Abstract
l-Threonine is an important branched-chain amino acid and could be applied in feed, drugs, and food. In this study, l-threonine production in an l-threonine-producing Escherichia coli strain TWF001 was significantly increased by overexpressing the gene cluster phaCAB from Ralstonia eutropha. TWF001/pFW01-phaCAB could produce 96.4-g/L l-threonine in 3-L fermenter and 133.5-g/L l-threonine in 10-L fermenter, respectively. In addition, TWF001/pFW01-phaCAB produced 216% more acetyl-CoA, 43% more malate, and much less acetate than the vector control TWF001/pFW01, and meanwhile, TWF001/pFW01-phaCAB produced poly-3-hydroxybutyrate, while TWF001/pFW01 did not. Transcription analysis showed that the key genes in the l-threonine biosynthetic pathway were up-regulated, the genes relevant to the acetate formation were down-regulated, and the gene acs encoding the enzyme which converts acetate to acetyl-CoA was up-regulated. The results suggested that overexpression of the gene cluster phaCAB in E. coli benefits the enhancement of l-threonine production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
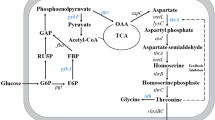
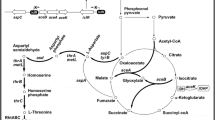
l-Threonine is an essential branched-chain amino acid, and has been widely used in food, feed, and pharmaceutical industries [7, 26]. l-Threonine production could be raised to a much higher level using Escherichia coli than using Corynebacterium glutamicum [6, 19, 42]. In E. coli, l-threonine biosynthesis involves six steps, starting from oxaloacetate (Fig. 1) [6, 45]. Oxaloacetate can be made from acetyl-CoA through tricarboxylic acid (TCA) cycle, and converted to L-aspartate. L-aspartate can be phosphorylated by aspartate kinases I, II, or III which are encoded by thrA, metL, or lysC, respectively [8]. Aspartate kinases I and II can also catalyze aspartyl semialdehyde to form homoserine. Homoserine can be phosphorylated by homoserine kinase encoded by thrB. Then, threonine synthase encoded by thrC converts homoserine phosphate into l-threonine. Intracellular l-threonine can be secreted by l-threonine export proteins RhtA, RhtB, or RhtC [15, 42, 43]. The strategy of overexpression of the thrABC cluster is frequently used to increase l-threonine production in E. coli [6, 16, 17, 28, 32, 45, 46].
Oxaloacetate is the initial metabolite of all l-aspartate family amino acids [6, 45]. It is a component of TCA cycle, which starts from reaction of acetyl-CoA and oxaloacetate [41]. It could be made from pyruvate or phosphoenol pyruvate through the anaplerotic reaction or from malate [2, 41, 46]. However, the metabolic flux towards pyruvate and acetyl-CoA can be easily directed to acetate by enzymes encoded by poxB and pta-ackA, respectively (Fig. 1) [10, 37]. Therefore, obtaining high amount of oxaloacetate and maintaining low concentration of acetate is important for l-threonine production in E. coli [45]. Acetate is one of the major by-products in l-threonine-producing E. coli, and its overproduction significantly retards the cell growth. Acetate can be converted into acetyl-CoA by acetyl-CoA synthetase encoded by acs (Fig. 1). The overexpression of acs could significantly improve l-threonine production [18]. Enhancing the glyoxylate shunt to accumulate oxaloacetate [46], increasing acetyl-CoA supply [18], overcoming acetate overflow [42], and improving the generation of cofactors [23] could also improve l-threonine production [42].
Poly-3-hydroxybutyrate (PHB) is a biopolymer consisting of linear chains of (R)-3-hydroxybutyrate units [20, 30, 47]. It can be biosynthesized from acetyl-CoA by enzymes β-ketothiolase, acetoacetyl-CoA reductase, and PHB synthase encoded by phaA, phaB, and phaC, respectively [1, 21]. PHB biosynthesis in E. coli could be improved by engineering the l-threonine bypass [22, 37]. PHB accumulation in E. coli could reduce the level of acetate and improve the production of succinate [12, 13], l-tryptophan [10], or 5-aminolevulinic acid [44]. Overexpression of the gene cluster phaCAB from Ralstonia eutropha in Corynebacterium glutamicum could increase l-glutamate production by 68% and reduce the accumulation of by-products α-ketoglutarate, glutamine, and lactic acid [24]. Overexpression of phaCAB in Corynebacterium crenatum could increase l-arginine production 20.6% [40]. Overexpression of phaCAB in l-isoleucine producing C. glutamicum WM001 produced 44% more l-isoleucine and 30% more acetyl-CoA [29]. Overexpression of phaCAB in E. coli could produce 11.6% more l-tryptophan and led to the up-regulation of the genes in the l-tryptophan operon [10]. Overexpression of phaCAB could also increase succinate in E. coli MG1655 [13] and 5-aminolevulinic acid production in E. coli DH5α [44]. However, increasing l-threonine production in E. coli by overexpression of the gene cluster phaCAB has not been reported.
In this study, the influence of overexpression of the gene cluster phaCAB on l-threonine production in E. coli TWF001 [46] has been investigated. The gene cluster phaCAB from pBHR68 [33] was inserted into the triclosan-resistant plasmid pFW01 [46], and introduced into TWF001. l-Threonine production and the acetate accumulation in TWF001/pFW01-phaCAB were investigated.
Materials and methods
Bacteria and culture conditions
Bacteria and plasmids used in this work are listed in Table 1. E. coli cells were grown at 37 °C in LB medium (5-g/L yeast extract, 10-g/L tryptone, and 10-g/L NaCl) with 200-rpm shaking. Triclosan (1 µM) was added when necessary. l-threonine-producing E. coli strain TWF001 (CCTCC no. M2017730) [46] was originally isolated from soil, and is closely related to E. coli MG1655, based on their 16S rDNA sequences.
