Abstract
Many shark species are at risk of overexploitation due to their high economic value, slow maturation, and low recruitment compared to most teleosts. However, there is insufficient knowledge about population structure at different spatial scales necessary to optimise fisheries models. We used single-nucleotide polymorphisms (SNPs) obtained through complexity-reduction genome sequencing to quantify the population structure of two highly mobile and commercially fished shark species: bronze whalers (Carcharhinus brachyurus) and dusky sharks (C. obscurus). We applied a comprehensive approach to test several population-structure hypotheses and signal consistency across methods and marker type. We found that C. obscurus was panmictic across Australia and Indonesia and across the Indian Ocean to South Africa based on neutral loci, whereas for C. brachyurus, the westernmost Australian samples appeared to be separate from the rest. The southernmost east Australian samples indicated some difference from the rest of Australia and New Zealand based on candidate loci for C. brachyurus, and potentially also C. obscurus; however, the lack of a reference genome makes the interpretation difficult. Despite similar patterns in both species, subtle and potentially important structure differences emphasise the importance of studying each target species independently rather than assuming similar patterns from closely related species with similar dispersal abilities, as well as considering different marker types in future studies. We found evidence of connectivity across the regions sampled, suggesting that the cumulative effects of regional fisheries and the potential for cross-jurisdictional fishery assessments and management should be considered for Australian, Indonesian, and New Zealand populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marine species generally exhibit high gene flow and connectivity across local, regional, and global scales. However, dispersal distances can vary substantially among species, making a unified approach to assessing connectivity and population structure difficult. Dispersal scales range from virtually zero in some sessile species, to thousands of kilometres in highly mobile elasmobranchs (e.g., Bonfil et al. 2005; Gore et al. 2008; Lea et al. 2015; Skomal et al. 2009; Veríssimo et al. 2017). This enables gene flow across vastly different spatial scales, which can lead to varied genetic population sub-structure. Outcomes can indicate: (1) panmixia, a common phenomenon in highly mobile species with large dispersal distances and home ranges (Quintela et al. 2014; Roy et al. 2014; but see Sellas et al. 2015), and is characterised by an absence of mating restrictions among individuals; (2) discrete populations and strong population sub-structure that are often found in species with limited dispersal capacity and narrow home ranges (Barbosa et al. 2013; Benestan et al. 2015), or (3) isolation-by-distance, where genetic similarity decreases as the distance between populations increases. Isolation-by-distance has been observed across all dispersal ranges (Wright et al. 2015) and is often established after the colonisation of new habitats (e.g., Junge et al. 2011). Although isolation-by-distance is a common pattern, distance alone sometimes only partially predicts gene flow and population connectivity (e.g., Ashe et al. 2015).
Quantifying population structure and connectivity is necessary to define units of management and conservation. Species particularly at risk of overexploitation and extinction are high priorities for management and conservation, and these species are often highly valued, slow to mature, and have sporadic or low recruitment (Dulvy et al. 2014; Field et al. 2009; Frisk et al. 2005). Such characteristics are typical of many elasmobranch species that are fished commercially and recreationally (Walker 1998), making it necessary to monitor their harvests. However, ongoing depletion of sharks makes their sustainable harvest a scientific and management challenge (Dulvy et al. 2017).
The sustainable harvest of sharks relies on effective management that requires quantifying population structure and connectivity. For example, a lack of population sub-structure calls for management of the species across its entire distribution, whereas highly structured populations might require different management strategies depending on specific conditions. However, assessing population connectivity and gene flow for many sharks and other elasmobranchs is difficult due to their large dispersal capacity and complex life histories, coupled with difficulties in observing and sampling individuals.
Several approaches to increase the power to detect population structure have recently become available for species that lack reference genomic information—so-called ‘non-model’ species. It is now feasible and affordable to sequence thousands of markers for hundreds or thousands of individuals. This is achieved via complexity reduction of the genome prior to sequencing through genotyping by sequencing (GBS; Deschamps et al. 2012), restriction-site associated DNA (RAD; Miller et al. 2007) sequencing, or diversity arrays technology (DArT; Wenzl et al. 2004).
Classic population-genetics studies based only on putatively neutral loci to quantify gene flow and genetic drift, and hence, assess population structure, are relevant only within a well-defined theoretical framework. Therefore, analysing loci under selection (hereafter ‘candidate loci’) might provide additional spatial resolution (Hess et al. 2013; Milano et al. 2014), or reveal associations with environmental variables (e.g., Gaggiotti et al. 2009; Lamichhaney et al. 2012) or life-history traits (Hemmer-Hansen et al. 2014). Such an approach is potentially a more effective way to integrate genetics into fisheries management (Dudgeon et al. 2012; Ovenden et al. 2015). There are now many marine studies using candidate loci to quantify population structure and divergence (e.g., Case et al. 2005; Guo et al. 2015; Milano et al. 2014). This helps to (1) determine weak population structure for highly mobile species (something traditional markers alone struggle to elucidate) and (2) detect markers potentially under selection more efficiently using outlier detection to reduce the background noise that can swamp weak population structure (Pérez-Figueroa et al. 2010). The patterns of neutral and ‘adaptive’ population structure (using candidate loci) might either be different—with one or the other marker type showing higher sub-structure (e.g., Hess et al. 2013; Limborg et al. 2012)—or the same, where both marker types show the same sub-structure (e.g., Moore et al. 2014) or a lack thereof.
We applied a comprehensive approach using a large number of neutral and sets of candidate, genome-wide single-nucleotide polymorphism (SNP) loci following the population-genomics approach outlined in Luikart et al. (2003) to quantify the population structure of two commercially fished elasmobranch species: the bronze whaler Carcharhinus brachyurus (Günther, 1870), and the dusky shark C. obscurus (Lesueur, 1818). At a global scale, both species are of conservation concern—C. brachyurus is currently listed as Near Threatened in the International Union for Conservation of Nature’s (IUCN) Red List of Threatened Species (Duffy and Gordon 2003b), and C. obscurus is listed as Vulnerable (Musick et al. 2009). Their life histories are characterised by low growth rates, late sexual maturity (C. brachyurus: 13–20 years, C. obscurus 17–32 years), and low fecundity (Drew et al. 2017; Dudley et al. 2005; Geraghty et al. 2013; McAuley et al. 2007; Romine et al. 2009; Walter and Ebert 1991), resulting in low recovery potential following depletion (Smith et al. 1998; Rogers et al. 2013a; Bradshaw et al. 2018). Both species are globally distributed: C. brachyurus occurs patchily in temperate regions (Duffy and Gordon 2003b), whereas C. obscurus occurs in tropical to warm temperate regions. However, both species are sympatric in temperate regions, where they can be misidentified as each other (Jones 2008).
Both species are targeted commercially and recreationally in many parts of their distribution (Bradshaw et al. 2018), as well as being taken with other more productive shark species in mixed-species fisheries. Landings from fisheries are often grouped together with other Carcharhinus species as ‘whaler sharks’, which means that population declines of an individual species are unlikely to be detected (Duffy and Gordon 2003b). Within Australasia and Indonesia, the major fisheries for C. brachyurus are in South Australia and upper North Island of New Zealand (Bradshaw et al. 2018), whereas C. obscurus are caught mainly in Australia: in southern Western Australia, New South Wales (Duffy and Gordon 2003b; Macbeth et al. 2009; Rogers et al. 2013b; Simpfendorfer et al. 1999) and to a lesser extent, South Australia; C. obscurus is rare in New Zealand (Roberts et al. 2015).
