Abstract
Halohydrin dehalogenases (HHDHs) are an important class of enzymes for preparing optically active haloalcohols, epoxides, and β-substituted alcohols. However, natural HHDHs rarely meet the requirements of industrial applications. Here, a novel high-throughput screening (HTS) methodology for directed evolution of HHDH was developed based on the colorimetric determination of azide. In this method, azide was involved in the HHDH-catalyzed ring-opening process and the decrease of azide was used to quantitatively evaluate HHDH activity. The HTS methodology was simple and sensitive (ε460 = 1.2173 × 104 L mol−1 cm−1) and could be performed in a microplate format using whole cells. To verify the efficiency of the HTS methodology, it was adopted to engineer a HHDH (HHDH-PL) from Parvibaculum lavamentivorans DS-1, which was applied in the process for ethyl (R)-4-cyano-3-hydroxybutanoate (HN) by the conversion of ethyl (S)-4-chloro-3-hydroxybutanoate ((S)-CHBE)). A random mutant library containing 2500 colonies was screened using the HTS methodology, and three beneficial mutants F176M, A187R, and A187S were obtained. By combining the beneficial mutated residues, the variant F176M/A187R was identified with 2.8-fold higher catalytic efficiency for preparation of HN. The high-throughput colorimetric assay would be very useful for directed evolution of HHDH for preparing β-substituted alcohols.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the past few years, the demand for single enantiomers in medicinal chemistry was becoming expanding and increasing. Enzyme-catalyzed reactions exhibit high efficiency, enantioselectivity, and stereoselectivity in synthetic chemistry, which are of high interest for synthesis of fine chemicals and pharmaceutical intermediates (Schmid et al. 2001; Schoemaker et al. 2003). Halohydrin dehalogenases (HHDHs, EC 4.5.1.X), also called halohydrin epoxidases and halohydrin hydrogen-halide-lyases, are able to catalyze the dehalogenation of vicinal halohydrins accompanying the formation of epoxides (de Vries and Janssen 2003). On the other hand, HHDHs could convert epoxides into corresponding β-substituted alcohols undergoing the nucleophile-mediated ring-opening processes (Fig. 1). Many nucleophiles such as CN−, NO2 −, N3 −, SCN−, OCN−, X−, and HCOO− lead to a diverse range of β-functionalized alcohols (de Vries and Janssen 2003; Spelberg et al. 2001).
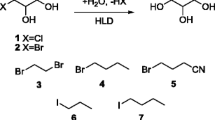
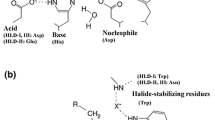
Catalytic mechanism investigations have revealed that HHDHs belong to short-chain dehydrogenase/reductase (SDR) enzyme superfamily and possess a conserved catalytic triad Ser-Tyr-Arg (de Jong et al. 2003). What sets HHDHs apart from a large number of SDR enzymes is that HHDHs do not have a NAD(P)+-binding site; instead, a spacious anion binding pocket is present. Recently, many novel HHDH genes have been identified by database mining which enrich the HHDH enzyme family (Schallmey et al. 2014; Xue et al. 2014). These HHDHs are classified into several phylogenetic subtypes based on the substrate specificity and sequence identity. Furthermore, several crystal structures have been resolved to give deeper insight into the structure-function relationship of HHDHs (de Jong et al. 2006, 2005; Schallmey et al. 2013). HheAAD2 from Arthrobacter sp. strain AD2 exhibits a more open substrate-binding pocket than HheC from Agrobacterium radiobacter AD1, which is in accord with that HheAAD2 probably prefers long-chain halohydrins. HHDHs and their variants have exhibited high regioselectivity and enantioselectivity in the dehalogenation and ring-opening processes. Owing to the especial promiscuity, HHDHs are becoming an attractive biocatalyst in organic chemistry (Vlieg et al. 2001; You et al. 2013). Examples of relevant applications are the preparation of optically active halohydrins (Elenkov et al. 2006b; Tang et al. 2012), epoxides (Spelberg et al. 2001), and β-functionalized alcohols (Elenkov et al. 2006a, 2007; Hasnaoui-Dijoux et al. 2008). An industrial application of HHDH was used for manufacturing HN, which is a key chiral building block for a salable cholesterol-lowering drug of atorvastatin (Fox et al. 2007; Liu et al. 2015; Ma et al. 2010; You et al. 2014). In this enzymatic process, (S)-CHBE is converted to corresponding epoxide and subsequently leading to HN in the presence of cyanide (Fig. 2). A high yield of HN has been achieved under the mild conditions, for which conventional chemical processes are rather challenging. Despite the worthwhile and potential for preparation of fine chemicals, natural HHDHs rarely meet the demands of industrial applications which require the high specific activity, stability, selectivity, and the high tolerance to the substrates and products.