DNA preparation and PCR techniques
Restrictions enzymes, T4 DNA ligase, and DNA Ladder were purchased from Sangon (Shanghai, China). PrimeSTAR™ HS DNA polymerase was purchased from TaKaRa (Dalian, China). Plasmid Minipreps Purification System B used for isolating the plasmid DNA from E. coli was from BioDev-Tech (Beijing, China). TIAnamp Bacteria DNA Kit used for isolating the genomic DNA from E. coli was from Tiangen (Beijing, China). EZ-10 Spin Column DNA Gel Extraction Kit used for purifying DNA from agarose gels was purchased from Bio Basic Inc. (Markham, Canada). DNA synthesis and sequencing were performed by Sangon (Shanghai, China).
PCR experiments were performed using Mastercycler from Eppendorf (Hamburg, Germany). The sequences of all primers used in this study are listed in Table 2.
Construction of plasmid pFW01-phaCAB
The gene cluster phaCAB for poly-3-hydroxybutyrate biosynthesis was cloned using pBHR68 [33] as a template and inserted in pFW01, resulting in the plasmid pFW01-phaCAB. Briefly, the phaCAB fragment was cloned with primers phaCAB-F/phaCAB-R, and then digested with EcoRI and HindIII, and pFW01 was digested with EcoRI and HindIII and then ligated together, resulting in the plasmid pFW01-phaCAB. The correct sequence in pFW01-phaCAB was checked by colony PCR, restriction enzyme digestion, and DNA sequencing.
The plasmids pFW01 and pFW01-phaCAB were transformed into E. coli strain TWF001, resulting in TWF001/pFW01 and TWF001/pFW01-phaCAB, respectively.
Quantification of mRNA using real-time PCR (RT-PCR)
RT-PCR was used to quantify mRNA levels of aceA, aceB, aspC, thrA, thrB, thrC, poxB, pta, and ackA in TWF001/pFW01-phaCAB using TWF001/pFW01 as the control. Total RNA was extracted from E. coli cells grown at the late-exponential phase using an RNA extraction kit (Bio Flux, Beijing, China). Residual DNA was removed from the RNA sample by DNase I. The quality and amount of RNA were judged and quantified by electrophoresis. Using random hexamer primers, 500-ng RNA was reversely transcribed into cDNA using a Revert Aid™ First Strand cDNA synthesis kit (Fermentas, Shanghai, China). RT-PCR was performed using an ABI Step One RT-PCR system (Applied Biosystems, San Mateo, CA, USA) with a ChamQ™ Universal SYBR qPCR Master Mix kit (Tiangen, Beijing, China). Primers for detection of various genes are listed in Table 2. The following RT-PCR procedure was used: 1 min at 94 °C, 40 cycles of 10 s at 94 °C, 30 s at 55 °C, and 15 s at 68 °C. The relative abundance of the targeted mRNAs was quantified based on the cycle threshold value, which is defined as the number of cycles required to obtain a fluorescent signal above the background and was calculated according to the published method [27, 31]. To standardize the results, the relative abundance of 16S rRNA was used as an internal standard control. The relative transcriptional levels were calculated by the − ΔΔCt method, as follows: 2^[(Ctnwt − Ct0wt) − (Ctn − Ct0)]. The Ctn and Ct0 are for the target gene and 16S rRNA in E. coli TWF001/pFW01-phaCAB, respectively; while Ctnwt and Ct0wt are for the target gene and 16 s rRNA in E. coli TWF001/pFW01, respectively. All assays were performed in triplicate.
Flask fermentation
The l-threonine-producing E. coli strains were grown on plates containing LB medium (5-g/L yeast extract, 10-g/L tryptone, and 10-g/L NaCl) for 24 h and transferred to a test tube containing 5-mL LB medium. After growing for 4 h, the OD600 of the culture was measured. Then, the culture was transferred to a flask-containing 25-mL LB medium, with the initial OD600 of 0.1. After growing for 4 h, 5-mL culture with the OD600 of 1.5 was transferred to a flask-containing 30-mL fermentation medium I [2-g/L yeast extract, 2-g/L citric acid, 25-g/L (NH4)2SO4, 7.46-g/L KH2PO4, 30-g/L glucose, 2-g/L MgSO4·7 H2O, 5-mg/L FeSO4·7H2O, 5-mg/L MnSO4·4 H2O, and 20-g/L CaCO3, pH 6.8 or pH 7.0] [17, 41, 46] and agitated for 36 h at 200 rpm.
Fed-batch fermentation
For fed-batch fermentation, a 3-L BioFlo310 fermenter (New Brunswick Scientific, USA) was applied. l-Threonine-producing E. coli strains were grown on LB plates for 16 h and transferred to a 250-mL flask-containing 25-mL LB medium. After growing for 4 h at 37 °C and 200 rpm, the culture was transferred to a 500-mL flask-containing 100-mL seed medium I (32.5-g/L glucose, 5-g/L (NH4)2SO4, 15-g/L yeast extract, 9.5-g/L KH2PO4, 24.35-g/L K2HPO4, 1-g/L MgSO4·7H2O, pH 7.0) with the initial OD600 of 0.1. After growing for 4 h at 37 °C, all the seed culture was transferred to the 3-L fermenter with 800-mL fermentation medium II (20-g/L glucose, 3-g/L yeast extract, 2-g/L KH2PO4, 10-g/L (NH4)2SO4, 0.5-g/L MgSO4·7H2O, 5-mg/L FeSO4·7H2O, and 5-mg/L MnSO4·4H2O). The inoculum ratio was 10% (v/v) and the initial glucose concentration was 20 g/L. When glucose concentration in the medium was below 5 g/L, a feeding solution-containing 800 g/L of glucose was supplied into the medium. The incubation temperature was set at 37 °C, and the pH was controlled at 6.8 with NH3·H2O. The dissolved oxygen was kept around 30% by adjusting the agitation speed and the aeration rate.
