Abstract
Serious antagonistic impacts of saline environment on plant growth, development, and yield are well established. In this regard, researchers and breeders have been utilizing many conventional as well as modern approaches to aid the process of developing salt-tolerant crops. Biotechnological tools have made the task of engineering salinity tolerance in plants easier. Currently, two major annexes are effectively employed to develop salt-tolerant crops, first, investigation of genetic variation via marker-assisted selection (MAS) and second the transgenic technology. Sustenance of plants under dynamically growth-limiting saline environment depends on alterations and/or switching between multiple biochemical pathways involved in response. A number of key regulatory genes have been successfully identified and characterized in this context which can be explored to serve the purpose of alleviation in salt-tolerant nature of plants. Several genomics-abetted approaches have been reported aiming toward improvement in growth and yield of crops under saline environment. Present chapter focuses on genomic roadmaps for augmentation of crop salt tolerance by various methods including MAS, transgenic breeding, manipulations in small non-coding RNAs, and genome editing. These approaches utilize key players involved in salinity-mediated plant defense mechanisms, such as ion transporters, osmolytes, antioxidants, transcription factors, signaling proteins, and microRNA. The chapter attempts to summarize the effective targets and exploration of these key entities to raise salt-tolerant plants through various genomics-related tools.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Marker assisted selection
- Ion transporters
- Osmolytes
- Antioxidants
- Transcription factors
- microRNA
- Transgenics
8.1 Introduction
Among the abiotic stresses, salinity stress is one of the most important environmental factors which considerably affect plant growth and productivity. Salinity affects about one third of the world’s irrigated land (Munns and Tester 2008), and it negatively influences water and nutrient homeostasis within living tissues. The deleterious effects on agricultural crops primarily include growth reduction and yield loss. In this context, both the conventional and modern crop improvement approaches are employed to facilitate development of novel genetic resources for use in direct or indirect breeding for improving salinity tolerance in crop plants. Currently, a wide range of mutational, biotechnological, and genomics-assisted tools are available which are more or less focused on gene discovery and boosting up the process of novel gene introduction or modification (Nongpiur et al. 2016).
Apparently, two main approaches are used to improve and impart salinity tolerance in crop plants. The first is through exploring natural genetic variation, either through selection under stress conditions or through quantitative trait loci (QTL) followed by marker-assisted selection (MAS). The other one is through transgenic technology by modifying the expression of endogenous genes or introducing novel genes (of plant or non-plant origin) to impart stress tolerance. Crop improvement via conventional breeding approaches has yielded limited success due to complexity of the trait since the process is time and labor intensive and requires well-characterized germplasm. In this regard, genetic engineering methods have become useful to develop transgenic crops tolerant to abiotic stresses (Yamaguchi and Blumwald 2005). The primary step before proceeding to make transgenics is the identification of functional and regulator genes serving to control different metabolic pathways, including ion homeostasis, antioxidant defense system, osmolyte synthesis, and other signaling pathways.
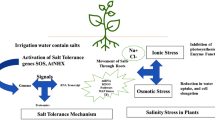
Salt stress increases ion toxicity and also affects uptake and movement of other essential nutrients such as potassium in the cell. This may occur either in a monophasic or biphasic manner depending on the duration and extent of exposure to saline conditions. A short exposure usually leads to osmotic or oxidative stress which would be followed by ionic stress upon long-term exposure (Munns and Tester 2008). To sustain under such dynamic growth-limiting situations, plants need to incur switching between multiple biochemical pathways that are much more complex when combined with other biotic and abiotic stresses. A general view of plant responses to salinity stress is presented in Fig. 8.1.
Significant progress has been made in the identification of genes involved in plant salt-stress responses (Hanin et al. 2016). Till date, a number of key genes involved in salinity tolerance have been isolated, characterized, and validated by using different transgenic methods. The candidate genes for salt tolerance are categorized into genes with functional and regulatory role (Shinozaki et al. 2003). The first group includes those involved in osmolyte biosynthesis, ion transporters, water channels, antioxidant systems, sugars, polyamines, heat shock proteins, and late embryogenesis abundant proteins. The second group are involved in the regulation of transcriptional and posttranscriptional machinery besides genes of signaling pathways. Some of these are transcription factors (TFs), protein kinases and phosphatases. In addition, there are several other strategies for attaining abiotic stress tolerance which are being tested for salt tolerance such as, using the stress-inducible promoters to avoid the pleiotropic effects (Checker et al. 2012), employing the protein post-translational modifications such as ubiquitination (Lyzenga and Stone 2012; Guo et al. 2008) and the use of halophyte gene resources.
Several state-of-the-art genomics-assisted approaches (Fig. 8.2), such as transgenic overexpression, RNAi, microRNA, genome editing, and genome-wide association studies, are being used for improving salt tolerance in crop plants (Mickelbart et al. 2015; Nongpiur et al. 2016). Overexpression of these genes has been shown as a successful strategy to improve plant tolerance to different abiotic stresses including salinity (Türkan and Demiral 2009; Cominelli et al. 2013; Hanin et al. 2016). In this article, we present an overview of the different genomics-based molecular genetic approaches (marker-assisted selection and transgenic breeding) that have contributed to improve the salt tolerance in crop plants.
8.2 Marker-Assisted Selection for Enhancement of Crop Salinity Stress Tolerance
Despite of availability of broad genetic resources, the slow progress in the genetic improvement for salt tolerance through conventional breeding is attributed to the complex nature of the trait accompanied with the high environmental influence and requirement of huge experimental fields (Flowers and Yeo 1997; Munns 2002; Thomson et al. 2010; Munns and Tester 2008). Further, phenotypic screening for salinity tolerance in the large sample population has remained a big challenge through the laborious conventional techniques (Mantri et al. 2014).
Marker-assisted selection (MAS) is a new precision breeding tool that allows the indirect and accurate selection for a desired trait from breeding population based on tightly linked molecular markers, viz., restriction fragment length polymorphisms (RFLP), amplified fragment length polymorphisms (AFLP), random amplified polymorphic DNA (RAPD), simple sequence repeats (SSR) or microsatellites, sequence-tagged microsatellite site (STMS), single nucleotide polymorphisms (SNPs), etc. It enables rapid and accurate screening for complex and polygenic traits which are difficult to score phenotypically through mapping or tagging of the trait linked quantitative trait loci (QTL). The successful applications of molecular marker-assisted breeding have proved its enhanced efficiency and accuracy in improved biotic and abiotic stress tolerance in rice and several other important crops (Singh et al. 2012; Ellur et al. 2016; Babu et al. 2017a, b). MAS offers advantage over the other genetic improvement tools as having relaxed biosafety regulations at development, field testing, commercial release, and import/export of the developed improved genotypes as well as their wider public acceptance.
Among field crops, rice being an important global staple food crop, considerable progress has been made for molecular breeding for improvement in tolerance to abiotic stresses such as salinity stress (Table 8.1). Through rigorous research on molecular breeding programs, the molecular marker maps for important agricultural crops have been constructed with varying density among the species. For development of molecular breeding tools, sources for abiotic stress tolerance have been identified through rigorous screening of genotypes in various crops. Ravikiran et al. (2017) identified two rice genotypes, CST 7–1 and Arvattelu as source for seedling-stage salinity tolerance, based on screening of 192 diverse genotypes under salinity stress (EC ∼ 12 dS m−1) using morphophysiological markers. Screening of the genotypes with SSR markers associated with Saltol region on chromosome 1 revealed RM 493 and RM 10793 as good candidates for marker-assisted selection of seedling-stage salinity tolerance. Linh et al. (2012) reported improved salt tolerance in high-yielding Bac Thom 7 rice cultivar through introgression of the Saltol QTL from donor parent FL478. The microsatellite markers, viz., RM493 and RM3412b tightly linked to the Saltol QTL, were used for foreground selection. The selected backcross lines displayed salt tolerance with agronomic performance similar to that of the original Bac Thom 7. The marker-assisted selection enabled rapid and efficient background (for the recurrent parent genome) and foreground (for target locus Saltol) selections in early generations with minimum linkage drag.
Another study by Babu et al. (2017a) reported use of marker-assisted backcrossing to transfer seedling-stage salt-stress tolerance by transferring a QTL, Saltol, into an elite salinity-sensitive rice cultivar Pusa Basmati 1121. RM 3412 STMS marker linked tightly to the QTL was used for indirect foreground selection. Back cross (BC) lines homozygous for the QTL were advanced to develop four improved near isogenic lines (NILs) of PB1121 with the salt tolerance. The field evaluation confirmed effect of QTL integration into the improved NILs in terms of greater salt tolerance at seedling stage and similar or better performance for other agronomic traits than the recurrent parent.
Recently, De Leon et al. (2017) utilized SSR and SNP markers to characterize introgression lines (ILs) of a high salinity-tolerant donor line Pokkali in an elite highly salt-sensitive rice cultivar Bengal and to further identify QTLs for traits contributing to salinity stress tolerance. As expected, because of abundance, more number of QTLs were detected using SNP markers than the SSR. The study emphasized marker-assisted breeding through introgression of salt injury score (SIS) QTLs, in addition to other major QTLs Saltol or qSKC1, for improved salinity tolerance. The identified tolerant ILs can be used as donor breeding lines for selective transfer of salinity tolerance without any linkage drag of undesirable traits from Pokkali to other recipient-sensitive varieties as well as for mapping and further positional cloning of the genes responsible for the trait.
Babu et al. (2017b) identified seedling-stage salt-tolerant indica landraces (Badami, Shah Pasand and Pechi Badam), Oryza rufipogon accessions (NKSWR2 and NKSWR17) and one each of Basmati rice (second Basmati) and japonica cultivars (Tompha Khau) based on phenotypic screening under hydroponics. The salt tolerance level was similar to that of high salt-tolerant genotypes Pokkali and FL478. Molecular diversity study of the diverse rice genotypes using polymorphic SSR markers linked with Saltol QTL revealed weak linkage disequilibrium-LD, suggesting its low usefulness in MAS, if the target foreground markers chosen are wide apart. LD mapping identified markers (RM10927, RM10871) linked with QTLs associated with salt tolerance traits. The study also identified Saltol marker, RM27, positioned on chromosome 10, associated with root Na/K ratio.
