Abstract
Biomass conversion to an utilizable energy sources such as monomer sugars using enzymatic hydrolysis has been emerged as the current technology which promises the future energy. In nature, bioconversion process of biomass is mediated by a group of biofunctional hydrolytic enzymes. These enzymes generally work in cooperative synergetic action to facilitate enhanced effective degradation of biomass. Xylanase is one of the crucial hydrolytic enzymes involved in hydrolysis of xylan, the hemicellulose which constitutes 15–30 % of the plant biomass. This chapter discusses in detail about the enzymatic hydrolysis of xylan by the xylolytic enzyme endo-1,4-β-xylanase, its occurrence in nature and mode of action, structure and classifications, current methods for its production, purification, and characterization. In addition, the major and recent industrial applications of this enzyme were highlighted as well.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Nowadays, the governments promote extensive researches for the development of an alternative transportation fuel from a renewable energy sources, realizing the upcoming major energy crisis due to the depletion of petroleum-derived fuels. As an initiative, the department of energy in US has started producing biofuels with a set target of 60 billion gallon per year by 2030. Europe also has the similar targets by that time to replace 25 % of petroleum-based liquid transportation fuel by biofuels (Himmel et al. 2007). However, it is a challenging target since the biofuel production using substrates such as sugar cane and corn has limited capacity to supply such a huge volumes. Apart from this, in many countries people protest against the use of food crops for biofuel production and many debates that it leads to a major food crisis. As a remedy, researchers found out that the lignocellulose biomass which consists of 75 % polysaccharide sugars can be used as a significant feedstock for biofuel production (Lynd et al. 1991). If the attraction is more, lignocellulose biomasses are renewable energy source and exist abundant in nature and it can be derived predominantly from agricultural wastes (Gomez et al. 2008). Lignocelluloses are heteropolysaccharides that give structural rigidity to all kinds of plant. They account for approximately more than 50 % of the total biomass in the nature, synthesized (estimated 10–50 × 1012 tons year−1) by plants through photosynthesis (Claassen et al. 1999). Lignocelluloses are composed of three major polymeric constituents such as cellulose (an unbranched linier polymers of long β-1-4 glucan between 2000 and 27,000 units), hemicellulose (a heterologous polysaccharides which are complex in nature with sugar monomeric subunits such as d-xylose, d-mannose, d-galactose, and l-arabinose) and lignin (complex phenolic structure act as linkage between cellulose and hemicellulose.
The name hemicellulose was first introduced by Schulze (1891). These heteropolysaccharides are abundant in plant cell wall and have complex structures when compared to cellulose due to the enormous number of sugar monomeric side chains (500–3000). It creates a cross-linked network between the cellulose microfibrils and lignin and plays a major role in structural integrity of plant cell walls. It also regulates the rate of expansion of primary cell wall and acts as a shield over cellulose to protect it from enzymatic hydrolysis. Hemicellulose is the second most abundant renewable biomass that is available after cellulose within the lignocellulose matrix representing about 20–35 % of total biomass (Saha 2003). Table 7.1 summarizes the approximate composition of various biomass materials. Classification of hemicellulose was made based on the types of sugar monomer units present on them. As mentioned above, hemicelluloses are composed of xylan, mannan, galactan, and arabinan polymers. Among these xylan is the most frequently occurring polymer composed mainly of β-d-xylopyranosyl residues which linked by β-1,4-glycosidic bonds (Beg et al. 2001).
In hardwoods plants, hemicellulose primarily consists of xylans and glucomannans whereas in softwood plants, it is composed of arabinoglucuronoxylans (xylans), xyloglucans, glucomannans, and arabinogalactans. Xylans are the major form of hemicellulose made of d-xylopyranosyl units which are linked by β-1,4-glycosidic bonds. β-mannan-based polymers are the second major form of cellulose which are made of β-1,4-linked mannose and/or glucose residues. Galactoglucomannans which consist of α-1,6-linked galactose side chains are also grouped under these polymers. They are found abundant in hardwood hemicellulose, which comprised varying ratios of d-mannopyranose units α-(1–6)-substituted galactopyranose side chains with O-acetyl side chains and β-1-4-linked-d-glucopyranose units. Arabinan and arabinogalactans are also considered as hemicelluloses which are generally composed of α-1,5-linked l-arabinofuranosyl units. Table 7.2 summarizes the major enzymes involved in the breakdown of cross-linked hemicelluloses. Those groups of enzymes which are involved in hydrolysis of xylan are generally known as xylanases. The current book focuses on one of the crucial xylolytic enzymes known as endo-1-4-β-xylanase.
2 Enzymatic Hydrolysis of Xylan
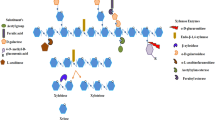
In most terrestrial plant species, secondary cell wall thickening takes place by the deposition of xylan and they are found in-between cellulosic fiber sheets and lignin with complex binding relationship through hydrogen bonding. Xylan thus forms a think wall over cellulose and enables the protection from degradation by different cellulose degrading enzymes. Xylan constitutes 15–30 % of the plant biomass in hardwood and 7–12 % of the plant biomass in softwood. Since they are the major hemicellulose group in most of the plant species, they have been considered as one among the renewable source of energy in the form of biomass. Structure of xylan is complex in nature as they are composed of β-1,4 linked-d-xylopyranose units with α-l-arabinofuranose and 4-O-methyl-α-d-glucuronopyranosyl acid side chains. Figure 7.1 shows the major enzymes involved and their site of action during the enzymatic hydrolysis is xylan (Polizeli et al. 2005).
Enzymatic hydrolysis of xylan (Polizeli et al. 2005)
Hardwood xylan (e.g., birchwood xylan) consists of more than 70 β-1,4 linked-d-xylopyranose units linked by β-1,4 glycosidic bonds and 4-O-methyl-α-d-glucuronopyranosyl acid side chains which are found attached at second carbon of every tenth xylopyranose units. Acetylation rate is higher in these groups and it is quite frequent at second and third carbon atom on xylopyranose ring. In softwood xylan, acetylation rate is zero but instead of an acetyl group they possess α-l-arabinofuranose linked by α-1-3 glycosidic bond at the third carbon atom on xylopyranose ring. Based on the side chains of the xylan backbone, they are classified into different groups such as linear homoxylan, arabinoxylan, glucuronoxylan, and glucuronoarabinoxylan. These four groups are heterogenic in terms of their degree and nature of branching. In many plants, xylans are found partially acetylated, which protect them from complete degradation by xylolytic enzymes. This could be the reason behind the fact that the complete degradation of xylan is accomplished only by synergetic interaction of acetyl xylan esterase and endo-xylanase enzymes.
Due to the heterogeneity and complex chemical structure of polymer, the complete breakdown of xylan requires the synergetic action of several hydrolytic enzymes which are specific in their mode of actions. Generally, xylans are not attacked randomly, but the bonds selected for hydrolysis depend on several biochemical factors that determine the nature of the substrate molecules such as the chain length, the degree of branching, and the presence of substituents.
The major xylolytic enzymes involved in xylan hydrolysis are as follows:
-
Endo-1,4-β-xylanase—cleaves long xylan backbone into short xylooligosaccharides.
-
β-xylosidases—also attacks the β-1,4-glycosidic linkages to liberate xylose from short oligosaccharides.
-
α-l-arabinofuranosidases—remove l-arabinofuranose side chains.
-
α-d-glucuronidases—hydrolyze the methyl glucuronate residues.
-
Acetyl xylan esterases—hydrolyze acetate groups on xylan backbone.
-
Ferulic acid esterases—hydrolyze the aromatic acids groups linked on arabinofuranoside residues.
Endo-1,4-β-xylanase: (EC 3.2.1.8) cleaves the glycosidic bonds between the d-xylopyranose units on the xylan backbone and as a result, short xylooligosaccharides are formed. β-d-xylosidases: (EC 3.2.1.37) liberate xylose from short xylooligosaccharides which are formed during xylan hydrolysis by endo-1,4-β-xylanase. These enzymes are classified based on their relative affinity for the xylooligosaccharides into distinct entities such as xylobiases and exo-1,4-β-xylanases. Even though they are two different enzymes, they have been recognized as β-d-xylosidases in general, and releases β-d-xylopyranosyl residues from small xylooligosaccharides and xylobiose. Their affinity for the xylooligosaccharides is inversely proportional to its complexity. Xylobiose was found to be the best substrate for these enzyme groups. These groups of enzymes play important role in xylan hydrolysis when there is endo-1,4-β-xylanase inhibition due to the accumulation of short oligomers of β-d-xylopyranosyl units, by hydrolyzing these products and remove the cause of inhibition thereby enhancing the xylan hydrolysis (Andrade et al. 2004). The molecular weights of these enzyme groups are relatively high, between 60 and 260 kDa and their pH optima is between 4.0 and 5.0. Generally, these enzymes are thermophilic in nature and their optimum temperature varies from 40° to 80°. However, in most cases they exhibit highest activity around 60 °C (Rizzatti et al. 2001). α-l-arabinases: These enzymes remove l-arabinose side chains which are substituted at second and third carbon on β-d-xylopyranosyl ring of arabino xylan. They have been classified as exo-α-l-arabinofuranosidase (EC 3.2.1.55) and endo-1,5-α-l-arabinase (EC 3.2.1.99), based on their distinct mode of action. The first group, which is most common, degrades branched arabinans and p-nitrophenyl-α-l-arabinofuranosidase, whereas the second group hydrolyzes only the linear arabinans (de Vries et al. 2000). The α-d-glucuronidases (EC 3.2.1.131) can hydrolyze the α-1,2 linkages between glucuronic acid and xylose residues found in glucuronoxylan. They are classified under glycoside hydrolase family 67 and rarely found in nature. During biodegradation of glucuronoxylan, these enzymes are able to release glucuronic acid. However, the enzyme activity is only limited on short xylooligomers due to the partial hindrance by acetyl group present in xylan backbone (Puls et al. 1991). Acetylxylan esterase: (EC 3.1.1.6) it acts by removing the O-acetyl groups from second and third carbon on β-d-xylopyranosyl ring of acetyl xylan. This enzyme plays important role in xylan hydrolysis by removing O-acetyl side chains, since O-acetyl groups in acetyl xylan interfere the activity of endo-1,4-β-xylanase and β-d-xylosidases on xylan backbone. Xylan extraction from hardwoods is usually mediated by alkali treatment, resulting in acetyls hydrolysis, and probably this could be the reason of the late discovery of this enzyme (Shao and Wiegel 1995; Blum et al. 1999). Ferulic acid esterase: Ferulic acid esterases (EC 3.1.1.-) act by hydrolyzing the bond between arabinose side chain and ferulic acid group attached (Crepin et al. 2004). The cooperative functioning among the above-mentioned xylolytic enzymes enhance the complete biodegradation of xylan. It was also observed that acetylxylan esterase activity on xylan results in deacetylation of xylan backbone and these deacetylated xylans are more easily hydrolyzed by endo-xylanase. Similarly, there was a significant increase in hydrolysis efficiency when arabinoxylan was pretreated by α-arabinofuranosidase. This was due to the removal of arabinan side chains which act as hindrance to endo-xylanase activity (de Vries et al. 1999).
3 Occurrence of Endo-1,4-β-xylanase
Endo-1,4-β-xylanase are also known as endo-(1→4)-β-xylan 4-xylanohydrolase, endo-1,4-xylanase, xylanase, β-1,4-xylanase, endo-1,4-xylanase, endo-β-1,4-xylanase, 1,4-β-xylan xylanohydrolase, β-xylanase, β-1,4-xylan xylanohydrolase, endo-1,4-β-xylanase, β-d-xylanase, 4-β-d-xylan xylanohydrolase, endo-(1→4)-beta-xylan 4-xylanohydrolase, beta-1,4-xylanase, endo-beta-1,4-xylanase, endo-1,4-beta-d-xylanase, 1,4-beta-xylan xylanohydrolase, beta-xylanase, beta-1,4-xylan xylanohydrolase, endo-1,4-beta-xylanase, beta-d-xylanase, and 4-beta-d-xylan xylanohydrolase. In nature, the endo-1,4-β-xylanase enzymes are widely distributed among eukaryotes and prokaryotes. The occurence of these enzymes was also reported in higher eukaryotes such as plants, protozoa, small insects, and several marine species. A Japanese pear fruit was reported producing endo-xylanase during its over-ripening period. Endo-xylanase of molecular weight 55 kDa was also isolated from wheat flour (Cleemput et al. 1997). Apart from these sources, members of higher animals such as insects and fresh water mollusc were also reported as xylanase producers (Yamura et al. 1997). However, bacteria, fungi, and actinomycetes were found to be the largest source of xylanase enzyme.
There have been several reports on microbial endo-1,4-β-xylanase since 1960. Nevertheless, most of these xylanase enzymes remained unnoticed. During 1980s first studies on xylanase-mediated bio-bleaching in pulp industry was reported and only since then the scientific world realized the great impact of xylanase enzymes in industrial applications. As mentioned above, a number of microorganisms including bacteria and fungi have the capacity for hydrolyzing xylans by synthesizing a variety of xylolytic enzymes. Early reports reveal that many of these microbial species are plant pathogens as they play an important role in degradation and invasion of plant tissues. It also shows that xylanases can elicit defense mechanisms in plants by a collective functioning with other cellulolytic enzyme groups.