For larger scale fed-batch fermentation, a stirred 10-L fermenter system (PV13140, Zhenjiang Dongfang Shenggong Equipment Technology Co., Ltd.) was applied. l-Threonine-producing E. coli strains were grown on LB plates for 16 h and transferred to a 500-mL flask-containing 100-mL seed medium II [1.2-g/L sucrose, 10-g/L peptone, 8-g/L yeast extract, 4-g/L (NH4)2SO4, 3-g/L K2HPO4, 0.4-g/L MgSO4, 0.01-g/L FeSO4, 0.01-g/L MnSO4, 5-g/L sodium glutamate, 0.2-g/L methionine, and pH 7.0] [34]. After growing for 4 h at 37 °C and 200 rpm, the culture was transferred to two 2-L flasks, each containing 500-mL seed medium III (30-g/L glucose, 2.86-g/L yeast extract, 5.7-g/L corn syrup, 2.86-g/L K2HPO4, 0.57-g/L MgSO4, 4.29-g/L amino acid mixture, 1-mg/L vitamin B1, 0.1-mg/L ATP, and pH 7.0) [34], and the initial OD600 was adjusted as 0.1. After growing for 4 h at 37 °C, all the culture was transferred to the 10-L fermenter-containing 4-L fermentation medium III (25-g/L glucose, 3.2-g/L corn syrup, 0.63-g/L betaine hydrochloride, 0.39-g/L MgSO4, 0.85-g/L KCl, 10-mg/L FeSO4, 10-mg/L MnSO4, 0.84-g/L H3PO4, 1.05-g/L amino acid mixture, 24-mg/L antifoaming oil, and pH 6.8) [34]. The inoculum ratio was 10% (v/v) and the initial glucose concentration was 25 g/L. The temperature was set at 37 °C, and the pH was controlled between 6.7 and 6.9 with NH3·H2O. The dissolved oxygen level was kept at 25–35% by agitation (200–800 rpm). The glucose concentration was kept at 5–20 g/L by feeding the concentrated glucose (600 g/L). When the dissolved oxygen level is over 30%, the glucose feeding pump was manually turned on. Sucrose was used instead of glucose in the seed medium, because sucrose was better than glucose for the growth of this E. coli strain according to the preliminary experiments. The addition of small amount of low-cost amino acid mixture, vitamin B1, and ATP in some media is also based on the preliminary experiments which showed that these additions could improve the cell viability for l-threonine production. Every gram amino acid mixture used in this study contains 0.026-g aspartate, 0.034-g glutamate, 0.025-g serine, 0.018-g glycine, 0.014-g threonine, 0.010-g arginine, 0.014-g alanine, 0.003-g tryptophan, 0.004-g cysteine, 0.021-g methionine, 0.011-g phenylalanine, 0.011-g isoleucine, 0.009-g leucine, and 0.00419-g lysine.
Determination of OD600, amino acids, glucose, and organic acids
Biomass was determined by measuring OD600 with UV-1800 spectrophotometer (Shimadzu, Japan). During fermentation, 1 mL of the culture broth was taken at different time points and centrifuged at 12,000 rpm for 5 min. The supernatant was used to analyze the levels of glucose, amino acids, and organic acids.
The glucose concentration was measured with an SBA-40C biosensor (Institute of Biology, Shan-dong Academy of Science, China). To determine the levels of amino acids, the supernatant was diluted 20–100 folds, and filtered and analyzed by the 1200 series HPLC system (Agilent Technology, USA); the separation and quantification of amino acids were performed on a Thermo ODS-2HYPERSIL C18 column (250 mm × 4.0 mm, USA) using the ortho-phthalaldehyde precolumn derivatization method [14]. All the solvents used in HPLC analysis were purchased from Sigma-Aldrich (Shanghai, China).
The intracellular levels of acetyl-CoA and malate were determined according to the previous publication [41]. Two-mL E. coli culture (OD600 around 9.0) was harvested, washed with filtered water at 4 °C, and disrupted by a ultrasonic cell disruptor (Scientz-II D) for 10 min. The supernatants were collected and analyzed immediately. One unit of OD600 corresponds to 1.7-g/L cells. For acetyl-CoA measurement, the acetyl-CoA Assay Kit (Solarbio, Beijing, China) was used. According to the OD at 340 nm, the acetyl-CoA concentrations were calculated with the following formula: acetyl-CoA (nmol/g wet cell) = (1640 × ΔA + 0.012)/(OD600 × 1.7 × 0.002); ΔA = A80s − A20s.
For malate determination, the above supernatants were subjected to Agilent 1260 HPLC equipped with Diamonsil C18 column (5 μm, 250 mm × 4.6 mm no. 99603) (DiKMA technology, Beijing, China). A linear gradient elution procedure was employed as methanol:H2O:phosphate (from 5:95:0.05 to 60:40:0.05) in 20 min. Samples were detected with an ultraviolet detector at emission wavelengths 210 nm with a flow rate of 0.8 mL/min.
Qualitative and quantitative analysis of PHB
For intracellular PHB granules analysis, cells were harvested and washed by phosphate buffer saline (pH7.4), and then prepared for further microscopic examination according to previous method [29]. One-µmol Nile Red dissolved in dimethyl sulfoxide (1 μg/μL) was added [33]. The mixtures were incubated in dark at 37 °C for 30 min after vigorous mixing. Finally, the stained cells were washed three times with deionized water to remove any residual dye and resuspended in phosphate buffer saline (pH 7.4), and 10 µL of the solution was prepared on the slides with cover glass and nail polish to visualization by fluorescence microscope (Leica TCS SP8, Leica, Germany). Cell excitation was accomplished using a 488 nm argon laser. Photographs were captured with a Leica TCS SP8 software (CellSens Standard 1.9).
Quantification of PHB was according to previous method [29]. GC-2010 plus system (Shimadzu, Japan) was employed with a DB-WAX column (30 m × 0.32 mm) (Agilent Technologies, Waldbronn, Germany) and a flame ionization detector, and the injection temperature was 250 °C. Cells were harvested by centrifugation at 10,000 rpm for 5 min, washed twice with pH 7.2 phosphate-buffered saline, and then lyophilized for 48 h. About 10-mg lyophilized cells, 2-mL methanol (with 3% H2SO4) and 2-mL chloroform were added to esterification tubes and treated in boiled water for 8 h. 1 mL distilled water was added to esterification tubes at room temperature, and rotary vibrated for 5 min, and then, 0.5 mL of organic phase was collected and filtrated with 0.22-µm filters (Satorious, Germany). Calibration curves were constructed with commercially available PHB (Sigma-Aldrich, Saint Louis, Missouri).