Demographic fishery models (e.g., Bradshaw et al. 2013, 2018; McAuley et al. 2007; Otway et al. 2004; Smart et al. 2017) generally require high-quality catch and effort data (Field et al. 2009; Walker 1998) and reliable information on demography and connectivity. Previous molecular genetic studies based on mitochondrial DNA showed that Australian C. obscurus has one of the lowest nucleotide diversities of any large, globally distributed shark (Benavides et al. 2011b), and they are weakly differentiated genetically between the east and west of Australia (Geraghty et al. 2014). Accordingly, they are currently considered and reported as distinct eastern and western biological stocks in Australia (Braccini et al. 2016). However, no investigation of genetic diversity or population structure of C. obscurus has yet included southern Australia populations that potentially connect the east and west via gene flow, and studies on C. brachyurus across Australia are lacking altogether. In each species, we therefore (1) tested whether the populations are panmictic, discrete, or show evidence of isolation-by-distance within Australasia and Indonesia and (2) compared the patterns shown by neutral versus candidate loci.
Materials and methods
Sampling, DNA extraction, and sequencing
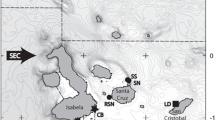
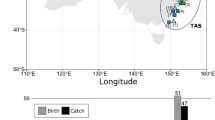
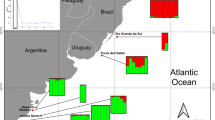
Tissue samples from C. brachyurus (n = 106 from seven areas; 43 males, 53 females, 10 unknown) and C. obscurus (n = 207 from 14 areas; 90 males, 110 females, 7 unknown) were collected from Australia, Indonesia, New Zealand, and South Africa between 2001 and 2013 by a combination of fisheries-dependent and -independent surveys, and from recreational fisheries, and shark bite-mitigation programs (Table 1). All samples were stored in 96% ethanol. We combined samples collected from each defined area here as ‘sample sets’ (Fig. 1; Electronic Supplementary Materials 1 and 2).
Sampling locations for Carcharhinus brachyurus (circles) and C. obscurus (closed upward triangles). Latitudes and longitudes are indicated. (top) Sampling locations across Australia and Indonesia, with insert for locations in the two gulfs in South Australia, and (bottom) overview sampling locations across the Indian and Southern Oceans for both species. WA Western Australia, NT Northern Territory, SA South Australia, QLD Queensland, NSW New South Wales, VIC Victoria
We extracted DNA using the DNeasy Blood & Tissue kit (Qiagen) following the manufacturer’s instructions. We used DArTseq™ technology (Cruz et al. 2013; Kilian et al. 2012) for single-nucleotide polymorphism (SNP) genotyping done by Diversity Arrays Technology (DArT, Canberra). We processed DNA samples in digestion/ligation reactions as described by Kilian et al. (2012), except we replaced the single PstI-compatible adaptor with two different adaptors corresponding to the PstI and SphI restriction-enzyme overhangs for a double digest. Details of the method are in the Electronic Supplementary Material 1 and Donnellan et al. (2015). We originally analysed libraries separately to check for batch effects, and only in the final step completely pooled all libraries and called the final SNPs. We also randomly divided samples from different sample sets across libraries and sequencing lanes to overcome batch effects, as suggested by Leigh et al. (2018).
Data filtering, species identification, and genetic diversity
We detected 50,608 SNPs among 313 individuals across both species with the DArT analytical pipeline. Due to strong morphological similarities of the two species, we first confirmed species identity with a discriminant analysis of principal components of all samples (DAPC) (Jombart et al. 2010) implemented in the R package ADEGENET 1.4-2 (Jombart 2008) R library prior to subsequent analyses. We ran steps 0–2 of our data-filtering pipeline (see Electronic Supplementary Material 3). We assigned individuals to the correct species based on unambiguous clustering results (Electronic Supplementary Material 4).
We then launched the filtering pipeline again, from the beginning with the corrected species IDs (Electronic Supplementary Material 3) including all data, and then, from step 2 onwards, we ran the pipeline independently for each species to retain the most intra-species SNPs.
The data-filtering pipeline comprised 10 quality filters, one outlier individual removal step (step 11), and one linked locus removal step (step 12; summary in Electronic Supplementary Material 3; details in Electronic Supplementary Material 1, section 1.2.). This produced two data sets (Table 2): brachyurus_ALL_loci: comprising 3766 loci from 106 C. brachyurus in Australia and New Zealand, and obscurus_ALL_loci: comprising 8886 loci from 207 C. obscurus from 13 Australian and Indonesian and one South African sample set (Table 2).
We used GENODIVE (Meirmans and Van Tienderen 2004) and ADEGENET 1.4-2 (Jombart 2008) to calculate heterozygosities, and allele frequencies, including distributions of minor allele frequency. We did all subsequent analyses for both species focussed on only samples from Australia, Indonesia and New Zealand, unless otherwise indicated.
Linkage disequilibrium, Hardy–Weinberg equilibrium, and outlier loci
We used the GENETICS R library (Warnes and Leisch 2006) to test for locus pairs that are potentially linked, i.e., those loci in linkage disequilibrium. We tested each species’ data set separately for pairwise linkage disequilibrium between genetic markers using the function LD. After sequential removal (see Electronic Supplementary Material 1), this resulted in two new unlinked data sets, one for each species: brachyurus_unlinked_loci with 520 loci and obscurus_unlinked_loci with 2828 loci (Table 2). We used these two unlinked data sets for all subsequent analyses (unless otherwise indicated). We also did the same analyses for the full data sets (i.e., unlinked plus linked loci) and did not observe differences in the resulting population-structure patterns. We present the results from the full data sets in the Electronic Supplementary Materials 5–8. We used GENEPOP 4.2.1 (Rousset 2008) to test for deviations from Hardy–Weinberg equilibrium (Electronic Supplementary Material 1).
We used two different FST outlier detection methods, LOSITAN (Antao et al. 2008) and BAYESCAN (Foll and Gaggiotti 2008) to identify candidate loci that are potentially under positive selection, and loci that could be considered neutral, because the interpretation of outlier loci should be done cautiously (Narum and Hess 2011). Both methods can be sensitive to low sample sizes, so we included only sample sets that had > 8 individuals (i.e., 4/7 for C. brachyurus, and 11/13 for C. obscurus) to achieve a representative number of alleles. We included only those loci safely identified as ‘neutral’ in the neutral data sets and excluded all other loci, including those identified as under balancing selection. We subsequently used the BLAST tool to search for sequence matches of the candidate loci that revealed some level of sub-structure.
Population structure
We separated all data sets into neutral and candidate loci, resulting in two data sets per species. We implemented simulations following the strategy of Krück et al. (2013) using POWSIM (Ryman and Palm 2006) to calculate the statistical power to detect genetic differentiation (details in Electronic Supplementary Material 1). We calculated global FST for the neutral loci for each species separately using GENEPOP and applied the implemented exact G test.
We investigated the population structure by two main approaches: (1) classic FST and population differentiation estimates and (2) cluster analysis. To estimate the differentiation, we calculated pairwise FST (Weir and Cockerham 1984) and ran exact G tests implemented in GENEPOP (details in Electronic Supplementary Material 1), correcting for multiple tests using a sequential Bonferroni method (Rice 1989).
The cluster analyses comprised of: (1) a principal component analysis using ADEGENET and (2) STRUCTURE 2.3.4 (Pritchard et al. 2000). In ADEGENET (Jombart 2008; Jombart and Ahmed 2011), we calculated principal components using the glPca function with default arguments and produced plots using the ggplot2 package (Wickham 2009). We did the same analysis including the South African sample set for C. obscurus. In STRUCTURE, we used the admixture model, assuming correlated allele frequencies (Falush et al. 2003; Pritchard et al. 2000) with a burn-in of 300,000 and 700,000 iterations, and the ‘locprior’ option (Hubisz et al. 2009), with 20 replicate runs for each k representing the number of sampled sites. We determined the optimal number of clusters following Pritchard et al. (2000) and Evanno et al. (2005) using STRUCTURE HARVESTER (Earl and vonHoldt 2012) and visualised the results in CLUMPAK (Kopelman et al. 2015).