3.1 Fungal Endo-1,4-β-xylanase
Many of the fungal species that are pathogenic to plants produce plant cell wall polysaccharide degrading enzymes. Endo-1,4-β-xylanase is one among the major group of such enzymes and they result in partial degradation of cell wall structures to the region of penetration (Subramaniyan 2000). A hypothetical model of fungal invasion on plant tissue is shown in Fig. 7.2.
Fungal invasion on plant tissue by xylolytic degradation (Prade 1996)
A number of fungal species have been used to produce xylanase since they are the major producer of xylanases in nature. Table 7.3 shows the major fungal species used to produce xylanases. Phanerochaete chrysosporium, a potent plant pathogen, reported to produce xylanase activity of 15–20 U mL−1 along with considerable amount of cellulose activity (Copa-Patino et al. 1993). Other study also showed that the thermophilic fungus Thermomyces lanuginosus has the capacity of high xylanase production up to 3576 U mL−1 (Singh et al. 2000). Many fungal species have been reported as xylanase producers; however, species with high xylanase activity and negligible cellulase activity are found to be very rare in nature. It was also observed that the pH optima of many of these fungal xylanases are between pH 5 and 6 even though they are stable at pH between 3 and 8. On the other hand, bacterial xylanases showed slightly higher pH optima that makes them suitable for many of the industrial applications. Thermal tolerance of the majority of fugal xylanase reported so far has been found below 50 °C and in most cases this particular property of fungal xylanase makes them less favorable for application in harsh industrial conditions. Reduced xylanase yield in fermenter studies is another major problem associated with fungi xylanase. This was due to poor oxygen transfer, shear force during fermentation, and the enzyme production process in fungi is highly regulated by growth morphology (El Enshasy et al. 1999, 2006; El Enshasy 2007). Generally, the fungal species has filamentous growth pattern, the fungal growth in fermenters is restricted due to the shear stress, and eventually this results in poor xylanase production (Dean et al. 1991).
3.2 Bacterial Endo-1,4-β-xylanase
Since 1980s, several bacterial strains which are capable of producing alkaline and highly thermophilic xylanase enzymes have been reported. Among these wide spectrum of bacteria, Bacilluss species are the most predominant producers of endo-1,4-β-xylanase enzyme with negligible amount of cellulose activity under optimized growth conditions. Other author showed a high yield of xylanase up to 506 IU mL−1 by Bacillus SSP-34 (Subramaniyan and Prema 2002). In general, bacterial endo-1,4-β-xylanase is suitable for industrial application since they have a wide pH optima and improved thermostability when compared to fungal xylanases. Ratto et al. (1992) reported Bacillus circulans strain with xylanase activity of 400 IU mL−1. They observed the optimal enzyme activity was at pH 7 and retained 40 % of activity at high pH up to 9.2. Many other researchers reported cellulose-free xylanase enzyme from Bacillus stearothermophilus strain T6 (Khasin et al. 1993; Lundgren et al. 1994). A relatively high xylanase activity was reported in Bacillus sp. strain NCL when it was grown in zeolite-induced medium (Balakrishnan et al. 2000). Thermophile bacteria such as Rhodothermus marinus produced approximately 1.8–4.03 IU mL−1 of thermostable xylanase along with detectable amount of cellulose activity. Other strain such as Bacillus circulans showed high yield of xylanase (400 IU mL−1) which has pH optima of 7 but with a high stability at alkaline pH (Beg et al. 2000). The concept of TCF (total chlorine free) bleaching of pulp was developed during this period and it was achieved with xylanase from Bacillus stearothermophilus strain T6 which has optimum activity at pH 6.5 (Kohli et al. 2001). A detailed description of major bacterial xylanase and their characteristics are given in Table 7.4.
4 Biosynthesis and Regulation of Microbial Endo-1,4-β-xylanase
Biosynthesis of xylanase enzyme by microorganisms and the phenomenon of xylanase induction are less investigated at molecular level due to the difficulty in setting up a cell-free system under experimental conditions. However, Srivastava and Srivastava (1993) introduced a hypothetical model of xylanase biosynthesis and regulation to describe similar conditions. Xylans are comparatively complex in structure with high molecular weight and thus it is usually hard to be utilized directly by microbes. Xylans are converted into smaller molecules such as xylose, xylobiose, xylotriose, or xylooligosaccharides by the constitutively produced small amount of endo-1,4-β-xylanase enzymes. These simpler molecules can be easily utilized by microbial cell as the carbon source (Zhao et al. 1997). These low molecular weight molecules can act as inducer for further xylanase expression and this has been one of the possible explanations for direct induction of xylanase gene.
Xylanases are usually secreted in the media which contains pure xylan or xylan residues in cultures of different species such as Aspergillus awamori, Trametestrogii, and Streptomyces sp. QG-11-3 (Beg et al. 2001). However, several exceptional cases have also been reported such as in Cellulomonas flavigena; xylan acts as a poor inducer of xylanase gene (Avalos et al. 1996). In some rare cases, for example, in yeast strain Trichosporon cutaneum, xylanase biosynthesis is induced by certain positional isomers. In many bacterial species, the xylanase induction is also possible with various sugars such as d-xylose, d-maltose, d-glucose, and d-arabinose. On the other hand, in many fungal species, the natural lignocelluloses such as corn cobs, rice straw, sugarcane bagasse, and wheat bran were found to be capable of triggering the xylanase induction (Gupta et al. 2000). Other study conducted by Kumar and Satyanarayana (2011) reported on the wheat bran-mediated xylanase induction in Bacillus halodurans TSEV1 strain. However, the hypothetical mechanism of direct induction is questionable when the transportation across the cell wall can be blocked by larger molecules. Based on the study of Gomes et al. (1994), there has been a universally supported concept based on intracellular β-xylosidases. They explains that the larger xylooligomers formed during xylan hydrolysis are directly transported into the cell matrix and it was further degraded into xylose residues by β-xylosidase which are present in the cytoplasm.
Hydrolysis of xylooligomers by hydrolytic transporter proteins during their transportation through cell membrane is another possible explanation for above phenomenon. There are also rare cases in which the high xylose concentration in the media inhibits the xylanase gene expression (Strauss et al. 1995). Exclusion of inducer transport across the cell membrane is another possible reason for poor gene expression. In similar cases such as in E. coli it has been observed that the presence of glucose in the media prevents the lactose transportation through the cell membrane which is inducer for lac operon (Borralho et al. 2002). It was also observed that the xylanase biosynthesis is mediated by complex metabolic pathway in which the inducer level and the level of repressor molecule to that particular inducer vary with the organism and their growth medium. For example, the xylanase production by Streptomycetes sp. was increased when cellulose substrates are used in the growth media, whereas in Cellulomonas favigena, sugarcane bagasse was found to be the best inducer of xylanase enzyme (Alejandro et al. 2007). Addition of xylose in fermentation media increased xylanase production to a significant amount in Bacillus sp. (Gupta and Kar 2009). However, other studies showed that the presence of readily utilizable sugars such as glucose, xylose, and galactose can suppress xylanase biosynthesis in other strains such as in Streptomyces sp. (Bajaj and Singh 2010) and in T. reesei (Mach-Aigner et al. 2010).
5 Classification and Structures of Xylanase
Xylanases are considered as one of the major hydrolyzing enzyme groups as they mediate the xylan degradation to simpler utilizable units. Activity of these enzymes depends on the substrate specificity as well as substrate complexity. Wong et al. (1988) classified xylanases into two major groups based on the end product of hydrolysis reaction. They were debranching enzymes which liberate arabinose and non-debranching enzymes which do not liberate arabinose residues from α,1,3,l-arabinofuranosyl. A few years later, another classification system was introduced based on the physiological properties of xylanase enzymes such as isoelectric point (PI) and molecular weight (MW). Therefore, xylanases were classified into two major groups: end xylanase enzymes with molecular weight lower than 30 kDa and basic pH and those with molecular weight higher than 30 kDa and acidic pH. However, few years later, this classification system was found incorrect since it matches for only a few xylanase enzymes.
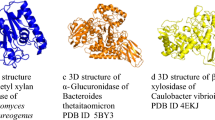
Henrissat and Bairoch (1993) proposed a broad classification system based on the similarities in amino acid sequence and catalytic module. The glycoside hydrolases are further classified into different families. This classification system has got wide acceptance since it gives information regarding the structural properties, the catalytic mechanisms of the enzyme, and their evolutionary relationship within the group. Xylanases were grouped under glycoside hydrolases (GH) 5, 8, 10, and 11 families. However, the enzymes with multi-domain that exhibit detectable amount of xylanase activity are grouped into GH7, 16, 43, and 62 families (Cantarel et al. 2009). Table 7.5 comprises major xylolytic enzymes, their classification into CE, GH families, and their current crystallographic structure. Some of these GH families are further classified into super families that represent a more distant common evolutionary ancestor. Major GH family xylanase and their general structures are shown in Fig. 7.3 (Table 7.6).
5.1 GH10 Xylanases
Most of the glycoside hydrolases which are classified under GH10 are endo-β-1,4-xylanases. Apart from these enzyme, G10 group also consists of a small number of endo-β-1,3-xylanases (EC 3.2.1.32), which cleave β-1,3-glycosidic linkages randomly in β-1,3-xylans backbone. A typical GH10 xylanases has a wide spectrum of substrate specificity as they are able to hydrolyze various forms of xylans in nature. These enzymes are capable to attack not only the linear forms of xylan but also the highly branched heteroxylans and xylosomes. Kolenova et al. (2006) analyzed the hydrolysis product from glucuronoxylan and confirmed that GH10 xylanases can attack xylan at its non-reducing end if there are two or more unbranched xylose residues which are present on backbone. It was also found that GH10 xylanases can even hydrolyse the highly branched arabinoxylan into small fragments. Plant xylanases which have been identified so far are grouped under GH10 family. In recent years many studies were carried out to characterize these plant xylanases and it was observed that these enzymes have limited substrate specificity (Chithra and Muralikrishna 2008). The study of Van Campenhout et al. (2007) was focused on the characterization of GH10 xylanase isolated from barley and they reported that the enzyme can release small xylose units from various substrates. GH10 xylanases, in addition to their xylolytic activity, are also active against glucose-derived substrates like aryl-cello-oligosaccharides (Charnock et al. 1997; Andrews et al. 2000).
Three-dimensional structure of xylan was first described by Atkins (1992). Subsequently, more studies were carried out using NMR spectrometry and X-ray crystallography techniques to obtain more carbohydrate structure details. It was found that under crystallized condition, the xylan backbone exhibits a threefold left-handed confirmation and the geometry showed that there is no effect on β-1,4 glycosidic bonds by the side chains of those attached. Electron diffraction pattern study confirmed the structure and the hexagonal morphology. The hydrogen at the fifth carbon atom on xylose ring has binding properties either intra- or interchain. The study also suggested that the structural confirmation of d-xylose ring indicated the di-equatorial binding property of both 1–4 and 1–3 glycosidic linkages.
The X-ray crystallographic image of endo-β-1,4-xylanases enzyme isolated from Thermoascus aurantiacus has been used to explain the general structure of GH family 10 xylanase (Fig. 7.4). It was found that the 32.5 kDa long polypeptide chain with a β/α-fold TIM barrel structure consists of eight major β strands which are arranged side by side and parallel, forming a cylinder in the center followed by eight major α helixes. In addition, around six short helixes are also found attached with the polypeptide chain. The catalytic domains of family 10 xylanases are found in cylindrical shape and the overall side view resembles a ‘Salad bowl’ (Fig. 7.5). In this configuration, the catalytic sites are seen relatively closer to the carboxyl terminus end of xylan backbone. Since the molecular weight of these xylanase is higher, they always exhibit low degree of oligosaccharide polymerization. Both the disulfide bond and salt bridges present on the xylan structure improve its thermostability. In the overall structure, the top phase of the molecule which is β-barrel side has larger than the bottom face, the α-β turns. This was due to the elaborate architecture of β–α loops (Natesh et al. 1999).
a A front view showing the (α/β) 8 TIM barrel fold, b salad bowl view exposing the substrate binding cleft of endo-1,4-β-xylanase from T. aurantiacus (Natesh et al. 1999)
Three-dimensional structure of NpXyn11A. The protein schematic is color ramped from the N terminus (blue) to the C terminus (red) (Vardakou et al. 2005)
5.2 GH11 Xylanases
GH11 xylanase unlike the other xylanase families, they only consist of endo-β-1,4-xylanases that are exclusive enzymes cleaves β-1,4-xylosidic bonds between xylose monomers. These enzymes are often sub-classified based on their PI values into acidic and alkaline xylanases (Joshi et al. 1997). Many studies demonstrated the correlation between the pH optima and the amino acid residues which are present adjacent to the catalytic site of these enzymes. In many cases, it was observed that the acidic xylanase (pH < 5) consists of asparagine, whereas in alkaline xylanase has arginine (Krengel and Dijkstra 1996; Fushinobu et al. 1998). This was also confirmed by other study which reported that arginine introduction to bacterial xylanase shifts the pH optimum to acidic range (Pokhrel et al. 2013). Heteroxylan, xylobiose, and xylotriose are the major subunits formed during GH11 xylanase-mediated xylan hydrolysis. Further hydrolytic activities on these subunits were found negligible; however, hydrolysis products such as xylotetrose, xylopentose, and xylohexose are further hydrolyzed by GH11 xylanases (Cervera-Tison et al. 2009). These enzymes cleave the xylose backbone at unsubstituted regions which are quite away from the branched xylose present at non-reducing end. It was also found that GH11 xylanases require minimum of three unsubstituted consecutive xylose residues for the primary binding and initiation of hydrolysis (Biely et al. 1997; Katapodis and Christakopoulos 2008).