Now, it is convenient to construct a mutant library by many genetic approaches such as DNA shuffling (Stemmer 1994a, b), error-prone PCR (Leung et al. 1989), MEGAWHOP (megaprimer PCR of whole plasmid) (Miyazaki and Takenouchi 2002), site-directed/site saturation mutagenesis (Zheng et al. 2004), and iterative saturation mutagenesis (Reetz and Carballeira 2007). By directed evolution approach, many beneficial enzyme variants with great improvement of capabilities are obtained and applied into the synthesis of pharmaceutical intermediates and fine chemicals on an industrial scale (DeSantis et al. 2003; Fox et al. 2007; Savile et al. 2010). Although directed evolution is an effective route to improve enzymatic properties, screening the large mutant libraries is time-consuming and laborious. Therefore, an accurate and rapid HTS methodology was critical to facilitate the screening procedure (Aharoni et al. 2006; Mastrobattista et al. 2005). Many successful HTS methods for screening enzyme mutants have been developed based on the properties of these specific enzymatic processes (Goddard and Reymond 2004; He et al. 2011; Olsen et al. 2000; Trollope et al. 2014; Zeng et al. 2013; Zheng et al. 2007). Two colorimetric assays have been developed for measuring HHDH activity by determining the released halide ions and protons in dehalogenation process (Schallmey et al. 2013; Tang et al. 2010). Another chromogenic assay for measuring HHDH activity requires the usage of the specific chromogenic substrates (para-nitro-2-bromo-1-phenylethanol and the epoxide para-nitrostyrene oxide) (Spelberg et al. 2002). An adrenaline-based colorimetric assay for monitoring HHDH-catalyzed epoxide ring-opening reactions has been reported recently, while the diol-producing substrate was required in this method (Tang et al. 2014).
HHDH-PL from P. lavamentivorans DS-1 has been cloned and applied to preparation of HN from (S)-CHBE (Wan et al. 2014). Unfortunately, the highly toxic cyanide made it difficult to perform directed evolution. In this study, an efficient and sensitive azide-based colorimetric assay was established and optimized to measure HHDH activity. Subsequently, a HTS methodology was developed and applied to directed evolution of HHDH-PL. After performing random mutation and site-directed mutagenesis, a variant F176M/A187R was obtained with up to 2.8-fold higher catalytic efficiency for synthesis of HN.
Materials and methods
Chemicals and reagents
All analytical reagents (sodium azide, ferric chloride, disodium hydrogen phosphate, monometallic sodium orthophosphate, boric acid, sodium borate, concentrated hydrochloric acid, concentrated sulfuric acid, and Tris base) were purchased from Aladdin Chemistry (Shanghai, China). (S)-CHBE and HN were purchased from J & K Chemical Technology (Shanghai, China). All microplate assays were performed on a Spectramax 5.4 microplate reader with SoftMax PRO version 5.2 software (Molecular Devices, Sunnyvale, CA, USA). Ninety-six well standard microplate (maximum volume 300 μL/well, Thermo Scientific, Shanghai, China) used for colorimetric assay and 96-deep-well plates (maximum volume 2 mL/well, Sangon Biotech, Shanghai, China) were employed to culture and conserve Escherichia coli cells. PrimerSTAR® HS DNA polymerase and restriction enzymes were from Takara (Dalian, China). GoTaq Green® Master Mix was from Promega (Madison, WI, USA). AxyPrep Plasmid Miniprep Kit, DNA Gel Extraction Kit, and PCR Cleanup Kit were from AXYGEN (Shanghai, China).
Establishment and optimization of the colorimetric assay
Ferric chloride solution (FCS) was selected to determine the concentration of azide. At first, several concentrations (25, 50, 100, and 200 mM) of FCS were tested to determine azide over the concentration ranges from 0 to 5 mM. Subsequently, three buffers of Na2HPO4-NaH2PO4, Na2B4O7-H2BO3, and Tris-H2SO4 (pH 8.0, concentrations range from 0 to 200 mM) were selected to imitate the enzymatic catalysis system. Different concentrations (from 0 to 200 mM) of HCl in Tris-H2SO4 buffer (50 mM and pH 8.0) and water were conducted to mimic the protons and halide ions that are released upon the dehalogenation process. For investigating their effects on the colorimetric assay, 10 μL of each buffer was added into 190 μL 50 mM FCS containing 1 mM sodium azide and recorded on Spectramax 5.4.
Cloning and expression of HHDH-PL
HHDH-PL gene (GenBank Accession Number, ABS64560.1) was amplified and ligated into NcoI/XhoI-digested pET32a (+) plasmids (Novagen, Darmstadt, Germany) with the trx-tag and his-tag flanking the gene N-terminal. The cloning primers are listed in Table S1. The recombinant plasmids were transformed into competent E. coli BL21 (DE3). Expression of the recombinant HHDH-PL was carried out in Luria-Bertani (LB) medium with 0.1 mM isopropyl-β-d-thiogalactopyranoside (IPTG) as the previously described procedure (Wan et al. 2014). Recombinant E. coli cells were harvested by centrifugation (BECKMAN COULTER Avanti J-26S XP, Brea, CA, USA) at 10,000 × g and 4 °C for 10 min. After discarding the supernatant, precipitated cells were washed with 50 mM Tris-H2SO4 buffer (pH 8.0).
Construction of a random mutant library
The random mutant library of HHDH-PL was generated using error-prone and MEGAWHOP PCR methods. The error-prone PCR was carried out in a total volume of 50 μL containing 25 μL 2× GoTaq Green® Master Mix, 5 μL 50 mM MgCl2 buffer, 5 μL 1 mM MnCl2, 0.5 μL (50 μM) of each forward and reverse primers (Table S2), 1 μL (100–200 ng) HHDH-PL plasmids, and 13 μL deionized water. The reaction mixtures were heated at 95 °C for 3 min, followed by 25 cycles of incubation at 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 60 s, and a final incubation at 72 °C for 10 min. The PCR products were analyzed by agarose gel electrophoresis assay and purified with PCR Cleanup Kit. The purified PCR products were used as primers for MEGAWHOP PCR. The reactions were carried out in a total volume of 50 μL containing 10 μL 5× PrimerStar buffer, 4 μL dNTP Mix (0.8 mM), 1 μL HHDH-PL plasmids, 0.5 μL PrimerStar, 8 μL mega primers (1 μg, the purified error-prone PCR products), and 26.5 μL deionized water. The PCR was performed as the following program: 72 °C for 10 min, 5 min at 95 °C followed by 30 cycles of 10 s at 98 °C, 15 s at 55 °C, 8 min at 72 °C, and a final incubation at 72 °C for 10 min. Ten-microliter PCR products were analyzed by agarose gel electrophoresis, and the remaining PCR products (40 μL) were digested with restriction enzyme DpnI (2 μL) at 37 °C for 3 h. The digested product was incubated at 65 °C for 30 min, transformed into the chemically competent E. coli BL21 (DE3), and plated on the agar/ampicillin LB plates.
