss of fitness over five generations, whereas the minimal threshold to retain long-term evolutionary potential is at least 1000 individuals (Frankham 2014). The Ne values estimated for elasmobranch populations suggest the need for long-term monitoring and can be informative for management decisions. Therefore, we believe that Ne is an important genetic parameter to be applied to future fisheries management plans because of its importance in assessing the level of genetic diversity (Willoughby et al. 2015) and because of its role as a proxy of abundance [(IUCN Red List criterion C) (Ovenden et al. 2016)], as determined by previous studies (e.g., Dudgeon et al. 2012; Frankham 2014; Ovenden et al. 2016).
Conservation policies have overlooked and neglected shark and ray genetic diversity
Many shark and ray species are highly migratory and overexploited; therefore, they may require international management efforts such as bilateral and multilateral fisheries management agreements (Musick et al. 2000; Herndon et al. 2010). The conservation status and management measures for many shark and ray species have been evaluated by international conservation organizations, and the policies for conservation are mainly based on retention bans, finning bans and trading bans intended to promote the recovery of shark populations (Tolotti et al. 2015). Like many other international policies these organizations do not have any initiatives that deal specifically with genetic diversity.
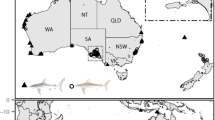
Recently, a consortium representing various organizations of experts, The Global Sharks and Rays Initiative (GSRI), launched the Global Strategy for the Conservation of Sharks and Rays (2015–2025). This document summarizes the global priorities for shark and ray conservation (Bräutigam et al. 2015) and highlights the urgent need to prevent the extinction of imperiled coastal sharks and rays in many diverse and endangered hotspots, including the coastal waters near Argentina, Australia, Brazil, Colombia, Indonesia, Japan, Madagascar, Mozambique, South African and Uruguay (Lucifora et al. 2011; Bräutigam et al. 2015). Similarly to the other conservation initiatives for shark and ray species, the GSRI does not recognize the importance of genetic diversity as a criterion for its management plan. Population genetics metrics, including genetic diversity levels, population structure and demographic history, could help the GSRI assess the genetic health of populations and determine priority areas for conservation. This approach could provide an excellent match between the use of common genetics tools in management and their relevant applications in the policy arena (Hoban et al. 2013a, b).
The same problem extends to the one of the most influential conservation organizations in the world, the International Union for Conservation of Nature (IUCN). Currently, the framework developed by the IUCN for assessing extinction risk of species is the most widely used even though the IUCN Red List does not hold any legal weight (Fung and Waples 2017). Although the IUCN considers genetic diversity as one of the three levels of biodiversity that must be conserved (McNelly et al. 1990), there are no specific genetic criteria listed that would categorize sharks and rays or other species as being under any level of threat (Laikre 2010; Rivers et al. 2014). Although the IUCN Red Lists take into account a range of quantitative species-specific criteria, such as distribution, number of individuals and declines in abundance, other than for rare exceptions, the categorization of a given species into Red List categories (IUCN 2001) largely ignores genetic diversity in its evaluation criteria. This shortcoming suggests that any shark and ray species listed under an IUCN Red List category could be overlooked in long-term management plans (Laikre et al. 2008; Willoughby et al. 2015).
According to Dulvy et al. (2014), 1041 shark and ray species are currently listed under the IUCN Red List threat categories. Of these, 181 shark and ray species fall into categories that represent varying degrees of threat (Dulvy et al. 2014). A Web of Science® search that we conducted, selecting data up to August, 2016, indicated that the number of shark and ray species for which genetic information is available has increased over the last 20 years, though these studies have mainly focused on sharks rather than rays (Fig. 1). However, some genetic data (genetic diversity metrics and population genetic structures) exist for only 10% of the 1041 shark and ray species currently listed by the IUCN, and approximately 25% of the species fall into some threat category (Fig. 2). Even with nearly half of all shark species listed by the IUCN as ‘Data Deficient’ (DD) due to incomplete data in terms of life history and population abundance dynamics (Hoffman et al. 2010), for some species in this category, there are genetic data that could be used for assessment when combined with other information. Moreover, the lowest values of observed heterozygosity are for species listed as DD and Least Concern (LC), whereas the lowest nucleotide diversity values are for species listed as LC (Fig. 3). This pattern is similar to those found by Willoughby et al. (2015) for bony fish, birds, mammals, and reptiles. For example, artisanal fisheries represent a main threat to nurse sharks (Ginglymostoma cirratum, Ginglymostomatidae), which inhabit coastal waters and are found throughout the Atlantic Ocean (Ebert et al. 2013). Although the IUCN lists nurse sharks as DD globally (Rosa et al. 2006), this species has been genetically analyzed, and these data could be used to help assess their populations. The nurse shark population in the western Atlantic Ocean shows low genetic diversity in the control region of the mtDNA (CR mtDNA) (h = 48 ± 5%; π = 0.08 ± 0.06%) and microsatellites (Ho = 0.58). Furthermore, nurse shark populations show significant and distinct genetic differences between offshore islands and the mainland in the western Atlantic Ocean, and there is a high degree of genetic variability (78.2%) within populations (Karl et al. 2012). These genetic isolation patterns and genetic diversity parameters are comparable to those of other shark species categorized by the IUCN as threatened, including the sand tiger shark and the narrownose shark (Mustelus schmitti, Triakidae) (Table 1). Although genetically depauperate shark populations result mainly from historical fluctuations in population size (O’Brien et al. 2013), these examples clearly highlight the usefulness of genetic diversity metrics, including haplotype and nucleotide diversity and heterozygosity, at least as indicators of the health of the population and the conservation status of a particular shark species. Specifically, high levels of genetic diversity can increase individual fitness and population resilience, and there is currently no framework for the direct use of genetic diversity metrics in management plans.
Boxplot of genetic diversity metrics of shark and ray species pooled for each IUCN category. a Haplotype diversity, b Nucleotide diversity, c Observed heterozygosity and d Expected heterozygosity. DD data deficient, LC least concern, NT near threatened, VU vulnerable, EN endangered, CR critically endangered
As advised in previous studies (Frankham 2010, 2014; Laikre 2010; Rivers et al. 2014; Willoughby et al. 2015), genetic diversity metrics imply that the IUCN should include genetic diversity as another criterion for categorizing threatened species. This could be completed using a novel approach for identifying vertebrate species with conservation needs based on the number of generations (t) until, using Ne as an index, the species loses significant genetic diversity (Willoughby et al. 2015). Obviously, we recognize that estimating Ne for shark and ray species can be difficult, mainly because these species tend to have overlapping generations; thus, estimated Ne must not be used as an autonomous criterion. However, this parameter is very informative in regard to the conservation and management of wildlife populations because it provides information regarding how quickly genetic diversity may be lost (Leberg 2005; Dudgeon and Ovenden 2015). From this prediction, it is possible to direct conservation efforts to mitigate this loss (Uzans et al. 2015). Therefore, given the potential association between Ne and the probability of extinction, estimates of Ne may be useful as an additional criterion in the assessment of species vulnerability (Leberg 2005; Willoughby et al. 2015).
Although the use of genetic parameters is largely overlooked and neglected in fisheries management plans and, consequently, in conservation policies for shark and ray species, there are a few good examples of their use. One such example is the Red List assessment of Stegostoma fasciatum (Stegostomatidae). In this case, the population genetic structure was used to analyze the two main populations, the Indian Ocean-Southeast Asian and Eastern Indonesian-Oceania populations, which were then assessed independently. Furthermore, Ne was used to estimate the approximate census size of the Eastern Indonesian-Oceania population. In another case, genetic data were used for stock delineation in the assessment of scalloped hammerhead populations (Miller et al. 2013), thus helping this shark species become the first shark species to be protected by the U.S. Endangered Species Act (Federal Register 80 FR 71774). For the scalloped hammerhead, stock delineation is important not only for increasing its population size but also for safeguarding its evolutionary dynamics. These examples clearly demonstrate the many advantages of adding genetic information to species assessments and management plans.
Limitations
Although the number of studies of shark and ray species has increased over recent decades, several significant limitations remain and can be highlighted, such as sampling protocols and population coverage, methodological issues, and appropriate use of molecular markers.
Sampling and geographical coverage
Similar to other marine apex predators and highly mobile species, there are many challenges to properly investigating the population structure and genetic diversity of sharks and rays. In general, sampling schemes are not always adequate in terms of sample size, geographical coverage, and collection method (Hindrikson et al. 2017; Letessier et al. 2017). A good strategy for solving this problem is the use of simple simulation software (e.g., POWSIM or SPOTG) that estimates statistical power (i.e., the probability of rejecting the null hypothesis when it is false) to simulate optimal combinations of sample size, number of loci, and allele frequency for any hypothetical degree of true differentiation (Ryman and Palm 2006; Hoban et al. 2013b). Additionally, recently, an initiative has been adopted to overcome these problems. Shark Share Global (SSG) (https://www.sharkshareglobal.org) provides an online database to which researchers can submit tissue samples, search, and request them from colleagues around the world. This allows researchers to obtain robust collections of tissues samples from various locations throughout the range of a species, enabling a better understanding of the population genetic structure and genetic diversity of the species studied. Despite aid from SSG to increase the sample size and geographical coverage, obtaining systematic and planned (as opposed to opportunistic) sampling for one specific location, sex, age, and time remains a challenge. Consequently, few studies based on such planned sampling have been performed to date (e.g., Chevolot et al. 2008; Veríssimo et al. 2017).