Ko et al. (1992) reported the first structural description of a GH11 xylanase that was isolated from Bacillus pumilus. But his analysis was not very precise and he failed to deposit the 3D structure in protein database. To date, more than 100 GH11 xylanase 3D structures from different fungal and microbial species have been solved and made available in CAZy database. Xylanases isolated from a thermophilic fungi Thermomyces lanuginosus are widely used to explain the general structure of GH11 xylanases. The structure shows a globular protein composed of two β-sheets (β-a, β-b) with a molecular weight of 25 kDa. The outer β-sheet (β-a) is composed of five antiparallel β strands with polar and uncharged amino acids such as threonine and serine. The inner β-b sheet is made up of nine antiparallel β strands of which the outer sides are developed to catalytic sites and the inner sides are found attached with β-a forming a hydrophilic core (Torronen et al. 1994). In general, most of the GH11 xylanases are made of only a single catalytic domain and found in β-jelly roll structure. It also resembles the shape of partially closed right hand in which two β-sheets represent the ‘fingers,’ the loops between β-strands of β-b such as B7 and B8 represent the ‘thumb,’ and the twisted inner sides of β-b and α-helix represent the ‘palm’ of the hand (Gruber et al. 1998).
While comparing these two major GH families, the GH11 family consists only of ‘true xylanases’ displaying substrate specificity toward d-xylose containing xylan substrates. Their catalytic versatility is lower when compared to GH10 xylanases and the products from its action such as xylobiose and xylotriose usually required further hydrolysis by other xylanase enzymes. The GH10 xylanases can mediate hydrolysis until the release of xylose as end product. Another interesting feature of GH10 xylanases is their ability to tolerate glucose-derived to a certain extend in addition to their xylanolytic activity that exhibits in addition to their xylanolytic activity (Collins et al. 2005a). It is generally accepted that GH10 can cleave decorated regions on AX backbone and its activity is less hampered by α-l-arabinofuranosyl and acetyl or 4-O-methyl-d-glucuronate side chains present on xylan backbone, whereas GH11 xylanases are very specific in their action as they cleave only at the unsubstituted regions (Biely et al. 1997). This property is also reflected in the shape of their active binding sites, where GH10 xylanase has a shallow groove active site which has less affinity toward the unsubstituted regions and GH11 xylanases possess a cleft-shaped active site which has higher affinity toward unsubstituted consecutive xylose.
5.3 GH8 and GH5 Xylanases
In addition to the above-mentioned GH families, GH8 family xylanase is another most studied xylanase which consists of both endo- and exo-xylanases enzymes. Exo-xylanases are those enzymes which reduce xylooligosaccharides (XOS) and release xylose subunits. A very few number of endo-xylanase have been identified so far that belong to GH8 family such as xylanases isolated from Pseudoalteromonas haloplanktis (Collins et al. 2005a); GH8 exo-oligoxylanases was also isolated from Bifidobacterium adolescentis (Lagaert et al. 2007) and from Bacillus halodurans (Honda and Kitaoka 2004). In general, the diversity in substrate specificity of these enzymes is large in nature and rXyn8 isolated from P. haloplanktis is the best example which shows highest activity on mixed linkage (β-1,3- and β-1,4-bonds) homoxylan (Pollet et al. 2010). Studies on three-dimensional structure of GH8 xylanase isolated from P. haloplanktis xylanase show that it is a single-domain molecule with six (α/α)-fold barrel which is found similar to GH48 enzymes (De Vos et al. 2006). Glycoside hydrolases 5 (GH5) is another largest enzyme family which consists of more than 1800 entries in the CAZy database. Under this family, several enzymes with apparent activity on xylan have been reported. Endo-xylanases from Aeromonas punctuate (Suzuki et al. 1997) and Meloidogyne incognita (Mitreva-Dautova et al. 2006) are the major reported GH5 xylanases so far. However, certain enzymes in this group such as xylanase from H. jecorina also exhibit exo-activity. Larson et al. (2003) was the first one who reported the structure of GH5 xylanase (XynA) isolated from E. chrysanthemi. In recent years, many other X-ray analysis of GH5 xylanase has been reported. St John et al. (2009) reported X-ray crystallographic analysis of XynC from B. subtilis, which shows a multidomain protein (consist of a catalytic domain along with a C-terminal CBM) with (α/β) 8-fold barrel.
6 Catalytic Mechanisms of Endo-1,4-β-xylanase
Hydrolysis of glycosidic bonds is carried out mainly by two different catalytic residues that are present in the active site of the enzyme. Hydrolysis takes place at the anomeric center of the substrate molecules either by inversion or retention of the configuration and this has been depended on factors such as the structure and position of catalytic residues on the enzymes (Collins et al. 2005b). Many of the endo-1,4-β-xylanase exhibit a double-displacement mechanism in their operation on specific substrate molecules (Fig. 7.6). In most cases, the first acid/base catalytic residue initiate the hydrolysis by protonating the xylopyranosyl linkages between the xylan monomers and this process is termed as glycosylation. The second catalytic residue which acts as a nucleophile attacks the same linkage and results in the formation of enzyme–glycosyl intermediate while passing through an oxocarbenium-ion-like transition state. In consecutive steps which are also termed as deglycosylation, the first catalytic residue exhibits basic properties by activating the incoming water molecule and abstracting a proton from it. The activated water molecule readily attacks the anomeric carbon of the enzyme–glycosyl complex formed in the previous step and release the product with an α-configuration at the anomeric carbon (Sidhu et al. 1999; Chiku et al. 2008).
A hypothetical model of catalytic mechanism of GH xylanase. The double-displacement reaction mechanism for retaining glycosidases (Koshland mechanism). Modified from Paës et al. (2012)
7 Factors Affecting Endo-1,4-β-xylanase Functionality
7.1 GH Family Origin
Among the above-mentioned families, endo-xylanases GH10 and GH11 are the most studied. Endo-xylanase enzymes with relatively higher molecular weight are grouped into family F/10 and they are found with cellulose-binding domain and a catalytic domain that are usually connected by linker peptide. Enzyme belonging to this family also found to have a (β/α)-fold TIM barrel structure (Biely et al. 1997), whereas the family G/11 xylanase is generally of low molecular weight and is further classified into two major groups based on their PI value. Similarly, β-xylosidases, another major xylolytic enzyme is grouped under GH classification system into various families such as 3, 39, 43, 52, and 54. Studies showed that β-xylosidases from the families 3, 39, 52, and 54 use a retaining mechanism to hydrolyze xylooligomers, while those from family 43 mediate the hydrolysis by inverting the anomeric configuration. Advanced bioinformatics tools such as Basic Local Alignment Search (BLAST) and pair-wise alignments of the protein sequences have been used to identify the xylanase enzymes and compared with closely related enzymes within the families. The sequences of family 10 xylanase are generally used to identify mutually exclusive enzymes using BLAST search. Further confirmations are also made using X-ray crystallographic studies. A continuous update to all these GH family classifications is provided by the carbohydrate-active enzyme database (CAZy) since there is a direct relationship between the protein folding and the sequence. This system is universally accepted as it can provide structural features of enzymes that help to predict their sole substrate specificity; it helps to identify and reveal the evolutionary relationships between the enzyme groups, and also act as a convenient tool to derive mechanistic information.
7.2 Carbohydrate-Binding Modules (CBMs)
The X-ray crystallographic studies on the xylanase structure reveal that most of them have a modular structure composed of a catalytic site and one or more carbohydrate-binding modules (CBMs) that are interconnected by flexible linkages (Kulkarni et al. 1999; Subramaniyan and Prema 2002; Collins et al. 2005b). A CBM is defined as a specially arranged amino acid sequence with high affinity to carbohydrate molecules within a carbohydrate-active hydrolytic enzyme. These unique sites are designed to bind with specific polysaccharides on plant tissue and mediate the structural damage by enzymatic hydrolysis (Bolam et al. 2001; Boraston et al. 2002). Similar to the catalytic modules of glycoside hydrolases, CBMs are classified into 64 families according to their amino acid sequence similarity and this classification is available in CAZy database (Tomme et al. 1994). A study of Boraston and his group reported a detailed overview about the structure and functions of CBMs (Boraston et al. 2000). However, a very few members of GH11 xylanases carry CMB. Among these only a few of them have been experimentally demonstrated, but others are hypothetically explained based on their sequential similarities with well-studied ones. It have been also reported that certain CBMs from Xyl-11 are specific for cellulose and this characteristic feature helps the xylanase to localize the substrates indirectly even when they are in close association with cellulose. Major CBMs of known structure which are linked to xylanases are presented in Table 7.7. Xylanases isolated from fungi such as Penicillium funiculosum XynB (Brutus et al. 2004) and Neocallimastix patriciarum XynS20 (Liu et al. 2008) are found to have CMB1 modules. It was also reported that the occurrence of a C-terminal CBM1 in Xyl-11 isolated from Podospora anserina found to have significantly improved ability to reduce the sugar during the hydrolysis of wheat straw (Couturier et al. 2010). Figure 7.7 shows the three-dimensional structure of NpXyn11A which belongs to GH11. Some GH11 xylanases, apart from the CBMs, also consist of dockerin domains that are key elements which help in the formation of cellulose-binding multi-enzyme complex known as cellulosomes. These enzyme complexes are originally observed in Clostridium thermocellum (Lamed et al. 1983). In addition to cellulolytic enzymes, cellulosomes are also found with hemicellulase such as xylosidases, xylanases, and arabinofuranosidases. Other research showed also the similar results in XynC isolated from Fibrobacter succinogenes S85 (Marrone et al. 2000).
Spatial conservation of residues paving the catalytic cleft of Xyl-11 (Paës et al. 2012)
7.3 Xylanase Enzyme Inhibitors
Almost all the plant species have developed a self-defense mechanism as response to the pathogenic microbial attack by hydrolases enzymes. Xylanase inhibition by the proteinaceous inhibitors is one among those defense response widely seen in both soft- and hardwood plants. A numerous studies have been carried out to understand this defense mechanism since 1990s and it was found that Triticum aestivum xylanase inhibitor (TAXI) and xylanase inhibitor protein (XIP) are the two major groups of proteins responsible for Xyl-11 in specific (Debyser et al. 1999; Rouau et al. 2006). Apart from the above two groups, the third group of inhibitor known as thaumatin-like xylanase inhibitor (TLXI) also been isolated from wheat. It was also found that none of these xylanase inhibitors are effective against plant origin xylanase enzymes (Dornez et al. 2010a, b). Occurrence of these inhibitors seems to be a significant hindrance in xylanase-mediated industrial process such as bread making and brewing as they inhibit effective hydrolysis of substrates (Sorensen and Sibbesen 2006). Therefore, it has been an important topic of research to understand the mode of action of these inhibitors. Sensitivity of GH11 xylanases to proteinaceous inhibitors (XIP-1, TAXI-1, and TLXI) are as given in Table 7.7 (Fig. 7.8).
a Structure of the inhibition complex between P. funiculosum Xyl-11 (in green) and XIP-I inhibitor (in blue). XIP-I inhibiting loop is in red, catalytic residues are in yellow. b Structure of the inhibition complex between B. subtilis Xyl 11 (in green) and TAXI-IA inhibitor (in blue). His374 residue is in red, catalytic residues are in yellow (Paës et al. 2012)
7.3.1 Triticum Aestivum Endo-xylanase Inhibitor (TAXI)
These are proteins with pI values of ≥8.0, which occur in two different molecular forms known as A and B (Gebruers et al. 2008). Form A is seen as full-length proteins with molecular mass near to 40 kDa, whereas the form B consists of two polypeptide chains (a longer chain and a shorter chain of size 28–30 and 10–12 kDa, respectively) that bound together by disulfide bonds (Croes et al. 2009). These inhibitors are seen largely in wheat grains during the early stages of grain development and specifically inhibit GH11 xylanase disrespect to their origin (Tables 7.8, 7.9). Further analysis of originally identified TAXI inhibitor proteins revealed that it is a mixture of two different proteins of different pI values. They are named as TAXI-I and TAXI-II and have different inhibition activities on GH11 endo-xylanase isolated from A. niger (Gebruers et al. 2004). Other research carried out by Raedschelders et al. (2004) identified and isolated the genes coding for these two inhibitor proteins from ray and barley grains and based on this study Fierens et al. (2004) developed a Pichia pastoris-based expression system for the large-scale production. By 2005, two more new TAXI class named TAXI-III and TAXI-IV were also identified (Igawa et al. 2005). TAXI-type inhibitors are found to be very specific in their inhibitory characteristics. The available research data shows that they inhibit only GH11 endo-xylanases of bacterial and fungal origin. However, the subclass TAXI-II was turned to be an exception, as they are unable to inhibit GH11 endo-xylanases such as XynBc1 and ExlA isolated from B. cinerea and A. niger, respectively (Brutus et al. 2005).
