Abstract
While a number of amphibian skin microbiomes have been characterized, it is unclear how these communities might vary in response to seasonal changes in the environment and the corresponding behaviors that many amphibians exhibit. Given recent studies demonstrating the importance of the skin microbiome in frog innate immune defense against pathogens, investigating how changes in the environment impact the microbial species present will provide a better understanding of conditions that may alter host susceptibility to pathogens in their environment. We sampled the bacterial skin microbiome of North American wood frogs (Rana sylvatica) from two breeding ponds in the spring, along with the bacterial community present in their vernal breeding pools, and frogs from the nearby forest floor in the summer and fall to determine whether community composition differs by sex, vernal pond site, or temporally across season (spring, summer, fall). Taxon relative abundance data reveals a profile of bacterial phyla similar to those previously described on anuran skin, with Proteobacteria, Bacteroidetes, and Actinobacteria dominating the wood frog skin microbiome. Our results indicate that sex had no significant effect on skin microbiota diversity; however, this may be due to our limited female frog sample size. Vernal pool site had a small but significant effect on skin microbiota, but skin-associated communities were more similar to each other than to the communities observed in the frogs’ respective pond water. Across seasons, diversity analyses suggest that there are significant differences between the bacterial skin microbiome of frogs from spring and summer/fall groups while the average α-diversity per frog remained consistent. These results illustrate seasonal variation in wood frog skin microbiome structure and highlight the importance of considering temporal trends in an amphibian microbiome, particularly for species whose life history requires recurrent shifts in habitat and behavior.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amphibians host unique communities of skin-dwelling microbes regulated by various mechanisms, including inoculation from the environment, skin-sloughing, and specific skin secretions [1, 2]. Studies of the bacterial microbiome of amphibian skin have revealed that while many bacterial phyla are present, species from Acidobacteria, Actinobacteria, Bacteroidetes, Cyanobacteria, Firmicutes, and Proteobacteria tend to dominate [1, 3,4,5]. Many families and genera within these phyla are present on a wide range of amphibian species, but the specific species tend to be unique to the host. Additionally, species which dominate the skin-associated community are not generally those which are dominant in the environment at large and therefore must be selected for by the skin microenvironment [1]. Further microbiome studies have shown that frogs of the same species sampled during different seasons or from different habitats can have large differences in the diversity of bacteria present on their skin [4, 6, 7], underscoring the influence of environmental factors on the microbial community. It is currently unclear how dramatic seasonal differences in climate and host behavior might influence the amphibian skin microbiome.
Although the functional significance of amphibian skin microbiota are not yet well defined, recent studies have highlighted the importance of the skin commensal bacterial microbiome to the amphibian host’s defense mechanisms against invading pathogens [2, 8,9,10]. For example, some bacterial species commonly found on the skin of amphibians inhibit the growth of the pathogenic fungi Batrachochytrium dendrobatidis [11,12,13], the causative agent of amphibian chytridiomycosis, and proximate cause of amphibian declines on multiple continents [14, 15]. These findings have spurred investigation into the utility of transferring or inoculating bacteria onto the skin of susceptible frogs as a means to mitigate or protect against B. dendrobatidis infection [16, 17]. These studies highlight the importance of understanding the characteristics of the amphibian skin microbiome and their contribution to skin innate immune functions in defense against pathogens in their environments. Thus, understanding how frog skin microbial community composition varies across environmental conditions will provide important information to aid in predicting environments where amphibians may be more or less susceptible to pathogens.
The North American wood frog (Rana sylvatica) is widespread with a range extending through most of Canada, Alaska, and the Northeastern USA [18]. R. sylvatica inhabits uplands environments and the far north where few, if any, other frog species inhabit. Wood frogs breed in temporary pools in the early spring, leave these pools to migrate upland into the terrestrial forest environment during warmer months [19], and hibernate on the forest floor in winter, routinely surviving multiple sustained bouts of whole body freezing [20, 21]. R. sylvatica is susceptible to infection by Frog virus 3 and B. dendrobatidis [22, 23], pathogens which threaten amphibian populations worldwide. Despite the wide range of R. sylvatica and its known susceptibility to pathogens of concern, no comprehensive study of skin microbiota has been performed on this species. Given the seasonal shifts in habitat and behavior experienced by R. sylvatica, they are well-suited to investigate whether seasonal changes affect skin microbiome composition and lay the foundation for investigations into the potential implications in susceptibility to amphibian pathogens, as has been explored in other species [24,25,26].
The objectives of this study are to (1) identify the composition of the R. sylvatica bacterial skin microbiome and (2) determine whether bacterial community structure varies with sex, vernal pool of origin or across seasons. We hypothesized that the wood frogs carry a range of bacterial phyla similar to those found on the skin of other frog species, with variation in representation and relative abundance of bacterial taxa between individuals to reflect separate ponds of origin and season of capture. To determine whether this is the case, we have analyzed the bacterial community present on the skin of R. sylvatica using 16S rRNA gene amplicon sequencing. Skin swabs were obtained from frogs captured from two spatially separated ponds in the spring and the surrounding forest in summer and fall so that skin-associated bacterial communities might be compared on a seasonal basis, as well as between different capture sites.
Materials and Methods
Experimental Design and Sample Collection
Wild R. sylvatica adults were sampled from a site in the Waterloo Region of Ontario, Canada, during the spring (April–May), summer (July–August), and fall (October). In the spring, individuals were captured from two vernal ponds spatially separated by ~ 200 m, herein referred to as pond 1 and pond 2, while individuals were captured from the surrounding forest floor during the summer and fall seasons. Frog skin microbiota sample sizes were dependent on the number of frogs that could be captured successfully. Overall, 27 frogs were captured in Spring 2018, 10 in Summer 2018, 7 in Fall 2018, and 22 in Spring 2019. Seasonal sample counts per sex and capture site are given in Table 1. Due to sampling limitations, few female frogs (n = 8) were included compared with the number of male frogs (n = 58). All female frogs were captured during 2018, with the majority (n = 6) captured during Spring 2018.
Individuals were captured by nets and each frog was handled with a new pair of sterile nitrile gloves. Wood frogs were gently rinsed with sterile distilled water to remove transient microbes. To collect resident microbes, frogs were swabbed with a sterile rayon-tipped applicator (Puritan Medical Products Company, LLC., Guilford, ME, USA), 12 times on both the dorsal and ventral surfaces, covering as much of the skin surface as possible. To control for environmental microbes that may have deposited onto the swab during the sampling process, “field control” swabs were produced by wetting a clean rayon swab with sterile distilled water and mimicking the swabbing action in the open air. Each rayon swab head was placed into a sterile 1.7-mL microfuge tube, and the applicator stick was cut just above the rayon tip using flame-sterilized scissors. Samples were transported on ice prior to storage at − 80 °C. Animal care and handling was performed in accordance with the guidelines of the University of Waterloo Animal Care Committee and the Canadian Council on Animal Care (Animal Utilization Projects #30008 and #40721), and animals were captured under the Ontario Ministry of Natural Resources and Forestry Wildlife Scientific Collectors Authorization Permits (#1088586 and #1092603) issued to Dr. B.A. Katzenback.
