Abstract
Animals host a wide diversity of symbiotic microorganisms that contribute important functions to host health, and our knowledge of what drives variation in the composition of these complex communities continues to grow. Microbiome studies at larger spatial scales present opportunities to evaluate the contribution of large-scale factors to variation in the microbiome. We conducted a large-scale field study to assess variation in the bacterial symbiont communities on adult frog skin (Pseudacris crucifer), characterized using 16S rRNA gene amplicon sequencing. We found that skin bacterial communities on frogs were less diverse than, and structurally distinct from, the surrounding habitat. Frog skin was typically dominated by one of two bacterial OTUs: at western sites, a Proteobacteria dominated the community, whereas eastern sites were dominated by an Actinobacteria. Using a metacommunity framework, we then sought to identify factors explaining small- and large-scale variation in community structure—that is, among hosts within a pond, and among ponds spanning the study transect. We focused on the presence of a fungal skin pathogen, Batrachochytrium dendrobatidis (Bd) as one potential driver of variation. We found no direct link between skin bacterial community structure and Bd infection status of individual frog hosts. Differences in pond-level community structure, however, were explained by Bd infection prevalence. Importantly, Bd infection prevalence itself was correlated with numerous other environmental factors; thus, skin bacterial diversity may be influenced by a complex suite of extrinsic factors. Our findings indicate that large-scale factors and processes merit consideration when seeking to understand microbiome diversity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Assembly of the host-associated microbiome is complex and influenced by a multitude of extrinsic and intrinsic factors. Broadly, the factors influencing host-associated microbiome diversity may be divided into two categories: local factors related to the host itself and regional factors operating at spatial scales beyond the host [1]. Host-associated factors can be measured on a per-host basis, and are typically regulated by host genetics and immunity (e.g., [2, 3]). Many of the factors that influence microbiome diversity, however, are not actually quantified on a per-host basis. Rather, these factors reflect a larger spatial or temporal scale experienced by a group of hosts, such as the biotic and abiotic conditions of the habitat in which hosts reside. These also include purely spatial factors, such as latitude, that can indicate a role for stochastic processes, dispersal, or historical factors on microbiome diversity [4,5,6]. As microbiome studies continue to increase in scale, so too does our ability to assess the contribution of these larger-scale factors and processes to microbiome community structure.
One approach to better understanding the contribution of large-scale factors to microbial symbiont diversity is to consider these systems as metacommunities [7,8,9]: collections of local communities interconnected by dispersal [10]. Key to the metacommunity concept is the idea that diversity is influenced by factors and processes that operate at different spatial and temporal scales, ranging from species interactions within a local community to dispersal across the landscape [10, 11]. Because metacommunity theory focuses on discerning the relative contributions of large-scale processes versus the local environment, it can lead to key insights about the factors driving local community structure. The theory has traditionally been applied to communities of free-living organisms, including microbes (e.g., [5]). However, it is proving amenable to the study of host-symbiont systems as well, for example, in determining the contribution of neutral processes to microbial symbiont community assembly [12, 13].
Amphibian skin harbors a diverse bacterial community that likely plays a role in protecting its hosts from pathogen infection [14]. Currently, there is great interest in understanding the factors influencing the diversity of the amphibian skin microbiome, in part because effective implementation of bioaugmentation as a conservation strategy may depend on it [15, 16]. Numerous studies have begun to investigate potential drivers of variation in the amphibian skin microbiome. These studies indicate that both host-associated factors, such as host genetics [2] or life stage [17], and larger-scale spatial and temporal factors (e.g., land use; [18, 19]; season; [17]; elevation [20, 21]) likely play an important role in shaping microbiome diversity.
Changes in skin microbiome structure have also been linked to the presence of the chytrid fungus Batrachochytrium dendrobatidis (Bd) [22,23,24,25,26,27,28,29], a pathogen that poses a critical threat to many amphibian species worldwide [30, 31]. Bd attacks amphibian skin, sometimes with lethal consequences [32]. Because symbiotic skin bacteria can play a role in protecting their hosts from the pathogen (e.g., [33]), it is not altogether surprising that Bd would be a strong selective agent on the structure of the amphibian skin microbiome. Bd infection drives temporal changes in the skin bacterial communities of the highly Bd-susceptible frog, Rana sierrae, that are apparent at the level of both the host and the population [22, 26]. However, in the more Bd-resistant species, Craugastor fitzingeri, historical effects of Bd exposure manifest at the population level in the absence of host-level responses to infection [23, 25]. Specifically, the skin bacterial communities of populations of C. fitzingeri in sites where Bd is endemic show reduced diversity overall, relative to Bd-naïve sites [25], even though Bd-infected frogs within endemic sites do not necessarily have less diverse bacterial communities compared to uninfected ones [23]. This indicates that it is possible for the influence of this pathogen to be uncoupled at the host and population levels, and underscores the need for approaches focusing on the effects of larger-scale factors on symbiont community structure.
In a previous study [34], we conducted a survey of spring peepers (Pseudacris crucifer), and other amphibian species, across a longitudinal transect in Virginia, U.S.A., and quantified variation in Bd infection intensity and prevalence in relation to pond water temperature. Here, we characterize the diversity of the skin bacterial microbiota of a subset of those spring peepers from nine ponds along the same transect. Using a metacommunity framework, we assessed the contribution of Bd infection parameters to differences in skin bacterial diversity at two spatial scales, focusing on host infection status and intensity, and pond prevalence. Furthermore, at three ponds spanning the west-east gradient of our study transect, we determined the composition of the free-living bacterial assemblages in the pond habitat to evaluate the extent to which skin bacterial diversity reflects the diversity of environmental bacterial source pools.