Results
l-Threonine production in E. coli TWF001 is enhanced by overexpression of the gene cluster phaCAB
Overexpression of the gene cluster phaCAB in l-isoleucine producing C. glutamicum WM001 could significantly enhance l-isoleucine production [29]. Since l-threonine is the intermediate of l-isoleucine, the overexpression of the gene cluster phaCAB might also improve l-threonine production in the l-threonine-producing E. coli TWF001 [46]. The gene cluster phaCAB from R. eutropha [33] was overexpressed in TWF001, resulting in the strain TWF001/pFW01-phaCAB. The pH of the medium used for l-threonine production is usually 7.0 [17, 39], but the solubility of the fermentation medium used in this study at pH 7.0 is not as good as at pH 6.8. Therefore, the cell growth, glucose consumption, l-threonine production, and PHB formation in TWF001/pFW01-phaCAB grown at pH 7.0 and pH 6.8 were investigated, using TWF001/pFW01 as a control (Fig. 2). TWF001/pFW01-phaCAB and TWF001/pFW01 grew better, consumed glucose faster, and produced more l-threonine when grown at pH 6.8 than at pH 7.0; but TWF001/pFW01-phaCAB always grew better, consumed glucose faster, and produced more l-threonine than TWF001/pFW01 when they were grown at the same pH condition. After 36-h fermentation at pH 6.8, TWF001/pFW01-phaCAB produced 17.0-g/L l-threonine from 30-g/L glucose, and its OD600 reached 18.0, while TWF001/pFW01 only produced 9.7-g/L l-threonine from 29.1-g/L glucose, and its OD600 reached 16.0 (Fig. 2). The yield from glucose to l-threonine was 0.567 g/g for TWF001/pFW01-phaCAB but only 0.334 g/g for TWF001/pFW01. Compared to TWF001/pFW01, the l-threonine production and the conversion efficiency of glucose to l-threonine in TWF001/pFW01-phaCAB increased by 75% and 70%, respectively. The results indicate that overexpression of the gene cluster phaCAB significantly improved l-threonine production in E. coli TWF001.
Comparison of flask fermentation of E. coli TWF001/pFW01-phaCAB and TWF001/pFW01 grown at pH 7.0 and pH 6.8. a Cell growth; b glucose consumption; cl-threonine production. d The extracellular levels of acetate accumulated by E. coli TWF001/pFW01-phaCAB and TWF001/pFW01 cells after 36-h flask fermentation. The error bars indicate the standard deviations from three independent experiments
Acetate accumulation in E. coli affects the l-threonine production, but it could be reduced by the intracellular PHB production [10, 12, 13, 24]. The levels of acetate accumulation in TWF001/pFW01-phaCAB and TWF001/pFW01 after 36-h fermentation (pH 6.8) were determined (Fig. 2d). TWF001/pFW01-phaCAB accumulated 2.83-g/L acetate, while TWF001/pFW01 accumulated 7.65-g/L acetate. This suggests that PHB biosynthesis could decrease the intracellular level of acetate, and the increased l-threonine production in TWF001/pFW01-phaCAB might be relevant to the decrease of intracellular acetate.
To investigate whether the overexpression of phaCAB in TWF001 could produce PHB, TWF001/pFW01-phaCAB and TWF001/pFW01 cells were stained with Nile Red and analyzed using fluorescence microscopy (Fig. 3). In the DIC mode, some long cells were observed in TWF001/pFW01-phaCAB (Fig. 3a), but not in TWF001/pFW01 cells (Fig. 3b). In the FITC mode, green particles were observed in TWF001/pFW01-phaCAB (Fig. 3a), but not in TWF001/pFW01 cells (Fig. 3b). This indicates that PHB was produced in TWF001/pFW01-phaCAB cells due to the overexpression of the gene cluster phaCAB, because Nile Red-stained PHB granules can emit green fluorescence. When the DIC and FITC spectra were merged, it can be seen that only part of the TWF001/pFW01-phaCAB cells produced PHB and most of these cells were larger than the ones without PHB production. The formation of intracellular PHB granules depends on cell physiology, carbon source, nitrogen source, and the plasmid stability. More l-threonine production could be expected if the heterogeneity for PHB production in cells can be resolved and all cells could produce PHB. The PHB production in TWF001/pFW01-phaCAB grown at pH 7.0 and pH 6.8 was also quantified (Fig. 3c). TWF001/pFW01-phaCAB cells grown at pH 6.8 could produce 12.9% and 12.2% (wt%) PHB after 24-h and 36-h fermentation, respectively. However, more l-threonine was produced in TWF001/pFW01-phaCAB after 36-h fermentation than after 24 h fermentation (Fig. 2c). The results indicate that overexpression of the gene cluster phaCAB in E. coli TWF001 could produce PHB at the early growth stage and significantly improved l-threonine production at the latter stage.
PHB accumulation determination. a PHB accumulation in E. coli TWF001/pFW01-phaCAB and TWF001/pFW01 grown at pH 7.0 and pH 6.8. b Fluorescence microscopy images of TWF001/pFW01-phaCAB cells grown at pH 6.8. c Fluorescence microscopy images of TWF001/pFW01 cells grown at pH 6.8. DIC differential interference contrast, FITC fluorescence contrast, Merged overlapped contrast of DIC and FITC. Scale bar in white is 25 μm
Acetyl-CoA and malate biosynthesis increased in E. coli TWF001 after overexpression of phaCAB
To better understand the influence of the phaCAB overexpression in E. coli on the l-threonine biosynthesis, the levels of a few key intermediate metabolites and by-product acetate as well as the relative transcriptional levels of relevant genes were determined. Acetyl-CoA is the precursor of PHB, and can be catalyzed by malate synthase encoded by aceA to form malate and then fluxing to the biosynthesis of l-threonine. The level of acetyl-CoA is a key factor for both PHB and l-threonine synthesis. The intracellular levels of acetyl-CoA and malate in TWF001/pFW01 and TWF001/pFW01-phaCAB grown at the log phase were determined (Fig. 4a). The levels of acetyl-CoA and malate were both increased in TWF001/pFW01-phaCAB. The level of acetyl-CoA reached to 4.51 µmol/g, which is 2.16 times higher than that of the control TWF001/pFW01. The level of malate in TWF001/pFW01-phaCAB increased by 43%, compared with the control TWF001/pFW01. The results indicate that the overexpression of phaCAB could contribute to the enhancement of carbon flux to the acetyl-CoA and malate, further enhancing l-threonine production.