7.3.2 Endo-xylanase Inhibiting Proteins (XIP)
Basically, these are glycosylated single-chained proteins with molecular weight of 30 kDa and its pI value falls between 8.0 and 9.0. The crystallography study shows that it has similar structural features of GH18 with typical (α/β) 8-fold TIM barrel shape (Payan et al. 2003). However, they are found to be passive on GH18 enzymes such as chitinases due to the high degree of mismatch at its enzyme biding site. In general, these inhibitors consist of two independent enzyme-binding sites that mediate binding with almost all GH10 and GH11 endo-xylanases that are fungal origin. However, GH11 endo-xylanase from Neocallimastix patriciarum (Payan et al. 2004) and GH10 endo-xylanase from Aspergillus aculeatus (Flatman et al. 2002) are found to be insensitive to XIP. Similar to TAXI-type inhibitors, XIP-type inhibitors are also categorized into different polymorphic families based on the recent proteomics studies. Followed by identification of Xip-I (the most common inhibitor in the XIP) class, Durand et al. (2005) identified Xip-II class in T. turgidum ssp. In addition to this, Xip-III type inhibitor encoding gene was discovered in T. aestivum and was observed that these genes are transcribed very rarely under stressed leaves. Later on, several other XIP-type genes were also identified known as Xip-R genes, OsXip that are of plant origin (Takahashi-Ando et al. 2007). Biely et al. (2008) confirmed the existence of XIP-type inhibitors in maize leaves and roots during the first week after germination.
7.3.3 Thaumatin-like Xylanase Inhibitor (TLXI)
Thaumatin-like endo-xylanase inhibitors (TLXI) consist of a single amino acid chain with molecular weight of 18 kDa. This type of inhibitor was first discovered in wheat by Fierens et al. (2007). These proteins have high pI and usually exhibit glycosylation at varying extents that found to have a very high stability (no denaturation was observed at 100 °C at extreme pH ranges even after 120 min). These inhibitors are grouped into class 5 pathogenesis-related proteins as they show very high sequential similarity with certain antifungal proteins (Trudel et al. 1998). Even though there are no crystallographic studies that have been reported on structure of TLXI inhibitor, the structure has been predicted and shows the existence of five disulfide bridges and three β-sheets (Fierens et al. 2009). The inhibitory mechanism of these proteins was found to be specific against Xyl-11. The xylanase inhibitors were recently reviewed by Dornez et al. (2010a, b).
8 Recombinant Endo-1,4-β-xylanase
The advances in molecular biology and genetic engineering opened up several new applications of recombinant DNA technology. The new opportunity for the construction of GM microbial strains with selected enzyme machinery was one of the greatest breakthroughs. On the basis of this, more efficient endo-1,4-β-xylanase-producing microorganisms were constructed to improve the biodegradation of hemicellulosic residues and thus find many industrial applications. In this section, we will highlight in detail the recent advance in recombinant xylanase production. Moreover, studies on xylanase gene regulation and biosynthesis at molecular level and various tools applied to enhance the novel characteristic of xylanase enzymes will be discussed.
Several xylanases have been isolated and identified from various sources. However, very limited numbers of these enzymes are found to be suitable for industrial applications due to their poor performance in harsh process conditions. Poor thermostability is the biggest issue associated with most of the xylanases. Enzymes with higher thermostability can enhance the efficiency of industrial process with an increased residual time. Therefore, there has been a huge demand for thermostable xylanases in pulp and paper industry as they are applied during pulp bleaching process which usually carried out at temperatures more than 90 °C. Most of the xylanases we have today are not able to withstand this high temperature. However, highly thermophilic xylanases have been reported from thermophilic bacteria such as G. thermoleovorans and Thermotoga sp. (Sharma et al. 2007). Alkali stability is also another major problem associated with industrial applications. It was observed that most of the highly thermophilic xylanases are less stable under alkaline conditions which limit their potential use in industries. Most of the xylanases reported so far have optimal activity at either acidic or neutral pH (Wang et al. 2010). Exceptional cases are also reported, such as xylanases from Bacillus halodurans which exhibit high alkali tolerance (Mamo et al. 2006), but these enzymes are found to have poor thermostability. Cellulose-free xylanase is another desirable characteristics required in pulp bleaching industries to mediate efficient hydrolysis without disturbing the cellulosic content of the pulp. Studies also shows that low molecular weight xylanases are more efficient in pulp bleaching as they can easily diffuse into pulp fibers. In practical and economical point of view, growing these extremophilic microorganisms in large scale is not feasible. Moreover, it is difficult to have a single enzyme with all desired characteristics. Conventional enzyme stabilization techniques such as helix capping and salt bridging (Puchkaev et al. 2003) have been practiced to improve the enzyme characteristics. However, these methods are found to be ineffective when the enzyme is required in large quantities. To overcome these problems and enhance the large-scale production for the industrial application, recombinant xylanases were introduced using gene cloning and expression techniques. rDNA technology has been widely used nowadays to make alteration or modifications in existing wild-type xylanase to improve their specific characteristics such as thermostability and alkaline stability. A number of xylanase coding genes from different microbial species have been isolated and expressed successfully in suitable expression hosts. The expression level of xylanase gene in host cell usually depends on several factors such as cloning strategies used, host cell, biosynthesis mechanism, gene regulation, fermentation media, and many bioprocessing parameters. Bacterial and fungal species are generally used to express xylanases in large scale. The cloning vector selection is usually carried out based on the expression host and a wide variety vectors have been introduced in last few decades. Even though the expression rate is higher in recombinant strain, the enzyme activity is usually less than in the native producer strain. Several reasons have been identified for this activity loss and the intercellular accumulation of the enzyme due to the lack of post-translational modification is suggested to be the key reason (Schlacher et al. 1996).
8.1 Cloning and Expression of Fungal Endo-1,4-β-xylanase Gene
It has been more than three decades since filamentous fungi are used in the production of xylanase enzyme. Fungal cells are the widely used biofactories in large-scale production of xylanases and have been considered as more potent xylanase producers as they are capable of secreting much higher amounts of xylanolytic enzymes in fermentation media than bacteria or yeast (Bergquist et al. 2002; Fang et al. 2008). Large numbers of fungal strains are widely used for xylanases production such as many species from Penicillium, Aspergillus, Fusarium, Trichoderma, and Disporotrichum. However, in many of these fungi, the difficulty in isolating pure form of xylanase was the major problem since they generally exhibit a low amount of cellulase activity. Recombinant DNA technology has been widely applied to overcome this drawback and production of xylanase expression systems which are free of cellulase activity which improved its acceptability in industrial applications. Apart from the bacterial expression hosts, filamentous fungi are also attractive hosts for xylanase expression. It is because of their ability to secrete the proteins into the fermentation media in large quantities. Aspergillus and Trichoderma species are the widely used expression hosts in homolog expression system. Kitamoto et al. (1999) successfully overexpressed XynF1 gene in A. oryzae under a strong TEF1 promoter in glucose-based submerged fermentation. XynF1, XynF3, and XynG2 are another important xylanase genes expressed efficiently in A. oryzae under stronger promoter called P-No 8142 in similar fermentation conditions. Aspergillus niger is another attractive host for recombinant xylanases production due to its ability to produce and secrete protein in high capacity up to 30 g L−1 in the fermentation media (Hessing et al. 1994). Xylanase coding gene from T. reesei was successfully expressed in A. niger cD15 stain under G6P-dehydrogenase promoter and glaA terminator which is isolated from Aspergillus awamori (Rose and van Zyl 2002). This recombinant xylanase retained its 75 % activity even after 3 h of incubation at 50 °C. The optimal condition of the enzyme activity was observed at temperature between 50 and 60 °C and pH 5–6. Levasseur et al. (2005) also reported on the potential use of Aspergillus niger as host to express xynB under promoter sequence isolated from Aspergillus nidulans. They showed that, A. nidulans can be used as efficient host for xylanase expression and production. Kimura et al. (1998) expressed xynG1 in A. nidulans under its own promoter showed that the xylanase expression was induced in the presence of glucose in the media. Filamentous fungi enable extracellular secretion of heterologous protein which facilitates the downstream processes and ease the purification and characterization of the produced enzyme. Moreover, most genes from fungi have introns and filamentous fungi are able to recognize them and express with minimal miscoding. In addition, fungal proteins are glycosylated and the protein expression in another filamentous fungi results in similar glycosylation pattern. For expressing highly hemophilic xylanase, many researches have been selected filamentous fungi because of above-mentioned advantages when compared to other expression hosts like bacteria. Table 7.10 shows major fungal xylanase expression system.
Yeast-based heterologous gene expression is considered as an excellent alternative system when based on their ease scaling up compared to fungi. The presence of eukaryotic post-translational modification mechanism is one of the main reasons behind the wide application of yeast expression systems in industry (Cheng et al. 2005). S. cerevisiae is one of the important hosts for xylanase production by cloning of xylanase cDNA obtained by RT-PCR through crossing over. The presence of intron sequence (lake of proper splicing of introns) that hinders the xylanase gene expression is the major disadvantage of these expression systems (Moreau et al. 1992). It has been also observed that a significant increase in xylanase expression in yeast transformed with the xynA gene without intron (26.2 U mL−1) when compared to the expression from the gene with intron (16.7 U mL−1) (Li and Ljungdahl 1996). It is also worthy to note that Chavez et al. (2004) made the first successful attempt to clone and express the xynA gene which is isolated from P. purpurogenum in S. cerevisiae. Isolated xynA gene that consists of eight introns regions was spliced correctly and was integrated on yeast chromosome. The Xln A gene was then expressed under transcriptional control (XlnR and CreA) where xylose or xylan acts as inducer and glucose acts as repressor (van Peij et al. 1997). The methanotropic yeast Pichia pastoris is another widely used xylanase expression host which has many advantages over S. cerevisiae such as improved secretion efficiency, ease to attain high cell density culture in inexpensive media, and relative ease in scaled up fermentation process (Cereghino and Cregg 2000). Moreover, the presence of strong and regulatory alcohol oxidase (AOX I and AOX 11) promoters which are involved in methanol metabolism promotes the overexpression of xylanase gene upon the addition of methanol in the fermentation media (Tsai and Huang 2008). Table 7.11 summarizes some examples of successful cloning of fungal xylanase in yeast.
8.2 Cloning and Expression of Bacterial Endo-1,4-β-xylanase Gene
La Grange et al. (2000) successfully co-expressed xylanase genes (xln B from Bacillus pumilus along with xyn2 isolated from T. reesei) in yeast S. cerevisiae. E coli-based expression systems are widely used to express the xylanase. Endo-1,4-β-xylanase coding genes isolated from A. usamii were cloned in E. coli BL-21 with the help of PET-28a plasmid vector and expressed successfully and a maximum xylanase activity of 49.4 U mg−1 was reported. The gene expression was induced by IPTG and the protein was expressed with 6-X histidine tags that facilitate the purification of expressed protein (Zhou et al. 2008a). Similarly, for the xylanase gene from B. circulans Teri-42 when cloned in E. coli DH5-alpha using pUC19 plasmid vector, the xylanase expression was found to be reduced. However, when the same gene is expressed in B. subtilis using plasmid pBA7, 14-fold increase in expression was observed. Therefore, Bacillus became more attractive hosts compared to E. coli and thus used widely in heterogeneous protein production. Bacillus sp. used in industrial fermentation are non-pathogenic in nature and in bacillus-based expression, the proteins are usually secreted out of the cell. Moreover, they are gram-positive bacteria and therefore, they do not produce endotoxins (lipopolysaccharides). These features could be of great advantage by reducing some purification steps in terms of breaking the cells and extraction from large number of intracellular proteins.
8.3 Overexpression in Escherichia coli
For the past several decades, E. coli has been recognized as the ideal platform for expression of recombinant proteins. Factors such as rapid growth on inexpensive media, non-pathogenicity, easy to make gene modification without effecting the cell growth, easy methods required for isolation and purification of expressed proteins made E. coli the most successful expression host for industrial use. In spite of these advantages, due to the lake of specific post-translational modification such as glycosylation and disulfide bond formation, many heterologous proteins were not successfully expressed in E. coli. Xylanase was one of those which require N-glycosylation during post-translation stage, whereas E. coli only can perform O-glycosylation. However, many exceptional cases such as successful expression of a glycosylated β-xylosidase gene isolated from thermophilic fungi P. thermophile have been reported (Teng et al. 2011). It was also observed that the endo-1,4-β-xylanase expression levels are significantly higher in gram-positive hosts such as Lactobacillus species and B. subtilis compared to E. coli as they can perform N-glycosylation.