Methodological issues
While fin clipping has been the most commonly used method for collecting genetic data from sharks and rays, this method has some drawbacks, such as stress, injury, and even death after release (Wasko et al. 2003). In the past decade, a variety of less invasive techniques, including noninvasive genetic sampling, have been developed especially for internationally protected shark and ray species, in order to minimize such drawbacks (Larson et al. 2017). For example, Lieber et al. (2013) tested the potential of mucus swabs from a vulnerable species, the basking shark (Cetorhinus maximus, Cetorhinidae), at three molecular markers (cytochrome oxidase I (COI), CR mtDNA, and ITS2). Similarly, Kashiwagi et al. (2015) evaluated the PCR success of mtDNA ND5 and nuclear DNA RAG1 for manta rays, as well as microsatellite loci from manta ray mucus collected underwater using toothbrushes. Such collection methods combined with new DNA technology, which require less representative sampling, show promise as a solution for more sustainable and less invasive genetics studies.
Molecular markers
Another way to increase the geographical coverage and understanding of the population genetic structure and genetic diversity of sharks and rays is the use of common methods and sets of genetic markers, which could be universally comparable between studies. To date, several methods and molecular markers have been used for shark and ray genetic studies (Table 1). However, CR mtDNA is the most commonly used, either in part (e.g., Duncan et al. 2006; Frodella et al. 2016; Domingues et al. 2017) or in whole (e.g., Clarke et al. 2015; Bernard et al. 2016, 2017). Similarly, a set of highly polymorphic microsatellite loci, such as those used by Daly-Engel et al. (2012), could be standardized and used for multiple shark and ray species. However, the rapidly developing field of genomics holds great promise developing other DNA markers (SNPs) for shark and ray population analysis.
Future challenges
Recently, many authors have claimed that the use of genetic diversity as should be at the forefront of conservation policy and management and not used only as supporting information (Laikre 2010; Hoban et al. 2013a, b). However, we note that making genetic diversity data more promptly useful to policymakers requires overcoming some challenges in either scientific or policy arenas, as described below:
-
Prioritize shark and ray species that have narrow geographic distributions and are currently overexploited
-
Conduct genetic monitoring by sampling in temporal series to assess genetic variations over time
-
Apply genomics to shark and ray genetic research
-
Include more conservation geneticists in developing conservation policies
-
Improve communication between scientists and policymakers
Prioritize shark and ray species that have narrow geographic distributions and are currently overexploited
From a conservation genetics perspective, the worst situation is the representation of an endangered species as a single population (Frankham et al. 2002). Frequently, small populations are most likely to be affected by the loss of genetic diversity due to overfishing, which affects their evolutionary potential and results in an elevated risk of extinction (Frankham et al. 2002; Allendorf et al. 2008). Therefore, obtaining information at the level of genetic diversity for a species either with a narrow geographic distribution or within an isolated population is important for indicate the population fragility of that species. For example, the narrownose shark is a species endemic to the Southwest Atlantic Ocean with a narrow geographic distribution, which extends from Rio de Janeiro, Brazil, to Patagonia, Argentina. This shark species has experienced intense overfishing along its entire geographic range, including its nursery grounds (Massa et al. 2006). Moreover, the genetic diversity of the narrownose shark is among the lowest among all sharks (Table 2), making this species highly susceptible to overfishing in the short term and to low genetic diversity in the long term. This situation could affect other elasmobranch species that have narrow geographic distributions and are currently overfished, including critically endangered elasmobranchs such as the daggernose shark (Lessa et al. 2016), the Brazilian guitarfish (Pseudobatos horkelii, Rhinobatidae) (Vooren et al. 2005, as Rhinobatos horkelii) and the common angel shark (Squatina squatina, Squatinidae) (Ferretti et al. 2015). On the other hand, widely distributed shark and ray species are rarely panmictic from one end of their distribution area to the other, and instead, they show partitioning genetics (Castro et al. 2007; Ahonen et al. 2009; Clarke et al. 2015). However, low population genetic differentiation may not be informative on the appropriate spatial scale for management decisions. For example, Schmidt et al. (2009) found only low levels of genetic differentiation between geographically distinct whale shark populations, suggesting that conservation efforts must target international protection for this species. Furthermore, the asymmetric dispersal (females non-roving and males roving), consistent with male-mediated gene flow, that is common in many shark and ray species (e.g., Feldheim et al. 2014; Sellas et al. 2015) is another factor that must be considered in management decisions. Therefore, obtaining information regarding the extent of gene flow among populations is important for determining whether a species requires the maintenance of genetic diversity through migration (Frankham et al. 2002; Allendorf et al. 2013).
Genetic monitoring by sampling in temporal series to assess genetic variations over time
Genetic monitoring, as defined by Schwartz et al. (2007), is the quantification of temporal changes in population genetic parameters or other population data generated using molecular markers. This technique can be performed using ancient DNA (aDNA) from the dried jaws and vertebrae of sharks and rays archived in museums and private collections and even kept as exotic souvenirs (Nielsen et al. 2016). These data allow for retrospective monitoring to assess historical conditions, such as the temporal stability of the population structure, the loss of genetic diversity, and changes in the Ne, which are difficult to determine using traditional methods (Schwartz et al. 2007; Nielsen and Hansen 2008). Good examples of the use of this approach are mainly demonstrated in bony fish (e.g., Hauser et al. 2002; Nielsen and Hansen 2008; Bonamoni et al. 2016). However, there are currently a few examples of the use of aDNA for shark species. Gubili et al. (2015) sequenced a small fragment (135–228 bp) of mtDNA (D-loop) from 34- to 129-year-old dried cartilage and skin samples from six Carcharodon carcharias individuals and found greater genetic diversity (number of haplotypes and nucleotide and haplotype diversity) in the historical samples than in contemporary samples found in the Mediterranean Sea. Moreover, Li et al. (2015) used the complete mitochondrial genome of aDNA to infer the phylogeny and gene flow of endangered river sharks (Glyphis spp., Carcharhinidae). Therefore, the management of genetically depauperate populations must embrace the identification of source founders from genetically diverse populations (Allendorf et al. 2013) and genetic monitoring through sampling in temporal series to assess genetic variation over time (Allendorf et al. 2008; Laikre et al. 2008).
Applications of genomics in elasmobranch genetic research
DNA sequences, especially the control regions of mitochondrial DNA and microsatellites, are the markers most widely used in elasmobranchs to date (Table 1). However, the availability of new high-resolution molecular markers such as single-nucleotide polymorphisms (SNPs), promises a marked advance in genetic studies in the future. Currently, with the advances in next-generation sequencing (NGS), there are many methods to uncover and genotype thousands of SNPs that cover the entire genome in a single step at minimal cost, thus making NGS feasible for most labs (Stapley et al. 2010). We will not attempt to describe NGS, its methods, or associated analyses in detail, as these have been covered in other reviews (e.g., Rocha et al. 2013; Goodwin et al. 2016). Instead, we intend to highlight the need for using NGS approaches to better answer questions pertaining to shark and ray population genetics in the near future.
Even though 10–20 microsatellites are estimated to be equivalent to 100 SNPs, with the recent development of new methods such as the use of restriction-site-associated DNA tags (RAD-tags), tens of thousands of SNPs can be recovered from multiple individuals at the same time, thereby increasing the statistical power of fine-scale detection in discrete populations (Nielsen et al. 2009; Davey and Blaxter 2011; Rocha et al. 2013). Consequently, SNP analysis requires relatively small numbers of samples from a given location, which in turn is an advantage due to the numbers of shark and ray species that are currently threatened. Another advantage of using SNPs is that this approach aids in determining which parts of the genome are responsible for local adaptation even in cases of high gene flow, thereby enabling the identification of priority areas to be conserved (Nielsen et al. 2009). However, future studies using approaches based on genomic population outliers must be conducted carefully because these approaches still pose several challenges, including genotyping errors, the underlying population structure and false positives, variation in the mutation rate and limited sensitivity (false negatives) (Narun and Hess 2011; Tiffin and Ross-Ibarra 2014; Hoban et al. 2016; Flanagan et al. 2017). In fact, NGS technologies are having substantial effects on many areas of biology, including the analysis of genetic diversity in populations, and they promise an abrupt advance in genetic studies in the coming years (review in Nielsen et al. 2009). However, the use of NGS technologies in developing countries may still be cost-prohibitive, due to limited funding for basic research and because they require sophisticated bioinformatics systems, fast data processing and large data storage capabilities (Willette et al. 2014; Puckett 2017). Furthermore, although the pitfalls of mtDNA and microsatellite studies are fairly well known and can usually be recognized and tested, the drawbacks of NGS approaches are still being identified (Bowen et al. 2014).
