Abstract
Non-obstructive azoospermia (NOA) and obstructive azoospermia (OA) are two common causes of infertility that affect a considerable number of men. However, few studies were performed to understand the molecular etiology of these disorders. Studies based on bioinformatics and genetic analyses in recent years, however, have yielded insightful information and have identified a number of genes that are involved in these disorders. In this review, we briefly summarize and evaluate these findings. We also discuss findings based on epigenetic modifications of sperm DNAs that affect a number of genes pertinent to NOA and OA. The information summarized in this Chapter should be helpful to investigators in future functional studies of NOA and OA.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Infertility is an emerging global health issue facing married couples [1, 2]. More than 10% of couples are unable to conceive their own child(ren), and approximately half of these cases are contributed by men [3, 4]. Azoospermia, defined as the complete absence of sperm from the ejaculate, is a major factor in male infertility. Azoospermia can be categorized into obstructive azoospermia (OA), comprising about 40% of the cases, and non-obstructive azoospermia (NOA), constitutes the other 60%. Obstruction in the ductal system is the cause of OA [5]. For NOA, the cause is the failure of the testicles to produce mature sperm so that no sperm are found in the ejaculate. NOA can be classified clinically into four types: NOA-I, no spermatozoa; NOA-II, no spermatids; NOA-III, no spermatocytes; and NOA-SCO (Sertoli cell-only), no spermatogenic cells of any types; in the ejaculate [6]. The definition of these four types of NOA is based on diagnostic analysis of the testes, hormonal analysis (e.g., FSH, testosterone) in plasma or serum, and physical examination [7]. OA patients are characterized by an obstructed flow of spermatozoa along the male genital tract. However, OA patients have normal spermatogenesis in the testis. The etiology of NOA is more complex, which can be divided into primary and secondary testicular failure. The primary testicular failure refers to pathology localized to the testis, including chromosomal/genetic abnormalities, Klinefelter’s syndrome, testicular tumor, undescended testes, and others. The secondary testis failure is caused by the abnormal secretion of gonadotropins from the pituitary gland, which contribute to insufficient stimulation of Leydig cells and Sertoli cells in the testis to maintain normal spermatogenesis. As a result, Sertoli cells are unable to secrete adequate factors including hormones (e.g., estradiol-17ß), and Leydig cells fail to provide enough local testosterone to support normal spermatogenesis [8, 9].
Genetic Mutation(s)
The diagnostic tools of male infertility are limited, and clinicians are racing to identify more predictive biomarkers. Several crucial fundamental laboratory analyses, including semen analysis, anti-sperm antibodies, Y-microdeletion analysis, Karyotype analysis, microarray technologies, and endocrine-based laboratory investigations are currently being used to support clinicians in diagnosing, categorizing, and treating male factor infertility [10]. To date, seminal fluid, containing the highest concentration of biomolecules, is often used as a standard biomarker for the evaluation of male infertility, including DNA fragmentation index, anti-sperm antibody and sperm fluorescence in situ hybridization (Table 1) [16]. Azoospermia often involves gene mutations. However, it is difficult to determine the genetic component of male infertility because more than 2000 genes have been shown to be involved in human spermatogenesis [17]. In recent years, novel high-throughput approaches have developed to study pertinent genes that are mutated in azoospermic men, including genomic hybridization-arrays (arrays-CGH), genome wide association studies (GWAS) and next generation sequencing (NGS). Whole genome sequencing (WGS), an advanced technology, has brought an unprecedented opportunity to explore the genetic basis of infertility, which also makes it possible to perform studies of large cohorts of patients [18]. Whole exomes sequencing is also widely used to identify potential pathogenic and novel genes pertinent to infertile males. Copy number variation (CNV) and single nucleotide polymorphisms are also the risk factors associated with male infertility [19]. Nonetheless, relatively few studies have provided functional and biological evidence to validate the variants as pathogenic genes.