a The intracellular levels of acetyl-CoA and malate in E. coli TWF001/pFW01-phaCAB and TWF001/pFW01 grown at the log phase; b relative transcription levels of poxB, pta, ackA, acs, aceA, aceB aspC, thrA, thrB, and thrC analyzed by RT-PCR. The error bars indicate the standard deviations from three independent experiments
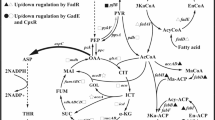
In TWF001/pFW01-phaCAB, acetyl-CoA could be used to biosynthesize PHB, acetate, and also flux to glyoxylate shunt. The enhanced glyoxylate shunt and the decreased acetate level are favorable for l-threonine biosynthesis. As expected, acetate accumulation could be reduced in TWF001/pFW01-phaCAB [10, 12, 13, 24]. In E. coli, acetate was synthesized either from pyruvate by PoxB or from acetyl-CoA by Pta-AckA; acetate could also be consumed to form acetyl-CoA by acetyl-CoA synthetase encoded by acs (Fig. 1). RT-PCR analysis showed that the transcriptional levels of poxB, pta, and ackA were down-regulated, but the transcriptional level of acs was up-regulated in TWF001/pFW01-phaCAB cells grown at the mid-log phase, compared to that in TWF001/pFW01 cells (Fig. 4b). In addition, the transcriptional levels of aceA, aceB, aspC, thrA, thrB, and thrC in TWF001/pFW01-phaCAB increased, compared to that in TWF001/pFW01 (Fig. 4b). This suggests that PHB formation could influence carbon flow distribution and enhance the glyoxylate shunt pathway, leading more carbon flux into acetyl-CoA and malate and eventually l-threonine production.
l-Threonine production was significantly increased TWF001/pFW01-phaCAB cells in fed-batch fermentation
The l-threonine production in the fed-batch fermentation of TWF001/pFW01-phaCAB was investigated using a 3-L fermenter (Fig. 5). TWF001/pFW01-phaCAB cells grew better than TWF001/pFW01. TWF001/pFW01-phaCAB reached the maximum OD600 of 49.7 at 28 h, while TWF001/pFW01 reached the maximum OD600 of 35.2 at 20 h (Fig. 5a). PHB granules in TWF001/pFW01-phaCAB might contribute to the high OD600. Similar patterns of glucose consumption were observed for TWF001/pFW01 and TWF001/pFW01-phaCAB during the fermentation (Fig. 5b). Similar levels of l-threonine production were observed in TWF001/pFW01-phaCAB and TWF001/pFW01 at the first 12 h, and then, more l-threonine were produced in TWF001/pFW01-phaCAB than in TWF001/pFW01. After 44 h, 96.4-g/L l-threonine were produced in TWF001/pFW01-phaCAB, but only 73.2-g/L l-threonine were produced in TWF001/pFW01 (Fig. 5c). The conversion ratio of glucose to l-threonine reached 0.54 g/g in TWF001/pFW01-phaCAB, but only reached 0.41 g/g in TWF001/pFW01. TWF001/pFW01-phaCAB produced 11.5% PHB and 6.3-g/L acetate, while TWF001/pFW01 produced 14.8-g/L acetate.
To further investigate the l-threonine production in larger scale fed-batch fermentation, E. coli strains were fermented in 10-L fermenter (Fig. 6). TWF001/pFW01-phaCAB cells grew slightly slower than TWF001/pFW01 before 6 h, but faster afterwards. After 36-h fermentation, TWF001/pFW01-phaCAB reached the maximum OD600 of 41.6, while TWF001/pFW01 reached the maximum OD600 of 33.2 (Fig. 6a). Much more l-threonine production was produced in TWF001/pFW01-phaCAB during the whole fermentation. After 36 h, 133.5-g/L l-threonine was produced in TWF001/pFW01-phaCAB, but only 94.3-g/L l-threonine was produced in TWF001/pFW01 (Fig. 6b). The conversion efficiency of glucose to l-threonine in TWF001/pFW01-phaCAB reached 0.50 g/g, while it only reached 0.38 g/g in the control TWF001/pFW01. In addition, TWF001/pFW01-phaCAB cells also accumulated approximately 13.0% PHB after 36-h fermentation.
In recent years, fermentation optimization has also been proved effective in improving l-threonine production. l-Threonine production in E. coli EC125 under the optimal feeding control reached 105.3 g/L [25], l-threonine production in E. coli TRFC with pseudo-exponential glucose-start feeding strategy reached 124.6 g/L [36], and l-threonine production in E. coli JLTHR reached 127.3 g/L when fed with 2-g/L betaine hydrochloride in 5-L fermenter [34]. In this study, 133.5-g/L l-threonine was produced in E. coli TWF001 when overexpressing phaCAB, strongly suggesting the importance of phaCAB expression. The PHB accumulation in TWF001/pFW01-phaCAB might rebalance metabolic flux distribution, decrease the acetate level, and increase the acetyl-CoA and malate levels [29], leading to the enhanced glyoxylate shunt and the increase of l-threonine production. The results in this study suggest that expressing the gene cluster phaCAB is an efficient strategy to improve l-threonine production in E. coli.
Discussion
In this study, the gene cluster phaCAB were introduced into the l-threonine-producing strain TWF001. The recombinant strain TWF001/pFW01-phaCAB produced 133.5 g/L after 36-h fed-batch fermentation in 10-L fermenter. Table 3 shows the comparison of production of l-threonine, acetyl-CoA, acetate, and PHB in TWF001/pFW01-phaCAB and TWF001/pFW01. The results suggest that the PHB accumulation could enhance the l-threonine production, possibly because the metabolic flux distribution was rebalanced [10, 12, 13, 44].