To date, only one study has used neutral and outlier SNPs to infer the local adaptation of sharks. Using neutral SNPs (648,035 SNPs), Portnoy et al. (2015) found differences in the population structure of the bonnethead shark between the North Atlantic (North Carolina) and the Gulf of Mexico (Florida Bay, Tampa Bay and Panama City), whereas the use of 30 outlier SNPs showed fine-scale differences in population structures among all locations except for Tampa Bay and Florida Bay, where the population structures were homogenous. The authors attributed this local adaptation to north–south (latitudinal) clinal patterns in allele frequencies. More recently, Pazmiño et al. (2017) conducted a genome-wide analysis using 8103 neutral SNPs to investigate the population structure of the Galapagos shark (Carcharhinus galapangesis) over a small geographic range (Galapagos Marine Reserve). Those authors found two differentiated populations and a low estimated Ne of 200, suggesting that these populations are susceptible to extinction and are of concern for long-term conservation. In another recent paper, Corrigan et al. (2017) performed a genome-wide analysis using 2152 SNPs to examine the patterns of genetic admixture between the Galapagos shark and the dusky shark (Carcharhinus obscurus), two closely related sharks. However, even with genomic data providing novel insights, its use in the analysis of shark and ray species remains limited (e.g., Feutry et al. 2015; Portnoy et al. 2015; Delser et al. 2016; Pazmiño et al. 2017; Corrigan et al. 2017).
Including more conservation geneticists in international developing conservation policy making
The importance of genetic criteria to guarantee long-term population viability and conservation is well known in academia. However, outside academia, genetics is still largely overlooked and neglected in practical management and in national and international policies, though there are some exceptions (Laikre 2010). The main agencies interested in the conservation of shark and ray species do not have many conservation genetics specialists integrated into their teams. Members of these organizations are mainly fisheries scientists interested in assessing stock abundance, which demonstrates the lack of concern for genetics in assessing species (Laikre 2010). Therefore, we recommend the inclusion of more conservation geneticists in conservation organizations. This inclusion would facilitate the addition of genetic criteria to assessments and future management plans for shark and ray species.
Bridging the gap between genetic science and shark and ray conservation policies
Currently, there remains a gap between the genetics research that generates knowledge about genetic data (e.g., genetic diversity, population genetic structure and demographic history) and conservation organizations that use these data to establish protection measures that aim to allow populations to recover (Laikre 2010; Hoban et al. 2013a, b; Haig et al. 2016). Typically, policymakers and managers are not geneticists, and they have difficulty interpreting genetic data correctly; consequently, these data are often used incorrectly in the creation of management plans (Hoban et al. 2013a, b; Haig et al. 2016). This issue could be resolved, for example, by providing training to national and international conservation organizations. Workshops, courses and lectures for non-genetics researchers, conservation practitioners and decision-makers interested in sharks and rays could be organized to show how genetic diversity can be effectively used in management plans. This approach would provide an excellent opportunity to show that genetic diversity data could reveal a wide variety of information for conservation policies. Furthermore, the inclusion of genetics experts among national and international policymakers could be a good start for incorporating genetic information into conservation policy (Fig. 4). For example, the IUCN Conservation Genetics Specialist Group (CGSG) (http://www.cgsg.uni-freiburg.de), whose mission is to promote the use of genetics in conservation management and decision-making, was recently created. However, the inclusion of geneticists among policymakers requires improvement in the communication between scientists and policymakers. To bridge the divide between conservation genetics research and practice, Conservation Genetic Resources for Effective Species Survival (ConGRESS), a freely available online resource (http://congressgenetics.eu) that increases access to current knowledge, facilities implementation of studies and interpretation of available data, and fosters collaboration between researchers and practitioners, was recently launched (Hoban et al. 2013c). Thus, geneticists’ research could help with elaborating on and revising regional and global reports on sharks and rays and proposals that incorporate genetic data for the management and research of these species.
The link between scientific knowledge and conservation policies aimed at preserving the genetic diversity of shark and ray species. Scientists must generate knowledge that identifies the main threats to genetic diversity and their deleterious effects on shark and ray populations, whereas policymakers must consider the information provided by scientists and apply this information to their conservation policies in order to effectively manage species populations
Conclusions
In light of the above considerations, we can assume that the increasing overexploitation of shark populations, mainly by fisheries, will have a long-term impact on the genetic variability of these populations and thus will reduce their fitness and responsiveness to environmental changes. Therefore, future assessments of shark and ray populations by conservation organizations should definitely include genetic parameters. Furthermore, as an important outcome of this synthesis, we highlighted the limitations and future challenges to overcoming the gap in current knowledge through genetic studies of sharks and rays. Finally, we argue that geneticists can inform policymakers when and where genetic diversity will be important for shark and ray conservation.
References
Ahonen H, Harcourt RG, Stow AJ (2009) Nuclear and mitochondrial DNA reveals isolation of imperiled grey nurse shark populations (Carcharias taurus). Mol Ecol 18:4409–4421
Allendorf FW, England PR, Ritchie PA, Ryman N (2008) Genetic effects of harvest on wild animal populations. Trends Ecol Evol 23:327–337
Allendorf FW, Luikart G, Aitken SN (2013) Conservation and the genetics of population. Blackwell, Oxford
Andreotti S, von der Heyden S, Henriques R, Rutzen M, Meÿer M, Oosthuizen H, Metthee CA (2015) New insights into the evolutionary history of white sharks, Carcharodon carcharias. J Biogeogr 43:328–339
Andreotti S, Rutzen M, van der Walt, von der Heyden S, Henriques R, Meÿer M, Oosthuizen H, Metthee CA (2016) An integrated mark-recapture and genetic approach to estimate the population size of white sharks in South Africa. Mar Ecol Prog Ser 552:241–253
Arlyza IS, Shen KN, Durand JD, Borsa P (2013) Mitochondrial haplotypes indicate parapatric-like phylogeographic structure in blue-spotted maskray (Neotrygon kuhlii) from the coral Triangle Region. J Hered 4:725–733
Ashe JL, Feldhei KA, Fields AT, Reyier EA, Brooks EJ, O’Connell MT, Skomal G, Gruber SH, Chapman DD (2015) Local population structure and context-dependent isolation by distance in a large coastal shark. Mar Ecol Prog Ser 520:203–216
Barker AM, Nosal AP, Lewallen EA, Burton RS (2015) Genetic structure of leopard shark (Triakis semifasciata) populations along the Pacific coast of North America. J Exp Mar Biol Ecol 472:151–157
Barreto R, Ferretti F, Flemming JM, Amorim A, Andrade H, Worm B, Lessa R (2016) Trends in the exploitation of South Atlantic shark populations. Cons Biol 30:792–804
Belgrano A, Fowler CW (2013) How fisheries affect evolution. Science 342:1176
Benavides MT, Feldheim KA, Duffy CA et al (2011a) Phylogeography of the copper shark (Carcharhinus brachyurus) in the southern hemisphere: implications for the conservation of a coastal apex predator. Mar Freshw Res 62:861–869
Benavides MT, Horn RL, Feldheim KA et al (2011b) Global phylogeography of the dusky shark Carcharhinus obscurus: implications for fisheries management and monitoring the shark fin trade. Endanger Species Res 14:13–22
Bernal MA, Sinai NL, Rocha C, Gaither MR, Dunker F, Rocha LA (2015) Long-term sperm storage in the brownbanded bamboo shark Chiloscyllium punctatum. J Fish Biol 86:1171–1176
