Abstract
Demands for clean and sustainable processes and products that are environmentally friendly are challenging biotechnologists to develop new strategies to produce fuels and chemicals. As the petroleum demands rise together with the concern of climatic and environmental changes, there is an increasing interest for renewable energy. Sugars present in the lignocellulosic biomass can be used as raw material in biotechnological processes employing yeasts as catalysts. Several known yeasts such as Saccharomyces cerevisiae assimilate glucose but lack the efficiency to consume xylose. Due to industrial interest, there has been an increasing effort to discover and construct new xylose-assimilating yeast strains. In this sense, due to the diversity and metabolic potential, several non-conventional yeasts species were isolated, identified, and physiologically and genetically characterized in the last years. The current review sought to summarize the main characteristics as well as the biotechnological applications of non-conventional yeasts for xylose utilization. First, it will present and discuss the data about non-conventional yeasts that naturally and efficiently assimilate xylose as Scheffersomyces, Meyerozyma, Candida, Spathaspora, and Kluyveromyces. Then the yeasts Komagataella, Yarrowia, and Ogataea that do not assimilate xylose or poorly assimilate xylose justifying genetic manipulation to increase xylose utilization will also be presented. In each case, basic information about yeast taxonomy, morphology, and physiology will be presented, and the clearest biotechnological application will be introduced.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
2.1 Introduction
2.1.1 Lignocellulosic Biomass
The energetic source most used nowadays is petroleum, which is employed as fuel and raw material for the production of chemicals with several industrial applications (Sun et al. 2013; Gallo and Trapp 2017). Since petroleum stocks are decreasing, prices are volatile and its burn is contributing to global pollution, development of sustainable and environmentally friendly processes to produce fuels and chemicals is required. Lignocellulosic biomass is a widely available resource that can be employed in such processes (Gallo and Trapp 2017).
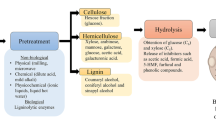
Biomass is the most abundant renewable feedstock resource available on the planet. Briefly, lignocellulosic biomass (vegetal cell wall) is composed of the sugar polymers cellulose and hemicellulose and the macromolecule lignin (Chandel et al. 2011). Cellulose, the most abundant polymer in biomass (Table 2.1), is composed of a homo-polysaccharide D-glucose (C6-hexose) linked by B-1,4-glucosidic bonds organized in complex microfibriles (Jørgensen et al. 2007). Hemicellulose is a heteropolysaccharide polymer composed of pentose and hexose sugars (C5 and C6, respectively). The nature of the main chain is formed by D-xylose or D-galactose, depending on the biomass source, which is ramified with other sugars, like L-arabinose, D-glucuronic acid, D-glucose, D-mannose, and other molecules, such as acids (van Wyk 2001). Lignin is a complex macromolecule composed of phenyl-propane units that protect cellulose-hemicellulose giving integrity to the lignocellulose structure (Canilha et al. 2010; Kim 2018).
Several biotechnological processes for the production of fuels and other chemicals from biomass have been developed and evaluated from laboratorial to industrial scale (Garrote et al. 2002). A common requirement in the processes is the hydrolysis of lignocellulose to release the monosaccharides present in the cellulose and hemicellulose fractions, which finally can be converted by the microorganism of choice to the desired product (Ji et al. 2017).
The composition and concentration of sugars in the lignocellulosic hydrolysate will vary according to biomass and pretreatment/hydrolysis conditions employed (Almeida et al. 2011). Glucose and xylose are, respectively, the most abundant hexose and pentose sugars in nature and consequently the most available in biomass hydrolysates (Table 2.1). Glucose is promptly metabolized and converted into a variety of fuels and chemicals by microorganisms (Elshahed 2010), whereas xylose consumption is limited to a reduced number of species (Jeffries 1983). Thus, xylose valorization processes have gained much attention lately.
2.1.2 Xylose Application Overview
The usage of xylose started to gain attention in the latest years of the 1920s when the National Bureau of Standards in the USA began to evaluate the possibility to aggregate value to crop wastes (Schreiber et al. 1930). After the discovery that some bacteria produced acetic and lactic acids from xylose fermentation, methods to recover xylose from different biomasses started to be investigated (Schreiber et al. 1930). Nowadays, D-xylose can be extracted from several different crops (Table 2.1) (Satish and Murthy 2010). Many efforts are being performed to recover as much xylose (and other sugars) as possible from the biomass with minimal generation of degradation products (Christopher 2012). The main process to obtain xylose is through a pretreatment process that hydrolyzes the hemicellulose with sulfuric acid (Zhang et al. 2014; Gallo and Trapp 2017).
Xylose can be applied in different industrial sectors, including food and medicine (Gunah 2011). It can be dehydrated by acid catalysis to produce furfural (Bozell and Petersen 2010), and it also can be reduced using nickel catalysts under high temperature and pressure to produce xylitol (Akinterinwa et al. 2008). Biologically, it can be converted to organic acids, lipids, alcohols, and other chemicals by pentose-assimilating microorganisms. For example it can be fermented to ethanol (Webb and Lee 1990; Bajwa et al. 2011), reduced to xylitol and arabitol (Gı́rio et al. 2000; Faria et al. 2014; Martins et al. 2018), or oxidized to xylonic and xylaric acid (Gallo and Trapp 2017) depending on the microorganism employed. With the increasing interest to produce chemicals from biomass through biotechnological processes, new xylose-assimilating microorganisms are being identified and/or constructed by bioprospecting and metabolic engineering strategies (Jeffries 2006; Mans et al. 2018).
2.1.3 Microbial Utilization of Xylose
Bacteria, eukaryotes (yeasts and fungus), and archaea are capable of using xylose as a carbon source (Sampaio et al. 2003; Johnsen et al. 2009; Kręgiel et al. 2017; Kim and Woo 2018), however, through specific pentose assimilation pathways (Martins et al. 2018). Most of the bacteria and some anaerobic fungi such as Piromyces (Harhangi et al. 2003) and Orpinomyces (Madhavan et al. 2009) use an isomerization pathway to assimilate xylose. In this pathway, a xylose isomerase (XI, EC 5.3.1.5) interconverts xylose to xylulose, which is then phosphorylated by the xylulokinase enzyme (XK, EC 2.7.1.17) to xylulose-5-phosphate, to enter the pentose phosphate pathway (PPP) (Kwak and Jin 2017). The archaea employ an oxidative pathway where xylose is oxidized to α-ketoglutarate (tricarboxylic acid cycle intermediate) by D-xylose dehydrogenase, xylonate dehydratase, 2-keto-3-deoxyxylonate dehydratase, and α-ketoglutarate semialdehyde dehydrogenase (Johnsen et al. 2009)
Yeast employs an oxidoreductase pathway (Fig. 2.1). In this pathway, xylose is reduced to xylitol by an NAD(P)H-dependent xylose reductase (XR, EC 1.1.1.30). Then, an NAD+-dependent xylitol dehydrogenase (XDH, EC 1.1.1.9) oxidates xylitol to D-xylulose, which in turn is phosphorylated to D-xylulose-5P by the enzyme xylulose kinase (XK, EC 2.7.1.17). D-xylulose-5P can be metabolized by the pentose phosphate pathway (PPP) and glycolysis (Wang et al. 1980).
Xylose assimilation by yeasts. Dashed arrow – summarize stretches of the metabolic pathway; names in black represent the preferred co-factor for the yeast. Numbers represent the product converted naturally by the yeasts using xylose as carbon source: 1Ethanol producer; 2Xylitol producer; 3Lipids producer
2.1.3.1 Xylose Conversion by Non-conventional Yeast (NCY)
Fermentation technology has continued to evolve in producing chemicals with industrial applications, and microbial physiology has become a fast-growing and increasingly impressive branch of science (Palsson et al. 1981). Saccharomyces cerevisiae, the yeast most employed in the industry nowadays, can convert efficiently glucose to ethanol and has been described as a producer of a large range of chemical compounds as biofuels (ethanol, farnese, isobutanol), pharmaceutical drugs, and carboxylic acids (Abbott et al. 2009). Its robustness makes it tolerant to contamination and inhospitable environment (Jeffries 2006). These phenotypes, among others, have led to the widespread study of S. cerevisiae and its development as a model eukaryotic host for chemical biosynthesis (Löbs et al. 2017). Although S. cerevisiae presents some limitations regarding metabolic profile, it is incapable of fermenting five-carbon sugars, such as xylose (Hou et al. 2017). Thus, several S. cerevisiae strains able to convert xylose to different chemicals have been constructed through genetically engineering strategies (Kim et al. 2013; Moysés et al. 2016). This choice is usually made because of the range of strains and genetic engineering tools available for this yeast. However, xylose metabolism by recombinant strains rarely reaches rates comparable to the glucose, due to different metabolic constraints (reviewed at Moysés et al. 2016).
On the other hand, several yeast species naturally capable of using xylose as a sole carbon source have been isolated and identified, and they present unique characteristics of industrial interest. Species-specific characteristics have been identified among such yeasts, some showing more fermentative metabolism and producing ethanol than others. For instance, Scheffersomyces stipitis and Spathaspora passalidarum are capable of fermenting xylose to ethanol with high yields and productivity (Veras et al. 2017), whereas Yarrowia lipolytica showed a respiratory metabolism, and it is capable of synthesizing and accumulate high levels of intracellular lipids (Beopoulos et al. 2009). In addition to the low number of xylose-consuming yeast strains reported among almost 1000 yeast taxa, there is still few information about physiology and genetics of many species (Nguyen et al. 2006; Kurtzman 2011b).
Due to the incomplete understanding of genetics, metabolism, and cellular physiology and availability of genome-editing tools, when compared with S. cerevisiae (Löbs et al. 2017), those yeasts have been named as non-conventional yeasts. The term “non-conventional” yeast is being used by microbiologists and biotechnologists to define yeasts that do not belong to Saccharomyces cerevisiae and Schizosaccharomyces pombe (Sreekrishna and Kropp 1996; Kręgiel et al. 2017). Different metabolic pathways, variability of fermentative profile, and growth physiology are some of the advantages of using non-conventional yeasts over S. cerevisiae (Kręgiel et al. 2017; Rebello et al. 2018)
As the molecular information is becoming available, the divergence of class, genus, and species is being made every year, and reclassification of non-conventional yeast species became common. Non-conventional yeasts present a huge, yet barely exploited, diversity that is an excellent opportunity to achieve some new strategies in biotechnologist research. Many of these non-conventional yeast species exhibit industrially relevant traits such as the ability to utilize complex substrates as nutrients and extreme tolerance against stress and fermentation inhibitors (Mukherjee et al. 2017). Such yeasts may be employed directly in biotechnological processes or used to identify traits of industrial interest for transferring these properties (genes that confers the specific advantage) to other yeasts (Mukherjee et al. 2017). Indeed, physiological, molecular, and genetic characterization and development of toolkits for non-conventional yeasts is readily increasing (Yaguchi et al. 2018), and the potential applications of non-conventional yeast species have been rising considerably (Radecka et al. 2015).
The current review sought to summarize the main characteristics as well as the biotechnological applications of non-conventional yeasts for xylose utilization. First, it will present and discuss the data about the most known non-conventional yeasts focusing on yeasts that naturally assimilate xylose in an efficient way such as Scheffersomyces, Meyerozyma, Candida, Spathaspora, and Kluyveromyces. Then, it will present data about the yeasts that do not assimilate xylose as Komagataella and poorly assimilate xylose as Yarrowia and Ogatae, which are being genetically modified with the purpose of generating and increasing the xylose incorporation. In each case, basic information about yeast taxonomy, morphology, and physiology will be presented as well as the clearest biotechnological application.
2.2 Yeast Descriptions and Biotechnological Applications
2.2.1 Scheffersomyces
Originally, some of the species assigned to Scheffersomyces belonged to Pichia genus because of its morphology, phenotype, and absence of nitrate assimilation. In an attempt to revise the polyphyletic genus Pichia, Kurtzman and Suzuki (2010) performed a comparative analysis of D1/D2 LSU and SSU rRNA gene sequences, resulting in a strong support for a change of genus of some yeasts within the Pichia clade, which included P. stipitis, P. segobiensis, P. spartinae, and some species of Candida (Kurtzman et al. 2010). Therefore, the ascosporic species of Pichia clade were transferred to the new genus Scheffersomyces and renamed as Scheffersomyces stipitis, S. segobiensis, and S. spartinae (Kurtzman et al. 2010).
Additional phylogenetic analysis suggests that the common ancestor of the Scheffersomyces genus probably was able to assimilate and ferment D-xylose and cellobiose. Currently, all the yeasts placed in the Scheffersomyces genus are either a D-xylose or a cellobiose fermenter; the only exception is the S. spartinae species, which does not ferment xylose or cellobiose (Urbina and Blackwell 2012). Thus, a subdivision of the Scheffersomyces genus into three smaller subclades was suggested to gather all the yeasts with the different characteristics (Table 2.2). Based on the D-xylose fermentation, and lack of bootstrap support in the D1/D2-SSU tree (52%), the placement of the S. spartinae in the Scheffersomyces genus is uncertain (Kurtzman et al. 2010).
Scheffersomyces yeasts have some traits that are interesting for several biotechnological applications. S. stipitis is one of the most efficient microorganism for xylose and lignocellulose fermentation, while S. amazonensis and S. stambukii have been highlighted as good xylitol producers (Jeffries et al. 2007; Lopes et al. 2018). S. stipitis is also capable of metabolizing several sugars found in lignocellulosic biomass, such as glucose, mannose, galactose, and cellobiose (Agbogbo and Coward-Kelly 2008; Jeffries and Van Vleet 2009). It is a predominantly haploid, homothallic, and hemiascomycetous yeast. During vegetative growth, budded cells are spherical to elongate and pseudohyphae are formed. Asci produce one or two hat-shaped ascospores (Jeffries et al. 2007; Kurtzman and Suzuki 2010). Phylogenetic analyses using the LSU D1/D2 sequences placed S. stipitis next to the Spathaspora and Lodderomyces clade (Fig. 2.2).
Phylogenetic tree based on LSU-D1/D2 sequence from different microorganism. Sequences were withdrawn from Yeast IP database and NCBI. Alignment was performed using Clustal W program. Tree was reconstructed using neighbor-joining analysis using 25 sequences. It was used default settings and 1000 bootstrap replicates. For a more detailed phylogeny analysis, see Riley et al. (2016)
Many studies have pointed the ethanol production from xylose by S. stipitis cells. It was found that batch fermentations of S. stipitis produce high ethanol amounts, reaching yields of 0.40 to 0.48 g/g xylose (Agbogbo and Coward-Kelly 2008; Veras et al. 2017), values that are close to the theoretical maximum yield. S. stipitis shows lower cell yields and higher ethanol yields when cultivated in xylose compared to that in glucose as sole carbon source and under low aeration condition (Su et al. 2015). The same behavior was also observed for the xylose-fermenting yeast S. passalidarum (Su et al. 2015). To explain this observation, Su et al. (2015) suggested that the net ATP yields are substantially lower when yeast is growing on xylose; consequently, cells need to consume more carbon to make the same amount of biomass. It is important to note that the energy expended per mole of sugar should be similar for a C5 or C6 sugar considering sugar uptake and xylulose phosphorylation but the C moles of carbon would be 17% less in each step.
Apart from the great potential of S. stipitis for 2G (second generation) bioethanol production, some critical points should be addressed to achieve the employment of this yeast at industrial scale, including improved conversion rates, tolerance to hydrolysate inhibitors, tolerance to high concentrations of ethanol, and necessity of controlled aeration conditions. S. stipitis has evolved in low-sugar environment prevailing in the beetle midgut; consequently, its glycolytic flux, ethanol tolerance, and productivity from glucose and xylose are lower than that obtained from S. cerevisiae when cultivated on glucose (Jeffries and Van Vleet 2009). In addition, glucose represses genes involved in xylose assimilation as XYL1, and XYL2, XYL3 (encoding XR, XDH, XK, respectively), and some genes from PPP (Jeffries et al. 2007; Jeffries and Van Vleet 2009). Indeed, acetic acid, furaldehydes, and phenolic compounds present in lignocellulosic hydrolysates can inhibit growth, viability, and fermentative performance by this yeast (Ma et al. 2017).
Several studies focus on solving some of these problems. Different pretreatment processes and detoxification steps have been suggested to alleviate the formation of toxic compounds or eliminate them before fermentation (Ferreira et al. 2011; Nakanishi et al. 2017; Artifon et al. 2018). Using cell recycling and decreasing temperature for each batch, S. stipitis showed increased xylose consumption and ethanol production during cultivation on a detoxified sugarcane bagasse hydrolysate, pretreated with NaOH/ anthraquinone (AQ), and then enzymatically hydrolyzed (Nakanishi et al. 2017). In turn, engineered S. stipitis strains with improved traits have been developed mainly by random mutagenesis approaches, adaptive evolution, protoplast fusion, and genome shuffling (Ma et al. 2017; Löbs et al. 2017). Through UV mutagenesis and evolutionary adaptation, Ma et al. (2017) isolated a S. stipitis mutant with improved fermentation performance and tolerance against ethanol and lignocellulosic derived inhibitors. Glucose and xylose consumption rates of the mutant were 4.18 g/L.h and 2.39 g/L.h, respectively, values approximately two times higher than those of the parental strain (2.09 g/L.h and 1.4 g/L.h, respectively) in batch fermentation using YP medium with 100 g/L sugar. Similar improvement was observed for ethanol productivity (mutant strain, 2.01 and 1.10 g/L.h; parental strain, 0.99 and 0.62 g/L.h on glucose and xylose, respectively). Besides that, employing membrane integrated continuous fermentation system and rice straw hydrolysate non-detoxified as feedstock, 43.2 g/L ethanol titer and 2.16 g/L/h ethanol productivity were achieved by the yeast (Ma et al. 2017). These values are one of the best results obtained so far (Table 2.3).