Efforts are also being made to identify novels QTLs for salinity tolerance from different sources in rice and other crops. The enhanced salt tolerance can be achieved through pyramiding of different novel QTLs in one genetic background through MAS. Bizimana et al. (2017) used Hasawi rice genotype, which conferred seedling-stage salinity tolerance due to novel QTLs other than Saltol, as a source to develop 300 recombinant inbred lines with high-yielding salt-sensitive cultivar-IR29. Further for identification of QTLs linked to salinity tolerance, a genetic linkage map was constructed using 194 polymorphic SNP markers. The study reported identification of 20 new QTLs on different chromosomes for salt tolerance through composite interval mapping.
In addition to rice, efforts are also being made for analysis of QTLs for breeding salt tolerance in other crops. In cotton, Zhao et al. (2016) identified salt-tolerant and salt-sensitive upland cotton cultivars through screening based on seedling emergence rates in response to 0.3% salt-NaCl. Seventy-four SSR markers were used to scan the genomes of these diverse cultivars, and eight markers associated with salt tolerance were identified through association analysis for further application in marker-assisted breeding. Similarly, Kere et al. (2017) screened salt-sensitive-11439S and salt-tolerant-11411S inbred parental lines with SSR markers to identify the QTLs for application in MAS for breeding salinity tolerance in cucumber. The analysis confirmed significant association of SSR markers with salt tolerance traits such as survival rate, relative leaf numbers, and percent green leaves, and salinity tolerance was evaluated by visual scoring. Recently, Luo et al. (2017) made efforts to map the critical QTLs contributing to salt tolerance in field-grown mature maize plants using a permanent doubled-haploid (DH) population and high-density SNP markers. Major QTLs responsible for salt tolerance and two candidate genes involving in ion homeostasis were mapped on chromosome 1. The mapped QTLs can be used in breeding salt-tolerant maize varieties through MAS.
Physiological and molecular studies on tolerance to various abiotic (ionic and/or osmotic stresses), viz., salinity, drought, etc., have revealed stress-specific as well as shared stress adaptation mechanisms, highlighting the complexity of stress response and adaptation in plants. In view of this, meta-QTL (MQTL) for tolerance to abiotic stresses including drought, salinity, and water logging through meta-analysis in barley has been recently projected (Zhang et al. 2017). The study conducted meta-analysis to detect and map the major QTL for drought, salinity, and water logging tolerance from different mapping populations on the barley physical map. Fine-mapped QTL for the stress tolerance were validated on MQTLs for further successful MAS in barley breeding.
8.3 Transgenic Breeding: Functional Genes Conferring Salinity Tolerance
8.3.1 Ion Transporters
In general, plants cannot withstand high salt concentration although the plant species differ in their mode of responses to the external salt exposure. Several distinct responses are generated in the plants to avoid high-salinity-induced harmful effects. One of the most distinguishing responses is avoiding salinity stress by compartmentation and the exclusion of detrimental ions like Na+ and Cl− from tissues that are very sensitive like the mesophyll and their relocation into the apoplast or vacuole (Sperling et al. 2014). Confinement of harmful ions within a root or apoplastic zone and maintainance of high K+/Na+ ratio are the major tolerance strategies for salt tolerance (Shabala and Cuin 2008).
Ion transporters are key players in maintaining ion homeostasis and in salt detoxification processes (Serrano et al. 1999; Hasegawa 2013). Various salts are present in soil out of which sodium chloride (NaCl) is the most significant. The Na+/H+ antiporter predominantly transports Na+ ion from cytoplasm to the vacuole. Therefore, overexpression of genes that are involved in Na+ transport was studied to a great extent with considerable success. Vacuolar Na+/H+ antiporter (AtNHX1) from the Arabidopsis was among the first and most studied gene. In Arabidopsis salt tolerance was conferred by overexpressing vacuolar Na+/H+ antiporter (Apse et al. 1999). Following with this initial success, many events were reported where transgenic plants exhibited higher potential for vacuolar sequestration of Na+ that subsequently avoid its harmful buildup into the cytoplasm. For example, overexpression of AtNHX1 and other NHX proteins from various hosts in many other plant species like tomato, B. napus, wheat, and cotton has been shown to increase salt tolerance (Zhang and Blumwald 2001; Zhang et al. 2001; Xue et al. 2004; He et al. 2005; Munns and Tester 2008). Vacuolar-type H+-ATPase and the vacuolar pyrophosphatase are the two types of H+ pumps that are present in vacuolar membrane (Dietz et al. 2001; Otoch et al. 2001; Wang et al. 2001). Overexpression of genes from wheat (Triticum aestivum) TaNHX1 and H+-pyrophosphatase (TVP1) resulted in improved salinity stress tolerance in Arabidopsis (Brini et al. 2007a). Similarly improved tolerance to salt stress was found in tobacco (Gouiaa et al. 2012) and tomato (Gouiaa and Khoudi 2015) by overexpression of Na(+)/H(+) antiporter H(+)- pyrophosphatase. On the other hand, the HKT gene family has a major role in preventing Na+ ion toxicity in shoots by root-to-shoot partitioning of Na+. The significant role of HKTs in Na+ transport in plants makes them promising candidates to enhance salinity tolerance. The identification of the wheat HKT1 (TaHKT2;1) gene (Schachtman and Schroeder 1994; Rubio et al. 1995) has steered the study of many HKT genes from several other crops (Horie et al. 2009). Moller et al. (2009) shown that targeted overexpression of AtHKT1;1 in the stele enhances salt tolerance in A. thaliana. Lately, Do et al. (2016) showed a close syndicate between the higher expression of the Ncl gene (homologous to the Na+/H+ antiporter gene family) in the root, the lesser buildup of Na+, K+, and Cl− in the shoot under salt stress. Overexpression of Ncl into a Japanese soybean cultivar Kariyutaka resulted in enhanced salt tolerance (Do et al. 2016).
Growing evidence was found on the role of salt overly sensitive (SOS) stress signaling pathway in ion homeostasis and salinity tolerance (Sanders 2000; Hasegawa et al. 2000). Most of the SOS signaling pathway was reported to be involved in exportation of Na+ out of the cell. The SOS signaling pathway includes three main proteins, namely, SOS1, SOS2, and SOS3. Plasma membrane-localized Na+/H+ antiporter (SOS1) is also known as NHX7 (Qiu et al. 2002). SOS2 (serine/threonine kinase) activated by salt stress elicited Ca+ signals, while SOS3 protein is a myristoylated Ca+ binding protein (Liu et al. 2000; Ishitani et al. 2000). Arabidopsis plants overexpressing genes for SOS1, SOS3, AtNHX1 + SOS3, SOS2 + SOS3, or SOS1 + SOS2 + SOS3 resulted in improved tolerance to salt stress (Yang et al. 2009). Similarly, Kumar et al. (2009) have shown that salinity stress tolerance in Brassica is correlated with transcript abundance of the genes related in SOS pathway.
8.3.2 Osmolytes
Osmolytes are small organic compounds having a protective role. Osmolytes are important for two functional roles: osmotic adjustment at high concentrations; and it plays unknown protective role at lower concentrations. Under salt-stress conditions, plant cell accumulates various osmolytes along with Na+ exclusion from the cytoplasm, to counter the osmotic pressure of harmful ions in vacuoles. Osmolytes like proline, glycine betaine, and sucrose accumulating upon salt stress in many plant species including halophytes are well studied and characterized (Flowers et al. 1977). Table 8.2 presents some of the successful examples of transgenic plants developed using different osmolyte genes.
Hu et al. (2015) found experimental evidences for the accumulation of sugars and amino acids. Particularly, sucrose and trehalose sugar and amino acids like proline, valine, glutamate, asparagine, glutamine, phenylalanine, and lysine accumulated under salt-stress conditions. Also, sugars like sucrose and pinitol are accumulated more in leaves, while starch accumulated in roots under salinity stress conditions. It has been found that these sugars (pinitol and sucrose) and starch can also increase in nodules under salt stress (Bertrand et al. 2015). Similarly, Boriboonkaset et al. (2013) found enrichment of soluble starch and soluble sugar in flag leaf of Pokkali genotype (salt tolerant) of rice which may have alternative role in osmotic adjustment in salt defense mechanism. In tomato plants, jasmonic acid and nitric oxide when applied exogenously, either individually or in combination, helped to boost the proline, flavonoid, and glycine betaine synthesis under NaCl salt treatments (Ahmad et al. 2018).