Bernard AM, Feldheim KA, Heithaus MR, Wintner SP, Wetherbee BM, Shivji MS (2016) Global population genetic dynamics of a highly migratory, apex predator shark. Mol Ecol 25:5312–5329
Bernard AM, Rebekah LH, Chapman DD, Feldheim KA, Garla RC, Brooks EJ, Gore MA, Shivji MS (2017) Genetic connectivity of a coral reef ecosystem predator: the population genetic structure and evolutionary history of the Caribbean reef shark (Carcharhinus perezi). J Biogeogr 44:2488–2500
Bitalo DN, Maduna SN, da Silva C, Roodt-Wilding R, van der Merwe AE (2015) Differential gene flow patterns for two commercially exploited shark species, tope (Galeorhinus galeus) and common smoothhound (Mustelus mustelus) along the south-west coast of South Africa. Fish Res 172:190–196
Blower DC, Pandolfi JM, Bruce BD, Gomez-Cabrera MD, Ovenden JR (2012) Population genetics of Australian white sharks reveals fine-scale spatial structure, transoceanic dispersal events and low effective population sizes. Mar Ecol Prog Ser 455:229–244
Bonamoni S, Therkildsen NO, Retzel A et al (2016) Historial DNA documents long distance natal homing in marine fish. Mol Ecol 25:2727–2734
Boomer JJ, Harcourt RG, Francis M, Stow AJ (2012) Genetic divergence, speciation and biogeography of Mustelus (sharks) in the central Indo-Pacific and Australasia. Mol Phylogenet Evol 64:697–703
Booth W, Schuett GW (2011) Molecular genetic evidence for alternative reproductive strategies in North American pitvipers (Serpentes: Viperidae): long-term sperm storage and facultative parthenogenesis. Biol J Linn Soc 104:934–942
Bowen BW, Shanker K, Yasuda N et al (2014) Phylogeography unplugged: comparative surveys in the genome era. Bull Mar Sci 90:13–46
Bräutigam A, Callow M, Campbell IR, Camhi MD, Cornish AS, Dulvy NK, Fordham SV, Fowler SL, Hood AR, McClennen C, Reuter EL, Sant G, Simpfendorfer CA, Welch DJ (2015) Global priorities for conserving sharks and rays: a 2015–2025 Strategy
Byrne RJ, Avise JC (2012) Genetic mating system of the brown smoothhound shark (Mustelus henlei), including a literature review of multiple paternity in other elasmobranch species. Mar Biol 159:749–756
Camargo SM, Coelho R, Chapman D, Howey-Jordan L et al (2016) Structure and genetic variability of the oceanic whitetip shark, Carcharhinus longimanus, determined using mitochondrial DNA. PLoS ONE 11:e0155623. https://doi.org/10.1371/journal.pone.0155623
Cardeñosa D, Hyde J, Caballero S (2014) Genetic diversity and population structure of the pelagic shark (Alopias pelagicus) in the Pacific Ocean: evidence for two evolutionary significant units. PLoS ONE 9:e110193. https://doi.org/10.1371/journal.pone.0110193
Castillo-Olguín E, Uribe-Alcocer M, Díaz-Jaimes P (2012) Assessment of the population genetic structure of Sphyrna lewini to identify conservation units in the Mexican Pacific. Cienc Mar 38:635–652
Castillo-Páez A, Sosa-Nishizaki O, Sandoval-Castillo J, Galván-Magaña F, Blanco-Parra MD, Rocha-Olivares A (2014) Strong population structure and shallow mitochondrial phylogeny in the banded guitarfish, Zapteryx exasperate (Jordan y Gilbert, 1880), from the Northen Mexican Pacific. J Hered 105:91–100
Castro ALF, Stewart BS, Wilson SG, Hueter RE, Meekan MG, Motta PJ (2007) Population genetic structure of Earth’s largest fish, the whale shark (Rhincodon typus). Mol Ecol 16:5183–5192
Catarino D, Knutsen H, Veríssimo A et al (2015) The pillars of Hercules as a bathymetric barrier to gene flow promoting isolation in a global deep-sea shark (Centroscymnus coelolepis). Mol Ecol 24:6061–6079
Chabot CL (2015) Microsatellite loci confirm a lack of population connectivity among globally distributed populations of the tope shark Galeorhinus galeus (Triakidae). J Fish Biol 87:371–385
Chabot CL, Allen LG (2009) Global population structure of the tope (Galeorhinus galeus) inferred by mitochondrial control region sequence data. Mol Ecol 18:545–552
Chabot CL, Espinoza M, Mascareñas-Osorio I, Rocha-Olivares A (2015) The effect of biogeographic and phylogeographic barriers on gene flow in the brown smoothhound shark, Mustelus henlei, in the northeastern Pacific. Ecol Evol 5:1585–1600
Chapman DD, Prodöhl PA, Gelsleichter J, Manire CA, Shivji MS (2004) Predominance of genetic monogamy by females in a hammerhead shark, Sphyrna tiburo: implications for shark conservation. Mol Ecol 13:1965–1974
Chapman DD, Shivji MS, Louis E, Sommer J, Fletcher H, Prodöhl PA (2007) Virgin birth in a hammerhead shark. Biol Lett 3:425–427
Chapman DD, Pinhal D, Shivji MM (2009) Tracking the fin trade: genetic stock identification in western Atlantic scalloped hammerhead sharks Sphyrna lewini. Endanger Species Res 9:221–228
Chapman DD, Simpfendorfer C, Wiley TR, Poulakis GR, Curtis C, Tringali M, Carlson J, Feldheim K (2011) Genetic diversity despite population collapse in a critically endangered marine fish: the smalltooth sawfish (Pristis pectinata). J Hered 102:643–652
Chapman DD, Feldheim KA, Papastamatiou YP, Hueter RE (2015) There and back again: a review of residency and return migrations in sharks, with implications for population structure and management. Ann Rev Mar Sci 7:547–570
Chevolot M, Hoarau G, Stam WT, Olsen J (2006) Phylogeography and population structure of thornback rays (Raja clavata L., Rajidae). Mol Ecol 15:3693–3705
Chevolot M, Wolfs PH, Pálsson K, Riijnsdorp AD, Wytze TS, Olsen JL (2007a) Population structure and historical demography of the thorny skate (Amblyraja radiata, Rajidae) in the North Atlantic. Mar Biol 151:1275–1286
Chevolot M, Ellis JR, Rijnsdorp AD, Stam WT, Olsen JL (2007b) Multiple paternity analysis in the thornback ray Raja clavata L. J Hered 98:712–715
Chevolot M, Ellis JR, Rijnsdorp AD, Stam WT, Olsen JL (2008) Temporal changes in allele frequencies but stable genetic diversity over the past 40 years in the Irish Sea population of thornback ray, Raja clavata. Heredity 101:120–126
Clarke SC, Magnussen JE, Abercrombie DL, McAllister MK, Shivji MS (2006) Identification of shark species composition and proportion in the Hong Kong shark fin market based on molecular genetics and trade records. Conserv Biol 20:201–211
Clarke SC, Harley SJ, Hoyle SD, Rice JS (2013) Population trends in pacific oceanic sharks and the utility of regulations on shark finning. Conserv Biol 27:197–209
Clarke CR, Karl SA, Horn RL, Bernard AM, Lea JS, Hazin FHZ, Prodöhl PA, Shivji MS (2015) Global mitochondrial DNA phylogeography and population structure of the silky shark, Carcharhinus falciformis. Mar Biol 162:945–955
Conrath CL, Musick JA (2012) Reproductive biology of elasmobranchs. In: Carrier JC, Musick JA, Heithaus MR (eds) Biology of sharks and their relatives, 2rd edn. CRC Press, Boca Raton, pp 291–312
Corrigan S, Delser PM, Eddy C, Duffy C, Yang L, Chenhong L, Bazinet AL, Mona S, Naylor GJP (2017) Historical introgression drivers pervasive mitochondrial admixture between two species of pelagic sharks. Mol Phylogenet Evol 110:122–126
Cortés E (2002) Incorporating uncertainty into demographic modeling: application to shark populations and their conservation. Conser Biol 16:1048–1062
Daly-Engel TS, Grubbs RD, Feldheim KA, Bowen B, Toonen RJ (2010) Is multiple mating beneficial or unavoidable? Low multiple paternity and genetic diversity in the shortspine spurdog Squalus mitsukurii. Mar Ecol Prog Ser 403:255–267
Daly-Engel TS, Seraphin KD, Holland KN, Coffey JP, Nance HA, Toonen RJ, Bowen BW (2012) Global phylogeography with mixed-marker analysis reveals male-mediated dispersal in the endangered scalloped hammerhead shark (Sphyrna lewini). PLoS ONE 7:e29986. https://doi.org/10.1371/journal.pone.0029986
Davey JW, Blaxter ML (2011) RADSeq: next-generation population genetics. Brief Funct Genomics 5:416–423
Delser PM, Corrigan S, Hale M, Li C, Veuille M, Planes S, Naylor G, Mona S (2016) Population genomics of C. melanopterus using target gene capture data: demographic inferences and conservation perspectives. Sci Rep 6:33753
DiBattista J (2008) Pattern of genetic variation in anthropogenically impacted populations. Conserv Genet 9:141–156
Domingues RR, Hilsdorf AWS, Mahmood MS, Hazin FVH, Gadig OBF (2017) Effects of the pleistocene on the mitochondrial population genetic structure and demographic history of the silky shark (Carcharhinus falciformis) in the western Atlantic Ocean. Rev Fish Biol Fish. https://doi.org/10.1007/s11160-017-9504-z
Dudgeon CL, Ovenden JR (2015) The relationship between abundance and genetic effective population size in elasmobranchs: an example from the globally threatened zebra shark Stegostoma fasciatum within its protected range. Conserv Gen 16:1443–1454
Dudgeon CL, Broderick D, Ovenden JR (2009) IUCN classification zones concord with, but underestimate, the population genetic structure of the zebra shark Stegostoma fasciatum in the Indo-West Pacific. Mol Ecol 18:248–261