In general, the recombinant xylanases expressed in E. coli are accumulated in the cytoplasm or in periplasmic place in the absence of a proper secretory channel on cell wall (Schlacher et al. 1996). However, in many cases, xylanase activity has also been detected outside the cell (Karlsson et al. 1997; Ebanks et al. 2000). Xylanase expression rate in E. coli mainly depends on the efficiency of transcription of xylanase gene which is cloned under promoter sequence. Even though there are several cloning vectors and host strains have been introduced, eukaryotic gene expression in E. coli is unable to carryout due to the absence of a functional promoter sequence. Basaran et al. (2001) isolated β-xylanase gene from Pichia stipites and successfully cloned and expressed in E. coli. The recombinant enzyme activity was low (4 U mg−1) when compared to the activity original enzyme activity (30 U mg−1) in wild stain. Although a variety of cloning vectors have been used, pET expression vector systems were reported to be the most efficient for protein expression in E. coli. Xylanase genes isolated from different sources are cloned under the control of a strong T7 promoter which is isolated from bacteriophage genome. This promoter remains silent until T7 RNA polymerase is produced by the host cell genome. Table 7.12 shows the major cloning vectors used in E. coli-based expression system.
Ogasawara et al. (2006) amplified xynIII gene from T. reesei genomic DNA using a RT-PCR and was cloned and expressed in E. coli JM109 using pAG9-3 plasmid vector. When the xylanase gene expression was induced by IPTG, they could produce 26 mU mL−1 of xylanase activity in cellular extract, which was very low. However, the XynIII activity increased by about 500-fold (13.2 U mL−1) compared to the activity in soluble supernatant, when the xylanase inclusion bodies were refolded in 8 mM urea solution. Zhou et al. (2008a) reported the maximum xylanase activity up to 49.4 U mg−1 when they cloned and expressed xynII cDNA isolated from A. usamii in E. coli BL21-Codon plus (DE3) RIL strain using pET-28a (+) expression vector. The xylanase protein expressed is attached to 6X-His-tag which ease protein purification by affinity chromatography. Jun et al. (2008) also used the same pET vector to clone and express a β-xylanase gene, xyn2, isolated from T. reesei in E. coli and reported the enhanced xylanase activity of 650 U mg−1. In some cases, the heterologous cloning and expression results in the change in original characteristics of xylanase gene. For example, a non-glycosylated XynA gene when expressed in E. coli exhibited significant loss in its substrate specificity when compared to the glycosylated xylanase from the wild strain P. stipites. The stability of the enzyme also decreased due to the change in optimum temperature rage. However, improved thermostability and pH stability have been reported when Xyn2 from T. reesei was expressed in E. coli. Table 7.13 shows the properties of major endo-β-1,4-xylanase enzyme expressed in E. coli.
9 Improvement of Endo-1,4-β-xylanase Characteristics
In general, enzymes are classified based on their conserved regions on them. Xylanases are specific in their identity due to the random variations in amino acid sequence and it was found that some very specific stretches of amino acids defines the structural and functional behavior of enzyme. Existing xylanases can be improved in terms of their activities and other novel characteristics such as substrate specificity, thermostability, and pH stability using protein engineering approaches. Critical amino acid exchanges are usually practiced in order to enhance the characteristics of existing xylanase enzymes. Therefore, before amino acid sequence modification a detailed studies on the native xylanase amino acid sequences are required to fully understand the structural and functional behaviors such as the presence of catalytic domains, specificity of substrate-binding sites, occurrence of thermo-stabilizing domains, and the presence of covalent/non-covalent interactions. The significance of a particular acid or a set of amino acids for novel enzyme characteristics can be identified easily using specific amino acid inhibitors such as N-bromosuccinimide (tryptophan), phenylmethylsulfonyl fluoride (serine), and iodo acetate (cysteine). It was also found that cysteine plays critical role in the formation of covalent glycosyl enzyme intermediate during most bacterial xylanase activity. Many of the family 10 xylanases possess three different amino acids such as Phe4, Trp6, and Tyr 343. They were identified as crucial in protein folding and enzyme stability even under extreme conditions. Torronen and Rouvinen (1997) studied the significance of asparagines and aspartic acid on the pH optima of family 11 xylanases. The studies conducted on xylanase isolated from B. circulans revealed that the presence of asparagine at 52nd position of amino acid sequence increased the thermal fluctuations at unstable regions. However, when it is substituted with tyrosine, the overall thermostability of the enzyme exhibited significant increased (Joo et al. 2011). It was also reported that the catalytic activity of B. circulans xylanase increased when the Asn at position 35 is mutated to Asp (Li and Wang 2011). Other study showed also a significant increase in thermostability (10–15 °C) as a result of point mutation (aspartic acid to asparagine) at position 56 (Yin et al. 2010). However, in many other cases, the point mutation was reported to have negative effects such as loss in substrate-binding affinity, alkaline stability, and thermostability. Simpson et al. (1999) reported the complete loss of carbohydrate-binding module affinity toward the xylan when the arginine was mutated to glycine. In case of B. circulans xylanase, change in pH optima was reported when R73V and Q167M were mutated (Yang et al. 2008). Table 7.14 comprises significant achievements made in xylanase protein engineering to increase enzyme stability.
10 Industrial Production of Endo-1,4-β-xylanase
Although many microbial species such as bacteria, fungi, and actinomycetes have been reported as xylanase producers, a few number of strains are acceptable by the industrial society. To meet the current and forecasting market demands, xylanase production need to be increased by many folds through efficient production strategies. Therefore, optimizations of production and purification processes are the key to obtain maximum xylanase yield either by solid-state fermentation (SSF) or submerged fermentation (SmF) (Narang and Satyanarayana 2001). Nowadays, the most of commercial xylanases production processes are carried out by SmF as shown in Table 7.15. Cultivation media plays important role in the production of xylanases in industrial scale. A suitable and economically viable media formulation which supports the maximum xylanase production is usually required for large-scale production. Therefore, abundantly available natural agriculture residues are widely used for xylanase production. Agricultural wastes such as corn cob, con stover, and wheat straw were successfully used as suitable substrates for fungal xylanases production (Damiano et al. 2003; Gupta et al. 2000).
10.1 Physical and Nutritional Parameters Affecting Endo-1,4-β-xylanase Production
In SSF, biomass and xylanase productions depend on several factors such as the type of microorganism, inoculums size, temperature, oxygen diffusion, the period of cultivation, the substrate used and its pretreatment, size, water activity, relative humidity, pH, and many other factors. On the other hand, for SmF, the temperature, pH, agitation, and dissolved O2 are the most critical physical parameters affecting xylanase production in SmF. Most of the xylanase-producing microorganisms reported so far are either mesophilic or thermophilic (Coughlan and Hazlewood 1993). It was observed that these microbes show highest xylanase activity and optimum growth at temperature between 40 and 70 °C. However, those extremophilic such as Thermotoga sp., Dictyoglomus, and Geobacillus exhibit even higher optimum temperature range of up to 80 °C (Ko et al. 2011; Sharma et al. 2007; Qureshy et al. 2002). Xylanases of fungal origin usually are found less thermophilic in nature. Even though some exceptional cases have been reported, the optimum temperature for most fungal xylanases is below 55 °C. This has been one of the major drawbacks of fungal xylanase in industrial application. On the other hand, bacterial xylanases exhibit higher thermostability. The pH of the fermentation media is another major factor effecting the xylanase production as well as the enzyme stability. Fungal xylanases usually are highly alkaliphilic as they can be active at wide ranges of pH, usually between 5.0 and 10.0, whereas the bacterial xylanases have quite narrow pH spectrum (Dhillon et al. 2000). Aeration and agitation rates during fermentation are the other crucial factors in case of xylanase production process. Studies show that xylanase production by Bacillus sp. is enhanced at an agitation rate of 200–250 rpm and 30 % and above dissolved O2 saturation (Anuradha et al. 2007; Sanghi et al. 2009). In cretin fungal species which exhibit mycelia growth pattern, higher agitation rate results in poor growth as well as less xylanase production due to the high sheer force. Studies made by Archana and Satyanarayana (1998) and Sharma et al. (2007) deal with the optimization of physical parameters effecting xylanase production.
In addition, carbon and nitrogen sources and their concentration in the fermentation media are found to be not only effecting the xylanase production but also the localization of enzymes. The fungal xylanase producers are able to utilize wide verity lignocellulosic-derived substrates such as corn cob, corn straw, wheat bran, rice straw, and sugarcane bagasse. For production in SSF system, these substrates are used as they are cheaper and easily available raw materials (Chauhan et al. 2007). However, in bacterial cells in SmF, simpler and readily utilizable nutrients are used. Organic nitrogen sources such as tryptone, peptone, yeast extract, and soybean meal are widely used as they enhance the xylanase production as well as biomass. Kapoor et al. (2008) observed a significant increase in xylanase production which was observed when the Bacillus sp. SSP-34 was provided with yeast extract in combination with peptone. In similar study, xylanase production by Bacillus sam-3 was increased up to 25-folds higher when the inorganic nitrogen source was replaced by soybean meal. Use of corn steep powder also significantly enhanced xylanase production by T. reesei (Lappalainen et al. 2000). Apart from these major nutrients, the incorporation of micronutrients such as trace elements, vitamins, and amino acids improved the xylanase production to great extend in cases of thermo-anaerobe-mediated xylanase fermentation. In cases of recombinant strains, the xylanase expression greatly depends on the gene inducers present in the fermentation media. IPTG is the most widely used chemical inducer for ‘lac’-based expression system (Mamo et al. 2007). However, in large-scale production, IPTG is not applicable due to its high cost.
In cases of recombinant xylanase, the expression of xylanase genes usually depends on several aspects such as biosynthesis mechanism, gene regulation, vector construction, expression host used, expression media, and other growth parameters during the fermentation. For commercial applications, xylanases have to be ideally produced quickly and in large quantities. The conventional production strategies have been followed are found to be less effective to meet the growing need in the present xylanase market. Therefore, many researches are going on for developing recombinant strains for xylanase production. However, a very limited number of these recombinant strains are studied further for the scaling up of xylanase production in pilot scale. Hence, more investigations on this area are required and it is very significant to have more efficient and economically viable process to reduce the cost of xylanase production.
10.2 Optimization of Endo-1,4-β-xylanase Production
Media screening and medium composition optimization are usually the preliminary steps involved in bioprocess optimization which are always carried out in shake flask level to select the most suitable medium formulation. Both physical and chemical parameters effecting the cell growth as well as xylanases production are monitored and optimized at this stage with a number of experiments. Basar et al. (2010) observed that the recombinant xylanase production in E. coli is a growth-associated process. They observed a rapid growth of E. coli, reaching the stationary phase after 16 h and the maximal xylanase production up to 324.72 U mL−1 was obtained in defined medium. They also reported the effects of different concentrations of glucose in growth media on cell growth and xylanase production. Farliahati et al. (2009) reported the xylanase expression in recombinant E. coli DH5α in different media compositions (defined, semi-defined, and complex). Optimization study of pH and agitation ranges also carried out in shake flask cultivation mode. From these studies, they observed the highest xylanase production (up to 2122 U mL−1) in defined media at initial pH 7 and agitation speed of 200 rpm. A suitable cultivation strategy for the production of two truncated thermostable recombinant xylanases (XynlAN and XynlANC), isolated from Rhodothermus marinas, was developed by Karlsson et al. (1998). They found that the fed-batch cultivations of E. coli strain BL21 (DE3) with a controlled exponential glucose feed led to high specific production of the recombinant proteins and addition of complex nutrients such as Tryptone Soya Broth (TSB) to the media exhibits positive impact on cell growth rate and cell productivity. Single addition of either lactose or isopropyl-thio-go-galactoside (IPTG) was used for induction. In lactose-induced cells, the production of recombinant xylanase was delayed for approximately 30 min compared to those induced with IPTG, but the specific product levels were comparable at 3 h after induction. Huang et al. (2006) cloned a xylanase gene from B. subtilis into E. coli and found that the xylanase distributions in extracellular, intracellular, and periplasmic fractions were in ratio 22.4, 28.0, and 49.6 %, respectively, with an optimal enzyme production at pH 6.0 and 50 °C. The superiority of defined medium for xylanase production using recombinant bacteria was also reported by Farliahati et al. (2009).
Catabolic repression of the xylanases biosynthesis is found to be a common phenomenon usually in the presence of glucose in culture medium. When glucose is abundant in the grown medium, the repression of catabolic enzyme synthesis takes place due to the gene regulation. When the microbe can easily absorb the readily available glucose in the medium, they will not utilize for other complex nutrients by spending energy to metabolize complex nutrients. Xylanase production reported to be reduced due to the catabolic repression that makes the xylanase coding gene in inactive mode. In another study by Basar et al. (2010) they enhanced xylanase production using recombinant E. coli by optimizing the media composition and other growth factors. Two basal mediums (defined media and complex media) have been used and found out that the defined media gives the highest growth as well as the xylanase production under experimental conditions. At 20 % enhanced dissolved oxygen tension (DOT) in lab scale bioreactor, they increased xylanase production up to 1784 mL−1.