When individuals were captured from vernal pools, a water sample was taken by pushing 50 mL of pond water, taken from just below the surface of the water, through a Sterivex-GP PES 0.22-μm filter (Millipore, Burlington, MA, USA) using a 50-mL syringe (Fisher). The filter units were disconnected from the syringe, placed in individual sterile 50 mL conical tubes (FroggaBio), and held on ice prior to storage at − 80 °C.
DNA Isolation and Amplicon Sequencing
DNA isolation was performed using the DNeasy PowerSoil Kit (QIAGEN Inc., Venlo, Netherlands) according to the manufacturer’s protocol. Frog skin swab samples or field controls were removed from − 80 °C and immediately transferred from their storage tubes into PowerBead tubes. DNA was isolated from vernal pool microbiota filtride by removing the filter paper from the cartridge and cutting the filter paper into thin strips using flame-sterilized scissors before addition to the PowerBead tubes. To control for bacterial contamination from the laboratory environment and/or the extraction kit components, a clean rayon swab was wet with sterile distilled water and cut into a labelled 1.7-mL microfuge tube with flame sterilized scissors to act as a “process control” and was processed alongside the samples. All samples (skin swabs, vernal pool filters, field controls, process controls) were immediately vortexed for 10 s after transfer to the PowerBead tubes prior to following the manufacturer’s protocol. Isolated DNA was eluted in the provided elution buffer and stored at − 80 °C.
To check for successful extraction of DNA from skin swabs prior to gene amplicon sequencing, an aliquot of each sample’s DNA was PCR amplified. Each reaction contained 18.875 μL of molecular biology grade water, 2.5 μL of 10× PCR buffer, 0.5 μL of 10 μM dNTPs, 1 μL of 10 μM forward primer 515FB 5′-GTGYCAGCMGCCGCGGTAA-3′ [27, 28], 1 μL of 10 μM reverse primer 926R 5′-CCGYCAATTYMTTTRAGTTT-3′ [27, 29], and 0.125 μL Taq DNA polymerase (5 units/μL) (GeneDirex) per 1 μL of sample, for a total reaction volume of 25 μL. PCR was performed with the following cycling conditions: 94 °C for 3 min, then 35 cycles of 94 °C for 45 s, 50 °C for 1 min, and 72 °C for 1.5 min, followed by extension at 72 °C for 5 min. PCR product were separated by electrophoresis at 130 V for 30 min on 2% agarose gel in TAE buffer and visualized using SafeRed dye and trans-UV (302 nm) imaging in a ChemiDoc XRS+ (Bio-Rad).
Once bacterial DNA extraction from skin swabs was verified by PCR, the original extracted DNA samples were sent for 16S rRNA gene amplicon sequencing by MetagenomBio (Waterloo, ON, Canada). PCR reactions were prepared in triplicate for each sample. Each 25 μL reaction mixture contained 1.6 μL of molecular grade water, 0.2 μL of BSA (20 mg/mL), 2.5 μL of 10× standard Taq buffer, 0.5 μL of 10 mM dNTPs, 5.0 μL of 1 μM forward primer 515FB 5′-GTGYCAGCMGCCGCGGTAA-3′ [27, 28], 5.0 μL of 1 μM reverse primer 806RB 5′-GGACTACNVGGGTWTCTAAT-3′ [27, 28], 0.2 μL of Taq DNA polymerase (5 units/μL), and 10 μL of sample DNA. PCR was performed with the following thermocycling conditions: 95 °C for 5 min, 35 cycles of 95 °C for 30 s, 50 °C for 30 s, and 72 °C for 60 s, followed by an extension at 72 °C for 10 min. The products of the triplicate reactions were pooled and resolved with 2% TAE agarose gel. PCR products of the correct amplicon size (~ 291 bp) were excised, pooled, gel purified, and quantified using the Invitrogen™ Qubit™ dsDNA HS Assay Kit (Thermo Fisher Scientific Inc., Waltham, MA, USA). For all 2018 samples, sequencing was performed using an Illumina MiSeq and the MiSeq Reagent Kit v2 (Illumina, Inc., San Diego, CA, USA) for 2 sets of 250 cycles. This was increased to 3 sets of 250 cycles for all 2019 samples due to an error at the sequencing center.
Amplicon Sequence Data Processing
Sequence data was obtained as FASTQ files in the CASAVA 1.8 paired-end demultiplexed format. Files from repeat sequencing runs were concatenated to create a FASTQ file containing all of the observed sequences for each sample. These files were imported into QIIME 2 v2019.1.0 [30], and all analysis was performed using QIIME 2 unless otherwise stated.
Using DADA2 [31], reads were trimmed by 25 bp on the 5′ end to remove the primer sequence and truncated to 245 bp to remove low-quality regions, filtering out any reads shorter than this length. The reads were dereplicated and denoised, and any chimeric sequences were removed. Paired forward and reverse reads were merged, generating the final amplicon sequence variants (ASVs). Each ASV was assigned taxonomy using a naïve Bayesian classifier trained on the SSU Ref NR 99 dataset from the SILVA 132 release [32], with sequences trimmed to include only the V4-V5 region. All unassigned ASVs and those assigned to chloroplast or mitochondria were filtered from the samples. To minimize erroneous ASVs, a minimum frequency of 22 (0.001% of the total sequence count) was set, and any ASVs with a total frequency less than 22 were filtered from the samples.
Taxonomic Assignment and Significance
A multiple sequence alignment (MSA) was produced from the ASVs using MAFFT [33]. Columns of the alignment which were ambiguously aligned were masked to avoid introducing error to the phylogenetic model. A phylogenetic tree was generated from the MSA using RAxML [34] with 100 bootstraps. The RAxML tree was assigned a midpoint root and used for all further phylogenetic analysis.
Core bacterial taxa were determined for each group of seasonal frog skin communities and for the overall frog skin microbiome, using the core-features command in QIIME2. Core taxa were defined as those found to be present in 90% or more of the samples within a given group, as this is a commonly used threshold [6, 35] which meets the definition of core microbiota as those which are commonly present within samples from a given environment [36]. Venn diagrams depicting the overlap of seasonally core microbiota were prepared using the online Venn diagram tool provided by the University of Ghent (http://bioinformatics.psb.ugent.be/webtools/Venn/).
Statistical Analyses
Differences in the relative abundance of core bacterial families were compared among seasonal groups using a Kruskal-Wallis test followed by Dunn’s multiple comparison test to determine which groups differed significantly.
For diversity analyses, samples were rarified to 10,000 sequences. Those containing less than 10,000 were omitted from diversity analyses. The phylogenetic tree and rarified ASV frequencies were used to calculate various α-diversity metrics (Faith’s phylogenetic diversity, Shannon diversity index, ASV richness). Diversity was compared among groups using a Kruskal-Wallis test followed by Dunn’s multiple comparison test.
To assess differences in bacterial community composition among samples, two β-diversity metrics were calculated (unweighted UniFrac and weighted UniFrac). These metrics were chosen as they each consider different aspects of the community, considering phylogenetic distance between observed ASVs as well as relative abundance of ASVs (weighted UniFrac) or the presence/absence of ASVs (unweighted UniFrac). An in-depth core microbiome analysis was performed to compare groups based on presence/absence of ASVs in the absence of phylogenetic distance information, and therefore, a distance metric accounting for these qualities was not included. β-diversity was visualized using principal coordinate analysis (PCoA). The adonis function from the R vegan package was used to perform permutational multivariate analysis of variance (PERMANOVA) with 999 permutations. To determine whether groups had significant differences in bacterial microbiome composition, pairwise PERMANOVA tests were applied to the dissimilarity matrix produced by each β-diversity metric and performed with 999 permutations.