Our previous studies showed that enhancing the glyoxylate shunt [41, 46] and reducing the acetate level [5] could significantly improve the l-threonine production. The sufficient supply of precursor malate and acetyl-CoA is important for the glyoxylate shunt and l-threonine overproduction in E. coli [41]. Acetyl-CoA is mainly derived from pyruvate. If the glyoxylate shunt operates actively and acetyl-CoA is actively used for the production of oxaloacetate, l-threonine production might be enhanced [23, 34]. PHB accumulation might rebalance the metabolic flux [31]. Overexpression of phaCAB could pull more metabolic flux to acetyl-CoA [38], resulting in a larger “acetyl-CoA pool” [29]. Consequently, the carbon flux to the malate was also increased 43%, the increased levels of acetyl-CoA and malate could further lead to the enhancement of glyoxylate shunt with the up-regulation of aceA and aceB. These all benefit the improvement of L-threonine production with the up-regulated aspC, thrA, thrB, and thrC transcriptional levels. In our previous study, overexpression of phaCAB in C. glutamicum WM001 could also significantly enhance the production of l-isoleucine, the downstream product of l-threonine [29]. These results showed that PHB biosynthesis in bacteria could enhance the biosynthesis of acetyl-CoA which benefits the l-threonine accumulation.
Acetate is a major by-product for l-threonine production in E. coli [23, 39]. Acetate accumulation consumes a large amount of pyruvate and acetyl-CoA, and thus affects the l-threonine production [23, 39]. Reducing acetate accumulation could enhance the l-threonine production and improve glucose conversion efficiency [23, 34, 39]. Overexpressing the gene cluster phaCAB has been used to reduce acetate accumulation and increase the production of L-glutamate in C. glutamicum [24], l-tryptophan in E. coli [10], and succinate in E. coli [13]. In addition, PHB biosynthesis pathway also directly exerts metabolic control over the carbon flux distribution [35]. The l-threonine-producing E. coli strain TWF001 could accumulate a large amount of acetate (7.65 g/L), and therefore, in this study, the gene cluster phaCAB was overexpressed in TWF001. The resulting strain TWF001/pFW01-phaCAB only accumulated 2.83-g/L acetate, but produced 216% more acetyl-CoA and 43% more malate. All these influences could benefit the enhancement of l-threonine production. The genes poxB, pta, and ackA were down-regulated, while the gene acs was up-regulated, suggesting that the carbon flux was directed to the production of PHB or l-threonine. In addition, PHB is an inclusion body and could improve the resistance of bacterial cells to the harsh environment [9], warrant normal carbon metabolism, and enhance the carbon metabolism, leading to more carbon flow to acetyl-CoA [3, 37, 47], and up-regulate the expression of GroEL, GroES, DnaK, and sigma 38 factor [4, 11]. Therefore, the influence of PHB accumulation on l-threonine production in E. coli TWF001 might be complex.
Conclusions
In this study, the PHB synthesis pathway was introduced into an l-threonine-producing E. coli strain TWF001, leading to a significant increase of l-threonine production as well as the PHB formation. The l-threonine production of TWF001/pFW01-phaCAB reached 133.5 g/L after fed-batch fermentation in 10-L fermenter. Transcription analysis showed that the key genes in the l-threonine biosynthetic pathway were up-regulated, the genes poxB, pta, and ackA relevant to the formation of acetate were down-regulated, and the gene acs encoding the enzyme which converts acetate to acetyl-CoA was up-regulated, further leading to the increased levels of acetyl-CoA and malate, while the decreased acetate accumulation. The results suggested that PHB formation in E. coli could enhance the carbon flux towards l-threonine production.
References
Anderson AJ, Dawes EA (1990) Occurrence, metabolism, metabolic role, and industrial uses of bacterial polyhydroxyalkanoates. Microbiol Rev 54:450–472
Borchert AJ, Downs DM (2018) Analyses of variants of the Ser/Thr dehydratase IlvA provide insight into 2-aminoacrylate metabolism in Salmonella enterica. J Biol Chem 293:19240–19249. https://doi.org/10.1074/jbc.RA118.005626
Chen D, Xu D, Li M, He J, Gong Y, Wu D, Sun M, Yu Z (2012) Proteomic analysis of Bacillus thuringiensis DeltaphaC mutant BMB171/PHB(-1) reveals that the PHB synthetic pathway warrants normal carbon metabolism. J Proteom 75:5176–5188. https://doi.org/10.1016/j.jprot.2012.06.002
de Almeida A, Catone MV, Rhodius VA, Gross CA, Pettinari MJ (2011) Unexpected stress-reducing effect of PhaP, a poly(3-hydroxybutyrate) granule-associated protein, in Escherichia coli. Appl Environ Microbiol 77:6622–6629. https://doi.org/10.1128/AEM.05469-11
Ding Z, Fang Y, Zhu L, Wang J, Wang X (2019) Deletion of arcA, iclR and tdcC in Escherichia coli to improve l-threonine production. Biotechnol Appl Biochem. https://doi.org/10.1002/bab.1789
Dong X, Quinn PJ, Wang X (2011) Metabolic engineering of Escherichia coli and Corynebacterium glutamicum for the production of l-threonine. Biotechnol Adv 29:11–23. https://doi.org/10.1016/j.biotechadv.2010.07.009