Developing a HTS methodology
The colonies on the agar plates were picked into 96-deep-well plates containing 1 mL LB medium and 50 μg/mL ampicillin. Three wild-type HHDH-PL (HHDH-PL WT) colonies were picked as controls. The plates were incubated at 37 °C and stirred for 5 h. Subsequently, each 100 μL culture was transferred into another 96-deep-well plate containing 100 μL of 30 % (w/v) sterile glycerol for conservation at −70 °C. One hundred-microliter fresh LB medium containing 50 μg/mL ampicillin and 1 mM IPTG was added into the remaining culture (900 μL) and induced at 28 °C for 12–14 h. The E. coli cells were collected by centrifugation at 3000 × g and 4 °C for 30 min, and the supernatants were discarded. Five hundred-microliter analysis buffer (50 mM Tri-H2SO4 buffer, 20 mM (S)-CHBE, and 20 mM NaN3) was added into each well to resuspend the cells. The plates were stirred at 40 °C, 150 rpm for 30 min. Ten-microliter reaction mixtures of each well were added into the standard 96-well plates containing 190 μL 50 mM FCS. Colorimetric assay was carried out at 460 nm on Spectramax 5.4 (Fig. S7). The relative activities were calculated using the following equation:
where OD460(c) and OD460(r) represented the optical density of the negative control (analysis buffer) and the sample catalyzed by HHDH, respectively, t represented the reaction time, 20 was the dilution factor, and 1.2173 was the coefficient of calibration curve (y = 1.2173 x + 0.2774, Fig. S2). The variants with higher relative activities than HHDH-PL WT were collected as beneficial mutants. The remaining reaction mixtures of beneficial mutants were extracted with 700 μL ethyl acetate and dried on anhydrous sodium sulfate for GC verification. GC analysis of (S)-CHBE, ethyl (S)-oxirane-2-acetate (OAE), and ethyl (R)-4-azido-3-hydroxybutanoate (AHE) was performed on GC-14C (Shimadzu, Japan) equipped with flame ionization detector (FID) detector and the Astec CHIRALDEX™ G-TA column (30 m × 0.25 mm, 0.12-μm film thickness, Supelco, USA). The temperature program was isothermal at 90 °C for 5 min, increase at 5 °C/min to 140 °C, and final staying at 140 °C for 10 min. Retention times were as follows: R t = 6.7 min for OAE, R t = 14.7 min for (S)-CHBE, and R t = 19.6 min for AHE (Fig. S3).
Recombination of beneficial mutants by site-directed mutagenesis
Three beneficial mutants F176M, A187S, and A187R were obtained after screening the random mutant library. Plasmid F176M was prepared using Plasmid Miniprep Kit and used as templates for site-directed mutagenesis PCR. The double mutants F176M/A187S and F176M/A187R were constructed using the corresponding primers given in Table S3. PCR reactions were carried out in a total volume of 50 μL using PrimerStar DNA polymerases with the following program: 95 °C for 5 min, followed by 30 cycles of 10 s at 98 °C, 15 s at 58 °C, 8 min at 72 °C, and a final incubation at 72 °C for 10 min. After digestion with DpnI at 37 °C for 3 h, the PCR products were transformed into E. coli BL21 (DE3) and plated on agar/ampicillin LB plates. Successful construction of these mutants was verified by sequencing.
Determination of the relative activities for HN
Mutants F176M, A187R, A187R, F176M/A187R, and F176M/A187S were tested to prepare HN and compared with HHDH-PL WT. Sodium dodecyl sulfate (SDS)-PAGE assay indicated that the expressions of these HHDHs were equivalent in the same conditions (Fig. S5), and the crude enzymes were used to determine relative activity. One-gram wet weight cells were resuspended in 10 mL Tris-H2SO4 buffer (200 mM, pH 7.5) for sonication for 20 min. After centrifugation at 12,000 × g and 4 °C for 20 min, the supernatants were used as crude enzymes to prepare HN. All the reactions were performed on 902 Titrando system (Metrohm, Switzerland) equipped with a magnetic stirrer (IKA, Staufen, Germany) and pH stat. In a 50-mL reactor, 30 mL Tris-H2SO4 buffer (200 mM, pH 7.5) and 400-μL 30 % (w/v) NaCN solution (caution: NaCN was highly toxic) were added, and the system pH was adjusted to 7.5 using 50 % diluted sulfuric acid. After heating the mixture up to 40 °C, 1.0 g (S)-CHBE and 10 mL of crude enzyme were added and stirred at 500 rpm. In this process, the protons released in the dehalogenation step would lower the pH, and 30 % NaCN solution as a base was injected to keep pH 7.5 by pH stat. After 30 min, 800 μL per reaction was extracted with 800 μL ethyl acetate for GC assay. GC analysis for (S)-CHBE, OAE, and HN was carried out on GC-14C equipped with FID detector and G-TA column. The temperature program was isothermal at 90 °C for 5 min, increase at 5 °C/min to 180 °C, and final 2 min at 180 °C. Retention times were as follows: R t = 7.9 min for OAE, R t = 13.9 min for (S)-CHBE, and R t = 18.6 min for HN (Fig. S4). The activities were calculated by using the yield of HN.