Results
Bacterial Microbiome Overview Statistics
We obtained microbial 16S rRNA amplicon sequences from each of the 66 frog swabs, 5 water samples, 5 field blanks, and 3 process blanks. This was despite a lack of visible bands in the initial PCR amplification for the field and process blanks, as well as faint bands for many samples. After filtering based on quality, taxonomy, and minimum frequency, a total of 1,937,866 sequences remained. Sequence counts varied between samples considerably, and in particular, those samples sequenced in the later set (Spring 2019 frog swabs, water samples and associated blanks) had much higher counts on average due to their increased sequencing depth. A total of 4325 ASVs were recognized, which ranged in frequency from 22 to 159,313 appearances, with a median frequency of 60. These ASVs were matched to 1384 unique taxonomic assignments, 1123 of which were specific to at least the genus level. While nearly all of the ASVs detected belonged to bacterial taxa, ASVs from two archaeal taxa were also present. Archaeal ASVs accounted for only 0.01% of the sequences detected, and these taxa appear to be rarities in an overwhelmingly bacterial community. The relative abundances of ASVs belonging to the various prokaryotic phyla detected in each sample are displayed in Fig. 1.
We found that DNA isolated from process controls yielded ASVs belonging to 9 bacterial phyla and contained 37 ASVs each on average, which equated to a total of 71 unique taxa when combined at the genus level (Supplementary Table 1). Unlike the communities found on frog skin and in pond water, the majority of ASVs found in the process controls belonged to a limited group of genera, with Curtobacterium being particularly dominant (43% relative abundance). The field controls generally had more diverse communities, with an average of 64 ASVs per blank. These ASVs belonged to a total of 15 phyla and 164 unique taxa when combined at the genus level (Supplementary Table 2). On average, 11% of amplicons detected in frog skin samples and 0.07% of amplicons detected in water samples were ASVs which were also found in the process controls. A group of 7 frog skin samples, including 2 samples from Summer 2018 and 5 from Spring 2019, had a considerably higher proportion of ASVs (> 50% relative abundance) overlapping with those found in the process control and were determined to be outliers using an iterative Grubb’s test (α = 0.05). These samples had a visibly different community composition from other samples taken from the same site and season and were regarded as contaminated or otherwise unlikely to represent an accurate depiction of the skin microbiome. These samples are included in Fig. 1, distinguished by an “x,” but were removed prior to further analyses.
Capture Site Influences the Structure of Bacterial Communities
Spring was the only season in which sampling occurred across two distinct sites, the two vernal pools. It was thus necessary to investigate how pond of origin might influence an individual’s skin bacterial community. We observed that the skin bacterial microbiota of wood frogs captured from pond 1 and pond 2 had very similar α-diversity metrics, and both of the skin-associated communities had higher average bacterial diversity than the bacterial communities present in the water samples taken from either vernal pond (Fig. 2). When we sorted frog skin microbiota and vernal pond microbiota samples from Spring 2018 and Spring 2019 into groups based on their pond of origin, there were significant differences in α-diversity (Kruskal-Wallis: ASV richness: p = 0.0087; Shannon diversity index: p = 0.0075; Faith’s phylogenetic diversity: p = 0.0259). Differences were between the pond 2 water and the pond 1 frog skin (Dunn’s: ASV richness: p = 0.1297; Shannon diversity index: p = 0.0.0495; Faith’s phylogenetic diversity: p = 0.0758) and between pond 2 water and pond 2 frog skin (pairwise ANOVA: ASV richness: p = 0.0128; Shannon diversity index: p = 0.0467; Faith’s phylogenetic diversity: p = 0.0267). The multiple comparison test indicated that there were no significant differences in α-diversity between pond 1 water and pond 1 frog skin, pond 1 water and pond 2 frog skin, pond 1 water and pond 2 water, or pond 1 frog skin and pond 2 frog skin (Fig. 2).
The microbial communities present in each group of frog skin samples and water samples included many of the same bacterial phyla, with 12 of the 32 phyla detected among the groups being present in at least one sample from each group. Samples from all groups had a high relative proportion of ASVs belonging to Proteobacteria, Bacteroidetes, and Actinobacteria on average (Fig. 1). These three phyla were the only phyla we found to have a high relative abundance in the water samples (Proteobacteria 73%, Bacteroidetes 18%, Actinobacteria 8%), while all other phyla present had a mean relative abundance of less than 1%. The frog skin communities from both ponds also had Acidobacteria and Verrucomicrobia ASVs present at greater than 1% mean relative abundance, and the pond 2 frog skin community additionally had Firmicutes ASVs present at more than 1% mean relative abundance. We employed a Kruskal-Wallis test and Dunn’s multiple comparison test to compare the relative abundance of each of these 6 common phyla between the water samples and frog skin samples from both ponds and found that the relative abundance of Firmicutes was significantly greater for frogs from pond 2 than water from both ponds (Dunn’s: P1 water, p = 0.0192; P2 water p = 0.0484) and that both groups of frogs had significantly higher relative abundance of Verrucomicrobia on their skin than water from pond 2 (Dunn’s: P1 frogs, p = 0.0436; P2 frogs p = 0.0138). Beyond the most abundant phyla, the frog skin bacterial microbiota exhibited a more diverse range of phyla compared with vernal pool bacterial microbiota; the skin-associated bacterial communities from frogs captured from pond 1 and pond 2 included ASVs from phyla that were not present in any pond water samples.
Sample type (frog skin, vernal pond), site (pond 1, pond 2), and year all had significant (p < 0.005) but small (R2 < 0.15) effects on community composition and relative abundance of the observed taxa when assessed using adonis (Table 2). Sample type had the largest effect size, as observed with unweighted UniFrac distance (Table 2a, adonis pseudo-F = 4.22, p < 0.001, R2 = 0.08) and weighted UniFrac distance (Table 2b, adonis pseudo-F = 7.97, p < 0.001, R2 = 0.14). Principal coordinate analysis of the β-diversity metrics indicated that many frog skin communities were similar despite different ponds of origin (Fig. 3). Water samples were not clearly distinguished from frog samples (Fig. 3a, b). Given that the effect of site was relatively minor and the bacterial communities of frogs from ponds 1 and 2 were largely similar, we combined these groups in further analysis.