Rational metabolic engineering of non-conventional yeasts requires a better understanding of their physiology, biochemistry, and genetics (Jeffries and Van Vleet 2009). Moreover, due to a lack of efficient genome-editing tools adapted to Scheffersomyces, relatively few studies have attempted the rational modification of these yeasts (Löbs et al. 2017). In an example, an engineered S. stipitis strain, expressing a heterologous LDH gene coding for lactate dehydrogenase, produced 41 or 58 g/L lactate from approximately 100 g/L of glucose or xylose, respectively (Ilmen et al. 2007). A more elaborate pathway was introduced in S. stipitis for fumaric acid production from xylose, based on the overexpression of a heterologous reductive pathway from Rhizopus oryzae. Additionally, codon optimization of the gene sequences introduced deletion of native fumarase genes, and overexpression of heterologous C4-dicarboxylic acid transporter led to 4.67 g/L fumaric acid titer from 20 g/L xylose (Wei et al. 2015). Table 2.3 summarizes some results of synthetic medium or lignocellulosic hydrolysate fermentation by S. stipitis strains.
Another interesting biotechnological application of S. stipitis in the biorefinery context was in the production of a biosurfactant (Franco Marcelino et al. 2017). Hemicellulosic sugarcane bagasse hydrolysate was used as raw material for the production of a green glycolipidic biosurfactant by S. stipitis, which shows significant larvicidal proprieties against Aedes aegypti, a vector of neglected tropical diseases. In addition, the biosurfactant showed emulsifying property in hydrophobic compounds that has potential for applications in bioremediation processes as well as cosmetic, agricultural, and food formulations.
To gain more information about the yeast S. stipitis, its genome was sequenced and analyzed. The genome was estimated in 15.4 Mb, accounting with 5841 predicted genes of which 72% have a single exon (Jeffries et al. 2007; Table 2.4). The species uses the alternative yeast nuclear codon that substitutes serine for leucine when CUG is specified, placing S. stipitis within the CUG-serine clade, together with other xylose-fermenting yeasts (Fig. 2.2) (Jeffries et al. 2007; Cadete and Rosa 2018). Kinases, helicases, transporters (sugar and major facilitator superfamily), and domains involved in transcriptional regulation are the most frequent domains characterized. Genes related to xylose assimilation, oxidative pentose phosphate pathway, glycolytic cycle, tricarboxylic acid cycle (TCA), and ethanol production were present in isoforms similar to those found in other yeasts (Jeffries et al. 2007).
In several yeasts, the redox imbalance concern to the first two steps of xylose metabolism, involving xylose reductase and xylitol dehydrogenase enzymes, has a significant impact on xylose fermentation and leads to higher xylitol production (Bruinenberg et al. 1983; de Albuquerque et al. 2014; Veras et al. 2017). However, S. stipitis presents various strategies for NAD+ and NADPH regeneration (Jeffries et al. 2007), and besides that its xylose reductase enzyme, encoded by XYL1 gene, can use both NADH and NADPH cofactors, although it shows higher affinity toward NADPH (Fig. 2.1) (Verduyn et al. 1985). All these factors may explain the efficient fermentation of xylose observed for S. stipitis yeasts.
Although S. stipitis is the most studied species from this clade, other Scheffersomyces yeasts also possess interesting biotechnological traits. Currently, around 20 yeast species belong to the Scheffersomyces genus (Table 2.2). Several of them have the ability to ferment D-xylose to ethanol, including S. insectosa, S. lignosus, S. parashehatae, S. quercinus, S titanus, S. xylosifermentans, and S. shehatae. However, all of them showed less efficient xylose fermentation capabilities than S. stipitis (du Preez and van der Walt 1983; Liu et al. 2016). In turn, other yeasts were reported as good xylitol producers. In a comparative analysis of two species of Scheffersomyces, S. amazonensis was able to produce 26.4 g/L xylitol, achieving a yield of 0.554 g/g and a volumetric productivity of 0.732 g/L/h, while S. stambukii produced 33.0 g/L xylitol, reaching a yield of 0.663 g/g and a volumetric productivity of 0.687 g/L/h (Cadete et al. 2016a; Lopes et al. 2018). These data reveal the great potential of these strains for the biotechnological production of xylitol (Table 2.3).
2.2.2 Meyerozyma
The genus Meyerozyma was proposed to englobe two closely related ascosporic species of yeasts belonging to the genus Pichia and, consequently, reduce the polyphyletic characteristic of the genus (Kurtzman et al. 2010). Pichia guilliermondii (anamorph Candida guilliermondii) and Pichia caribbica (anamorph Candida fermentati) were reclassified to Meyerozyma guilliermondii and Meyerozyma caribbica, respectively (Wrent et al. 2016), because of its phylogenetic proximity and common production of Coenzyme Q-9 (Kurtzman et al. 2010).
Such classifications were proposed through analysis of the similarity between the DNA sequences of the domains D1/D2 LSU rRNA and the sequence SSU rRNA of the yeasts that have the common ability to produce coenzyme Q-9 (CoQ-9) (Yurkov et al. 2017). Those analyses provided enough data for allowing the redistribution of the CoQ-9-producing species in a distinct genus and also reuniting those species in a different group, named M. guilliermondii species complex (Kurtzman et al. 2010).
Like Scheffersomyces yeasts, the Meyerozyma genus belongs to the Saccharomycotina CTG clade and currently englobe two sexual species, M. guilliermondii and M. caribbica (Fig. 2.2) (Yurkov et al. 2017). The phylogenetic structure of the yeasts that belongs to the clade CTG has been studied for more than four decades and is still not fully understood, in part due to the morphological and metabolic similarity between the yeasts but is also due to the large range of synonyms for each species, what is related with the different names given to the same yeasts but in different sexual states (anamorph/teleomorph) (De Marco et al. 2018a). For example, the yeast M. guilliermondii is the sexual state of C. guilliermondii (Wickerham and Burton 1954), and a very few wild strains of C. guilliermondii are sexually reactive, what represents a teleomorph-anamorph pair (Kurtzman et al. 2010).
Most species belonging to the Meyerozyma genus have been found in insects or related environments like fruits and soil (Yurkov et al. 2017). The type yeast of the genus M. guilliermondii has been found for the first time in eggs of insects in the USA, and the other component of the genus M. caribbica was found in sugarcane juice, in Cuba (Kurtzman et al. 2010). In addition to M. guilliermondii and M. caribbica, it was recently proposed the yeast Meyerozyma amylolytica A. M. Yurkov and Péter sp. Nov. (Yurkov et al. 2017) as a new species into the Meyerozyma genus (Yurkov et al. 2017). This species was found in Germany trees, and like other members of the genus, its origin is related to insects (Yurkov et al. 2017).
Some studies have shown that the yeast M. guilliermondii displays antimicrobial activity. This yeast has the ability to inhibit the growth of fungi (Coda et al. 2013), bacteria (Zhao et al. 2010), and even protozoa (Dantán-González et al. 2015). Thus it can be used as a biocontrol agent in the food and agriculture industries, acting on the preservation of fruits and vegetables after the harvest (Yan et al. 2018). But even if M. guilliermondii is not reported as a pathogen in laboratory tests, the anamorph state of the yeast M. guilliermondii (Candida guilliermondii) is an opportunistic pathogen (Corte et al. 2015). Because of the proximity with C. guilliermondii (more than 98% of nuclear DNA compatibility) (Kurtzman et al. 2010), M. guilliermondii have a questionable GRAS (generally recognized as safe) status, confusing the health legislation in many countries (Corte et al. 2015).
M. guilliermondii is used in industrial production of vitamin B2 (riboflavin), due to its flavinogenic potential (Romi et al. 2014). Indeed, most yeasts from the Candida genus have the potential to overproduce riboflavin under iron-limitation conditions. Three genes that regulate the riboflavin production in M. guilliermondii (P. guilliermondii) were identified. Although the specific role of the genes in the cell metabolism is not fully understood, it was also shown that the regulation of iron and riboflavin production is closely related in this yeast, (Protchenko et al. 2011).
Strains of Meyerozyma genus have also been evaluated for biotechnological application for production of chemicals, due to their natural ability to assimilate and utilize pentose in its metabolism. When the subject is xylose fermentation, the strains of the anamorph pair C. guilliermondii FTI 20037 has received more attention, because it can produce xylitol, with higher yields and productivity than the most strains in its complex, as shown in Table 2.5 (Vaz de Arruda et al. 2017). Xylitol is an important sugar-alcohol with five carbons derived from xylose that can be used as a substitute for the currently used sugar (sucrose). It has many healthy properties, such as it prevents tooth decay and airway inflammations and can also be an intermediate in the production of other chemicals of interest, like ethylene glycol, an important lubricant (Venkateswar Rao et al. 2015).
The Meyerozyma species are also able to assimilate the xylose present in lignocellulosic hydrolysates, and even in the presence of inhibitors, they were capable of producing xylitol and/or ethanol (Table 2.5) (Martini et al. 2016; Vaz de Arruda et al. 2017).
M. caribbica is another member of Meyerozyma genus but is less studied. M. caribbica is also a hexose and pentose naturally ferment yeast, and it can be used as a biocontrol microorganism as well. In addition to that, this species has a nonpathogenic anamorph state, known as Candida fermentati, which may facilitate its biotechnological utilization in food and health industries (Romi et al. 2014). Phylogenetically M. caribbica is closely related with M. guilliermondii (Fig. 2.2), and these species are often misidentified by the classic molecular differentiation method employing the sequence of the D1/D2 domains (Romi et al. 2014). Romi et al. (2014) proposed a new protocol to differentiate the species using ITS-RFLP combined with an in silico selection of restriction enzymes with further in vitro validations that proved work for M. guilliermondii and M. caribbica species differentiation (Romi et al. 2014).
The Meyerozyma species have been studied for a long time, but the biotechnological utilization is currently around the riboflavin production and prevention of mold decay in fruits and vegetables (Corte et al. 2015). Research from the past decade are focusing in the great natural potential of xylose assimilation and fermentation capability, shown by the Meyerozyma yeasts, for industrial uses of biomass in biorefineries (Hernández-Pérez et al. 2016).
2.2.3 Candida
The genus Candida belongs to the Ascomycota phylum, Hemiascomycetes class, and Saccharomycetales order and covers all imperfect ascomycetes species whose phylogenetic relationships have not been defined (Hommel and Ahnert 1999). Candida is a dimorphic fungus that can present as yeast globose, ellipsoidal, cylindroidal or elongate, occasionally ogival, triangular, or lunate (Kurtzman 2011b). In addition, this genus presents a great variety of CoQ types, with the species that were found to sporulate having teleomorphic counterparts in 11 different genera (Schauer and Hanschke 1999).
Candida is phylogenetically heterogeneous and covers more than 310 species. The identification of Candida species is complex and based on morphological, physiological, chemotaxonomic characteristics and in the sequence analysis of the D1/D2 domain of rDNA 26S (Odds 2010). Candida species show a wide range of physiological properties and are widespread in natural habitats with a high content of organic matter, with low or high temperature, and high osmolarity. The majority is mesophilic and grow well in temperatures between 25 and 30 °C (Spencer and Spencer 1997; Odds 2010). The yeasts in this genus are aerobic, not growing in an anaerobic condition, but some strains survive and reproduce under microaerophilic conditions (Lachance 2011).
Candida species are mostly found in nutrient-rich habitats such as soil, rotting vegetation (fruit peel and decaying fruit), plants (leaf surfaces, nectaries, and nectar of flowers, flower petals, and other flower parts), and insects that feed on plants (Hommel and Ahnert 1999). They are able to assimilate and ferment xylose (from wood degradation) and cellobiose, and also a smaller number of species oxidize aliphatic hydrocarbons (components of the cuticle plant), degrade starch, or use methanol as a possible metabolite of pectin catabolism (Schauer and Hanschke 1999). Naturally, most of them require individual vitamins produced mainly in plant materials.
Like other yeasts, Candida species are able to metabolize xylose through via oxido-reductive pathway (Fig. 2.1) (Chakravorty et al. 1962). Among the xylitol-producing yeasts, Candida strains have shown high XR-specific activity, associated with dependence on the NADPH cofactor, which favors the production of xylitol (Veras et al. 2017). Candida strain application on fermentative processes has gained attention due to their capacity to produce xylitol with high yields, up to 65 and 85% of theoretical maximum (Bier et al. 2007). Recently, different Candida species have been considered for the production of chemicals, mainly ethanol and xylitol, using different substrates, such as horticultural waste (Zhang et al. 2012), olive tree pruning (Mateo et al. 2015), rice straw (Swain and Krishnan 2015), corncob (Kumar et al. 2018), and sugarcane bagasse (Tizazu et al. 2018). The most common Candida species evaluated in biotechnological processes are discussed in the next topics.
2.2.3.1 Candida magnoliae
Candida magnoliae is a member of the of Starmerella clade, presents CoQ 9 (Yamada and Kondo 1972) and % mol GC of 60% (Nakase and Komagata 1971). Based on D1/D2 LSU rRNA gene sequences, C. magnoliae is a close relative to C. sorbosivorans and C. geochares (Rosa et al. 2003). They were collected and isolated from bees and their habitats such as flower of Magnolia sp. (Magnoliaceae), concentrated orange juice, and gut of a bee (Kurtzman 2011b). This species is interesting in biotechnology mainly in the production of substitute sweeteners such as erythritol from glucose with productivity of 0.54 g/L.h and 43% of conversion yield based on glucose (Yang et al. 1999), mannitol in the presence of fructose and glucose as carbon sources reaching a productivity of 1.03 g/L.h and 83% of conversion (Song et al. 2002), and xylitol from xylose-containing lignocellulosic hydrolysates with productivity of 0.21 g/L.h and 54.5% of conversion yield based on xylose (Wannawilai et al. 2017).
2.2.3.2 Candida tropicalis
Candida tropicalis, a member of the phylogenetic placement of Lodderomyces-Spathaspora, also presents CoQ 9 (Yamada and Kondo 1972) with % mol GC between 34.4 and 36.1% (Stenderup and Bak 1968; Nakase and Komagata 1971; Meyer et al. 1975). The first strains of this species were found in Jamaica from fruits of Stenocereus hystrix (Cactaceae) and Royen cactus flower (Cephalocereus royenii, Cactaceae) and as a contaminant of a clinical specimen of Candida albicans in Brazil (Kurtzman 2011b). C. tropicalis has a high degree of similarity with related species and exhibits variability among lineages, so their identification by growth characteristics is problematic. Despite this being one of the most frequently found clinical yeast species after C. albicans (Moran 2004), in recent years many kinds of research have been carried out using C. tropicalis in the fermentation of xylose to produce xylitol (Table 2.6).
C. tropicalis has been applied in different bioprocesses, especially for the production of ethanol or xylitol. It usually shows better productivity and yields when compared to other Candida strains. C. shehatae produced ethanol with a productivity of 0.12 g/L.h and yield of 0.50 g/g in a medium containing rice straw hydrolysate with 20 g/L of xylose and 40 g/L of glucose (Yuvadetkun et al. 2018). On the other hand, C. tropicalis produced ethanol in a medium containing rice straw hydrolysate presenting productivity 2 times and yields 12 times higher than C. shehatae (Table 2.6) (Swain and Krishnan 2015). The differences are significant even if a direct comparison cannot be made due to the variations in the experimental conditions.
In relation to the production of xylitol, C. tropicalis has been outstanding for achieving better productivity and yields, especially when applied in a medium containing lignocellulosic hydrolysates. C. tropicalis reached xylitol productivity and yields up to 0.80 g/L.h and 0.85 g/g in a medium containing lignocellulosic hydrolysates (Table 2.6). C. tropicalis’ ability to produce xylitol efficiently has been associated with its high NADPH-XR-specific activity, which favors higher xylose consumption rate and xylitol secretion (Cadete et al. 2016a; unpublished data).
2.2.4 Spathaspora
The genus Spathaspora was described by Nguyen et al. (2006) to accommodate the teleomorphic species Spathaspora passalidarum, which form specific allantoid asci, with a single, elongated, and curve-ended ascospore, very distinct from any other known yeast. The first species was isolated from the gut of the wood-boring beetle Odontotaenius disjunctus (Coleoptera: Passalidae), collected in Louisiana (USA). The vegetative cells are predominantly globose, formed by budding, and septate hyphae and pseudohyphae are present. In sexual reproduction, an allantoid ascus is formed without conjugation and contains the single ascospore surrounded by a persistent membrane (Nguyen et al. 2006; Kurtzman 2011b). Supported by phylogenetic analyses of D1/D2 LSU, the genus also included other related taxa as the anamorphic species Candida jeffriesii and Candida materiae (Nguyen et al. 2006; Barbosa et al. 2009). The phylogeny in Fig. 2.2 shows the placement of the Spathaspora and C. materiae strains in the same clade.