10.3 Application of Response Surface Methodology (RSM) in Optimization of Xylanase Production
The conventional media optimization method based on step-by-step process varying one variable at a time (OFAT) while keeping the other variables constant is a time-consuming process and less reliable in many cases. To overcome this problem, statistical approach using experimental designs was applied. Response surface methodology (RSM) is the most reliable statistical application that has been used for optimization of fermentation media. Optimizations of physiological and nutritional growth factors are found to be very important procedures to reduce the production cost of xylanases in industrial scale. Many researches have been carried out based on RSM which used to understand and evaluate the significant interaction between the physiological and nutritional factors effecting the cell growth and product formation. A prior knowledge about these growth factors and their interactions is necessary for the designing of a more realistic model. Hence, many of the RSM studies are found to be based on the results obtained from OFAT method. Selection of these growth parameters has been done by checking their significance level using a 2FFD Pareto chart before proposing an experimental design. Most significant factors were then used as variables in central composite design (CCD) (Farliahati et al. 2010). The effect of various growth parameters in production of xylanase enzyme was studied by Mullai et al. (2010). In his study, statistical experimental designs (CCD) were constructed with the help of statistical application MINITAB 14. Plackett–Burman (PB) design for seven different variables, NH4NO3, KH2PO4, CaCl2, MgSO4·7H2O, FeSO4·7H2O, MnSO4·7H2O, and NaCl at two levels +1 and −1 (higher and lower level, respectively) and the statistical significance were studied by determining the F-value. The optimum levels of selected variables from the Plackett–Burman design were investigated using central composite design (CCD). The response was measured in terms of xylanase activity. Counter plots and response surface 3D graphical representations generated could show the interactions of different variables under experimental conditions. Garai and Kumar (2012) optimized the xylanase production by A. candidus in SSF. They studied the effect of various nitrogen sources in the production media and they found that xylanase production was enhanced when the ammonium nitrate was used as sole nitrogen source. In the same study, applying RSM-based BoxBehnken design, the researcher also optimized the physical parameters such as time of incubation, temperature, and moisture content which effects xylanase activity. The crude xylanase recovered at the end of solid-state fermentation was used for the scarification of ammonia-treated lignocellulose materials to study the substrate-binding assay. Among the various substrates used, corn comb was found releasing the maximum reduced sugar (438.47 mg g−1) when it was treated with crude xylanase enzyme. In RSM-based optimization, xylanase production was optimized by Box–Behnken design. Initially, Planket–Berman design was employed to identify the key ingredients and process conditions which exhibit the most significant effect on the xylanase expression. In the later stage, using Box–Behnken design, the optimal level of the identified key ingredients in production media as well as the optimum range of process parameters such as cultivation time, pH, temperature, and agitation are identified. Using this approach 6.83-fold increase in xylanase production was obtained under optimized conditions when compared to that of initial un-optimized conditions.
Solid-state fermentation (SSF) is also widely used for large-scale production of xylanase enzymes. It is basically designed to mimic the natural habitat of microbes that are less likely to grow in submerged fermentation (SMF) conditions. In SSF, the solid bed is used as nutrient source as well as the mechanical support for cell growth. A. niger, T. reesei, and T. koningii are the major fungal species used to produce xylanase under SSF conditions (Dhiman et al. 2008; Garg et al. 1998). However, xylanase production by bacteria is commonly carried out in submerged fermentation conditions where the physical parameters such as temperature, pH, and dissolved O2 level involved are controlled to the optimum levels with more ease, and therefore an enhanced production can be achieved. Esteban et al. (1982) reported that the large-scale production of xylanase in SmF is the growth-associated process. However, several exceptional cases were also been reported such as xylanase production using Bacillus sp. (Dey et al. 1992). Later study by Bocchini et al. (2002) reported the production of highly thermostable cellulase-free xylanase in SMF using B. circulans. Similar study was also carried out by Sharma et al. (2007) who used Geobacillus thermoleovorans AP7 to produce highly thermo-alkali-stable xylanase. The switch to xylanase-producing recombinant E. coli is seen as an economic strategy toward higher productivity and easier downstream purification. Sandhu and Kennedy (1984) reported a large-scale production of xylanase by transforming E. coli with recombinant plasmids carrying a 1,4-β,d-xylanase gene from B. polymyxa. In another study, xylanase gene isolated from A. usmani was cloned in pET-28a expression vector and when it is expressed in E. coli BL21, the maximum xylanase activity was found to be 49.4 U mg−1. The gene expression was induced by IPTG and the xylanase was found expressed with 6-X histidine tag that facilitates an easy purification of expressed protein (Zhou et al. 2008b). A very successful cultivation strategy for the production of two truncated thermostable recombinant xylanases (XynIAN and XynIANC), isolated from Rhodothermus marinas, was developed by Karlsson et al. (1999). They found that the fed-batch cultivations of E. coli strain BL21 (DE3) with a controlled exponential glucose feed led to high specific production of the recombinant proteins with additional complex medium. Highest xylanase production in defined medium was achieved by Farliahati et al. (2009). In this study, xylanase production reached up to 2122.5 U mL−1 at pH 7.4 and when (NH4)2HPO4 was used as the main nitrogen source. They also reported that the xylanase production was a growth-associated process as the enzyme secretion was greatly dependent on biomass concentration for strain E. coli DH5α.
Biswas et al. (2010) used mutant IITD3A of Melanocarpus albomyces for the production of high titer cellulose-free xylanase in 14-L stirred tank bioreactor where an optimized (RSM based) production media and process parameters are used. The authors reported the xylanase activity reached up to 415 U mL−1 after 24 h by pH recycling strategy (between 7.8 and 8.2) during the production phase. They also reported that the xylanase production is significantly influenced by aeration and agitation. A 5.2-fold increase (overall volumetric productivity of 22,000 U L−1 h−1) in activity was observed when the cells exhibited pellet form at an agitation speed of 600 RPM.
Kumar et al. (2009) optimized the xylanase production in lab scale stirred tank bioreactor using mycelial fungal strain T. lanuginosus MC 134. Fermentation media contains corn cob as a sole source of carbon which was inoculated with T. lanuginosus MC 134 and observed the production in terms of xylanase activity over a period of 9 days at various process parameters. A highest xylanase activity, 2009 U mL−1, was obtained after 6 days of fermentation. However, a significant decrease (40 %) in productivity was observed in bioreactors when compared to shake flask culture. Similarly, xylanase production by several microbial strains using corn cob was reported and from these studies it was clear that the production largely depends on rheological factors in bioreactors and the type of extraction methods used. Ruanglek et al. (2007) successfully cloned and expressed XylB (564 bp) encoding endo-1,4-β-xylanase obtained from A. niger BCC14405 in Pichia pastoris under AOX1 promoter. A maximum of 7352 U mg−1 of xylanase-specific activity was obtained in 2 L lab scale bioreactor with BMGY/BSM medium under exponential feeding strategy. During the batch phase which was aimed to achieve HCDC, the temperature and pH were maintained at 30 °C and 5.0, respectively. Agitation was maintained in the range of 600–1000 rpm. Fed-batch phase was started after 16–20 h by feeding 67 % (v/v) glycerol plus 1 % (v/v) trace mineral solution in exponential manner. Table 7.16 shows examples for xylanase production in lab and pilot-scale submerged fermentation by different fungal strains.
10.4 pH Optima and Stability of Endo-1,4-β-xylanase Enzymes
In general, endo-1,4-β-xylanases are active at wide pH ranges and their pH optima depend on the producer organism and the physiochemical properties of the substrate molecules. Fungal xylanases are usually acidophilic in nature. However, the optimal pH for bacterial xylanases is usually above 5.5. Endo-1,4-β-xylanase from A. kawachii reported as the most acidophilic xylanase which has a very low pH optimum of 2. On the other hand, xylanase from Bacillus sp. 41 M1 reported as the most alkalophilic xylanase which has pH optimum 9.0 (Nakamura et al. 1993). In certain cases, different pH optimums were observed in different xylanases secreted by same organism. For example, XynI and XynII produced by H. jecorina were reported to have pH optimums 3–4 and 5–5.5, respectively. This showed that the pI values of the enzymes are correlated with the amino acid residues at various positions on xylanase. However, among Xyl-11 family, those are acidophilic in nature possess an Asp residues at position 33 even though there are very rare exceptions such as A. kawachii IFO 4308 xylanase which has pH optimum close to 5.0 (Zhu et al. 1994). However, many alkalophilic xylanases such as Xyn11 from A. versicolor (Carmona et al. 1998), XynC from Uccinogenes sp. (Zhu et al. 1994) also displayed an Asp at the same position. Similarly, studies on P. griseofulvum Xyl-11 reported that it displays a Ser residue at position 44 and mutating this residue resulted significant changes in pH optimum of the enzyme (Andre-Leroux et al. 2008).
There have been many different explanations for the factors determining the optimal pH value of xylanases. However, it believes that certain amino acid residues of xylanase structure play key role in the determination of the optimal pH value of the enzyme. In case of acidic pH optima, the Asp/Glu residues near the catalytic cleft become protonated and it decreases the electrostatic repulsion between the substrate and enzyme, whereas in alkaline xylanases, instead of acidic residues more alkaline residues such as arginine are seen. de Lemos Esteves et al. (2005) experimented various mutations in Streptomyces sp. S38, and observed a pH optimum shift from 6.0 to 7.5, which enhanced the xylanase enzyme activity several folds higher than the enzyme from the wild type. The pI values of catalytic domain present on xylanase play a key role in its pH activity profile. Many recent studies reveals that, among GH11 xylanase, the residues those contribute positive charges and hydrogen bonds to the catalytic domain help in increasing or decreasing the pI values depending upon the electrostatic interaction between the substrate molecules (Joshi et al. 1997). Apart from these residues, factors such as occurrence of salt bridges are also reported to have significant effects on pH optima (Hakulinen et al. 1998).
10.5 Endo-1,4-β-xylanase of Extremophilic Origin
Considerable progress has been made in the isolation of extremophilic microorganisms and their successful cultivation in the laboratory. Commercial applications of the xylanases demand highly thermostable enzyme which can work effectively under elevated temperature. Many advantages such as reduced contamination risk and faster reaction rates have been proposed for the use of thermophiles in biotechnology processes. In general, parameters such as temperature, pH, and chemical and enzymatic stability are important for the industrial applicability of any enzyme. Recently, a simple model, based on the thermodynamic and kinetic parameters for inactivation of the enzyme, has been presented for predicting (1) thermostabilities in the presence of substrate and (2) residual activities of enzymes in harsh processing environments. The study of extremophiles and their enzymes can extend the present understanding of protein chemistry in addition to expanding the potential applications of biocatalysts. Studies of alkaliphiles have led to the discovery of a variety of enzymes which exhibit some unique properties. Biological detergents contain enzymes such as alkaline proteases and/or alkaline cellulases from alkaliphiles. Commercial production became economical only after the discovery of alkaliphilic Bacillus-producing CGT with enhanced pH stability. Alkaline xylanases have gained importance due to their application for the development of eco-friendly technologies used in paper and pulp industries. The enzymes are able to hydrolyze xylan which is soluble in alkaline solutions.
10.5.1 Alkaliphilic and Acidophilic Endo-1,4-β-xylanase
The pH tolerance of microbial endo-1,4-β-xylanase enzymes depends on the natural growing environment of the microbial species. Endo-1,4-β-xylanase-producing alkalophilic microbes grow optimally at higher pH above 9.0 and those produce acidophilic xylanases grow between pH 1.0–5.0 (Kimura et al. 2000). These organisms are usually isolated from various natural environments such as soil, decomposing organic matters, paper pulp industry wastes, and many other sources. Bacillus sp. C-59-2 was the first reported xylanase-producing alkaliphilic microorganism (Nakamura et al. 1993). Since then, there have been a number of reports on xylanases production using alkaliphilic and acidophilic microbes. However, the studies indicated that the intracellular xylanase produced by many of these microbes may not be adapted to extreme environmental growth conditions of their hosts. A large number of alkaliphilic microbes found to produce xylanase at optimal pH close to neutral although their activity can be retained in alkaline conditions. Many exceptional cases are also reported such as XylB from Bacillus sp. which has pH optima of 9.0–10 (Gessesse and Mamo 1998). Many acidophilic xylanases of fungal and bacterial origin have also been reported in last two decades. Cryptococcus sp. S-2 and Penicillium sp. 40 (Kimura et al. 2000) are the most studied extremely acidophilic (pH optima 2–3) xylanase producers.