Purification of HHDH-PL WT and F176M/A187R
The variant F176M/A187R and HHDH-PL WT were cultured and expressed in a shaking flask (5 × 100 mL of LB medium). The crude enzymes of F176M/A187R and HHDH-PL WT were prepared as described above. The supernatants were applied to a Nickel-NTA column (MC/20, 16 × 100, A&B Applied Biosystems) preequilibrated with buffer A (20 mM imidazole, 50 mM Tris sulfate, 300 mM NaCl, pH 8.0) at flow rate of 2.0 mL/min. Non-tagged containing proteins were washed off with ten column volumes of buffer A and ten column volumes of buffer B (50 mM imidazole, 50 mM Tris sulfate, 300 mM NaCl, pH 8.0). Bound proteins were eluted with buffer C (200 mM imidazole, 50 mM Tris sulfate, 500 mM NaCl, pH 8.0) (Liu et al. 2009). The eluted fractions showing HHDH activity were pooled, concentrated, and buffer exchanged in 50 mM Tris sulfate (pH 8.0) using an ultrafiltration membrane. SDS-PAGE assay using a 12 % gel was carried out to determine the purity of the purified HHDHs (Fig. S6). The concentration of the purified enzyme was determined by measuring the absorbance at 590 nm with BCA Protein Assay Kit (KeyGen Biotech, China).
Determination of kinetic constants of HHDH-PL WT and F176M/A187R
The steady-state kinetic parameters of HHDH-PL WT and F176M/A187R were examined using substrate (S)-CHBE. The initial velocities of the formation of OAE and HN were determined by varying the concentrations of substrate (S)-CHBE ranging from 10 to 100 mM. The initial rates obtained were fitted with the Michaelis-Menten equation (V = V max · [S]/K m + [S], where V max was the maximum rate, [S] was the substrate concentration, and K m was the Michaelis constant) using Origin 8.0. The enzymatic reactions to determine the formation of OAE were performed in 850 μL Tris-H2SO4 (100 mM, pH 8.0) buffer with 50 μL purified enzyme and 100 μL substrate solution (0.1 to 1 M of (S)-CHBE in 100 mM Tris-H2SO4 buffer, pH 8.0). The enzymatic reactions to determine the formation of HN were performed in 850 μL Tris-H2SO4 (100 mM, pH 8.0) buffer containing 117 mM NaCN (100 mM for the final 1-mL mixture). One hundred-microliter substrate solution and 50 μL purified enzyme were added to the mixture for incubation at 40 °C. All the reactions were carried out on Thermomixer comfort (Eppendorf, Hamburg, Germany) at 500 rpm. After the reactions were stopped, 1000 μL ethyl acetate was added to extract the products and the organic extracts were analyzed by GC. The yields of OAE and HN were determined by GC-14 using the program as described above.
Molecular homology modeling and docking
The modeling structures of HHDH-PL WT and mutant F176M/A187S were constructed by Modeller software (version 9.12) using the crystal structure of HheAAD2 (PDB code: 1ZMO, 2.00 Å) (de Jong et al. 2006) and HheC (PDB code: 1PWX, 1.80 Å) (de Jong et al. 2005) as templates. The generated models were sent to Procheck (Laskowski et al. 1993) and UCLA-DOE server for analyzing and validation (Khajamohiddin et al. 2006). The optimal models were selected to perform docking study for (S)-CHBE using AutoDocK (version 4.20) (Morris et al. 2009). (S)-CHBE was treated with the program LigPrep of Maestro 9.0 (Schrodinger LLC) to generate the lowest energy conformation before performing docking studies. The docking results were analyzed and visualized using PyMoL software (academic version).
Results
Establishment and optimization of the colorimetric assay
HHDH was able to catalyze the dehalogenation of vicinal halohydrins and the ring opening of epoxides in the presence of nucleophiles. Azide was a frequently used nucleophile for kinetic resolution of epoxides by HHDHs (Elenkov et al. 2008; Hasnaoui-Dijoux et al. 2008; Molinaro et al. 2010; Spelberg et al. 2001). In this study, azide was used as a nucleophile and involved in the continuous dehalogenation and ring-opening processes (Fig. 3). In this case, halohydrin was converted into corresponding epoxide and followed by ring opening with azide to form β-azidoalcohol which could be monitored by the decrease of azide. The aim of this study was to develop an efficient assay to monitor the changes of azide in the ring-opening process. A spectrophotometric determination of azide by ferric perchlorate solution has been reported previously (Anton et al. 1960), which was based on the blood red complex (iron(III)/azide) with a characteristic absorbance at 454 nm. Herein, ferric chloride solution (FCS) was used and optimized to establish a proper colorimetric assay for determining HHDH activity. SCN- was not chosen because it exhibited a high inhibition constant and low activity in HHDH-catalyzed ring-opening process even if it had a convenient colorimetric assay (Bergmann and Sanik 1957; Hasnaoui-Dijoux et al. 2008). Nucleophiles including CN−, OCN−, and NO2 − were also eliminated due to the inconvenient test methods for measuring HHDH activity.
The different concentrations of azide were analyzed in 25 mM FCS, and the generated iron(III)/azide complexes were scanned at the wavelength ranges from 390 to 750 nm (Fig. S1). With the raise of azide, the absorbance was gradually increased at 460 nm. In order to gain the feasible colorimetric conditions for monitoring HHDH activity, the concentrations of azide and FCS were systematically investigated (Fig. 4). Absorbances reached the maximum 3.0 at 3 mM azide. At the same concentration of azide, the higher concentration of FCS gave the higher absorbance. The low concentration (25 mM) of FCS was hard to observe the color change, and the high concentration of FCS (100 and 200 mM) had a high intrinsic absorbance. All things considered, 50 mM FCS was selected to determine the azide. The detection calibration curve (y = 1.2173x + 0.2774, R 2 = 0.999) for azide over the concentration from 0 to 2 mM was plotted from the mean extinction values (Fig. S2). The molar absorptivity, ε460 = 1.2173 × 104 L mol−1 cm−1, was determined from the slope of the calibration curve.