Accumulating evidence has shown that TEX11 and TEX15 are mutated in infertile men of NOA and also meiotic arrest, analogous to the mutation mouse model [20, 21]. In one study of 40 Japanese patients with idiopathic NOA by conducting sequence analysis of 25 known disease-associated genes using next-generation sequencing and genome-wide copy-number analysis. The results revealed that oligogenic mutations, including SOHLH1 and TEX11 and monogenic mutation, which accounted for more than 10% of cases of idiopathic NOA (Table 2). Furthermore, submicroscopic copy-number variations (CNVs) on the autosomes and X chromosome may contribute to NOA, which require additional validation [22]. In 2014, truncating mutations in TAF4B and ZMYND15 were reported in the azoospermic brothers of two families by exome sequencing. The two genes were shown to have an important role in spermatogenesis in mice, and they were also the first genes identified in the azoospermic men [24]. Also, using exome analysis and Sanger sequencing, a splice site mutation in SYCE1 was found in two NOA patients in a consanguineous Iranian Jewish family [25]. SYCE1, a component of the central element of the synaptonemal complex, was shown to be a crucial interacting/regulatory protein between proteins SYCE2 and RAD51, and its deletion led to meiosis arrest in SYCE1 null mouse [26]. He et al. discovered DMC1, a meiosis-related gene, was crucial to support meiosis since its missense mutation led to meiotic arrest at the zygotene stage during human spermatogenesis. This DMC1 mutation was identified from the male patient’s family by whole-exome sequencing [27]. Cystic fibrosis transmembrane conductance regulator (CFTR), a crucial gene in supporting spermatogenesis, was recently found to be involved in NOA [28]. In another study of five azoospermic infertile NOA patients, recessive deleterious mutation in TDRD9 was identified which contributed to sperm maturation arrest. Similar clinical phenotype was also observed in the Tdrd9 global knockout mice [29]. Additionally, mutations and polymorphisms in HIWI2 were detected in idiopathic NOA, which were crucial for piRNA biogenesis and function in supporting spermatogenesis [47]. piRNA pathway is a fundamental component of spermatogenesis which ensures male fertility and genome integrity [48]. Furthermore, PABPC1, PABPC3, and EPAB, the poly(A)-binding protein genes, are differentially expressed in NOA patients when compared to normal men, implying their involvement in NOA [34]. It is known that development of the spermatogonial stem cells (hSSC) is essential for human spermatogenesis, and FOXP3 pathogenic variants affected the proliferation and apoptosis of hSSC, causing male infertility [30]. Similar to FOXP3, a reduced expression of PAK was noted in NOA patients, which thus inhibited apoptosis and promoted proliferation of hSSC through PDK1/KDR/ZNF367 and ERK1/2 and AKT pathways [31]. Also, genetic variants in STAG3, a meiosis-specific protein, has been reported to cause meiotic arrest in both male and female mice; however, its genetic variants in humans led to premature ovarian failure in women, but not in infertility in men [49] (Table 2).
DNA Methylation
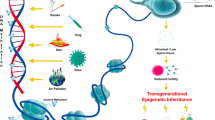
Emerging evidence has shown that DNA methylation, sperm-borne and epigenetic abnormalities in chromatin dynamics, may contribute to male infertility [50]. Epigenetic modifications take place frequently during spermatogenesis, including large-scale demethylation of the genome to allow for sex-specific resetting by DNA methylation and histone modifications. DNA methylation, a heritable epigenetic modification, and a widely investigated epigenetic marker plays an essential role in regulating genes during human spermatogenesis, which mainly takes place in the fifth position of cytosine bases and followed by guanine (CpG). Male germ cells acquire DNA methylation beginning at mitotic and meiotic germ cells, and it is completed at the stage of the pachytene phase of meiosis [51, 52]. Abnormal DNA methylation in sperm may contribute to male infertility and pass onto offspring, who may become more susceptible of developing illnesses later on in life [53]. Several studies have reported that there is a significant difference on DNA methylation levels between normal and infertile men, which also leads to lower sperm count, reduced semen volume and lower sperm progressive motility [54]. Studies