Principal coordinate analysis of β-diversity of spring frog skin and pond water microbiome samples. Principal coordinate analysis plots were created using Emperor from distance matrices calculated using a sampling depth of 10,000. Plots were limited to representing the two dimensions with the highest percent variation explained and were calculated for a unweighted UniFrac distances and b weighted UniFrac distances
Host Sex May Not Affect Diversity and Structure of Bacterial Communities
Due to the low number of female frogs, most of which were captured during spring, it is difficult to come to definitive answers regarding sex-dependent microbiome characteristics and in particular to distinguish sex-dependent effects from seasonal differences. We found that the sex of the frog did not significantly affect diversity or structure of the skin bacterial community by most metrics. Comparing the α-diversity metrics of all male and female frogs using a Mann-Whitney test resulted in no significant difference for any metric observed (ASV richness: p = 0.3561; Shannon diversity index: p = 0.8359; Faith’s phylogenetic diversity: p = 0.4861). Additionally, sex was not a significant driver of community structure when assessed using adonis for the unweighted UniFrac β-diversity metric (pseudo-F = 1.13, p = 0.238, R2 = 0.022). Sex did appear to have a very small, but significant effect when considering weighted UniFrac (pseudo-F = 2.17, p = 0.035, R2 = 0.041).
Season of Capture Influences the Structure of Bacterial Communities
To assess the influence of season on the bacterial skin microbiome, the frog samples were divided into groups based on capture date (Spring 2018, Summer 2018, Fall 2018, and Spring 2019). The bacterial communities of frog skin from each seasonal group were similarly rich and even (Fig. 4). Of the three α-diversity metrics we considered, two differed significantly between seasonal sample groups (Kruskal-Wallis: ASV richness, p < 0.0002; Faith’s phylogenetic diversity, p < 0.0038). In both cases, the mean diversity of the Spring 2019 group was significantly higher than that of Spring 2018 (Dunn’s: ASV richness, p < 0.0001; Faith’s phylogenetic diversity, p < 0.0033), while no other groups had significantly different means.
Comparison of seasonal wood frog skin microbiota α-diversity metrics. Sample α-diversity was calculated using a sampling depth of 10,000. Mean seasonal value and standard deviation of each group is shown. Results are given for a ASV richness, b Shannon diversity index, and c Faith’s phylogenetic diversity
We found that community structure differed between seasonal groups, including observable trends at the highest taxonomic levels (Fig. 1). While the majority of ASVs present in any given sample were typically Proteobacteria, the relative abundances of ASVs belonging to other major phyla such as Bacteroidetes, Actinobacteria, Verrucomicrobia, and Acidobacteria varied considerably (Fig. 1). We assessed the variation in relative abundances of the 5 major phyla between seasonal groups using a Kruskal-Wallis test. All groups exhibited some seasonal variation (Acidobacteria, p < 0.0001; Actinobacteria, p < 0.0001; Bacteroidetes, p = 0.0026; Proteobacteria, p < 0.0001; Verrucomicrobia, p = 0.0106). Acidobacteria and Actinobacteria were both present at significantly higher relative abundances in Summer and Fall 2018 than in either Spring 2018 or 2019 (Dunn’s: Acidobacteria, p = 0.0074–0.0026; Actinobacteria, p = 0.0162–0.0001). Acidobacteria had a mean relative abundance of 5.9% in Summer 2018 and 6.4% in Fall 2018, contrasting with 1.9% in Spring 2018 and 0.7% in Spring 2019. Similarly, while Actinobacteria had a mean relative abundance of 18% in Summer 2018 and 15% in Fall 2018, only relative abundances of 3.5% in Spring 2018 and 4.7% in Spring 2019 were observed. Additionally, Bacteroidetes had a significantly lower mean relative abundance in Fall 2018 as compared with both Spring 2018 and 2019 groups (Dunn’s: p = 0.0445 and 0.0012, respectively) and Proteobacteria had a significantly lower mean relative abundance in Summer 2018 as compared with both Spring 2018 and 2019 groups (Dunn’s: p < 0.0001 and 0.0033, respectively), but no other clear trends existed.
Season was a significant source of variation in community structure when measured using unweighted UniFrac (Table 3a, adonis pseudo-F = 4.01, p < 0.001, R2 = 0.14) and weighted UniFrac (Table 3b, adonis pseudo-F = 8.82, p < 0.001, R2 = 0.26). We performed pairwise PERMANOVAs to determine which seasonal groups had significantly different community structures and every pairwise comparison revealed a significant difference across all β-diversity metrics (PERMANOVA: p = 0.001), with the exception of the Summer and Fall 2018 frog skin microbiome groups which were not significantly different when considering unweighted UniFrac distance (PERMANOVA: pseudo-F = 1.36, p = 0.108) or weighted UniFrac distance (PERMANOVA: pseudo-F = 1.54, p = 0.260). This result was reflected in the principle coordinate analysis results (Fig. 5), where skin microbiota from frogs collected during the summer and fall tended to be similar, but were distinct from the spring samples when unweighted UniFrac (Fig. 5a) was considered, although summer and fall samples were not clearly distinct from spring samples when visualizing weighted UniFrac distances (Fig. 5b).
Principal coordinate analysis of β-diversity of frog skin microbiome samples. Principal coordinate analysis plots were created using Emperor from distance matrices calculated using a sampling depth of 10,000. Plots were limited to representing the two dimensions with the highest percent variation explained and were calculated for a unweighted UniFrac distances and b weighted UniFrac distances
Few Bacterial Taxa Are Core to All Seasons
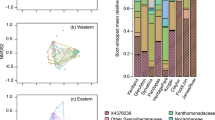
We identified core microbiota for all frog skin samples as a whole, as well as for each seasonal group separately. No 16S rRNA gene ASV was present in ≥ 90% of all frog skin microbiome samples, so ASVs were collapsed at the genus and family level to consider a more inclusive core microbiome. A group of 6 genera were present in ≥ 90% of all frog samples, accounting for approximately 10% of ASVs on the average individual (Table 4). A group of 12 families were present in ≥ 90% of all frog samples, accounting for 60% of ASVs on the average individual (Table 5). We analyzed the seasonal variation in relative abundance the ten core families which had a mean relative abundance greater than 1% (Fig. 6), revealing five bacterial families with significantly different mean relative abundance across seasonal groups: Burkholderiaceae, Chitinophagaceae, Sphingobacteriaceae, Spirosomaceae, and Xanthobacteracea (Dunn’s: p < 0.05). In all cases, Summer and Fall 2018 relative abundances did not differ significantly. Similarly, Spring 2018 and 2019 significantly differed only in the relative abundance of Chitinophagaceae (Dunn’s: p = 0.0462) and Sphingobacteriaceae (Dunn’s: p = 0.0006).
