Abstract
It is estimated that one in 100 men have azoospermia, the complete lack of sperm in the ejaculate. Currently, ~ 20% of azoospermia cases remain idiopathic. Non-obstructive azoospermia (NOA) is mostly explained by congenital factors leading to spermatogenic failure, such as chromosome abnormalities. The knowledge of the monogenic causes of NOA is very limited. High genetic heterogeneity due to the complexity of spermatogenesis and testicular function, lack of non-consanguineous familial cases and confirmatory studies challenge the field. The reported monogenic defects cause syndromic NOA phenotypes presenting also additional congenital problems and isolated NOA cases, explained by spermatogenic defects. The established and recently reported NOA genes (n = 38) represent essential guardians of meiosis, transcriptional and endocrine regulators of reproduction. Despite the list being short, 92% of these loci are predicted to functionally interact with each other (STRING analysis: average 5.21 connections/gene, enrichment P < 10–16). Notably, ~ 50% of NOA genes have also been implicated in primary ovarian insufficiency, amenorrhea and female genital anomalies, referring to overlapping mechanisms. Considering the knowledge from respective female phenotypes and animal models, exploring the scenarios of di/oligogenic and de novo mutations represent perspective directions in the genetic research of NOA. Knowing the exact genetic cause in each patient improves the management of infertility and other health risks (e.g., cancer), and facilitates the counseling of family members about their reproductive health. Uncovering the loci and biological processes implicated in NOA will also broaden the understanding of etiologies behind spermatogenic failure and promote the development of novel non-invasive treatments for male infertility.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Male infertility can be caused by either quantitative or qualitative spermatogenic impairment. The former refers to the condition, where mature sperm cells are few in number, but have no apparent defects in their structure, motility and fertilization capacity. In the latter case, sperm cells have morphological malformations either in the head or tail and thus, are characterized by defective motility or inability to give rise to a viable embryo. In clinical practice, quantitative or qualitative impairment of spermatogenesis in andrology patients may also appear hand-in-hand. The most severe form of male infertility is azoospermia, referring to the complete lack of sperm in the ejaculate. Despite the extreme phenotype, the estimated prevalence of azoospermia in the general population is surprisingly high, one in 100 men (Stephen and Chandra 2006). Among patients with male factor infertility [< 39 million sperm per ejaculate (WHO 2010)], azoospermia cases represent 10–20% (Jarow et al. 1989; Olesen et al. 2017; Punab et al. 2017; Tüttelmann et al. 2011).
Azoospermia—established knowledge and current challenges
Etiology of azoospermia
Although the primary diagnosis of azoospermia is straightforward and explicit using semen analysis and hormonal evaluation, there is a high heterogeneity of clinical sub-phenotypes complicating the assessment of underlying disease etiology in each patient (Fig. 1, Table 1). The current routine andrological workup is able to assign the primary cause to ~ 80% of azoospermia patients (Punab et al. 2017; Tüttelmann et al. 2011). Up to 30% of cases represent obstructive azoospermia (OA) due to physical blockage in their genital tract without directly affecting sperm production. OA can be suspected if testicular volume and serum FSH levels are within a normal range. The majority of OA cases are caused by acquired conditions (Fig. 1). Diagnostic workup of azoospermia patients includes screening mutations in the CFTR gene that are known to cause congenital OA due to abnormal formation or bilateral absence of the vas deferens [~ 3% of azoospermia (Punab et al. 2017; Tüttelmann et al. 2011)].
Etiology of azoospermia (based on data from Punab et al. 2017)
The rest of the azoospermia patients (> 70%) represent non-obstructive azoospermia (NOA), a spectrum of testicular disorders resulting in spermatogenic failure and typically also reduced testes size. The majority of NOA patients have primary testicular failure due to an intrinsic defect in the initiation or normal progression of spermatogenesis that is often reflected in elevated serum FSH levels. A minor fraction of NOA cases present secondary testicular failure caused by endocrine disturbances or other pre-testicular factors, e.g., developmental defects. In contrast to OA, the majority of NOA cases represent various congenital conditions and a notable proportion of patients are diagnosed with already known genetic causes, such as abnormal karyotype (up to 17% of the patients) and pathogenic Y-chromosomal microdeletions (2–10%; Fig. 1) (Krausz et al. 2014; Olesen et al. 2017; Punab et al. 2017; Tournaye et al. 2017; Tüttelmann et al. 2011). The most commonly detected genetic abnormality is 47, XXY karyotype causing the Klinefelter syndrome and accounting for ~ 15% of all azoospermia cases (Punab et al. 2017; Vockel et al. 2019). Tests for karyotype abnormalities and Y-chromosomal microdeletions are routinely offered to andrology patients for 20 years already. However, the knowledge of monogenic causes of NOA is limited and none of the current clinical guidelines include mutational analysis of any NOA genes (Jarvi et al. 2015; Jungwirth et al. 2012; Practice Committee of the American Society for Reproductive Medicine in collaboration with the Society for Male Reproduction and Urology 2018).
Challenges in identifying the monogenic causes of spermatogenic failure
NOA is not a single genetic condition, but rather a clinical endpoint of a spectrum of alternative pathological processes and sub-phenotypes (Krausz and Riera-Escamilla 2018). Given the large number of genes implicated in spermatogenesis and testicular function (Chalmel et al. 2012; Soraggi et al. 2020), a high heterogeneity in monogenic defects that may cause NOA is expected. Alternative forms of genetic inheritance of the condition are to be considered. NOA may manifest itself through autosomal recessive (AR) mutations inherited from fertile parents and combined to a pathogenic genotype in the homozygous, compound heterozygous or hemizygous form. Alternatively, NOA can be caused by maternally inherited or de novo mutations in X-chromosomal or dosage sensitive autosomal dominant (AD) genes. In rare occasions, an AD mutation with reduced penetrance or a de novo Y-chromosomal microdeletion can be inherited from a fertile father. Thus, a complete medical examination of the patient and his longitudinal health records is strongly recommended to complement the testicular and hormonal assessment during a routine andrology workup.
A specific challenge in research of the monogenic causes of NOA is the lack of familial cases. Most NOA patients are singleton, sporadic cases in their families. Due to this restriction, it is impossible to perform a proper familial segregation and linkage analysis that has been the key tool to uncover inborn errors of other Mendelian phenotypes (Posey et al. 2019). To further complicate matters, mutations in AD genes implicated in syndromic forms of NOA often exhibit incomplete penetrance and variable phenotype, including non-affected carriers reported in the familial studies. This sets an additional challenge in interpreting the genetic tests and making conclusions about the causative nature of identified variants.
Finally, although the majority of the NOA cases are expected to be sporadic, analysis of the genomes of parents and siblings along with the proband will have a clear benefit in excluding candidate variants carried by fertile male family members and in identifying de novo variants as possible causes of infertility. Still, in many cases the motivation and psychological readiness of either the patient or the family restrict the recruitment and genetic analysis of the whole pedigree. Family planning and related issues are usually considered highly private matters, and having difficulties conceiving a child is typically not discussed among relatives.
Monogenic causes of NOA
Approach for data extraction from the available literature resources
To reach a high-confidence list of genes implicated in monogenic NOA, a search was conducted in August 2019 in the following databases: Human Phenotype Ontology (HPO) (Köhler et al. 2019), Online Mendelian Inheritance in Man (OMIM, https://omim.org/) and PubMed (https://www.ncbi.nlm.nih.gov/pubmed). From the HPO database, all 39 genes listed under the term HP:0011961 (non-obstructive azoospermia) were considered (Supplementary Table 1). OMIM was queried for “non-obstructive azoospermia”, listing 26 candidate genes. In parallel, a literature search was conducted in PubMed with the following search term: ("nonobstructive azoospermia"[All Fields] OR "non-obstructive azoospermia"[All Fields]) AND ("mutation"[MeSH Terms] OR "mutation"[All Fields]) NOT ("review"[Publication Type] OR "review literature as topic"[MeSH Terms] OR "review"[All Fields]), which resulted in 214 publications. All extracted HPO and OMIM genes were manually assessed along with the supporting literature reports to show the evidence for the causative link between gene mutations and monogenic NOA. Among publications retrieved from the PubMed search, only studies reporting monogenic mutations in NOA patients were considered, whereas all genetic associations (both SNP and CNV based) and reports on deletions/duplication involving multiple genes were excluded.
The final manually assessed gene list was divided into three categories: established causative NOA genes with support from at least two independent studies (n = 22; Tables 2, 3); promising candidate genes reported in a single study and supported by in vitro or in vivo experimental data (n = 16; Table 4); genes that currently lack explicit evidence for the monogenic causative link to NOA (n = 29; Supplementary Table 2). While the established monogenic causes of NOA represent diverse modes of inheritance including AR, AD and X-linked genes, mutations in the majority of novel proposed genes are expressed in the AR form. For some proposed candidate genes, the inheritance mode is still unclear due to limited number of reported NOA cases and conflicting data in different sources of information.
Isolated and syndromic forms of monogenic NOA
Based on the critical assessment of available genotype–phenotype data, the known and recently proposed candidate genes were assigned to either an isolated form of NOA presenting no other major health complications or a syndromic form of NOA characterized by several concurrent clinical symptoms with variable phenotypic expressivity, including NOA in male mutation carriers. Among the well-established genes implicated in NOA, 10 have been linked to isolated NOA and 12 reported to cause syndromic disease phenotypes (Tables 2, 3). The recently proposed additional candidate genes represent 13 isolated and three syndromic NOA loci (Table 4).