Dudgeon CL, Blower DC, Broderick D, Giles JL, Holmes BJ, Kashiwagi T, Krück NC, Morgan JAT, Tillett BJ, Ovenden JR (2012) A review of the application of molecular genetics for fisheries management and conservation of sharks and rays. J Fish Biol 80:1789–1843
Dulvy NK, Fowler SL, Musick JA, Cavanagh RD et al (2014) Extinction risk and conservation of the world’s sharks and rays. eLife 3:e00590
Duncan KM, Martin AP, Bowen BW, Couet HG (2006) Global phylogeography of the scalloped hammerhead shark (Sphyrna lewini). Mol Ecol 15:2239–2251
Ebert DA, Fowler S, Compagno L (2013) Sharks of the world: a fully illustrated guide. Wild Nature Press, Plymouth
Escatel-Luna E, Adams DH, Uribe-Alcocer M, Islas-Villanueva V, Díaz-Jaimes P (2015) Population genetic structure of the bonnethead shark, Sphyrna tiburo, from the Western North Atlantic Ocean based on mtDNA sequences. J Hered 106:355–365
Federal Register Volume 80, Issue 221 (Nov 17, 2015)
Feldheim KA, Gruber SH, Ashley MV (2001) Population genetic structure of the lemon shark (Negaprion brevirostris) in the western Atlantic: DNA microsatellite variation. Mol Ecol 10:295–303
Feldheim KA, Gruber SH, Dibattista JD et al (2014) Two decades of genetic profiling yields first evidence of natal philopatry and long-term fidelity to parturition sites in sharks. Mol Ecol 23:110–117
Ferretti F, Worm B, Britten GL, Heithaus MR, Lotze HK (2010) Patters and ecosystem consequences of shark declines in the ocean. Ecol Lett 13:1055–1071
Ferretti BLS, Mendonça FF, Coelho R, Vasconcelos PG, Hazin FVH, Romanov EV, Oliveira C, Santos MN, Foresti F (2015) High connectivity of the crocodile shark between Atlantic and Southwest Indian Oceans: highlights for conservation. PLoS ONE 10:e0117549. https://doi.org/10.1371/journal.pone.0117549
Feutry P, Kyne PM, Pillans RD, Chen X, Marthick JR, Morgan DL, Grewe PM (2015) Whole mitogenome sequencing population structure of the critically endangered sawfish Pristis pristis. Mar Ecol Prog Ser 533:237–244
Fields AT, Feldheim KA, Poulakis GR, Chapman DD (2015) Facultative parthenogenesis in a critically endangered wild vertebrate. Curr Biol 25:446–447
Flanagan SP, Forester BR, Latch EK, Aitken SN, Hoban S (2017) Guidelines for planning genomic assessment and monitoring of locally adaptive variation to inform species conservation. Evol Appl. https://doi.org/10.1111/eva.12569
Forselledo R, Pons M, Miller P, Domingos A (2008) Distribution and population structure of the pelagic stingray, Pteroplatytrygon violacea (Dasyatidae), in the south-western Atlantic. Aquat Living Resour 21:357–363
Frankham R (2010) Challenges and opportunities of genetic approaches to biological conservation. Biol Cons 143:1919–1927
Frankham R (2014) Genetics in conservation management: revised recommendations for the 50/500 rules, Red List criteria and population viability analyses. Biol Conserv 170:56–63
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Frederico RG, Farias I, Araújo MLG, Charvet-Almeida P, Alves-Gomes JA (2012) Phylogeography and conservation genetics of the Amazonian freshwater stingray Paratrygon aiereba Müller & Henle, 1841 (Chondrichthyes: Potamotrygonidae). Neotrop Ichtyol 10:71–80
Frodella N, Cannas R, Velonà A et al (2016) population connectivity and phylogeography of the Mediterranean endemic skate Raja polystigma and evidence of its hybridization with the parapatric sibling R. montagui. Mar Ecol Prog Ser 554:99–113
Fu M, Wnag J, Ding S, Du J, Su Y (2010) Studies on the genetic structure and genetic subdividion of white spotted bamboo shark, Chiloscyllium plagiosum, by analyzing mitochondrial Cyt b genes. J Trop Ocean 29:86–91
Fung HC, Waples RS (2017) Perfomance of IUCN proxies for generation length. Cons Biol 31:739–961
Gaida IH (1997) Population structure of the pacific angel shark, Squatina californica (Squatiniformes: Squatinidae), around the California Channel Islands. Copeia 4:738–744
Gallagher AJ, Hammerschlag N, Shiffman DS, Giert S (2014) Evolved for extinction: the cost and conservation implications of specialization in hammerhead sharks. Bioscience 67:619–624
Galván-Tirado C, Díaz-Jaimes P, García-de León FJ, Galván-Magaña FG, Uribe-Alcocer M (2013) Historical demography and genetic differentiation inferred from the mitochondrial DNA of the silky shark (Carcharhinus falciformis) in the Pacific Ocean. Fish Res 147:36–46
Garber AF, Tringali EM, Franks EJS (2005) Population genetic and phylogeographic structure of wahoo, Acanthocybium solandri, from the western central Atlantic and central Pacific Oceans. Mar Biol 147:205–214
Garcia G, Pereyra S, Gutierrez V, Oviedo S, Miller P, Domingo A (2015) Population structure of Squatina guggenheim (Squatiniformes, Squatinidae) from the south-western Atlantic Ocean. J Fish Biol 86:186–202
Gardner MG, Ward RD (1998) Taxonomic affinities within Australian and New Zealand Mustelus sharks (Chondrichthyes: Triakidae) inferred from allozymes, mitochondrial DNA and precaudal vertebrae counts. Copeia 2:356–363
Garza JC, Williamson EG (2001) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10:305–318
Geraghty PT, Williamson JE, Macbeth WG, Wintner SP, Harry AV, Ovenden JR, Gillings MR (2013) Population expansion genetic structure in Carcharhinus brevipinna in the southern Indo-Pacific. PLoS ONE 8:e75169
Geraghty PT, Williamson JE, Macbeth WG, Blower DC, Morgan JAT, Johnson G, Ovenden JR, Gillings MR (2014) Genetic structure and diversity of two highly vulnerable Carcharhinids in Australian waters. Endanger Species Res 24:45–60
Giles JL, Ovenden JR, Almojil D, Garvilles E, Khampetch K, Manjebrayakath H, Riginos C (2014) Extensive genetic population structure in the Indo-West Pacific spot-tail shark, Carcharhinus sorrah. Bull Mar Sci 90:427–454
Giles JL, Riginos C, Naylor GJP, Ovenden JR (2016) Genetic and phenotypic diversity in the wedgefish Rhynchobatus australiae, a threatened ray of high value in the shark fin trade. Mar Ecol Prog Ser 548:165–180
Goodwin S, McPherson JD, McCombie WR (2016) Coming of age: ten years of next-generation sequencing technologies. Nat Rev Genet 17:333–351
Grant WS, Cheng W (2012) Incorporating deep and shallow components of genetic structure into the management of Alaskan red king crab. Evol Appl 5:820–837
Grogan ED, Lund R, Greenfest-Allen E (2012) The origin and relationship of early Chondrichthyes. In: Carrier JC, Musick JA, Heithaus MR (eds) Biology of sharks and their relatives, 2nd edn. CRC Press, Boca Raton, pp 3–29
Gubili C, Robinson CEC, Cliff G et al (2015) DNA from historical and trophy samples provides insights into white shark population origins and genetic diversity. Endang Species Res 27:233–241
Haig SM, Miller MP, Bellinger R, Draheim HM, Mercer DM, Mullins TD (2016) The conservation genetics juggling act: integrating genetics and ecology, science and policy. Evol Appl 9:181–195
Hare MT, Nunney L, Schwartz MK, Ruzzante DE, Burford M, Waples RS, Palstra F (2011) Understanding and estimating effective population size for practical application in marine species management. Conserv Biol 3:438–449
Hauser L, Carvalho GR (2008) Paradigm shifts in marine fisheries genetics: ugly hypotheses slain by beautiful facts. Fish Fish 9:333–362
Hauser L, Adcock GJ, Smith PJ, Ramírez JHB, Carvalho GR (2002) Loss of microsatellite diversity and low effective population size in an overexploited population of New Zealand snapper (Pagrus auratus). Proc Natl Acad Sci USA 99:11742–11747
Heino M, Pauli DB, Dieckmann U (2015) Fisheries-induced evolution. Annu Rev Ecol Evol Syst 46:461–480
Heist EJ, Graves JE, Musick JA (1995) Population genetics of the sandbar shark (Carcharhinus plumbeus) in the Gulf of Mexico and Mid-Atlantic Bight. Copeia 3:555–562
Heist EJ, Musick JA, Graves JE (1996a) Genetic population structure of the shortfin mako (Isurus oxyrinchus) inferred from restriction fragment length polymorphism analysis of mitochondrial DNA. Can J Fish Aquat 53:583–588
Heist EJ, Musick JA, Graves JE (1996b) Mitochondrial DNA diversity and divergence among sharpnose sharks, Rhizoprionodon terraenovae, from the Gulf of Mexico and Mid-Atlantic Bight. Fish Bull 94:664–668
Heithaus MR, Frid A, Wirsing AJ, Worm B (2008) Predicting ecological consequences of marine top predator declines. Trends Ecol Evol 23:202–210
Hernández S, Daley R, Walker T, Braccini M, Varela A, Francis MP, Ritchie (2015) Demographic history and the South Pacific dispersal barrier for school shark (Galeorhinus galeus) inferred by mitochondrial DNA and microsatellite DNA mark. Fish Res 176:132–142