Subsequently, we tested if there was any structure related to region, distance or sex (detailed in Electronic Supplementary Material 1).
Results
Carcharhinus brachyurus
Descriptive statistics
The expected and observed heterozygosities for the neutral loci ranged from 0.23 to 0.29 and 0.21 to 0.37, respectively; observed heterozygosity was higher than expected in the westernmost Australian sample set (Table 1). The median observed heterozygosity was 0.259 when all and 0.235 when the westernmost samples were excluded. When we analysed all loci together, little heterozygosity was evident (Fig. 2b). However, after removing potentially linked loci, average heterozygosity increased (Fig. 2d), and we used the unlinked loci for all subsequent analyses. Removing the confounding effect of genetic linkage produced the following data set: brachyurus_unlinked with 520 loci (an 86% reduction) (Table 2). All sample sets were in Hardy–Weinberg equilibrium in both the neutral and the candidate loci data sets. We detected 13 candidate loci with LOSITAN (Table 2), whereas BAYESCAN detected none, irrespective of the prior settings (Electronic Supplementary Material 5). The BLAST search did not reveal any explanatory sequence matches (data not shown).
Observed heterozygosity distribution within species for both species. Shown as densities (= percentage of loci with a given heterozygosity) for a, c C. obscurus; b, d C. brachyurus, including a, b) all SNPs, and c, d only unlinked SNPs. Inset: observed versus expected heterozygosity. The red line represents the y = x relation between expected heterozygosity and observed heterozygosity, i.e., the expected 1:1 relationship
Population structure
Global FST for the neutral loci considering all sample sets was 0.0078 (CI 95% from 10,000 permutations = [0.0043, 0.0113], p value from 1000 permutations < 0.05), and 0.0034 (CI 95% from 10,000 permutations = [5e−04, 0.0064], p value from 1000 permutations > 0.05) if the westernmost sample set (3-SW_WEST) was removed. We found no evidence for population differentiation between pairs of sample sets for neutral loci, whereas the candidate loci showed differentiation in 6 out of 21 pairwise comparisons, involving sample sets within and between regions; half of them involved the Australian EAST sample set (9-EAST_South; Electronic Supplementary Material 6). For each of those 13 candidate loci individually, 0 to approximately 30% of all pairwise comparisons revealed differences. Pairwise FST ranged from − 0.014 to 0.294 in the candidate locus data set, and from − 0.014 to 0.112 in the neutral one (Electronic Supplementary Material 6); in the latter, all comparisons involving the WEST sample set (3-WEST_SW1) had the highest FST. Power simulations concluded that we have sufficient power to detect differences (for details, see Electronic Supplementary Material 1).
For the neutral loci, STRUCTURE revealed evidence for two population clusters—WEST (3-WEST_SW1) and westernmost SOUTH (5-SOUTH_GAB) formed one cluster, and all other individuals irrespective of sample sets were assigned to the second (Fig. 3b); a cluster possibility for K = 3 did not reveal any further conclusive sub-structure (Fig. 3c), although it was the most likely number when analysing the unlinked neutral loci only (see Electronic Supplementary Material 7). The same individuals also clustered in the principal components analysis (Fig. 3d). The STRUCTURE analysis of the candidate loci revealed two clusters, separating the EAST (9-EAST_South) from all other (Fig. 3a), whereas the principal components analysis revealed no clear separation (Electronic Supplementary Material 7). We detected no differentiation signal based on distance or sex, but found a differentiation between the EAST and SOUTH regions based on the candidate loci (details in the Electronic Supplementary Material 1).

(modified from Duffy and Gordon 2003a) with sample sets indicated: 3–W_SW1, 5–S_GAB, 6–S_SG, 7–S_GSV, 8–S_Vic, 9–E_South, and 10–E_NZ (see Table 1 for abbreviation definitions)
Cluster analyses for Carcharhinus brachyurus. Shown for unlinked loci using STRUCTURE (a–c) for a candidate loci, and b, c neutral loci [shown for multiple clusters as recommended Meirmans (2015)] for K = 2 (b) and K = 3 clusters (c) and (d) principal component analysis based on 2842 neutral loci from the entire set; e distribution of C. brachyurus
Carcharhinus obscurus
Descriptive statistics
The expected and observed heterozygosity for the neutral loci ranged from 0.19 to 0.20 and 0.16 to 0.20, respectively. The median observed heterozygosity was 0.186. The general pattern of heterozygosity was similar to C. brachyurus (i.e., low but increasing after removing linked loci; Fig. 2a, c). Removing the confounding effect of genetic linkage produced the following data set: obscurus_unlinked with 2828 loci, resulting in a 68% reduction of loci (Table 2). All sample sets were in Hardy–Weinberg equilibrium based on the neutral loci. For candidate loci, eight out of 13 sample sets did not conform to Hardy–Weinberg equilibrium. LOSITAN revealed 24 loci and BAYESCAN between two and six loci when using 20 or 10 as prior odds for the neutral model, respectively (Electronic Supplementary Material 5). None of these loci was detected by both programs. However, when testing for candidate loci using all loci (including the linked ones), 10 loci were identified by both programs (see Table 2).
Population structure
Global FST for the neutral loci was 0.0014 (CI 95% from 10,000 permutations = [7e−04, 0.0021], p value from 1000 permutations > 0.05). None of the pairwise population comparisons showed evidence for differentiation in the neutral data set (pairwise FST range: − 0.008 to 0.011; Electronic Supplementary Material 8). For candidate loci, 38 pairwise comparisons out of 78 showed differentiation (FST range: − 0.032 to 0.210; Electronic Supplementary Material 8). In the 10 candidate loci identified by both LOSITAN and BAYESCAN, all pairwise comparisons involving one EAST sample set (11-EAST_Mid) were differentiated, and approximately half of the comparisons involving one SOUTH sample set (6-SOUTH_GSV) were as well, totalling 18 out of 78. Power simulations concluded that we have sufficient power to detect differences (for details, see Electronic Supplementary Material 1).
There was neither population sub-structure within the Australia–Indonesia region, nor between that region and South Africa, based on FST and principal component analysis using neutral or candidate loci (Electronic Supplementary Materials 7, 8, and 9; Fig. 4c). No clustering was evident with STRUCTURE for neutral loci (Electronic Supplementary Material 7). For the candidate loci, STRUCTURE revealed K = 2, but most sample sets showed mixed ancestry without any geographic pattern (Electronic Supplementary Material 7). For the 10 loci jointly detected by LOSITAN and BAYESCAN, there was evidence for K = 2 or 3 (Fig. 4a, b). One EAST (11-EAST_Mid), one SOUTH (7-SOUTH_SG) sample set and potentially some individuals from neighbouring SOUTH (6-SOUTH_GSV) were separated from the remaining sample sets. We detected no differentiation signal based on distance or sex.