The STRING analysis of physical and functional protein–protein interactions (Szklarczyk et al. 2019) demonstrated that the established and novel candidate genes for monogenic NOA belong to a dense network of ‘predicted functional partners’ (Fig. 2a). Although the number of shortlisted genes is not extensive, the majority of them (35 of 38; 92%) are functionally linked with an average of 5.21 active connections per locus. Overall, there is a statistically highly significant enrichment of interactions among the loci in the network (observed 99, expected 7; P < 10–16; Supplementary Table 3). Interestingly, the proteins form two separate clusters of interactions that largely overlap with the gene lists representing either the isolated or syndromic NOA condition. This further underlies different etiologies of the two forms of NOA and their accompanying health consequences. The following chapters introduce the three broader categories of genes implicated in NOA, representing essential guardians of meiosis, transcriptional regulators of male gonadal development and function, and endocrine regulators of the reproductive system.
a STRING network analysis of 38 established and novel proposed NOA genes. The analysis of physical and functional protein–protein interactions was performed using the default settings (Szklarczyk et al. 2019). Edge colors correspond to interactions according to the shown legend. The network consists of 99 inter–locus interactions, whereas the expected number of by-chance interactions between 38 proteins is 7 (an enrichment P value < 1 × 10–16). Details on each protein–protein pairwise interaction are provided in Supplementary Table 3. b The most significant results from the functional enrichment analysis of the 38 NOA genes. Top 25 terms from Gene Ontology ‘Biological processes’ category are shown (FDR ≤ 4.62 × 10–8). GO terms are ordered on the X-axis based on the significance of gene enrichment in this category, from left to right. Detailed results are presented in Supplementary Tables 4A–C
Genetic defects affecting meiosis and DNA repair: isolated and syndromic NOA
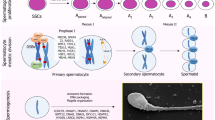
The process of spermatogenesis is inherently complex, consisting of various stages of mitosis, meiosis and spermiogenesis to transform haploid spermatids into mature sperm. So far, the largest category of monogenic defects detected in NOA patients comprises 19 genes involved in different stages of spermatogenesis, mostly functioning in the prophase of the first meiotic division (Fig. 3). Eight of these are established NOA genes (Table 2), whereas 11 still require independent confirmation of the link to monogenic NOA (Table 4). Mutations in the majority of these genes (16/19) result in isolated NOA with maturation arrest (MA) or Sertoli cell-only syndrome (SCOS).
Established (green) and novel proposed (orange) NOA genes (n = 19) implicated in human spermatogenesis, and in the maintenance of genomic integrity in mitosis and meiosis. Some loci function only in specific spermatogenic stages, whereas others are involved in multiple steps. Defects in nearly half (n = 8; underlined) of these genes have also been reported in female patients with primary ovarian insufficiency
In the spermatogenic dynamics, balance between the maintenance of mitotic quiescence and meiotic entry of germ cells is crucial. Mutations in NANOS2 that contribute to the maintenance of the spermatogonial stem cell population and suppression of meiotic entry (Saba et al. 2014), were recently reported to co-segregate with SCOS (Fakhro et al. 2018). Taking into account the high number of mitotic cell divisions in the pre-meiotic phase, defects in the proteins regulating DNA replication and repair, and maintaining overall genomic integrity are strong candidates for spermatogenic failure. During the last couple of years, loss-of-function (LoF) or pathogenic missense variants in the FANCM, FANCA, XRCC2, MCM8, and SETX genes involved in these processes have been reported in either isolated or syndromic NOA patients (Becherel et al. 2019; Catford et al. 2019; Kasak et al. 2018; Krausz et al. 2014; Tenenbaum-Rakover et al. 2015; Yang et al. 2018a; Yin et al. 2019). Importantly, as all these proteins are also involved in regulating meiotic recombination, double-stranded break (DSB) repair and ultimate chromosomal crossing over, they represent essential guardians of genome stability through spermatogenesis (Fig. 3). Three of these highlighted NOA genes—FANCM, FANCA, and XRCC2 belong to the Fanconi anemia (FA) pathway (Niraj et al. 2019), and MCM8 has been suggested to interact with members of the FA pathway in cross-link repair during replication (Griffin and Trakselis 2019). FANCM is a testis-enhanced gene that fulfills the most diverse palette of functions in the pathway, including interstrand cross-link removal, anti-crossover function, and protection against replication interference by RNA–DNA hybrids (Basbous and Constantinou 2019). Whereas FANCM is one of the few genes in the pathway that does not cause the FA phenotype, FANCA is the most commonly mutated gene in the genetically heterogeneous FA disorder with variable age of onset (Krausz et al. 2014; Shimamura and Alter 2010). Importantly, infertility is a fairly common clinical feature of male as well as female FA patients (Tsui and Crismani 2019). A novel NOA-linked gene is SETX encoding sentaxin acting as a DNA/RNA helicase involved in diverse aspects of RNA metabolism and genomic integrity (Andrews et al. 2018; Bennett and La Spada 2018). Although SETX is primarily linked with Ataxia with Oculomotor Apraxia Type 2 (AOA2) (Moreira et al. 2004) and amyotrophic lateral sclerosis (Chen et al. 2004), recent independent reports have shown that AOA2 male patients also exhibit meiotic arrest at the primary spermatocyte stage (Table 2).
Centriole duplication, involving the PLK4 protein, is a further critical process to be completed before primary spermatocytes can undergo meiosis (Habedanck et al. 2005). Although homozygous LoF variants in PLK4 cause microcephaly and chorioretinopathy (Martin et al. 2014; Shaheen et al. 2014), a recent study reported a heterozygous LoF variant as a novel candidate to explain a NOA case (Miyamoto et al. 2016).
Creation of DSBs by a specific DNA topoisomerase called SPO11 is essential to initiate meiotic recombination and formation of the synaptonemal complex between homologous chromosomes (Tock and Henderson 2018). Fakhro et al. 2018 detected a homozygous missense mutation in SPO11 in two brothers with meiotic arrest. MEI1 represents a further gene that is implicated in DSB formation and has been reported to be mutated in NOA patients exhibiting MA at the spermatocyte stage (Ben Khelifa et al. 2018; Nguyen et al. 2018). Mei1 null mice fail to complete the first meiotic division (Libby et al. 2003). MEI1 along with other established NOA genes (MEIOB, TEX15, and TEX11) also contributes to the successful formation and maintenance of the synaptonemal complex and crossovers between homologous chromosomes. Tex11-deficient spermatocytes show abnormal synapsis and undergo apoptosis at the pachytene stage (Yang et al. 2008). In men, TEX11 mutations were first described in NOA patients in two parallel reports (Yang et al. 2015; Yatsenko et al. 2015) and have been henceforth referred in several studies being strongly associated with the manifestation of NOA due to MA (Sha et al. 2018). More recently, recessive missense or frameshift variants in the MEIOB gene were reported in seven patients (from three families) also presenting the testicular MA phenotype (Gershoni et al. 2017, 2019). Consistently, Meiob-deficient mice fail to form crossovers and repair DSBs in germ cells and are infertile due to meiotic arrest (Luo et al. 2013). The functional failure of synaptonemal complex to explain a subset of isolated NOA cases is further supported by fresh reports on novel defective genes CCDC155, SYCE1, and RNF212 identified in patients diagnosed with MA (Fakhro et al. 2018; Maor-Sagie et al. 2015; Riera-Escamilla et al. 2019). The respective mutant mouse models are supportive to the human genetic data (Bolcun-Filas et al. 2009; Horn et al. 2013; Reynolds et al. 2013).
A critical step in the completion of crossing overs is DSB repair, involving a large complex of additional critical proteins (e.g., TEX15, DMC1), that have been identified to harbor pathogenic mutations in NOA patients (Colombo et al. 2017; He et al. 2018; Okutman et al. 2015). For example, TEX15 is responsible for the recruitment of DNA repair proteins onto DSB locations and DMC1 is a meiosis-specific recombinase interacting with several DNA repair proteins in the FA pathway, such as BRCA2 (Thorslund et al. 2007). The most recent addition to the list of confident isolated NOA genes is STAG3 that was reported within a short timeframe in two separate studies to be implicated in MA (Riera-Escamilla et al. 2019; van der Bijl et al. 2019). In addition to meiotic DSB repair, STAG3 is involved in the formation of chromosomal axis and cohesion of sister chromatids after DNA replication. Disruption of this protein leads to the persistence of meiotic DSBs and a failure to complete chromosome pairing (Riera-Escamilla et al. 2019). The dramatic chromosomal deficiencies in human (van der Bijl et al. 2019) were observed also in Stag3−/− mice (Winters et al. 2014).
Additional mechanism disrupted in NOA is the maintenance of stable intercellular bridges, thought to enable rapid ‘communication’ between gametes (Greenbaum et al. 2011). Mutations in TEX14 have proposed to cause impaired spermatogenesis due to failure in maintaining these intercellular cytoplasmatic connections (Fakhro et al. 2018; Gershoni et al. 2017). Interestingly, a pathogenic variant in the TDRD9 gene essential for retrotransposon silencing in meiosis has lately been shown to segregate with NOA in a large consanguineous Bedouin family (Arafat et al. 2017). Expression of Line1 transcripts has been shown to be increased in the Tdrd9−/− mice and although spermatocytes initiate the early DNA recombination pathway, spermatogenesis arrests at the stage of zygotene due to failed synapsis (Shoji et al. 2009).
Taken together, prime candidates for defective genes in patients with isolated NOA presenting MA and SCOS are loci implicated in genomic integrity, regulation of meiotic progression, DNA recombination and repair. In addition, there is growing evidence that these pathways are also involved in syndromic NOA caused by mutations in pleiotropic genes.
Mutations in transcriptional regulators of testicular function: syndromic and isolated NOA
The second category of genes implicated in NOA represents transcriptional regulators of testicular development and spermatogenesis. Testicular gene expression across the life span follows a functionally and temporally conserved pattern, including critical time windows during the fetal, neonatal and pubertal development, and the highly conserved process of spermatogenesis (Cardoso-Moreira et al. 2019; Chalmel et al. 2012; Zimmermann et al. 2015). Although the testicular expression of each locus is expected to be tightly controlled by a high number of regulators and interlinked cellular processes (Soumillon et al. 2013), the list of identified transcription modulating genes mutated in NOA patients is rather short. This includes specific transcription factors (NR5A1, WT1) and regulators of transcriptional (ZMYND15) or translational activity (TDRD7) in human testis, and a ubiquitously expressed component of the transcription initiation complex, TAF4B (Tables 2, 4).
The two well-known and functionally interacting transcription factors implicated in gonadal development in both sexes are NR5A1 encoding a Steroidogenic factor 1 (SF1), and WT1 encoding Wilms’ tumor protein (Hanley et al. 1999). In fact, WT1 modulates SF1 expression in a sex-specific manner (Nachtigal et al. 1998). Mutations in NR5A1 and WT1 primarily cause AD syndromic phenotypes, including monogenic disorders of sex development [DSD; reviewed in (Cools et al. 2018)]. In andrology practice, defects in the WT1 or NR5A1 genes could be suspected in patients presenting hypospadias, cryptorchidism, and other signs of congenital testicular damage (Eggers et al. 2016; Köhler et al. 2011). Defects in both genes are characterized by variable expressivity and incomplete penetrance, including asymptomatic family members (Kaneko et al. 2015; Lourenco et al. 2009).