Engineering plants by overexpressing the osmolytes was considered as one of the ways to enhance salt tolerance in plants. In Arabidopsis, knockout of the P5CS1, a key gene in proline biosynthesis, which encodes a delta1-pyrroline-5-carboxylate synthetase (P5CS), impairs proline synthesis resulting in salt hypersensitivity (Székely et al. 2008). P5CS transformed in tobacco and rice has shown increased proline production, linked with increased salt-stress tolerance (Kishor et al. 1995; Su and Wu 2004). Also, transgenic rice expressing the moth bean P5CS gene showed enhanced tolerance to higher dose of NaCl (Su and Wu 2004). Recently, mutated P5CS (P5CSF129A) gene was overexpressed in Sorghum and found that transgenic plants accumulated more proline and showed salt-stress tolerance. Moreover overproduction of proline through transfer of a P5CSF129A gene conferred protection of photosynthetic and antioxidant enzyme activities (Reddy et al. 2015). Glycine betaine is yet another important osmolyte that helps to balance the osmotic potential of intracellular ions under salinity. Under high salinity, glycine betaine accumulation increased in lamina leaves and bladder hairs of Atriplex gmelini (Tsutsumi et al. 2015). Overexpressing choline oxidase in rice plant showed increased levels of glycine betaine and improved tolerance to salt and cold stress (Sakamoto et al. 1998). It was found that transgenic Arabidopsis and tobacco plants transformed with bacterial mtlD gene which encodes for mannitol-1-phosphate dehydrogenase conferred salt tolerance and thereby maintained normal growth and development under high salt-stress growth conditions (Binzel et al. 1998; Thomas et al. 1995). Ectopic expression of bacterial gene mannitol-1-phosphate dehydrogenase (mt1D), an enzyme involved in mannitol biosynthesis, in tobacco successfully enhanced salt tolerance (Tarczynski et al. 1992). Genes for trehalose biosynthesis have also been employed in improving salt tolerance by developing transgenic plants for overproduction of trehalose (Penna 2003; Turan et al. 2012). Garg et al. (2002) demonstrated tolerance to salt and drought stress in rice by using tissue-specific or stress-inducible expression of a bifunctional trehalose-6-phosphate synthase/phosphatase (TPSP) fusion gene (comprising the E. coli trehalose biosynthetic genes). Li et al. (2011) reported that transgenic plants overexpressing rice trehalose-6-phosphate synthase (OsTPS1) showed improved salinity tolerance without much alteration in plant phenotype. It is also suggested that stress-inducible solute accumulation by using stress-specific or stress-inducible promoters may be better to achieve salt-specific expression of genes for osmotic adjustment.
8.3.3 Antioxidants and Protective Proteins
Abiotic stress causes the accumulation of reactive oxygen species (ROS) that can damage sensitive plant tissues during high salt stress by disturbing cell wall, enzymes, and membrane functions. Antioxidant enzymes and nonenzymatic compounds play a crucial role in detoxifying salinity stress-induced ROS. Salt-stress tolerance is positively correlated with antioxidant enzyme activity and with the accumulation of nonenzymatic antioxidant compounds (Gupta et al. 2005). Antioxidants include superoxide dismutase (SOD), catalase (CAT), glutathione peroxidase (GPX), ascorbate peroxidase (APX), glutathione reductase (GR), etc. Hence, overexpressing ROS-scavenging enzymes is shown to induce salinity tolerance in plants (Roxas et al. 1997; Badawi et al. 2004; Miller et al. 2008). Overexpression of ascorbate peroxidase (APX) in tobacco chloroplasts enhances the tolerance to salinity and drought stress (Badawi et al. 2004). The alternate oxidase (AOX) pathway plays a role under stress conditions. Smith et al. (2009) constitutively overexpressed an AOX1a gene in Arabidopsis plants and demonstrated superior salt tolerance than wild-type plants suggesting that genes of the AOX pathway can be useful to improve tolerance to stressful environmental conditions including salinity.
Other proteins like polyamines, osmotin, and LEA proteins mitigate salt stress by protecting macromolecules like nucleic acids, proteins, and carbohydrates from damages caused by ion toxicity and by drought conditions. Polyamines play a critical role in salinity and other abiotic stress tolerance by increasing level of polyamines which shows positive correlation of increased level of polyamines with stress tolerance in plants (Yang et al. 2007; Groppa and Benavides 2008; Gupta et al. 2013). Overproduction of spermidine and spermine in rice enhances salt tolerance (Roy and Wu 2002). Xu et al. (1996) found that HVA7, a LEA from barley, when transferred to rice, confers tolerance to drought and salinity stress. Dehydrins, another LEA protein, were shown to enhance plant tolerance to various stresses (Hanin et al. 2011). Brini et al. (2007b) found positive correlation between wheat dehydrin DHN-5 and salt tolerance. The study showed that the expression of the wheat dehydrin DHN-5 in Arabidopsis led to an increase in tolerance to salt and osmotic stresses. Some of the transgenics where functional genes overexpressed to impart salt tolerance in different plant species have been shown in Table 8.2.
8.4 Transgenic Breeding: Regulatory Genes Controlling Salinity Stress Responses
8.4.1 Transcription Factors
Plant response to salt stress is a complex process and involves a vast array of genes working in different or overlapping regulatory pathways. As stress response is complex process and regulated by multi-genes, it is very challenging to achieve success in improving plant stress tolerance with the single functional gene approach (Mittler and Blumwald 2010; Varshney et al. 2011). Thus, instead of manipulating single functional gene, engineering regulatory genes or master regulators can be potential strategy for controlling stress responses. Transcription factors (TFs) and signaling proteins are master regulators of many genes involved in stress responses; hence, they are possible candidates for genetic engineering to obtain salinity-tolerant crops (Table 8.3). Transcription factors from various families like AP2/ERF, NAC, MYB, MYC, DREB, Cys2/His2 zinc finger, bZIP, and WRKY have been reported to be involved salt-stress tolerance (Golldack et al. 2011, 2014).
NAC TF family found to be involved in abiotic stress responses along with other important functions in plants (Nakashima et al. 2012). Hu et al. (2008) reported that overexpression of SNAC1 and SNAC2 genes from NAC TF family helps the survival of transgenic Oryza sativa plants under high-salinity conditions. Similarly, overexpression of TF gene OsNAC04 leads to drought and salinity stress tolerance in O. sativa (Zheng et al. 2009). The AP2/ERF another family of plant-specific TFs which is known to play key role against various abiotic stresses (Mizoi et al. 2012). The transgenic plants for gene GmDREB2 from soybean showed enhanced salinity and drought tolerance (Chen et al. 2007). Overexpression Oryza sativa MYB2 (TF from MYB family) exhibited salt tolerance by variation in expression levels of various stress responsive genes (Yang et al. 2012).
Several groups reported a key role of WRKY TFs in responses to various abiotic stresses including salinity stress (Banerjee and Roychoudhury 2015). Li et al. (2013) reported ZmWRKY33 (WRKY TF family) enhanced tolerance to salinity stress in Arabidopsis while overexpressing GmWRKY54 exhibited salt tolerance, probably through the regulation of another TF DREB2A and STZ/Zat10 (Zhou et al. 2008). bZIP is also another family of TFs having an important role in response to various abiotic stresses including salinity stress (Jakoby et al. 2002). Wang et al. (2010) demonstrated that overexpression of ThbZIP1 gene of Tamarix hispida from TF bZIP contributes to salinity tolerance by enhancing the activity of antioxidant enzymes such as peroxidase and superoxide dismutase and by accumulating compatible osmolytes like soluble sugars and soluble proteins. SlAREB1 and SlAREB2 are two members of bZIP TF in Solanum lycopersicum. Transgenic tomato overexpressing SlAREB1 plants showed improved salinity and drought tolerance (Orellana et al. 2010).
8.4.2 Signaling Proteins
In addition to TFs, genetic engineering of signaling proteins has also become one of the feasible approaches. Some of the examples of transgenics where regulatory genes are overexpressed to impart salt tolerance are presented in Table 8.3. Several studies have reported that abiotic stresses (cold, high salt, and drought) trigger rapid increase in plant cells calcium (Ca2+) levels (Sanders et al. 2002). Calcium signaling is often coupled with protein phosphorylation and dephosphorylation mediated by protein kinases and phosphatases, respectively. Changes in cellular Ca2+ level are being mediated by different Ca2+ binding proteins like calmodulin (CaM) and CaM-related proteins (CML), calcium-dependent protein kinases (CDPKs), and calcineurin B-like proteins (CBL) (Bouché et al. 2005). CDPK is one of the best studied protein kinases in the Ca2+ signaling pathway. Another family of protein kinases that function in stress tolerance is mitogen-activated protein kinases (MAPKs) (Zhang and Klessig 2001).
Overexpression of ShCML44, cold-responsive calmodulin-like gene in tomato, showed enhanced tolerance to salinity stress with higher germination rate and better growth of seedling (Munir et al. 2016). Similarly, transgenic rice overexpressing OsMSR2, a novel calmodulin-like gene, enhanced salinity tolerance with altered expression pattern of genes related to stress (Xu et al. 2013). Transgenic rice plant expressing calcineurin A subunit from mouse exhibited a higher level of salinity stress tolerance; also it has been observed that Na+ content is higher in roots of untransformed wild-type plants than that of transgenic roots (Ma et al. 2005).
The OsCPK4 gene is a member of calcium-dependent protein kinases in rice. Recently Campo et al. (2014) showed that transgenic rice plants overexpressing of OsCPK4 significantly enhances salt and drought tolerance. Mitogen-activated protein kinase (MAPK) cascades also play crucial regulatory roles in various stress responses other than plant development processes. Zhang et al. (2011) reported ectopic expression of cotton GhMPK2 in Nicotiana tabacum and found elevated levels of proline and induced expression of several genes related to stress, and as a result transgenic Nicotiana tabacum exhibited enhanced drought and salt tolerance. Similarly, overexpression MAPK from rice (OsMAPK5) exhibited increased kinase activity along with increased tolerance for salinity and other abiotic stresses like drought and cold (Xiong and Yang 2003). Overexpression of mitogen-activated protein kinase kinase 5 (MKK5) in Arabidopsis wild-type plants improved their tolerance level against various salt treatments (Xing et al. 2015). In another study overexpression of mitogen-activated protein kinase kinase 4 from Populus trichocarpa (PtMAPKK4) shows improved salt tolerance in tobacco. Specifically, under salt-stress condition, PtMAPKK4 overexpressing lines showed improved germination and growth and development (Yang et al. 2017). However, some MAPKs can also have contrary effects particularly in case of rice; overexpression of OsMAPK33 caused increased sensitivity to salinity and drought stress compared to wild-type plants (Lee et al. 2011). Receptor-like kinases (RLKs), another type of kinase, also have an important role in stress responses. Overexpressing OsSIK1 (OsSIK1-ox), one of the putative RLKs, showed greater tolerance to salt stress as compared to control plants, gene-silenced plants by RNA interference (RNAi), knockout mutants sik1 in rice (Ouyang et al. 2010). Sucrose non-fermenting 1 (SNF1)-related protein kinases (SnRKs) is one type of well-characterized protein kinase involved in stress responses (Halford and Hey 2009). In one study PtSnRK2.5 and PtSnRK2.7genes (SnRKs from Poplar) heterologously overexpressed in Arabidopsis and found that overexpression of PtSnRK2 leads to enhanced tolerance level for salt stress.