10.5.2 Psychrophilic Endo-1,4-β-xylanase
Even though the xylanase-producing microorganisms are abundant in nature, only a very less number of endo-1,4-β-xylanase-producing psychrophilies have been identified so far. Gram-negative bacteria such as Pseudoalteromonas haloplanktis TAH3a (Collins et al. 2005a) and Flavobacterium frigidarium (Humphry et al. 2001); gram-positive Clostridium sp. PXYL1 (Akila and Chandra 2002); fungal species such as Alternaria alternate, Penicillium sp., and Phoma sp.; and yeast sp Cryptococcus adeliae (Petrescu et al. 2000) are the major psychrophilic organisms reported to be endo-1,4-β-xylanase producers. Almost all these microbial species are isolated from Antarctic regions. The best studied two psychrophilic xylanases, which are from Pseudoalteromonas haloplanktis and Cryptococcus adeliae, were classified under GH 8 and GH10, respectively. The (α/α) 6 barrel structure of these structures is described by Van Petegem et al. (2003). The limited studies revealed that these enzyme groups have very poor stability due to the reduced number of salt bridges present on it but higher catalytic activity even at very low temperature. Georlette et al. (2004) also observed that psychrophilic xylanases are characterized by up to 10-folds higher activity even at mild cool temperature of 5 °C when compared to mesophilic xylanase activity. Moreover, the psychrophilic xylanase displayed higher catalytic and retained 60 % of maximum activity at very low temperature. However, the psychrophilic xylanases are totally unstable at high temperature (temperature 55 °C) and exhibited 12 times shorter half-life compared to mesophilic xylanases (Collins et al. 2005b).
10.5.3 Thermophilic Endo-1,4-β-xylanase
A large number of thermophilic and hyper-thermophilic (optimal growth temperature 50–80 °C and more than 80 °C, respectively) microbes which are able to produce xylanases have been isolated from various sources such as thermal hot springs, deep marine fields, self-heating organic debris, etc. Almost all of these enzymes were grouped under families 10 and 11. Extremophile microbial species such as Caldicellulosiruptor sp. (Luthi et al. 1990), Rhodothermus marinus (Abou Hachem et al. 2000), and Bacillus stearothermophilus (Khasin et al. 1993) are the major producers of thermophilic family 10 xylanases, whereas family 11 thermophilic xylanases are less common and have been isolated from extremophiles such as Bacillus strain D3 (Connerton et al. 1999), Chaetomium thermophilum growing medicinal mushrooms in Egypt as a new strategy for cancer treatment, dictyoglomus thermophilum (McCarthy et al. 2000), paecilomyces variotii (Kumar et al. 2000), and thermomyces lanuginosus (Schlacher et al. 1996).
Optimum activity range of these thermophilic xylanases was found between 65 and 80 °C, whereas the xylanases from Thermotoga sp. exhibit optimal activity at 105 °C and pH 5.5. X-ray crystallographic structure and amino acid sequential analysis of many of these thermophilic xylanase indicated very close similarities with mesophilic xylanase and probable reason for the higher thermostability could be the minor changes in amino acid alignment. Many mutagenic studies also have indicated that minor modifications in mesophilic aminoacid array enhanced the thermostability to many folds (Winterhalter and Liebl 1995).
10.5.4 Features Responsible for Endo-1,4-β-xylanase Thermostability
Thermostability comparison made between the enzymes are usually misleading since in most cases, the enzymes compared do not share the same experimental conditions such as incubation time, concentration of protein, nature, and concentration of the substrate. Many researchers believe that the high thermostability of xylanases is due to the presence of a network of certain charged amino acid residues near to the catalytic sites. These residues are usually found coupled with the disulfide bridges between β-strand and the α-helix. However, many recombinant thermostable xylanases are expressed in E. coli and are found to have thermostability ranging from 70 to 80 °C. When analyzing their protein structure, comparatively longer N-terminal ends are observed which may account for its higher thermostability (Morris et al. 1996). Many other analyzes also propose that the thermostability is due to the presence of highly flexible α-helix and C-terminal regions where the protein unfolding is generally initiated (Kumar et al. 2000). However, several other factors have also been identified that explain the xylanase thermostability such as follows:
-
Presence of disulfide bridges which may strengthen the whole protein framework would give more stability to the enzyme.
-
Presence of ion and aromatic pairs and their interactions may favor to more stable framework.
-
Strategic position of water molecules on the framework, since the readily available water bindings are required for the catalytic activity even at higher temperature.
-
Complex folding nature of enzyme. The higher the folding complexity, the higher the stability.
-
Occurrence of oligomerization and the presence of certain aromatic residues result in the hydrophobic patches which allows stronger monomeric interactions enhancing the enzyme stability.
Certain GH11 endo-1,4-β-xylanases display disulfide bridges at position between β-strand and α-helix. However, these enzymes do not exhibit higher thermostability, whereas endo-1,4-β-xylanase produced by Dictyoglomus thermophilum exhibits thermostability even though that do not have any disulfide bridges on their structure. Therefore, the presence of disulfide bridges does not necessarily account for the thermophilic characters of the enzymes (Table 7.17).
10.5.5 Improvement of Endo-1,4-β-xylanase Thermostability
In last two decades, thermostability of xylanase enzymes was continuously improved through the protein engineering approaches. In most of these studies, the researches considered to modify the mesophilic xylanase enzyme structure to design a new structural model similar to the thermophilic xylanase. Arase et al. (1993) used chemical random mutagenesis in XYNA from B. pumilus and this strategy was successful to increasing the enzymes’ half-life from 4 to 12 min at 57 °C. In a similar study, the specific activity and optimum temperature of Xyl1 from Streptomyces sp. S38 was enhanced from 2100 to 2500 IU mg−1 with the shift of the optimal temperature from 57 to 66 °C (Georis et al. 2000). Other study showed that the replacement of the N-terminal end of mesophilic xylanase xlnB from Streptomyces lividans with those from thermostable xylanases TfxA from T. fusca by random gene shuffling and observed a significant increase in thermostability (Shibuya et al. 2000). Introduction of two disulfide bridges, one between B9 strand and the α-helix and another between N and C-terminal ends, also enhanced the thermostability of Tx-xyl (Paës and O’Donohue 2006). The authors also reported that the occurrence of residues that met at N-terminal end brought negative impact on xylanase thermostability and temperature optimum which are reduced by 10-folds and 5 °C, respectively. From these engineering strategies applied in xylanases protein engineering, researchers concluded that:
-
The α-helix on xylanase structure seems very sensitive to thermal unfolding—making these are more rigid and may enhance the thermostability of the enzyme. However, it may also diminish the catalytic activity (Muilu et al. 1998).
-
N-terminal end plays key role in thermostabilization in GH11 xylanase—linking the N-terminal extremity to A2 strand or C-terminal extremity enhanced the thermostability of xylanase. Introduction of an additional complete β strand by mutagenesis also beneficial for improved thermostability.
-
Presence of disulfide bridges does not necessarily enhance the thermophilic behavior of the enzymes. Therefore, addition of these bridges does not always bring positive impact on enzyme thermostability.
11 Industrial Applications of Endo-1,4-β-xylanase
Industrial production of xylanases has been grown significantly during last few decades based on their increased application. Xylan can be converted into many useful products such as xylitol and furfurol through enzymatic hydrolysis (Subramaniyan and Prema 2002). The hydrolysis could be also carried out through acid treatment process which is relatively easy, fast, and cheap. However, the application of this method in industries is limited due to the formation of toxic end substances that may have cross reactions with product of interest (Beg et al. 2001). Therefore, enzymatic hydrolysis method is widely applied even though it is a slow process but more efficient and eco-friendly. During industrial application, the xylolytic enzymes are not commonly applied in pure form but in combination with other hydrolytic enzymes to achieve better results. The industrial applications of xylanases cover different areas and the list of applications increased by time and thus increases the market demand of xylanases accordingly.
11.1 Animal Feed Industry
Xylanases are widely used in animal feed manufacturing. They are usually applied in combination of other hydrolytic enzymes such as pectinase, protease, cellulose, lipase, glucanase, alginase, and phytase. These enzymes are used to reduce the viscosity of raw materials used in animal feeds. The complex arabinoxylan that is present on the cell wall of most grains may reduce the effective digestion and absorption of nutrients in poultry. Studies on this area have reported that the xylanase-treated feed has better digestibility and nutrient absorption rate when compared to raw feeds (Bedford and Classen 1992). In animal feed industries, the feed conversion rate (FCR) is defined as the ratio of feed consumed to the body weight gained by the animal. A lower FCR means improved feed digestibility and nutrient absorption. Xylanase enzymes are widely incorporated in poultry feed to reduce the FCR. It have been reported that the addition of endo-xylanase is isolated from Acidothermus cellulolyticus in poultry feed and observed a significant decrease in FCR, indicating an improved digestibility of feed (Clarkson et al. 2001). Similar results were observed when endo-xylanase is isolated from Neocallimastix patriciarum added to canola seed after oil extraction and applied for cattle feeding. The digestibility and stability of nutritive value of silage for cattle are found to be enhanced when it was treated with xylanase along with probiotic lactobacillus. The xylose formed by the de-polymerization of hemicellulose was fermented by the probiotic lactobacillus and thus enhanced the feed nutritive value. It was also reported that xylanase supplementation in the animal diet improved the animal growth through the better nutrition absorbability and enhancement of immune response as well as the gut micro-flora (Gao et al. 2008). In other study, Vandeplas et al. (2010) reported that when T. viride xylanase is incorporated in poultry feed, it increased nutrient digestion by 60 % and resulted in a significant increase in chicken weight. Pretreatment of forage crops with xylanase leads to better digestion in cattle and these enzymes are often found in synergy with other cellulolytic enzymes. The assimilation of ruminant feed was also improved many folds by xylanases addition (Yang and Xie 2010).
11.2 Food and Beverages Industry
Food and beverages industries use also xylanases in wide range of applications. Xylanases are capable of hydrolyzing the hemicellulose in wheat that enhances the water absorption of dough at leavening level in bread making process. This could possibly enhance the fermentation rate and as a result the softness of the bread and volume was increased (Rouau 1993). Xylanase cocktails are often used to reduce the stickiness of the dough thus enhance the crumb (Goesaert et al. 2005; Butt et al. 2008). Other study showed also that under specific process condition, incorporation of xylanase in the dough increased the shelf life of bread and also improved the volume up to 24 % (Verjans et al. 2010). However, enzyme selection is critical in these conditions since the cereals used for bread making often exhibit the xylanase inhibition by TAXI, XIP, or TLXI (Gebruers et al. 2004). Pasta processing is another area where xylanases are widely used as solubilizing agent of durum wheat semolina without changing its rheological properties (Ingelbrecht et al. 2001; Brijs et al. 2004). It was also reported that xylanase enzymes can improve the separation of wheat flour into starch and gluten (Frederix et al. 2004). However, in certain cases, action of Xyl-11 does not favor the separation due to high viscosity caused by the un-extractable arabino xylanase hydrolysis. In such conditions, synergetic dosages of Xyl-10 and Xyl-11 are used as it can mediate-improved agglomeration of gluten protein even though the wheat quality is very poor (Van Der Borght et al. 2005). Table 7.18 shows some examples of commonly used xylanase preparations for the baking industry. Xylanases are widely used also in juice, beer, and wine making industries. The enzymatic breakdown of fruits and vegetables hemicellulose can increase the clarity and stability of juice and wine. Xylanases can be used along with several other enzymes such as pectinase and amylase to improve the quality of beverage products in terms of aroma, taste, clarity, and stability (Wong et al. 1988). It was reported that up to 27 % decrease in insoluble fibers was achieved in juice clarification by addition of xylanase enzyme isolated from Sclerotinia sclerotiorum (Olfa et al. 2007). In beer production, partial hydrolysis of arabinoxylan found in the barley cell wall gives a middy viscous appearance to the fermented beer. In such cases, various xylanase enzyme groups are used to increase the filtration performance of beer wort by hydrolysis of viscous AXs fractions. Xylanase inhibitors present in the malt are the major hinderance to xylanase action. The xylanase activity is retained with the help of other synergetic enzymes used in the process (Debyser et al. 1997).
11.3 Paper and Pulp Industries
Endo-1,4-β-xylanase enzymes play significant role in paper and pulp industry since they facilitate an eco-friendly bio-bleaching of wood pulp by reducing the use of chemical bleaching agents such as chlorine and chlorine dioxide. For the past few decades, kraft pulp bleaching industry is recognized as the biggest area of xylanase application. Several studies reported that the removal of lignin content can be achieved by xylanase enzymes far better than by other lignin-degrading enzymes (Lundgren et al. 1994). Use of chemical bleaching agents in conventional pulp bleaching methods cause serious environmental pollution issues and this drove the researcher’s attention on xylanase enzymes (Subramaniyam and Prema 2002). Table 7.19 shows major endo-1,4-β-xylanase supplier in pulp bleaching industry. Application of endo-1,4-β-xylanase enzymes in pulp bleaching can reduce using chlorine as well as the total energy requirement for this process. Kulkarni et al. (1999) observed that xylanase pretreatment of kraft pulp lowers the usage of chemicals by 10–20 %. The concept of TCF (total chlorine free) bleaching technologies was also recently introduced to make the process more eco-friendly. However, cellulase-free endo-1,4-β-xylanase is required in this process since the cellulose enzyme may degrade the cellulose which is the main target of the industry (Bajpai and Bajpai 2001). Relatively lower molecular weight of xylanase enzymes helps them to penetrate more deep into the lignocelluloses substrates and facilitate better hydrolytic actions (Beaugrand et al. 2005; Beg et al. 2001). Higher thermostability and alkaline stability are other key features that make xylanase more favorable for bio-bleaching industry (Buchert et al. 1995). The mechanisms of pulp bleaching by xylanase have been studied and many researchers believe that xylanase hydrolysis takes place at lignin xylan interaction and thus results in a loosely packed cellulosic framework (Shatalov and Pereira 2007). However, it was recently reported the potential use of Aspergillus oryzae MDU-4-based fungal xylanase for effective deinking of newspaper pulp where a significant improvement in optical properties such as brightness up to 57.9 % ISO with decreased (211 ppm) residual ink concentration (Chutani and Sharma 2015).