have also shown that DNA methylation can be induced by environmental factors, including exposure to endocrine disrupting chemicals [e.g., perfluorooctanesulfonate (PFOS), phthalates) and heavy metals (e.g., cadmium, mercury, lead), nutritional status, air pollution, smoking, and unhealthy lifestyle, since these are contributing factors to gene-specific and global DNA methylation [55, 56]. For example, cadmium, an environment toxicant that exists as CdCl2 in the environment has been shown to reduce fetal growth by hypomethylation of the PCDHAC1 promoter region, which leads to a positive expression of PCDHAC1 [57]. Air pollutants, containing massive different environmental exposures, was also found to alter DNA methylation levels of the genes encoding the mitogen-activated protein kinase (MAPK) regulatory network and other blood-based proteins [58, 59]. Male infertility is also influenced by smoking via epigenetic pathways [60]. Smoking was also shown to have a strong influence on DNA methylation, which alters the CpG methylation patterns in the regions of MAPK8IP and TKR genes, leading to reduced sperm count, motility, and defects in sperm morphology [61]. In some azoospermia sperm samples, there was a considerable increase in the levels of KCNQ1OT1 (KCNQ1 Opposite Strand/Antisense Transcript 1, is a long non-coding RNA gene found in the KCNQ1 locus), compared to the normal group [62]. Also, global methylation level of sperm DNA did not affect the pregnancy rate in IVF, but it affected embryo development when global DNA methylation level was below a threshold value [63]. Furthermore, smokers displayed hypomethylation of reproductive related genes, including Nme2, Trim27, ICR, H19, SNRPN, Sort and Pebp1, which negatively impeded spermatogenesis and sperm motility [64,65,66,67].
Also, distinctive DNA methylation modifications are noted in promoters and repeat elements during spermatogenesis [68]. As a crucial transcription factor for mitochondrial biogenesis, nuclear respiratory factor (NRF)-1 cooperates with DNA methylation to directly regulate the expression of various germ cell-specific genes, including Asz1 [69]. The hypermethylation at the promoter of SOX30 contributes to its silencing of expression in NOA, and the decreasing level of SOX30 is related to the severity of NOA disease. Furthermore, the absence of Sox30 in mice led to male infertility with a complete lack of spermatozoa, which impaired testis development and spermatogenesis. However, Sox30 does not influence ovary development and female fertility [6]. Hypermethylation of the MAEL promoter increased the expression of the transposable element LINE-1, leading to a decrease in the appearance of MAEL, and the methylation of the MALE promoter in infertility men correlates with the severity of spermatogenic failure [70]. On the other hand, aberrant methylation of the GTF2A1L promoter did not affect fertilization rates, but its expression was reduced in NOA patients.
A recent case-control study in NOA and OA patients by investigating the differences and conservations in DNA methylation based on genome-wide DNA methylation and bulk RNA-Seq between these groups for transcriptome profiling. These results have shown that the genome modification of testicular cells from NOA patients is disordered, and the reproductive related gene expression is considerably different [71]. Four functional regions (CGI, gene body, promoter, and TEs) were identified and it was noted that the NOA patient’s entropy values in these regions were considerably lower versus the OA group. Meanwhile, the methylation level of the OA patients was lower, and the gene expression level was higher than the NOA patients. Likewise, the methylation level of Dazl gene, an RNA binding protein deleted in azoospermia [72], in OA patients was lower than that of the NOA patients. A series of reproductive genes, including testis and ovary-specific PAZ domain gene1 (Topaz1), the nuclear receptor NR5A1, and the vertebrate-conserved RNA-binding protein gene DND1, all displayed lower NDA methylation level and higher gene expression level in OA patients compared to NOA, which are related to the development of spermatogonia that may contribute to male infertility. Transposons are often silenced by DNA methylation, and some functional transposons exhibited higher enrichment scores in NOA patients, including ALU, ERV1, HAT, and MIR. These findings are summarized in Table 3.