Other isolates of S. passalidarum were collected later from rooting wood samples (Cadete et al. 2012; Ren et al. 2014), wood-boring beetles, and galleries (Souza et al. 2017) or soil (Rodrussamee et al. 2018). The higher number of isolates obtained from decaying wood and soil/gallery samples suggests that the species might be more associated with this environment rather than the gut of beetles. However, the hypothesis of a symbioses yeast-insect cannot be discarded until further and extensive ecological studies are carried out (Nguyen et al. 2006; Cadete and Rosa 2018).
The mainly metabolic trait of S. passalidarum is its high capacity to ferment xylose. Until now, only few yeast strains have been described as natural xylose fermenters, among them are Scheffersomyces (Pichia) stipitis, Candida tenuis, and Pachysolen tannophilus (Slininger et al. 1982; Suh et al. 2003; Wohlbach et al. 2011a). Together with S. stipitis, S. passalidarum has the highest native capacity for xylose fermentation (Jeffries and Van Vleet 2009; Veras et al. 2017; Cadete and Rosa 2018). Batch cultivation reaches ethanol yields up to 0.48 g/g xylose, showing higher specific ethanol productivity under oxygen-limited condition than aerobic condition (Table 2.7) (Cadete et al. 2016a, b; Veras et al. 2017). In addition, S. passalidarum strains have the advantageous ability to co-utilize different substrates like glucose, xylose, and cellobiose and to produce higher ethanol yields on xylose than on glucose (Long et al. 2012; Su et al. 2015).
The high ethanol yield and productivity of S. passalidarum make clear the great potential of this strain for 2G bioethanol processes. However, like other microorganisms, S. passalidarum are inhibited by toxic compounds present in the lignocellulosic hydrolysates (Almeida et al. 2011; Su et al. 2018). Therefore, research efforts have focused on surpassing the toxicity of hydrolysate to achieve the practical use of this strain in biotechnological processes. Previous studies have shown that S. passalidarum was able to co-ferment glucose, xylose, and cellobiose mixtures in synthetic medium (ethanol yield of 0.42g/g) or hardwood hydrolysates without acetic acid or furaldehydes (ethanol yield of 0.34 g/g). Nevertheless, a significant inhibition and delay in the fermentation were observed when another kind of hydrolysate (AFEX) was used, which contains approximately 1.5 g/L of acetic acid (Long et al. 2012). Changing pretreatment process, a sugarcane bagasse hydrolysate with 43 g/L of glucose and 15 g/L of xylose, with no acetic acid, furfural, or hydroxymethylfurfural was obtained (Nakanishi et al. 2017). Employing this hydrolysate, the maximum yield achieved for fed-batch fermentation by S. passalidarum with cell recycle was 0.46 g/g, with a productivity of 0.81 g/L/h.
Evolutionary strategies have been successfully applied to develop S. passalidarum strains with improved traits. An evolved strain obtained by Morales et al. (2017) showed increased tolerance to acetic acid, achieving an ethanol yield of 0.48 g/g in the presence of 4.5 g/l acetic acid or 0.36 g/g in a medium containing 100% of non-detoxified Eucalyptus autohydrolysate. In another study, a S. passalidarum strain (E11) was selected after several rounds of batch adaptation, cell mating, and high-throughput screening, with traits that confer resistance to toxins found in hydrolysates and improved xylose fermentation abilities (Su et al. 2018).
The second species of the genus Spathaspora was isolated from rooting wood collected in the Brazilian Atlantic Rainforest and Cerrado biomes, and it was named as S. arborariae (Cadete et al. 2009). The species present similar asci and ascospore to the type strain S. passalidarum. Differently from this one, the new species can assimilate L-sorbose and produce both ethanol and xylitol as major products from xylose (Cadete et al. 2009, 2016b). Recently, other ten species were identified as belonging to Spathaspora genus. Cadete et al. (2013) described the species S. brasiliensis, S. roraimensis, S. suhii, and S. xylofermentans, which were mainly xylitol producers, albeit some of them are also able to produce ethanol from xylose. Wang et al. (2016) proposed the new species S. allomyrinae for a yeast isolated from the gut of beetles in China. Lopes et al. (2016) reported three novel strains S. girioi, S. gorwiae, and S. hagerdaliae, which are also isolated from rotting wood in Brazil. Finally, other two species were isolated from rotting wood samples in Brazil, S. boniae and S. piracicabensis, reported by Morais et al. (2017) and Varize et al. (2018).
Among the Spathaspora strains already characterized, S. passalidarum, S. arborariae, S. gorwiae, S. hagerdaliae, and S. piracicabensis are mainly ethanol producers from xylose, while other Spathaspora are mostly xylitol producers (Cadete et al. 2016b; Cadete and Rosa 2018; Varize et al. 2018). In addition to Candida tropicalis strains that are characterized as a great xylitol producers (de Albuquerque et al. 2014), Spathaspora species have shown an interesting potential for xylitol production, reaching yields in the range of 0.21–0.61 g/g xylose depending on fermentation medium and process conditions (Cadete et al. 2012, 2016b). Until now, the best xylitol production by Spathaspora strains on xylose fermentation medium was obtained for the S. roraimanensis UFMG-CM-Y477T (27.4 g/L xylitol, Y = 0.56), S. xylofermentans UFMG-CM-Y478T (24.4 g/L xylitol, Y= 0.51) (Cadete et al. 2012, 2016b), and for a not completely identified strain Spathaspora sp. JA1 (22.6 g/L xylitol, Y= 0.75) (Trichez et al. unpublished data). In the case of detoxified sugarcane hemicellulosic hydrolysate as feedstock, an uncharacterized Spathaspora UFMG-XMD-23.2 strain showed significant xylitol yield (17.1 g/L xylitol, Y = 0.61 g/g) (Cadete et al. 2012, 2016b).
Genomic information about native xylose-assimilating yeasts may contribute to understand xylose metabolism and be a source of genes to improve xylose fermentation. The genomic sequence of S. passalidarum was reported by Wohlbach et al. (2011a), and more recently other Spathaspora genomes have been deposited (Lobo et al. 2014a; Lopes et al. 2017; Morais et al. 2017; Varize et al. 2018) (Table 2.4).
Like other yeasts, the Spathaspora strains show the genes involved in the xylose oxido-reductase pathway XYL1, XYL2, and XYL3 that encodes for a xylose reductase (XR), a xylitol dehydrogenase (XDH), and a xylulokinase (XK), respectively (Wohlbach et al. 2011a; Lobo et al. 2014a; Cadete et al. 2016b; Lopes et al. 2017; Morais et al. 2017). Genome sequencing revealed that differently from other Spathaspora spp., S. passalidarum contains two copies of xylose reductase-encoding gene. The second copy (XYL1.2) encodes an NADH-preferred XR greatly expressed in oxygen-limited conditions, while XYL1.1 encodes a strictly NADPH-dependent enzyme (Wohlbach et al. 2011a; Cadete et al. 2016b; Cadete and Rosa 2018). Previous studies have suggested that the better balance between cofactor supply and demand through the XR and XDH leads to higher ethanol yields and efficient xylose fermentation by S. passalidarum cells (Hou 2012; Cadete et al. 2016b). In addition to S. passalidarum, apparently, S. arborariae, S. boniae, and S. gorwiae also have XR enzymes that use both cofactors NADH and NADPH, although showing a preference for the last one (Fig. 2.1) (Lopes et al. 2016; Cadete et al. 2016b; Morais et al. 2017). In turn, xylitol producers, as S. brasiliensis, S. roraimanensis, S. suhii, and S. xylofermentans, have genes encoding strictly NADPH-dependent XR enzymes (Cadete et al. 2016b). Thus, since all Spathaspora spp. characterized until now have shown XDH activities strictly NAD+-dependent, the imbalance of cofactors, in relation to the first steps of xylose metabolism, apparently favors the high xylitol yields observed in these strains.
In summary, Spathaspora species are able to use xylose and lignocellulosic hydrolysates in the fermentation process. S. passalidarum is one of the best ethanol producers from xylose. The high yields and productivities show the great potential of this yeast for 2G ethanol production. In addition, the capacity of some Spathaspora to produce xylitol opens the possibility of the employment of these yeasts in production processes for other bio-products. Bioprospection of novel yeasts, data from genome sequencing, metabolic engineering approaches, and process optimization may contribute to the development of yeast strains or better process conditions to an effective industrial application of these yeasts.
2.2.5 Kluyveromyces
The Kluyveromyes genus, located in the Saccharomycetes class, was created in 1956 by van der Walt in order to fit the yeast Kluyveromyces polysporus (currently a member of the Vanderwaltozyma genus). In the following years, it was noted that the type yeast K. polysporus presented some resemblances features to species previously accepted in the Saccharomyces genus like ascus deliquescence, ascospores, and robust fermentation (van der Walt 1965; Lachance 2007). That resulted in the transfer of important yeasts like S. fragilis, S. marxianus, and S. lactis to the Kluyveromyces genus (van der Walt 1965; Fonseca et al. 2008). Since then, many changes in the number of accepted species followed as taxonomic studies were being made. These changes kept happening until advances in genetic sequencing and multigene analysis tools enabled more accurate studies, decreasing the number of species for the current six members, K. aestuarii, K. dobzhanskii, K. lactis, K. marxianus, K. nonfermentans, and K. wickerhamii, distributing the species previously attributed to Kluyveromyces in the genera Kazachstania, Nakaseomyces, Tetrapisispora, Vanderwaltozyma, and Lachancea (Kurtzman and Robnett 2003; Kurtzman 2011b). The current type species is Kluyveromyces marxianus (Lachance 2007).
As described by Lachance (2011), generally, Kluyveromyces cells can be ovoid, ellipsoid, cylindrical, or, elongate. Its members can reproduce by asexual means, with multilateral budding, where pseudohyphae may be formed but never true hyphae. Sexual reproduction also occurs with or without the production of ascus preceding conjugation (Lachance 2011). They are considered to be thermotolerant yeasts, with the maximum growing temperature ranging from 35 to 52°C. With the exception of K. nonfermentans, all species are capable of fermenting glucose (Lane and Morrissey 2010). Genomic analysis of the Kluyveromyces genus representative members shows that genome sizes, in general, vary from 9.5 to 11 Mb, having in mind that other information like chromosome number and protein count are more available for K. marxianus and K. lactis (Table 2.4).
Among the six species present in the genus, K. marxianus and K. lactis are the ones largely used in biotechnological studies and industrial processes. A rare trait present in both species is lactose assimilation, enabled by the presence of genes LAC12, which encodes a permease responsible for lactose uptake, and LAC4, encoding for β-galactosidase, the enzyme responsible for hydrolyzing lactose molecules to glucose and galactose. In addition, the recurrent isolation of both yeasts from dairy products, like cheese, yogurt, and kefir, along with many years of secure use, has granted them the GRAS and QPS status (generally regarded as safe and qualified presumption of safety) (Morrissey et al. 2015). Moreover, more details about K. marxianus and K. lactis are presented, and the possible application of these yeasts on the biotechnological valorization of xylose is discussed.
2.2.5.1 Kluyveromyces marxianus
Despite not having many studies involving matters like physiology and metabolism, the type species Kluyveromyces marxianus is largely applied in industries (Lane and Morrissey 2010). Some features like the assimilation of inulin, by the activity of the INU1 gene; high thermotolerance (up to 52°C), one of the fastest growth rates among eukaryote organisms (Groeneveld et al. 2009); and its respiro-fermentative nature are differentials that make this yeast very attractive for use in industrial processes (Morrissey et al. 2015). For this reason, not only different applications for this species are being largely studied but also biotechnological tools are being developed in order to overcome bottlenecks related to this yeast and to establish it as a new synthetic biology platform (Cernak et al. 2018).
Industrial applications of K. marxianus are mainly related to protein expression, enzyme production, and ethanol production (Fonseca et al. 2008; Lane and Morrissey 2010). Having that in mind, among the most important commercial applications are the production of its native enzymes. One of them is inulinase, encoded by the previously mentioned INU1 gene. Inulinase (β-2,1-D-fructan fructanohydrolase) targets the β-2,1 linkage of inulin, a polyfructan consisting of linear β-2,1-linked fructose, and hydrolyzes it into fructose. Thus, this enzyme can be used for the production of syrups with high fructose content. Another possible application of K. marxianus is the production of ethanol using whey permeate and Jerusalem artichokes as substrate (Yang et al. 2015). As mentioned before, tolerance to high temperatures is an interesting trait, once it allows this yeast to be utilized together with enzymes responsible for liberating monomers from the lignocellulosic feedstocks during simultaneous saccharification and fermentation (SSF). Another use of K. marxianus that has been recently reviewed and drawn attention to is the production of high alcohols, such as 2-phenylethanol (Morrissey et al. 2015). Characterized by having a strong rose scent, 2-PE can be used in cosmetics and ffragrance applications, and as a flavor additive in the food industry (Morrissey et al. 2015). The biological production, via the Ehrlich pathway, is an important alternative for the chemical route production, as the latter involves the use of benzene, which can be hazardous when applied in food and cosmetics.
The metabolism of xylose is present in K. marxianus once it possesses XYL1, XYL2, and XKS1 genes. The transport of pentoses to the intracellular environment is also possible in this yeast, but its kinetic parameters depend on substrates and conditions applied. In 2003, Stambuk et al. demonstrated that not only the presence of glucose in the medium is responsible for inhibiting the uptake of xylose but also that under anaerobic conditions, the transport presents low affinity and high capacity, while aerobic conditions induce the production of the high-affinity xylose transporter system (Stambuk et al. 2003). Although the necessary genes for xylose conversion to ethanol are present, unbalanced redox reactions in the xylose reductase pathway (XR and XDH) usually result in low ethanol yield and formation of undesired co-products under low-oxygen conditions (Varela et al. 2017). Therefore, metabolic engineering strategies have been employed to optimize xylose conversion through XR-XDH or XI pathways. One of these studies has constructed the strain K. marxianus YZJ088 by the combination of XR from N. crassa and XDH from S. stipitis, achieving high productivity (2.49 g/L/h) and yield (0.38 g/g) of ethanol at high temperatures with xylose as the only carbon source (Zhang et al. 2015). It was also reported that the recombinant K. marxianus YZJ119 strain was able to produce 44.58 g/L of ethanol and 32.03 g/L of xylitol from detoxified pre-treated corncob lignocellulosic hydrolysate (Table 2.8) (Zhang et al. 2016).
Xylitol, a sugar alcohol considered a very promising bio-based chemical, is also one of the possible products originated from xylose utilization by K. marxianus. Many genetic engineering strategies had also been applied for this end. Among them, the overexpression of heterologous transporters allowed to generate K. marxianus YZJ074 strain, which produces 101.30 xylitol with a productivity of 2.81 g/L/h even at temperatures as high as 45°C (Table 2.8) (Zhang et al. 2014).
2.2.5.2 Kluyveromyces lactis
K. lactis is considered one of the most studied yeasts and a model organism among the “non-conventional” group, making it unavoidable to compare it with S. cerevisiae. A major difference between them is the preference for respiration in K. lactis over sugar fermentation in S. cerevisiae (Dias et al. 2012). The low-cost production of bovine chymosin by K. lactis is considered to be a milestone for the expression of high-eukaryote enzymes in microorganisms (van Ooyen et al. 2006). By 2006 more than 40 proteins were already produced by K. lactis, and this number increased to around 100 proteins in 2016 (van Ooyen et al. 2006; Spohner et al. 2016). The applications of this yeast and its products also have a wide range, like the production of β-galactosidase and chymosin in the food industry and the production of human interleukin 1-β and interferon-α for the pharmaceutical industry (Rodicio and Heinisch 2013; Spohner et al. 2016).
Not many studies were made with K. lactis to analyze its potential of using lignocellulosic biomass in bioprocesses. However, it was already observed that this yeast is capable of metabolizing xylose and obtaining products like d-xylonate and xylitol (Nygård et al. 2011). In this work, a xylose dehydrogenase gene from Trichoderma reesei was expressed, and the resulting strain produced xylonic acid with higher productivity and yield than a S. cerevisiae strain expressing the same gene (Nygård et al. 2011). This study can indicate not only possible advantages of this yeast over S. cerevisiae, such as better xylose transport and tolerance to acid accumulation, but also points out that these traits represent the potential of the application K. lactis in studies involving xylose and lignocellulosic biomass (Nygård et al. 2011).
Although there are not many studies regarding xylose utilization involving the remaining species of this genus, interesting findings involving them can be observed in recent years. As examples, we can cite K. aestuarii being depicted as a quality bioindicator (Araujo and Hagler 2011) and the application of a toxin produced by K. wickerhamii for controlling spoilage yeasts in winemaking (Comitini and Ciani 2011).