In the HHDH-catalyzed reactions, the released protons would lower the pH and a buffer was usually used to keep the optimal catalytic pH (7.8–8.0). Here, three buffers Na2HPO4-NaH2PO4, Na2B4O7-H2BO3, and Tris-H2SO4 (pH 8.0) were selected and their effects on this colorimetric method were investigated. Figure 5 shows that Na2B4O7-H2BO3 and Tris-H2SO4 buffers had no effect on the absorbance, but Na2HPO4-NaH2PO4 buffer irregularly changed the absorbance owing to the formed sediment ironic phosphate. In addition, the liberated protons and halide ions were mimicked using different concentrations of HCl, and their influences on the absorbance were also examined. Figure 6 shows that the HCl in water and Tris-H2SO4 buffer (pH 8.0, 50 mM) gave the similar influences on the absorbance. The HCl concentration of up to 1.3 mM had insignificant effect on the absorbance. With continual increase of the HCl, the absorbance was gradually reduced, and a distinct decrease was found at the concentration higher than 10 mM. For the accurate absorbance, it was better to control the final proton concentration within 2.5 mM by adjusting the concentrations of enzyme and substrates, reaction time or the mixture volume that is used for colorimetric assay.
Based on the results obtained above, a colorimetric assay for measuring HHDH activity was established: the enzymatic reaction was carried out in the Tris-H2SO4 buffer (100 mM, pH 8.0) containing 20 mM substrate (halohydrins or epoxides) and 20 mM NaN3. After incubation at 40 °C for a specific time, 10 μL of the reaction mixture was added into 190 μL 50 mM FCS for colorimetric assay at 460 nm. The relative activity could be calculated by the decrease rate of azide.
Screening mutant library using HTS methodology
The developed colorimetric assay was adopted to engineer HHDH-PL in HTS format for higher catalytic efficiency of HN. Figure 7 is a brief summary of the HTS methodology for directed HHDH-PL evolution. After screening a mutant library of 2500 colonies in 2 weeks (3 days for constructing random mutagenesis library, every 2 days for 500 colonies screening), three beneficial mutants F176M, A187S, and A817R were identified with the improvement of the catalytic efficiency for AHE. For getting a higher activity, the beneficial mutated residues combined by site-directed mutagenesis were used to generate the mutants F176M/A187R and F176M/A187S. Subsequently, the capabilities of these mutants for preparing HN were tested and compared to HHDH-PL WT. The absolute activity for HHDH-PL WT was 1521 U/mL (crude enzyme). Figure 8 reveals that mutants F176M, A187S, and A187R gave the 2.1-, 1.6-, and 1.8-fold higher activity than HHDH-PL WT, respectively. The double mutant F176M/A187S was identified with 2.3-fold higher activity than HHDH-PL WT, but the increase was not significant compared with F176M. The relative activity of the best mutant F176M/A187R was 3.1-fold higher than HHDH-PL WT.
Determination of kinetic constants of HHDH-PL WT and F176M/A187R
The kinetic constants, K m, k cat, and k cat/K m, for the formation of OAE and HN were determined and are summarized in Table 1. The enzymatic efficiency (k cat/K m, 533 ± 10 mM−1 s−1) of mutant F176M/A187R for formation of OAE was 1.75-fold higher than HHDH-PL WT (k cat/K m, 305 ± 6 mM−1 s−1). On the other hand, the enzymatic efficiency (k cat/K m, 207 ± 3 mM−1 s−1) of mutant F176M/A187R for formation of HN was 2.8-fold higher than HHDH-PL WT (k cat/K m, 74 ± 2 mM−1 s−1). The results indicated that the catalytic efficiencies for ring closure and ring-opening processes were both improved by the mutant F176M/A187R.
Molecular homology modeling and docking
To gain deeper insights into the structural changes, the docking studies of HHDH-PL WT and mutant F176M/A187R for (S)-CHBE were performed using the model structures (Fig. 9). S133 was involved in substrate binding, and Y146 acquired a proton from (S)-CHBE to form OAE. The substitution of F176M and A187R changed the conformation of (S)-CHBE in the active site, which made (S)-CHBE closer to the catalytic triad S133-Y146-R150. The mutation of F176M and A187R had no effect on the conformation of catalytic triad. When the small amino acid A187 mutated to large amino acid R187, the steric hindrance was enhanced to (S)-CHBE, which changed the conformation of (S)-CHBE to the catalytic triad. As a result, the substrate (S)-CHBE was closed to S133 and Y146 which made it easier for substrate binding and proton transfer in dehalogenation and cyanation processes. The affinity of mutant F176M/A187R to (S)-CHBE was increased, which was reflected by the decrease of K m from 7.3 ± 0.3 mM (HHDH-PL WT) to 5.9 ± 0.2 mM (F176M/A187R). The two residue substitutions contributed to the 2.8-fold improvement of catalytic efficiency for preparing HN.