Chromosomal Aberrations and Y chromosome (Yq) Microdeletions
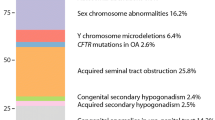
Genetic disorders are one of the primary causes of azoospermia, including chromosomal abnormalities, monogenic disorder, multifactorial genetic diseases, and epigenetic disorders, which also constitute the genetic basis of reproductive failure [73]. The prevalence of chromosomal aberrations in azoospermic patients was between 15% and 25% [28, 74,75,76]. Klinefelter syndrome and its variants (47, XXY and mosaics 46, XY/47, XXY) is the most frequent chromosomal anomaly in NOA, whereas oligozoospermia is more prevalent in men with autosomal structural defects [77]. Klinefelter syndrome (KS), identified over 70 years ago, also remains one of the prevalent causes of infertility, which is typified by small testes, hypogonadotropic hypogonadism, and cognitive impairment. As a syndromic disease, KS is associated with cardiovascular abnormalities, autoimmune diseases, metabolic disorders and cognitive or psychiatric health problems, which may also increase the risk of death [78, 79]. The average prevalence of KS is 152 per 100,000 newborn males, based on several lager cytogenetic chromosome surveys in countries around the world as noted in 2017 [80]. However, KS is often insufficiently diagnosed, and treatment is limited mostly to testosterone therapy, which overcomes some but not all comorbidities [77]. Y chromosome harbors a large number of genes that are necessary for testis development and function. The azoospermia factor (AZF) deletions impaired spermatogenesis, which is also a major molecular cause of male infertility [81]. Meanwhile, Y chromosome (Yq) microdeletions constitute a significant cause of male infertility. European infertile men are less susceptible to Yq microdeletions compared to East Asian and Americans infertile men. Y chromosome is composed of a short arm (Yp), a long arm (Yq), and two pseudo autosomal regions (PARs), which are separated by a centromere [82]. Studies have demonstrated that the deletion of human PARs in men reduced recombination in PARs, leading to sterility [83, 84]. This thus increases the frequency of sex chromosome aneuploidy in sperm, contributing to X-chromosome monosomy (Turner syndrome) or XXY (Klinefelter syndrome) in the offspring [85, 86]. X-chromosome monosomy (Turner syndrome), accounted for approximately 2% of all conceptions, is due to a partial or total loss of the second sexual chromosome, leading to an abnormal development phenotype, including typical dysmorphic stigmata, sexual infantilism, short stature, and partial organs and metabolic abnormalities [87]. However, the TS phenotype may be associated with a genomic imbalance from the absence of genes linked to the second sex chromosome and altered regulation of gene expression that triggered by epigenetic factors. Thus, both copy number variations and epigenetic changes are crucial contributing factors in the TS phenotype [87]. Epigenetic alterations in pericentromeric heterochromatin may also contribute to reconstructing of chromatin conformation, leading to chromosomes that have defects in their ability to align, attach to mitotic spindle fibers, and segregate during mitosis [88].
Concluding Remarks and Future Perspectives
DNA methylation as an epigenetic marker which plays an important role in male spermatogenesis. In this chapter, we discuss the importance of DNA methylation and gene expression that contribute to NOA and OA. We also summarized the several important reproductive genes in NOA and OA that show different DNA methylation and expression level. Also, we discuss findings based on the use of advanced technology to detect genetic mutations in NOA vs. OA that lead to male infertility. More studies are needed by increasing the sample sizes to integrate multiple epigenomic and RNA-seq analysis, which will help in the identification of epigenetic markers and genes pertinent to the regulation of fertility and infertility. Future investigations using single cell (sc) RNA-seq, scATAC-seq and epigenomics will be important to define the etiology and pathogenesis in NOA and OA [89, 90].
References
de Kretser, D. M. (1997). Male infertility. Lancet (London, England), 349(9054), 787–790.
Lotti, F., & Maggi, M. (2018). Sexual dysfunction and male infertility. Nature Reviews. Urology, 15(5), 287–307.
Maduro, M. R., & Lamb, D. J. (2002). Understanding new genetics of male infertility. The Journal of Urology, 168(5), 2197–2205.
Huynh, T., Mollard, R., & Trounson, A. (2002). Selected genetic factors associated with male infertility. Human Reproduction Update, 8(2), 183–198.
Practice Committee of American Society for Reproductive Medicine in collaboration with Society for Male Reproduction and Urology. (2008). The management of infertility due to obstructive azoospermia. Fertility and Sterility, 90(5 Suppl), S121–S124.
Han, F., et al. (2020). Epigenetic inactivation of SOX30 is associated with male infertility and offers a therapy target for non-obstructive azoospermia. Molecular Therapy--Nucleic Acids, 19, 72–83.