2.2.6 Komagataella
Komagataella belongs to Saccharomycetales order and Saccharomycetaceae family. The genus was proposed by Yamada et al (1994) when the partial sequences of 18S and 26S rRNAs subunits of methanol-assimilating yeasts were phylogenetically analyzed. The study exhibited that partial sequences of Pichia pastoris were different when compared to other strains. Therefore, it was proposed the new genus Komagataella for P. pastoris that was phylogenetically distinct from the other genera examined. The proposal was not initially accepted because just a few numbers of strains were used in the comparison. However, Kurtzman and Robnett (1998) analyzed the divergence of the D1/D2 LSU rRNA gene sequences among approximately 500 species of Ascomycetous yeasts. When they compared D1/D2 sequences, the divergences found in P. pastoris were confirmed. Subsequently, multigenic sequence analysis supported the phylogenetic difference position of Pichia pastoris (renamed to Komagataella pastoris). Thus, the genus became accepted and currently has seven species.
The majority of Komagataella species was isolated from habitats such as tree exudates and decomposing woods, sources capable of supplying methanol for the growth of these yeasts. The type species of K. pastoris was first isolated in 1919 from a chestnut tree in France by Guilliermond. Furthermore, most strains of K. pastoris and K. pseudopastoris were found in exudates and decomposing wood in Hungary. The species K. phaffii, K. populi, K. ulmi, K. kurtzmanii, and K. mondaviorum were isolated from exudates of different trees at the USA in the states of California, Arizona, and Illinois. The growth and isolation of these species were mostly done in YM agar medium at 25 °C, as described by Yarrow (1998) and Kurtzman (2011a).
The colonies of Komagataella sp. have spherical to ovoid shapes; the coloration can be white to cream and may have moderately lobate margins. In the asexual reproduction, haploid cells divide by multilateral budding and do not present pseudohyphae or true hyphae (Kurtzman 2011b). For sexual reproduction, the formation of ascospore is hat-shaped which may range from 1 to 4 and can be conjugated or not. Also, known species are homothallic. An important characteristic of the genus is the ability to grow at high cell densities and ferment glucose. Because those species are methylotrophic, they can use methanol as the only carbon source (Kurtzman 2011b). The nitrate cannot be used as the only source for nitrogen during growth. The seven species have very similar phenotypic and fermentative characteristics. Therefore, they cannot be differentiated with tests that are usually used for yeast taxonomy. In this way, multigenic sequence analysis, such as D1/D2 LSU rRNA, ITS, EF-1α, and other gene regions, are required for species identification of the genus (Kurtzman 2011b).
Komagataella species play a major rule in biotechnological applications. These yeasts provide an efficient host system for heterologous protein production, having both industrial and pharmaceutical importance. In addition, metabolic engineering has become an interesting application for these yeasts to produce different chemicals. On account of that, the genome sequencing of strains K. pastoris DMSZ and K. phaffii GS115 were performed (Mattanovich et al. 2009; De Schutter et al. 2009). The approximate size of the genome of both species is 9.4 Mb, containing 4 chromosomes and more than 5000 genes (Table 2.4). The information provided by genome data highlights the advantages of Komagataella sp. as a heterologous expression system, and it is essential for metabolic engineering studies.
Initially, the Phillips Petroleum Company used Komagataella (Pichia) pastoris as single-cell protein (SCP) because of its ability to assimilate methanol as a carbon source and ability to grow in high cell densities. The SCP was marketed for high protein animal additive. Posteriorly, the oil crisis in the 1970s increased the methanol price, and the SCP system was no longer favorable. In the following years, the genetic manipulation of K. pastoris strains and the development of protocols, vectors, and isolation of the AOX gene led to the creation of the heterologous expression system known today. Invitrogen Company developed the “Pichia Expression Kit,” which is widely commercialized and used in laboratories for heterologous protein expression. The strains used in the expression kit were phenotypically identified as K. (Pichia) pastoris. However, multigene sequence analysis determined that strains used in the Invitrogen Expression Kit were from K. phaffii (Kurtzman 2009; Ahmad et al. 2014).
The success of the Komagataella sp. expression system is due to established genetic manipulation techniques, with recombinant proteins that can be expressed both extracellular and intracellular. In addition, the expressed proteins may undergo post-translational eukaryotic modifications such as glycosylation, disulfide bonds, and proteolytic processing. The vectors integrated into its genome facilitate the genetic stability of the recombinant elements even in continuous and large-scale fermentation processes. Furthermore, the species have the “generally recognized as safe” (GRAS), status, and for this reason, more than 500 biopharmaceutical proteins and a great number of other recombinant enzymes have been produced since 2009 (Yang and Zhang 2018). Many studies have been reporting the expression of recombinant proteins that can be useful in food, feed, detergent, clothing industries, and more (Table 2.9).
There is some research showing the potential of some Komagataella strains to use xylose and glucose from lignocellulosic biomass to achieve the production of value-added products. In this context, K. pastoris DSM 70877 was cultivated on mixtures of glucose and xylose to produce chitin-glucan complex (CGC) (Araújo et al. 2017). This biopolymer is a component of the inner cell wall of yeast and has several applications in food, cosmetics, and pharmaceutical industries. When cultivated in the glucose/xylose mixture, the CGC and xylitol were produced. The xylose started to be assimilated after the glucose depletion, but no cell growth was detected. However, the production of xylitol reached a final concentration of 7.64 g/L. Therefore, this work presents great potential for co-production of bioproducts using the lignocellulosic biomass.
According to Li et al. (2015), K. phaffii GS115 strain is able to use the xylose at low rates for specific growth (0.0075 h−1). In order to enhance the efficiency in xylose assimilation, the host strain GS115 was engineered with XI (xylose isomerase). After the introduction of XI pathway, an evolutionary engineering strategy was also applied, resulting in a strain (GS-XISB50) capable to assimilate xylose at higher rates (specific growth: 0.0193 h−1). Also, this engineered strain was used for heterologous expression of β-mannanase enzyme produced in a medium containing xylose. Finally, this engineered strain is able to grow in xylose medium producing enzymes with industrial importance.
A few numbers of microorganisms have been engineered to produce xylonic acid by the overexpression of xylose dehydrogenases either from Caulobacter crescentus or Trichoderma reesei. K. phaffii is a promising yeast for production of xylonic acid because of its tolerance to low pHs and ability to grow at very high cell densities. The xylonic acid can be used as a precursor for other chemicals and has several industrial applications such as building blocks of polymers. Recently, new putative xylose dehydrogenase (XDH) genes from bacteria and fungi were identified by phylogenetic analysis. Then, three of those were chosen for genetic engineering of K. phaffii X33. Recombinant strains expressing each gene were evaluated by their ability to produce xylonic acid, and the best candidate genes were chosen for further analysis. Strains were able to produce up to 36.2 g/L of xylonic acid from 40 g/L xylose, which accounts for a yield of 0.95 g/g, under the best fermentative evaluated conditions (patent under subscription by the number BR102018001359-9_870180005782). Finally, strain’s capability to produce xylonic acid on sugarcane biomass hydrolysate was demonstrated.
K. phaffii was also engineered for production of acid lactic. This acid has applications such as in food, pharmaceutical, and textile industries. Also, it is a monomer of a biodegradable plastic called poly-lactic acid (PCA). In that way, de Lima et al. (2016) genetically modified strains of K. phaffii (X-33 and GS115) that were capable to produce lactic acid using glycerol as the only carbon source at limited oxygen conditions. They used a bovine lactate dehydrogenase (LDH) and lactate transporter to make the production of this acid possible. The best strain (named GLS) obtained by the introduction of both genes reached a yield of 0.7 g/g that was really close to the maximum theoretical yield (1 g/g). However, the production of lactic acid from xylose using recombinant K. phaffii strains was not reported.
As a conclusion, Komagataella species can be explored in a range of biotechnological applications (Mattanovich et al. 2009). The main use for those species is the heterologous expression of recombinant proteins that have important uses in several industry sectors. Also, with the genome sequencing, new studies are showing potential as a platform of metabolic engineering and synthetic biology. In that way, new approaches and methodologies are being explored to achieve future developments of heterologous expression, metabolic engineering, and synthetic biology of these yeasts.
2.2.7 Yarrowia
Yarrowia (van der Walt and von Arx 1980), a fungal genus of the family Diposdascaceae, constructed from the previously ascosporic state of Candida lipolytica, is composed of 13 species. This genus was, prior DNA comparisons, considered monotypic, containing the single species Yarrowia lipolytica (Dujon et al. 2004; Kurtzman 2005; Kurtzman et al. 2010). Some descriptions of new Yarrowia species clade were recognized in the last years based on D1/D2 sequence comparisons (Nagy et al. 2013). For example, Groenewald and Smith (2013) have described two novel species, Y. deformans and Y. yakushimensi, Nagy et al. (2013) described Y. divulgata, and Crous et al. (2017) described another species called Y. parophonii. These new species are phenotypically indistinguishable from Y. lipolytica. Kurtzman (2005) and Crous et al. (2017) based on phylogenetic analysis of nucleotide sequences from domains D1/D2 26S rDNA have proposed new relocation in the genus Yarrowia, such as Y. galli (basionym C. galli) and Y. oslonensis (basionym C. oslonensis). The discovery of new species is very recent, so Y. lipolytica is still considered the most important yeast of the Yarrowia genus. The other species have not yet been widely studied, and absence of growth is recorded for xylose (Kurtzman 2005; Nagy et al. 2013, 2014; Crous et al. 2017). Therefore, only Y. lipolytica description will be covered in this section. But it is important to note that some yeast isolates identified as Y. lipolytica based on phenotypic characteristics and used in industries may really be a member of other Yarrowia clade yet incorrectly identified (Nagy 2015).
Yarrowia lipolytica is strictly an aerobic nonpathogenic ascomycetous yeast. This yeast shares some properties common to filamentous fungi being distantly associated with other yeasts (Kurtzman and Robnett 1998; Dujon et al. 2004; Gonçalves et al. 2014). It was originally isolated from lipid-rich materials as rancid butter and first identified, in the 1960s, as Candida lipolytica and reclassified as Endomycopsis lipolytica, Saccharomycopsis lipolytica, and finally Yarrowia lipolytica (Liu et al. 2015). Strains of this species are usually isolated as contaminant or spoilage organisms from lipid-rich or protein-rich environments such as dairy products, meat, poultry, and olive oil (Najjar et al. 2011; Nagy et al. 2013; Liu et al. 2015; Li and Alper 2016). Y. lipolytica has a haplodiplontic cycle. The most of natural isolated strains are haploid. A diploid strain has little occurrence, but both states are stable in laboratory conditions (Dujon et al. 2004; Liu et al. 2015). The process of genetic regulation that leads to dimorphism is still unknown (Liu et al. 2015).
Y. lipolytica is classified as oleaginous yeast and has the ability to assimilate different carbon sources that can be soluble in water, such as glucose, glycerol, alcohols, and acetate, or insoluble in water, such as fatty acids, triacylglycerols, and alkanes. Besides, it is tolerant to environmental stress like presence of salt, low temperatures, and acidic and alkaline pH (Gonçalves et al. 2014; Liu et al. 2015). These characteristics would allow the use of low-cost raw materials, such as wastewaters, industrial fat, glycerol co-product of biodiesel production, and molasses. Y. lipolytica is also exceptionally resistant to inhibitors associated with lignocellulosic biomass pretreatments (Ryu and Trinh 2017).
Thus, Y. lipolytica has multiple biotechnological and industrial applications such as the production of citric, isocitric, α-ketoglutaric and succinic acids, biomass to be used as single-cell protein, biosurfactants, and flavoring lactones to be used in the food industry (Groenewald and Smith 2013; Nagy et al. 2013). It can also secrete enzymes with functional applications, like lipases, acid or alkaline proteases, and RNase (Kurtzman et al. 2010; Najjar et al. 2011). Currently, one of the main interests in Y. lipolytica is its ability to produce and store lipids (Fig. 2.1) (up to 36% of its dry cell weight), which can be used for biodiesel production (Abghari and Chen 2014).
There have been inconsistent reports regarding xylose utilization by Y. lipolytica strains (Ledesma-Amaro et al. 2016; Spagnuolo et al. 2018). Some studies claiming native xylose metabolism demonstrated that xylose can be converted into lipids when cells of Y. lipolytica Po1g are transferred in the stationary phase into high xylose concentrations. Even in this condition, xylose was not a primary source for biomass production (Blazeck et al. 2014). In addition, Tsigie et al. (2011) showed that the maximum growth of Y. lipolytica Po1g (derivative of the wild-type strain W29 with a series of genetic modification) (Yeastern Biotech Co. Ltd.) in a medium containing xylose as only carbon source is 9.89 g/L with a short lag period and without adaptation time and have produced 6.68 g/L of lipids using sugarcane bagasse hydrolysate as a carbon source. On the other hand, the majority of studies suggests that Y. lipolytica is unable to grow with xylose as a primary carbon source without subjecting it to adaptation or starvation periods (Kurtzman 2011b; Stephanopoulos 2013; Zhao et al. 2015). These reports conflicted even among identical strains.
The xylose pathway in Y. lipolytica was not proven to be functional. Even though this yeast has putative genes coding for XR, XDH, and pentose transport genes, XK was the only enzyme with activity in this pathway (Ryu et al. 2016; Li and Alper 2016; Ryu and Trinh 2017). Then, in order to improve xylose catabolic phenotype in Y. lipolytica, pathway engineering is necessary. Overexpression of XR and XDH of S. stipitis in Y. lipolytica allowed moderate growth in xylose. When cultured in co-fermentation of 20 g/L glycerol and 80 g/L xylose, the strain produced 18 g/L of biomass, 7.64 g/L of lipids, and 9.13 g/L of citrate (Stephanopoulos 2013; Li and Alper 2016). Li and Alper (2016), introducing heterologous XR and XDH genes from S. stipitis, established a Y. lipolytica strain that is able to use xylose as sole carbon source and produce over 15 g/L of intracellular lipids, with a productivity of 0.19 g/L.h. Recently, Niehus et al. (2018) showed that overexpression of native XDH, XR, and XK genes allowed Y. lipolytica to grow on xylose as a sole carbon source and produced 16.5 g/L of lipids in high yield (3.44 g/g sugars) on a non-detoxified agave bagasse hydrolysate.
The activation of the native xylose pathway in Y. lipolytica ATCC MYA-2613 could be achieved by adaptation of the strain to grow on xylose as sole carbon source through serial culture transfers (Ryu et al. 2016). In the first transfer, the cell growth was very low and started only after 3 days. After 15 generations, Y. lipolytica grew in xylose with a specific growth rate of 0.10 h-1 and consumed 2.23 g/L xylose, producing 0.18 g/L xylitol, in 72 h. By transcriptomic analysis of the adapted strain, it was shown that Y. lipolytica has 16 putative xylose transporters and xylose-degrading metabolic enzymes (XR, XDH, and XK) required for xylose assimilation. In comparison with growth on glucose, Y. lipolytica reached a much lower OD, producing a high yield of xylitol, and did not completely consume xylose. This phenotype suggests that xylose consumption is not efficient because the XDH step was limiting. In addition, it was demonstrated that XDH is transcriptionally repressed by glucose. Ryu and Trinh (2017) have reported that overexpression of pentose transporters YALI0C04730p and YALI0B00396p and rate-limiting D-xylitol dehydrogenases (XDH) allowed activation of the dormant pentose metabolism of Y. lipolytica ATCC MYA-2613 and improved xylose assimilation approximately 50% in comparison with the parental strain. However, the improved strain grows on xylose ten times slower than on glucose.
The overexpression of XR and XDH of S. stipitis is necessary but not sufficient to permit Y. lipolytica’s growth on xylose. Thus, to improve oils and citric acid production from lignocellulosic materials by Y. lipolytica, an additional overexpression of the endogenous xylulokinase (XK) was necessary (Ledesma-Amaro et al. 2016). The recombinant strain was able to produce high titers of lipids, up to 20 g/L, on a medium containing xylose as sole carbon source and 50 g/L of lipids on xylose medium co-fed with glycerol using high xylose/nitrogen ratio. When lower xylose/nitrogen ratio and higher pH were used, citric acid was produced up to 80 g/L.
It can be observed that the mechanisms of many enzymes involved in different biochemical reactions to xylose assimilation remain unclear in Y. lipolytica (Liu et al. 2015; Spagnuolo et al. 2018). However, the fully sequenced genome, physiological capacity, and particular genetic advantages are what make Y. lipolytica a promising platform to produce added-value chemicals and biofuels (Table 2.10) (Dujon et al. 2004; Stephanopoulos 2013; Li and Alper 2016). Among the advantages of Y. lipolytica, the following can be mentioned: (1) protein is secreted primarily by the co-transcription pathway; (2) it has a high secretion capacity and low glycosylation modification; (3) it is a non-pathogenic yeast; (4) versatility of metabolites is produced; (5) it is robust in culture; (6) it is an obligate aerobe; (7) it is able to grownon a variety of substrates; (8) simple alterations in the fermentative process can modulate metabolite production (Gonçalves et al. 2014; Liu et al. 2015; Spagnuolo et al. 2018).
2.2.8 Ogataea
First, the genus Hansenula included species with ascopores in pellicle shape, hat shape, or saturn shape. After analyzing the yeast with saturn form, it was transferred to the Williopsis genus (Yamada et al. 1994). Nuclear DNA analyses revealed that 75% of the Hansenula minuta DNA was related to Pichia lindneri, the Pichia-type species. Kurtzman et al. (2011b) concluded with these analyses that the nitrate assimilation ability was not enough to separate the genera Hansenula and Pichia. After the analyses, all ascopores species with hat shape and nitrate assimilation ability were classified as Pichia (previously classified as Hansenula) (Yamada et al. 1994).