Dong X, Quinn PJ, Wang X (2012) Microbial metabolic engineering for l-threonine production. Subcell Biochem 64:283–302. https://doi.org/10.1007/978-94-007-5055-5_14
Dong X, Zhao Y, Zhao J, Wang X (2016) Characterization of aspartate kinase and homoserine dehydrogenase from Corynebacterium glutamicum IWJ001 and systematic investigation of l-isoleucine biosynthesis. J Ind Microbiol Biotechnol 43:873–885. https://doi.org/10.1007/s10295-016-1763-5
Eggers J, Steinbuchel A (2014) Impact of Ralstonia eutropha’s poly(3-hydroxybutyrate) (PHB) depolymerases and phasins on PHB storage in recombinant Escherichia coli. Appl Environ Microbiol 80:7702–7709. https://doi.org/10.1128/AEM.02666-14
Gu P, Kang J, Yang F, Wang Q, Liang Q, Qi Q (2013) The improved l-tryptophan production in recombinant Escherichia coli by expressing the polyhydroxybutyrate synthesis pathway. Appl Microbiol Biotechnol 97:4121–4127. https://doi.org/10.1007/s00253-012-4665-0
Han MJ, Yoon SS, Lee SY (2001) Proteome analysis of metabolically engineered Escherichia coli producing poly(3-hydroxybutyrate). J Bacteriol 183:301–308. https://doi.org/10.1128/JB.183.1.301-308.2001
Kang Z, Du L, Kang J, Wang Y, Wang Q, Liang Q, Qi Q (2011) Production of succinate and polyhydroxyalkanoate from substrate mixture by metabolically engineered Escherichia coli. Bioresour Technol 102:6600–6604. https://doi.org/10.1016/j.biortech.2011.03.070
Kang Z, Gao C, Wang Q, Liu H, Qi Q (2010) A novel strategy for succinate and polyhydroxybutyrate co-production in Escherichia coli. Bioresour Technol 101:7675–7678. https://doi.org/10.1016/j.biortech.2010.04.084
Kőrös Á, Varga Z, Molnár-Perl I (2008) Simultaneous analysis of amino acids and amines as their o-phthalaldehyde-ethanethiol-9-fluorenylmethyl chloroformate derivatives in cheese by high-performance liquid chromatography. J Chromatogr A 1203:146–152. https://doi.org/10.1016/j.chroma.2008.07.035
Kruse D, Kramer R, Eggeling L, Rieping M, Pfefferle W, Tchieu JH, Chung YJ, Jr Saier MH, Burkovski A (2002) Influence of threonine exporters on threonine production in Escherichia coli. Appl Microbiol Biotechnol 59:205–210. https://doi.org/10.1007/s00253-002-0987-7
Lee JH, Lee DE, Lee BU, Kim HS (2003) Global analyses of transcriptomes and proteomes of a parent strain and an l-threonine-overproducing mutant strain. J Bacteriol 185:5442–5451. https://doi.org/10.1128/JB.185.18.5442-5451.2003
Lee JH, Sung BH, Kim MS, Blattner FR, Yoon BH, Kim JH, Kim SC (2009) Metabolic engineering of a reduced-genome strain of Escherichia coli for l-threonine production. Microb Cell Fact 8:2. https://doi.org/10.1186/1475-2859-8-2
Lee KH, Park JH, Kim TY, Kim HU, Lee SY (2007) Systems metabolic engineering of Escherichia coli for l-threonine production. Mol Syst Biol 3:149. https://doi.org/10.1038/msb4100196
Lee MH, Lee HW, Park JH, Ahn JO, Jung JK, Hwang YI (2006) Improved l-threonine production of Escherichia coli mutant by optimization of culture conditions. J Biosci Bioeng 101:127–130. https://doi.org/10.1263/jbb.101.127
Leong YK, Show PL, Ooi CW, Ling TC, Lan JC (2014) Current trends in polyhydroxyalkanoates (PHAs) biosynthesis: insights from the recombinant Escherichia coli. J Biotechnol 180:52–65. https://doi.org/10.1016/j.jbiotec.2014.03.020
Lin JH, Lee MC, Sue YS, Liu YC, Li SY (2017) Cloning of phaCAB genes from thermophilic Caldimonas manganoxidans in Escherichia coli for poly(3-hydroxybutyrate) (PHB) production. Appl Microbiol Biotechnol 101:6419–6430. https://doi.org/10.1007/s00253-017-8386-2
Lin Z, Zhang Y, Yuan Q, Liu Q, Li Y, Wang Z, Ma H, Chen T, Zhao X (2015) Metabolic engineering of Escherichia coli for poly(3-hydroxybutyrate) production via threonine bypass. Microb Cell Fact 14:185. https://doi.org/10.1186/s12934-015-0369-3
Liu J, Li H, Xiong H, Xie X, Chen N, Zhao G, Caiyin Q, Zhu H, Qiao J (2019) Two-stage carbon distribution and cofactor generation for improving l-threonine production of Escherichia coli. Biotechnol Bioeng 116:110–120. https://doi.org/10.1002/bit.26844
Liu Q, Ouyang SP, Kim J, Chen GQ (2007) The impact of PHB accumulation on l-glutamate production by recombinant Corynebacterium glutamicum. J Biotechnol 132:273–279. https://doi.org/10.1016/j.jbiotec.2007.03.014
Liu S, Liang Y, Liu Q, Tao T, Lai S, Chen N, Wen T (2013) Development of a two-stage feeding strategy based on the kind and level of feeding nutrients for improving fed-batch production of l-threonine by Escherichia coli. Appl Microbiol Biotechnol 97:573–583. https://doi.org/10.1007/s00253-012-4317-4
Liu Y, Li Q, Zheng P, Zhang Z, Liu Y, Sun C, Cao G, Zhou W, Wang X, Zhang D, Zhang T, Sun J, Ma Y (2015) Developing a high-throughput screening method for threonine overproduction based on an artificial promoter. Microb Cell Fact 14:121. https://doi.org/10.1186/s12934-015-0311-8
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Livshits VA, Zakataeva NP, Aleshin VV, Vitushkina MV (2003) Identification and characterization of the new gene rhtA involved in threonine and homoserine efflux in Escherichia coli. Res Microbiol 154:123–135. https://doi.org/10.1016/S0923-2508(03)00036-6
Ma W, Wang J, Li Y, Yin L, Wang X (2018) Poly(3-hydroxybutyrate-co-3-hydroxyvalerate) co-produced with l-isoleucine in Corynebacterium glutamicum WM001. Microb Cell Fact 17:93. https://doi.org/10.1186/s12934-018-0942-7