Schlegel, P. N. (2004). Causes of azoospermia and their management. Reproduction, Fertility, and Development, 16(5), 561–572.
Behre, H. M., et al. (2000). Primary testicular failure. In K. R. Feingold et al. (Eds.), Endotext. MDText.com.
Krausz, C., Escamilla, A. R., & Chianese, C. (2015). Genetics of male infertility: From research to clinic. Reproduction (Cambridge, England), 150(5), R159–R174.
Kovac, J. R., Pastuszak, A. W., & Lamb, D. J. (2013). The use of genomics, proteomics, and metabolomics in identifying biomarkers of male infertility. Fertility and Sterility, 99(4), 998–1007.
Turek, P. J. (2012). Male reproductive physiology. In A. Wein (Ed.), Campbell-Walsh urology (Vol. 1, pp. 591–615). Elsevier-Saunders.
Francavilla, F., et al. (2007). Naturally-occurring antisperm antibodies in men: Interference with fertility and clinical implications. An update. Frontiers in Bioscience : A Journal and Virtual Library, 12, 2890–2911.
Lee, R., et al. (2009). Value of serum antisperm antibodies in diagnosing obstructive azoospermia. The Journal of Urology, 181(1), 264–269.
Johnson, A., et al. (1995). A quality control system for the optimized sperm penetration assay. Fertility and Sterility, 64(4), 832–837.
Smith, R. G., et al. (1987). Functional tests of spermatozoa. Sperm penetration assay. The Urologic Clinics of North America, 14(3), 451–458.
Bieniek, J. M., Drabovich, A. P., & Lo, K. C. (2016). Seminal biomarkers for the evaluation of male infertility. Asian Journal of Andrology, 18(3), 426–433.
Kumari, A., et al. (2015). Copy number variation and microdeletions of the Y chromosome linked genes and loci across different categories of Indian infertile males. Scientific Reports, 5, 17780.
Bracke, A., et al. (2018). A search for molecular mechanisms underlying male idiopathic infertility. Reproductive Biomedicine Online, 36(3), 327–339.
Araujo, T. F., et al. (2020). Sequence analysis of 37 candidate genes for male infertility: Challenges in variant assessment and validating genes. Andrology, 8(2), 434–441.
Yang, F., et al. (2015). TEX11 is mutated in infertile men with azoospermia and regulates genome-wide recombination rates in mouse. EMBO Molecular Medicine, 7(9), 1198–1210.
Okutman, O., et al. (2015). Exome sequencing reveals a nonsense mutation in TEX15 causing spermatogenic failure in a Turkish family. Human Molecular Genetics, 24(19), 5581–5588.
Nakamura, S., et al. (2017). Next-generation sequencing for patients with non-obstructive azoospermia: Implications for significant roles of monogenic/oligogenic mutations. Andrology, 5(4), 824–831.
Yatsenko, A. N., et al. (2015). X-linked TEX11 mutations, meiotic arrest, and azoospermia in infertile men. The New England Journal of Medicine, 372(22), 2097–2107.
Ayhan, O., et al. (2014). Truncating mutations in TAF4B and ZMYND15 causing recessive azoospermia. Journal of Medical Genetics, 51(4), 239–244.
Maor-Sagie, E., et al. (2015). Deleterious mutation in SYCE1 is associated with non-obstructive azoospermia. Journal of Assisted Reproduction and Genetics, 32(6), 887–891.
Bolcun-Filas, E., et al. (2009). Mutation of the mouse Syce1 gene disrupts synapsis and suggests a link between synaptonemal complex structural components and DNA repair. PLoS Genetics, 5(2), e1000393.
He, W. B., et al. (2018). DMC1 mutation that causes human non-obstructive azoospermia and premature ovarian insufficiency identified by whole-exome sequencing. Journal of Medical Genetics, 55(3), 198–204.
Jiang, L., et al. (2017). CFTR gene mutations and polymorphism are associated with non-obstructive azoospermia: From case-control study. Gene, 626, 282–289.
Arafat, M., et al. (2017). Mutation in TDRD9 causes non-obstructive azoospermia in infertile men. Journal of Medical Genetics, 54(9), 633–639.