(modified from Musick et al. 2009) with the sample sets indicated: 1–W_North, 2–W_Perth, 3–W_SW1, 4–W_SW2, 5–S_GAB, 6–S_SG, 7–S_GSV, 9–E_South, 11–E_Mid, 12–E_North, 13–E_NORF, 14–N_IDN, and 15–N_NT) (see Table 1 for abbreviation definitions)
Cluster analyses for C. obscurus using STRUCTURE. Shown for unlinked loci using STRUCTURE (a, b) consensus candidate loci (10) identified by both, LOSITAN and BAYESCAN, programs for K = 2–3 clusters [shown for multiple clusters as recommended Meirmans 2015)]; c principal component analysis plot based on 7969 neutral loci from the entire set, d distribution range of C. obscurus
Discussion
Using a comprehensive approach involving 3000–9000 genome-wide SNPs, neutral and candidate loci, and several different analytical methods, we found no evidence for population sub-structure across Australia and Indonesia for the dusky shark (Carcharhinus obscurus), but some support for a separation of the westernmost Australian samples for the bronze whaler (C. brachyurus) from the rest of Australia and New Zealand based on neutral loci only (albeit based on few samples). For C. obscurus, we therefore propose panmixia within Australia and Indonesia as well as across the Indian Ocean, thus rejecting the hypotheses of discrete populations and isolation-by-distance. Although candidate loci generally revealed higher differentiation among populations, we could not identify any consistent population sub-structure in C. obscurus. However, C. brachyurus from the east of Australia showed weak differentiation from the rest of Australia and New Zealand using candidate loci, although a clustering could not be conclusively confirmed using principal components analysis. Interestingly, some individuals from the same EAST location showed some admixture as well in C. obscurus using STRUCTURE, but we could not confirm a distinct clustering.
Recent studies on the movement and residency of both species in Australia using acoustic and satellite tags have demonstrated some connectivity between South Australia, Victoria, and Western Australia, with several individuals moving between these states (Huveneers et al. 2014; Rogers et al. 2013a, b). Some C. obscurus have travelled up to 2736 km between South Australia and Western Australia (Rogers et al. 2013b), and similar large-scale movements have also been recorded in the Northwest Atlantic (Kohler et al. 1998) and off South Africa (Dudley et al. 2005; Hussey et al. 2009); however, there is no direct evidence for movements across the Indian Ocean. Large-scale movements of C. brachyurus have also been documented in South Africa (Cliff and Dudley 1992).
Neutral versus candidate loci
The lack of population genetic structure for C. brachyurus based on mitochondrial DNA between Australia and New Zealand (Benavides et al. 2011a) suggests that these sharks have substantial gene flow. Our results based on neutral loci confirm panmixia between the two countries, but also show some support for a separation of the western Australia samples, albeit based on a low sample size from the west. This is confirmed by a significant global FST when including all sample sets but a non-significant result when removing the westernmost samples (3-SW_WEST). We checked those and the other western samples (5-SOUTH_GAB) with respect to DNA quality, sequence quality as well as amount of missing data and we have no reason to conclude that their separation from the rest of the Australian and New Zealand sample sets is due to a sample problem. In contrast, the candidate loci suggest that the east Australian samples (9-EAST_South) differed from all other sample sets, including New Zealand (10-EAST_NZL), the latter being more similar to southern and western Australia samples. However, the reason for this difference requires understanding the function of these candidate loci based on future genome mapping.
Although C. brachyurus from EAST Australia was larger than those from SOUTH Australia, a previous study found genetic structure across populations around the globe (Benavides et al. 2011a, b), suggesting that it is unlikely that large C. brachyurus in the EAST originated from areas not included in our study. In addition, studies of the movement of C. brachyurus using acoustic tags and the Integrated Marine Observing System (IMOS) animal-tracking network (Hoenner et al. 2018) and pop-up satellite archival tags (Huveneers and Drew, unpublished data) show some limited, large-scale movements of C. brachyurus between the WEST, the SOUTH, and the south-eastern SOUTH (Victoria). However, these movements occurred across life stages, and predominantly between the SOUTH and the WEST, with no records of sharks mixing between the SOUTH and the EAST. For C. obscurus, neutral and candidate loci revealed the same lack of clear population sub-structure. However, the candidate loci showed more variation that was not consistent with geography.
Different gene flow patterns in C. obscurus
Previous molecular studies of C. obscurus have revealed differing patterns of population structure between eastern and western Australia, from genetic homogeneity using mitochondrial DNA and microsatellites (Benavides et al. 2011b; Ovenden et al. 2009) to weak population structure based on mitochondrial DNA (Geraghty et al. 2014). These have also suggested, but not demonstrated conclusively, limited or no dispersal between Indonesia and Australia (Geraghty et al. 2014; Ovenden et al. 2009). Our results confirm genetic connectivity across the Sunda Shelf Barrier (Timor Trench), a widely recognised biogeographic barrier (Dudgeon et al. 2012), and high gene flow around Australia, including Bass Strait, another biogeographic border for many marine species (e.g., Barnes et al. 2015).
We found no population sub-structure for C. obscurus across Australia when analysing almost 3000 neutral loci across differentiation and clustering approaches and irrespective of sex, which was confirmed by a non-significant global FST. Our finding contrasts with mitochondrial DNA evidence that demonstrated weak differentiation between the east and west of Australia (Geraghty et al. 2014). We observed no evidence of differences between northern Australia (NORTH), situated midway between the east and west coast, and the eastern and western samples. This incongruence between mtDNA and nuclear DNA variation could be due to female philopatry [reviewed in Chapman et al. (2015)]—if females mate and/or give birth close to their natal sites, this can lead to a pattern of local or regional population structuring with gene flow mediated more by males. However, we found no evidence for male-biased dispersal in C. obscurus. For candidate loci, we detected many pairwise population differences among males from different samples (29%) compared to the female–female comparisons (5%). This implies that male-biased dispersal is not the driver for population structuring in the Australian–Indonesian C. obscurus. Another possibility is that differentiation of mtDNA is driven by selection based on environmental factors. Under this scenario, mtDNA might be an indicator for demographic connectivity, but this requires confirmation.
Increasing the number of molecular markers is expected to increase the chances of detecting population structure, whereby new technologies enable the sequencing of thousands of loci across the genome to resolve suggestions of weak population structure. However, here we found that even using between 2000 and 8000 neutral loci, there was no sub-structure within Australia and Indonesia for C. obscurus, and there were no obvious barriers to gene flow between these areas and South Africa.
Linkage disequilibrium
Although violation of linkage-disequilibrium assumptions can introduce bias when interpreting population genetic data, few genomic studies have incorporated statistical steps and strict testing in their analysis to identify and account for linked markers and the effect of removing such potentially linked markers on their results. We detected many linked loci and reduced our data sets accordingly until only one member per linkage group was retained. Low heterozygosity and minor allele frequencies could mimic linkage between markers due to the prevalence of the most common allele across markers. This is not necessarily a signal of linkage, but the two possibilities might be indistinguishable. When we removed ‘linked’ loci, there was a reduction of loci with low heterozygosity in both species (most prominent in C. obscurus). Removing potentially linked loci resulted in a large reduction of the number of loci, especially for C. brachyurus, and it would therefore be worthwhile to test in a future mapping study if pairs showing evidence of linkage disequilibrium are in fact physically linked.
Management implications
Our results provide evidence for high genetic connectivity across Australia and Indonesia for C. obscurus, and Australia and New Zealand for C. brachyurus. The practical implications of this are that fisheries management needs to be considered across jurisdictional boundaries by taking catches from other states and countries into account in fisheries models (e.g., Bradshaw et al. 2018). For C. brachyurus, we found some genetic separation between Western Australia (WEST) and South Australia (SOUTH), with the Great Australian Bight (5-SOUTH_GAB) samples appearing to be connected more to Western Australia rather than to South Australia; however, larger samples sizes are still required to measure the full extent of connectivity between those regions. From the perspective of fisheries management, C. brachyurus from the major fisheries regions in South Australia and New Zealand are panmictic and, therefore, represent a single management stock. Interestingly, C. brachyurus showed an overall higher genetic diversity compared to C. obscurus across the sampling areas we investigated. More samples from the western and eastern regions for C. brachyurus would contribute to an understanding of any underlying biological relevance, but unfortunately, few samples were available from those areas.