Materials and Methods
Sample Collection and Processing
We sampled the skin bacterial communities of 116 spring peepers from nine ponds on a west-east gradient of 342 km across Virginia, U.S.A., during March–April 2012 (N = 7–19 individuals per pond; subset of individuals from the Bd survey in Hughey et al. [34]). Spring peepers are abundant and widespread throughout eastern North America. In early spring, they congregate at ponds to breed. This species can become infected with Bd, and both the probability of being infected with Bd and individual infection loads tend to be greater at warmer ponds [34]. However, as far as is known, spring peepers do not develop chytridiomycosis, the disease caused by Bd [35].
Bacterial community diversity and Bd infection were assessed using non-lethal skin swabs [23, 36, 37]. Frogs were caught by hand between 2000 and 2300 h, using new nitrile gloves for each one, placed in a sterile Whirl-Pak bag, and swabbed within ~ 1 h of capture. Before swabbing, we rinsed each frog with 50 ml of sterile, deionized water. To standardize, we swabbed the ventral surface 20 times, each thigh 5 times, and each hind foot 5 times for a total of 40 strokes per frog using sterile rayon swabs (MW113, Medical Wire Equipment & Co. Ltd., Corsham, UK). After swabbing, all animals were released at the site of capture.
To quantify the relationship between the diversity of free-living environmental bacteria to that of the host-associated bacterial communities, we sampled the bacterial community of the habitat in which the frogs were breeding at three sites across our study transect (Sylvatica Pond, Claytor Pond, and James River Wetlands). We passed swabs through two microhabitats—pond water and pond substrate—for approximately 5 s/swab (N = 3 replicates/habitat type; as in [38]). All swabs (frog and habitat) were placed in sterile 1.5-ml microcentrifuge tubes on ice in the field and then transferred to − 20 or − 80 °C for laboratory storage prior to DNA extraction.
Pond water temperature was measured in three haphazardly selected locations near calling frogs using a YSI meter (model 63). The average of these three measurements represented pond environmental conditions. Additionally, we recorded the latitude, longitude, and elevation of each site using a hand-held GPS device. To determine each individual’s Bd infection status (infected or not infected) and infection intensity (measured as the number of zoospore equivalents), we used a quantitative, real-time qPCR assay developed by Boyle et al. [39], as described in Hughey et al. [34]. In brief, we included a negative control and a series of five dilution standards, using Bd strain JEL404 (isolated from Rana catesbeiana in Maine, USA), in each run. Standards ranged in concentration from 0.1 to 1000 zoospore equivalents. Bd prevalence for each pond was defined as the proportion of Bd-positive frogs.
To characterize the bacterial communities on amphibian skin and in the habitat, we used 16S rRNA gene amplicon sequencing [23]. DNA was extracted from swabs using Qiagen’s DNeasy Blood & Tissue Kit (Qiagen, Inc., Valencia, CA, USA). We followed the manufacturer’s Quick-Start protocol, except we incubated each sample with 180 μl lysis buffer solution (20 mg lysozyme/1 ml lysis buffer) at 37 °C for 1 h, and we added 25 μl proteinase K in addition to 200 μl buffer AL and incubated at 70 °C for 30 min. Our final elution volume was 100 μl.
We amplified the V4 region of the 16S rRNA gene [40] to maintain consistency with previous work on the amphibian skin microbiome, although sampling this region may miss certain taxa (e.g., Propionibacterium; [41]). PCRs were run in triplicate for each sample, which were combined after amplification. Each 25 μl reaction contained the following: 11.5 μl PCR water, 10 μl 5 PRIME Hot Master Mix, 0.5 μl 515f forward primer, 0.5 μl 806r reverse primer including a 12-base barcode sequence, and 2.5 μl genomic DNA. Thermocycler conditions consisted of a denaturation step of 94 °C for 3 min, followed by 34 cycles at 94 °C for 45 s, 50 °C for 60 s, and 72 °C for 90 s, and a final extension at 72 °C for 10 min.
Amplified DNA was quantitated using a Qubit® 2.0 Flourometer and dsDNA HS assay kit according the manufacturer’s guidelines (Life Technologies, Carlsbad, CA, USA). Samples were pooled by combining 180 ng of each amplicon into a single tube. The pooled sample was cleaned using the QIAquick PCR purification kit according to the manufacturer’s instructions (Qiagen, Inc., Valencia, CA, USA). Fifty microliters of the pooled sample were sent to the Molecular Biology Core Facilities of the Dana Farber Cancer Institute at Harvard University (Cambridge, MA, USA) for sequencing on an Illumina MiSeq instrument using a 250 bp paired-end strategy. To compensate for the low base diversity of the amplicon pool, the sample was run with a 10% PhiX control. Version 1.18.42 of the MiSeq Real-Time Analysis software (Illumina) was used to perform base calling and quality scoring. Sequence data is deposited in the NCBI database (Bioproject accession number PRJNA339746).