While the action of WT1 in utero is critical in several stages in the development of the urogenital system, its postnatal activity is limited to the renal glomerulus. Mutated or deleted WT1 leads to a spectrum of congenital defects in kidneys and genitalia (e.g., Denys-Drash syndrome, Wilms’ tumor, nephropathy etc.) (Hastie 2017). Molecular diagnostics is critical as the majority of patients with the initial diagnosis of DSD will develop Wilms' tumor, nephropathy and/or gonadal tumor (Köhler et al. 2011). However, pathogenic WT1 missense variants have also been reported in patients with the primary diagnosis of NOA without malformations in the genitourinary tract (Seabra et al. 2015; Wang et al. 2013; Xu et al. 2017).
NR5A1 represents the second most commonly mutated gene among DSD patients (Eggers et al. 2016), and in most extreme cases its defects may cause sex reversal in both, XY- and XX-subjects (Achermann et al. 1999; Bashamboo et al. 2016). There is solid literature evidence that some pathogenic missense variants may cause ‘only’ spermatogenic failure, including NOA, without any clearly identifiable developmental defects in the testis (Bashamboo et al. 2010; Ferlin et al. 2015; Zare-Abdollahi et al. 2015). As SF1 is also implicated in the adrenal, spleen and pituitary function, the syndromic phenotype in the patients may include additional congenital defects that have to be considered in the clinical management (Cools et al. 2018).
Recently, LoF variants in the TDRD7 gene were reported in two analyzed consanguineous families to cause a rare syndrome combining congenital cataract in both sexes and NOA in affected men, supported by respective mouse models (Tan et al. 2019). TDRD7 is a component of chromatoid bodies contributing to the post-transcriptional regulation of specific mRNAs and has dual roles in the development of lens in utero and haploid spermatids in adulthood (Lachke et al. 2011; Tanaka et al. 2011).
Two transcriptional regulators have been proposed to explain isolated NOA (Ayhan et al. 2014). Homozygous truncating variants in a testis-specific transcriptional repressor ZMYND15 (Yan et al. 2010) and ubiquitous coactivator TAF4B were reported in azoospermic brothers in consanguineous families. As the patients shared large homozygous regions, oligogenic contributors to NOA cannot be ruled out and the claims require further supportive data.
Taken together, given the broad spectrum of downstream targets, and pre- and postnatal expressional dynamics, the potential target group to screen known or undescribed congenital defects in transcriptional regulators represents NOA patients with mostly syndromic phenotypes. However, isolated NOA cannot be excluded.
Congenital hormonal defects in reproductive physiology: syndromic and isolated NOA
The third broader functional category of congenital defects causing the NOA phenotype includes genes that act in the hypothalamic–pituitary–gonadal (HPG) axis. Mutations in genes involved in this pathway lead to secondary testicular failure due to impaired central hormonal regulation of the testis function, referred as congenital hypogonadotropic hypogonadism (CHH) (Boehm et al. 2015; Swee and Quinton 2019; Young et al. 2019). Patients with CHH may show syndromic phenotypes characterized by endocrine disturbances, pubertal absence or delay, cryptorchidism, micropenis, small testis size (bitesticular volume < 8 ml), gynaecomastia and variable accompanying developmental defects (e.g., anosmia, renal agenesis, cleft-lip palate, anomalies of digits, hearing impairment) (Boehm et al. 2015). However, normosmic CHH may present delayed or completely absent puberty as an isolated phenotype (Young et al. 2019). CHH is a rare clinical condition [prevalence 1/8000; (Dode and Hardelin 2010)]. The majority of andrological patients with CHH diagnosis present NOA and among all azoospermia patients, normosmic and anosmic CHH cases represent ~ 2.0–2.4% (Fig. 1; Punab et al. 2017; Tüttelmann et al. 2011). There is a wealth of literature reporting heterogeneous pathogenic variants in at least 30 genes causative to isolated, syndromic or both forms of CHH (Table 3; Maione et al 2018; Young et al. 2019). Notably, the mutations in genes implicated in CHH vary in regards to inheritance mode, penetrance and tolerance to asymptomatic carriers. There is also a solid body of data showing that pathogenic variants in several CHH genes are expressed only in oligogenic background (Supplementary Table 2; Boehm et al. 2015; Maione et al 2018; Pitteloud et al. 2007; Sykiotis et al. 2010).
Classical CHH results from the abnormal development, migration or function of specific neurons responsible for the secretion of hypothalamic gonadotropin releasing hormone (GnRH) that stimulates the expression of FSH and LH in the pituitary. Due to absent or delayed puberty, the majority of CHH cases are clinically recognized, diagnosed and managed already in adolescence or already during the minipuberty in infancy (Swee and Quinton 2019; Young et al. 2019). The first described CHH gene was ANOS1 (KAL1) encoding a secreted glycoprotein anosmin-1 regulating cellular adhesion and migration of nerve cell precursors, including olfactory and GnRH neurons. The primary phenotype resulting from the defective ANOS1 is Kallmann syndrome characterized by CHH combined with anosmia due to maldevelopment of the olfactory bulb (Bick et al. 1992; Hardelin et al. 1992). Further CHH genes are implicated in other syndromic phenotypes with variable expressivity. For example, mutations in orphan nuclear receptor gene NR0B1 (DAX1) cause congenital adrenal hypoplasia with/without CHH or NOA (Muscatelli et al. 1994; Suntharalingham et al. 2015). In extreme cases, NR0B1 mutations may cause sex reversal (Suntharalingham et al. 2015). Defects in brain-expressed helicase CHD7 cause either milder CHH or more severe CHARGE syndrome phenotype (Marcos et al. 2014). Other major CHH genes encode either morphogenic proteins or their receptors that are critical in GnRH neuron fate specification (FGFR1), development (PROK2, PROKR2), GnRH secretion or action (GNRH1/GNRHR, KISS1R, TACR3) (Table 3).
Genetic defects affecting primarily the function of gonadotropins, follicle stimulating (FSH) and luteinizing hormone (LH) are extremely rare (Nagirnaja et al. 2010). Most of the patients reported in the literature originate from consanguineous families. During the past 20 years, only three male patients have been described with homozygous pathogenic mutations in the FSHB gene (Table 3). All cases showed low FSH, impaired sexual development, hypogonadism and NOA. Likewise for the LHB gene, only five azoospermia patients with homozygous or compound heterozygous pathogenic variants have been reported since the seminal publication in 1992. Other clinical features of congenital absence of LH include delayed/absent puberty and sexual infantilism, gynecomastia and micropenis, low testicular volume. In contrast to the defects in gonadotropin-encoding genes, mutations in the testicular receptors of FSH and LH do not cause NOA. FSHR mutations lead to premature ovarian insufficiency (POI) in women (Tapanainen et al. 1997). Inactivating or activating LHGCR mutations cause other male reproductive disorders, such as Leydig cell hypoplasia or precocious puberty, respectively (Segaloff 2009).
Taken together, there is a broad spectrum of pathogenic variants and phenotypic variation in patients with hypogonadotropic hypogonadism, implicated in syndromic or isolated NOA. Although diagnosis of CHH among NOA patients is infrequent (~ 2%), the major genetic defects behind the condition are rather well known, the diagnostic yield of molecular tests exceeds 50% and clinical management options are well-established (Boehm et al. 2015; Tournaye et al. 2017; Maione et al. 2018). Spermatogenic failure in patients diagnosed with CHH is often treatable using either gonadotropin hormone injections or pulsatile GnRH administration (Frapsauce et al. 2011; Pitteloud et al. 2002; Swee and Quinton 2019; Young et al. 2019).
Other reported rare monogenic defects in NOA patients
Mutations in the NOA patients have been reported in additional candidate genes involved in novel biological pathways. Two NOA brothers and a sporadic SCOS patient were identified as carriers of hemizygous missense variants in WNK3 that functions as a serine/threonine kinase in regulating intracellular chloride concentrations and cellular volume (Fakhro et al. 2018). Two brothers from a consanguineous family carried a homozygous LoF variant in the SPINK2 gene encoding a serine protease inhibitor preventing premature activation of proacrosin to acrosin (Kherraf et al. 2017). In mice, deficiency of SPINK2 results in the fragmentation of the Golgi apparatus and arrest of cell proliferation. These novel NOA candidate genes and pathways require further confirmation in independent studies and clinical cases.
Discussion on the state-of-the-art and beyond
Summary of the current knowledge on the monogenic causes of NOA
The reported monogenic defects causing NOA represent either distinct syndromic phenotypes that concern a small proportion of andrology patients or rare isolated NOA cases, explained by errors in various stages of spermatogenesis (Tables 2, 3 and 4, Fig. 3). However, due to the limited number of published studies, the current list of genes under the category of non-syndromic NOA is not explicit as some of them may have unreported pleiotropic effects that cause other health-related issues. When further data on the patients’ phenotype become available, these loci may be later re-classified as syndromic. For example, the STRING analysis of protein–protein interactions linked FANCM that has been currently assigned to the category of isolated NOA with FANCA, XRCC2, CHD7 and SETX implicated in syndromic cases (Fig. 2a).
Despite the list of established and novel NOA genes being rather short (n = 38), the functional enrichment analysis showed highly significant clustering of loci to several Gene Ontology (GO) categories and provided support to their relevance to the NOA phenotype. The five most significantly enriched (FDR < 6.2 × 10–18) biological processes are ‘reproductive process’ (GO:0022414′; 30 of 38 genes), ‘multicellular organismal reproductive process’ (GO:0048609; 23), ‘meiotic nuclear division’ (GO:0140013; 15), ‘gamete generation’ (GO:0007276; 21), and ‘meiosis I’ (GO:0007127; 13) (Fig. 2b; Supplementary Table 4A). Among molecular functions, the top enrichment (FDR = 0.002) was detected for ‘helicase activity’ (GO:0004386; 5 genes) and ‘ATP binding’ (GO:0005524; 12 genes) (Supplementary Table 4B). NOA genes also cluster into cellular components that point to the tight link between the defective genome dynamics and integrity, and congenital spermatogenic failure (e.g., ‘chromosome’ GO:0000794, 12 genes; ‘synaptonemal complex’ GO:0000795, 5 genes) (FDR < 7.6 × 10–6; Supplementary Table 4C). Interestingly, when inspecting the distribution of currently reported NOA genes implicated in meiosis, there is an enrichment of loci contributing to chromosomal pairing and crossing over, whereas there are only few reports on genetic defects in other stages in spermatogenesis. Mapping the involved biological processes and uncovering new ones will promote filling the gaps with ‘missing NOA genes’.