Herndon A, Gallucci VF, DeMaster D, Burke W (2010) The case for an international commission for the conservation and management of sharks (ICCMS). Mar Policy 34:1239–1248
Hindrikson M, Remm J, Pilot M et al (2017) Wolf population in Europe: a systematic review, meta-analysis and suggestions for conservation and management. Biol Rev 92:1601–1629
Hobab SM, Arntzen W, Bertorelle G et al (2013c) Conservation genetic for effective species survival (ConGRESS): bridging the divide between conservation research and practice. J Nat Conserv 21:433–437
Hoban SM, Hauffe HC, Pérez-Espona S et al (2013a) Bringing genetic diversity to the forefront of conservation policy and management. Conserv Genet Resour 5:593–598
Hoban S, Gaggiotti O, Bertorelle G (2013b) Sample planning optimization tool for conservation and population genetics (SPOTG): a software for choosing the appropriate number of markers and samples. Methods Ecol Evol 4:299–303
Hoban S, Kelley JL, Lotterhos KE et al (2016) Finding the genomic basis of local adaptation: pitfalls, practical solutions, and future directions. Am Nat 188:379–397
Hoelzel AR, Shivji MS, Magnussen J, Francis MP (2006) Low worldwide genetic diversity in the basking sharks (Cetorhinus maximus). Biol Lett 2:639–642
Hoffman M, Hilton-Taylor C, Angulo A et al (2010) The impact of conservation on the status of the world’s vertebrates. Science 330:1503–1509
Jorgensen SJ, Reeb CA, Chapple TK, Anderson S, Perle C, Van Sommeran SR, Fritz-Cope C, Brown AC, Klimley AP, Block BA (2009) Philopatry and migration of Pacific white sharks. Proc R Soc B 277:679–688
Karl SA (2008) The effect of multiple paternity on the genetically effective size of a population. Mol Ecol 17:3973–3977
Karl SA, Castro ALF, Garla RC, Lopez JA, Charvet P, Burgess GH (2011) Phylogeography and conservation of the bull shark (Carcharhinus leucas) inferred from mitochondrial and microsatellite DNA. Conserv Gen 12:371–382
Karl SA, Castro ALF, Garla RC (2012) Population genetics of the nurse shark (Ginglymostoma cirratum) in the western Atlantic. Mar Biol 159:489–498
Kashiwagi T, Maxwell EA, Marshall AD, Christensen AB (2015) Evaluating manta ray mucus as an alternative DNA source for population genetics study: underwater-sampling, dry-storage and PCR success. PeerJ 3:e1188. https://doi.org/10.7717/peerj.1188
Keeney DB, Heist EJ (2006) Worldwide phylogeography of the blacktip shark (Carcharhinus limbatus) inferred from mitochondrial DNA reveals isolation of western Atlantic populations coupled with recent Pacific dispersal. Mol Ecol 15:3669–3679
Keeney DB, Heupel ME, Hueter RE, Heist EJ (2003) Genetic heterogeneity among blacktip shark, Carcharhinus limbatus, continental nurseries along the U.S. Atlantic and Gulf of Mexico. Mar Biol 143:1039–1046
Keeney DB, Heupel M, Hueter RE, Heist EJ (2005) Microsatellite and mitochondrial analyses of the genetic structure of blacktip shark (Carcharhinus limbatus) nurseries in the northwestern Atlantic, Gulf of Mexico, and Caribbean Sea. Mol Ecol 14:1911–1923
Kenchington E, Heino M, Nielsen EE (2003) Managing marine genetic diversity: time for action? ICES J Mar Sci 60:1172–1176
King JR, Wetklo M, Supernaul J, Taguchi M, Yokawa K, Sosa-Nishizaki O, Withler RE (2015) Genetic analysis of stock structure of blue shark (Prionace glauca) in the north Pacific. Fish Res 172:181–189
Kousteni V, Kasapidis P, Kotoulas G, Megalofonou P (2015) Strong population genetic structure and contrasting histories for the small-spotted catshark (Scyliorhinus canicula) in the Mediterranean Sea. Heredity 114:1–11
Laikre L (2010) Genetic diversity is overlooked in international conservation policy implementation. Conserv Genet 11:349–354
Laikre L, Larsson LC, Palmé A, Charlier J, Josefsson M, Ryman N (2008) Potentials for monitoring gene level biodiversity: using Sweden as an example. Biodivers Conserv 17:893–910
Laikre L, Schwartz MK, Waples RS et al (2010b) Compromising genetic diversity in the wild: unmonitored large-scale release of plants and animals. Trends Ecol Evol 25:520–529
Larson S, Farrer D, Lowry D, Ebert DA (2015) Preliminary observations of population genetics and relatedness of the broadnose sevengill shark, Notorynchus cepedianus, in two Northeast Pacific estuaries. PLoS ONE 10:e0129278. https://doi.org/10.1371/journal.pone.0129278
Larson SE, Daly-Engel TS, Phillips NM (2017) Review of current conservation genetic analyses of Northeast Pacific sharks. Adv Mar Biol 77:79–110
Last PR, White WT, Carvalho MR, Séret B, Stehmann MFW, Naylor GJP (2016) Rays of the world. Cornell University Press, Ithaca
Lavery S, Shaklee JB (1989) Population genetics of two tropical sharks, Carcharhinus tilstoni and C. sorrah in Northern Australia. Mar Freshw Res 40:541–547
Le Port A, Lavery S (2012) Population structure and phylogeography of the short-tailed stingray, Dasyatis brevicaudata (Hutton 1875), in the southern Hemisphere. J Hered 103:1–12
Leberg P (2005) Genetic approaches for estimating the effective size of populations. J Wildl Res 69:1385–1399
Lessa R, Batista VS, Santana FM (2016) Close to extinction? The collapse of the endemic daggernose shark (Isogomphodon oxyrhynchus) off Brazil. Global Ecol Evol 7:70–81
Letessier TB, Bouchet PJ, Meeuwig JJ (2017) Sampling mobile oceanic fishes and sharks: implications for fisheries and conservation planning. Biol Rev 92:627–646
Lewallen EA, Anderson TW, Bohonak AJ (2007) Genetic structure of leopard shark (Triakis semifasciata) populations in California waters. Mar Biol 152:599–609
Li N, Song N, Cheng G, Gao T (2013) Genetic diversity and population structure of the red stingray, Dasyatis akajei inferred by AFLP marker. Biochem Syst Ecol 51:130–137
Li N, Chen X, Sun D, Song N, Lin Q, Gao T (2015) Phylogeography and population structure of the red stingray, Dasyatis akajei inferred by mitochondrial control region. Mitochondr DNA 26:505–513
Li W, Dai X, Zhu J, Tian S, He S, Wu F (2016) Genetic differentiation in blue shark, Prionace glauca, from the central Pacific Ocean, as inferred by mitochondrial cytochrome b region. Mitochondr DNA 28:575–578
Lieber L, Berrow S, Johnston E, Graham H, Hall J, Gubili C, Sims DW, Jones CS, Noble LR (2013) Mucus: aiding elasmobranch conservation through non-invasive genetic sampling. Endanger Species Res 21:215–222
Lucifora LO, García VB, Worm B (2011) Global diversity hotspots and conservation priorities for sharks. PLoS ONE 6:e19356
Maduna SN, da Silva C, Wintner SP, Roodt-Wilding R, van der Merwe AE (2016) When two oceans meet: regional population genetics of an exploited coastal shark, Mustelus mustelus. Mar Ecol Prog Ser 544:183–196
Martin AP, Naylor GJP, Palumbi SR (1992) Rate of mitochondrial evolution is low in sharks compared to mammals. Nature 357:153–155
Massa A, Hozbor N, Chiaramonte GE, Balestra AD, Vooren CM (2006) Mustelus schmitti. The IUCN Red List of threatened species 2006: e.T60203A12318268. http://dx.doi.org/10.2305/IUCN.UK.2006.RLTS.T60203A12318268.en. Accessed 07 Nov 2016
McClenachan L, Cooper AB, Dulvy NK (2016) Rethinking trade-driven extinction risk in marine and terrestrial megafauna. Curr Biol 26:1–7
McCusker MR, Bentzen P (2010) Positive relationships between genetic diversity and abundance in fishes. Mol Ecol 19:4852–4862
MacDonald CM (1988) Genetic variation, breeding structure and taxonomic status of the gummy shark Mustelus antarticus in Southern Australian waters. Mar Freshw Res 39:641–648
McNelly JA, Miller KR, Reid WV, Mittermeier RA, Werner TB (1990) World Conservation Union. World Resources Institute, Conservation International, World Wildlife Fund–US, and the World Bank, Washington, D.C
Mendonça FF, Oliveira C, Gadig OBF, Foresti F (2011) Phylogeography and genetic population structure of Caribben sharpnose shark Rhizoprionodon porosus. Rev Fish Biol Fish 21:799–814
Mendonça FF, Oliveira C, Gadig OBF, Foresti F (2013) Diversity and genetic population of the Brazilian sharpnose shark Rhizoprionodon lalandii. Aquat Conserv Mar Fresh Ecosyst 23:850–9857
Miller MH, Carlson J, Cooper P, Kobayashi D, Wilson J (2013) Status review report: scalloped hammerhead shark (Sphyrna lewini). Report to National Marine Fisheries Service, Office of Protected Resources, p 131
Momigliano P, Harcourt R, Robbins WD, Stow A (2015) Connectivity in grey reef sharks (Carcharhinus amblyrhynchos) determined using empirical and simulated genetic data. Sci Rep 5:13229
Mourier J, Planes S (2013) Direct genetic evidence for reproductive philopatry and associated fine-scale migration in female blacktip reef sharks (Carcharhinus melanopterus) in French Polynesia. Mol Ecol 22:201–214