For C. obscurus, our confirmation of a seemingly shared stock across the Timor Trench also has important implications for management. The Australian C. obscurus samples seem to form some part of the Indonesian fishery and vice versa, which is not unexpected given the reported overlap in areas of the shark fisheries (Blaber et al. 2009). This could affect the management of the cross-jurisdictional stock of mature dusky sharks that is protected from fishing in Western Australia. The largest Australian catches of C. obscurus occur in Western Australia, where management relies on effort control, spatial closures, gear restrictions, and size limits designed to direct fishing-related mortality on the youngest age classes and protect large juveniles and adults. No such size limits exist in other regions (e.g., New South Wales and South Australia), where large juveniles and/or adults can be caught (Geraghty et al. 2015; Rogers et al. 2013a). However, to restrict what was considered unsustainable catches during 2007, New South Wales implemented a total allowable catch (Macbeth et al. 2009). Given that our findings and various tagging studies indicate that stocks are shared across jurisdictional boundaries (Huveneers et al. 2014; Rogers et al. 2013a, b), catches in Indonesia, eastern Australia, and South Australia would affect the stock of C. obscurus in Western Australia, and vice versa, if these populations are also demographically linked. Fisheries catching C. obscurus should therefore consider the cumulative effects of each fishery and the potential need for cross-jurisdiction fishery assessments and management.
We had access to only seven Indonesian samples, of which the exact origin was uncertain. However, the fish market from where the samples came and the size of boats landing sharks suggest that sharks were most likely caught in Indonesia. The samples also came from the same region/market as described in Geraghty et al. (2014), and described as Indonesian as well. More samples from Indonesia would help clarify structure between Indonesia and Australia.
Using genetics to manage shark fisheries is plagued with vagueness in the eyes of managers (Waples et al. 2008). Managers are primarily interested in determining whether there are demographically independent units that can be modelled separately; however, it is still unknown how much migration is required to produce demographic coupling. Populations linked by a 10% permanent immigration rate between them can be considered ‘demographically coupled’ (sensu Hastings 1993); regardless, immigration rates lower than this could still mean that populations are demographically ‘linked’. However, genetic methods struggle to distinguish the degree of connectivity requiring separate stock management if migration rates are high (Waples et al. 2008), because demographically linked populations might still be connected genetically over many generations (Ovenden 2013). Nevertheless, genetic assessments are the gold standard for determining stock structure, because they allow enduring patterns of connectivity to be captured by measuring it across multiple generations (Dichmont et al. 2012; Flood et al. 2014); however, they are most powerful in conjunction with other stock-differentiation methods.
A recent study from Bailleul et al. (2018) on blue sharks proposed the concept of a “population grey zone” for similarly mobile species and scenarios, explaining that such widespread genetic interdependence could in fact camouflage a wide range of demographic situations. They point to a common problem highly relevant to managers, which refers to the inferences that can be made about genetic and demographic independence if differentiation is found, and how on the other hand, a lack thereof (panmixia) does not necessarily imply demographic unity. In such cases, the authors suggest a conservative and global management strategy until demographic homogeneity is either confirmed or can be ruled out.
Both dusky sharks and bronze whalers, seem largely panmictic across the areas we sampled. However, recreational catch records, conventional tagging, and preliminary acoustic data (Huveneers et al. 2014; Huveneers, unpublished data; Izzo et al. 2016; Rogers et al. 2013b) suggest some residency and site fidelity [defined by Chapman et al. (2015)]. Accurately scaled stock assessments are therefore needed to ensure that the effect of each local fishery is considered (Bradshaw et al. 2018). We also recommend more studies examining residency, site fidelity, and potential philopatry to determine the requirement for and effectiveness of protective legislation (e.g., spatial closures) intended to prevent local extirpation (Chapman et al. 2015). A logical next step would also be to collect more samples from difficult to access, but nevertheless important areas, like the western parts of Australia for C. brachyurus, and to include a seascape genomic approach with a balanced sample design integrating genomic and oceanographic data.
References
Antao T, Lopes A, Lopes RJ, Beja-Pereira A, Luikart G (2008) LOSITAN: a workbench to detect molecular adaptation based on a Fst-outlier method. BMC Bioinform 9:323
Ashe JL, Feldheim KA, Fields AT, Reyier EA, Brooks EJ, O’Connell MT, Skomal G, Gruber SH, Chapman CC (2015) Local population structure and context-dependent isolation by distance in a large coastal shark. Mar Ecol Prog Ser 520:203–216. https://doi.org/10.3354/meps11069
Bailleul D, Mackenzie A, Sacchi O, Poisson F, Bierne N, Arnaud-Haond S (2018) Large-scale genetic panmixia in the blue shark (Prionace glauca): a single worldwide population, or a genetic lag-time effect of the “grey zone” of differentiation? Evol Appl 11:614–630
Barbosa SS, Klanten SO, Puritz JB, Toonen RJ, Byrne M (2013) Very fine-scale population genetic structure of sympatric asterinid sea stars with benthic and pelagic larvae: influence of mating system and dispersal potential. Biol J Linn Soc 108:821–833. https://doi.org/10.1111/bij.12006
Barnes TC, Junge C, Myers SA, Taylor MD, Rogers PJ, Ferguson GJ, Lieschke JA, Donnellan SC, Gillanders BM (2015) Population structure in a wide-ranging coastal teleost (Argyrosomus japonicus, Sciaenidae) reflects marine biogeography across southern Australia. Mar Freshw Res 67:8. https://doi.org/10.1071/MF15044
Benavides MT, Feldheim KA, Duffy CA, Wintner S, Braccini M, Boomer J, Huveneers C, Rogers P, Mangel JC, Alfaro-Shigueto J, Cartamil DP, Chapman DD (2011a) Phylogeography of the copper shark (Carcharhinus brachyurus) in the southern hemisphere: implications for the conservation of a coastal apex predator. Mar Freshw Res 62:861–869. https://doi.org/10.1071/mf10236
Benavides MT, Horn RL, Feldheim KA, Shivji MS, Clarke SC, Wintner S, Natanson L, Braccini M, Boomer JJ, Gulak SJB, Chapman DD (2011b) Global phylogeography of the dusky shark Carcharhinus obscurus: implications for fisheries management and monitoring the shark fin trade. Endanger Species Res 14:13–22. https://doi.org/10.3354/esr00337