Recent studies have revealed novel protein families implicated in NOA. An example is the FA pathway critical in DNA replication and repair, investigated mostly in the context of cancer development (Niraj et al. 2019). So far, LoF variants in NOA patients have been reported for the FANCA, FANCM, and XRCC2 genes. The demonstration of the involvement of Tudor Domain proteins in spermatogenic failure is an additional interesting finding. Both reported genes, TDRD9 and TDRD7 are responsible for the suppression of LINE1 retrotransposons in the male germline to guarantee its integrity (Tanaka et al. 2011). Supported by the reports on NANOS2 mutations, a novel biological etiology behind NOA was highlighted—defects in the pathway ‘cytoplasmic ribonucleoprotein granule’ (GO:0036464).
Caution in interpreting the published literature
A large proportion of the recently reported monogenic forms of NOA represent homozygous AR mutations identified in consanguineous families (Table 4). A strength of these studies is the availability of several affected and ideally, also non-affected family members. However, as the genomes of inbred subjects have long tracks of homozygosity, this sets a limitation to confidently define the reported genetic variant as a monogenic cause to their condition. Additionally, the reports on the pleiotropic effects of these mutations to cause other clinical symptoms apart from NOA have to be taken with caution until confirmed by an independent source. Genetic ‘matchmaking’ is necessary to establish an explicit link between a particular monogenic defect and NOA. However, it has to be also considered that particular genes may carry ultra-rare mutations found only in one or a few consanguineous families worldwide and may seldom or even never be identified among NOA patients in the outbred populations.
Warningly, in depth assessment of published literature in the field has revealed that ‘all that glitters is not gold’ and not all reported loci immediately qualify to be utilized for diagnostic purposes. A recent careful systematic analysis of the available literature on monogenic causes of male infertility reached the conclusion that only 92/521 (17.6%) reported gene–disease relationships were based on actual adequate scientifically supportive evidence (Oud et al. 2019). Critical assessment of NOA genes for the current review revealed that even in medical genetics databases, OMIM and HPO, there are misclassified loci regarding the evidence to be implicated in monogenic NOA (Supplementary Table 2). An example of an uncertain NOA gene is SOHLH1 (MIM: 618115; spermatogenic failure 32). From the first glance it fits perfectly as a candidate gene for spermatogenic failure, encoding a testis-specific transcription factor that induces spermatogonial differentiation during testicular development in mice (Ballow et al. 2006; Suzuki et al. 2012). Two studies have claimed SOHLH1 variants to cause AD form of NOA (Choi et al. 2010; Nakamura et al. 2017). However, the two highlighted missense variants (rs199935200, rs201142743; Choi et al. 2010) are defined as functionally ‘benign/tolerated’ according to all current in silico prediction tools. Furthermore, the c.346-1G>4A splice variant reported in two Korean and two Japanese NOA patients (Choi et al. 2010; Nakamura et al. 2017) is rather common among the Finns (gnomAD database: minor allele frequency 1.5%). This is not consistent with the scenario of a truly pathogenic mutation causing monogenic dominant form of NOA.
Other frequently claimed loci that still require further supportive evidence as monogenic NOA genes are DMRT1 and Androgen Receptor (AR). DMRT1 encodes a testis-specific transcriptional regulator for mammalian postnatal sex determination and testis differentiation (Kim et al. 2007; Macdonald et al. 2018; Raymond et al. 1998, 2000). Dmrt1 null male mice undergo sex reversal after birth (Matson and Zarkower 2012) and human heterozygous microdeletions involving DMRT1 cause ambiguous genitalia and also sex reversal (Bennett et al. 1993; Tannour-Louet et al. 2010). Deletions encompassing DMRT1 have also been reported in five azoospermia cases from Utah and China in one study (Lopes et al. 2013), but the follow-up investigations have not reached clear conclusions about the monogenic causative link of DMRT1 mutations to NOA (Tewes et al. 2014). The X-linked AR gene encoding a nuclear transcription factor is the most frequently mutated gene in patients with various DSD symptoms (cryptorchidism, hypospadias, micropenis, ambiguous genitalia, 46,XY females) (Eggers et al. 2016). AR mutations cause androgen insensitivity—cellular inability to respond to androgens, and their effect on the phenotype and fertility status depends on the patient’s karyotype, variant type and penetrance (Gottlieb et al. 2012; O'Hara and Smith 2015). So far, confident literature evidence is missing that AR could also be classified as a gene causing monogenic NOA.
Taking together, there is a need for international coordination to develop joint approaches and molecular diagnostic guidelines for the testing of NOA-linked genes and critical interpretation of the identified variants in the clinical context.
Approaches to detect and verify novel genetic causes of NOA
Current state-of-the art methodology in medical genetics discovery research is whole exome sequencing (WES) and with increasing applications, also whole genome sequencing (WGS). The strength of these methods is the detection of all genetic variants in a patient either in the coding region or across the genome. However, as NOA is a phenotype with a limited number of known causative genes, the analysis and interpretation of patients’ WES or WGS data sets require smart approaches. Identification of the true causative relationship between the gene mutation and the phenotype is also dependent on the quality and depth of clinical evaluation.
The classical approach to perform a family-based segregation analysis for the NOA phenotype has biological restrictions. For sporadic NOA cases, confirmation of the claimed genotype–phenotype link by an independent study is an absolute necessity. Recent years have slowly increased the list of confident loci for both isolated and syndromic NOA with back-to-back independent studies reporting novel genes such as TEX11 (in 2015), FANCM (2018), STAG3 (2019), SETX (2019) (Table 2 and references therein). An attractive option to shortlist most relevant genetic variants observed in the patients’ WES/WGS data set is utilization of knowledge from relevant animal models. A recent study by Riera-Escamilla et al. 2019 targeting 175 genes representing the ‘mouse azoospermia’ panel identified two new candidate genes RNF212 and STAG3, and the latter was shortly confirmed by an independent study (van der Bijl et al. 2019). An additional option to narrow down the list of considered variants is a trio-based analysis incorporating parental genetic data that can pinpoint rare AD pathogenic variants inherited from the mother and confirm compound heterozygosity of AR mutations, as well as reveal de novo variants as candidates for the sporadic condition.
Compared to the diversity of genes expressed in human testes or being critical to testicular function, the established NOA genes are restricted to a rather narrow spectrum of functions or include only a few genes from a particular biological pathway. It is highly likely that NOA could be caused by defective genes in biological pathways and protein families that have not been considered so far. Targeted transcriptome/proteome data sets mapping the genes implicated in spermatogenesis, testis development and function represent valuable resources to uncover the specific roles of novel candidate genes and their compatibility to human phenotypes (Chalmel et al. 2012; Darde et al. 2019; Lecluze et al. 2018).
Options to expand the view
A debated issue is whether the genetic causes behind NOA (and its clinical subtypes) are explicit or overlap with other phenotypes of quantitative or even qualitative spermatogenic failure. There is increasing support in the scientific literature to the latter scenario. For example, men with TEX14 pathogenic variants have been reported to exhibit variable testicular histology, either sperm maturation arrest or SCOS (Fakhro et al. 2018; Gershoni et al. 2017). This could be explained by the fact that patients diagnosed with MA and SCOS carry TEX14 missense and LoF variants, respectively. It is generally known that protein truncating variants compared to amino acid substitutions are more likely to cause a severe effect on the phenotype, e.g., missense mutations in CHD7 cause CHH, while LoFs lead to CHARGE syndrome (Marcos et al. 2014). Recently reported biallelic LoF mutations in the FANCM gene were detected in patients with either NOA caused by SCOS or oligoasthenospermia (Kasak et al. 2018; Yin et al. 2019). Also, mutations in the TDRD9 gene were identified not only in azoospermia, but also in a cryptozoospermia case (Arafat et al. 2017). Recessive mutations in the TEX15, SETX, TAF4B, and LHB genes have been reported not only in NOA patients, but also in oligozoospermia cases (Ayhan et al. 2014; Becherel et al. 2019; Okutman et al. 2015; Valdes-Socin et al. 2004). It has to be reminded that the sub-phenotyping and sampling are usually based on the records documented upon the first clinical visit and it is not possible to retrieve retrospective andrological data to properly assess the dynamics of spermatogenic impairment throughout the patient’s life course. In some occasions, also a continuum of phenotypes from the most extreme NOA cases to the milder forms of spermatogenic failure could be possibly explained by pathogenic variants with variable dosage effect within the same gene. For example, SPINK2 homozygosity was shown to cause NOA, whereas heterozygous mutation carriers exhibited oligozoospermia demonstrating an incomplete penetrance of spermatogenic failure (Kherraf et al. 2017). The phenotypic spectrum can also be modulated by the location of the pathogenic variant in the gene as defects in different functional domains may have variable consequences (Roca et al. 2018). These issues have also been recently highlighted and discussed in the context of other genetic disorders (Clark et al. 2019; Kasak et al. 2019a, b).