Qiu, Q., et al. (2019). pathogenic variants cause male infertility through affecting the proliferation and apoptosis of human spermatogonial stem cells. Aging, 11(24), 12581–12599.
Fu, H., et al. (2018). PAK1 promotes the proliferation and inhibits apoptosis of human spermatogonial stem cells via PDK1/KDR/ZNF367 and ERK1/2 and AKT pathways. Molecular Therapy--Nucleic Acids, 12, 769–786.
van der Bijl, N., et al. (2019). Mutations in the stromal antigen 3 (STAG3) gene cause male infertility due to meiotic arrest. Human Reproduction, 34(11), 2112–2119.
Riera-Escamilla, A., et al. (2019). Sequencing of a ‘mouse azoospermia’ gene panel in azoospermic men: Identification of RNF212 and STAG3 mutations as novel genetic causes of meiotic arrest. Human Reproduction (Oxford, England), 34(6), 978–988.
Ozturk, S., et al. (2016). The poly(A)-binding protein genes, EPAB, PABPC1, and PABPC3 are differentially expressed in infertile men with non-obstructive azoospermia. Journal of Assisted Reproduction and Genetics, 33(3), 335–348.
Khan, M. J., et al. (2018). X-linked ADGRG2 mutation and obstructive azoospermia in a large Pakistani family. Scientific Reports, 8(1), 16280.
Gershoni, M., et al. (2017). A familial study of azoospermic men identifies three novel causative mutations in three new human azoospermia genes. Genetics in Medicine : Official Journal of the American College of Medical Genetics, 19(9), 998–1006.
Tan, Y.-Q., et al. (2019). Loss-of-function mutations in TDRD7 lead to a rare novel syndrome combining congenital cataract and nonobstructive azoospermia in humans. Genetics in Medicine : Official Journal of the American College of Medical Genetics, 21(5), 1209–1217.
Wang, X. N., et al. (2013). The Wilms tumor gene, Wt1, is critical for mouse spermatogenesis via regulation of sertoli cell polarity and is associated with non-obstructive azoospermia in humans. PLoS Genetics, 9(8), e1003645.
Zhang, Y., et al. (2015). Association of single nucleotide polymorphisms in the USF1, GTF2A1L and OR2W3 genes with non-obstructive azoospermia in the Chinese population. Journal of Assisted Reproduction and Genetics, 32(1), 95.
Miyamoto, T., et al. (2003). Azoospermia in patients heterozygous for a mutation in SYCP3. Lancet (London, England), 362(9397), 1714–1719.
Kherraf, Z.-E., et al. (2017). SPINK2 deficiency causes infertility by inducing sperm defects in heterozygotes and azoospermia in homozygotes. EMBO Molecular Medicine, 9(8), 1132–1149.
Ma, Q., et al. (2016). A novel missense mutation in USP26 gene is associated with nonobstructive azoospermia. Reproductive Sciences (Thousand Oaks, Calif.), 23(10), 1434–1441.
Lu, C., et al. (2021). Human X chromosome exome sequencing identifies BCORL1 as contributor to spermatogenesis. Journal of Medical Genetics, 58(1), 56–65.
Lv, M., et al. (2020). Homozygous mutations in DZIP1 can induce asthenoteratospermia with severe MMAF. Journal of Medical Genetics, 57(7), 445–453.
Schilit, S. L. P., et al. (2020). SYCP2 translocation-mediated dysregulation and frameshift variants cause human male infertility. American Journal of Human Genetics, 106(1), 41–57.
Wang, W., et al. (2019). Biallelic mutations in lead to severe asthenoteratospermia due to acrosome hypoplasia and flagellum malformations. Journal of Medical Genetics, 56(11), 750–757.
Kamaliyan, Z., et al. (2018). Investigation of piwi-interacting RNA pathway genes role in idiopathic non-obstructive azoospermia. Scientific Reports, 8(1), 142.
Chuma, S., & Nakano, T. (2013). piRNA and spermatogenesis in mice. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 368(1609), 20110338.
van der Bijl, N., et al. (2019). Mutations in the stromal antigen 3 (STAG3) gene cause male infertility due to meiotic arrest. Human Reproduction (Oxford, England), 34(11), 2112–2119.