Benestan L, Gosselin T, Perrier C, Sainte-Marie B, Rochette R, Bernatchez L (2015) RAD-genotyping reveals fine-scale genetic structuring and provides powerful population assignment in a widely distributed marine species; the American lobster (Homarus americanus). Mol Ecol 24:3299–3315. https://doi.org/10.1111/mec.13245
Blaber SJM, Dichmont CM, White W, Buckworth R, Sadiyah L, Iskandar B, Nurhakim S, Pillans R, Andamari R, Dharmadi Fahmi (2009) Elasmobranchs in southern Indonesian fisheries: the fisheries, the status of the stocks and management options. Rev Fish Biol Fisher 19:367–391
Bonfil R, Meyer M, Scholl MC, Johnson R, O’Brien S, Oosthuizen H, Swanson S, Kotze D, Paterson M (2005) Transoceanic migration, spatial dynamics, and population linkages of white sharks. Science 310:100–103. https://doi.org/10.1126/science.1114898
Braccini M, Johnson G, Rogers P, Hansen S, Peddemors V (2016) Dusky Whaler (Carcharhinus obscurus). In: Stewardson C, Andrews J, Ashby C, Haddon M, Hartmann K, Hone P, Horvat P, Mayfield S, Roelofs A, Sainsbury K, Saunders T, Stewart J, Stobutzki I, Wise B (eds) 2016, Status of Australian fish stocks reports 2016. FRDC, Canberra
Bradshaw CJA, Field IC, McMahon CR, Johnson GJ, Meekan MG, Buckworth RC (2013) More analytical bite in estimating targets for shark harvest. Mar Ecol Prog Ser 488:221–232. https://doi.org/10.3354/meps10375
Bradshaw CJA, Prowse TAA, Drew M, Gillanders BM, Donnellan SC, Huveneers C (2018) Predicting sustainable shark harvests when stock assessments are lacking. ICES J Mar Sci 75:1591–1601. https://doi.org/10.1093/icesjms/fsy031
Case RAJ, Hutchinson WF, Hauser L, Van Oosterhout C, Carvalho GR (2005) Macro- and micro-geographic variation in pantophysin (PanI) allele frequencies in NE Atlantic cod Gadus morhua. Mar Ecol Prog Ser 301:267–278. https://doi.org/10.3354/meps301267
Chapman DD, Feldheim KA, Papastamatiou YP, Hueter RE (2015) There and back again: a review of residency and return migrations in sharks, with implications for population structure and management. Annu Rev Mar Sci 7:547–570. https://doi.org/10.1146/annurev-marine-010814-015730
Cliff G, Dudley SFJ (1992) Sharks caught in the protective gill nets off Natal, South Africa. 6. The copper shark Carcharhinus brachyurus (Günther). S Afr J Marine Sci 12:663–674. https://doi.org/10.2989/02577619209504731
Cruz VMV, Kilian A, Dierig DA (2013) Development of DArT marker platforms and genetic diversity assessment of the US collection of the new oilseed crop Lesquerella and related species. PLoS One 8:e64062. https://doi.org/10.1371/journal.pone.0064062
Deschamps S, Llaca V, May GD (2012) Genotyping-by-sequencing in plants. Biology 1:460–483
Dichmont CM, Ovenden JR, Berry O, Welch DJ, Buckworth RC (2012) Scoping current and future genetic tools, their limitations and their applications for wild fisheries management. CSIRO, Brisbane, p 129
Donnellan SC, Foster R, Junge C, Huveneers C, Rogers P, Kilian A, Bertozzi T (2015) Fiddling with the proof: the magpie fiddler ray is a colour pattern variant of the common southern fiddler ray (Rhinobatidae: Trygonorrhina). Zootaxa 3981:367–384
Drew M, Rogers P, Huveneers C (2017) Slow life-history traits of a neritic predator, the bronze whaler (Carcharhinus brachyurus). Mar Freshw Res 68:461–472
Dudgeon CL, Blower DC, Broderick D, Giles JL, Holmes BJ, Kashiwagi T, Krück NC, Morgan AT, Tillett BJ, Ovenden JR (2012) A review of the application of molecular genetics for fisheries management and conservation of sharks and rays. J Fish Biol 80:1789–1843. https://doi.org/10.1111/j.1095-8649.2012.03265.x
Dudley SFJ, Cliff G, Zungu MP, Smale MJ (2005) Sharks caught in the protective gill nets off KwaZulu-Natal, South Africa. 10. The dusky shark Carcharhinus obscurus (Lesueur 1818). Afr J Mar Sci 27:107–127. https://doi.org/10.2989/18142320509504072
Duffy C, Gordon I (2003a) Carcharhinus brachyurus (SSG Australia & Oceania Regional Workshop, March 2003). The IUCN Red List of Threatened Species version 20143 online: http://iucnredlist.org/
Duffy C, Gordon I (2003b) Carcharhinus brachyurus. In: Cavanagh Rachel D, Kyne PM, Fowler SL, Musick JA, Bennett MB (ed) The conservation status of Australian chondrichthyans: report of the IUCN Shark Specialist Group Australia and Oceania Regional Red List Workshop. The University of Queensland, School of Biomedical Sciences, Brisbane, Australia, vol x + 170, pp 106–109
Dulvy NK, Fowler SL, Musick JA, Cavanagh RD, Kyne PM, Harrison LR, Carlson JK, Davidson LNK, Fordham SV, Francis MP, Pollock CM, Simpfendorfer CA, Burgess GH, Carpenter KE, Compagno LJV, Ebert DA, Gibson C, Heupel MR, Livingstone SR, Sanciangco JC, Stevens JD, Valenti S, White WT (2014) Extinction risk and conservation of the world’s sharks and rays. eLife 3:e00590
Dulvy NK, Simpfendorfer CA, Davidson LNK, Fordham SV, Bräutigam A, Sant G, Welch DJ (2017) Challenges and priorities in shark and ray conservation. Curr Biol 27:R565–R572
Earl D, vonHoldt B (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Field IC, Meekan MG, Buckworth RC, Bradshaw CJ (2009) Susceptibility of sharks, rays and chimaeras to global extinction. Adv Mar Biol 56:275–363
Flood MSI, Andrews J, Ashby C, Begg G, Fletcher R, Gardner C, Georgeson L, Hansen S, Hartmann K, Hone P, Horvat P, Maloney L, McDonald B, Moore A, Roelofs A, Sainsbury K, Saunders T, Smith T, Stewardson C, Stewart J, Wise B (eds) (2014) Status of key Australian fish stocks reports 2014. FRDC, Canberra
Foll M, Gaggiotti O (2008) A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180:977–993. https://doi.org/10.1534/genetics.108.092221
Frisk MG, Miller TJ, Dulvy NK (2005) Life histories and vulnerability to exploitation of elasmobranchs: inferences from elasticity, perturbation and phylogenetic analyses. J Northwest Atl Fish Sci 35:27–45
Gaggiotti OE, Bekkevold D, Jørgensen HBH, Foll M, Carvalho GR, Andre C, Ruzzante DE (2009) Disentangling the effects of evolutionary, demographic, and environmental factors influeing genetic structure of natural populations: Atlantic herring as a case study. Evolution 63:2939–2951. https://doi.org/10.1111/j.1558-5646.2009.00779.x
Geraghty PT, Macbeth WG, Harry AV, Bell JE, Yerman MN, Williamson JE (2013) Age and growth parameters for three heavily exploited shark species off temperate eastern Australia. ICES J Mar Sci 71:559–573. https://doi.org/10.1093/icesjms/fst164
Geraghty PT, Williamson JE, Macbeth WG, Blower DC, Morgan JAT, Johnson G, Ovenden JR, Gillings MR (2014) Genetic structure and diversity of two highly vulnerable carcharhinids in Australian waters. Endanger Species Res 24:45–60. https://doi.org/10.3354/esr00580
Geraghty P, Macbeth W, Williamson J (2015) Aspects of the reproductive biology of dusky, spinner and sandbar sharks (Family Carcharhinidae) from the Tasman Sea. Mar Freshw Res 67:513–525
Gore MA, Rowat D, Hall J, Gell FR, Ormond RF (2008) Transatlantic migration and deep mid-ocean diving by basking shark. Biol Lett 4:395–398
Guo BC, DeFaveri J, Sotelo G, Nair A, Merilä J (2015) Population genomic evidence for adaptive differentiation in Baltic Sea three-spined sticklebacks. BMC Biol 13:19. https://doi.org/10.1186/s12915-015-0130-8
Hastings A (1993) Complex interactions between dispersal and dynamics: lessons from coupled logistic equations. Ecology 74:1362–1372