8.4.3 Manipulating the miRNAs
In the past decade, miRNAs have become major players in the regulation of plant response to environmental abiotic stresses (Zhang 2015, Shriram et al. 2016). There has been great interest in exploring regulatory roles of microRNAs against different stresses including salt for their exploitation in genetic engineering for higher stress tolerance, biomass, and yield (Zhang and Wang 2016, Patel et al. 2018). Transgenic plants overexpressing miR319 showed significantly higher plant tolerance to drought and salinity stress in creeping bentgrass (Agrostis stolonifera) (Zhou et al. 2013). In another study, expression of miR408 was shown to improve higher tolerance to salinity, cold, and oxidative stress in Arabidopsis seedlings (Ma et al. 2015). A rice microRNA osa-miR393 was overexpressed in Arabidopsis plant resulting in enhanced salt tolerance (Gao et al. 2011a; b). In several other studies, novel microRNAs have been identified suggesting that these well-characterized candidates could become targets for plant genetic engineering investigations as successful in silico predictions could result in finding the target genes involved in pathways of signaling, ion homeostasis besides sustained plant growth under salt stress.
8.5 Halophyte Genes for Improving Salt-Stress Tolerance of Crops
In plants, gene expression and regulation decides the fate of plants from growth and development to stress tolerance. Modification/ manipulation in the regulation of these entities can dramatically change the fate of plant’s life. In this sense, stress tolerance of plants can be improved by manipulating particular genes. In terms of stress tolerance, it is proved that the tolerant and sensitive plants possess same set of genes, but their efficient regulation or subtle changes in gene sequence can make one plant sensitive and other plant tolerant to the same environmental condition. This phenomenon is also true for salt-sensitive glycophytes and salt-tolerant halophytic plants. The halophytes are naturally tolerant to high salinity. Their genetic analysis revealed that differences in promoter activities and gene duplication in halophytes as compared to their glycophytic relative is responsible for their high salt tolerance (Nikalje et al. 2017). For example, the NHX8 showed stress-induced expression in Arabidopsis while in Thellungiella, it showed constitutive expression. However Arabidopsis possess a single copy of CBL10, while T. parvula contain three copies; such changes make Thellungiella more salt tolerant. In addition, the efficient post-translational modifications are highly efficient in halophytes (Bose et al. 2015), and the halophytic gene sequences are more complex with presence of extra transposons and intergenic sequences (Rui et al. 2007). Therefore, for genetic improvement of crops, it may be important to choose genes from halophytic origin. Overexpression of NHX1 gene of Aeluropus littoralis in soybean resulted in less sodium accumulation in aerial parts than underground parts, increased potassium ion content under salt stress and increased salt tolerance up to 150 mM NaCl (Liu et al. 2014). Shabala and Potossin (2014) opined that retention of potassium ions under salt stress is a key factor for salt tolerance in plants and specialty of halophytes. Further, Bose et al. (2015) have confirmed this by showing that maintenance of negative water potential because of high H+ ATPase activity is important for halotropism. Similarly different halophytic antiporters were overexpressed in rice, and the transgenic plants showed high salt tolerance. PtNHA1 and PtNHX from Puccinellia tenuiflora were transformed into rice, and the resulting transgenic rice showed improved tolerance to NaCl and NaHCO3. Transgenic rice harboring AgNHX1 from Atriplex gmelinii increased vacuolar antiporter activity by almost eightfold and improved its tolerance up to 300 mM NaCl (Ohta et al. 2002).
8.6 New Research on the Salt Pan
Genome editing tools have opened up new avenues for specific and targeted modifications in the crop plants (de Wiel et al. 2017). The method enables the introduction of targeted precise genomic changes using customized nucleases (Jain 2015). Genes associated with salt tolerance such as those involved in signaling, ion homeostasis, osmolyte synthesis, and transporters can be the suitable candidates for editing based manipulation. The plasma membrane ATPase plays a critical role in the regulation of ion homeostasis under salt stress and hence has been used as the target gene in a recent study. Osakabe et al. (2016) induced mutation of an abiotic stress tolerance gene encoding OPEN STOMATA 2 (OST2) (AHA1) – a major plasma membrane H + -ATPase via the precise site modification by using truncated gRNAs (tru-gRNAs) in the CRISPR-Cas9 system (Table 8.3).
High-throughput screening methods have advanced our knowledge about the genomes and phenomes. Plant stress biology research depends on robust screening methods for contrasting salt-stress-responsive phenotypes at different levels of tissue, organ, and whole-plant level. This branch of research, plant phenomics, is now being applied to facilitate efficient and reliable evaluation of stress (and salt) tolerant lines. Several such platforms for phenotyping are now available. Some of these include the High Resolution Plant Phenomics Centre (http://www.plantphenomics.org.au/HRPPC), Plant AccelatorTM (http://www.plantaccelerator.org.au/), Jülich Plant Phenotyping Centre- JPPC (http://www.fz-juelich.de/ibg/ibg-2/EN/_organisation/JPPC/JPPC_node.html) and Deep Plant Phenomics (Ubbens and Stavness 2017). Campbell et al. (2015) have developed a novel approach to analyze the dynamic plant responses to salt stress and studied the genetic basis of salt stress associated, genetically determined changes using a longitudinal genome-wide association model. This study highlights the use of image-based phenomics platforms combined with genome-wide association studies (GWAS) for dissecting the plant stress responses and should enable to establish liaison between expressed phenotypes with related genomic regions and environmental conditions. Further research into plant genetic manipulation via precise genetic tools will benefit from efficient phenotyping screens and high-throughput analysis tools.
8.7 Conclusions
Increasing salinity severely affects crop productivity and is becoming threat to world agriculture. The development of several genomics-assisted approaches including genetically modified plants has been advocated to circumvent this problem. Toward this goal, several stress-responsive genes have been identified and successfully introduced into other crops to create transgenic crops with enhanced stress tolerance. The most impressive results were obtained when manipulating transcription and signaling factors, as they control a broad range of downstream events, which results in superior tolerance to multiple stresses. However, challenges still lie ahead before successfully improving crop yield under saline conditions as most methods have been limited by the problem of yield penalty. Salinity tolerance involves a complex of responses at molecular, cellular, metabolic, physiological, and whole-plant levels. The marker-assisted selection as the molecular breeding method has begun to deliver its expected benefits in commercial breeding programs for salinity stress tolerance. For this, in addition to the key loci identified for salt tolerance traits majorly in rice, emphasis should also be given on identification and validation of other new loci in rice and other crops and their pyramiding in elite genetic background for enhanced salt tolerance through molecular marker-assisted breeding. Generation of salinity-tolerant transgenic varieties should necessarily involve gene stacking where multiple genes need to be overexpressed using advanced genetic engineering tools. Furthermore, the critical step is the field trials required to evaluate the transgenic plants, especially focusing on their growth and tolerance in the whole life period. New and novel information is generated through omics methods such as metabolomics and proteomics, and it is expected to develop more understanding of the salt-stress responses. It is also equally important that further understanding how plants perceive stress signals (salt sensors, osmosensors), transmit, and trigger a cascade of genetic mechanisms is necessary to develop crop plants that can tolerate extreme environments. With the current renewed interest in stress genomics, fast-forward approaches of phenomics, allele mining, and stress-metabolite profiling, it is expected to gain thorough understanding of salt-adaptive diversity for use in crop breeding for salt tolerance. Continued research should be aimed at development of salt-tolerant crop germplasm to expand the utilization of saline soils for enhancing agricultural productivity and environmental sustainability.