A further horizon to explore in the genetic landscape to explain idiopathic NOA cases are di- or oligogenic causes that have been highlighted already for a number of genetic diseases, including andrological conditions (Schäffer 2013). The phenotype resulting from digenic recessive mutations in functionally closely linked genes may mimic a typical biallelic monogenic defect (Papadimitriou et al. 2019). Disease-enhancing modifier genes may modulate the penetrance of a major dominant mutation. Approximately 20% of hypogonadotropic hypogonadism cases have been estimated to be expressed due to the combined effect of two or more mutations in different genes (Boehm et al. 2015; Cassatella et al. 2018; Miraoui et al. 2013; Pitteloud et al. 2007; Sykiotis et al. 2010). Recently, several teams have also proposed that digenic/oligogenic effects may explain variable penetrance observed among carriers of mutations in DSD genes (Camats et al. 2018; Robevska et al. 2018; Wang et al. 2018). A systematic analysis in the budding yeast Saccharomyces cerevisiae has demonstrated that both, di- and trigenic interactions are enriched among genes annotated to the same biological process (Kuzmin et al. 2018). Several of the top-interacting gene categories in this lower eukaryotic species are also relevant in the context of NOA, e.g., mRNA and tRNA processing, mitosis and chromosome segregation, DNA replication and repair, transcription and chromatin organization. The protein–protein interaction network analysis of already reported NOA genes showed a highly significant enrichment of active connections and complementary functions among loci implicated in NOA (Figs. 2, 3, 4, Supplementary Tables 4A–C). In perspective, a scenario of di-/oligogenic contribution to the NOA phenotype has to be potentially considered and explored, especially for the genetic defects in spermatogenesis in isolated NOA cases.
Mouse knockout models have revealed 125 genes that are crucial for fertility regardless of the sex (Schimenti and Handel 2018), but not enough attention has been paid to the potential overlap between human male and female reproductive phenotypes. For genes with a conserved role in the gonadal development and gametogenesis in both sexes, pairing up with the knowledge on the genetic causes of female congenital reproductive disorders and infertility may provide further supportive evidence for the candidate NOA gene. Examples of ‘NOA-coupled’ phenotypes reported in female family members carrying the disease mutations are primary ovarian insufficiency (POI) (FANCM, STAG3, MCM8, SYCE1, FANCA, DMC1, MEIOB, SETX), amenorrhea due to missing or late puberty (FSHB, LHB, KISS1R, GNRHR, GNRH1, FGFR1, TACR3, CHD7), and female genital anomalies (NR5A1, WT1, CHD7) (representative references in Supplementary Table 5).
Gradually changing and improving the clinical management
Determination of the precise cause for NOA is critical in the clinical management decisions, assessment of potential accompanying health-related issues of patients and counseling about their options in family planning. The current knowledge indicates that genetic causes and their broader clinical implications behind isolated and syndromic NOA are different (Tables 2, 3 and 4, Fig. 2). Although some attempts have been made to establish a diagnostic gene panel including either all proposed genes implicated in male infertility (including NOA genes) or selected candidate genes for NOA, the current yield of diagnostic mutation detection in the clinical practice is modest (Oud et al. 2017; Tüttelmann et al. 2018). All the accumulated evidence shows that there is no ‘recurrently’ mutated NOA gene that can be straightforwardly included into a widely applicable diagnostic gene panel and daily workup of andrology patients. As the majority of idiopathic NOA cases are most probably explained by yet unknown congenital factors, the key molecular diagnostics is still relying on time-consuming WES/WGS analysis. Due to high expected genetic heterogeneity, there is a need for systematic research and quality criteria for reporting novel NOA-related genes.
However, there are already signs for a ‘light at the end of the tunnel’. Current leadership in translating the findings of monogenic causes of congenital reproductive disorders for the immediate patient benefit concerns rare conditions affecting both sexes, DSD and CHH. For 46,XY subjects with CHH or DSD conditions, the yield of genetic testing is already over 50%. Personalized treatment and management schemes supported by the genetic finding are well established and summarized in the European consensus statements assembled by multidisciplinary expert groups (Boehm et al. 2015; Cools et al. 2018).
What would be the benefit of genetic testing for the rest (majority) of the NOA patients? At first glance, NOA is an extreme and mostly irreversible condition. Still, there is a clear added value in the determination of an exact genetic cause for each patient:
-
Improved management of infertility and counseling about the health risks to the offspring
Genetic finding may assist in making the decision to consider (or not) an invasive and expensive procedure Testicular/Epididymal Sperm Extraction—IntraCytoplasmic Sperm Injection (TESE-ICSI). Recent large-scale analysis of 714 NOA patients reported that only 13.4% of men embarking for TESE eventually become a biological father, although 40.5% have successful sperm retrieval at their first TESE procedure (Vloeberghs et al. 2015). Genetic test result can be highly predictive of the NOA sub-phenotype (e.g., SCOS, MA) and the probability to succeed in the extraction of immature sperm cells from the testis for the ICSI application. Another aspect is capacity of the immature sperm cells to fertilize the egg and give rise to a healthy offspring. This concerns especially congenital developmental defects with variable penetrance and genetic mutations affecting genome integrity, increasing the risk to childhood cancers (Hanson et al. 2018).
-
Clinical management of the patient’s general health
All azoospermia patients exhibit increased risk to various cancers due to defects in biological pathways regulating genomic integrity (Chalmel et al. 2007; Eisenberg et al. 2013; Hanson et al. 2018; Kasak et al. 2018; Krausz et al. 2014; Nagirnaja et al. 2018; Yin et al. 2019). Clinical management of syndromic NOA cases has to be preferentially conducted in collaboration with other medical specialties due to a possible overlap with other health issues, risk to chronic diseases and comorbidities.
-
Counseling the family members about potential congenital health-related issues
Even though NOA usually manifests as a sporadic case in the family, it is important that the family members are included into genetic testing. A special attention has to be paid to female relatives of NOA patients as nearly 50% of the established and novel proposed NOA genes (18/38) are also implicated in either POI, amenorrhea or female genital anomalies (Fig. 3, Supplementary Table 5). Whereas gonadal ambiguities are usually documented at birth and amenorrhea in puberty, POI can also manifest with age and may not be present in a severe form in young women. Additional topic to be considered is the presence of asymptomatic carriers of pathogenic mutations, for example dominant mutations with reduced penetrance or female carriers of X-linked recessive diseases.
Finally, although monogenic causes of NOA are estimated to represent up to 20% of azoospermia cases and only 2–4% of all male infertility patients, the knowledge of underlying genetic defects behind NOA is highly valuable to understand the etiology of spermatogenic failure. First, it could be utilized for the genetic research (and in perspective, in diagnostics) of oligozoospermia, a less severe form of quantitative sperm defects that concerns nearly 70% of cases diagnosed with male factor infertility. Today, 75% of oligozoospermia cases remain unexplained (Punab et al. 2017). Second, uncovering novel loci, protein families and biological pathways implicated in NOA may promote clinical research aiming to develop novel and preferentially non-invasive treatment targets and options for spermatogenic failure.
References
Achermann JC, Ito M, Ito M, Hindmarsh PC, Jameson JL (1999) A mutation in the gene encoding steroidogenic factor-1 causes XY sex reversal and adrenal failure in humans. Nat Genet 22:125–126. https://doi.org/10.1038/9629
Andrews AM, McCartney HJ, Errington TM, D'Andrea AD, Macara IG (2018) A senataxin-associated exonuclease SAN1 is required for resistance to DNA interstrand cross-links. Nat Commun 9:2592. https://doi.org/10.1038/s41467-018-05008-8
Arafat M et al (2017) Mutation in TDRD9 causes non-obstructive azoospermia in infertile men. J Med Genet 54:633–639. https://doi.org/10.1136/jmedgenet-2017-104514
Ayhan O, Balkan M, Guven A, Hazan R, Atar M, Tok A, Tolun A (2014) Truncating mutations in TAF4B and ZMYND15 causing recessive azoospermia. J Med Genet 51:239–244. https://doi.org/10.1136/jmedgenet-2013-102102
Ballow D, Meistrich ML, Matzuk M, Rajkovic A (2006) Sohlh1 is essential for spermatogonial differentiation. Dev Biol 294:161–167. https://doi.org/10.1016/j.ydbio.2006.02.027
Basbous J, Constantinou A (2019) A tumor suppressive DNA translocase named FANCM. Crit Rev Biochem Mol Biol 54:27–40. https://doi.org/10.1080/10409238.2019.1568963
Basciani S et al (2012) Hypogonadism in a patient with two novel mutations of the luteinizing hormone beta-subunit gene expressed in a compound heterozygous form. J Clin Endocrinol Metab 97:3031–3038. https://doi.org/10.1210/jc.2012-1986