Binding poses of (S)-CHBE in HHDH-PL WT and mutant F176M/A187R. HHDH-PL WT and mutant F176M/A187 were painted with magenta and green. The (S)-CHBE binding in HHDH-PL WT was painted with magenta, and the (S)-CHBE binding in F176M/A187R was painted with green. Catalytic triad Ser133-Tyr146-Arg150, Phe176, Ala187, Met176, Arg187, and (S)-CHBE were shown as sticks
Discussion
Recently, HHDHs have exhibited excellent practicability in the synthesis of optically active pharmaceutical intermediates and fine chemicals such as enantiopure halohydrins, epoxides, and β-substituted alcohols (Elenkov et al. 2006a, 2008; Molinaro et al. 2010; Tang et al. 2012). However, the reported natural HHDHs with low activity could not meet the requirements of industrial application, and most studies were still in the stage of experiment (de Vries and Janssen 2003). In the last few decades, many gene techniques were developed to construct mutant libraries, which made it convenient for engineering enzymes in vitro. Due to the large mutant library in evolution procedure, an effective and accurate HTS methodology was crucial to rapidly find a desired excellent variant.
In this study, a colorimetric assay for measuring HHDH activity was established based on the existing spectrophotometric assay for the determination of azide with FCS. In view of the accuracy and visualized color change, 50 mM FCS was adopted to determine the azide and analyze HHDH activity. The enzymatic reaction should be carried out in Tris-H2SO4 buffer or Na2B4O7-H2BO3 buffer. In the case of using a halohydrin substrate, the high concentration of proton released in dehalogenation process would affect the absorbance. Hence, it was better to control the final proton concentration lower than 2.5 mM in colorimetric process. However, when a epoxide substrate was used in this colorimeric assay, it could be performed without consideration of the effect of proton. The detection calibration curve (y = 1.2173 × + 0.2774, R 2 = 0.999) for azide was linear with the molar absorptivity 1.2173 × 104 L mol−1 cm−1, which indicated that this colorimetric was sensitive and accurate for determining HHDH activity. Because of the high molar absorptivity of iron(III)/azide complex, the final azide concentration in FCS should be controlled in 0–2 mM by altering the sample volume that used for colorimetric assay.
In addition, a HTS methodology was established based on the colorimetric assay and used to screen mutant library of HHDH-PL. After quickly screening 2500 colonies, two variants F176M and A187R with 2.1- and 1.8-fold higher relative activities than HHDH-PL WT were obtained. By combining the two mutated residues, the double mutant F176M/A187R exhibited 2.8-fold improvement of catalytic efficiency for HN. Although the improvement was not remarkable, several additional potential applications regarding the colorimetric assay were worth noting. Firstly, the colorimetric assay could be used to screen the HHDH-producing strains. The previous reported colorimetric assay for measuring HHDH activity was based on the liberated halide ions or protons in dehalogenation process. In some cases, halohydrins such as 1,3-dibromo-2-propanol and 1,3-dichloro-2-propanol could spontaneously generate the corresponding epoxides in alkalescent condition, and the β-substituted alcohols were difficult to form in the absence of HHDH. Hence, the colorimetric assay could remove the background activity in screening process. Secondly, the colorimetric assay was not limited to the substrates. Both halohydrin and epoxide substrates were applicable in this colorimetric assay. Thirdly, the HTS methodology based on the colorimetric assay could be performed on a 96-well microplate format using whole cells, which was suitable for directed evolution of HHDH for preparing β-substituted alcohols. The halide-based colorimetric assay was sensitive but complicated, which involves two solutions: solution I [NH4Fe(SO4)2 (0.25 M) in HNO3 (9 M)] and solution II [saturated solution of Hg(SCN)2 in absolute ethanol] (Bergmann and Sanik 1957; Schallmey et al. 2013). In addition, the Hg(SCN)2 solution was highly toxic which was unsafe for a HTS procedure. The pH-based high-throughput colorimetric assay for screening HHDH mutant library has been reported, while it was inconvenient to establish the optimal concentrations of HEPES and enzyme (Tang et al. 2012). Furthermore, the two colorimetric methods were limited to dehalogenation process and not suitable for ring-opening reaction. Another chromogenic assay for measuring HHDH activity is needed to use the chromogenic substrates para-nitro-2-bromo-1-phenylethanol (ε310 = 4289 L mol−1 cm−1) and the epoxide para-nitrostyrene oxide (ε310 = 3304 L mol−1 cm−1). This colorimetric assay was limited to the chromogenic substrates, and the sensitivity was lower than the azide-based colorimetric assay (ε460 = 1.2173 × 104). The adrenaline-based colorimetric assay was limited to the diol-producing substrates (Tang et al. 2014). In summary, the azide-based HTS methodology was simple, sensitive, and no substrate limit.
In conclusion, a sensitive and accurate colorimetric assay was presented to determine HHDH activity based on the determination of azide. In addition, a novel HTS methodology was developed by means of the colorimetric assay and applied to improving the catalytic efficiency of HHDH-PL for preparing HN via conversion of (S)-CHBE. By performing random mutation and site-directed mutagenesis, the variant F176M/A187R was identified with 2.8-fold improvement of catalytic efficiency for HN. This efficient HTS methodology would be very useful for the directed evolution of HHDHs for preparing β-substituted alcohols.