Abbreviations
- AFLP:
-
Amplified fragment length polymorphisms
- AOX:
-
Alternate oxidase
- APX:
-
Ascorbate peroxidase
- AtNHX1:
-
Na+/H+ antiporter
- CaM:
-
calmodulin
- CAT:
-
Catalase
- CBL:
-
Calcineurin B-like proteins
- CDPKs:
-
Calcium-dependent protein kinases
- CML:
-
CaM-related proteins
- GPX:
-
Glutathione peroxidase
- ILs:
-
Introgression lines
- MAS:
-
Marker-assisted selection
- MQTL:
-
Meta-QTL
- mt1D:
-
Mannitol-1-phosphate dehydrogenase
- P5CS:
-
delta1-pyrroline-5-carboxylate synthetase
- QTL:
-
Quantitative trait loci
- RAPD:
-
Random amplified polymorphic DNA
- RFLP:
-
Restriction fragment length polymorphisms
- RNAi:
-
RNA interference
- SNPs:
-
Single nucleotide polymorphisms
- SOD:
-
Superoxide dismutase
- SOS:
-
Salt overly sensitive
- SSR:
-
Simple sequence repeats
- STMS:
-
Sequence-tagged microsatellite site
- TFs:
-
Transcription factors
- TPSP:
-
Trehalose-6-phosphate synthase/phosphatase
References
Abebe T, Guenzi AC, Martin B, Cushman JC (2003) Tolerance of mannitol-accumulating transgenic wheat to water stress and salinity. Plant Physiol 131:1748–1755
Ahmad P, Ahanger MA, Alyemeni MN, Wijaya L, Alam P, Ashraf M (2018) Mitigation of sodium chloride toxicity in Solanum lycopersicum L. by supplementation of jasmonic acid and nitric oxide. J Plant Inter 13(1):64–72
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by over expression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285:1256–1258
Augustine SM, Ashwin Narayan J, Syamaladevi DP, Appunu C, Chakravarthi M, Ravichandran V, Tuteja N, Subramonian N (2015) Overexpression of EaDREB2 and pyramiding of EaDREB2 with the pea DNA helicase gene (PDH45) enhance drought and salinity tolerance in sugarcane (Saccharum spp. hybrid). Plant Cell Rep 34:247–263
Babu NN, Krishnan SG, Vinod KK, Krishnamurthy SL, Singh VK, Singh MP, Singh R, Ellur RK, Rai V, Bollinedi H, Bhowmick PK, Yadav AK, Nagarajan M, Singh NK, Prabhu KV, Singh AK (2017a) Marker Aided Incorporation of Saltol, a Major QTL Associated with Seedling Stage Salt Tolerance, into Oryza sativa ‘Pusa Basmati 1121’. Front Plant Sci 8:41
Babu NN, Vinod KK, Krishnamurthy SL, Gopala Krishnan S, Yadav A, Bhowmick PK, Nagarajan M, Singh NK, Prabhu KV, Singh AK (2017b) Microsatellite based linkage disequilibrium analyses reveal Saltol haplotype fragmentation and identify novel QTLs for seedling stage salinity tolerance in rice (Oryza sativa L.). J Plant Biochem Biotechnol 26(3):310–320
Badawi GH, Kawano N, Yamauchi Y, Shimada E, Sasaki R, Kubo A, Tanaka K (2004) Over-expression of ascorbate peroxidase in tobacco chloroplasts enhances the tolerance to salt stress and water deficit. Physiol Plant 121:231–238
Banerjee A, Roychoudhury A (2015) WRKY proteins: signaling and regulation of expression during abiotic stress responses. Sci World J 2015:807560
Bertrand A, Dhont C, Bipfubusa M, Chalifour FP, Drouin P, Beauchamp CJ (2015) Improving salt stress responses of the symbiosis in alfalfa using salt-tolerant cultivar and rhizobial strain. Appl Soil Eco 87:108–117
Binzel ML, Hess FD, Bressan RA, Hasegawa PM (1998) Intracellular compartmentation of ions in salt adapted tobacco cells. Plant Physiol 86:607–614
Bizimana JB, Luzi-Kihupi A, Murori RW, Singh RK (2017) Identification of quantitative trait loci for salinity tolerance in rice (Oryza sativa L.) using R29/Hasawi mapping population. J Genet 96(4):571–582
Boriboonkaset T, Theerawitaya C, Yamada N, Pichakum A, Supaibulwatana K, Cha-Um S, Takabe T, Kirdmanee C (2013) Regulation of some carbohydrate metabolism-related genes, starch and soluble sugar contents, photosynthetic activities and yield attributes of two contrasting rice genotypes subjected to salt stress. Protoplasma 250:1157–1167
Bose J, Rodrigo-Moreno A, Lai D, Xie Y, Shen W, Shabala S (2015) Rapid regulation of the plasma membrane H+-ATPase activity is essential to salinity tolerance in two halophyte species, Atriplex lentiformis and Chenopodium quinoa. Annals of Botany 115(3):481–494
Bouaziz D, Pirrello J, Charfeddine M, Hammami A, Jbir R, Dhieb A, Bouzayen M, Gargouri-Bouzid R (2013) Overexpression of StDREB1 transcription factor increases tolerance to salt in transgenic potato plants. Mol Biotechnol 54:803–817
Bouché N, Yellin A, Snedden WA, Fromm H (2005) Plant-specific calmodulin-binding proteins. Annu Rev Plant Biol 56:435–466
Brini F, Hanin M, Mezghani I, Berkowitz GA, Masmoudi K (2007a) Overexpression of wheat Na+/H+ antiporter TNHX1 and H+-pyrophosphatase TVP1 improve salt-and drought stress tolerance in Arabidopsis thaliana plants. J Exp Bot 58:301–308
Brini F, Hanin M, Lumbreras V, Amara I, Khoudi H, Hassairi A, Pagès M, Masmoudi K (2007b) Overexpression of wheat dehydrin DHN-5 enhances tolerance to salt and osmotic stress in Arabidopsis thaliana. Plant Cell Rep 26:2017–2026
Cai R, Zhao Y, Wang Y, Lin Y, Peng X, Li Q, Chang Y, Jiang H, Xiang Y, Cheng B (2014) Overexpression of a maize WRKY58 gene enhances drought and salt tolerance in transgenic rice. Plant Cell Tissue Organ Cult 119:565–577
Campbell MT, Knecht AC, Berger B, Brien CJ, Wang D, Walia H (2015) Integrating image-based phenomics and association analysis to dissect the genetic architecture of temporal salinity responses in rice. Plant Physiol 168(4):1476–1489
Campo S, Baldrich P, Messeguer J, Lalanne E, Coca M, Segundo BS (2014) Overexpression of a calcium-dependent protein kinase confers salt and drought tolerance in Rice by preventing membrane lipid peroxidation. Plant Physiol 165(2):688–704
Checker VG, Chhibbar AK, Khurana P (2012) Stress-inducible expression of barley Hva1 gene in transgenic mulberry displays enhanced tolerance against drought, salinity and cold stress. Transgenic Res 21:939–957
Chen M, Wang QY, Cheng XG, Xu ZS, Li LC, Ye XG, Xia LQ, Ma YZ (2007) GmDREB2, a soybean DRE-binding transcription factor, conferred drought and high salt tolerance in transgenic plants. Biochem Biophys Res Commun 353:299–305
Chen JB, Yang JW, Zhang ZY, Feng XF, Wang SM (2013) Two P5CS genes from common bean exhibiting different tolerance to salt stress in transgenic Arabidopsis. J Genet 92(3):461–469
Chen X, Wang Y, Lv B, Li J, Luo L, Lu S, Zhang X, Ma H, Ming F (2014) The NAC family transcription factor OsNAP confers abiotic stress response through the ABA pathway. Plant Cell Physiol 55:604–619
Cheng L, Li S, Hussain J, Xu X, Yin J, Zhang Y, Chen X, Li L (2013) Isolation and functional characterization of a salt responsive transcriptional factor, LrbZIP from lotus root (Nelumbo nucifera Gaertn). Mol Biol Rep 40:4033–4045
Choi JY, Seo YS, Kim SJ, Kim WT, Shin JS (2011) Constitutive expression of CaXTH3, a hot pepper xyloglucan endotransglucosylase/hydrolase, enhanced tolerance to salt and drought stresses without phenotypic defects in tomato plants (Solanum lycopersicum cv. Dotaerang). Plant Cell Rep 30(5):867–877
Cominelli E, Conti L, Tonelli C, Galbiati M (2013) Challenges and perspectives to improve crop drought and salinity tolerance. Nat Biotechnol 30:355–361
De Leon TB, Linscombe S, Subudhi PK (2017) Identification and validation of QTLs for seedling salinity tolerance in introgression lines of a salt tolerant rice landrace ‘Pokkali’. PLoS One 12(4):e0175361
Dietz KJ, Tavakoli N, Kluge C, Mimura T, Sharma SS, Harris GC, Chardonnens AN, Golldack D (2001) Significance of the V type ATPase for the adaptation to stressful growth conditions and its regulation on the molecular and biochemical level. J Exp Bot 52(363):1969–1980
Do TD, Chen H, Vu HTT, Hamwieh A, Yamada T, Sato T, Yan Y, Cong H, Shono M, Suenaga K, Xu D (2016) Ncl synchronously regulates Na+, K+, andCl-in soybean greatly increases the grain yield in saline field conditions. Sci Rep 6:19147
Ellur RK, Khanna A, Yadav A, Pathania S, Rajashekara H, Singh VK et al (2016) Improvement of Basmati rice varieties for resistance to blast and bacterial blight diseases using marker assisted backcross breeding. Plant Sci 242:330–341
Faize M, Burgo SL, Faize L, Piqueras A, Nicolas E, Barba-Espin G, Clemente-Moreno MJ, Alcobendas R, Artlip T, Hernandez JA (2011) Involvement of cytosolic ascorbate peroxidase and Cu/Zn-superoxide dismutase for improved tolerance against drought stress. J Exp Bot 62:2599–2613
Flowers TJ, Yeo AR (1997) Breeding for salt resistance in plants. In: Jaiwal PK, Singh PR, Gulaati A (eds) Strategies for improving salt tolerance in higher plants. Science Publishers Inc, Enfield, pp 247–264
Flowers TJ, Troke PF, Yeo AR (1977) The mechanism of salt tolerance in halophytes. Ann Rev Plant Physiol 28:89–121
Gao P, Bai X, Yang L, Lv D, Pan X, Li Y (2011a) Osa-MIR393: a salinity and alkaline stress-related microRNA gene. Mol Biol Rep 38:237–242
Gao SQ, Chen M, Xu ZS, Zhao CP, Li L, Xu HJ, Tang YM, Zhao X, Ma YZ (2011b) The soybean GmbZIP1 transcription factor enhances multiple abiotic stress tolerances in transgenic plants. Plant Mol Biol 75:537–553
Garg AK, Kim J-K, Owens TG, Ranwala AP, Choi YD, Kochian LV, Wu R (2002) Trehalose accumulation in rice plants confers high tolerance levels to different abiotic stresses. Proc Natl Acad Sci U S A 99:15898–15903
Goel D, Singh AK, Yadav V, Babbar SB, Bansal KC (2010) Overexpression of osmotin gene confers tolerance to salt and drought stresses in transgenic tomato (Solanum lycopersicum L.). Protoplasma 245:133–141