Mahishi LH, Tripathi G, Rawal SK (2003) Poly(3-hydroxybutyrate) (PHB) synthesis by recombinant Escherichia coli harbouring Streptomyces aureofaciens PHB biosynthesis genes: effect of various carbon and nitrogen sources. Microbiol Res 158:19–27. https://doi.org/10.1078/0944-5013-00161
Nolden L, Farwick M, Krämer R, Burkovski A (2001) Glutamine synthetases of Corynebacterium glutamicum: transcriptional control and regulation of activity. FEMS Microbiol Lett 201:91–98. https://doi.org/10.1111/j.1574-6968.2001.tb10738.x
Posfai G, Plunkett G 3rd, Feher T, Frisch D, Keil GM, Umenhoffer K, Kolisnychenko V, Stahl B, Sharma SS, de Arruda M, Burland V, Harcum SW, Blattner FR (2006) Emergent properties of reduced-genome Escherichia coli. Science 312:1044–1046. https://doi.org/10.1126/science.1126439
Spiekermann P, Rehm BH, Kalscheuer R, Baumeister D, Steinbuchel A (1999) A sensitive, viable-colony staining method using Nile red for direct screening of bacteria that accumulate polyhydroxyalkanoic acids and other lipid storage compounds. Arch Microbiol 171:73–80
Su Y, Guo QQ, Wang S, Zhang X, Wang J (2018) Effects of betaine supplementation on l-threonine fed-batch fermentation by Escherichia coli. Bioprocess Biosyst Eng 41:1509–1518. https://doi.org/10.1007/s00449-018-1978-0
Tyo KE, Fischer CR, Simeon F, Stephanopoulos G (2010) Analysis of polyhydroxybutyrate flux limitations by systematic genetic and metabolic perturbations. Metab Eng 12:187–195. https://doi.org/10.1016/j.ymben.2009.10.005
Wang J, Cheng LK, Chen N (2014) High-level production of l-threonine by recombinant Escherichia coli with combined feeding strategies. Biotechnol Biotechnol Equip 28:495–501. https://doi.org/10.1080/13102818.2014.927682
Wang Q, Zhuang Q, Liang Q, Qi Q (2013) Polyhydroxyalkanoic acids from structurally-unrelated carbon sources in Escherichia coli. Appl Microbiol Biotechnol 97:3301–3307. https://doi.org/10.1007/s00253-013-4809-x
Wang RY, Shi ZY, Chen JC, Wu Q, Chen GQ (2012) Enhanced co-production of hydrogen and poly-(R)-3-hydroxybutyrate by recombinant PHB producing E. coli over-expressing hydrogenase 3 and acetyl-CoA synthetase. Metab Eng 14:496–503. https://doi.org/10.1016/j.ymben.2012.07.003
Xie X, Liang Y, Liu H, Liu Y, Xu Q, Zhang C, Chen N (2014) Modification of glycolysis and its effect on the production of l-threonine in Escherichia coli. J Ind Microbiol Biotechnol 41:1007–1015. https://doi.org/10.1007/s10295-014-1436-1
Xu M, Qin J, Rao Z, You H, Zhang X, Yang T, Wang X, Xu Z (2016) Effect of polyhydroxybutyrate (PHB) storage on l-arginine production in recombinant Corynebacterium crenatum using coenzyme regulation. Microb Cell Fact 15:15. https://doi.org/10.1186/s12934-016-0414-x
Yang J, Fang Y, Wang J, Wang C, Zhao L, Wang X (2019) Deletion of regulator-encoding genes fadR, fabR and iclR to increase l-threonine production in Escherichia coli. Appl Microbiol Biotechnol 103:4549–4564. https://doi.org/10.1007/s00253-019-09818-8
Yuzbashev TV, Vybornaya TV, Larina AS, Gvilava IT, Voyushina NE, Mokrova SS, Yuzbasheva EY, Manukhov IV, Sineoky SP, Debabov VG (2013) Directed modification of Escherichia coli metabolism for the design of threonine-producing strains. Appl Biochem Microbiol 49:723–742. https://doi.org/10.1134/S0003683813090056
Zakataeva NP, Aleshin VV, Tokmakova IL, Troshin PV, Livshits VA (1999) The novel transmembrane Escherichia coli proteins involved in the amino acid efflux. FEBS Lett 452:228–232
Zhang X, Zhang J, Xu J, Zhao Q, Wang Q, Qi Q (2018) Engineering Escherichia coli for efficient coproduction of polyhydroxyalkanoates and 5-aminolevulinic acid. J Ind Microbiol Biotechnol 45:43–51. https://doi.org/10.1007/s10295-017-1990-4
Zhang Y, Meng Q, Ma H, Liu Y, Cao G, Zhang X, Zheng P, Sun J, Zhang D, Jiang W, Ma Y (2015) Determination of key enzymes for threonine synthesis through in vitro metabolic pathway analysis. Microb Cell Fact 14:86. https://doi.org/10.1186/s12934-015-0275-8
Zhao H, Fang Y, Wang X, Zhao L, Wang J, Li Y (2018) Increasing l-threonine production in Escherichia coli by engineering the glyoxylate shunt and the l-threonine biosynthesis pathway. Appl Microbiol Biotechnol 102:5505–5518. https://doi.org/10.1007/s00253-018-9024-3
Zhuang Q, Wang Q, Liang Q, Qi Q (2014) Synthesis of polyhydroxyalkanoates from glucose that contain medium-chain-length monomers via the reversed fatty acid beta-oxidation cycle in Escherichia coli. Metab Eng 24:78–86. https://doi.org/10.1016/j.ymben.2014.05.004
Acknowledgements
This study was supported by the National Key R&D Program of China (2018YFA0900302), the National First-Class Discipline Program of Light Industry Technology and Engineering (LITE2018-10), the post-graduate research and practice innovation program of Jiangsu Province (KYLX15_1141), and the Collaborative Innovation Center of Jiangsu Modern Industrial Fermentation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, J., Ma, W., Fang, Y. et al. Increasing l-threonine production in Escherichia coli by overexpressing the gene cluster phaCAB. J Ind Microbiol Biotechnol 46, 1557–1568 (2019). https://doi.org/10.1007/s10295-019-02215-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-019-02215-0