The results of 18s RNA analyses of P. membraefariens and P. anomala showed that the two species were phylogenetically different. Therefore, was proposed the creation of three new genera: Ogataea, Kuraishia, and Kakazawea (Yamada et al. 1994). The genus Ogataea was proposed in 1994 by Yamada, which was composed of yeasts previously identified as Pichia and Hansenula. At the beginning, five species and two varieties were proposed to the Ogataea genus: Ogataea clucozyma, Ogataea minuta minuta variatie, Ogataea minuta nonfermentans variatie, Ogataea philodendra, Ogataea polymorpha, and Ogataea henricie (Yamada et al. 1994). In the last edition of the book The Yeasts: A Taxonomic Study, 31 species of Ogataea were accepted to the genus (Kurtzman 2011b).
The genus belongs to Saccharomycetables family. Colonies are butyroid to mucoid. The cells can be globose, ellipsoid, ovoid, or cylindrical. In the asexual reproduction, cell division is by multilateral budding on a narrow base, and budded cells are spherical to ellipsoidal. True hyphas are not formed, but pseudohyphae if formed consist of few elongated T cells (Kurtzman 2011a).
Among the several species of Ogatae, O. methanolica, O. minuta, O. thermomethanolica, and O. polymorpha are being studied and applied in biotechnology processes. The first three species are employed mainly as heterologous expression systems (Kuroda et al. 2008; Tsai and Huang 2008; Puseenam et al. 2018).
After the yeast Kluveromyces marxianus, Ogataea polymorpha is the second yeast thermotolerant that can grow in temperature of 50o C, an advantage of using simultaneous saccharification and fermentation (SSF). O. polymorpha can grow using glucose, cellobiose, and xylose as carbon source; the yeast can also produce ethanol using glycerol. The ethanol yield and productivity of O. polymorpha using xylose are very low. But it could be increased raising the fermentation temperature (from 30oC to 37oC) in a simultaneous saccharification and fermentation (SSF) process (Ryabova et al. 2003) and through genetic engineering techniques such a riboflavin-deficient mutant under suboptimal supply with flavins. The flavin limitation apparently makes the pyruvate be redistributed via a flavin-independent pathway to ethanol production (Ryabova et al. 2003).
In a different approach, ethanol yield and productivity of O. polymorpha were improved by engineering the enzyme xylose reductase (XR). Firstly, the native XR was engineered to reduce affinity toward NADPH. After genes coding for modified XR, native XDH and XK were overexpressed in strain CBS4732. Then, the strain CBS4732 was modified to be unable to utilize ethanol as a carbon source, and the gene PDC1, coding for pyruvate decarboxylase (PDC) was cloned and overexpressed. The strain resulting was named 2EthOH-. Finally the strain 2EthOH- was able to produce up to 10 g of ethanol per liter at 45 oC using xylose as carbon source (Kurylenko et al. 2014).
2.3 Conclusions
Production of fuels and chemicals through environmentally friendly and cost-effective processes has been proposed and evaluated based on the utilization of sugars present in the lignocellulosic biomass as raw material and yeast as the catalyst. As yeasts promptly assimilate hexose sugars (glucose), much effort has been employed to obtain strains with high efficiency in pentose (xylose) assimilation and conversion. In this sense, due to the diversity and metabolic potential, several non-conventional yeasts species were isolated, identified, and physiologically and genetically characterized in the last years.
Bioprospecting of novel species and increasing the knowledge about non-conventional yeast genome and physiology opened new opportunities to explore the available biodiversity as well as identify new industrially relevant features. In addition, metabolic and evolutionary engineering strategies allowed further improvement and diversification of the bioconversion efficiency of xylose-containing hydrolysates to products of interest.
The non-conventional yeasts can be divided into two groups, the first with species naturally capable of efficiently metabolize xylose, belonging to Scheffersomyces, Meyerozyma, Candida, and Spathaspora. The second group, with species that have limited capacity to naturally metabolize xylose, but efficient strains have mainly been obtained mainly by genetic engineering strategies, belonging to Komagataella, Ogatae, and Yarrowia. Yeasts from Komagataella, Ogatae, and Kluyveromyces have been mostly used for the production of heterologous proteins of industrial application, in the pharmaceutical, food, feeding, detergents, and clothing industries. But recent developments allowed production of chemicals, such as xylonic acid, ethanol, and xylitol, with K. phaffii, Ogatae, and Kluyvermomyces. S. passalidarum and S. stipitis show great potential for production of ethanol 2G due to their high efficiency to ferment xylose, whereas other species as Spathaspora, Scheffersomyces, Candida, and Meyerozyma shows great potential for xylitol production. Finally, Yarrowia strains have been mostly employed for the production of lipids and organic acids.
In conclusion, robust strains for several applications have been obtained and evaluated in the most varied conditions. Even if this field has space to explore the improvement of strain’s performance before industrial use, new products based on non-conventional yeast processes should be in the market in the coming years.
References
Abbott DA, Zelle RM, Pronk JT, van Maris AJA (2009) Metabolic engineering of Saccharomyces cerevisiae for production of carboxylic acids: current status and challenges. FEMS Yeast Res 9:1123–1136. https://doi.org/10.1111/j.1567-1364.2009.00537.x
Abdulrachman D, Thongkred P, Kocharin K, Nakpathom M, Somboon B, Narumol N, Champreda V, Eurwilaichitr L, Suwanto A, Nimchua T, Chantasingh D (2017) Heterologous expression of Aspergillus aculeatus endo-polygalacturonase in Pichia pastoris by high cell density fermentation and its application in textile scouring. BMC Biotechnol 17. https://doi.org/10.1186/s12896-017-0334-9
Abghari A, Chen S (2014) Yarrowia lipolytica as an oleaginous cell factory platform for production of fatty acid-based biofuel and bioproducts. Front Energy Res 2. https://doi.org/10.3389/fenrg.2014.00021
Agbogbo FK, Coward-Kelly G (2008) Cellulosic ethanol production using the naturally occurring xylose-fermenting yeast, Pichia stipitis. Biotechnol Lett 30:1515–1524. https://doi.org/10.1007/s10529-008-9728-z
Ahmad M, Hirz M, Pichler H, Schwab H (2014) Protein expression in Pichia pastoris: recent achievements and perspectives for heterologous protein production. Appl Microbiol Biotechnol 98:5301–5317. https://doi.org/10.1007/s00253-014-5732-5
Akinterinwa O, Khankal R, Cirino PC (2008) Metabolic engineering for bioproduction of sugar alcohols. Curr Opin Biotechnol 19:461–467. https://doi.org/10.1016/j.copbio.2008.08.002
Almeida JRM, Runquist D, Sànchez Nogué V, Lidén G, Gorwa-Grauslund MF (2011) Stress-related challenges in pentose fermentation to ethanol by the yeast Saccharomyces cerevisiae. Biotechnol J 6:286–299. https://doi.org/10.1002/biot.201000301
Araujo FV, Hagler AN (2011) Kluyveromyces aestuarii, a potential environmental quality indicator yeast for mangroves in the State of Rio de Janeiro, Brazil. Braz J Microbiol 42:954–958
Araújo D, Freitas F, Sevrin C, Grandfils C, Reis MAM (2017) Co-production of chitin-glucan complex and xylitol by Komagataella pastoris using glucose and xylose mixtures as carbon source. Carbohydr Polym 166:24–30. https://doi.org/10.1016/j.carbpol.2017.02.088
Artifon W, Bonatto C, Bordin ER, Bazoti SF, Dervanoski A, Alves SL, Treichel H (2018) Bioethanol Production From Hydrolyzed Lignocellulosic After Detoxification Via Adsorption With Activated Carbon and Dried Air Stripping. Front Bioeng Biotechnol 6. https://doi.org/10.3389/fbioe.2018.00107
Bajwa PK, Phaenark C, Grant N, Zhang X, Paice M, Martin VJJ, Trevors JT, Lee H (2011) Ethanol production from selected lignocellulosic hydrolysates by genome shuffled strains of Scheffersomyces stipitis. Bioresour Technol 102:9965–9969. https://doi.org/10.1016/j.biortech.2011.08.027
Ballesteros I, Negro MJ, Oliva JM, Cabañas A, Manzanares P, Ballesteros M (2006) Ethanol production from steam-explosion pretreated wheat straw. In: McMillan JD, Adney WS, Mielenz JR, Klasson KT (eds) Twenty-seventh symposium on biotechnology for fuels and chemicals. Humana Press, Totowa, pp 496–508
Barbosa AC, Cadete RM, Gomes FCO, Lachance M-A, Rosa CA (2009) Candida materiae sp. nov., a yeast species isolated from rotting wood in the Atlantic Rain Forest. Int J Syst Evol Microbiol 59:2104–2106. https://doi.org/10.1099/ijs.0.009175-0
Beopoulos A, Cescut J, Haddouche R, Uribelarrea J-L, Molina-Jouve C, Nicaud J-M (2009) Yarrowia lipolytica as a model for bio-oil production. Prog Lipid Res 48:375–387. https://doi.org/10.1016/j.plipres.2009.08.005
Bier MCJ, Maranho LT, Azevedo JAM, Junios LS d S (2007) Crescimento e consumo de Xilose de Candida guilliermondii na fermentação submersa utilizando-se bagaço de cana-de-açúcar. Evidência 7:119–130
Blazeck J, Hill A, Liu L, Knight R, Miller J, Pan A, Otoupal P, Alper HS (2014) Harnessing Yarrowia lipolytica lipogenesis to create a platform for lipid and biofuel production. Nat Commun 5. https://doi.org/10.1038/ncomms4131
Bozell JJ, Petersen GR (2010) Technology development for the production of biobased products from biorefinery carbohydrates—the US Department of Energy’s “Top 10” revisited. Green Chem 12:539. https://doi.org/10.1039/b922014c
Bruinenberg PM, Bot PHM, Dijken JP, Scheffers WA (1983) The role of redox balances in the anaerobic fermentation of xylose by yeasts. Eur J Appl Microbiol Biotechnol 18:287–292. https://doi.org/10.1007/BF00500493
Butler G, Rasmussen MD, Lin MF, Santos MAS, Sakthikumar S, Munro CA, Rheinbay E, Grabherr M, Forche A, Reedy JL, Agrafioti I, Arnaud MB, Bates S, Brown AJP, Brunke S, Costanzo MC, Fitzpatrick DA, de Groot PWJ, Harris D, Hoyer LL, Hube B, Klis FM, Kodira C, Lennard N, Logue ME, Martin R, Neiman AM, Nikolaou E, Quail MA, Quinn J, Santos MC, Schmitzberger FF, Sherlock G, Shah P, Silverstein KAT, Skrzypek MS, Soll D, Staggs R, Stansfield I, Stumpf MPH, Sudbery PE, Srikantha T, Zeng Q, Berman J, Berriman M, Heitman J, Gow NAR, Lorenz MC, Birren BW, Kellis M, Cuomo CA (2009) Evolution of pathogenicity and sexual reproduction in eight Candida genomes. Nature 459:657–662. https://doi.org/10.1038/nature08064
Cadete RM, Rosa CA (2018) The yeasts of the genus Spathaspora : potential candidates for second-generation biofuel production: Spathaspora yeasts as candidates for 2G bioethanol production. Yeast 35:191–199. https://doi.org/10.1002/yea.3279
Cadete RM, Santos RO, Melo MA, Mouro A, DL Gça, Stambuk BU, Gomes FCO, Lachance M-A, Rosa CA (2009) Spathaspora arborariae sp. nov., a d-xylose-fermenting yeast species isolated from rotting wood in Brazil. FEMS Yeast Res 9:1338–1342. https://doi.org/10.1111/j.1567-1364.2009.00582.x
Cadete RM, Melo MA, Dussán KJ, Rodrigues RCLB, Silva SS, Zilli JE, Vital MJS, Gomes FCO, Lachance M-A, Rosa CA (2012) Diversity and Physiological Characterization of D-Xylose-Fermenting Yeasts Isolated from the Brazilian Amazonian Forest. PLoS One 7:e43135. https://doi.org/10.1371/journal.pone.0043135
Cadete RM, Melo MA, Zilli JE, Vital MJS, Mouro A, Prompt AH, Gomes FCO, Stambuk BU, Lachance M-A, Rosa CA (2013) Spathaspora brasiliensis sp. nov., Spathaspora suhii sp. nov., Spathaspora roraimanensis sp. nov. and Spathaspora xylofermentans sp. nov., four novel d-xylose-fermenting yeast species from Brazilian Amazonian forest. Antonie Van Leeuwenhoek 103:421–431. https://doi.org/10.1007/s10482-012-9822-z
Cadete RM, de las Heras AM, Sandström AG, Ferreira C, Gírio F, Gorwa-Grauslund M-F, Rosa CA, Fonseca C (2016a) Exploring xylose metabolism in Spathaspora species: XYL1.2 from Spathaspora passalidarum as the key for efficient anaerobic xylose fermentation in metabolic engineered Saccharomyces cerevisiae. Biotechnol Biofuels 9. https://doi.org/10.1186/s13068-016-0570-6
Cadete RM, Melo-Cheab MA, Viana AL, Oliveira ES, Fonseca C, Rosa CA (2016b) The yeast Scheffersomyces amazonensis is an efficient xylitol producer. World J Microbiol Biotechnol 32. https://doi.org/10.1007/s11274-016-2166-5
Canilha L, Milagres AMF, Silva SS, Almeida e Silva JB, Felipe MGA, Roche GJM, Ferraz A, Carvalho W (2010) Sacarificação da Biomassa Lignocelulósica através de pré-hidrólise ácida seguida por hidrólise enzimática: uma estratégia de “desconstrução” da fibra vegetal. Revista Analytica 44
Cernak P, Estrela R, Poddar S, Skerker JM, Cheng Y-F, Carlson AK, Chen B, Glynn VM, Furlan M, Ryan OW, Donnelly MK, Arkin AP, Taylor JW, Cate JHD (2018) Engineering Kluyveromyces marxianus as a robust synthetic biology platform host. mBio 9. https://doi.org/10.1128/mBio.01410-18
Chakravorty M, Veiga LA, Bacila M, Horecker BL (1962) Pentose Metabolism in Candida. II. The diphosphopyridine nucleotide-specific polyol dehydrogenase of candida utilis. J Biol Chem 237:1014–1020
Chandel AK, Singh OV, Rao LV, Chandrasekhar G, Narasu ML (2011) Bioconversion of novel substrate Saccharum spontaneum, a weedy material, into ethanol by Pichia stipitis NCIM3498. Bioresour Technol 102:1709–1714. https://doi.org/10.1016/j.biortech.2010.08.016
Chen W-H, Pen B-L, Yu C-T, Hwang W-S (2011) Pretreatment efficiency and structural characterization of rice straw by an integrated process of dilute-acid and steam explosion for bioethanol production. Bioresour Technol 102:2916–2924. https://doi.org/10.1016/j.biortech.2010.11.052
Christopher L (2012) Adding value prior to pulping: bioproducts from hemicellulose. In: Okia DCA (ed) Global perspectives on sustainable forest management. InTech
Coda R, Rizzello CG, Di Cagno R, Trani A, Cardinali G, Gobbetti M (2013) Antifungal activity of Meyerozyma guilliermondii: Identification of active compounds synthesized during dough fermentation and their effect on long-term storage of wheat bread. Food Microbiol 33:243–251. https://doi.org/10.1016/j.fm.2012.09.023
Comitini F, Ciani M (2011) Kluyveromyces wickerhamii killer toxin: purification and activity towards Brettanomyces/Dekkera yeasts in grape must: Kwkt killer toxin purification. FEMS Microbiol Lett 316:77–82. https://doi.org/10.1111/j.1574-6968.2010.02194.x