McSwiggin, H. M., & O’Doherty, A. M. (2018). Epigenetic reprogramming during spermatogenesis and male factor infertility. Reproduction (Cambridge, England), 156(2), R9–R21.
Kobayashi, H., et al. (2007). Aberrant DNA methylation of imprinted loci in sperm from oligospermic patients. Human Molecular Genetics, 16(21), 2542–2551.
Trasler, J. M. (2009). Epigenetics in spermatogenesis. Molecular and Cellular Endocrinology, 306(1–2), 33–36.
Gutierrez-Arcelus, M., et al. (2013). Passive and active DNA methylation and the interplay with genetic variation in gene regulation. eLife, 2, e00523.
Laqqan, M., Solomayer, E.-F., & Hammadeh, M. (2017). Aberrations in sperm DNA methylation patterns are associated with abnormalities in semen parameters of subfertile males. Reproductive Biology, 17(3), 246–251.
Marcho, C., Oluwayiose, O. A., & Pilsner, J. R. (2020). The preconception environment and sperm epigenetics. Andrology, 8(4), 924–942.
Martin, E. M., & Fry, R. C. (2018). Environmental influences on the epigenome: Exposure-associated DNA methylation in human populations. Annual Review of Public Health, 39, 309–333.
Everson, T. M., et al. (2016). Maternal cadmium, placental PCDHAC1, and fetal development. Reproductive Toxicology, 65, 263–271.
Bind, M.-A., et al. (2012). Air pollution and markers of coagulation, inflammation, and endothelial function: Associations and epigene-environment interactions in an elderly cohort. Epidemiology (Cambridge, Mass.), 23(2), 332–340.
Carmona, J. J., et al. (2014). Short-term airborne particulate matter exposure alters the epigenetic landscape of human genes associated with the mitogen-activated protein kinase network: A cross-sectional study. Environmental Health: A Global Access Science Source, 13, 94.
Fragou, D., et al. (2019). Smoking and DNA methylation: Correlation of methylation with smoking behavior and association with diseases and fetus development following prenatal exposure. Food and Chemical Toxicology, 129, 312–327.
Laqqan, M., et al. (2017). Aberrant DNA methylation patterns of human spermatozoa in current smoker males. Reproductive Toxicology (Elmsford, N.Y.), 71, 126–133.
Laurentino, S., et al. (2015). Epigenetic germline mosaicism in infertile men. Human Molecular Genetics, 24(5), 1295–1304.
Benchaib, M., et al. (2005). Influence of global sperm DNA methylation on IVF results. Human Reproduction, 20(3), 768–773.
Gu, Y., et al. (2016). Nicotine induces Nme2-mediated apoptosis in mouse testes. Biochemical and Biophysical Research Communications, 472(4), 573–579.
Dong, H., et al. (2017). Abnormal methylation of imprinted genes and cigarette smoking: Assessment of their association with the risk of male infertility. Reproductive Sciences (Thousand Oaks, Calif.), 24(1), 114–123.
Xu, W., et al. (2013). Cigarette smoking exposure alters pebp1 DNA methylation and protein profile involved in MAPK signaling pathway in mice testis. Biology of Reproduction, 89(6), 142.
Dai, J., et al. (2016). Protein profile screening: Reduced expression of Sord in the mouse epididymis induced by nicotine inhibits tyrosine phosphorylation level in capacitated spermatozoa. Reproduction (Cambridge, England), 151(3), 227–237.
Liu, Y., et al. (2019). Distinct H3K9me3 and DNA methylation modifications during mouse spermatogenesis. The Journal of Biological Chemistry, 294(49), 18714–18725.
Wang, J., et al. (2017). NRF1 coordinates with DNA methylation to regulate spermatogenesis. FASEB Journal : Official Publication of the Federation of American Societies for Experimental Biology, 31(11), 4959–4970.
Cheng, Y. S., et al. (2017). MAEL promoter hypermethylation is associated with de-repression of LINE-1 in human hypospermatogenesis. Human Reproduction, 32(12), 2373–2381.
Wu, X., et al. (2020). Unraveling epigenomic abnormality in azoospermic human males by WGBS, RNA-Seq, and transcriptome profiling analyses. Journal of Assisted Reproduction and Genetics, 37(4), 789–802.