Golldack D, Luking I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30:1383–1391
Golldack D, Li C, Mohan H, Probst N (2014) Tolerance to drought and salt stress in plants: unraveling the signaling networks. Front Plant Sci 5:151
Gouiaa S, Khoudi H (2015) Co-expression of vacuolar Na+/H+ antiporter and H+-pyrophosphatase with an IRES-mediated dicistronic vector improves salinity tolerance and enhances potassium biofortification of tomato. Phytochemistry 117:537–546
Gouiaa S, Khoudi H, Leidi EO, Pardo JM, Masmoudi K (2012) Expression of wheat Na(+)/H(+) antiporter TNHXS1 and H(+)- pyrophosphatase TVP1 genes in tobacco from a bicistronic transcriptional unit improves salt tolerance. Plant Mol Biol 79:137–155
Groppa MD, Benavides MP (2008) Polyamines and abiotic stress: recent advances. Amino Acids 34(1):35–45
Guo Q, Zhang J, Gao Q, Xing S, Li F, Wang W (2008) Drought tolerance through overexpression of monoubiquitin in transgenic tobacco. J Plant Physiol 165(16):1745–1755
Gupta KJ, Stoimenova M, Kaiser WM (2005) In higher plants, only root mitochondria, but not leaf mitochondria reduce nitrite to NO, in vitro and in situ. J Exp Bot 56(420):2601–2609
Gupta K, Dey A, Gupta B (2013) Polyamines and their role in plant osmotic stress tolerance. In: Tuteja N, Gill SS (eds) Climate change and plant abiotic stress tolerance. Wiley-VCH, Weinheim, pp 1053–1072
Halford NG, Hey SJ (2009) Snf1-related protein kinases (SnRKs) act with in an intricate network that links metabolic and stress signaling in plants. Biochem J 419:247–259
Hanin M, Brini F, Ebel C, Toda Y, Takeda S, Masmoudi K (2011) Plant dehydrins and stress tolerance: versatile proteins for complex mechanisms. Plant Signal Behav 6:1503–1509
Hanin M, Ebel C, Ngom M, Laplaze L, Masmoudi K (2016) New insights on plant salt tolerance mechanisms and their potential use for breeding. Front Plant Sci 7
Hasegawa PM (2013) Sodium (Na+) homeostasis and salt tolerance of plants. Environ Exp Bot 92:19–31
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Ann Rev Plant Biol 51:463–499
He CX, Yan JQ, Shen GX, Fu LH, Holaday AS, Auld D, Blumwald E, Zhang H (2005) Expression of an Arabidopsis vacuolar sodium/proton antiporter gene in cotton improves photosynthetic performance under salt conditions and increases fiber yield in the field. Plant Cell Physiol 46:1848–1854
Hong Z, Lakkineni K, Zhang Z, Verma DPS (2000) Removal of feedback inhibition of 1 pyrroline-5-carboxylase synthetase (P5CS) results in increased proline accumulation and protection of plants from osmotic stress. Plant Physiol 122:1129–1136
Hong Y, Zhang H, Huang L, Li D, Song F (2016) Overexpression of a stress-responsive NAC transcription factor gene ONAC022 improves drought and salt tolerance in rice. Front Plant Sci 7:4
Hoque ABMZ, Haque MA, Sarker MRA, Rahman MA (2015) Marker-assisted introgression of saltol locus into genetic background of BRRI Dhan-49. Int J Biosci 6:71–80
Horie T, Hauser F, Schroeder JI (2009) HKT transporter-mediated salinity resistance mechanisms in Arabidopsis and monocot crop plants. Trends Plant Sci 14:660–668
Hu H, You J, Fang Y, Zhu X, Qi Z, Xiong L (2008) Characterization of transcription factor gene SNAC2 conferring cold and salt tolerance in rice. Plant Mol Biol 67:169–181
Hu L, Zhang P, Jiang Y, Fu J (2015) Metabolomic analysis revealed differential adaptation to salinity and alkalinity stress in Kentucky bluegrass (Poa pratensis). Plant Mol Bio Report 33:56–68
Ishitani M, Liu J, Halfter U, Kim CS, Shi W, Zhu JK (2000) SOS3 function in plant salt tolerance requires N-myristoylation and calcium binding. Plant Cell 12(9):1667–1677
Jain M (2015) Function genomics of abiotic stress tolerance in plants: a CRISPR approach. Front Plant Sci 6
Jakoby M, Weisshaar B, Dröge-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parcy F (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7(3):106–111
Jamoussi RJ, Elabbassi MM, Jouira HB, Hanana M, Zoghlami N, Ghorbel A, Mliki A (2014) Physiological responses of transgenic tobacco plants expressing the dehydration responsive RD22 gene of Vitis vinifera to salt stress. Turk J Bot 38:268–280
Kere GM, Chen C, Guo Q, Chen J (2017) Genetics of salt tolerance in cucumber (Cucumis sativus L.) revealed by quantitative traits loci analysis. Sci Lett 5(1):22–30
Kishor PBK, Hong Z, Miao GH, Hu CAA, Verma DPS (1995) Overexpression of Δ-pyrroline-5-carboxilase synthetase increases proline production and confers osmotolerance in transgenic plants. Plant Physiol 108:1387–1394
Kong X, Pan J, Zhang M, Xing X, Zhou Y, Liu Y, Li D, Li D (2011) ZmMKK4, a novel group C mitogen-activated protein kinase kinase in maize (Zea mays), confers salt and cold tolerance in transgenic Arabidopsis. Plant Cell Environ 34(8):1291–1303
Kumar K, Sinha AK (2013) Overexpression of constitutively active mitogen activated protein kinase kinase 6 enhances tolerance to salt stress in rice. Rice 6:25
Kumar G, Purty RS, Sharma MP, Singla-Pareek SL, Pareek A (2009) Physiological responses among Brassica species under salinity stress show strong correlation with transcript abundance for SOS pathway-related genes. J Plant Physiol 166:507–520
Lee SK, Kim BG, Kwon TR, Jeong MJ, Park SR, Lee JW, Byun MO, Kwon HB, Matthews BF, Hong CB, Park SC (2011) Overexpression of the mitogen-activated protein kinase gene OsMAPK33 enhances sensitivity to salt stress in rice (Oryza sativa L.). J Biosci 36(1):139–151
Li HW, Zang BS, Deng XW, Xi-Ping W (2011) Overexpression of the trehalose-6-phosphate synthase gene OsTPS1 enhances abiotic stress tolerance in rice. Planta 234:1007
Li H, Gao Y, Xu H, Dai Y, Deng D, Chen J (2013) ZmWRKY33, a WRKY maize transcription factor conferring enhanced salt stress tolerances in Arabidopsis. Plant Growth Regul 70:207–216
Linh LH, Khanh TD, Luanl NV, Cuc DTK, Duc LD, Linh TH, Ismail AM, Ham LH (2012) Application of marker assisted backcrossing to pyramid salinity tolerance (Saltol) into Rice Cultivar- Bac Thom 7. VNU J Sci Nat Sci Technol 28:87–99
Liu J, Ishitani M, Halfter U, Kim CS, Zhu JK (2000) The Arabidopsis thaliana SOS2 gene encodes a protein kinase that is required for salt tolerance. Proc Natl Acad Sci U S A 97(7):3730–3734
Liu H, Zhou X, Dong N, Liu X, Zhang H, Zhang Z (2011) Expression of a wheat MYB gene in transgenic tobacco enhances resistance to Ralstonia solanacearum, and to drought and salt stresses. Funct Integr Genomics 11:431–443
Liu C, Mao B, Ou S, Wang W, Liu L, Wu Y, Chu C, Wang X (2014) OsbZIP71, abZIP transcription factor, confers salinity and drought tolerance in rice. Plant Mol Biol 84:19–36
Liu L, Zhang Z, Dong J, Wang T (2016) Overexpression of MtWRKY76 increases both salt and drought tolerance in Medicago truncatula. Environ Exp Bot 123:50–58
Luo M, Zhao Y, Zhang R, Xing J, Duan M, Li J, Wang N, Wang W, Zhang S, Chen Z, Zhang H, Shi Z, Song W, Zhao J (2017) Mapping of a major QTL for salt tolerance of mature field-grown maize plants based on SNP markers. BMC Plant Biol 17:140
Lyzenga WJ, Stone SL (2012) Abiotic stress tolerance mediated by protein ubiquitination. J Exp Bot 63(2):599–616
Ma X, Qian Q, Zhu D (2005) Expression of a calcineurin gene improves salt stress tolerance in transgenic rice. Plant Mol Bio 58:483–495
Ma C, Burd S, Lers A (2015) miR408 is involved in abiotic stress responses in Arabidopsis. Plant J Cell Mol Biol 84:169–187
Mallikarjuna G, Mallikarjuna K, Reddy MK, Kaul T (2011) Expression of OsDREB2A transcription factor confers enhanced dehydration and salt stress tolerance in rice (Oryza sativa L.). Biotechnol Lett 33:1689–1697
Mantri N., Patade V., Pang E. (2014) Recent Advances in Rapid and Sensitive Screening For Abiotic Stress Tolerance. In: Tran LP (ed) Improvement of Crops in the Era of Climate Changes, vol 2 (trans: Ahmad P, Wani MR, Azooz MM). Springer, New York. ISBN 978-1-4614-8823-1
Mickelbart MV, Hasegawa PM, Bailey-Serres J (2015) Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat Rev Genet 16:237–251
Miller G, Shulaev V, Mittler R (2008) Reactive oxygen signaling and abiotic stress. Physiol Plant 133:481–489
Mittler R, Blumwald E (2010) Genetic engineering for modern agriculture: challenges and perspectives. Ann Rev Plant Biol 61:443–462
Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) AP2/ERF family transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819(2):86–96
Moller IS, Gilliham M, Jha D, Mayo GM, Roy SJ, Coates JC, Haseloff J, Tester M (2009) Shoot Na+ exclusion and increased salinity tolerance engineered by cell type-specific alteration of Na+ transport in Arabidopsis. Plant Cell 21(7):2163–2178
Munir S, Liu H, Xing Y, Hussain S, Ouyang B, Zhang Y, Li H, Ye Z (2016) Overexpression of calmodulin-like (ShCML44) stress-responsive gene from Solanum habrochaites enhances tolerance to multiple abiotic stresses. Sci Rep 6:31772
Munns R (2002) Comparative physiology of salt and water stress. Plant Cell Environ 25:239–250
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Ann Rev Plant Biol 59:651–681
Nakashima K, Takasaki H, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) NAC transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819:97–103
Nikalje GC, Nikam TD, Suprasanna P (2017) Looking at halophytic adaptation to high salinity through genomics landscape. Curr Genom 18(6). https://doi.org/10.2174/1389202918666170228143007.