Mourier J, Burray J, Schultz JK, Clua E, Planes S (2013) Genetic network and breeding of a sicklefin lemon shark (Negaprion acutidens) population in the Society Islands, French Polynesia. PLoS ONE 8:e73899. https://doi.org/10.1371/journal.pone.0073899
Murray BW, Wang JY, Yang SC, Stevens JD, Fisk A, Svavarsson J (2008) Mitochondrial cytochrome b variation in sleeper sharks (Squaliformes: Somniosidae). Mar Biol 153:1015–1022
Musick JA, Burgess G, Cailliet G, Camhi M, Fordham S (2000) Management of sharks and their relatives (Elasmobranchii). Fisheries 25:9–13
Nance HA, Klimley P, Galván-Magaña F, Martínez-Ortíz J, Marko PB (2011) Demographic process underlying subtle pattern of population structure in the scalloped hammerhead shark, Sphyrna lewini. PLoS ONE 6:e21459. https://doi.org/10.1371/journal.pone.0021459
Narun SR, Hess JE (2011) Comparison of F ST outlier tests for SNP loci under selection. Mol Ecol Resour 11:184–194
Nelson JS, Grande TC, Wilson MHV (2016) Fishes of the world. Wiley, Hoboken
Newby JN, Darden T, Shedlock AM (2014) Population genetic structure of spotted eagle rays, Aetobatus narinari, off Sarasota, Florida and the Southeastern United States. Copeia 3:503–512
Nielsen EE, Hansen MM (2008) Waking the dead: the value of population genetic analyses of historical samples. Fish Fish 9:450–461
Nielsen EE, Hemmer-Hansen J, Larsen PF, Bekkevold D (2009) Population genomics of marine fishes: identifying adaptive variation in space and time. Mol Ecol 18:3128–3150
Nielsen EE, Morgan JAT, Maher SL, Edson J, Gauthier M, Pepperell J, Holmes BJ, Bennett MB, Ovenden JR (2016) Extracting DNA from ‘jaws’: high yield and quality from archived tiger shark (Galeocerdo cuvier) skeletal material. Mol Ecol Res 17:431–442
O’Leary SJ, Feldheim KA, Fields AT, Natanson LJ, Wintner S, Hussey N, Shivji MS, Chapman DD (2015) Genetic diversity of white sharks, Carcharodon carcharias, in the Northwest Atlantic and Southern Africa. J Hered 106:1–8
Oñate-González EC, Rocha-Olivares A, Saavedra-Sotelo N, Sosa-Nishizaki O (2015) Mitochondrial genetic structure and matrilineal origin of white sharks, Carcharodon carcharias in the Northeastern Pacific: implications for their conservation. J Hered 106:347–354
Ovenden JR, Kashiwagi T, Broderick D, Giles J, Salini J (2009) The extent of population genetic subdivision differs among four co-distributed shark species in the Indo-Australian archipelago. BMC Evol Biol 9:40
Ovenden JR, Morgan JA, Street R, Tobin A, Simpfendorfer C, Macbeth W, Welch D (2011) Negligible evidence species Rhizoprionodon acutus (Rüppell, 1837) and Sphyrna lewini (Griffith & Smith, 1834) with contrasting biology. Mar Biol 158:1497–1509
Ovenden JR (2013) Crinkles in connectivity: combining genetics and other types of biological data to estimate movement and interbreeding between populations. Mar Freshwater Res 64:201–207
Ovenden JR, Berry O, Welch DJ, Buckworth R, Dichmont CM (2013) Ocean’s eleven: a critical evaluation of the role of population, evolutionary and molecular genetics in the management of wild fisheries. Fish Fish 16:125–159
Ovenden JR, Blower DC, Jones AT, Moore A, Bustamante C, Buckworth RC, Bennett MB, Dudgeon CL (2016) Can estimates of genetic effective population size contribute to fisheries stock assessments? J Fish Biol 89:2505–2518
O’Brien SM, Gallucci VF, Hauser L (2013) Effects of species biology on the historical demography of sharks and their implications for likely consequences of contemporary climate change. Conserv Genet 14:125–144
Pardini AT, Jones CS, Noble LR, Kreiser B, Malcolm H, Bruce BD, Stevens JD, Cliff G, Scholl MC, Francis M, Clinton CAJ, Duffy P, Martin AP (2001) Sex-biased dispersal of great white sharks. Science 412:139–140
Parsolini P, Ragazzini C, Zaccaro Z, Cariani A, Ferrara G, Gonzalez EG, Landi M (2011) Quartenary geographical sibling speciation and population structuring in the eastern Atlantic skates (suborder Rajoidea) Raja clavata and R. straeleni. Mar Biol 158:2173–2186
Pazmiño Da, Maes GE, Simpfendorfer CA, Salinas-de-León P, van Herwerden L (2017) Genome-wide SNPs reveal low effective population size within confined management units of the highly vagile Galapagos shark (Carcharhinus galapagensis). Conserv Genet 18:1151–1163
Pereyra S, Garcia G, Miller P, Oviedo S, Domingo A (2010) Low genetic diversity and population structure of the narrownose shark (Mustelus schmitti). Fish Res 106:468–473
Phillips NM, Chaplin JA, Morgan DL, Peverell SC (2011) Population genetic structure and genetic diversity of three critically endangered Pristis sawfishes in Australian waters. Mar Biol 158:903–915
Pichler FB, Baker CS (2000) Loss of genetic diversity in the endemic Hector’s dolphin due to fisheries-related mortality. Proc Biol Sci 267:97–102
Pinsky ML, Palumbi SR (2014) Meta-analysis reveals lower genetic diversity in overfished populations. Mol Ecol 23:29–39
Plank SM, lowe CG, Feldheim KA, Wilson RR Jr, Brusslan JA (2010) Population genetic structure of the round stingray Urobatis halleri (Elasmobranchii: Rajiformes) in southern California and the Gulf of California. J Fish Biol 77:329–340
Portnoy DS, McDowell JR, McCandless CT, Musick JA, Graves JE (2009) Effective size closely approximates the census size in the heavily exploited western Atlantic population of the sandbar shark, Carcharhinus plumbeus. Conserv Genet 10:1697–1705
Portnoy DS, McDowell JR, Heist EJ, Musick JA, Graves JE (2010) World phylogeography and male-mediated gene flow in the sandbar shark, Carcharhinus plumbeus. Mol Ecol 19:1994–2010
Portnoy DS, Hollenbeck CM, Johnston JS, Casman HM, Gold JR (2014a) Parthenogenesis in a whitetip reef shark Triaenodon obesus involves a reduction in ploidy. J Fish Biol 85:502–508
Portnoy DS, Hollenbeck CM, Belcher CN, Driggers WB III, Frazier BS, Gelsleichter J, Grubbs RD, Gold JR (2014b) Contemporary population structure and post-glacial genetic demography in a migratory marine species, the blacknose shark, Carcharhinus acronotus. Mol Ecol 23:5480–5495
Portnoy DS, Puritz JB, Hollenbeck CM, Gelsleichter J, Chapman D, Gold JR (2015) Selection and sex-biased dispersal in a coastal shark: the influence of philopatry on adaptive variation. Mol Ecol 24:5877–5885
Portnoy DS, Hollenbeck CM, Bethe DM, Frazier BS, Gelsleichter J, Gold JR (2016) Population structure, gene flow, and historical demography of a small coastal shark (Carcharhinus isodon) in US waters of the Western Atlantic Ocean. ICES J Mar Sci 73:2322–2332
Puckett EE (2017) Variability in total project and per sample genotyping costs under varying study designs including with microsatellites or SNPs to answer conservation genetic questions. Conserv Genet Resour 9:289–304
Quintanilha S, Gómez A, Mariño-Ramírez C, Sorzano C, Bessudo S, Soler G, Bernal JE, Caballero S (2015) Conservation genetics of the scalloped hammerhead shark in the Pacific coast od Colombia. J Hered 106:448–458
Ramakrishnan U, Storz JF, Taylor BL, Lande R (2004) Estimation of genetically effective breeding numbers using a rejection algorithm approach. Mol Ecol 13:3283–3292
Richards VP, Suzuki H, Stanhope MJ, Shivji MS (2013) Characterization of the heart transcriptome of the shark (Carcharodon carcharias). BMC Genom 4:697
Rivers MC, Brummitt NA, Lughadha EN, Meagher TR (2014) Do species conservation assessments capture genetic diversity? Global Ecol Evol 2:81–87
Rocha L, Bernal MA, Gaither MR, Alfaro ME (2013) Massively parallel DNA sequencing: the new frontier in biogeography. Front Biogeogr 5:67–77
Rosa RS, Castro ALF, Furtado M, Monzini J, Grubbs RD (2006) Ginglymostoma cirratum. The IUCN Red List of Threatened Species 2006: e.T60223A12325895. http://dx.doi.org/10.2305/IUCN.UK.2006.RLTS.T60223A12325895.en. Acessed 14 April 2016
Ryman N, Palm S (2006) POWSIM: a computer for assessing statistical power when testing for genetic differentiation. Mol Ecol Resour 6:600–602
Ryman N, Utter F, Laikre L (1995) Protection of intraspecific biodiversity of exploited fishes. Rev Fish Biol Fish 5:417–446
Sandoval-Castillo J, Beheregaray LB (2015) Metapopulation structure informs conservation management in a heavily exploited coastal shark (Mustelus henlei). Mar Ecol Prog Ser 533:191–203
Sandoval-Castillo J, Rocha-Olivares A (2011) Deep mitochondrial divergence in Baja California populations of an aquilopelagic elasmobranch: the golden cownose ray. J Hered 102:269–274
Sandoval-Castillo J, Rocha-Olivares A, Villavicencio-Garayzar C, Balart E (2004) Cryptic isolation of Gulf of California shovelnose guitarfish evidence by mitochondrial DNA. Mar Biol 145:983–988
Schluessel V, Broderick D, Collin SP, Ovenden JR (2010) Evidence for extensive population structure in the white-spotted eagle ray within the Indo-Pacific inferred from mitochondrial DNA gene sequences. J Zool 281:46–55