Bashamboo A et al (2016) A recurrent p.Arg92Trp variant in steroidogenic factor-1 (NR5A1) can act as a molecular switch in human sex development. Hum Mol Genet 25:5286. https://doi.org/10.1093/hmg/ddw390
Bashamboo A et al (2010) Human male infertility associated with mutations in NR5A1 encoding steroidogenic factor 1. Am J Hum Genet 87:505–512. https://doi.org/10.1016/j.ajhg.2010.09.009
Becherel OJ, Fogel BL, Zeitlin SI, Samaratunga H, Greaney J, Homer H, Lavin MF (2019) Disruption of Spermatogenesis and Infertility in Ataxia with Oculomotor Apraxia Type 2 (AOA2). Cerebellum 18:448–456. https://doi.org/10.1007/s12311-019-01012-w
Ben Khelifa M et al (2018) A MEI1 homozygous missense mutation associated with meiotic arrest in a consanguineous family. Hum Reprod 33:1034–1037. https://doi.org/10.1093/humrep/dey073
Bennett CL, La Spada AR (2018) Senataxin, A novel helicase at the interface of RNA transcriptome regulation and neurobiology: from normal function to pathological roles in motor neuron disease and cerebellar degeneration. Adv Neurobiol 20:265–281. https://doi.org/10.1007/978-3-319-89689-2_10
Bennett CP, Docherty Z, Robb SA, Ramani P, Hawkins JR, Grant D (1993) Deletion 9p and sex reversal. J Med Genet 30:518–520. https://doi.org/10.1136/jmg.30.6.518
Bick D et al (1992) Brief report: intragenic deletion of the KALIG-1 gene in Kallmann's syndrome. N Engl J Med 326:1752–1755. https://doi.org/10.1056/NEJM199206253262606
Boehm U et al (2015) Expert consensus document: European Consensus Statement on congenital hypogonadotropic hypogonadism–pathogenesis, diagnosis and treatment. Nat Rev Endocrinol 11:547–564. https://doi.org/10.1038/nrendo.2015.112
Bolcun-Filas E et al (2009) Mutation of the mouse Syce1 gene disrupts synapsis and suggests a link between synaptonemal complex structural components and DNA repair. PLoS Genet 5:e1000393. https://doi.org/10.1371/journal.pgen.1000393
Bult CJ, Blake JA, Smith CL, Kadin JA, Richardson JE, Mouse Genome Database G (2019) Mouse genome database (MGD) 2019. Nucleic Acids Res 47:D801–D806. https://doi.org/10.1093/nar/gky1056
Camats N, Fernandez-Cancio M, Audi L, Schaller A, Fluck CE (2018) Broad phenotypes in heterozygous NR5A1 46, XY patients with a disorder of sex development: an oligogenic origin? Eur J Hum Genet 26:1329–1338. https://doi.org/10.1038/s41431-018-0202-7
Cardoso-Moreira M et al (2019) Gene expression across mammalian organ development. Nature 571:505–509. https://doi.org/10.1038/s41586-019-1338-5
Cassatella D et al (2018) Congenital hypogonadotropic hypogonadism and constitutional delay of growth and puberty have distinct genetic architectures. Eur J Endocrinol 178:377–388. https://doi.org/10.1530/EJE-17-0568
Catford SR, O'Bryan MK, McLachlan RI, Delatycki MB, Rombauts L (2019) Germ cell arrest associated with aSETX mutation in ataxia oculomotor apraxia type 2. Reprod Biomed Online 38:961–965. https://doi.org/10.1016/j.rbmo.2018.12.042
Chalmel F, Lardenois A, Primig M (2007) Toward understanding the core meiotic transcriptome in mammals and its implications for somatic cancer. Ann N Y Acad Sci 1120:1–15. https://doi.org/10.1196/annals.1411.010
Chalmel F et al (2012) Global human tissue profiling and protein network analysis reveals distinct levels of transcriptional germline-specificity and identifies target genes for male infertility. Hum Reprod 27:3233–3248. https://doi.org/10.1093/humrep/des301
Chen YZ et al (2004) DNA/RNA helicase gene mutations in a form of juvenile amyotrophic lateral sclerosis (ALS4). Am J Hum Genet 74:1128–1135. https://doi.org/10.1086/421054
Choi Y et al (2010) Mutations in SOHLH1 gene associate with nonobstructive azoospermia. Hum Mutat 31:788–793. https://doi.org/10.1002/humu.21264
Clark WT et al (2019) Assessment of predicted enzymatic activity of alpha-N-acetylglucosaminidase variants of unknown significance for CAGI 2016. Hum Mutat 40:1519–1529. https://doi.org/10.1002/humu.23875
Colombo R, Pontoglio A, Bini M (2017) Two novel TEX15 mutations in a family with nonobstructive azoospermia. Gynecol Obstet Invest 82:283–286. https://doi.org/10.1159/000468934
Cools M et al (2018) Caring for individuals with a difference of sex development (DSD): a Consensus Statement. Nat Rev Endocrinol 14:415–429. https://doi.org/10.1038/s41574-018-0010-8
Darde TA et al (2019) The ReproGenomics Viewer: a multi-omics and cross-species resource compatible with single-cell studies for the reproductive science community. Bioinformatics 35:3133–3139. https://doi.org/10.1093/bioinformatics/btz047
Dode C, Hardelin JP (2010) Clinical genetics of Kallmann syndrome. Ann Endocrinol (Paris) 71:149–157. https://doi.org/10.1016/j.ando.2010.02.005
Eggers S et al (2016) Disorders of sex development: insights from targeted gene sequencing of a large international patient cohort. Genome Biol 17:243. https://doi.org/10.1186/s13059-016-1105-y
Eisenberg ML, Betts P, Herder D, Lamb DJ, Lipshultz LI (2013) Increased risk of cancer among azoospermic men. Fertil Steril 100:681–685. https://doi.org/10.1016/j.fertnstert.2013.05.022
Fakhro KA et al (2018) Point-of-care whole-exome sequencing of idiopathic male infertility. Genet Med 20:1365–1373. https://doi.org/10.1038/gim.2018.10
Ferlin A, Rocca MS, Vinanzi C, Ghezzi M, Di Nisio A, Foresta C (2015) Mutational screening of NR5A1 gene encoding steroidogenic factor 1 in cryptorchidism and male factor infertility and functional analysis of seven undescribed mutations. Fertil Steril 104(163–169):e161. https://doi.org/10.1016/j.fertnstert.2015.04.017
Frapsauce C et al (2011) Birth after TESE-ICSI in a man with hypogonadotropic hypogonadism and congenital adrenal hypoplasia linked to a DAX-1 (NR0B1) mutation. Hum Reprod 26:724–728. https://doi.org/10.1093/humrep/deq372
Gershoni M et al (2017) A familial study of azoospermic men identifies three novel causative mutations in three new human azoospermia genes. Genet Med 19:998–1006. https://doi.org/10.1038/gim.2016.225
Gershoni M, Hauser R, Barda S, Lehavi O, Arama E, Pietrokovski S, Kleiman SE (2019) A new MEIOB mutation is a recurrent cause for azoospermia and testicular meiotic arrest. Hum Reprod 34:666–671. https://doi.org/10.1093/humrep/dez016
Gottlieb B, Beitel LK, Nadarajah A, Paliouras M, Trifiro M (2012) The androgen receptor gene mutations database: 2012 update. Hum Mutat 33:887–894. https://doi.org/10.1002/humu.22046
Greenbaum MP, Iwamori T, Buchold GM, Matzuk MM (2011) Germ cell intercellular bridges. Cold Spring Harb Perspect Biol 3:a005850. https://doi.org/10.1101/cshperspect.a005850
Griffin WC, Trakselis MA (2019) The MCM8/9 complex: a recent recruit to the roster of helicases involved in genome maintenance. DNA Repair (Amst) 76:1–10. https://doi.org/10.1016/j.dnarep.2019.02.003
Habedanck R, Stierhof YD, Wilkinson CJ, Nigg EA (2005) The Polo kinase Plk4 functions in centriole duplication. Nat Cell Biol 7:1140–1146. https://doi.org/10.1038/ncb1320
Hanley NA et al (1999) Expression of steroidogenic factor 1 and Wilms' tumour 1 during early human gonadal development and sex determination. Mech Dev 87:175–180. https://doi.org/10.1016/s0925-4773(99)00123-9
Hanson BM, Eisenberg ML, Hotaling JM (2018) Male infertility: a biomarker of individual and familial cancer risk. Fertil Steril 109:6–19. https://doi.org/10.1016/j.fertnstert.2017.11.005
Hardelin JP et al (1992) X chromosome-linked Kallmann syndrome: stop mutations validate the candidate gene. Proc Natl Acad Sci U S A 89:8190–8194. https://doi.org/10.1073/pnas.89.17.8190
Hastie ND (2017) Wilms' tumour 1 (WT1) in development, homeostasis and disease. Development 144:2862–2872. https://doi.org/10.1242/dev.153163
He WB et al (2018) DMC1 mutation that causes human non-obstructive azoospermia and premature ovarian insufficiency identified by whole-exome sequencing. J Med Genet 55:198–204. https://doi.org/10.1136/jmedgenet-2017-104992
Horn HF, Kim DI, Wright GD, Wong ES, Stewart CL, Burke B, Roux KJ (2013) A mammalian KASH domain protein coupling meiotic chromosomes to the cytoskeleton. J Cell Biol 202:1023–1039. https://doi.org/10.1083/jcb.201304004
Jarow JP, Espeland MA, Lipshultz LI (1989) Evaluation of the azoospermic patient. J Urol 142:62–65. https://doi.org/10.1016/s0022-5347(17)38662-7
Jarvi K et al (2015) The workup and management of azoospermic males. Can Urol Assoc J 9:229–235. https://doi.org/10.5489/cuaj.3209
Jungwirth A et al (2012) European Association of Urology guidelines on male infertility: the 2012 update. Eur Urol 62:324–332. https://doi.org/10.1016/j.eururo.2012.04.048
Kaneko Y et al (2015) A high incidence of WT1 abnormality in bilateral Wilms tumours in Japan, and the penetrance rates in children with WT1 germline mutation. Br J Cancer 112:1121–1133. https://doi.org/10.1038/bjc.2015.13
Kasak L et al (2018) Bi-allelic recessive loss-of-function variants in FANCM cause non-obstructive azoospermia. Am J Hum Genet 103:200–212. https://doi.org/10.1016/j.ajhg.2018.07.005
Kasak L et al (2019a) Assessing computational predictions of the phenotypic effect of cystathionine-beta-synthase variants. Hum Mutat 40:1530–1545. https://doi.org/10.1002/humu.23868
Kasak L et al (2019b) CAGI SickKids challenges: assessment of phenotype and variant predictions derived from clinical and genomic data of children with undiagnosed diseases. Hum Mutat 40:1373–1391. https://doi.org/10.1002/humu.23874
Kherraf ZE et al (2017) SPINK2 deficiency causes infertility by inducing sperm defects in heterozygotes and azoospermia in homozygotes. EMBO Mol Med 9:1132–1149. https://doi.org/10.15252/emmm.201607461