Hemmer-Hansen J, Therkildsen NO, Meldrup D, Nielsen EE (2014) Conserving marine biodiversity: insights from life-history trait candidate genes in Atlantic cod (Gadus morhua). Conserv Genet 15:213–228. https://doi.org/10.1007/s10592-013-0532-5
Hess JE, Campbell NR, Close DA, Docker MF, Narum SR (2013) Population genomics of Pacific lamprey: adaptive variation in a highly dispersive species. Mol Ecol 22:2898–2916. https://doi.org/10.1111/mec.12150
Hoenner X, Huveneers C, Steckenreuter A, Simpfendorfer C, Tattersall K, Jaine F, Atkins N, Babcock R, Brodie S, Burgess J (2018) Australia’s continental-scale acoustic tracking database and its automated quality control process. Sci Data 5:170206
Hubisz MJ, Falush D, Stephens M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour 9:1322–1332
Hussey NE, McCarthy ID, Dudley SF, Mann BQ (2009) Nursery grounds, movement patterns and growth rates of dusky sharks, Carcharhinus obscurus: a long-term tag and release study in South African waters. Mar Fresh Res 60:571–583
Huveneers C, Rogers PJ, Drew M (2014) Monitoring shark species of conservation concern within the Adelaide metropolitan and Gulf St Vincent regions. Final report to the Adelaide and Mount Lofty ranges natural resources management board. SARDI research report series No. 754, SARDI Aquatic Sciences, Adelaide, Australia
Izzo C, Huveneers C, Drew M, Bradshaw CJ, Donnellan SC, Gillanders BM (2016) Vertebral chemistry demonstrates movement and population structure of bronze whaler. Mar Ecol Prog Ser 556:195–207
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Jombart T, Ahmed I (2011) adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics 27:3070–3071. https://doi.org/10.1093/bioinformatics/btr521
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics 11:94. https://doi.org/10.1186/1471-2156-11-94
Jones GK (2008) Review of the fishery status for whaler sharks (Carcharhinus spp.) in South Australian and adjacent waters. FRDC Project 2004/067. SARDI Aquatic Sciences
Junge C, Vøllestad LA, Barson NJ, Haugen TO, Otero J, Sætre G-P, Primmer CR (2011) Strong gene flow and lack of stable population structure in the face of rapid adaptation to local temperature in a spring-spawning salmonid, the European grayling (Thymallus thymallus). Heredity 106:460–471. https://doi.org/10.1038/hdy.2010.160
Kilian A, Wenzl P, Huttner E, Carling J, Xia L, Blois H, Caig V, Heller-Uszynska K, Jaccoud D, Hopper C, Aschenbrenner-Kilian M, Evers M, Peng K, Cayla C, Hok P, Uszynski G (2012) Diversity arrays technology: a generic genome profiling technology on open platforms. In: Pompanon F, Bonin A (eds) Data production and analysis in population genomics, vol 888. Methods in Molecular Biology. Humana Press, New York, pp 67–89. https://doi.org/10.1007/978-1-61779-870-2_5
Kohler NE, Casey JG, Turner PA (1998) NMFS cooperative shark tagging program, 1962-93: an atlas of shark tag and recapture data. Mar Fish Rev 60:1–87
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15:1179–1191. https://doi.org/10.1111/1755-0998.12387
Krück NC, Innes DI, Ovenden JR (2013) New SNPs for population genetic analysis reveal possible cryptic speciation of eastern Australian sea mullet (Mugil cephalus). Mol Ecol Resour 13:715–725. https://doi.org/10.1111/1755-0998.12112
Lamichhaney S, Martinez Barrio A, Rafati N, Sundström G, Rubin C-J, Gilbert ER, Berglund J, Wetterbom A, Laikre L, Webster MT, Grabherr M, Ryman N, Andersson L (2012) Population-scale sequencing reveals genetic differentiation due to local adaptation in Atlantic herring. Proc Natl Acad Sci USA 109:19345–19350. https://doi.org/10.1073/pnas.1216128109
Lea JSE, Wetherbee BM, Queiroz N, Brunie N, Aming C, Sousa LL, Muceintes GR, Humphries NE, Harvey GM, Sims DW, Shivji MS (2015) Repeated, long-distance migrations by a philopatric predator targeting highly contrasting ecosystems. Sci Rep 5:11202. https://doi.org/10.1038/srep11202
Leigh DM, Lischer HEL, Grossen C, Keller LF (2018) Batch effects in a multiyear sequencing study: false biological trends due to changes in read lengths. Mol Eco Resour 18(4):778–788
Limborg MT et al (2012) Environmental selection on transcriptome-derived SNPs in a high gene flow marine fish, the Atlantic herring (Clupea harengus). Mol Ecol 21:3686–3703. https://doi.org/10.1111/j.1365-294X.2012.05639.x
Luikart G, England PR, Tallmon D, Jordan S, Taberlet P (2003) The power and promise of population genomics: from genotyping to genome typing. Nat Rev Genet 4:981–994
Macbeth WG, Geraghty PT, Peddemors VM, Gray CA (2009) Observer-based study of targeted commercial fishing for large shark species in waters off northern New South Wales. Industry & Investment NSW—Fisheries Final Report Series 114
McAuley RB, Simpfendorfer CA, Hall NG (2007) A method for evaluating the impacts of fishing mortality and stochastic influences on the demography of two long-lived shark stocks. ICES J Mar Sci 64:1710–1722. https://doi.org/10.1093/icesjms/fsm146
Meirmans PG (2015) Seven common mistakes in population genetics and how to avoid them. Mol Ecol 24:3223–3231. https://doi.org/10.1111/mec.13243
Meirmans PG, Van Tienderen PH (2004) Genotype and genodive: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794. https://doi.org/10.1111/j.1471-8286.2004.00770.x
Milano I, Babbucci M, Cariani A, Atanassova M, Bekkevold D, Carvalho GR, Espiñeira M, Florentino F, Garofalo G, Geffen AJ, Hansen JH, Helyar SJ, Nielsen EE, Ogden R, Patarnello T, Stagioni M, FishPopTrace Consortium, Tinti F, Bargelloni L (2014) Outlier SNP markers reveal fine-scale genetic structuring across European hake populations (Merluccius merluccius). Mol Ecol 23:118–135
Miller MR, Dunham JP, Amores A, Cresko WA, Johnson EA (2007) Rapid and cost-effective polymorphism identification and genotyping using restriction site associated DNA (RAD) markers. Genome Res 17:240–248. https://doi.org/10.1101/gr.5681207
Moore J-S, Bourret V, Dionne M, Bradbury I, O’Reilly P, Kent M, Chaput G, Bernatchez L (2014) Conservation genomics of anadromous Atlantic salmon across its North American range: outlier loci identify the same patterns of population structure as neutral loci. Mol Ecol 23:5680–5697. https://doi.org/10.1111/mec.12972
Musick JA, Grubbs RD, Baum J, Cortés E (2009) Carcharhinus obscurus. The IUCN red list of threatened species 2009:e.T3852A10127245. https://doi.org/10.2305/IUCN.UK.2009-2.RLTS.T3852A10127245.en
Narum SR, Hess JE (2011) Comparison of FST outlier tests for SNP loci under selection. Mol Ecol Resour 11:184–194
Otway NM, Bradshaw CJA, Harcourt RG (2004) Estimating the rate of quasi-extinction of the Australian grey nurse shark (Carcharias taurus) population using deterministic age- and stage-classified models. Biol Conserv 119:341–350. https://doi.org/10.1016/j.biocon.2003.11.017
Ovenden JR (2013) Crinkles in connectivity: combining genetics and other types of biological data to estimate movement and interbreeding between populations. Mar Freshw Res 64:201–207. https://doi.org/10.1071/mf12314