We produced a single OTU table, including all samples (N = 116 frog and 18 habitat samples). Forward and reverse reads from the raw Illumina files were joined using Fastq-join v. 0.1 [42]. Joined sequences were de-multiplexed and filtered using the Quantitative Insights Into Microbial Ecology pipeline (MacQIIME, v. 1.8.0; [43]). We used the default settings with the following exceptions: we allowed for no errors in the barcode during demultiplexing; we set the minimum fraction of consecutive high quality base calls required to include a read at half the total read length; and we set the maximum number of consecutive low-quality base calls allowed before truncating a read at 10. Using Geneious v 8.0.4, any remaining PhiX sequences were filtered out, and all sequences between 250 and 255 bp in length were extracted. In QIIME, sequences were assigned to operational taxonomic units (OTUs) using the UCLUST method at 97% sequence similarity [44]. The most abundant sequence from each cluster was used to represent each OTU. Representative sequences were aligned to the Greengenes v. 13.5 reference database [45] using PyNAST [46], and assigned taxonomy using the RDP classifier [47]. OTUs assigned as chloroplast, mitochondria, and Archaea, as well as OTUs with fewer than 0.01% of the total number of reads (following the recommendations in [48]), were removed from the dataset. OTU abundances were rarefied to a sequencing depth of 5000 sequences/sample (minimum sequencing depth = 5378 sequences). The final dataset consisted of 134 samples and 663 OTUs.
Influence of Bd Infection Parameters on Microbiome Diversity
Statistical analyses were run in R v. 3.1.2 [49] using the package vegan [50], unless specified otherwise. To visualize variation in bacterial community diversity, we used non-metric multidimensional scaling (NMDS) on a community dissimilarity matrix based on Euclidean distances of Hellinger-transformed relative abundance data. We tested for general separation of samples based on pond of origin using permutational analysis of variance (PERMANOVA; [51]). We repeated this analysis using Jaccard, Bray-Curtis, and weighted and unweighted Unifrac dissimilarity indices and obtained similar results (not shown).
Potential explanatory variables represented one of two spatial scales: some factors corresponded directly to individual hosts (e.g., Bd infection intensity), whereas others corresponded to a population of hosts at a given pond (e.g., Bd infection prevalence). To understand if these factors were associated with bacterial community structure, we drew upon the concept of metacommunities [10] to interpret diversity calculated for different scales of ecological organization [52]. Thus, we defined the “local community” at both the host scale and the pond scale, and we conducted analyses on responses for each of these scales.
For host-level analyses, we used the skin community data associated with each frog. We determined if Bd infection status or Bd infection intensity explained variation in community richness (OTU richness) or evenness (Shannon index and Simpson index calculated based on OTU relative abundances) using generalized linear mixed models. We included “pond” as a random effect in each model to control for among-pond variation in alpha diversity. OTU richness and Shannon Index were log-transformed to better meet assumptions of normality. The Simpson index was fit to beta regression models (package glmmADMB; [53, 54]), which were developed to accommodate data like the Simpson index that are limited to a range of 0 to 1. Additionally, we determined if Bd infection status and/or intensity explained variation in host community composition using redundancy analysis (RDA; [55]). Redundancy analysis is a form of multivariate linear regression that can be used with community datasets to assess how much of the variation in community composition can be explained by one or more explanatory variables [56, 57]. Because ponds varied in composition, and accounting for among-pond differences using random effects is technically not possible for permutation-based approaches such as RDA in the vegan package, we analyzed each pond separately. Bacterial community data were Hellinger-transformed prior to RDA because the dataset contained a high number of zeros that otherwise would have been interpreted in the analysis as indicating similarity between sites [58].
For pond-level analyses, we combined the data for all frogs within a pond. Pond alpha diversity was calculated as the mean of each diversity measure (OTU richness, Shannon index, Simpson index) across all individuals at a given pond. For pond community composition, we used a bootstrapping approach to obtain an average frog skin community for each pond as follows: We generated relative abundance distributions for each OTU using the data for all frogs sampled at each pond, resampling 999 times with replacement. We selected the median value to represent the relative abundance of each OTU at each pond. OTUs whose median values were equal to zero at all sites were removed from the dataset prior to analysis (N = 50). R code for this approach is available as Supplementary Material. We tested for a relationship between Bd prevalence and each of the three measures of alpha diversity (mean OTU richness, Shannon index, and Simpson index) using linear models for OTU richness and Shannon index, and beta regression models for Simpson index (similar to above but with no random effects). Additionally, we determined if Bd prevalence explained variation in pond community composition using the boot-strapped community data, acknowledging that Bd prevalence was highly correlated with several other variables that also varied across the study transect: pond water temperature, longitude, and elevation (Pearson’s correlations: Bd prevalence and water temperature r = 0.77, P = 0.01; and longitude r = 0.84, P = 0.005; and elevation r = − 0.72, P = 0.03). We chose Bd prevalence as the representative variable to maintain the focus on disease and its potential relationship to the microbiome. As with the host-level analyses, pond-level community data were Hellinger-transformed prior to RDA.
Because we detected significant relationships at the pond level, we then tested for variation in the relative abundance of specific OTUs in relation to Bd prevalence across ponds. We focused on dominant OTUs, here defined as OTUs representing, on average, 1% or more of the relative abundance of the entire metacommunity ([59, 60]; N = 12 OTUs). For each OTU, we tested for a relationship between pond means for relative abundance and Bd prevalence using beta regression. P values were adjusted for false discovery rate using the Benjamini Hochberg correction.
Frog-Environment Comparisons
To determine if the composition of the skin bacterial communities on frogs was similar to that of the habitat (which could indicate high rates of dispersal between hosts and the local environment), we used PERMANOVA with Hellinger-transformed relative abundance data that were converted to dissimilarity matrices based on Euclidean distances. Analyses were limited to the three sites where both sample types were collected: Sylvatica Pond, Claytor Pond, and James River Wetlands. We tested if sample source (i.e., frog, pond water, or pond substrate) was a significant predictor of bacterial community composition, restricting the permutations so that observations were permuted within pond. We repeated these analyses using Jaccard, Bray-Curtis, and weighted and unweighted Unifrac dissimilarity indices and obtained similar results (not shown). To further evaluate how closely the structure of the frog skin bacterial community matched that of the environment, we compared the mean relative abundance of OTUs shared across sample types (frog to water or frog to substrate), separately for each pond where both sample types were collected.