Corte L, di Cagno R, Groenewald M, Roscini L, Colabella C, Gobbetti M, Cardinali G (2015) Phenotypic and molecular diversity of Meyerozyma guilliermondii strains isolated from food and other environmental niches, hints for an incipient speciation. Food Microbiol 48:206–215. https://doi.org/10.1016/j.fm.2014.12.014
Crous PW, Wingfield MJ, Burgess TI, Carnegie AJ, Hardy GESJ, Smith D, Summerell BA, Cano-Lira JF, Guarro J, Houbraken J, Lombard L, Martín MP, Sandoval-Denis M, Alexandrova AV, Barnes CW, Baseia IG, Bezerra JDP, Guarnaccia V, May TW, Hernández-Restrepo M, Stchigel AM, Miller AN, Ordoñez ME, Abreu VP, Accioly T, Agnello C, Agustin Colmán A, Albuquerque CC, Alfredo DS, Alvarado P, Araújo-Magalhães GR, Arauzo S, Atkinson T, Barili A, Barreto RW, Bezerra JL, Cabral TS, Camello Rodríguez F, Cruz RHSF, Daniëls PP, da Silva BDB, de Almeida DAC, de Carvalho Júnior AA, Decock CA, Delgat L, Denman S, Dimitrov RA, Edwards J, Fedosova AG, Ferreira RJ, Firmino AL, Flores JA, García D, Gené J, Giraldo A, Góis JS, Gomes AAM, Gonçalves CM, Gouliamova DE, Groenewald M, Guéorguiev BV, Guevara-Suarez M, Gusmão LFP, Hosaka K, Hubka V, Huhndorf SM, Jadan M, Jurjević Ž, Kraak B, Kučera V, Kumar TKA, Kušan I, Lacerda SR, Lamlertthon S, Lisboa WS, Loizides M, Luangsa-ard JJ, Lysková P, Mac Cormack WP, Macedo DM, Machado AR, Malysheva EF, Marinho P, Matočec N, Meijer M, Mešić A, Mongkolsamrit S, Moreira KA, Morozova OV, Nair KU, Nakamura N, Noisripoom W, Olariaga I, Oliveira RJV, Paiva LM, Pawar P, Pereira OL, Peterson SW, Prieto M, Rodríguez-Andrade E, Rojo De Blas C, Roy M, Santos ES, Sharma R, Silva GA, Souza-Motta CM, Takeuchi-Kaneko Y, Tanaka C, Thakur A, Smith MT, Tkalčec Z, Valenzuela-Lopez N, van der Kleij P, Verbeken A, Viana MG, Wang XW, Groenewald JZ (2017) Fungal Planet description sheets: 625–715. Persoonia Mol Phylog Evolut Fungi. https://doi.org/10.3767/persoonia.2017.39.11
da Cunha-Pereira F, Hickert LR, Rech R, Dillon AP, Záchia Ayub MA (2017) Fermentation of hexoses and pentoses from hydrolyzed soybean Hull into ethanol and xylitol BY Candida guilliermondii BL 13. Braz J Chem Eng 34:927–936
Dantán-González E, Quiroz-Castañeda RE, Cobaxin-Cárdenas M, Valle-Hernández J, Gama-Martínez Y, Tinoco-Valencia JR, Serrano-Carreón L, Ortiz-Hernández L (2015) Impact of Meyerozyma guilliermondii isolated from chickens against Eimeria sp. protozoan, an in vitro analysis. BMC Vet Res 11:1–11. https://doi.org/10.1186/s12917-015-0589-0
de Albuquerque TL, da Silva IJ, de Macedo GR, Rocha MVP (2014) Biotechnological production of xylitol from lignocellulosic wastes: a review. Process Biochem 49:1779–1789. https://doi.org/10.1016/j.procbio.2014.07.010
de Lima PBA, Mulder KCL, Melo NTM, Carvalho LS, Menino GS, Mulinari E, de Castro VH, dos Reis TF, Goldman GH, Magalhães BS, Parachin NS (2016) Novel homologous lactate transporter improves l-lactic acid production from glycerol in recombinant strains of Pichia pastoris. Microb Cell Factories 15. https://doi.org/10.1186/s12934-016-0557-9
De Marco L, Epis S, Capone A, Martin E, Bozic J, Crotti E, Ricci I, Sassera D 2018a The genomes of four Meyerozyma caribbica isolates and novel insights into the Meyerozyma guilliermondii species complex. G3 (Bethesda) 8(3):755–759. https://doi.org/10.1534/g3.117.300316
De Marco L, Epis S, Capone A, Martin E, Bozic J, Crotti E, Ricci I, Sassera D (2018b) The genomes of four Meyerozyma caribbica isolates and novel insights into the Meyerozyma guilliermondii Species Complex. G3 (Bethesda) 8(3):755–759. https://doi.org/10.1534/g3.117.300316
De Schutter K, Lin Y-C, Tiels P, Van Hecke A, Glinka S, Weber-Lehmann J, Rouzé P, Van de Peer Y, Callewaert N (2009) Genome sequence of the recombinant protein production host Pichia pastoris. Nat Biotechnol 27:561–566. https://doi.org/10.1038/nbt.1544
de Souza G, Luana TCNV, Samila RPN, Maxwel AA (2017) Efficient production of second generation ethanol and xylitol by yeasts from Amazonian beetles (Coleoptera) and their galleries. Afr J Microbiol Res 11:814–824. https://doi.org/10.5897/AJMR2017.8522
Dias O, Gombert AK, Ferreira EC, Rocha I (2012) Genome-wide metabolic (re-) annotation of Kluyveromyces lactis. BMC Genomics 13:517. https://doi.org/10.1186/1471-2164-13-517
du Preez JC, van der Walt JP (1983) Fermentation of D-xylose to ethanol by a strain ofCandida shehatae. Biotechnol Lett 5:357–362. https://doi.org/10.1007/BF01141138
Duan H, Wang H, Ma B, Jiang P, Tu P, Ni Z, Li X, Li M, Ma X, Wang B, Wu R, Li M (2015) Codon optimization and expression of irisin in Pichia pastoris GS115. Int J Biol Macromol 79:21–26. https://doi.org/10.1016/j.ijbiomac.2015.04.030
Dujon B, Sherman D, Fischer G, Durrens P, Casaregola S, Lafontaine I, de Montigny J, Marck C, Neuvéglise C, Talla E, Goffard N, Frangeul L, Aigle M, Anthouard V, Babour A, Barbe V, Barnay S, Blanchin S, Beckerich J-M, Beyne E, Bleykasten C, Boisramé A, Boyer J, Cattolico L, Confanioleri F, de Daruvar A, Despons L, Fabre E, Fairhead C, Ferry-Dumazet H, Groppi A, Hantraye F, Hennequin C, Jauniaux N, Joyet P, Kachouri R, Kerrest A, Koszul R, Lemaire M, Lesur I, Ma L, Muller H, Nicaud J-M, Nikolski M, Oztas S, Ozier-Kalogeropoulos O, Pellenz S, Potier S, Richard G-F, Straub M-L, Suleau A, Swennen D, Tekaia F, Wésolowski-Louvel M, Westhof E, Wirth B, Zeniou-Meyer M, Zivanovic I, Bolotin-Fukuhara M, Thierry A, Bouchier C, Caudron B, Scarpelli C, Gaillardin C, Weissenbach J, Wincker P, Souciet J-L (2004) Genome evolution in yeasts. Nature 430:35–44. https://doi.org/10.1038/nature02579
Elshahed MS (2010) Microbiological aspects of biofuel production: Current status and future directions. J Adv Res 1:103–111. https://doi.org/10.1016/j.jare.2010.03.001
Faria NT, Santos MV, Fernandes P, Fonseca LL, Fonseca C, Ferreira FC (2014) Production of glycolipid biosurfactants, mannosylerythritol lipids, from pentoses and d-glucose/d-xylose mixtures by Pseudozyma yeast strains. Process Biochem 49:1790–1799. https://doi.org/10.1016/j.procbio.2014.08.004
Ferreira AD, Mussatto SI, Cadete RM, Rosa CA, Silva SS (2011) Ethanol production by a new pentose-fermenting yeast strain, Scheffersomyces stipitis UFMG-IMH 43.2, isolated from the Brazilian forest. Yeast 28:547–554. https://doi.org/10.1002/yea.1858
Fonseca GG, Heinzle E, Wittmann C, Gombert AK (2008) The yeast Kluyveromyces marxianus and its biotechnological potential. Appl Microbiol Biotechnol 79:339–354. https://doi.org/10.1007/s00253-008-1458-6
Franco Marcelino PR, da Silva VL, Rodrigues Philippini R, Von Zuben CJ, Contiero J, dos Santos JC, da Silva SS (2017) Biosurfactants produced by Scheffersomyces stipitis cultured in sugarcane bagasse hydrolysate as new green larvicides for the control of Aedes aegypti, a vector of neglected tropical diseases. PLoS One 12:e0187125. https://doi.org/10.1371/journal.pone.0187125
Gallo R, Trapp A (2017) The chemical conversion of biomass-derived saccharides: an overview. J Braz Chem Soc. https://doi.org/10.21577/0103-5053.20170009
García-Cubero MT, González-Benito G, Indacoechea I, Coca M, Bolado S (2009) Effect of ozonolysis pretreatment on enzymatic digestibility of wheat and rye straw. Bioresour Technol 100:1608–1613. https://doi.org/10.1016/j.biortech.2008.09.012
Garrote G, Dominguez H, Parajo JC (2002) Autohydrolysis of corncob: study of non-isothermal operation for xylooligasaccharide production. J Food Eng 52(3):211–218
Gı́rio F, Amaro C, Azinheira H, Pelica F, Amaral-Collaço M (2000) Polyols production during single and mixed substrate fermentations in Debaryomyces hansenii. Bioresour Technol 71:245–251. https://doi.org/10.1016/S0960-8524(99)00078-4
Gonçalves FAG, Colen G, Takahashi JA (2014) Yarrowia lipolytica and its multiple applications in the biotechnological industry. Sci World J 2014:1–14. https://doi.org/10.1155/2014/476207
Groeneveld P, Stouthamer AH, Westerhoff HV (2009) Super life – how and why ‘cell selection’ leads to the fastest-growing eukaryote: Control of highest eukaryotic growth rate. FEBS J 276:254–270. https://doi.org/10.1111/j.1742-4658.2008.06778.x
Groenewald M, Smith MT (2013) The teleomorph state of Candida deformans Langeron & Guerra and description of Yarrowia yakushimensis comb. nov. Antonie Van Leeuwenhoek 103:1023–1028. https://doi.org/10.1007/s10482-013-9882-8
Guan D, Li Y, Shiroma R, Ike M, Tokuyasu K (2013) Sequential incubation of Candida shehatae and ethanol-tolerant yeast cells for efficient ethanol production from a mixture of glucose, xylose and cellobiose. Bioresour Technol 132:419–422. https://doi.org/10.1016/j.biortech.2012.12.040
Gunah P (2011) Optimization of xylose production from sugarcane bagasse using response surface methodology (RSM). University of Malayia Pahang, Bachelor
Harhangi HR, Akhmanova AS, Emmens R, van der Drift C, de Laat WTAM, van Dijken JP, Jetten MSM, Pronk JT, Op den Camp HJM (2003) Xylose metabolism in the anaerobic fungus Piromyces sp. strain E2 follows the bacterial pathway. Arch Microbiol 180:134–141. https://doi.org/10.1007/s00203-003-0565-0
Hernández-Pérez AF, de Arruda PV, Felipe M das G de A (2016) Sugarcane straw as a feedstock for xylitol production by Candida guilliermondii FTI 20037. Braz J Microbiol 47:489–496. https://doi.org/10.1016/j.bjm.2016.01.019
Hommel RK, Ahnert P (1999) Candida. In: Encyclopedia of food microbiology, 1st edn. Elsevier Ltd, London
Hou X (2012) Anaerobic xylose fermentation by Spathaspora passalidarum. Appl Microbiol Biotechnol 94:205–214. https://doi.org/10.1007/s00253-011-3694-4
Hou J, Qiu C, Shen Y, Li H, Bao X (2017) Engineering of Saccharomyces cerevisiae for the efficient co-utilization of glucose and xylose. FEMS Yeast Res 17. https://doi.org/10.1093/femsyr/fox034
Ilmen M, Koivuranta K, Ruohonen L, Suominen P, Penttila M (2007) Efficient production of L-lactic acid from xylose by Pichia stipitis. Appl Environ Microbiol 73:117–123. https://doi.org/10.1128/AEM.01311-06
Jeffries TW (1983) Utilization of xylose by bacteria, yeasts, and fungi. In: Pentoses and lignin. Springer, Berlin/Heidelberg, pp 1–32
Jeffries TW (2006) Engineering yeasts for xylose metabolism. Curr Opin Biotechnol 17:320–326. https://doi.org/10.1016/j.copbio.2006.05.008
Jeffries TW, Van Vleet JRH (2009) Pichia stipitis genomics, transcriptomics, and gene clusters. FEMS Yeast Res 9:793–807. https://doi.org/10.1111/j.1567-1364.2009.00525.x
Jeffries TW, Grigoriev IV, Grimwood J, Laplaza JM, Aerts A, Salamov A, Schmutz J, Lindquist E, Dehal P, Shapiro H, Jin Y-S, Passoth V, Richardson PM (2007) Genome sequence of the lignocellulose-bioconverting and xylose-fermenting yeast Pichia stipitis. Nat Biotechnol 25:319–326. https://doi.org/10.1038/nbt1290
Ji X, Ma H, Tian Z, Lyu G, Fang G, Chen J, Saeed HAM (2017) Production of xylose from diluted sulfuric acid hydrolysis of wheat straw. Bioresources 12:7084–7095
Johnsen U, Dambeck M, Zaiss H, Fuhrer T, Soppa J, Sauer U, Schönheit P (2009) d-xylose degradation pathway in the halophilic Archaeon Haloferax volcanii. J Biol Chem 284:27290–27303. https://doi.org/10.1074/jbc.M109.003814
Jørgensen H, Kristensen JB, Felby C (2007) Enzymatic conversion of lignocellulose into fermentable sugars: challenges and opportunities. Biofuels Bioprod Biorefin 1:119–134. https://doi.org/10.1002/bbb.4
Kim D (2018) Physico-chemical conversion of lignocellulose: inhibitor effects and detoxification strategies: a mini review. Molecules 23:309. https://doi.org/10.3390/molecules23020309
Kim D, Woo HM (2018) Deciphering bacterial xylose metabolism and metabolic engineering of industrial microorganisms for use as efficient microbial cell factories. Appl Microbiol Biotechnol 102:9471–9480. https://doi.org/10.1007/s00253-018-9353-2
Kim TH, Taylor F, Hicks KB (2008) Bioethanol production from barley hull using SAA (soaking in aqueous ammonia) pretreatment. Bioresour Technol 99:5694–5702. https://doi.org/10.1016/j.biortech.2007.10.055
Kim SR, Park Y-C, Jin Y-S, Seo J-H (2013) Strain engineering of Saccharomyces cerevisiae for enhanced xylose metabolism. Biotechnol Adv 31:851–861. https://doi.org/10.1016/j.biotechadv.2013.03.004
Kim D, Ximenes EA, Nichols NN, Cao G, Frazer SE, Ladisch MR (2016) Maleic acid treatment of biologically detoxified corn stover liquor. Bioresour Technol 216:437–445. https://doi.org/10.1016/j.biortech.2016.05.086
Kim D, Orrego D, Ximenes EA, Ladisch MR (2017) Cellulose conversion of corn pericarp without pretreatment. Bioresour Technol 245:511–517. https://doi.org/10.1016/j.biortech.2017.08.156
Kręgiel D, Pawlikowska E, Antolak H (2017) Non-conventional yeasts in fermentation processes: potentialities and limitations. In: Lucas C, Pais C (eds) Old yeasts – new questions. InTech
Kumar V, Krishania M, Preet Sandhu P, Ahluwalia V, Gnansounou E, Sangwan RS (2018) Efficient detoxification of corn cob hydrolysate with ion-exchange resins for enhanced xylitol production by Candida tropicalis MTCC 6192. Bioresour Technol 251:416–419. https://doi.org/10.1016/j.biortech.2017.11.039
Kuroda K, Kobayashi K, Kitagawa Y, Nakagawa T, Tsumura H, Komeda T, Shinmi D, Mori E, Motoki K, Fuju K, Sakai T, Nonaka K, Suzuki T, Ichikawa K, Chiba Y, Jigami Y (2008) Efficient antibody production upon suppression of O mannosylation in the yeast Ogataea minuta. Appl Environ Microbiol 74:446–453. https://doi.org/10.1128/AEM.02106-07
Kurtzman CP (2005) New species and a new combination in the Hyphopichia and Yarrowia yeast clades. Antonie Van Leeuwenhoek 88:121–130. https://doi.org/10.1007/s10482-005-2495-0
Kurtzman CP (2009) Biotechnological strains of Komagataella (Pichia) pastoris are Komagataella phaffii as determined from multigene sequence analysis. J Ind Microbiol Biotechnol 36:1435–1438. https://doi.org/10.1007/s10295-009-0638-4
Kurtzman CP (2011a) Yarrowia van der Walt & von Arx (1980). In: The yeasts. Elsevier, Amsterdam, pp 927–929
Kurtzman CP (ed) (2011b) The yeasts: a taxonomic study 5 Elsevier, Amsterdam
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek 73:331
Kurtzman C, Robnett C (2003) Phylogenetic relationships among yeasts of the ? complex? determined from multigene sequence analyses. FEMS Yeast Res 3:417–432. https://doi.org/10.1016/S1567-1356(03)00012-6
Kurtzman CP, Suzuki M (2010) Phylogenetic analysis of ascomycete yeasts that form coenzyme Q-9 and the proposal of the new genera Babjeviella, Meyerozyma, Millerozyma, Priceomyces, and Scheffersomyces. Mycoscience 51:2–14. https://doi.org/10.1007/S10267-009-0011-5