Kim S, Bardwell VJ, Zarkower D (2007) Cell type-autonomous and non-autonomous requirements for Dmrt1 in postnatal testis differentiation. Dev Biol 307:314–327. https://doi.org/10.1016/j.ydbio.2007.04.046
Köhler B et al (2011) Analysis of the Wilms' tumor suppressor gene (WT1) in patients 46, XY disorders of sex development. J Clin Endocrinol Metab 96:E1131–1136. https://doi.org/10.1210/jc.2010-2804
Köhler S et al (2019) Expansion of the Human Phenotype Ontology (HPO) knowledge base and resources. Nucleic Acids Res 47:D1018–D1027. https://doi.org/10.1093/nar/gky1105
Krausz C, Riera-Escamilla A (2018) Genetics of male infertility. Nat Rev Urol 15:369–384. https://doi.org/10.1038/s41585-018-0003-3
Krausz C, Hoefsloot L, Simoni M, Tuttelmann F, European Academy of A, European Molecular Genetics Quality N (2014) EAA/EMQN best practice guidelines for molecular diagnosis of Y-chromosomal microdeletions: state-of-the-art 2013. Andrology 2:5–19. https://doi.org/10.1111/j.2047-2927.2013.00173.x
Kuzmin E et al (2018) Systematic analysis of complex genetic interactions. Science. https://doi.org/10.1126/science.aao1729
Lachke SA et al (2011) Mutations in the RNA granule component TDRD7 cause cataract and glaucoma. Science 331:1571–1576. https://doi.org/10.1126/science.1195970
Lecluze E, Jegou B, Rolland AD, Chalmel F (2018) New transcriptomic tools to understand testis development and functions. Mol Cell Endocrinol 468:47–59. https://doi.org/10.1016/j.mce.2018.02.019
Libby BJ, Reinholdt LG, Schimenti JC (2003) Positional cloning and characterization of Mei1, a vertebrate-specific gene required for normal meiotic chromosome synapsis in mice. Proc Natl Acad Sci U S A 100:15706–15711. https://doi.org/10.1073/pnas.2432067100
Lopes AM et al (2013) Human spermatogenic failure purges deleterious mutation load from the autosomes and both sex chromosomes, including the gene DMRT1. PLoS Genet 9:e1003349. https://doi.org/10.1371/journal.pgen.1003349
Lourenco D et al (2009) Mutations in NR5A1 associated with ovarian insufficiency. N Engl J Med 360:1200–1210. https://doi.org/10.1056/NEJMoa0806228
Luo M et al (2013) MEIOB exhibits single-stranded DNA-binding and exonuclease activities and is essential for meiotic recombination. Nat Commun 4:2788. https://doi.org/10.1038/ncomms3788
Macdonald J et al (2018) DMRT1 repression using a novel approach to genetic manipulation induces testicular dysgenesis in human fetal gonads. Hum Reprod 33:2107–2121. https://doi.org/10.1093/humrep/dey289
Maione L, Dwyer AA, Francou B, Guiochon-Mantel A, Binart N, Bouligand J, Young J (2018) Genetics in endocrinology: genetic counseling for congenital hypogonadotropic hypogonadism and Kallmann Syndrome: new challenges in the era of oligogenism and next-generation sequencing. Eur J Endocrinol 178:R55–R80. https://doi.org/10.1530/EJE-17-0749
Maor-Sagie E et al (2015) Deleterious mutation in SYCE1 is associated with non-obstructive azoospermia. J Assist Reprod Genet 32:887–891. https://doi.org/10.1007/s10815-015-0445-y
Marcos S et al (2014) The prevalence of CHD7 missense versus truncating mutations is higher in patients with Kallmann Syndrome than in typical CHARGE patients. J Clin Endocrinol Metab 99:E2138–E2143. https://doi.org/10.1210/jc.2014-2110
Martin CA et al (2014) Mutations in PLK4, encoding a master regulator of centriole biogenesis, cause microcephaly, growth failure and retinopathy. Nat Genet 46:1283–1292. https://doi.org/10.1038/ng.3122
Matson CK, Zarkower D (2012) Sex and the singular DM domain: insights into sexual regulation, evolution and plasticity. Nat Rev Genet 13:163–174. https://doi.org/10.1038/nrg3161
Miraoui H et al (2013) Mutations in FGF17, IL17RD, DUSP6, SPRY4, and FLRT3 are identified in individuals with congenital hypogonadotropic hypogonadism. Am J Hum Genet 92:725–743. https://doi.org/10.1016/j.ajhg.2013.04.008
Miyamoto T et al (2016) A PLK4 mutation causing azoospermia in a man with Sertoli cell-only syndrome. Andrology 4:75–81. https://doi.org/10.1111/andr.12113
Moreira MC et al (2004) Senataxin, the ortholog of a yeast RNA helicase, is mutant in ataxia-ocular apraxia 2. Nat Genet 36:225–227. https://doi.org/10.1038/ng1303
Muscatelli F et al (1994) Mutations in the DAX-1 gene give rise to both X-linked adrenal hypoplasia congenita and hypogonadotropic hypogonadism. Nature 372:672–676. https://doi.org/10.1038/372672a0
Nachtigal MW, Hirokawa Y, Enyeart-VanHouten DL, Flanagan JN, Hammer GD, Ingraham HA (1998) Wilms' tumor 1 and Dax-1 modulate the orphan nuclear receptor SF-1 in sex-specific gene expression. Cell 93:445–454. https://doi.org/10.1016/s0092-8674(00)81172-1
Nagirnaja L, Rull K, Uuskula L, Hallast P, Grigorova M, Laan M (2010) Genomics and genetics of gonadotropin beta-subunit genes: Unique FSHB and duplicated LHB/CGB loci. Mol Cell Endocrinol 329:4–16. https://doi.org/10.1016/j.mce.2010.04.024
Nagirnaja L, Aston KI, Conrad DF (2018) Genetic intersection of male infertility and cancer. Fertil Steril 109:20–26. https://doi.org/10.1016/j.fertnstert.2017.10.028
Nakamura S et al (2017) Next-generation sequencing for patients with non-obstructive azoospermia: implications for significant roles of monogenic/oligogenic mutations. Andrology 5:824–831. https://doi.org/10.1111/andr.12378
Nguyen NMP et al (2018) Causative mutations and mechanism of androgenetic hydatidiform moles. Am J Hum Genet 103:740–751. https://doi.org/10.1016/j.ajhg.2018.10.007
Niraj J, Farkkila A, D'Andrea AD (2019) The Fanconi anemia pathway in cancer. Annu Rev Cancer Biol 3:457–478. https://doi.org/10.1146/annurev-cancerbio-030617-050422
O'Hara L, Smith LB (2015) Androgen receptor roles in spermatogenesis and infertility. Best Pract Res Clin Endocrinol Metab 29:595–605. https://doi.org/10.1016/j.beem.2015.04.006
Okutman O et al (2015) Exome sequencing reveals a nonsense mutation in TEX15 causing spermatogenic failure in a Turkish family. Hum Mol Genet 24:5581–5588. https://doi.org/10.1093/hmg/ddv290
Olesen IA, Andersson AM, Aksglaede L, Skakkebaek NE, Rajpert-de Meyts E, Joergensen N, Juul A (2017) Clinical, genetic, biochemical, and testicular biopsy findings among 1,213 men evaluated for infertility. Fertil Steril 107(74–82):e77. https://doi.org/10.1016/j.fertnstert.2016.09.015
Oud MS et al (2017) Validation and application of a novel integrated genetic screening method to a cohort of 1,112 men with idiopathic azoospermia or severe oligozoospermia. Hum Mutat 38:1592–1605. https://doi.org/10.1002/humu.23312
Oud MS, Volozonoka L, Smits RM, Vissers L, Ramos L, Veltman JA (2019) A systematic review and standardized clinical validity assessment of male infertility genes. Hum Reprod 34:932–941. https://doi.org/10.1093/humrep/dez022
Papadimitriou S et al (2019) Predicting disease-causing variant combinations. Proc Natl Acad Sci U S A 116:11878–11887. https://doi.org/10.1073/pnas.1815601116
Pitteloud N, Hayes FJ, Dwyer A, Boepple PA, Lee H, Crowley WF Jr (2002) Predictors of outcome of long-term GnRH therapy in men with idiopathic hypogonadotropic hypogonadism. J Clin Endocrinol Metab 87:4128–4136. https://doi.org/10.1210/jc.2002-020518
Pitteloud N et al (2007) Digenic mutations account for variable phenotypes in idiopathic hypogonadotropic hypogonadism. J Clin Invest 117:457–463. https://doi.org/10.1172/JCI29884
Posey JE et al (2017) Resolution of disease phenotypes resulting from multilocus genomic variation. N Engl J Med 376:21–31. https://doi.org/10.1056/NEJMoa1516767
Posey JE et al (2019) Insights into genetics, human biology and disease gleaned from family based genomic studies. Genet Med 21:798–812. https://doi.org/10.1038/s41436-018-0408-7
Practice Committee of the American Society for Reproductive Medicine in collaboration with the Society for Male Reproduction and Urology (2018) Evaluation of the azoospermic male: a committee opinion. Fertil Steril 109:777–782. https://doi.org/10.1016/j.fertnstert.2018.01.043
Punab M et al (2017) Causes of male infertility: a 9-year prospective monocentre study on 1737 patients with reduced total sperm counts. Hum Reprod 32:18–31. https://doi.org/10.1093/humrep/dew284
Raymond CS, Shamu CE, Shen MM, Seifert KJ, Hirsch B, Hodgkin J, Zarkower D (1998) Evidence for evolutionary conservation of sex-determining genes. Nature 391:691–695. https://doi.org/10.1038/35618
Raymond CS, Murphy MW, O'Sullivan MG, Bardwell VJ, Zarkower D (2000) Dmrt1, a gene related to worm and fly sexual regulators, is required for mammalian testis differentiation. Genes Dev 14:2587–2595. https://doi.org/10.1101/gad.834100
Reynolds A et al (2013) RNF212 is a dosage-sensitive regulator of crossing-over during mammalian meiosis. Nat Genet 45:269–278. https://doi.org/10.1038/ng.2541
Riera-Escamilla A et al (2019) Sequencing of a 'mouse azoospermia' gene panel in azoospermic men: identification of RNF212 and STAG3 mutations as novel genetic causes of meiotic arrest. Hum Reprod 34:978–988. https://doi.org/10.1093/humrep/dez042
Robevska G et al (2018) Functional characterization of novel NR5A1 variants reveals multiple complex roles in disorders of sex development. Hum Mutat 39:124–139. https://doi.org/10.1002/humu.23354
Roca I, Fernández-Marmiesse A, Gouveia S, Segovia M, Couce ML (2018) Prioritization of variants detected by next generation sequencing according to the mutation tolerance and mutational architecture of the corresponding genes. Int J Mol Sci 19(6):1584. https://doi.org/10.3390/ijms19061584