Results
Spring peepers harbored diverse skin bacterial communities. In total, we identified 660 unique operational taxonomic units (OTUs) from 13 phyla on spring peeper skin (Table 1). The phyla Proteobacteria and Bacteroidetes accounted for the vast majority of OTU diversity, whereas the phyla Proteobacteria and Actinobacteria accounted for the vast majority of OTU relative abundance (Table 1). The number of OTUs on individual frogs ranged from 56 to 359 (mean ± sd = 181 ± 80 OTUs), and mean values for Shannon and Simpson indices were 2.83 ± 1.01 (range 0.81–4.89) and 0.80 ± 0.16 (range 0.27–0.98), respectively.
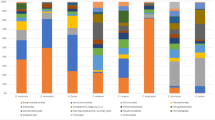
Twelve OTUs dominated the spring peeper metacommunity (Table 2). However, the relative abundance of these OTUs differed among individuals, contributing to among-pond variation in community structure (PERMANOVA pseudoF = 9.08, R2 = 0.40, P = 0.001; Fig. 1a–c). Most notably, we observed a marked shift in the structure of the skin bacterial communities on frogs between western and eastern sites (Fig. 1d). The skin bacterial communities of individuals from Claytor, Food Lion, and James River in the east were particularly enriched in a single OTU classified as a Proteobacteria in the family Alcaligenaceae (X820379, further classified to Bordetella at the genus level by a BLAST search; Fig. 1d). By contrast, frogs from Kentland, Kroger, Pandapas, and Glen Alton ponds in the west had a much higher relative abundance of Actinobacteria, especially an OTU in the genus Sanguibacter (X4378239; Fig. 1d). Frogs from Heritage Park and Sylvatica ponds in the west were not enriched in Sanguibacter (OTU X4378239); however, they also lacked the higher relative abundances of the Proteobacteria (OTU X820379) characteristic of eastern ponds (Fig. 1d).
Variation in the structure of the bacterial communities on the skin of spring peepers (Pseudacris crucifer) from nine sites across the state of Virginia. a–c Ordination was constructed using non-metric multidimensional scaling based on Euclidean distances of Hellinger-transformed relative abundance data (stress = 0.16). Data are colored according to pond of origin. Points represent the community of a single individual. Polygons enclose all points for a given pond. Polygons for ponds from b western and c eastern sites are presented separately for comparison, but the underlying data are exactly the same in all three plots. d Pond-level variation in the spring peeper (Pseudacris crucifer) skin microbiome. Ponds are ordered longitudinally from west to east along the X axis. Colored bars represent the boot-strapped mean relative abundance of the ten most abundant families on spring peeper skin. Hatch marked regions (within their respective families) indicate the mean relative abundance of the two most relatively abundant OTUs (X4378239 and X820379) in the spring peeper microbiome. Numbers listed in bold above each bar indicate the proportion of individuals that were infected with Bd (data from [34])
Thirty-one out of 116 frogs were infected with Bd. Infection intensity of these frogs ranged from 1 to 421 zoospore equivalents (mean ± sd = 69 ± 119; [34]). Bd infection prevalence ranged from 0 to 86% across ponds (Fig. 1d; [34]). Ponds in western Virginia generally had low Bd prevalence, whereas ponds to the east had higher prevalence. At the individual frog level, neither Bd infection status (infected/not infected) nor infection intensity were important predictors of skin bacterial diversity (GLMs, Richness, status: χ2 = 2.6, P = 0.1, intensity: χ2 = 0.4, P = 0.5; Shannon, status: χ2 = 1.1, P = 0.3, intensity: χ2 = 0.2, P = 0.6; Simpson, status: χ2 = 0.01, P = 0.9, intensity: χ2 = 0.3, P = 0.6) nor community structure (RDA, status: P > 0.1 for all ponds; intensity: P > 0.1 for all ponds). At the pond level, however, Bd infection prevalence explained variation in the Simpson index (i.e., dominance in the community; χ2 = 7.3, P = 0.007; Fig. 2a), although not in OTU richness (F1,7 = 0.05, P = 0.8) or evenness (Shannon index: F1,7 = 0.83, P = 0.4). In addition, prevalence of Bd infection explained variation in pond-level community structure (RDA, R2adj = 29%, F1,7 = 4.21, P = 0.02). In particular, skin bacterial communities from eastern ponds showed an increase in the dominance of certain taxa—and thus reduced diversity—relative to those from western ponds (Figs. 1 and 2). Eight out of the 12 dominant OTUs exhibited changes in relative abundance in relation to Bd prevalence (Table 2; Fig. 2b–m). One OTU, the Proteobacteria X820379, increased dramatically in abundance, whereas most of the other OTUs decreased in abundance as Bd prevalence increased (Fig. 2b–m).