References
Aharoni A, Thieme K, Chiu CPC, Buchini S, Lairson LL, Chen HM, Strynadka NCJ, Wakarchuk WW, Withers SG (2006) High-throughput screening methodology for the directed evolution of glycosyltransferases. Nat Methods 3:609–614
Anton A, Dodd JG, Harvey AE (1960) Spectrophotometric determination of azide with ferric perchlorate. Anal Chem 32:1209–1210
Bergmann JG, Sanik J (1957) Determination of trace amounts of chlorine in naphtha. Anal Chem 29:241–243
de Jong RM, Tiesinga JJW, Rozeboom HJ, Kalk KH, Tang L, Janssen DB, Dijkstra BW (2003) Structure and mechanism of a bacterial haloalcohol dehalogenase: a new variation of the short-chain dehydrogenase/reductase fold without an NAD(P)H binding site. Embo J 22:4933–4944
de Jong RM, Tiesinga JJW, Villa A, Tang LX, Janssen DB, Dijkstra BW (2005) Structural basis for the enantioselectivity of an epoxide ring opening reaction catalyzed by halo alcohol dehalogenase HheC. J Am Chem Soc 127:13338–13343
de Jong RM, Kalk KH, Tang L, Janssen DB, Dijkstra BW (2006) The X-ray structure of the haloalcohol dehalogenase HheA from Arthrobacter sp. strain AD2: insight into enantioselectivity and halide binding in the haloalcohol dehalogenase family. J Bacteriol 188:4051–4056
de Vries EJ, Janssen DB (2003) Biocatalytic conversion of epoxides. Curr Opin Biotechnol 14:414–420
DeSantis G, Wong K, Farwell B, Chatman K, Zhu ZL, Tomlinson G, Huang HJ, Tan XQ, Bibbs L, Chen P, Kretz K, Burk MJ (2003) Creation of a productive, highly enantioselective nitrilase through gene site saturation mutagenesis (GSSM). J Am Chem Soc 125:11476–11477
Elenkov MM, Hauer B, Janssen DB (2006a) Enantioselective ring opening of epoxides with cyanide catalysed by halohydrin dehalogenases: a new approach to non-racemic beta-hydroxy nitriles. Adv Synth Catal 348:579–585
Elenkov MM, Tang LX, Hauer B, Janssen DB (2006b) Sequential kinetic resolution catalyzed by halohydrin dehalogenase. Org Lett 8:4227–4229
Elenkov MM, Hoeffken HW, Tang L, Hauer B, Janssen DB (2007) Enzyme-catalyzed nucleophilic ring opening of epoxides for the preparation of enantiopure tertiary alcohols. Adv Synth Catal 349:2279–2285
Elenkov MM, Tang LX, Meetsma A, Hauer B, Janssen DB (2008) Formation of enantiopure 5-substituted oxazolidinones through enzyme-catalysed kinetic resolution of epoxides. Org Lett 10:2417–2420
Fox RJ, Davis SC, Mundorff EC, Newman LM, Gavrilovic V, Ma SK, Chung LM, Ching C, Tam S, Muley S, Grate J, Gruber J, Whitman JC, Sheldon RA, Huisman GW (2007) Improving catalytic function by ProSAR-driven enzyme evolution. Nat Biotechnol 25:338–344
Goddard JP, Reymond JL (2004) Enzyme assays for high-throughput screening. Curr Opin Biotechnol 15:314–322
Hasnaoui-Dijoux G, Elenkov MM, Spelberg JHL, Hauer B, Janssen DB (2008) Catalytic promiscuity of halohydrin dehalogenase and its application in enantioselective epoxide ring opening. Chem Bio Chem 9:1048–1051
He YC, Ma CL, Xu JH, Zhou L (2011) A high-throughput screening strategy for nitrile-hydrolyzing enzymes based on ferric hydroxamate spectrophotometry. Appl Microbiol Biotechnol 89:817–823
Khajamohiddin S, Babu PS, Chakka D, Merrick M, Bhaduri A, Sowdhamini R, Siddavattam D (2006) A novel meta-cleavage product hydrolase from flavobacterium sp. ATCC27551. Biochem Biophys Res Commun 351:675–681
Laskowski RA, Macarthur MW, Moss DS, Thornton JM (1993) Procheck—a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291
Leung DW, Chen E, Goeddel DV (1989) A method for random mutagenesis of a defined DNA segment using a modified polymerase chain reaction. Technique 1:1–15
Liu ZQ, Gosser Y, Baker PJ, Ravee Y, Lu ZY, Alemu G, Li HG, Butterfoss GL, Kong XP, Gross R, Montclare JK (2009) Structural and functional studies of Aspergillus oryzae cutinase: enhanced thermostability and hydrolytic activity of synthetic ester and polyester degradation. J Am Chem Soc 131:15711–15716
Liu ZQ, Ye JJ, Shen ZY, Hong HB, Yan JB, Lin Y, Chen ZX, Zheng YG, Shen YC (2015) Upscale production of ethyl (S)-4-chloro-3-hydroxybutanoate by using carbonyl reductase coupled with glucose dehydrogenase in aqueous-organic solvent system. Appl Microbiol Biotechnol 99:2119–2129
Ma SK, Gruber J, Davis C, Newman L, Gray D, Wang A, Grate J, Huisman GW, Sheldon RA (2010) A green-by-design biocatalytic process for atorvastatin intermediate. Green Chem 12:81–86
Mastrobattista E, Taly V, Chanudet E, Treacy P, Kelly BT, Griffiths AD (2005) High-throughput screening of enzyme libraries: in vitro evolution of a β-galactosidase by fluorescence-activated sorting of double emulsions. Chem Biol 12:1291–1300
Miyazaki K, Takenouchi M (2002) Creating random mutagenesis libraries using megaprimer PCR of whole plasmid. Biotechniques 33:1033–1037
Molinaro C, Guilbault AA, Kosjek B (2010) Resolution of 2,2-disubstituted epoxides via biocatalytic azidolysis. Org Lett 12:3772–3775
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Olsen M, Iverson B, Georgiou G (2000) High-throughput screening of enzyme libraries. Curr Opin Biotechnol 11:331–337