Saba R, Kato Y, Saga Y (2014) NANOS2 promotes male germ cell development independent of meiosis suppression. Dev Biol 385:32–40. https://doi.org/10.1016/j.ydbio.2013.10.018
Schäffer AA (2013) Digenic inheritance in medical genetics. J Med Genet 50:641–652. https://doi.org/10.1136/jmedgenet-2013-101713
Schimenti JC, Handel MA (2018) Unpackaging the genetics of mammalian fertility: strategies to identify the "reproductive genome". Biol Reprod 99:1119–1128. https://doi.org/10.1093/biolre/ioy133
Seabra CM et al (2015) The mutational spectrum of WT1 in male infertility. J Urol 193:1709–1715. https://doi.org/10.1016/j.juro.2014.11.004
Segaloff DL (2009) Diseases associated with mutations of the human lutropin receptor. Prog Mol Biol Transl Sci 89:97–114. https://doi.org/10.1016/S1877-1173(09)89004-2
Sha Y et al (2018) A novel TEX11 mutation induces azoospermia: a case report of infertile brothers and literature review. BMC Med Genet 19:63. https://doi.org/10.1186/s12881-018-0570-4
Shaheen R, Al Tala S, Almoisheer A, Alkuraya FS (2014) Mutation in PLK4, encoding a master regulator of centriole formation, defines a novel locus for primordial dwarfism. J Med Genet 51:814–816. https://doi.org/10.1136/jmedgenet-2014-102790
Shimamura A, Alter BP (2010) Pathophysiology and management of inherited bone marrow failure syndromes. Blood Rev 24:101–122. https://doi.org/10.1016/j.blre.2010.03.002
Shoji M et al (2009) The TDRD9-MIWI2 complex is essential for piRNA-mediated retrotransposon silencing in the mouse male germline. Dev Cell 17:775–787. https://doi.org/10.1016/j.devcel.2009.10.012
Soumillon M et al (2013) Cellular source and mechanisms of high transcriptome complexity in the mammalian testis. Cell Rep 3:2179–2190. https://doi.org/10.1016/j.celrep.2013.05.031
Soraggi S, Riera M, Rajpert‑De Meyts E, Schierup MH, Almstrup K (2020) Evaluating genetic causes of azoospermia: What can we learn from a complex cellular structure and single‑cell transcriptomics of the human testis? Hum Genet. https://doi.org/10.1007/s00439-020-02116-8
Stephen EH, Chandra A (2006) Declining estimates of infertility in the United States: 1982–2002. Fertil Steril 86:516–523. https://doi.org/10.1016/j.fertnstert.2006.02.129
Suntharalingham JP, Buonocore F, Duncan AJ, Achermann JC (2015) DAX-1 (NR0B1) and steroidogenic factor-1 (SF-1, NR5A1) in human disease. Best Pract Res Clin Endocrinol Metab 29:607–619. https://doi.org/10.1016/j.beem.2015.07.004
Suzuki H, Ahn HW, Chu T, Bowden W, Gassei K, Orwig K, Rajkovic A (2012) SOHLH1 and SOHLH2 coordinate spermatogonial differentiation. Dev Biol 361:301–312. https://doi.org/10.1016/j.ydbio.2011.10.027
Swee DS, Quinton R (2019) Managing congenital hypogonadotrophic hypogonadism: a contemporary approach directed at optimizing fertility and long-term outcomes in males. Ther Adv Endocrinol Metab 10:2042018819826889. https://doi.org/10.1177/2042018819826889
Sykiotis GP et al (2010) Oligogenic basis of isolated gonadotropin-releasing hormone deficiency. Proc Natl Acad Sci U S A 107:15140–15144. https://doi.org/10.1073/pnas.1009622107
Szklarczyk D et al (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613. https://doi.org/10.1093/nar/gky1131
Tan YQ et al (2019) Loss-of-function mutations in TDRD7 lead to a rare novel syndrome combining congenital cataract and nonobstructive azoospermia in humans. Genet Med 21:1209–1217. https://doi.org/10.1038/gim.2017.130
Tanaka T et al (2011) Tudor domain containing 7 (Tdrd7) is essential for dynamic ribonucleoprotein (RNP) remodeling of chromatoid bodies during spermatogenesis. Proc Natl Acad Sci USA 108:10579–10584. https://doi.org/10.1073/pnas.1015447108
Tannour-Louet M et al (2010) Identification of de novo copy number variants associated with human disorders of sexual development. PLoS ONE 5:e15392. https://doi.org/10.1371/journal.pone.0015392
Tapanainen JS, Aittomaki K, Min J, Vaskivuo T, Huhtaniemi IT (1997) Men homozygous for an inactivating mutation of the follicle-stimulating hormone (FSH) receptor gene present variable suppression of spermatogenesis and fertility. Nat Genet 15:205–206. https://doi.org/10.1038/ng0297-205
Tenenbaum-Rakover Y et al (2015) Minichromosome maintenance complex component 8 (MCM8) gene mutations result in primary gonadal failure. J Med Genet 52:391–399. https://doi.org/10.1136/jmedgenet-2014-102921
Tewes AC, Ledig S, Tuttelmann F, Kliesch S, Wieacker P (2014) DMRT1 mutations are rarely associated with male infertility. Fertil Steril 102(816–820):e813. https://doi.org/10.1016/j.fertnstert.2014.05.022
Thorslund T, Esashi F, West SC (2007) Interactions between human BRCA2 protein and the meiosis-specific recombinase DMC1. EMBO J 26:2915–2922. https://doi.org/10.1038/sj.emboj.7601739
Tock AJ, Henderson IR (2018) Hotspots for initiation of meiotic recombination. Front Genet 9:521. https://doi.org/10.3389/fgene.2018.00521
Tournaye H, Krausz C, Oates RD (2017) Novel concepts in the aetiology of male reproductive impairment. Lancet Diabetes Endocrinol 5:544–553. https://doi.org/10.1016/S2213-8587(16)30040-7
Tsui V, Crismani W (2019) The Fanconi anemia pathway and fertility. Trends Genet 35:199–214. https://doi.org/10.1016/j.tig.2018.12.007
Tüttelmann F, Werny F, Cooper TG, Kliesch S, Simoni M, Nieschlag E (2011) Clinical experience with azoospermia: aetiology and chances for spermatozoa detection upon biopsy. Int J Androl 34:291–298. https://doi.org/10.1111/j.1365-2605.2010.01087.x
Tüttelmann F, Ruckert C, Ropke A (2018) Disorders of spermatogenesis: perspectives for novel genetic diagnostics after 20 years of unchanged routine. Med Genet 30:12–20. https://doi.org/10.1007/s11825-018-0181-7
Uhlen M et al (2015) Proteomics Tissue-based map of the human proteome. Science 347:1260419. https://doi.org/10.1126/science.1260419
Valdes-Socin H et al (2004) Hypogonadism in a patient with a mutation in the luteinizing hormone beta-subunit gene. N Engl J Med 351:2619–2625. https://doi.org/10.1056/NEJMoa040326
Vockel M, Riera-Escamilla A, Tüttelmann F, Krausz C (2019) The X chromosome and male infertility. Hum Genet. https://doi.org/10.1007/s00439-019-02101-w
van der Bijl N et al (2019) Mutations in the stromal antigen 3 (STAG3) gene cause male infertility due to meiotic arrest. Hum Reprod 34:2112–2119. https://doi.org/10.1093/humrep/dez204
Vloeberghs V, Verheyen G, Haentjens P, Goossens A, Polyzos NP, Tournaye H (2015) How successful is TESE-ICSI in couples with non-obstructive azoospermia? Hum Reprod 30:1790–1796. https://doi.org/10.1093/humrep/dev139
Wang XN et al (2013) The Wilms tumor gene, Wt1, is critical for mouse spermatogenesis via regulation of sertoli cell polarity and is associated with non-obstructive azoospermia in humans. PLoS Genet 9:e1003645. https://doi.org/10.1371/journal.pgen.1003645
Wang H et al (2018) Next-generation sequencing reveals genetic landscape in 46, XY disorders of sexual development patients with variable phenotypes. Hum Genet 137:265–277. https://doi.org/10.1007/s00439-018-1879-y
WHO (2010) WHO laboratory manual for the examination and processing of human semen, 5th edn. World Health Organization, Geneva, p 2010
Winters T, McNicoll F, Jessberger R (2014) Meiotic cohesin STAG3 is required for chromosome axis formation and sister chromatid cohesion. EMBO J 33:1256–1270. https://doi.org/10.1002/embj.201387330
Xu J et al (2017) A novel functional variant in Wilms' Tumor 1 (WT1) is associated with idiopathic non-obstructive azoospermia. Mol Reprod Dev 84:222–228. https://doi.org/10.1002/mrd.22768
Yan W et al (2010) Zmynd15 encodes a histone deacetylase-dependent transcriptional repressor essential for spermiogenesis and male fertility. J Biol Chem 285:31418–31426. https://doi.org/10.1074/jbc.M110.116418
Yang F et al (2008) Meiotic failure in male mice lacking an X-linked factor. Genes Dev 22:682–691. https://doi.org/10.1101/gad.1613608
Yang F et al (2015) TEX11 is mutated in infertile men with azoospermia and regulates genome-wide recombination rates in mouse. EMBO Mol Med 7:1198–1210. https://doi.org/10.15252/emmm.201404967
Yang Y et al (2018) XRCC2 mutation causes meiotic arrest, azoospermia and infertility. J Med Genet 55:628–636. https://doi.org/10.1136/jmedgenet-2017-105145
Yang X et al (2018) Homozygous nonsense mutation Trp28X in the LHB gene causes male hypogonadism. J Assist Reprod Genet 35:913–919. https://doi.org/10.1007/s10815-018-1133-5
Yatsenko AN et al (2015) X-linked TEX11 mutations, meiotic arrest, and azoospermia in infertile men. N Engl J Med 372:2097–2107. https://doi.org/10.1056/NEJMoa1406192
Yin H et al (2019) A homozygous FANCM frameshift pathogenic variant causes male infertility. Genet Med 21:62–70. https://doi.org/10.1038/s41436-018-0015-7
Young J et al (2019) Clinical Management of Congenital Hypogonadotropic Hypogonadism. Endocr Rev 40:669–710. https://doi.org/10.1210/er.2018-00116
Zare-Abdollahi D et al (2015) Mutational screening of the NR5A1 in azoospermia. Andrologia 47:395–401. https://doi.org/10.1111/and.12274
Zimmermann C et al (2015) Research resource: the dynamic transcriptional profile of sertoli cells during the progression of spermatogenesis. Mol Endocrinol 29:627–642. https://doi.org/10.1210/me.2014-1356
Acknowledgements
M. Laan and L. Kasak are supported by Estonian Research Council grant IUT34-12.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kasak, L., Laan, M. Monogenic causes of non-obstructive azoospermia: challenges, established knowledge, limitations and perspectives. Hum Genet 140, 135–154 (2021). https://doi.org/10.1007/s00439-020-02112-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-020-02112-y