Changes in alpha diversity and OTU relative abundance in relation to Bd prevalence across ponds. All data are pond means. a Changes in Simpson index. b–m Changes in the relative abundance of the 12 most dominant OTUs, defined as OTUs representing at least 1% of the mean relative abundance of the entire metacommunity. OTUs are presented in the same order as in Table 2. Asterisks (*) indicate OTUs that significantly increased or decreased in relative abundance with increasing Bd prevalence (see Table 2 for results of statistical tests)
The bacterial communities associated with the skin of spring peepers were distinct from that of their habitat—including both water and substrate samples—and water and substrate samples were also distinct from one another (adonis, overall model: pseudoF = 6.68, R2 = 0.22, P = 0.001; frog–water: pseudoF = 8.12, R2 = 0.17, P = 0.001; frog–substrate: pseudoF = 7.25, R2 = 0.15, P = 0.001; water–substrate: pseudoF = 1.90, R2 = 0.11, P = 0.001; Fig. 3a). However, most OTUs that were found on spring peepers were also found in environmental samples (Fig. 3b). At the three sites where we sampled both spring peepers and habitat, 545 (84%) of the OTUs were shared across frog and habitat samples (either water or substrate), 33 OTUs (5%) were present in habitat samples but were not recovered from spring peepers, and 72 OTUs (11%) were associated only with spring peepers. The mean relative abundance of most OTUs that were shared by frogs and the surrounding water or substrate was low in each sample type (i.e., mean relative abundance on frogs and in water or substrate, both < 0.05; Fig. 3c–h). For those OTUs that were more abundant, they were usually abundant in habitat samples (water or substrate) or on frogs, but not both. For example, X820379 (Proteobacteria: Alcaligenaceae) displayed low relative abundance in all habitat samples (mean relative abundance ≤ 0.00007 for habitat samples at all sites), even though it was the most abundant OTU (on average, 0.33 ± 0.17) on frogs at all three sites. Just two OTUs, at two different ponds, demonstrated a near 1:1 match in mean relative abundance between the frogs and the surrounding habitat (specifically, the substrate; Fig. 3d): X4378239 (Actinobacteria: Sanguibacter) at Sylvatica Pond and X3219861 (Proteobacteria: Crenothrix) at James River (Fig. 3h).
a Diversity of the bacterial communities on the skin of spring peepers (Pseudacris crucifer) as compared to the water and substrate in the surrounding pond environment. Only data for the three sites where all three sample types were collected are shown (N = 3 swabs of water and substrate from all ponds; N = 10, 14, and 9 frogs for Sylvatica, Claytor, and James River, respectively). Ordination was constructed using non-metric multidimensional scaling based on Euclidean distances of Hellinger-transformed relative abundance data (stress = 0.16). Points represent the community of a single sample. Sample type is denoted by color and pond of origin by symbol type. b Venn diagram showing overlap of OTUs associated with the skin bacterial communities of spring peepers and the surrounding pond water and pond substrate. c–h Relative abundances of 545 OTUs that were shared between spring peeper skin and the surrounding habitat, separated by habitat type (substrate or water). Data are the mean relative abundance of each OTU in each sample type. The dashed line is a 1:1 line; OTUs that fall along this line display similar relative abundances on frogs and in the surrounding habitat
Discussion
As evidence of spatial variation in the diversity of host-associated microbial communities continues to grow—in amphibian systems and others—this underscores the need for understanding causes of microbiome turnover at spatial scales beyond that of individual hosts. Here, we investigate how both host- and pond-scale Bd infection parameters potentially influence the diversity of the bacterial communities inhabiting spring peeper skin. Because Bd invades the skin where the bacterial symbionts live, we predicted that the presence of Bd would be associated with variation in bacterial community structure among hosts. We found no relationship between Bd infection and individual hosts’ microbiomes. However, we did find that the prevalence of Bd infection across ponds explained regional variation in the spring peeper skin microbiome. This observation of emergent patterns of microbiome diversity at a scale beyond the level of the host is compelling, and we hope it inspires thinking about novel ways in which pathogens may interact with host populations.
One of the most striking patterns we observed was between western and eastern ponds. At four out of six ponds that we sampled in the west, many individuals harbored high relative abundances of OTUs that were essentially absent from eastern ponds. In the east, the bacterial communities shifted to favor high relative abundances of a specific Proteobacteria in the family Alcaligenaceae, likely in the genus Bordetella. Several other studies have documented changes in amphibian microbiome diversity and/or composition in relation to Bd under both natural and experimental settings [22,23,24,25,26,27,28,29]. From these studies, which encompass a phylogenetically diverse and geographically widespread set of amphibian hosts, it appears that responses of alpha diversity to Bd may be quite variable. Bd infection has been associated with increases [28], reductions [25, 29], or little change in alpha diversity [22, 24, 26]. In all of these studies, however, compositional changes in the microbiota resulting from Bd exposure have been observed. Responses to the pathogen are consistent across populations within a host species (this study; [22, 25, 29]); however, the identity of the bacterial taxa that respond most strongly to the presence of Bd seems to vary considerably among amphibian species. Ultimately, these patterns suggest that interactions between Bd and the microbiome are host species specific, and any attempts to develop bioaugmentation strategies to protect vulnerable species of amphibians will likely need to be tailored to specific hosts.
Another important consideration for assessing results of Bd-driven correlations in field surveys is that many other environmental factors often vary along essentially the same gradient as Bd prevalence. Thus, Bd prevalence may be representative of some other factor that is actually driving the large-scale patterns we observed. For example, the three easternmost ponds along our longitudinal gradient were substantially lower in elevation than the remainder of our study sites. These sites generally experience milder winters and an earlier spring, and so, at the time of our sampling, were warmer than western ponds. Furthermore, spring peepers at eastern sites were nearing the end of their breeding season (J.R. Vonesh and G.K. Eaton, pers. comm.), whereas those in western sites were just beginning. Because breeding activity is energetically demanding, the general condition of hosts from western and eastern sites at the time of sampling were likely very different.