Nongpiur RC, Singla-Pareek SL, Pareek A (2016) Genomics approaches for improving salinity stress tolerance in crop plants. Curr Genomics 17(4):343–357
Ohta M, Hayashi Y, Nakashima A, Hamada A, Tanaka A, Nakamura T, Hayakawa T (2002) Introduction of a Na /H antiporter gene from confers salt tolerance to rice. FEBS Letts 532(3):279–282
Orellana S, Yañez M, Espinoza A, Verdugo I, González E, Ruiz-Lara S, Casaretto JA (2010) The transcription factor SlAREB1 confers drought, salt stress tolerance and regulates biotic and abiotic stress-related genes in tomato. Plant Cell Environ 33:2191–2208
Osakabe Y, Watanabe T, Sugano SS, Ueta R, Ishihara R, Shinozaki K, Osakabe K (2016) Optimization of CRISPR/Cas9 genome editing to modify abiotic stress responses in plants. Sci Rep 6(1)
Otoch MLO, Sobreira ACM, Aragão MEF, Orellano EG, Lima MGS, Melo DF (2001) Salt modulation of vacuolar H+-ATPase and H+-Pyrophosphatase activities in Vigna unguiculata. J Plant Physiol 158:545–551
Ouyang SQ, Liu YF, Liu P, Lei G, He SJ, Ma B, Zhang WK, Zhang JS, Chen SY (2010) Receptor-like kinase OsSIK1 improves drought and salt stress tolerance in rice (Oryza sativa) plants. Plant J 62:316–329
Park HY, Seok HY, Park BK, Kim SH, Goh CH, Lee BH, Lee CH, Moon YH (2008) Overexpression of Arabidopsis ZEP enhances tolerance to osmotic stress. Biochem Biophys Res Commun 375:80–85
Pasapula V, Shen G, Kuppu S, Paez-Valencia J, Mendoza M, Hou P, Chen J, Qiu X, Zhu L, Zhang X, Auld D, Blumwald E, Zhang H, Gaxiola R, Payton P (2011) Expression of an Arabidopsis vacuolar H+-pyrophosphatase gene (AVP1) in cotton improves drought-and salt tolerance and increases fibre yield in the field conditions. Plant Biotechnol J 9:88–99
Patel P, Yadav K, Ganapathi TR and Suprasanna Penna (2018) Plant miRNAome: cross talk in abiotic stressful times. In: Rajpal VR, Sehgal D, Raina SN (eds) Genomics-assisted breeding for crop improvement: abiotic stress tolerance. Springer (In Press)
Penna S (2003) Building stress tolerance through over-producing trehalose in transgenic plants. Trends Plant Sci 8(8):355–357
Prashanth SR, Sadhasivam V, Parida A (2008) Overexpression of cytosolic copper/zinc superoxide dismutase from a mangrove plant Avicennia marina in indica rice var Pusa Basmati-1 confers abiotic stress tolerance. Transgenic Res 17:281–291
Qi L, Yang J, Yuan Y, Huang L, Chen P (2015) Overexpression of twoR2R3-MYB genes from Scutellaria baicalensis induces phenylpropanoid accumulation and enhances oxidative stress resistance in transgenic tobacco. Plant Physiol Biochem 94:235–243
Qiu QS, Guo Y, Dietrich MA, Schumaker KS, Zhu J-K (2002) Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc Natl Acad Sci U S A 99:8436–8441
Quan R, Wang J, Hui J, Bai H, Lyu X, Zhu Y, Zhang H, Zhang Z, Li S, Huang R (2018) Improvement of salt tolerance using wild rice genes. Front Plant Sci 8:2269
Ravikiran KT, Krishnamurthy SL, Warraich AS, Sharma PC (2017) Diversity and haplotypes of rice genotypes for seedling stage salinity tolerance analyzed through morpho-physiological and SSR markers. Field Crops Res. Online published. https://doi.org/10.1016/j.fcr.2017.04.006
Reddy PS, Jogeswar G, Rasineni GK, Maheswari M, Reddy AR, Varshney RK, Kishor PBK (2015) Proline over-accumulation alleviates salt stress and protects photosynthetic and antioxidant enzyme activities in transgenic sorghum [Sorghum bicolor (L.) Moench]. Plant Physiol Biochem 94:104–113
Rong W, Qi L, Wang A, Ye X, Du L, Liang H, Xin Z, Zhang Z (2014) The ERF transcription factor TaERF3 promotes tolerance to salt and drought stresses in wheat. Plant Biotechnol J 12:468–479
Roxas VP, Smith RK Jr, Allen ER, Allen RD (1997) Overexpression of glutathione S-transferase/glutathione peroxidase enhances the growth of transgenic tobacco seedlings during stress. Nat Biotechnol 15:988–991
Roy M, Wu R (2002) Overexpression of S-adenosyl methionine decarboxylase gene in rice increases polyamine level and enhances sodium chloride-stress tolerance. Plant Sci 163(5):987–992
Roy SJ, Negrão S, Tester M (2014) Salt resistant crop plants. Curr Opin Biotechnol 26:115–124
Rubio F, Gassmann W, Schroeder JI (1995) Sodium-driven potassium uptake by the plant potassium transporter HKT1 and mutations conferring salt tolerance. Science 270:1660–1663
Rui A, Qi-Jun C, Mao-Feng C, Ping-Li L, Zhao S, Zhi-Xiang Q, Jia C, Xue-Chen W (2007) AtNHX8, a member of the monovalent cation:proton antiporter-1 family in Arabidopsis thaliana, encodes a putative Li+/H+ antiporter. Plant J 49(4):718–728
Saijo Y, Hata S, Kyozuka J, Shimamoto K, Izui K (2000) Overexpression of a single Ca2+ dependent protein kinase confers both cold and salt/drought tolerance on rice plants. Plant J 23:319–327
Sakamoto A, Murata A, Murata N (1998) Metabolic engineering of rice leading to biosynthesis of glycine betaine and tolerance to salt and cold. Plant Mol Biol 38:1011–1019
Sanders D (2000) Plant biology: the salty tale of Arabidopsis. Curr Biol 10(13):R486–R488
Sanders D, Pelloux J, Brownlee C, Harper JF (2002) Calcium at the crossroads of signaling. Plant Cell 14:401–417
Schachtman DP, Schroeder JI (1994) Structure and transport mechanism of a high-affinity potassium uptake transporter from higher plants. Nature 370:655–658
Schilling RK, Marschner P, Shavrukov Y, Berger B, Tester M, Roy SJ, Plett DC (2013) Expression of the Arabidopsis vacuolar H+-pyrophosphatase gene (AVP1) improves the shoot biomass of transgenic barley and increases grain yield in a saline field. Plant Biotechnol J 12:378–386
Serrano R, Mulet JM, Rios G, Marquez JA, de Larrinoa IF, Leube MP, Mendizabal I, Pascual-Ahuir A, Proft M, Ros R, Montesinos C (1999) A glimpse of the mechanisms of ion homeostasis during salt stress. J Exp Bot 50:1023–1036
Shabala S, Cuin TA (2008) Potassium transport and plant salt tolerance. Physiol Plantarum 133:651–669
Shabala S, Pottosin I (2014) Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol Plant 151(3):257–279
Shen G, Wei J, Qiu X, Hu R, Kuppu S, Auld D, Blumwald E, Gaxiola R, Payton P, Zhang H (2014) Co-overexpression of AVP1 and AtNHX1 in cotton further improves drought and salt tolerance in transgenic cotton plants. Plant Mol Biol Rep 33:167–177
Sheveleva E, Chmara W, Bohnert HJ, Jensen RG (1997) Increased salt and drought tolerance by D-Ononitol production in transgenic Nicotiana tabacum. Plant Physiol 115:1211–1219
Shi W, Liu D, Hao L, Wu CA, Guo X, Li H (2014) GhWRKY39, a member of the WRKY transcription factor family in cotton, has a positive role in disease resistance and salt stress tolerance. Plant Cell Tissue Organ Cult 118:17–32
Shinozaki K, Yamaguchi-Shinozakiy K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant 6(5):410–417
Shriram V, Kumar V, Devarumath RM, Khare TS, Wani SH (2016) MicroRNAs as potential targets for abiotic stress tolerance in plants. Front Plant Sci 7
Singh A, Singh VK, Singh SP, Pandian RTP, Ellur RK, Singh D, et al. (2012) Molecular breeding for the development of multiple disease resistance in Basmati rice. AoB Plants 2012 ls029
Smith CA, Melino VJ, Sweetman C, Soole KL (2009) Manipulation of alternative oxidase can influence salt tolerance in Arabidopsis thaliana. Physiol Plant 137:459–447
Song X, Yu X, Hori C, Demura T, Ohtani M, Zhuge Q (2016) Heterologous overexpression of poplar SnRK2 genes enhanced salt stress tolerance in Arabidopsis thaliana. Front Plant Sci 7:612
Sperling O, Lazarovitch N, Schwartz A, Shapira O (2014) Effects of high salinity irrigation on growth, gas-exchange, and photoprotection in date palms (Phoenix dactylifera L., cv. Medjool). Environ and Exp Bot. 99:100–109
Su J, Wu R (2004) Stress-inducible synthesis of proline in transgenic rice confers faster growth under stress conditions than that with constitutive synthesis. Plant Sci 166:941–948
Székely G, Abrahám E, Cséplo A, Rigó G, Zsigmond L, Csiszár J, Ayaydin F, Strizhov N, Jásik J, Schmelzer E, Koncz C, Szabados L (2008) Duplicated P5CS genes of Arabidopsis play distinct roles in stress regulation and developmental control of proline biosynthesis. Plant J 53:11–28