Zagore, L. L., et al. (2018). DAZL regulates germ cell survival through a network of polyA-proximal mRNA interactions. Cell Reports, 25(5), 1225.
Hamada, A. J., Esteves, S. C., & Agarwal, A. (2013). A comprehensive review of genetics and genetic testing in azoospermia. Clinics (São Paulo, Brazil), 68(Suppl 1), 39–60.
Donker, R. B., et al. (2017). Chromosomal abnormalities in 1663 infertile men with azoospermia: The clinical consequences. Human Reproduction (Oxford, England), 32(12), 2574–2580.
Jungwirth, A., et al. (2012). European Association of Urology guidelines on male infertility: The 2012 update. European Urology, 62(2), 324–332.
Corona, G., et al. (2017). Sperm recovery and ICSI outcomes in Klinefelter syndrome: A systematic review and meta-analysis. Human Reproduction Update, 23(3), 265–275.
Krausz, C., & Riera-Escamilla, A. (2018). Genetics of male infertility. Nature Reviews. Urology, 15(6), 369–384.
Belling, K., et al. (2017). Klinefelter syndrome comorbidities linked to increased X chromosome gene dosage and altered protein interactome activity. Human Molecular Genetics, 26(7), 1219–1229.
Calogero, A. E., et al. (2017). Klinefelter syndrome: Cardiovascular abnormalities and metabolic disorders. Journal of Endocrinological Investigation, 40(7), 705–712.
Gravholt, C. H., et al. (2018). Klinefelter Syndrome: Integrating genetics, neuropsychology, and endocrinology. Endocrine Reviews, 39(4), 389–423.
Krausz, C., & Casamonti, E. (2017). Spermatogenic failure and the Y chromosome. Human Genetics, 136(5), 637–655.
Colaco, S., & Modi, D. (2018). Genetics of the human Y chromosome and its association with male infertility. Reproductive Biology and Endocrinology, 16(1), 14.
Gabriel-Robez, O., et al. (1990). Deletion of the pseudoautosomal region and lack of sex-chromosome pairing at pachytene in two infertile men carrying an X;Y translocation. Cytogenetics and Cell Genetics, 54(1–2), 38–42.
Mohandas, T. K., et al. (1992). Role of the pseudoautosomal region in sex-chromosome pairing during male meiosis: Meiotic studies in a man with a deletion of distal Xp. American Journal of Human Genetics, 51(3), 526–533.
Hassold, T. J., et al. (1991). XY chromosome nondisjunction in man is associated with diminished recombination in the pseudoautosomal region. American Journal of Human Genetics, 49(2), 253–260.
Shi, Q., & Martin, R. H. (2001). Aneuploidy in human spermatozoa: FISH analysis in men with constitutional chromosomal abnormalities, and in infertile men. Reproduction (Cambridge, England), 121(5), 655–666.
Alvarez-Nava, F., & Lanes, R. (2018). Epigenetics in Turner syndrome. Clinical Epigenetics, 10, 45.
Allshire, R. C., & Karpen, G. H. (2008). Epigenetic regulation of centromeric chromatin: Old dogs, new tricks? Nature Reviews. Genetics, 9(12), 923–937.
Guo, J., et al. (2018). The adult human testis transcriptional cell atlas. Cell Research, 28(12), 1141–1157.
Pijuan-Sala, B., et al. (2020). Single-cell chromatin accessibility maps reveal regulatory programs driving early mouse organogenesis. Nature Cell Biology, 22(4), 487–497.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Wu, X., Lin, D., Sun, F., Cheng, C.Y. (2021). Male Infertility in Humans: An Update on Non-obstructive Azoospermia (NOA) and Obstructive Azoospermia (OA). In: Cheng, C., Sun, F. (eds) Molecular Mechanisms in Spermatogenesis. Advances in Experimental Medicine and Biology, vol 1381. Springer, Cham. https://doi.org/10.1007/978-3-030-77779-1_8
Download citation
DOI: https://doi.org/10.1007/978-3-030-77779-1_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-77778-4
Online ISBN: 978-3-030-77779-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)