The large-scale spatial turnover observed in this study could also indicate that historical or biogeographical processes shape spring peeper bacterial microbiome diversity. Spring peepers are genetically diverse, clustering into four distinct lineages across their range, two of which occur on either side of the Appalachian Mountains [61]. Furthermore, populations along the coast of Virginia represent a distinct subclade of the lineage east of the Appalachians [61]. Given that our study transect began in the Appalachians in the west and continued eastward down onto the Coastal Plain, it is possible that we sampled distinct spring peeper lineages with uniquely adapted microbiomes, although the findings of Griffiths et al. [2] indicate that factors other than host population genetic structure may play a stronger role in influencing regional microbiome variation.
Turnover in skin bacterial communities along a broad longitudinal gradient can also indicate that dispersal and transmission dynamics influence microbiome diversity. However, the existence of a common set of dominant OTUs suggests a relatively consistent regional source pool throughout the study area. Furthermore, we identified only ten bacterial OTUs on frogs that were specific to a single pond, indicating that most of the bacteria present on spring peeper skin are likely not dispersal-limited at the regional scale. However, it is possible that dispersal contributes to changes in relative abundance (more so than turnover in composition) at the spatial scales we examined. Metacommunity theory has enormous potential to enhance our understanding of the role that dispersal plays in generating patterns of among-host and among-pond diversity such as observed in this study. However, use of a metacommunity framework has thus far been underutilized for host-symbiont systems despite being emphasized in several recent reviews [1, 7,8,9, 62]. Future research could use a combination of modeling and experimental approaches to evaluate the relative roles of dispersal versus host characteristics in generating the patterns in symbiont diversity observed in this study (e.g., [63]).
Given the large number of OTUs that were shared between frog and habitat samples, it is likely that many members of the spring peeper skin microbiota are sourced from the environment [25, 38, 64]. Yet based on our comparison of free-living and host-associated communities across sites, we do not think that among-pond variation simply reflects variation in the bacterial pool of potential colonists. Were that the case, we would have expected to see greater clustering of frog and habitat samples by pond of origin; instead, frogs from all three ponds clustered distinctly from water and substrate samples. Furthermore, we found very little evidence of matching, in terms of mean relative abundance, between OTUs on frogs and in the habitat. Indeed, one of the most abundant members of the skin bacterial community, the Proteobacteria (OTU X820379), was nearly absent from our environmental samples. Its low relative abundance in the environment could indicate mass effects from the frog skin habitat. Alternatively, some skin bacteria may be derived from other sources not sampled as part of this study, such as other microhabitats occupied by spring peepers (e.g., adults are also terrestrial and arboreal), or vertical transfer from parents [65,66,67].
This study demonstrates the potential for large-scale factors and processes to structure bacterial symbiont diversity, but it is still just the first step in identifying and disentangling the specific factors that interact to shape the amphibian skin bacterial community. Much among-host and among-pond variation remains to be explained. For amphibians and other organisms whose microbial symbionts support immune function, a better understanding of how different factors affect microbial community structure could be key to predicting pathogen dynamics in the host or the success of bioaugmentation efforts. Although unraveling all of the factors structuring amphibian skin bacterial communities will require many more empirical tests, this research highlights the importance of large-scale influences on the structure of host-symbiont assemblages and offers one starting point for assessing the contribution of large-scale factors to microbial symbiont diversity.
References
Adair KL, Douglas AE (2017) Making a microbiome: the many determinants of host-associated microbial community composition. Curr Opin Microbiol 35:23–29
Griffiths SM, Harrison XA, Weldon C, Wood MD, Pretorius A, Hopkins K, Fox G, Preziosi RF, Antwis RE (2018) Genetic variability and ontogeny predict microbiome structure in a disease-challenged montane amphibian. ISME J 12:2506–2517
SanMiguel A, Grice EA (2015) Interactions between host factors and the skin microbiome. Cell Mol Life Sci 72:1499–1515
Ricklefs RE (1987) Community diversity: relative roles of local and regional processes. Science 235:167–171
Lindström ES, Langenheder S (2012) Local and regional factors influencing bacterial community assembly. Environ Microbiol Rep 4:1–9
Nemergut DR, Schmidt SK, Fukami T, O’Neill SP, Bilinski TM, Stanish LF, Knelman JE, Darcy JL, Lynch RC, Wickey P, Ferrenberg S (2013) Patterns and processes of microbial community assembly. Microbiol Mol Biol Rev 77:342–356
Costello EK, Stagaman K, Dethlefsen L, Bohannan BJM, Relman DA (2012) The application of ecological theory toward an understanding of the human microbiome. Science 336:1255–1262