Schmidt JV, Schmidt CL, Ozer F, Ernst RE, Feldheim KA, Ashley MV, Levine M (2009) Low genetic differentiation across three major ocean populations of the whale shark, Rhincodon typus. PLoS ONE 4:e4988. https://doi.org/10.1371/journal.pone.0004988
Schrey AW, Heist EJ (2003) Microsatellite analysis of population structure in the shortfin mako (Isurus oxyrinchus). Can J Fish Aquat Sci 60:670–675
Schultz JK, Feldheim KA, Gruber SH, Ashley MV, McGovern TM, Bowen BW (2008) Global phylogeography and seascape genetics of the lemon sharks (genus Negaprion). Mol Ecol 17:5336–5348
Schwartz MK, Luikart G, Waples RS (2007) Genetic monitoring as a promising tool for conservation and management. Trends Ecol Evol 22:25–33
Seitz JC, Poulakis GR (2006) anthropogenic effects on the smalltooth sawfish (Pristis pectinata) in the United States. Mar Pollut Bull 52:1533–1540
Sellas AB, Bassos-Hull K, Pérez-Jiménez JC, Angulo-Valdés JA, Bernal MA, Hueter RE (2015) Population structure and seasonal migration of the spotted eagle ray, Aetobatus narinari. J Hered 106:266–275
Simpfendorfer CA, Heupel MR, White WT, Dulvy NK (2011) The importance of research and public opinion to conservation management of sharks and rays: a synthesis. Mar Freshw Res 62:518–527
Sodré D, Rodrigues-Filho LFS, Souza RFC, Rêgo PS, Schneider H, Sampaio I, Vallinoto (2012) Inclusion of South American samples reveals new population structuring of the blacktip shark (Carcharhinus limbatus) in the western Atlantic. Genet Mol Biol 4:752–760
Solé-Cava AM, Vooren CM, Levy JA (1983) Isozymic differentiation of two sibling species of Squatina (Condrichthyes) in southern Brazil. Comp Biochem Physiol 75:355–358
Spaet JLY, Jabado RW, Henderson AC, Moore ABM, Berumen ML (2015) Population genetics of four heavily exploited shark species around the Arabian Peninsula. Ecol Evol 5:2317–2332
Stapley J, Reger J, Feulner PGD, Smadja C, Galindo J, Ekblom R, Bennison C, Ball AD, Beckerman AP, Slate J (2010) Adaptation genomics: the next generation. Trends Ecol Evol 25:705–712
Stevens JD, Bonfil R, Dulvy NK, Walker PA (2000) The effects of fishing on sharks, rays, and chimaeras (Chondrichthyans), and the implications for marine ecosystems. ICES J Mar Sci 57:476–494
Stow A, Zenger K, Briscoe D, Gillings M, Peddemors V, Otway N, Harcourt R (2006) Isolation and genetic diversity of endangered grey nurse shark (Carcharias taurus) populations. Biol Lett 2:308–311
Suárez-Moo J, Rocha-Olivares A, Zapata-Pérez O, Quiroz-Moreno A, Sánchez-Teyer F (2013) High genetic connectivity in the Atlantic sharpnose shark, Rhizoprionodon terraenovae, from the southeast Gulf of Mexico inferred from AFLP fingerprinting. Fish Res 147:338–343
Taguchi M, King JR, Wetklo M, Withler RE, Yokawa (2015) Population genetic structure and demographic history of Pacific blue sharks (Prionace glauca) inferred from mitochondrial DNA analysis. Mar Freshw Res 66:267–275
Tavares W, Rodrigues-Filho LFS, Sodré D, Souza RFC, Schneider H, Sampaio I (2013) Multiple substitutions and reduced genetic variability in sharks. Biochem Syst Ecol 49:21–29
Tiffin P, Ross-Ibarra (2014) Advances and limits of using population genetics to understand local adaptation. Trends Ecol Evol 29:673–680
Tillet BJ, Meekan MG, Broderick D, Field IC, Cliff G, Ovenden JR (2012a) Pleistocene isolation, secondary introgression and restricted contemporary gene flow in the pig-eye shark, Carcharhinus amboinensis across northern Australia. Conserv Genet 13:99–115
Tillet BJ, Meekan MG, Field IC, Thorburn DC, Ovenden JR (2012b) Evidence fro reproductive philopatry in the bull shark Carcharhinus leucas. J Fish Biol 80:2140–2158
Tolotti MT, Filmalter JD, Bach P, Travassos P, Seret B, Dagorn L (2015) Banning is not enough: the complexities of oceanic shark management by tuna regional fisheries management organizations. Glob Ecol Evol 4:1–7
Uzans AJ, Lucas Z, McLeod BA, Frasier TR (2015) Small Ne of the isolated and unmanaged horse population on Sable Island. J Hered 106:660–665
Veríssimo A, McDowell JR, Graves JE (2010) Global population structure of the spiny dogfish Squalus acanthias, a temperate shark with an antitropical distribution. Mol Ecol 19:1651–1662
Veríssimo A, McDowell JR, Graves JE (2011) Population structure of a deep-water squaloid shark, the Portuguese dogfish (Centroscymnus coelolepis). ICES J Mar Sci 68:555–563
Veríssimo A, McDowell JR, Graves JE (2012) Genetic population structure and connectivity in a commercially exploited and wide-ranging deepwater shark, the leafscale gulper (Centrophorus squamosus). Mar Freshw Res 63:505–512
Veríssimo A, Sampaio Í, McDowell JR, Alexandrino P, Mucientes G, Queiroz N, da Silva C, Jones CS, Noble LR (2017) World without borders—genetic population structure of a highly migratory marine predator, the blue shark (Prionace glauca). Ecol Evol 7:4768–4781
Vignaud T, Clua E, Mourier J, Maynard J, Planes S (2013) Microsatellite analyses of blacktip reef sharks (Carcharhinus melanopterus) in a fragmented environment show structured clusters. PLoS ONE 8:e61067. https://doi.org/10.1371/journal.pone.0061067
Vignaud TM, Mourier J, Maynard J, Leblois R, Azquez-Juárez RV, Ramírez-Macías D, Pierce SJ, Rowat D, Berumen ML, Beeravolu C, Baksay S, Planes S (2014a) Genetic structure of populations of whale sharks among ocean basins and evidence for their historic rise and recent decline. Mol Ecol 23:2590–2601
Vignaud TM, Maynard J, Leblois R, Meekan M, Spaet JLY, Clua E, Neglia V, Planes S (2014b) Blacktip reef sharks, Carcharhinus melanopterus, have high genetic structure and varying demographic histories in their Indo-Pacific range. Mol Ecol 23:5193–5207
Vooren CM, Lessa RP, Klippel S (2005) Biologia e status de conservação da viola Rhinobatos horkelli. In: Vooren CM, Klippel S (eds) Ações para a conservação de tubarões e raias no sul do Brasil. Igaré, Porto Alegre, pp 32–56
Walker TI, Taylor BL, Russell JH, Cottier JP (1998) The phenomenon of apparent change of growth rate in gummy shark (Mustelus antarcticus) harvested off southern Australia. Fish Res 39:139–163
Waples R (1995) Evolutionary significant units and the conservation of biological diversity under the endangered’s species. Am Fish Soc Symp 17:8–27
Waples RS, Gaggiotti O (2006) What is a population? An empirical evaluation of some genetic methods for identifying the number of gene pools and their degree of connectivity. Mol Ecol 15:1419–1439
Wasko AP, Martins C, Oliveira C, Foresti F (2003) Non-destructive genetic sampling in fish. An improved DNA extraction from fish fins and scales. Hereditas 138:161–165
Watts PC, Buley KR, Sanderson S, Boardman W, Ciofi C, Gibson R (2006) Parthenogenesis in Komodo dragons. Nature 444:1021–1022
Weigmann S (2016) Annotated checklist of the living sharks, batoids and chimaeras (Chondrichthyes) of the world, with focus on biogeographical diversity. J Fish Biol 88:837–1037
Whitney NM, Robbins WD, Schultz JK, Bowen BW, Holland KN (2012) Oceanic dispersal in a sedentary reef shark (Triaenodon obesus): genetic evidence for extensive connectivity without a pelagic larval stage. J Biogeogr 39:1144–1156
Willette DA, Allendorf FW, Barber PH et al (2014) So, you want to use next-generation sequencing in marine systems? Insight from the Pan-Pacific Advanced Studies Institute. Bull Mar Sci 90:79–122
Willoughby JR, Sundaram M, Wijayawardena BK, Kimble SJA, Yanzhu J, Fernandez NB, Antonides JD, Lamb MC, Marra NJ, DeWoody JA (2015) The reduction of genetic diversity in threatened vertebrates and new recommendations regarding IUCN conservation rankings. Biol Cons 191:495–503
Worm B, Davis B, Kettemer L, Ward-Paige CA, Chapman D, Heithaus MR, Kessel S, Gruber S (2013) Global catches, exploitation rates, and rebuilding options for sharks. Mar Policy 40:194–204
Zeh JA, Zeh DW (2003) Toward a new sexual selection paradigm: polyandry, conflict and incompatibility. Ethology 109:929–950
Acknowledgements
The authors thank three anonymous reviewers and the editors of Conservation Genetics for their valuable suggestions for strengthening the manuscript during peer review. This work was supported by the São Paulo Research Foundation (FAPESP #2009/54660-6 and #2013/08675-7) and by a grant from the Brazilian National Council for Scientific and Technological Development to OBFG (CNPq #308430-8).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Domingues, R.R., Hilsdorf, A.W.S. & Gadig, O.B.F. The importance of considering genetic diversity in shark and ray conservation policies. Conserv Genet 19, 501–525 (2018). https://doi.org/10.1007/s10592-017-1038-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-017-1038-3