Relative abundance of core microbial families in wood frog skin microbiota. Families included were present in ≥ 90% of all frog skin samples and had a mean relative abundance ≥ 1%. Relative abundance of each family was compared between seasonal groups using a Kruskal-Wallis test and Dunn’s multiple comparison test, letters are used to indicate significant inter-seasonal variation for a given family. Seasons marked with the same letter do not significantly differ
When considering each seasonal group of skin associated communities separately, it was revealed that 25 of the 50 core genera and 18 of the 45 core families were considered core during only one season (Fig. 7). Only two genera were core to all seasonal groups (Fig. 7a), indicating that the six core genera observed when considering all frogs together were not reflective of the core genera present during each individual season. The Spring 2018 and Spring 2019 frog skin bacterial communities had the most core genera in common (20 genera core to both groups), while Summer 2018 and Fall 2018 had the second highest level of overlap (5 genera core to both groups). Additionally, the frog skin microbiota from Spring 2018 and Spring 2019 each had a larger number of in-season core genera, with 20 in Spring 2018 and 35 in Spring 2019 as compared with 12 in Summer 2018 and 14 in Fall 2018. In comparison, a group of 8 families were core across all seasonal groups (Fig. 7b), but the consistent prevalence of these families was not always matched with a high relative abundance. We found that Burkholderiaceae was the only family to consistently have a mean relative abundance above 5% on wood frog skin, ranging from 9% in Summer 2018 to 44% in Spring 2018. Interestingly, there were 9 core families unique to the Fall 2018 wood frog skin group, while other seasonal groups had only 2–4 uniquely core families. Fall 2018 wood frog bacterial skin microbiomes also had the most core families overall at 26, followed by Spring 2019 with 25, Summer 2018 with 21, and Spring 2018 with 19. In all seasons except for Spring 2018, the core families present on wood frog skin included members of the phyla Actinobacteria, Bacteroidetes, Proteobacteria, and Verrucomicrobia. In Summer 2018, a single family belonging to Acidobacteria was considered core. The core families present on the skin of wood frogs in Spring 2018 lacked any member of Actinobacteria but included a single representative from the Gemmatimonadetes. In all cases, the majority of core families present on wood frog skin were from the Proteobacteria, which ranged from 41 to 68% summed relative abundance. In general, community structure appeared to be more consistent for bacterial families than for individual genera.
Discussion
As we continue to improve our understanding of the skin microbiome and its role in maintaining the health of amphibians, it is important to consider the inherent variability in bacterial microbial communities and the factors that drive this variability. Many amphibians experience major seasonal fluctuations in environmental conditions, and environmental conditions are known to influence the microbiome [4, 6, 7], potentially affecting the hosts ability to resist infection by a pathogen [37, 38]. In this study, we have reported on the bacterial community associated with R. sylvatica skin over the course of multiple seasons to observe the changes in community composition and structure. To provide perspective for these seasonal effects, we have compared them to the effects of host sex and vernal pool site of capture within a single season (spring).
Bacterial Community Structure and Core Taxa
We observed that the bacterial community associated with R. sylvatica skin has much in common with those found on other frog species. ASVs belonging to Proteobacteria, Actinobacteria, Bacteroidetes, Firmicutes, Verrucomicrobia, and Acidobacteria made up the vast majority of sequences observed from all samples, suggesting that these phyla dominate the bacterial microbiome. These phyla are commonly present on the skin of other frog species [3,4,5, 8], and all but Verrucomicrobia are abundant elements of the microbial communities associated with the related frog species Rana pipiens and Rana catesbeiana [1, 39]. While these phyla varied in relative abundance on the skin of individual wood frogs, there were very few cases in which any of the above bacterial phyla were found to be absent from the wood frog skin microbiota. Communities were much less consistent at finer taxonomic levels, and among the hundreds of bacterial genera observed, very few were prevalent enough to be considered core taxa. The most abundant of these core genera, Sphingomonas, is widespread in the environment at large and has been shown to be similarly abundant on the skin of other frogs [1, 6, 39].
Effect of Vernal Pool Site and Host Sex on the Wood Frog Bacterial Skin Microbiome
While R. sylvatica are generally terrestrial and solitary, wood frogs converge on vernal pools during the spring thaw to seek mates and reproduce [19]. In this study, the two temporary ponds from which frogs were captured during the spring served as the only truly distinct sampling sites, since the surrounding area was fairly uniform mixed woodland. As R. sylvatica are known to venture as far as 1 km from their breeding pond and the ponds sampled are ~ 200 m apart, it is unlikely that they harbor genetically distinct populations [40], and therefore, any variation in the skin-associated bacterial community is better attributed to the environmental conditions of the site. This is an important distinction, as it is not well understood to what degree host phylogenetics and environment affect the microbiome of amphibians, and examples exist which emphasize the role of both factors [41,42,43]. Pond of origin was found to explain a small, but significant amount of the variation observed in the bacterial communities on frogs captured during the spring. This effect was less pronounced than the variation between frogs captured in spring of 2018 and 2019 however, and the largest differences were observed between frog swabs and water samples. This was expected, as previous studies have established that amphibians have communities of skin-associated microbiota distinct from their environment [1, 39, 44]. It is worth noting that samples taken from the water column would likely not include the entire microbial community found in a pond environment, and the communities associated with pond sediment or vegetation may better reflect the community found on frog skin. There was no clear trend linking the bacterial communities of frog skin microbiomes to the microbiota found in the water of their pond of capture. There was no significant difference in ASV richness, evenness, or phylogenetic diversity between the skin-associated bacterial communities of frogs in pond 1 and pond 2. Despite water samples from pond 2 having a lower mean ASV richness than those of pond 1, frogs from pond 2 had more ASVs on average, suggesting that seeding of bacteria from the water to the skin was not a major driver of community diversity. Frogs from both ponds hosted bacterial phyla that were uncommon, or not present, in the pond water and exhibited more diverse and even communities, while water samples were almost entirely populated by Proteobacteria, Bacteroidetes, and Actinobacteria. Given the increased relative abundance of many of these phyla on frogs captured during the summer and fall, it seems that they must either be stable members of the microbiome or are seeded from rich microbial communities found in the soil and leaf litter of the surrounding forest. It is unclear whether seeding from soil environments might occur while buried during winter hibernation, and a study of the microbial communities present in frog hibernacula, although challenging, would be an interesting avenue of future research.
The effect of frog sex on the bacterial microbiome was also considered, as sex of an individual has been shown to influence the microbiome in other frog species and was observed to interact with environmental factors to produce significant effects [45]. These findings from studies on other frog species coincide with previous observations of sex-dependent microbiome characteristics in humans [46] and other vertebrates [47]. In the present study, we found that sex did not have a significant effect, or had only a very small effect, on the structure or diversity of the microbial community for the metrics considered and observed no clearly sex-linked trends. Several previous studies considering the effect of sex on amphibian microbiomes have also failed to find significant differences between males and females [41, 48], and our work corroborates these findings, although it is limited by the low number of female frogs and does not consider the effect of sex in the context of interaction with other factors.
Effect of Season on the Wood Frog Bacterial Skin Microbiome
Season was also associated with significant variation in the structure and composition of the bacterial communities present on wood frog skin. The effect of season was more pronounced than the effects of site, year, or sample type observed in the spring samples and was evidenced by shifts in the relative abundances of major phyla on wood frog skin. Most notably, relative abundances of Acidobacteria and Actinobacteria were significantly higher among frogs captured in the summer and fall than those captured during spring. Given the particularly high relative abundance of these phyla in soils [49, 50], it is not surprising that members would be highly abundant on frogs active in soil and leaf litter. The seasonality of Acidobacteria and Actinobacteria on frog skin matches a proposal that Acidobacteria are transiently associated with the human skin microbiome [51]. While the composition of the frog skin microbiome varied, average diversity of individual R. sylvatica bacterial microbiomes remained fairly constant across seasons. When considering ASV richness and phylogenetic diversity, the Spring 2019 wood frog skin microbiota group was determined to have a mean diversity significantly higher than Spring 2018 and Summer 2018. However, when considering the Shannon diversity index, wood frog skin microbiomes sampled during Spring 2019 fall well within the range of these groups. This suggests that the Spring 2019 wood frog bacterial communities were not as even, and a larger proportion of uncommon ASVs were contributing to their diversity. This is likely at least partially the result of the Spring 2019 samples undergoing an additional sequencing replicate. The resultant greater sequencing depth would increase the number of rare ASVs [52] and contribute to the increase in observed diversity. Rarefaction prior to diversity analyses was conducted to mitigate this issue, but it is not a perfect method [53].