Fierer N, Ferrenberg S, Flores GE, González A, Kueneman J, Legg T, Lynch RC, McDonald D, Mihaljevic JR, O’Neill SP, Rhodes ME, Song SJ, Walters WA (2012) From animalcules to an ecosystem: application of ecological concepts to the human microbiome. Annu Rev Ecol Evol Syst 43:137–155
Mihaljevic JR (2012) Linking metacommunity theory and symbiont evolutionary ecology. Trends Ecol Evol 27:323–329
Leibold MA, Holyoak M, Mouquet N, Amarasekare P, Chase JM, Hoopes MF, Holt RD, Shurin JB, Law R, Tilman D, Loreau M, Gonzalez A (2004) The metacommunity concept: a framework for multi-scale community ecology. Ecol Lett 7:601–613
Holyoak M, Leibold MA, Holt RD (eds.) (2005) Metacommunities: spatial dynamics and ecological communities. University of Chicago Press
Venkataraman A, Bassis CM, Beck JM, Young VB, Curtis JL, Huffnagle GB, Schmidt TM (2015) Application of a neutral community model to assess structuring of the human lung microbiome. MBio 6:e02284–e02214
Burns AR, Stephens WZ, Stagaman K, Wong S, Rawls JF, Guillemin K, Bohannan BJ (2016) Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J 10:655–664
Bletz MC, Loudon AH, Becker MH, Bell SC, Woodhams DC, Minbiole KPC, Harris RN (2013) Mitigating amphibian chytridiomycosis with bioaugmentation: characteristics of effective probiotics and strategies for their selection and use. Ecol Lett 16:807–820
Rebollar EA, Antwis RE, Becker MH, Belden LK, Bletz MC, Brucker RM, Harrison XA, Hughey MC, Kueneman JG, Loudon AH, McKenzie V (2016) Using “omics” and integrated multi-omics approaches to guide probiotic selection to mitigate chytridiomycosis and other emerging infectious diseases. Front Microbiol 7:68
Woodhams DC, Bletz M, Kueneman J, McKenzie V (2016) Managing amphibian disease with skin microbiota. Trends Microbiol 24:161–164
Longo AV, Savage AE, Hewson I, Zamudio KR (2015) Seasonal and ontogenetic variation of skin microbial communities and relationships to natural disease dynamics in declining amphibians. R Soc Open Sci 2:140377
Krynak KL, Burke DJ, Benard MF (2016) Landscape and water characteristics correlate with immune defense traits across Blanchard’s cricket frog (Acris blanchardi) populations. Biol Conserv 193:153–167
Becker CG, Longo AV, Haddad CF, Zamudio KR (2017) Land cover and forest connectivity alter the interactions among host, pathogen and skin microbiome. Proc R Soc B 284:20170582
Hughey MC, Pena JA, Reyes R, Medina D, Belden LK, Burrowes PA (2017) Skin bacterial microbiome of a generalist Puerto Rican frog varies along elevation and land use gradients. PeerJ 5:e3688
Muletz Wolz CR, Yarwood SA, Campbell Grant EH, Fleischer RC, Lips KR (2018) Effects of host species and environment on the skin microbiome of Plethodontid salamanders. Anim Ecol 87:341–353
Jani AJ, Briggs CJ (2014) The pathogen Batrachochytrium dendrobatidis disturbs the frog skin microbiome during a natural epidemic and experimental infection. 2014. Proc Natl Acad Sci 111:E5049–E5058
Belden LK, Hughey MC, Rebollar EA, Umile TP, Loftus SC, Burzynski EA, Minbiole KPC, House LL, Jensen RV, Becker MH, Walke JB, Medina D, Ibáñez R, Harris RN (2015) Panamanian frog species host unique skin bacterial communities. Front Microbiol 6:1171
Walke JB, Becker MH, Loftus SC, House LL, Teotonio TL, Minbiole KP, Belden LK (2015) Community structure and function of amphibian skin microbes: an experiment with bullfrogs exposed to a chytrid fungus. PLoS One 10:e0139848
Rebollar EA, Hughey MC, Medina D, Harris RN, Ibáñez R, Belden LK (2016) Skin bacterial diversity of Panamanian frogs is associated with host susceptibility and presence of Batrachochytrium dendrobatidis. ISME J 10:1682–1695
Jani AJ, Knapp RA, Briggs CJ (2017) Epidemic and endemic pathogen dynamics correspond to distinct host population microbiomes at a landscape scale. Proc R Soc B 284:20170944
Longo AV, Zamudio KR (2017) Environmental fluctuations and host skin bacteria shift survival advantage between frogs and their fungal pathogen. ISME J 11:349–361
Longo AV, Zamudio KR (2017) Temperature variation, bacterial diversity and fungal infection dynamics in the amphibian skin. Mol Ecol 26:4787–4797
Bates KA, Clare FC, O’Hanlon S, Bosch J, Brookes L, Hopkins K, McLaughlin EJ, Daniel O, Garner TW, Fisher MC, Harrison XA (2018) Amphibian chytridiomycosis outbreak dynamics are linked with host skin bacterial community structure. Nat Commun 9:693
Kilpatrick AM, Briggs CJ, Daszak P (2010) The ecology and impact of chytridiomycosis: an emerging disease of amphibians. Trends Ecol Evol 25:109–118
Lips KR (2016) Overview of chytrid emergence and impacts on amphibians. Philos Trans R Soc B 371:20150465
Voyles J, Rosenblum EB, Berger L (2011) Interactions between Batrachochytrium dendrobatidis and its amphibian hosts: a review of pathogenesis and immunity. Microbes Infect 13:25–32
Harris RN, Brucker RM, Walke JB, Becker MH, Schwantes CR, Flaherty DC, Lam BA, Woodhams DC, Briggs CJ, Vredenburg VT, Minbiole KPC (2009) Skin microbes on frogs prevent morbidity and mortality caused by a lethal skin fungus. ISME J 3:818–824
Hughey MC, Becker MH, Walke JB, Swartwout MC, Belden LK (2014) Batrachochytrium dendrobatidis in Virginia amphibians: within and among site variation in infection. Herpetol Rev 45:428–438
Gahl MK, Longcore JE, Houlahan JE (2012) Varying responses of northeastern north American amphibians to the chytrid pathogen Batrachochytrium dendrobatidis. Conserv Biol 26:135–141
Hyatt AD, Boyle DG, Olsen V, Boyle DB, Berger L, Obendorf D, Dalton A, Kriger K, Hero M, Hines H, Phillott R, Campbell R, Marantelli G, Gleason F, Colling A (2007) Diagnostic assays and sampling protocols for the detection of Batrachochytrium dendrobatidis. Dis Aquat Org 73:175–192