Reetz MT, Carballeira JD (2007) Iterative saturation mutagenesis (ISM) for rapid directed evolution of functional enzymes. Nat Protoc 2:891–903
Savile CK, Janey JM, Mundorff EC, Moore JC, Tam S, Jarvis WR, Colbeck JC, Krebber A, Fleitz FJ, Brands J, Devine PN, Huisman GW, Hughes GJ (2010) Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture. Science 329:305–309
Schallmey M, Floor RJ, Hauer B, Breuer M, Jekel PA, Wijma HJ, Dijkstra BW, Janssen DB (2013) Biocatalytic and structural properties of a highly engineered halohydrin dehalogenase. Chem Bio Chem 14:870–881
Schallmey M, Koopmeiners J, Wells E, Wardenga R, Schallmey A (2014) Expanding the halohydrin dehalogenase enzyme family: identification of novel enzymes by database mining. Appl Environ Microbiol 80:7303–7315
Schmid A, Dordick JS, Hauer B, Kiener A, Wubbolts M, Witholt B (2001) Industrial biocatalysis today and tomorrow. Nature 409:258–268
Schoemaker HE, Mink D, Wubbolts MG (2003) Dispelling the myths—biocatalysis in industrial synthesis. Science 299:1694–1697
Spelberg JHL, Vlieg JETV, Tang LX, Janssen DB, Kellogg RM (2001) Highly enantioselective and regioselective biocatalytic azidolysis of aromatic epoxides. Org Lett 3:41–43
Spelberg JHL, Tang LX, van Gelder M, Kellogg RM, Janssen DB (2002) Exploration of the biocatalytic potential of a halohydrin dehalogenase using chromogenic substrates. Tetrahedron-Asymmetr 13:1083–1089
Stemmer WPC (1994a) DNA shuffling by random fragmentation and reassembly in-vitro recombination for molecular evolution. Proc Natl Acad Sci U S A 91:10747–10751
Stemmer WPC (1994b) Rapid evolution of a protein in-vitro by DNA shuffling. Nature 370:389–391
Tang LX, Li Y, Wang XO (2010) A high-throughput colorimetric assay for screening halohydrin dehalogenase saturation mutagenesis libraries. J Biotechnol 147:164–168
Tang LX, Zhu XC, Zheng HY, Jiang RX, Elenkov MM (2012) Key residues for controlling enantioselectivity of halohydrin dehalogenase from Arthrobacter sp. strain AD2 revealed by structure-guided directed evolution. Appl Environ Microbiol 78:2631–2637
Tang L, Liu Y, Jiang R, Zheng Y, Zheng K, Zheng H (2014) A high-throughput adrenaline test for the exploration of the catalytic potential of halohydrin dehalogenases in epoxide-ring opening reactions. Biotechnol Appl Biochem. doi:10.1002/bab.1278
Trollope KM, Nieuwoudt HH, Gorgens JF, Volschenk H (2014) Screening a random mutagenesis library of a fungal beta-fructofuranosidase using FT-MIR ATR spectroscopy and multivariate analysis. Appl Microbiol Biotechnol 98:4063–4073
Vlieg JETV, Tang LX, Spelberg JHL, Smilda T, Poelarends GJ, Bosma T, van Merode AEJ, Fraaije MW, Janssen DB (2001) Halohydrin dehalogenases are structurally and mechanistically related to short-chain dehydrogenases/reductases. J Bacteriol 183:5058–5066
Wan NW, Liu ZQ, Huang K, Shen ZY, Xue F, Zheng YG, Shen YC (2014) Synthesis of ethyl (R)-4-cyano-3-hydroxybutyrate at high concentration using a novel halohydrin dehalogenase HHDH-PL from Parvibaculum lavamentivorans DS-1. RSC Adv 109:64027–64031
Xue F, Liu ZQ, Wan NW, Zheng YG (2014) Purification, gene cloning, and characterization of a novel halohydrin dehalogenase from Agromyces mediolanus ZJB120203. Appl Biochem Biotechnol 174:352–364
You ZY, Liu ZQ, Zheng YG (2013) Properties and biotechnological applications of halohydrin dehalogenases: current state and future perspectives. Appl Microbiol Biotechnol 97:9–21
You ZY, Liu ZQ, Zheng YG (2014) Chemical and enzymatic approaches to the synthesis of optically pure ethyl (R)-4-cyano-3-hydroxybutanoate. Appl Microbiol Biotechnol 98:11–21
Zeng W, Lin YS, Qi ZX, He YY, Wang DY, Chen GG, Liang ZQ (2013) An integrated high-throughput strategy for rapid screening of poly(gamma-glutamic acid)-producing bacteria. Appl Microbiol Biotechnol 97:2163–2172
Zheng L, Baumann U, Reymond JL (2004) An efficient one-step site-directed and site-saturation mutagenesis protocol. Nucleic Acids Res 32:e115
Zheng RC, Zheng YG, Shen YC (2007) A screening system for active and enantioselective amidase based on its acyl transfer activity. Appl Microbiol Biotechnol 74:256–262
Acknowledgments
This work was financially supported by the National High Technology Research and Development Program of China (No. 2012AA022201B), National Natural Science Foundation of China (No. 21176224), and Natural Science Foundation of Zhejiang Province of China (Nos. Z4080032 and R3110155).
Conflict of interest
The authors declare that they have no financial or non-financial conflict of interest in the publication of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Yu-Guo Zheng holds a PhD, Zhejiang University of Technology.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 266 kb)
Rights and permissions
About this article
Cite this article
Wan, NW., Liu, ZQ., Xue, F. et al. An efficient high-throughput screening assay for rapid directed evolution of halohydrin dehalogenase for preparation of β-substituted alcohols. Appl Microbiol Biotechnol 99, 4019–4029 (2015). https://doi.org/10.1007/s00253-015-6527-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6527-z