Due to the much higher number of wood frog microbiome samples collected during the spring months (75% of total frog swab samples), any analysis considering overall prevalence of taxa was heavily biased toward taxa which were common during the spring. To better represent the bacterial communities present on wood frog skin during summer and fall samples, core taxa were considered for each seasonal group individually and overlaps in seasonal groups’ core taxa were determined. As core taxa represent the microbes which are most commonly found in the frog skin environment and often represent key members of the microbial community [36], common core taxa should reflect similar community dynamics. The skin of R. sylvatica hosted only a small number of core microbiota, particularly at lower taxonomic levels. Several of the core taxa are known to be core to other frog skin communities, Pseudomonas being one of the most commonly represented [6, 35, 39], but the majority of core taxa appeared to be fairly unique across seasons. Additionally, many of the most prevalent taxa experienced significant changes in relative abundance between seasons. Xanthobacteraceae were significantly more relatively abundant on the frog skin in summer and fall, while Spirosomaceae significantly increased in relative abundance on frog skin during the spring seasons only. The variability in the core bacterial taxa observed on R. sylvatica skin suggests that the skin microbiome is a highly dynamic environment, where seasonal factors can re-shape the core structure and few microbes are suited to inhabit the skin year-round.
While this study did not attempt to isolate the specific factors driving seasonal change in the frog skin microbiome, there are a number of potential reasons that shifts would occur. As amphibians are ectothermic, seasonal changes in temperature undoubtedly play a role. Variation in temperature has been shown to have community-scale effects on the amphibian microbiome [54, 55], the effects of which are complex, as varying temperatures also affect the microbial communities of soil [56] and the immune function of frogs [57, 58]. Additionally, the skin microbiome of several amphibian species has been observed to vary significantly between populations inhabiting different environments [6, 45], and in some cases, association with particular microhabitats may contribute to the skin microbial community significantly more than host phylogeny does [44]. Clearly, the skin-associated microbial community is a complex system and seasonal trends are likely to be multifactorial.
Conclusions
Our results indicate that season has a significant effect on the structure of the North American wood frog bacterial skin microbiome and has a proportionally greater effect than spring breeding pond association. Frogs captured during summer and fall were the most similar in terms of β-diversity distances and could be distinguished from spring frogs by their increased relative abundance of Acidobacteria and Actinobacteria, as well as other soil-associated bacterial families. It remains unclear whether the shift toward increased relative abundance of soil-associated bacteria on frog skin in the summer and fall is a result of transient colonization from frequent exposure or a stable equilibrium shift in the community. Skin-associated bacterial communities had consistent structural similarities at the highest taxonomic levels but displayed a high degree of diversity at finer levels, and the few core genera identified were not a dominant component of the community. R. sylvatica is a widespread species, and further study of populations from varied environments (Boreal shield, montane forest, etc.) could reveal related trends. While the effect of season has been explored in other temperate frog species [4], this study provides insight into the seasonality of skin microbiome structure on amphibian species inhabiting northern environments and establishes foundational knowledge for further study of species which experience dramatic shifts in habitat and behavior between seasons.
Data Availability
16S rRNA gene amplicon sequence data for skin microbiome samples are deposited in the NCBI Sequence Read Archive (Bioproject PRJNA603391).
References
McKenzie VJ, Bowers RM, Fierer N et al (2012) Co-habiting amphibian species harbor unique skin bacterial communities in wild populations. ISME J 6:588–596. https://doi.org/10.1038/ismej.2011.129
Colombo BM, Scalvenzi T, Benlamara S, Pollet N (2015) Microbiota and mucosal immunity in amphibians. Front Immunol 6:1–15. https://doi.org/10.3389/fimmu.2015.00111
Walke JB, Becker MH, Hughey MC, Swartwout MC, Jensen RV, Belden LK (2015) Most of the dominant members of amphibian skin bacterial communities can be readily cultured. Appl Environ Microbiol 81:6589–6600. https://doi.org/10.1128/AEM.01486-15
Longo AV, Savage AE, Hewson I, Zamudio KR (2015) Seasonal and ontogenetic variation of skin microbial communities and relationships to natural disease dynamics in declining amphibians. R Soc Open Sci 2:140377. https://doi.org/10.1098/rsos.140377
Belden LK, Hughey MC, Rebollar EA, Umile TP, Loftus SC, Burzynski EA, Minbiole KPC, House LL, Jensen RV, Becker MH, Walke JB, Medina D, Ibáñez R, Harris RN (2015) Panamanian frog species host unique skin bacterial communities. Front Microbiol 6:1171. https://doi.org/10.3389/fmicb.2015.01171
Ellison S, Rovito S, Parra-Olea G, Vásquez-Almazán C, Flechas SV, Bi K, Vredenburg VT (2018) The influence of habitat and phylogeny on the skin microbiome of amphibians in Guatemala and Mexico. Microb Ecol 1–11:257–267. https://doi.org/10.1007/s00248-018-1288-8
Costa S, Lopes I, Proença DN, Ribeiro R, Morais PV (2016) Diversity of cutaneous microbiome of Pelophylax perezi populations inhabiting different environments. Sci Total Environ 572:995–1004. https://doi.org/10.1016/j.scitotenv.2016.07.230
Varga JFA, Bui-Marinos MP, Katzenback BA (2019) Frog skin innate immune defences: sensing and surviving pathogens. Front Immunol 10:3128. https://doi.org/10.3389/fimmu.2018.03128
Woodhams DC, Vredenburg VT, Simon MA, Billheimer D, Shakhtour B, Shyr Y, Briggs CJ, Rollins-Smith LA, Harris RN (2007) Symbiotic bacteria contribute to innate immune defenses of the threatened mountain yellow-legged frog, Rana muscosa. Biol Conserv 138:390–398. https://doi.org/10.1016/j.biocon.2007.05.004
Harris RN, Brucker RM, Walke JB, Becker MH, Schwantes CR, Flaherty DC, Lam BA, Woodhams DC, Briggs CJ, Vredenburg VT, Minbiole KPC (2009) Skin microbes on frogs prevent morbidity and mortality caused by a lethal skin fungus. ISME J 3:818–824. https://doi.org/10.1038/ismej.2009.27