Tarczynski MC, Jensen RG, Bohnert HJ (1992) Expression of a bacterial mtlD gene in transgenic tobacco leads to production and accumulation of mannitol. Proc Natl Acad Sci U S A 89:2600–2604
Thomas JC, Sepahi M, Arendall B, Bohnert HJ (1995) Enhancement of seed germination in high salinity by engineering mannitol expression in Arabidopsis thaliana. Plant Cell Environ 18(7):801–806
Thomson MJ, Ocampo M, Egdane J, Rahman MA, Sajise AG, Adorada DL et al (2010) Characterizing the Saltol quantitative trait locus for salinity tolerance in rice. Rice 3:148–160
Tsutsumi K, Yamada N, Cha-Um S, Tanaka Y, Takabe T (2015) Differential accumulation of glycine betaine and choline monooxygenase in bladder hairs and lamina leaves of Atriplex gmelini, under high salinity. J Plant Physiol 176:101–107
Turan S, Katrina C, Kumar S (2012) Salinity tolerance in plants: breeding and genetic engineering. Aust J Crop Sci 6(9):1337–1348
Türkan I, Demiral T (2009) Recent developments in understanding salinity tolerance. Environ Exp Bot 67:2–9
Ubbens JR, Stavness I (2017) Deep plant phenomics: a deep learning platform for complex plant phenotyping tasks. Front Plant Sci 8:1190. https://doi.org/10.3389/fpls.2017.01190
van de Wiel CCM, Schaart JG, Lotz LAP, Smulders MJM (2017) New traits in crops produced by genome editing techniques based on deletions. Plant Biotechnol Rep 11(1):1–8
Varshney RK, Bansal KC, Aggarwal PK, Datta SK, Craufurd PQ (2011) Agricultural biotechnology for crop improvement in a variable climate: hope or hype? Trends Plant Sci 16:363–371
Wang B, Lüttge U, Ratajczak R (2001) Effects of salt treatment and osmotic stress on V-ATPase and V-PPase in leaves of the halophyte Suaeda salsa. J Exp Bot 52(365):2355–2365
Wang Y, Gao C, Liang Y, Wang C, Yang C, Liu G (2010) A novel bZIP gene from Tamarix hispida mediates physiological responses to salt stress in tobacco plants. J Plant Physiol 167:222–230
Wang C, Deng P, Chen L, Wang X, Ma H, Hu W, Yao N, Feng Y, Chai R, Yang G, He G (2013) A wheat WRKY transcription factor TaWRKY10 confers tolerance to multiple abiotic stresses in transgenic tobacco. PLoS One 8(6):e65120
Wang RK, Cao ZH, Hao YJ (2014) Overexpression of a R2R3MYB gene MdSIMYB1 increases tolerance to multiple stresses in transgenic tobacco and apples. Physiol Plant 150:76–87
Wang H, Wang H, Shao H, Tang X (2016) Recent advances in utilizing transcription factors to improve plant abiotic stress tolerance by transgenic technology. Front Plant Sci 7:67. https://doi.org/10.3389/fpls.2016.00067
Xing Y, Chen W-H, Jia W, Zhang J (2015) Mitogen-activated protein kinase kinase 5 (MKK5)-mediated signalling cascade regulates expression of iron superoxide dismutase gene in Arabidopsis under salinity stress. J Exp Bot 66(19):5971–5981
Xiong L, Yang Y (2003) Disease resistance and abiotic stress tolerance in rice are inversely modulated by an abscisic acid–inducible mitogen-activated protein kinase. Plant Cell 15:745–759
Xu D, Duan X, Wang B, Hong B, Ho THD, Wu R (1996) Expression of a late embryogenesis abundant protein gene, HVA1, from barley confers tolerance to water deficit and salt stress in transgenic rice. Plant Physiol 110:249–257
Xu G, Cui Y, Li M, Wang M, Yu Y, Zhang B, Huang L, Xia X (2013) OsMSR2, a novel rice calmodulin-like gene, confers enhanced salt tolerance in rice (Oryza sativa L.). AJCS 7(3):368–373
Xue ZY, Zhi DY, Xue GP, Zhang H, Zhao YX, Xia GM (2004) Enhanced salt tolerance of transgenic wheat (Triticum aestivum L.) expressing a vacuolar Na+/H+ antiporter gene with improved grain yields in saline soils in the field and a reduced level of leaf Na+. Plant Sci 167:849–859
Yamaguchi T, Blumwald E (2005) Developing salt-tolerant crop plants: challenges and opportunities. Trends Plant Sci 10:615–620
Yang J, Zhang J, Liu K, Wang Z, Liu L (2007) Involvement of polyamines in the drought resistance of rice. J Exp Bot 58(6):1545–1555
Yang Q, Chen ZZ, Zhoua XF, Yina HB, Lia X, Xina XF, Honga XH, Zhu JK, Gong Z (2009) Over-expression of SOS (salt overly sensitive) genes increases salt tolerance in transgenic Arabidopsis. Mol Plant 2:22–31
Yang A, Dai X, Zhang WH (2012) AR2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J Exp Bot 63:2541–2556
Yang C, Wang R, Gou L, Si Y, Guan Q (2017) Overexpression of Populus trichocarpa mitogen-activated protein Kinase Kinase4 enhances salt tolerance in tobacco. Int J Mol Sci 18:2090
Yusuf MA, Kumar D, Rajwanshi R, Strasser RJ, Govindjee MTM, Sarin NB (2010) Overexpression of γ-tocopherol methyl transferase gene in transgenic Brassica juncea plants alleviates abiotic stress: physiological and chlorophyll a fluorescence measurements. Biochim Biophys Acta 1797:1428–1438
Zhai Y, Wang Y, Li Y, Lei T, Yan F, Su L, Li X, Zhao Y, Sun X, Li J, Wang Q (2013) Isolation and molecular characterization of GmERF7, a soybean ethylene-response factor that increases salt stress tolerance in tobacco. Gene 513:174–183
Zhang B (2015) MicroRNA: a new target for improving plant tolerance to abiotic stress. J Exp Bot 66:1749–1761
Zhang HX, Blumwald E (2001) Transgenic salt tolerant tomato plants accumulate salt in the foliage but not in the fruits. Nat Biotechnol 19:765–768
Zhang S, Klessig DF (2001) MAPK cascades in plant defense signaling. TRENDS Plant Sci 6:520–527
Zhang B, Wang Q (2016) MicroRNA, a new target for engineering new crop cultivars. Bioengineered 7(1):7–10
Zhang HX, Hodson JN, Williams JP, Blumwald E (2001) Engineering salt-tolerant Brassica plants: characterization of yield and seed oil quality in transgenic plants with increased vacuolar sodium accumulation. Proc Natl Acad Sci U S A 98:12832–12836
Zhang L, Xi D, Li S, Gao Z, Zhao S, Shi J, Wu C, Guo X (2011) A cotton group C MAP kinase gene, GhMPK2, positively regulates salt and drought tolerance in tobacco. Plant Mol Biol 77:17–31
Zhang X, Liu X, Wu L, Yu G, Wang X, Ma H (2015) The SsDREB transcription factor from the succulent halophyte Suaeda salsa enhances abiotic stress tolerance in transgenic tobacco. Int J Genomics 2015:875497
Zhang X, Shabala S, Koutoulis A, Shabala L, Zhou M (2017) Meta-analysis of major QTL for abiotic stress tolerance in barley and implications for barley breeding. Planta 245(2):283–295
Zhao YL, Wang HM, Shao BX, Chen W, Guo ZJ, Gong HY, Sang XH, Wang JJ, Ye WW (2016) SSR-based association mapping of salt tolerance in cotton (Gossypium hirsutum L.). Genet Mol Res 15(2):gmr.15027370
Zheng X, Chen B, Lu G, Han B (2009) Overexpression of a NAC transcription factor enhances rice drought and salt tolerance. Biochem Biophys Res Commun 379:985–989
Zhou QY, Tian AG, Zou HF, Xie ZM, Lei G, Huang J, Wang CM, Wang HW, Zhang JS, Chen SY (2008) Soybean WRKY-type transcription factor genes, GmWRKY13, GmWRKY21, and GmWRKY54, confer differential tolerance to abiotic stresses in transgenic Arabidopsis plants. Plant Biotechnol J 6:486–503
Zhou M, Li D, Li Z, Hu Q, Yang C, Zhu L, Luo H (2013) Constitutive expression of a miR319 gene alters plant development and enhances salt and drought tolerance in transgenic creeping bentgrass. Plant Physiol 161:1375–1391
Zhu N, Cheng S, Liu X, Du H, Dai M, Zhou DX, Yang W, Zhao Y (2015) The R2R3-type MYB gene OsMYB91 has a function in coordinating plant growth and salt stress tolerance in rice. Plant Sci 236:146–156
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Suprasanna, P., Ghuge, S.A., Patade, V.Y., Mirajkar, S.J., Nikalje, G.C. (2018). Genomic Roadmaps for Augmenting Salinity Stress Tolerance in Crop Plants. In: Kumar, V., Wani, S., Suprasanna, P., Tran, LS. (eds) Salinity Responses and Tolerance in Plants, Volume 2. Springer, Cham. https://doi.org/10.1007/978-3-319-90318-7_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-90318-7_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-90317-0
Online ISBN: 978-3-319-90318-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)