McKenzie VJ, Bowers RM, Fierer N, Knight R, Lauber CL (2012) Co-habiting amphibian species harbor unique skin bacterial communities in wild populations. ISME J 6:588–596
Walke JB, Becker MH, Loftus SC, House LL, Cormier G, Jensen RV, Belden LK (2014) Amphibian skin may select for rare environmental microbes. ISME J 8:2207–2217
Boyle DG, Boyle DB, Olsen V, Morgan JAT, Hyatt AD (2004) Rapid quantitative detection of chytridiomycosis (Batrachochytrium dendrobatidis) in amphibian samples using real-time Taqman PCR assay. Dis Aquat Org 60:141–148
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M, Gormley N, Gilbert JA, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624
Meisel JS, Hannigan GD, Tyldsley AS, SanMiguel AJ, Hodkinson BP, Zheng Q, Grice EA (2016) Skin microbiome surveys are strongly influenced by experimental design. J Invest Dermatol 136:947–956
Aronesty E. (2011) Ea-utils: “command-line tools for processing biological sequencing data”. http://code.google.com/p/ea-utils
Caporaso JG, Kuczynski J, Stombaugh JJ, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R, Mills DA, Caporaso JG (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10:57–59
R Core Team (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna http://www.R-project.org
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2015) Vegan: community ecology package. http://CRAN.R-project.org/package=vegan
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Aust Ecol 26:32–46
Whittaker RH (1972) Evolution and measurement of species diversity. Taxon 21:213–251
Fournier DA, Skaug HJ, Ancheta J, Ianelli J, Magnusson A, Maunder M et al (2012) AD model builder: using automatic differentiation for statistical inference of highly parameterized complex nonlinear models. Optim Methods Softw 27:233–249
Skaug H, Fournier D, Bolker B, Magnusson A, Nielsen A (2014) Generalized linear mixed models using AD model builder. R package version 0.8.0
Legendre P, Legendre L (2012) Numerical ecology. 3rd English edition. Elsevier
Tuomisto H, Ruokolainen K (2006) Analyzing or explaining beta diversity? Understanding the targets of different methods of analysis. Ecology 87:2697–2708
Paliy O, Shankar V (2016) Application of multivariate statistical techniques in microbial ecology. Mol Ecol 25:1032–1057
Legendre P, Gallagher ED (2001) Ecologically meaningful transformations for ordination of species data. Oecologia 129:271–280
Campbell BJ, Yu L, Heidelberg JF, Kirchman DL (2011) Activity of abundant and rare bacteria in a coastal ocean. Proc Natl Acad Sci U S A 108:12776–12781
Walke JB, Becker MH, Hughey MC, Swartwout MC, Jensen RV, Belden LK (2017) Dominance-function relationships in the amphibian skin microbiome. Environ Microbiol 19:3387–3397
Austin JD, Lougheed SC, Neidrauer L, Chek AA, Boag PT (2002) Cryptic lineages in a small frog: the post-glacial history of the spring peeper, Pseudacris crucifer (Anura: Hylidae). Mol Phylogenet Evol 25:316–329
Christian N, Whitaker BK, Clay K (2015) Microbiomes: unifying animal and plant systems through the lens of community ecology theory. Front Microbiol 6:869
Burns AR, Miller E, Agarwal M, Rolig AS, Milligan-Myhre K, Seredick S, Guillemin K, Bohannan BJ (2017) Interhost dispersal alters microbiome assembly and can overwhelm host innate immunity in an experimental zebrafish model. Proc Natl Acad Sci 114:11181–11186
Fitzpatrick BM, Allison AL (2014) Similarity and differentiation between bacteria associated with skin of salamanders (Plethodon jordani) and free-living assemblages. FEMS Microbiol Ecol 88:482–494
Banning JL, Weddle AL, Wahl GW, Simon MA, Lauer A, Walters RL, Harris RN (2008) Antifungal skin bacteria, embryonic survival, and communal nesting in four-toed salamanders, Hemidactylium scutatum. Oecologia 156:423–429
Walke JB, Harris RN, Reinert LK, Rollins-Smith LA, Woodhams DC (2011) Social immunity in amphibians: evidence for vertical transmission of innate defenses. Biotropica 43:396–400
Hughey MC, Delia J, Belden LK (2017) Diversity and stability of egg-bacterial assemblages: the role of paternal care in the glassfrog Hyalinobatrachium colymbiphyllum. Biotropica 49:792–802
Acknowledgments
We thank P. Shirk, J. Vonesh, J. Wyderko, and S. Zemmer for assistance in the field; P. Sattler at Liberty University, G. Eaton at Claytor Nature Study Center, and J. Vonesh at Virginia Commonwealth University for recommending sites in Lynchburg, Bedford, and Richmond, respectively; J. Touchon for R code consultations; C. Herbold for stats advice; and Z. Herbert at the Dana Farber Cancer Institute Molecular Biology Core Facility for Illumina MiSeq amplicon sequencing. We are also thankful for the comments of anonymous reviewers that improved the manuscript.
Funding
This work was supported by the Morris Animal Foundation [D10ZO-028] and the National Science Foundation [DEB-1136640].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This research was conducted with approval from Virginia Tech’s Institutional Animal Care and Use Committee (protocol 10-029-BIOL) and with permission from the Virginia Department of Game and Inland Fisheries (permit 44303).
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
ESM 1
(R 3 kb)
Rights and permissions
About this article
Cite this article
Hughey, M.C., Sokol, E.R., Walke, J.B. et al. Ecological Correlates of Large-Scale Turnover in the Dominant Members of Pseudacris crucifer Skin Bacterial Communities. Microb Ecol 78, 832–842 (2019). https://doi.org/10.1007/s00248-019-01372-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-019-01372-0