Kurtzman M, Suzuki M, Kurtzman CP, Clade C, Clade C, U CY-T, Ay MY-T, Ay CY-T, Ay CY-T, Ay CY-T, U DY-T (2010) Meyerozyma Kurtzman & M. Suzuki (2010). Elsevier BV, London
Kurylenko OO, Ruchala J, Hryniv OB, Abbas CA, Dmytruk KV, Sibirny AA (2014) Metabolic engineering and classical selectionof the methylotrophic thermotolerant yeast Hansenulapolymorpha for improvement of high-temperature xylose alcoholicfermentation. Microb Cell Factories 13:122. https://doi.org/10.1186/s12934-014-0122-3
Kwak S, Jin Y-S (2017) Production of fuels and chemicals from xylose by engineered Saccharomyces cerevisiae: a review and perspective. Microb Cell Factories 16. https://doi.org/10.1186/s12934-017-0694-9
Lachance M-A (2007) Current status of Kluyveromyces systematics. FEMS Yeast Res 7:642–645. https://doi.org/10.1111/j.1567-1364.2006.00197.x
Lachance M-A (2011) Kluyveromyces van der Walt (1971). In: The yeasts. Elsevier, Amsterdam, pp 471–481
Lane MM, Morrissey JP (2010) Kluyveromyces marxianus: A yeast emerging from its sister’s shadow. Fungal Biol Rev 24:17–26. https://doi.org/10.1016/j.fbr.2010.01.001
Ledesma-Amaro R, Lazar Z, Rakicka M, Guo Z, Fouchard F, Coq A-MC-L, Nicaud J-M (2016) Metabolic engineering of Yarrowia lipolytica to produce chemicals and fuels from xylose. Metab Eng 38:115–124. https://doi.org/10.1016/j.ymben.2016.07.001
Li H, Alper HS (2016) Enabling xylose utilization in Yarrowia lipolytica for lipid production. Biotechnol J 11:1230–1240. https://doi.org/10.1002/biot.201600210
Li C, Knierim B, Manisseri C, Arora R, Scheller HV, Auer M, Vogel KP, Simmons BA, Singh S (2010) Comparison of dilute acid and ionic liquid pretreatment of switchgrass: Biomass recalcitrance, delignification and enzymatic saccharification. Bioresour Technol 101:4900–4906. https://doi.org/10.1016/j.biortech.2009.10.066
Li P, Sun H, Chen Z, Li Y, Zhu T (2015) Construction of efficient xylose utilizing Pichia pastoris for industrial enzyme production. Microb Cell Factories 14. https://doi.org/10.1186/s12934-015-0206-8
Liu Y, Wu C, Wang J, Mo W, Yu M (2013) Codon optimization, expression, purification, and functional characterization of recombinant human IL-25 in Pichia pastoris. Appl Microbiol Biotechnol 97:10349–10358. https://doi.org/10.1007/s00253-013-5264-4
Liu H-H, Ji X-J, Huang H (2015) Biotechnological applications of Yarrowia lipolytica: Past, present and future. Biotechnol Adv 33:1522–1546. https://doi.org/10.1016/j.biotechadv.2015.07.010
Liu X-J, Cao W-N, Ren Y-C, Xu L-L, Yi Z-H, Liu Z, Hui F-L (2016) Taxonomy and physiological characterisation of Scheffersomyces titanus sp. nov., a new D-xylose-fermenting yeast species from China. Sci Rep 6. https://doi.org/10.1038/srep32181
Lobo FP, Goncalves DL, Alves SL, Gerber AL, de Vasconcelos ATR, Basso LC, Franco GR, Soares MA, Cadete RM, Rosa CA, Stambuk BU (2014a) Draft genome sequence of the D-xylose-fermenting yeast Spathaspora arborariae UFMG-HM19.1AT. Genome Announ 2. https://doi.org/10.1128/genomeA.01163-13
Lobo FP, Goncalves DL, Alves SL, Gerber AL, de Vasconcelos ATR, Basso LC, Franco GR, Soares MA, Cadete RM, Rosa CA, Stambuk BU (2014b) Draft genome sequence of the D-xylose-fermenting yeast Spathaspora arborariae UFMG-HM19.1AT. Genome Announ 2. https://doi.org/10.1128/genomeA.01163-13
Löbs A-K, Schwartz C, Wheeldon I (2017) Genome and metabolic engineering in non-conventional yeasts: Current advances and applications. Synthet Syst Biotechnol 2:198–207. https://doi.org/10.1016/j.synbio.2017.08.002
Long TM, Su Y-K, Headman J, Higbee A, Willis LB, Jeffries TW (2012) Cofermentation of glucose, xylose, and cellobiose by the beetle-associated yeast Spathaspora passalidarum. Appl Environ Microbiol 78:5492–5500. https://doi.org/10.1128/AEM.00374-12
Lopes MR, Morais CG, Kominek J, Cadete RM, Soares MA, Uetanabaro APT, Fonseca C, Lachance M-A, Hittinger CT, Rosa CA (2016) Genomic analysis and D-xylose fermentation of three novel Spathaspora species: Spathaspora girioi sp. nov., Spathaspora hagerdaliae f. a., sp. nov. and Spathaspora gorwiae f. a., sp. nov. FEMS Yeast Res 16:fow044. https://doi.org/10.1093/femsyr/fow044
Lopes DD, Cibulski SP, Mayer FQ, Siqueira FM, Rosa CA, Hector RE, Ayub MAZ (2017) Draft genome sequence of the d-xylose-fermenting yeast Spathaspora xylofermentans UFMG-HMD23.3. Genome Announ 5. https://doi.org/10.1128/genomeA.00815-17
Lopes MR, Batista TM, Franco GR, Ribeiro LR, Santos ARO, Furtado C, Moreira RG, Goes-Neto A, Vital MJS, Rosa LH, Lachance M-A, Rosa CA (2018) Scheffersomyces stambukii f.a., sp. nov., a d-xylose-fermenting species isolated from rotting wood. Int J Syst Evol Microbiol 68:2306–2312. https://doi.org/10.1099/ijsem.0.002834
Ma K, He M, You H, Pan L, Hu G, Cui Y, Maeda T (2017) Enhanced fuel ethanol production from rice straw hydrolysate by an inhibitor-tolerant mutant strain of Scheffersomyces stipitis. RSC Adv 7:31180–31188. https://doi.org/10.1039/C7RA04049K
Madhavan A, Tamalampudi S, Ushida K, Kanai D, Katahira S, Srivastava A, Fukuda H, Bisaria VS, Kondo A (2009) Xylose isomerase from polycentric fungus Orpinomyces: gene sequencing, cloning, and expression in Saccharomyces cerevisiae for bioconversion of xylose to ethanol. Appl Microbiol Biotechnol 82:1067–1078. https://doi.org/10.1007/s00253-008-1794-6
Mans R, Daran J-MG, Pronk JT (2018) Under pressure: evolutionary engineering of yeast strains for improved performance in fuels and chemicals production. Curr Opin Biotechnol 50:47–56. https://doi.org/10.1016/j.copbio.2017.10.011
Martín C, Klinke HB, Thomsen AB (2007) Wet oxidation as a pretreatment method for enhancing the enzymatic convertibility of sugarcane bagasse. Enzym Microb Technol 40:426–432. https://doi.org/10.1016/j.enzmictec.2006.07.015
Martini C, Tauk-Tornisielo SM, Codato CB, Bastos RG, Ceccato-Antonini SR (2016) A strain of Meyerozyma guilliermondii isolated from sugarcane juice is able to grow and ferment pentoses in synthetic and bagasse hydrolysate media. World J Microbiol Biotechnol 32. https://doi.org/10.1007/s11274-016-2036-1
Martins GM, Bocchini-Martins DA, Bezzerra-Bussoli C, Pagnocca FC, Boscolo M, Monteiro DA, da Silva R, Gomes E (2018) The isolation of pentose-assimilating yeasts and their xylose fermentation potential. Braz J Microbiol 49:162–168. https://doi.org/10.1016/j.bjm.2016.11.014
Mateo S, Puentes JG, Moya AJ, Sánchez S (2015) Ethanol and xylitol production by fermentation of acid hydrolysate from olive pruning with Candida tropicalis NBRC 0618. Bioresour Technol 190:1–6. https://doi.org/10.1016/j.biortech.2015.04.045
Mattanovich D, Graf A, Stadlmann J, Dragosits M, Redl A, Maurer M, Kleinheinz M, Sauer M, Altmann F, Gasser B (2009) Genome, secretome and glucose transport highlight unique features of the protein production host Pichia pastoris. Microb Cell Factories 8:29. https://doi.org/10.1186/1475-2859-8-29
Mellitzer A, Weis R, Glieder A, Flicker K (2012) Expression of lignocellulolytic enzymes in Pichia pastoris. Microb Cell Factories 11:61. https://doi.org/10.1186/1475-2859-11-61
Meyer SA, Anderson K, Brown RE, Smith MT, Yarrow D, Mitchell G, Ahearn DG (1975) Physiological and DNA characterization of Candida maltose, a hydrocarbon-utilizing yeast. Arch Microbiol 104:225–231
Morais CG, Batista TM, Kominek J, Borelli BM, Furtado C, Moreira RG, Franco GR, Rosa LH, Fonseca C, Hittinger CT, Lachance M-A, Rosa CA (2017) Spathaspora boniae sp. nov., a D-xylose-fermenting species in the Candida albicans/Lodderomyces clade. Int J Syst Evol Microbiol 67:3798–3805. https://doi.org/10.1099/ijsem.0.002186
Morales P, Gentina JC, Aroca G, Mussatto SI (2017) Development of an acetic acid tolerant Spathaspora passalidarum strain through evolutionary engineering with resistance to inhibitors compounds of autohydrolysate of Eucalyptus globulus. Ind Crop Prod 106:5–11. https://doi.org/10.1016/j.indcrop.2016.12.023
Moran G (2004) Comparative genomics using Candida albicans DNA microarrays reveals absence and divergence of virulence-associated genes in Candida dubliniensis. Microbiology 150:3363–3382. https://doi.org/10.1099/mic.0.27221-0
Morrissey JP, Etschmann MMW, Schrader J, de Billerbeck GM (2015) Cell factory applications of the yeast Kluyveromyces marxianus for the biotechnological production of natural flavour and fragrance molecules. Yeast 32:3–16
Mosier N (2005) Features of promising technologies for pretreatment of lignocellulosic biomass. Bioresour Technol 96:673–686. https://doi.org/10.1016/j.biortech.2004.06.025
Moysés D, Reis V, Almeida J, Moraes L, Torres F (2016) Xylose Fermentation by Saccharomyces cerevisiae: Challenges and Prospects. Int J Mol Sci 17:207. https://doi.org/10.3390/ijms17030207
Mukherjee V, Radecka D, Aerts G, Verstrepen KJ, Lievens B, Thevelein JM (2017) Phenotypic landscape of non-conventional yeast species for different stress tolerance traits desirable in bioethanol fermentation. Biotechnol Biofuels 10. https://doi.org/10.1186/s13068-017-0899-5
Nagy E (2015) Biodiversity of food spoilage Yarrowia group in different kinds of food. Dissertation, University of Budapest
Nagy E, Niss M, Dlauchy D, Arneborg N, Nielsen DS, Peter G (2013) Yarrowia divulgata f.a., sp. nov., a yeast species from animal-related and marine sources. Int J Syst Evol Microbiol 63:4818–4823. https://doi.org/10.1099/ijs.0.057208-0
Nagy E, Dlauchy D, Medeiros AO, Péter G, Rosa CA (2014) Yarrowia porcina sp. nov. and Yarrowia bubula f.a. sp. nov., two yeast species from meat and river sediment. Antonie Van Leeuwenhoek 105:697–707. https://doi.org/10.1007/s10482-014-0125-4
Najjar A, Robert S, Guérin C, Violet-Asther M, Carrière F (2011) Quantitative study of lipase secretion, extracellular lipolysis, and lipid storage in the yeast Yarrowia lipolytica grown in the presence of olive oil: analogies with lipolysis in humans. Appl Microbiol Biotechnol 89:1947–1962. https://doi.org/10.1007/s00253-010-2993-5
Nakanishi SC, Soares LB, Biazi LE, Nascimento VM, Costa AC, Rocha GJM, Ienczak JL (2017) Fermentation strategy for second generation ethanol production from sugarcane bagasse hydrolyzate by Spathaspora passalidarum and Scheffersomyces stipitis: Fermentation Strategy for Second Generation Ethanol Production. Biotechnol Bioeng 114:2211–2221. https://doi.org/10.1002/bit.26357
Nakase T, Komagata K (1971) Significance of DNA base composition in the classification of the yeast genus Candida. J Gen Appl Microbiol 17:259–179
Nguyen NH, Suh S-O, Marshall CJ, Blackwell M (2006) Morphological and ecological similarities: wood-boring beetles associated with novel xylose-fermenting yeasts, Spathaspora passalidarum gen. sp. nov. and Candida jeffriesii sp. nov. Mycol Res 110:1232–1241. https://doi.org/10.1016/j.mycres.2006.07.002
Niehus X, Crutz-Le Coq A-M, Sandoval G, Nicaud J-M, Ledesma-Amaro R (2018) Engineering Yarrowia lipolytica to enhance lipid production from lignocellulosic materials. Biotechnol Biofuels 11. https://doi.org/10.1186/s13068-018-1010-6
Nygård Y, Toivari MH, Penttilä M, Ruohonen L, Wiebe MG (2011) Bioconversion of d-xylose to d-xylonate with Kluyveromyces lactis. Metab Eng 13:383–391. https://doi.org/10.1016/j.ymben.2011.04.001
Odds FC (2010) Molecular phylogenetics and epidemiology of Candida albicans. Future Microbiol 5:67–79. https://doi.org/10.2217/fmb.09.113
Okada N, Tanimura A, Hirakawa H, Takashima M, Ogawa J, Shima J (2017) Draft Genome Sequences of the Xylose-Fermenting Yeast Scheffersomyces shehatae NBRC 1983 T and a Thermotolerant Isolate of S. shehatae ATY839 (JCM 18690). Genome Announ 5. https://doi.org/10.1128/genomeA.00347-17
Palsson BO, Fathi-Afshar S, Rudd DF, Lightfoot EN (1981) Biomass as a Source of Chemical Feedstocks: An Economic Evaluation. Science 213:513–517. https://doi.org/10.1126/science.213.4507.513
Protchenko O, Philpott CC, Sibirny AA (2011) Insertion mutagenesis. doi:https://doi.org/10.1111/j.1567-1364.2011.00720.x
Puseenam A, Kocharin K, Tanapongpipat S, Eurwilaichitr L, Ingsriswang S, Roongsawang N (2018) A novel sucrose-based expression system for heterologous proteins expression in thermotolerant methylotrophic yeast Ogataea thermomethanolica. FEMS Microbiol Lett 365. https://doi.org/10.1093/femsle/fny238
Radecka D, Mukherjee V, Mateo RQ, Stojiljkovic M, Foulquié-Moreno MR, Thevelein JM (2015) Looking beyond Saccharomyces : the potential of non-conventional yeast species for desirable traits in bioethanol fermentation. FEMS Yeast Res 15:fov053. https://doi.org/10.1093/femsyr/fov053
Rebello S, Abraham A, Madhavan A, Sindhu R, Binod P, Babu AK, Aneesh EM, Pandey A (2018) Non-conventional Yeast cell factories for sustainable bioprocesses. FEMS Microbiol Lett. https://doi.org/10.1093/femsle/fny222
Ren Y, Chen L, Niu Q, Hui F (2014) Description of Scheffersomyces henanensis sp. nov., a new D-xylose-fermenting yeast species isolated from rotten wood. PLoS One 9:e92315. https://doi.org/10.1371/journal.pone.0092315
Riley R, Haridas S, Wolfe KH, Lopes MR, Hittinger CT, Göker M, Salamov AA, Wisecaver JH, Long TM, Calvey CH, Aerts AL, Barry KW, Choi C, Clum A, Coughlan AY, Deshpande S, Douglass AP, Hanson SJ, Klenk H-P, LaButti KM, Lapidus A, Lindquist EA, Lipzen AM, Meier-Kolthoff JP, Ohm RA, Otillar RP, Pangilinan JL, Peng Y, Rokas A, Rosa CA, Scheuner C, Sibirny AA, Slot JC, Stielow JB, Sun H, Kurtzman CP, Blackwell M, Grigoriev IV, Jeffries TW (2016) Comparative genomics of biotechnologically important yeasts. Proc Natl Acad Sci 113:9882–9887. https://doi.org/10.1073/pnas.1603941113
Rodicio R, Heinisch JJ (2013) Yeast on the milky way: genetics, physiology and biotechnology of Kluyveromyces lactis: Kluyveromyces lactis. Yeast 30:165–177. https://doi.org/10.1002/yea.2954
Rodrussamee N, Sattayawat P, Yamada M (2018) Highly efficient conversion of xylose to ethanol without glucose repression by newly isolated thermotolerant Spathaspora passalidarum CMUWF1–2. BMC Microbiol 18. https://doi.org/10.1186/s12866-018-1218-4
Romi W, Keisam S, Ahmed G, Jeyaram K (2014) Reliable differentiation of Meyerozyma guilliermondii from Meyerozyma caribbica by internal transcribed spacer restriction fingerprinting. BMC Microbiol 14:52. https://doi.org/10.1186/1471-2180-14-52
Rosa C, Lachance M, Silva J, Teixeira A, Marini M, Antonini Y, Martins R (2003) Yeast communities associated with stingless bees. FEMS Yeast Res 4:271–275. https://doi.org/10.1016/S1567-1356(03)00173-9