Brucker RM, Baylor CM, Walters RL, Lauer A, Harris RN, Minbiole KPC (2008) The identification of 2,4-diacetylphloroglucinol as an antifungal metabolite produced by cutaneous bacteria of the salamander Plethodon cinereus. J Chem Ecol 34:39–43. https://doi.org/10.1007/s10886-007-9352-8
Lauer A, Simon MA, Banning JL, André E, Duncan K, Harris RN (2007) Common cutaneous bacteria from the eastern red-backed salamander can inhibit pathogenic fungi. Copeia 2007:630–640. https://doi.org/10.1643/0045-8511(2007)2007[630:CCBFTE]2.0.CO;2
Lauer A, Simon MA, Banning JL, Lam BA, Harris RN (2008) Diversity of cutaneous bacteria with antifungal activity isolated from female four-toed salamanders. ISME J 2:145–157. https://doi.org/10.1038/ismej.2007.110
Daszak P, Cunningham AA, Hyatt AD (2003) Infectious disease and amphibian population declines. Divers Distrib 9:141–150. https://doi.org/10.1046/j.1472-4642.2003.00016.x
Kilpatrick AM, Briggs CJ, Daszak P (2010) The ecology and impact of chytridiomycosis: an emerging disease of amphibians. Trends Ecol Evol 25:109–118. https://doi.org/10.1016/j.tree.2009.07.011
Harris RN, Lauer A, Simon MA, Banning JL, Alford RA (2009) Addition of antifungal skin bacteria to salamanders ameliorates the effects of chytridiomycosis. Dis Aquat Org 83:11–16. https://doi.org/10.3354/dao02004
Kueneman JG, Woodhams DC, Harris R, Archer HM, Knight R, McKenzie VJ (2016) Probiotic treatment restores protection against lethal fungal infection lost during amphibian captivity. Proc R Soc B Biol Sci 283:20161553. https://doi.org/10.1098/rspb.2016.1553
Berven KA, Gill DE (1983) Interpreting geographic variation in life-history traits. Integr Comp Biol 23:85–97. https://doi.org/10.1093/icb/23.1.85
Berven KA (1990) Factors affecting population fluctuations in larval and adult stages of the wood frog (Rana sylvatica). Ecology 71:1599–1608. https://doi.org/10.2307/1938295
Matutte B, Storey KB, Knoop FC, Conlon JM (2000) Induction of synthesis of an antimicrobial peptide in the skin of the freeze-tolerant frog, Rana sylvatica, in response to environmental stimuli. FEBS Lett 483:135–138. https://doi.org/10.1016/S0014-5793(00)02102-5
Storey KB, Storey JSKM (1992) Natural freeze tolerance in ectothermic vertebrates. Annu Rev Physiol 54:619–637. https://doi.org/10.1146/annurev.physiol.54.1.619
Gahl MK, Longcore JE, Houlahan JE (2012) Varying responses of northeastern north American amphibians to the chytrid pathogen Batrachochytrium dendrobatidis. Conserv Biol 26:135–141. https://doi.org/10.1111/j.1523-1739.2011.01801.x
Forzán MJ, Jones KM, Ariel E, Whittington RJ, Wood J, Markham RJF, Daoust PY (2017) Pathogenesis of frog virus 3 (Ranavirus, Iridoviridae) infection in wood frogs (Rana sylvatica). Vet Pathol 54:531–548. https://doi.org/10.1177/0300985816684929
Walke JB, Becker MH, Loftus SC, House LL, Teotonio TL, Minbiole KPC, Belden LK (2015) Community structure and function of amphibian skin microbes: an experiment with bullfrogs exposed to a chytrid fungus. PLoS One 10:e0139848. https://doi.org/10.1371/journal.pone.0139848
Becker MH, Walke JB, Cikanek S, Savage AE, Mattheus N, Santiago CN, Minbiole KPC, Harris RN, Belden LK, Gratwicke B (2015) Composition of symbiotic bacteria predicts survival in Panamanian golden frogs infected with a lethal fungus. Proc R Soc B Biol Sci 282:20142881–20142881. https://doi.org/10.1098/rspb.2014.2881
Rebollar EA, Gutiérrez-Preciado A, Noecker C, Eng A, Hughey MC, Medina D, Walke JB, Borenstein E, Jensen RV, Belden LK, Harris RN (2018) The skin microbiome of the neotropical frog Craugastor fitzingeri: inferring potential bacterial-host-pathogen interactions from metagenomic data. Front Microbiol 9:466. https://doi.org/10.3389/fmicb.2018.00466
Walters W, Hyde ER, Berg-Lyons D et al (2016) Improved bacterial 16S rRNA gene (V4 and V4-5) and fungal internal transcribed spacer marker gene primers for microbial community surveys. mSystems 1:e00009-15. https://doi.org/10.1128/mSystems.00009-15
Apprill A, Mcnally S, Parsons R, Weber L (2015) Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat Microb Ecol 75:129–137. https://doi.org/10.3354/ame01753
Parada AE, Needham DM, Fuhrman JA (2016) Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol 18:1403–1414. https://doi.org/10.1111/1462-2920.13023
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodríguez AM, Chase J, Cope EK, da Silva R, Diener C, Dorrestein PC, Douglas GM, Durall DM, Duvallet C, Edwardson CF, Ernst M, Estaki M, Fouquier J, Gauglitz JM, Gibbons SM, Gibson DL, Gonzalez A, Gorlick K, Guo J, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley GA, Janssen S, Jarmusch AK, Jiang L, Kaehler BD, Kang KB, Keefe CR, Keim P, Kelley ST, Knights D, Koester I, Kosciolek T, Kreps J, Langille MGI, Lee J, Ley R, Liu YX, Loftfield E, Lozupone C, Maher M, Marotz C, Martin BD, McDonald D, McIver LJ, Melnik AV, Metcalf JL, Morgan SC, Morton JT, Naimey AT, Navas-Molina JA, Nothias LF, Orchanian SB, Pearson T, Peoples SL, Petras D, Preuss ML, Pruesse E, Rasmussen LB, Rivers A, Robeson II MS, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song SJ, Spear JR, Swafford AD, Thompson LR, Torres PJ, Trinh P, Tripathi A, Turnbaugh PJ, Ul-Hasan S, van der Hooft JJJ, Vargas F, Vázquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan Y, Wang M, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang Y, Zhu Q, Knight R, Caporaso JG (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Yilmaz P, Parfrey LW, Yarza P, Gerken J, Pruesse E, Quast C, Schweer T, Peplies J, Ludwig W, Glöckner FO (2014) The SILVA and “all-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res 42:D643–D648. https://doi.org/10.1093/nar/gkt1209
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066. https://doi.org/10.1093/nar/gkf436
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Medina D, Hughey MC, Becker MH, Walke JB, Umile TP, Burzynski EA, Iannetta A, Minbiole KPC, Belden LK (2017) Variation in metabolite profiles of amphibian skin bacterial communities across elevations in the Neotropics. Microb Ecol 74:227–238. https://doi.org/10.1007/s00248-017-0933-y
Shade A, Handelsman J (2012) Beyond the Venn diagram: the hunt for a core microbiome. Environ Microbiol 14:4–12. https://doi.org/10.1111/j.1462-2920.2011.02585.x
Muletz-Wolz CR, Almario JG, Barnett SE, DiRenzo GV, Martel A, Pasmans F, Zamudio KR, Toledo LF, Lips KR (2017) Inhibition of fungal pathogens across genotypes and temperatures by amphibian skin bacteria. Front Microbiol 8:1551. https://doi.org/10.3389/fmicb.2017.01551
Ellison S, Knapp RA, Sparagon W, Swei A, Vredenburg VT (2019) Reduced skin bacterial diversity correlates with increased pathogen infection intensity in an endangered amphibian host. Mol Ecol 28:127–140. https://doi.org/10.1111/mec.14964