Ovenden J, Kashiwagi T, Broderick D, Giles J, Salini J (2009) The extent of population genetic subdivision differs among four co-distributed shark species in the Indo-Australian archipelago. BMC Evol Biol 9:1–15. https://doi.org/10.1186/1471-2148-9-40
Ovenden JR, Berry O, Welch DJ, Buckworth RC, Dichmont CM (2015) Ocean’s eleven: a critical evaluation of the role of population, evolutionary and molecular genetics in the management of wild fisheries. Fish Fish 16:125–159. https://doi.org/10.1111/faf.12052
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Pérez-Figueroa A, García-Pereira MJ, Saura M, Rolán-Alvarez E, Caballero A (2010) Comparing three different methods to detect selective loci using dominant markers. J Evol Biol 23:2267–2276. https://doi.org/10.1111/j.1420-9101.2010.02093.x
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Quintela M, Skaug HJ, Ølen N, Haug T, Seliussen BB, Solvang HK, Pampoulie C, Kanda N, Pastene LA, Glover KA (2014) Investigating population genetic structure in a highly mobile marine organism: the minke whale Balaenoptera acutorostrata in the North East Atlantic. PLoS One 9:e108640. https://doi.org/10.1371/journal.pone.0108640
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Roberts CD, Stewart AL, Struthers CD (eds) (2015) The fishes of New Zealand. In four volumes. Te Papa Press, Wellington
Rogers PJ, Huveneers C, Goldsworthy SD, Cheung WWL, Jones GK, Mitchell JG, Seuront L (2013a) Population metrics and movement of two sympatric carcharhinids: a comparison of the vulnerability of pelagic sharks of the southern Australian gulfs and shelves. Mar Freshw Res 64:20–30. https://doi.org/10.1071/MF11234
Rogers PJ, Huveneers C, Goldsworthy SD, Mitchell JG, Seuront L (2013b) Broad-scale movements and pelagic habitat of the dusky shark Carcharhinus obscurus off southern Australia determined using pop-up satellite archival tags. Fish Oceanogr 22:102–112. https://doi.org/10.1111/fog.12009
Romine J, Musick J, Burgess G (2009) Demographic analyses of the dusky shark, Carcharhinus obscurus, in the Northwest Atlantic incorporating hooking mortality estimates and revised reproductive parameters. Environ Biol Fish 84:277–289. https://doi.org/10.1007/s10641-008-9435-6
Rousset F (2008) genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Resour 8:103–106. https://doi.org/10.1111/j.1471-8286.2007.01931.x
Roy D, Hardie DC, Treble MA, Reist JD, Ruzzante DE (2014) Evidence supporting panmixia in Greenland halibut (Reinhardtius hippoglossoides) in the Northwest Atlantic. Can J Fish Aquat Sci 71:763–774. https://doi.org/10.1139/cjfas-2014-0004
Ryman N, Palm S (2006) POWSIM: a computer program for assessing statistical power when testing for genetic differentiation. Mol Ecol Notes 6:600–602
Sellas AB, Bassos-Hull K, Pérez-Jiménez JC, Angulo-Valdés JA, Bernal MA, Hueter RE (2015) Population structure and seasonal migration of the spotted eagle ray, Aetobatus narinari. J Hered 106:266–275. https://doi.org/10.1093/jhered/esv011
Simpfendorfer CA, McAuley R, Chidlow J, Lenanton R, Hall N, Bastow T (1999) Biology and stock assessment of western Australia’s commercially important shark species. Final Report. FRDC, ISBN 0 7309 8448 6, Canberra, Australia
Skomal GB, Zeeman SI, Chisholm JH, Summers EL, Walsh HJ, McMahon KW, Thorrold SR (2009) Transequatorial migrations by basking sharks in the Western Atlantic Ocean. Curr Biol 19:1019–1022. https://doi.org/10.1016/j.cub.2009.04.019
Smart JJ, Chin A, Tobin AJ, White WT, Kumasi B, Simpfendorfer CA (2017) Stochastic demographic analyses of the silvertip shark (Carcharhinus albimarginatus) and the common blacktip shark (Carcharhinus limbatus) from the Indo-Pacific. Fish Res 191:95–107
Smith SE, Au DW, Show C (1998) Intrinsic rebound potentials of 26 species of Pacific sharks. Mar Freshwater Res 49:663–678. https://doi.org/10.1071/MF97135
Veríssimo A, Sampaio Í, McDowell JR, Alexandrino P, Mucientes G, Queiroz N, da Silva C, Jones CS, Noble LR (2017) World without borders—genetic population structure of a highly migratory marine predator, the blue shark (Prionace glauca). Ecol Evol 7:4768–4781
Walker TI (1998) Can shark resources be harvested sustainably? A question revisited with a review of shark fisheries. Mar Freshw Res 49:553–572
Walter JP, Ebert DA (1991) Preliminary estimates of age of the bronze whaler Carcharhinus brachyurus (Chondrichthyes: Carcharhinidae) from southern Africa, with a review of some life history parameters. S Afr J Mar Sci 10:37–44. https://doi.org/10.2989/02577619109504617
Waples RS, Punt AE, Cope JM (2008) Integrating genetic data into management of marine resources: how can we do it better? Fish Fish 9:423–449. https://doi.org/10.1111/j.1467-2979.2008.00303.x
Warnes G, Leisch F (2006) Genetics: population genetics. R Package, version 1.2.1
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wenzl P, Carling J, Kudrna D, Jaccoud D, Huttner E, Kleinhofs A, Kilian A (2004) Diversity arrays technology (DArT) for whole-genome profiling of barley. Proc Natl Acad Sci USA 101:9915–9920. https://doi.org/10.1073/pnas.0401076101
Wickham H (2009) ggplot2: elegant graphics for data analysis. Springer Science & Business Media, Berlin
Wright D, Bishop JM, Matthee CA, von der Heyden S (2015) Genetic isolation by distance reveals restricted dispersal across a range of life histories: implications for biodiversity conservation planning across highly variable marine environments. Divers Distrib 21:698–710. https://doi.org/10.1111/ddi.12302
Acknowledgements
We thank W White, M Scott, S Tindale, J Ovenden, and Auckland museum for providing samples, R Keane for the distance estimations, A Loi for help in the lab, and JCA Pistevos for the shark artwork in Figs. 3 and 4. The project received funding from the Australian Research Council (LP120100652), the Neiser Foundation, and the Nature Foundation of South Australia. BMG and CJAB were supported by ARC Future Fellowships FT100100767 and FT110100306, respectively). Unpublished acoustic tracking data was sourced from the Acoustic Tracking Database (animaltracking.aodn.org.au) of the Integrated Marine Observing System (IMOS; www.imos.org.au)—IMOS is a national collaborative research infrastructure supported by Australian Government.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
No sharks were sacrificed for this study. All tissue samples were approximately 2 × 2 cm and were taken from pectoral fins (unless otherwise indicated). All samples were obtained legally (see Electronic Supplementary Material 10).
Conflict of interest
All authors declare that they have no conflict of interest and all applicable international, national and/or institutional guidelines for the care and use of animals were followed.
Additional information
Responsible Editor: J. Raeymaekers.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Reviewed by P. Hablützel and an undisclosed expert.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Junge, C., Donnellan, S.C., Huveneers, C. et al. Comparative population genomics confirms little population structure in two commercially targeted carcharhinid sharks. Mar Biol 166, 16 (2019). https://doi.org/10.1007/s00227-018-3454-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-018-3454-4