Ruiz E, Cara C, Manzanares P, Ballesteros M, Castro E (2008) Evaluation of steam explosion pre-treatment for enzymatic hydrolysis of sunflower stalks. Enzym Microb Technol 42:160–166. https://doi.org/10.1016/j.enzmictec.2007.09.002
Ryabova O, Chmil O, Sibirny A (2003) Xylose and cellobiose fermentation to ethanol by the thermotolerant methylotrophic yeast. FEMS Yeast Res 4:157–164. https://doi.org/10.1016/S1567-1356(03)00146-6
Ryu S, Trinh CT (2017) Understanding functional roles of native pentose-specific transporters for activating dormant pentose metabolism in Yarrowia lipolytica. Appl Environ Microbiol 84. https://doi.org/10.1128/AEM.02146-17
Ryu S, Hipp J, Trinh CT (2016) Activating and Elucidating Metabolism of Complex Sugars in Yarrowia lipolytica. Appl Environ Microbiol 82:1334–1345. https://doi.org/10.1128/AEM.03582-15
Sampaio FC, da Silveira WB, Chaves-Alves VM, Passos FML, Coelho JLC (2003) Screening of filamentous fungi for production of xylitol from D-xylose. Braz J Microbiol 34:321–324. https://doi.org/10.1590/S1517-83822003000400007
Satish T, Murthy NYS (2010) Optimisation of xylose production using xylanase. Intl J Sci 8:909–913
Schauer F, Hanschke R (1999) Taxonomy and ecology of the genus Candida. Mycosis 42:12–21
Schreiber WT, Geib NV, Wingfield B, Acree SF (1930) Semi-commercial production of xylose. Ind Eng Chem 22:497–501. https://doi.org/10.1021/ie50245a020
Shu M, Shen W, Yang S, Wang X, Wang F, Wang Y, Ma L (2016) High-level expression and characterization of a novel serine protease in Pichia pastoris by multi-copy integration. Enzym Microb Technol 92:56–66. https://doi.org/10.1016/j.enzmictec.2016.06.007
Slininger PJ, Bothast RJ, Van Cauwenberge JE, Kurtzman CP (1982) Conversion of D-xylose to ethanol by the yeastPachysolen tannophilus. Biotechnol Bioeng 24:371–384. https://doi.org/10.1002/bit.260240210
Song KH, Song JY, Hong SG, Baek H, Kim SY, Hyun HH (2002) Production of mannitol by a novel strain of Candida magnoliae. Biotechnol Lett 24:9–12
Spagnuolo M, Shabbir Hussain M, Gambill L, Blenner M (2018) Alternative substrate metabolism in Yarrowia lipolytica. Front Microbiol 9. https://doi.org/10.3389/fmicb.2018.01077
Spencer JFT, Spencer DM (eds) (1997) Yeasts in natural and artificial habitats. Springer, Berlin/Heidelberg
Spohner SC, Schaum V, Quitmann H, Czermak P (2016) Kluyveromyces lactis: an emerging tool in biotechnology. J Biotechnol 222:104–116. https://doi.org/10.1016/j.jbiotec.2016.02.023
Sreekrishna K, Kropp KE (1996) Pichia pastoris. In: Nonconventional yeasts in biotechnology. Springer, Berlin/Heidelberg, pp 203–253
Stambuk BU, Franden MA, Singh A, Zhang M (2003) D-xylose transport by Candida succiphila and Kluyveromyces marxianus. Appl Biochem Biotechnol 106:255–264. https://doi.org/10.1385/ABAB:106:1-3:255
Stenderup A, Bak AL (1968) Deoxyribonucleic acid base composition of some species within the genus Candida. J Gen Microbiol 52:231–236
Stephanopoulos G TM (2013) Engineered microbes and methods for microbial oil overproduction from cellulosic materials
Su Y-K, Willis LB, Jeffries TW (2015) Effects of aeration on growth, ethanol and polyol accumulation by Spathaspora passalidarum NRRL Y-27907 and Scheffersomyces stipitis NRRL Y-7124: Aeration Effects on Xylose Fermenting Yeasts. Biotechnol Bioeng 112:457–469. https://doi.org/10.1002/bit.25445
Su Y-K, Willis LB, Rehmann L, Smith DR, Jeffries TW (2018) Spathaspora passalidarum selected for resistance to AFEX hydrolysate shows decreased cell yield. FEMS Yeast Res 18. https://doi.org/10.1093/femsyr/foy011
Suh S-O, Marshall CJ, Mchugh JV, Blackwell M (2003) Wood ingestion by passalid beetles in the presence of xylose-fermenting gut yeasts: gut yeasts of passalid beetles. Mol Ecol 12:3137–3145. https://doi.org/10.1046/j.1365-294X.2003.01973.x
Sun J, Ding S-Y, Doran-Peterson J (2013) Chapter 1: biomass and its biorefinery: novel approaches from nature-inspired strategies and technology. In: Sun J, Ding S-Y, Peterson JD (eds) Energy and environment series. Royal Society of Chemistry, Cambridge, pp 1–13
Swain MR, Krishnan C (2015) Improved conversion of rice straw to ethanol and xylitol by combination of moderate temperature ammonia pretreatment and sequential fermentation using Candida tropicalis. Ind Crop Prod 77:1039–1046. https://doi.org/10.1016/j.indcrop.2015.10.013
Tizazu BZ, Roy K, Moholkar VS (2018) Ultrasonic enhancement of xylitol production from sugarcane bagasse using immobilized Candida tropicalis MTCC 184. Bioresour Technol 268:247–258. https://doi.org/10.1016/j.biortech.2018.07.141
Tsai CT, Huang C-T (2008) Overexpression of the Neocallimastix frontalis xylanase gene in the methylotrophic yeasts Pichia pastoris and Pichia methanolica. Enzym Microb Technol 42:459–465. https://doi.org/10.1016/j.enzmictec.2008.01.018
Tsigie YA, Wang C-Y, Truong C-T, Ju Y-H (2011) Lipid production from Yarrowia lipolytica Po1g grown in sugarcane bagasse hydrolysate. Bioresour Technol 102:9216–9222. https://doi.org/10.1016/j.biortech.2011.06.047
Urbina H, Blackwell M (2012) Multilocus phylogenetic study of the scheffersomyces yeast clade and characterization of the N-terminal region of xylose reductase gene. PLoS ONE 7. https://doi.org/10.1371/journal.pone.0039128
Vallejos ME, Chade M, Mereles EB, Bengoechea DI, Brizuela JG, Felissia FE, Area MC (2016) Strategies of detoxification and fermentation for biotechnological production of xylitol from sugarcane bagasse. Ind Crop Prod 91:161–169. https://doi.org/10.1016/j.indcrop.2016.07.007
van der Walt JP (1965) The emendation of the genus Kluyveromyces v. d. Walt. Antonie Van Leeuwenhoek 31:341–348
van der Walt JP, von Arx JA (1980) The yeast genus Yarrowia gen. nov. Antonie Van Leeuwenhoek 46:517–521. https://doi.org/10.1007/BF00394008
van Ooyen AJJ, Dekker P, Huang M, Olsthoorn MMA, Jacobs DI, Colussi PA, Taron CH (2006) Heterologous protein production in the yeast Kluyveromyces lactis. FEMS Yeast Res 6:381–392. https://doi.org/10.1111/j.1567-1364.2006.00049.x
van Wyk JP (2001) Biotechnology and the utilization of biowaste as a resource for bioproduct development. Trends Biotechnol 19:172–177
Varela JA, Gethins L, Stanton C, Ross P, Morrissey JP (2017) Applications of Kluyveromyces marxianus in biotechnology. In: Satyanarayana T, Kunze G (eds) Yeast diversity in human welfare. Springer, Singapore, pp 439–453
Varize CS, Cadete RM, Lopes LD, Christofoleti-Furlan RM, Lachance M-A, Rosa CA, Basso LC (2018) Spathaspora piracicabensis f. a., sp. nov., a d-xylose-fermenting yeast species isolated from rotting wood in Brazil. Antonie Van Leeuwenhoek 111:525–531. https://doi.org/10.1007/s10482-017-0974-8
Vaz de Arruda P, dos Santos JC, de Cássia Lacerda Brambilla Rodrigues R, da Silva DDV, Yamakawa CK, de Moraes Rocha GJ, Júnior JN, da Cruz Pradella JG, Vaz Rossell CE, das Graças de Almeida Felipe M (2017) Scale up of xylitol production from sugarcane bagasse hemicellulosic hydrolysate by Candida guilliermondii FTI 20037. J Ind Eng Chem 47:297–302. https://doi.org/10.1016/j.jiec.2016.11.046
Venkateswar Rao L, Goli JK, Gentela J, Koti S (2015) Bioconversion of lignocellulosic biomass to xylitol: an overview. Bioresour Technol 213:299–310. https://doi.org/10.1016/j.biortech.2016.04.092
Veras HCT, Parachin NS, Almeida JRM (2017) Comparative assessment of fermentative capacity of different xylose-consuming yeasts. Microb Cell Factories 16. https://doi.org/10.1186/s12934-017-0766-x
Verduyn C, Van Kleef R, Frank J, Schreuder H, Van Dijken JP, Scheffers WA (1985) Properties of the NAD(P)H-dependent xylose reductase from the xylose-fermenting yeast Pichia stipitis. Biochem J 226:669–677
Wang PY, Shopsis C, Schneider H (1980) Fermentation of a pentose by yeasts. Biochem Biophys Res Commun 94:248–254. https://doi.org/10.1016/S0006-291X(80)80213-0
Wang Y, Ren Y-C, Zhang Z-T, Ke T, Hui F-L (2016) Spathaspora allomyrinae sp. nov., a d-xylose-fermenting yeast species isolated from a scarabeid beetle Allomyrina dichotoma. Int J Syst Evol Microbiol 66:2008–2012. https://doi.org/10.1099/ijsem.0.000979
Wannawilai S, Chisti Y, Sirisansaneeyakul S (2017) A model of furfural-inhibited growth and xylitol production by Candida magnoliae TISTR 5663. Food Bioprod Process 105:129–140. https://doi.org/10.1016/j.fbp.2017.07.002
Webb SR, Lee H (1990) Regulation of d-xylose utilization by hexoses in pentose-fermenting yeasts. Biotechnol Adv 8:685–697. https://doi.org/10.1016/0734-9750(90)91991-O
Wei L, Liu J, Qi H, Wen J (2015) Engineering Scheffersomyces stipitis for fumaric acid production from xylose. Bioresour Technol 187:246–254. https://doi.org/10.1016/j.biortech.2015.03.122
Wickerham LJ, Burton KA (1954) A clarification of the relationship of Candida Guilliermondii to other yeasts by a study of their mating types. J Bacteriol 68(5):594–597
Wohlbach DJ, Kuo A, Sato TK, Potts KM, Salamov AA, LaButti KM, Sun H, Clum A, Pangilinan JL, Lindquist EA, Lucas S, Lapidus A, Jin M, Gunawan C, Balan V, Dale BE, Jeffries TW, Zinkel R, Barry KW, Grigoriev IV, Gasch AP (2011a) Comparative genomics of xylose-fermenting fungi for enhanced biofuel production. Proc Natl Acad Sci 108:13212–13217. https://doi.org/10.1073/pnas.1103039108
Wohlbach DJ, Kuo A, Sato TK, Potts KM, Salamov AA, LaButti KM, Sun H, Clum A, Pangilinan JL, Lindquist EA, Lucas S, Lapidus A, Jin M, Gunawan C, Balan V, Dale BE, Jeffries TW, Zinkel R, Barry KW, Grigoriev IV, Gasch AP (2011b) Comparative genomics of xylose-fermenting fungi for enhanced biofuel production. Proc Natl Acad Sci 108:13212–13217. https://doi.org/10.1073/pnas.1103039108
Wrent P, Rivas EM, Peinado JM, de Silóniz MI (2016) Development of an affordable typing method for Meyerozyma guilliermondii using microsatellite markers. Int J Food Microbiol 217:1–6. https://doi.org/10.1016/j.ijfoodmicro.2015.10.008
Yaguchi A, Spagnuolo M, Blenner M (2018) Engineering yeast for utilization of alternative feedstocks. Curr Opin Biotechnol 53:122–129. https://doi.org/10.1016/j.copbio.2017.12.003
Yamada Y, Kondo K (1972) Taxonomic significance of coenzyme Q system in yeasts and yeast-like fungi (1). Yeast 363:373
Yamada Y, Maeda K, Mikata K (1994) The Phylogenetic Relationships of the Hat-shaped Ascospore-forming, Nitrate-assimilating Pichia Species, Formerly Classified in the Genus Hansenula S ydow et S ydow , Based on the Partial Sequences of 18S and 26S Ribosomal RNAs (Saccharomycetaceae): The Proposals of Three New Genera, Ogataea , Kuraishia , and Nakazawaea. Biosci Biotechnol Biochem 58:1245–1257. https://doi.org/10.1271/bbb.58.1245
Yan Y, Zhang X, Zheng X, Apaliya MT, Yang Q, Zhao L, Gu X, Zhang H (2018) Control of postharvest blue mold decay in pears by Meyerozyma guilliermondii and it’s effects on the protein expression profile of pears. Postharvest Biol Technol 136:124–131. https://doi.org/10.1016/j.postharvbio.2017.10.016
Yang Z, Zhang Z (2018) Engineering strategies for enhanced production of protein and bio-products in Pichia pastoris: a review. Biotechnol Adv 36:182–195. https://doi.org/10.1016/j.biotechadv.2017.11.002
Yang SW, Park JB, Han NS, Ryu YW, Seo JH (1999) Production of erythritol from glucose by an osmophilic mutant of Candida magnolia. Biotechnol Lett 21:887–890
Yang L, He QS, Corscadden K, Udenigwe CC (2015) The prospects of Jerusalem artichoke in functional food ingredients and bioenergy production. Biotechnol Rep 5:77–88. https://doi.org/10.1016/j.btre.2014.12.004
Yarrow D (1998) Methods for the isolation, maintenance and identification of yeasts. In: The yeasts. Elsevier, Amsterdam, pp 77–100
Yurkov AM, Dlauchy D, Péter G (2017) Meyerozyma amylolytica sp. nov. from temperate deciduous trees and the transfer of five candida species to the genus meyerozyma. Int J Syst Evol Microbiol 67:3977–3981. https://doi.org/10.1099/ijsem.0.002232
Yuvadetkun P, Reungsang A, Boonmee M (2018) Comparison between free cells and immobilized cells of Candida shehatae in ethanol production from rice straw hydrolysate using repeated batch cultivation. Renew Energy 115:634–640. https://doi.org/10.1016/j.renene.2017.08.033
Zhang J, Geng A, Yao C, Lu Y, Li Q (2012) Xylitol production from d-xylose and horticultural waste hemicellulosic hydrolysate by a new isolate of Candida athensensis SB18. Bioresour Technol 105:134–141. https://doi.org/10.1016/j.biortech.2011.11.119
Zhang H-J, Fan X-G, Qiu X-L, Zhang Q-X, Wang W-Y, Li S-X, Deng L-H, Koffas MAG, Wei D-S, Yuan Q-P (2014) A novel cleaning process for industrial production of xylose in pilot scale from corncob by using screw-steam-explosive extruder. Bioprocess Biosyst Eng 37:2425–2436. https://doi.org/10.1007/s00449-014-1219-0
Zhang J, Zhang B, Wang D, Gao X, Sun L, Hong J (2015) Rapid ethanol production at elevated temperatures by engineered thermotolerant Kluyveromyces marxianus via the NADP(H)-preferring xylose reductase-xylitol dehydrogenase pathway. Metab Eng 31:140–152. https://doi.org/10.1016/j.ymben.2015.07.008
Zhang B, Zhang J, Wang D, Han R, Ding R, Gao X, Sun L, Hong J (2016) Simultaneous fermentation of glucose and xylose at elevated temperatures co-produces ethanol and xylitol through overexpression of a xylose-specific transporter in engineered Kluyveromyces marxianus. Bioresour Technol 216:227–237. https://doi.org/10.1016/j.biortech.2016.05.068
Zhao J, Mou Y, Shan T, Li Y, Zhou L, Wang M, Wang J (2010) Antimicrobial metabolites from the endophytic fungus pichia guilliermondii Isolated from Paris polyphylla var. yunnanensis. Molecules 15:7961–7970. https://doi.org/10.3390/molecules15117961
Zhao C, Gu D, Nambou K, Wei L, Chen J, Imanaka T, Hua Q (2015) Metabolome analysis and pathway abundance profiling of Yarrowia lipolytica cultivated on different carbon sources. J Biotechnol 206:42–51. https://doi.org/10.1016/j.jbiotec.2015.04.005
Zhou W-J, Yang J-K, Mao L, Miao L-H (2015) Codon optimization, promoter and expression system selection that achieved high-level production of Yarrowia lipolytica lipase in Pichia pastoris. Enzym Microb Technol 71:66–72. https://doi.org/10.1016/j.enzmictec.2014.10.007
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bergmann, J.C. et al. (2019). Biotechnological Application of Non-conventional Yeasts for Xylose Valorization. In: Sibirny, A. (eds) Non-conventional Yeasts: from Basic Research to Application. Springer, Cham. https://doi.org/10.1007/978-3-030-21110-3_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-21110-3_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-21109-7
Online ISBN: 978-3-030-21110-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)