Walke JB, Becker MH, Loftus SC, House LL, Cormier G, Jensen RV, Belden LK (2014) Amphibian skin may select for rare environmental microbes. ISME J 8:2207–2217. https://doi.org/10.1038/ismej.2014.77
Berven KA, Grudzien TA (1990) Dispersal in the wood frog (Rana sylvatica): implications for genetic population structure. Evolution (N Y) 44:2047–2056. https://doi.org/10.2307/2409614
Prado-Irwin SR, Bird AK, Zink AG, Vredenburg VT (2017) Intraspecific variation in the skin-associated microbiome of a terrestrial salamander. Microb Ecol 74:745–756. https://doi.org/10.1007/s00248-017-0986-y
Muletz-Wolz CR, Yarwood SA, Campbell Grant EH et al (2018) Effects of host species and environment on the skin microbiome of Plethodontid salamanders. J Anim Ecol 87:341–353. https://doi.org/10.1111/1365-2656.12726
Bletz MC, Archer H, Harris RN, McKenzie VJ, Rabemananjara FCE, Rakotoarison A, Vences M (2017) Host ecology rather than host phylogeny drives amphibian skin microbial community structure in the biodiversity hotspot of Madagascar. Front Microbiol 8:1530. https://doi.org/10.3389/fmicb.2017.01530
Albecker MA, Belden LK, McCoy MW (2018) Comparative analysis of anuran amphibian skin microbiomes across inland and coastal wetlands. Microb Ecol 78:348–360. https://doi.org/10.1007/s00248-018-1295-9
Krynak KL, Burke DJ, Benard MF (2016) Landscape and water characteristics correlate with immune defense traits across Blanchard’s cricket frog (Acris blanchardi) populations. Biol Conserv 193:153–167. https://doi.org/10.1016/j.biocon.2015.11.01948
Fierer N, Hamady M, Lauber CL, Knight R (2008) The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci 105:17994–17999. https://doi.org/10.1073/pnas.0807920105
Saag P, Tilgar V, Mänd R, Kilgas P, Mägi M (2011) Plumage bacterial assemblages in a breeding wild passerine: relationships with ecological factors and body condition. Microb Ecol 61:740–749. https://doi.org/10.1007/s00248-010-9789-0
Campbell LJ, Garner TWJ, Hopkins K, Griffiths AGF, Harrison XA (2019) Outbreaks of an emerging viral disease covary with differences in the composition of the skin microbiome of a wild United Kingdom amphibian. Front Microbiol 10:1245. https://doi.org/10.3389/fmicb.2019.01245
Chase AB, Karaoz U, Brodie EL, Gomez-Lunar Z, Martiny AC, Martiny JBH (2017) Microdiversity of an abundant terrestrial bacterium encompasses extensive variation in ecologically relevant traits. MBio 8. https://doi.org/10.1128/mBio.01809-17
Kielak AM, Barreto CC, Kowalchuk GA, van Veen JA, Kuramae EE (2016) The ecology of Acidobacteria: moving beyond genes and genomes. Front Microbiol 7:744. https://doi.org/10.3389/fmicb.2016.00744
Grönroos M, Parajuli A, Laitinen OH, Roslund MI, Vari HK, Hyöty H, Puhakka R, Sinkkonen A (2019) Short-term direct contact with soil and plant materials leads to an immediate increase in diversity of skin microbiota. MicrobiologyOpen 8:e00645. https://doi.org/10.1002/mbo3.645
Zaheer R, Noyes N, Ortega Polo R, Cook SR, Marinier E, van Domselaar G, Belk KE, Morley PS, McAllister TA (2018) Impact of sequencing depth on the characterization of the microbiome and resistome. Sci Rep 8:5890. https://doi.org/10.1038/s41598-018-24280-8
McMurdie PJ, Holmes S (2014) Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput Biol 10:e1003531. https://doi.org/10.1371/journal.pcbi.1003531
Longo AV, Zamudio KR (2017) Temperature variation, bacterial diversity and fungal infection dynamics in the amphibian skin. Mol Ecol 26:4787–4797. https://doi.org/10.1111/mec.14220
Muletz-Wolz CR, Fleischer RC, Lips KR (2019) Fungal disease and temperature alter skin microbiome structure in an experimental salamander system. Mol Ecol 28:mec.15122. https://doi.org/10.1111/mec.15122
Zhou J, Deng Y, Shen L, Wen C, Yan Q, Ning D, Qin Y, Xue K, Wu L, He Z, Voordeckers JW, Nostrand JDV, Buzzard V, Michaletz ST, Enquist BJ, Weiser MD, Kaspari M, Waide R, Yang Y, Brown JH (2016) Temperature mediates continental-scale diversity of microbes in forest soils. Nat Commun 7:12083. https://doi.org/10.1038/ncomms12083
Maniero GD, Carey C (1997) Changes in selected aspects of immune function in the leopard frog, Rana pipiens, associated with exposure to cold. J Comp Physiol B 167:256–263. https://doi.org/10.1109/LARS
Katzenback BA, Holden HA, Falardeau J, Childers C, Hadj-Moussa H, Avis TJ, Storey KB (2014) Regulation of the Rana sylvatica brevinin-1SY antimicrobial peptide during development and in dorsal and ventral skin in response to freezing, anoxia and dehydration. J Exp Biol 217:1392–1401. https://doi.org/10.1242/jeb.092288
Acknowledgments
The authors thank Nicole Wang for the generous contribution of a trained taxonomic classifier for 16S rRNA gene sequences and Maxwell P. Bui-Marinos, Joseph F.A. Varga, and Nathanael B. J. Harper for their technical assistance in collecting frog skin swabs.
Funding
This study was funded by a Natural Sciences and Engineering Research Council of Canada Discovery Grant (NSERC DG) to BAK (Grant # RGPIN-2017-04218), a Tier II Canada Research Chair to LAH, and salary support to AJD through a Natural Sciences and Engineering Research Council of Canada Undergraduate Summer Research Assistantship (NSERC USRA), the University of Waterloo Undergraduate Research Internship (URI) funding initiative, as well as a Graduate Research Studentship, Science Graduate Award, and UW Graduate Scholarship awarded by the University of Waterloo, Department of Biology.
Author information
Authors and Affiliations
Contributions
AJD, LAH, and BAK conceived the study; AJD and BAK performed field sampling; AJD performed the experiments and analyzed the data; AJD, LAH, and BAK wrote and critically revised the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. All procedures performed in studies involving animals were in accordance with the ethical standards of the institution at which the studies were conducted (University of Waterloo Animal Care Committee and the Canadian Council on Animal Care, Animal Utilization Projects #30008 and #40721), and animals were captured under the Ontario Ministry of Natural Resources and Forestry Wildlife Scientific Collectors Authorization Permits (#1088586 and #1092603 issued to Dr. B.A. Katzenback). This article does not contain any studies with human participants performed by any of the authors.
Rights and permissions
About this article
Cite this article
Douglas, A.J., Hug, L.A. & Katzenback, B.A. Composition of the North American Wood Frog (Rana sylvatica) Bacterial Skin Microbiome and Seasonal Variation in Community Structure. Microb Ecol 81, 78–92 (2021). https://doi.org/10.1007/s00248-020-01550-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-020-01550-5