11.4 Biofuel Production
The next-generation fuel production is based on renewable sources such as lignocellulosic biomass. As first step, the complex lignocellulosic biomass needs to be breakdown to a mixture of relatively smaller units made of hexoses and pentoses or their monosaccharides that are further fermented to ethanol or other biofuels (Margeot et al. 2009). The process of converting the lignocellulosic feedstocks into biofuels usually involves the following steps:
-
Enzyme hydrolysis: the critical step involved where the hydrolysis of lignocelluloses into various sugar monomers by the action of cocktail of hydrolytic enzymes.
-
Fermentation: production of biofuel by the utilization of sugar monomers formed by various microbial species, usually carried out by different bacterial and yeast strains depends on the desired type of biofuel.
-
Distillation: separation and purification of biofuel produced.
The controlled hydrolysis of lignocelluloses using enzymatic cocktail to produce the desirable products is the challenge involved in this process. Xylolytic enzymes, such as Endo-1,4-β-xylanase in particular, are very essential in biomass hydrolysis as they initiate progressive cell wall in the assembly and thus mediate the cell wall penetration by other cellulolytic enzymes (Beaugrand et al. 2005).
11.5 Other Applications
Apart from the traditional applications in pulp beaching industry, xylanases have been used in several other areas as well. The broad spectrum of xylanase application includes the saccharification of xylan to xylose, plant cell protoplasting, coffee mucilage liquefaction, vegetable softening, extraction of plant pigments and oils, etc. (Kulkarni et al. 1999). In laundry industries, endo-1,4-β-xylanase was effectively used to remove the plant-based stains. The stain-removing capacity of family 11 endo-xylanase was improved when it chemically linked with cellulose-binding domain of cellulases. In textile industries, xylanase enzymes can be used to process the plant fiber as they can decrease the lignin content and improve the color and quality of fibers (Csiszár et al. 2001; Battan et al. 2012). Until now, xylanase applications in pharmaceutical industries are very limited. However, it is used in some dietary supplements in combination with other enzymes (Polizeli et al. 2005). Commercial products such as chewing gum contain xylitol and it can be easily produced by hydrolysis of xylan using xylanase enzymes.
11.6 Future Perspectives
The current list of highly diversified applications of xylanases clearly demonstrates the increased demand of this type of enzymes. In addition, the continuous improvement of the enzyme catalytic properties increases the potential uses of this enzyme by adding many new industrial applications. The improvement of catalytic properties of the xylanases by shifting the optimal operation conditions of the enzyme and by isolation of new potential xylanases from extremophilic microbes are further expressed in suitable mesophilic host to industrialize the process. In addition, with the increased knowledge about protein engineering, it become possible also to increase enzyme stability and kinetic properties using different approaches such as changing of amino acid sequence, creation of sulfide bridge within the molecule, and changing the enzyme 3D configuration. All these will help to increase the potential application of xylanases under extreme operation conditions.
However, some recent studies were skeptic about the future application of xylanases in biofuel sector based on the current dramatic drop of oil price which make biofuel non-competitive alternative to fossil fuel in economic point of view. However, xylan biorefinery is not limited to fuel market but the biohydrolysis products of xylan such as pentose oligomers and monomers are also used as intermediate in the production of many biochemicals. Moreover, the increased public awareness about the negative impacts of chemicals on the environment, ecosystem, and human and animal health increased the pressure on chemical industries to replace the current chemical-based catalytic processes to a biobased and eco-friendly processes especially in paper/pulp, leather, and detergent industries. In addition, there is significant increased demand for xylanases application in food and feed industries for the production of safe and high-quality product. These all together support our current estimation of the continuous increase of xylanases demand in many industrial applications.
References
Abou Hachem M, Karlsson EN, Bartonek-Roxa E, Raghothama S, Simpson PJ, Gilbert HJ, Williamson MP, Holst O (2000) Carbohydrate-binding modules from a thermostable Rhodothermus marinus xylanase: cloning, expression and binding studies. Biochem J 345:53–60
Alejandro SH, Jesús VE, del María CMH, María EHL (2007) Purifcation and characterization of two sugarcane bagasse-absorbable thermophilic xylanases from the mesophilic Cellulomonas Xavigena. Microbiol Biotechnol 34:331–338
Akila G, Chandra TS (2002) A novel cold-tolerant Clostridium strain PXYL1 isolated from a psychrophilic cattle manure digester that secretes thermolabile xylanase and cellulase. FEMS Microbiol Lett 219:63–67
Andrade SV, Polizeli MLTM, Terenzi HF, Jorge JA (2004) Effect of carbon source on the biochemical properties of β-xylosidases produced by Aspergillus versicolor. Process Biochem 39:931–938
Andre-Leroux G, Berrin JG, Georis J, Arnaut V, Juge V (2008) Structure-based mutagenesis of Penicillium griseofulvum xylanase using computational design. Proteins 72(4):1298–1307
Andrews SR, Charnock SJ, Lakey JH, Davies GJ, Claeyssens M, Nerinckx V, Underwood M, Sinnott ML, Warren RA, Gilbert HJ (2000) Substrate specificity in glycoside hydrolase family 10. Tyrosine 87 and leucine 314 play a pivotal role in discriminating between glucose and xylose binding in the proximal active site of Pseudomonas cellulosa xylanase 10A. J Biol Chem 275(30):23027–23033
Anuradha P, Vijayalakshmi K, Prasanna ND, Sridevi K (2007) Production and properties of alkaline xylanases from Bacillus sp. isolated from sugarcane fields. Curr Sci 92(9):1283–1286
Arase A, Yomo T, Urabe I, Hata Y, Katsube Y, Okada H (1993) Stabilization of xylanase by random mutagenesis. FEBS Lett 316(2):123–127
Archana A, Satyanarayana T (1998) Cellulase-free xylanase production by thermophilic Bacillus licheniformis A99. Indian J Microbiol 38:135–139
Atkins EDT (1992) Three-dimensional structure, interactions and properties of xylans. Xylanas Xylanases 7:39–50
Avalos OP, Noyola TP, Plaza IM, Torre M (1996) Induction of xylanase and β-xylosidase in Cellulomonas flavigena growing on different carbon sources. Appl Microbiol Biotchnol 46:405–409
Bai Y, Wang J, Zhang Z, Yang P, Shi P, Luo H, Meng K, Huang H, Yao B (2010) A new xylanase from thermoacidophilic Alicyclobacillus sp. A4 with broad-range pH activity and pH stability. J Ind Microbiol Biotechnol 37(2):187–194
Bajpai P, Bajpai PK (2001) Development of a process for the production of dissolving kraft pulp using xylanase enzyme. Appita J 54(4):381–384
Bajaj BK, Singh NP (2010) Production of xylanase from an alkali tolerant Streptomyces sp. 7b under solid-state fermentation, its purification, and characterization. Appl Biochem Biotechnol 162(6):1804–1818
Balakrishnan H, Srinivasan MC, Rele MV, Chaudhari K, Chandwadkar AJ (2000) Effect of synthetic zeolites on xylanase production from an alkalophilic Bacillus sp. Curr Sci 79:95–97
Basar B, Shamzi MM, Rosfarizan M, Puspaningsih NNT, Ariff AB (2010) Enhanced production of thermophilic xylanase by recombinant Escherichia coli DH5α through optimization of medium and dissolved oxygen level. Int J Agric Biotechnol 12:321–328
Basaran P, Hang YD, Basaran N, Worobo RW (2001) Cloning and heterologous expression of xylanase from Pichia stipitis in Escherichia coli. J Appl Microbiol 90(2):248–255
Battan B, Dhiman SS, Ahlawat S, Mahajan R, Sharma J (2012) Application of thermostable xylanase of Bacillus pumilus in textile processing. Indian J Microbiol 52(2):222–229
Beaugrand J, Paës G, Reis D, Takahashi M, Debeire P, O’Donohue MJ, Chabbert B (2005) Probing the cell wall heterogeneity of micro-dissected wheat caryopsis using both active and inactive forms of a GH11 xylanase. Planta 222:246–257
Bedford MR, Classen HL (1992) Reduction of intestinal viscosity through manipulation of dietary rye and pentosanase concentration is effected through changes in the carbohydrate composition of the intestinal aqueous phase and results in improved growth rate and food conversion efficiency of broiler chicks. J Nutr 122:560–569
Beg QK, Bhushan B, Kappor M, Hoondal G (2000) Production and characterization of thermostable xylanase and pectinase from Streptomyces sp. QG-11-3. J Ind Microbiol Biotechnol 24:396–402
Beg QK, Kapoor M, Mahajan L, Hoondal GS (2001) Microbial xylanases and their industrial applications: a review. Appl Microbiol Biotechnol 56(3–4):326–338
Bergquist P, Te’o V, Gibbs M, Cziferszky A, de Faria FP, Azevedo M, Nevalainen H (2002) Expression of xylanase enzymes from thermophilic microorganisms in fungal hosts. Extremophiles 6(3):177–184
Berrin JG, Juge N (2008) Factors affecting xylanase functionality in the degradation of arabinoxylans. Biotechnol Lett 30(7):1139–1150
Berrin JG, Williamson G, Puigserver A, Chaix JC, McLauchlan WR, Juge N (2000) High-level production of recombinant fungal endo-beta-1,4-xylanase in the methylotrophic yeast Pichia pastoris. Protein Expr Purif 19(1):179–187
Bhalerao J, Patki AH, Bhave V, Khurana I, Deobagkar DN (1990) Molecular-cloning and expression of a xylanase gene from Cellulomonas sp. into Escherichia coli. Appl Microbiol Biotechnol 34(1):71–76
Bhardwaj A, Leelavathi S, Mazumdar-Leighton S, Ghosh A, Ramakumar S, Reddy VS (2010) The critical role of N- and C-terminal contact in protein stability and folding of a family 10 xylanase under extreme conditions. PLoS ONE 5(6):e11347
Biely P, Vrsanska M, Kluepfel M, Tenkanen D (1997) Endo-beta-1,4-xylanase families: differences in catalytic properties. J Biotechnol 57(1–3):151–166
Biely P, Leathers TD, Cziszarova M, Vrsanska M, Cotta MA (2008) Endo-beta-1,4-xylanase inhibitors in leaves and roots of germinated maize. J Cereal Sci 48(1):27–32
Biswas R, Sahai V, Mishra S, Bisaria VS (2010) Bioprocess strategies for enhanced production of xylanase by Melanocarpus albomyces IITD3A on agro-residual extract. J Biosci Bioeng 110:702–708
Blum DL, Li XL, Chen H, Ljungdahl LG (1999) Characterization of an acetyl xylan esterase from the anaerobic fungus Orpinomyces sp. strain PC-2. Appl Environ Microbiol 65(9):3990–3995
Bocchini DA, Alves-Prado HF, Baida LC, Roberto IC, Gomes E, Da-Silva R (2002) Optimization of xylanase production by Bacillus circulans D1 in submerged fermentation using response surface methodology. Process Biochem 38:727–731
Bolam DN, Xie HF, White P, Simpson PJ, Hancock SM, Williamson MP, Gilbert HJ (2001) Evidence for synergy between family 2b carbohydrate binding modules in Cellulomonas fimi xylanase 11A. Biochemistry 40(8):2468–2477
Boraston AB, Tomme P, Amandoron EA, Kilburn DG (2000) A novel mechanism of xylan binding by a lectin-like module from Streptomyces lividans xylanase 10A. Biochem J 350(Pt 3):933–941
Boraston AB, McLean BW, Chen V, Li A, Warren RA, Kilburn DG (2002) Co-operative binding of triplicate carbohydrate-binding modules from a thermophilic xylanase. Mol Microbiol 43(1):187–194
Borralho T, Chang Y, Jain P, Lalani M, Parghi K (2002) Lactose induction of the lac operon in Escherichia coli B23 and its effect on the o-nitrophenyl β-galactoside assay. J Exp Microbiol Immunol 2:117–123
Bridgeman TG, Jones JM, Shield I, Williams PT (2008) Torrefaction of reed canary grass, wheat straw and willow to enhance solid fuel qualities and combustion properties. Fuel 87(6):844–856
Brijs K, Ingelbrecht JA, Courtin CM, Schlichting L, Marchylo BA, Delcour JA (2004) Combined effects of endoxylanases and reduced water levels in pasta production. Cereal Chem 81:361–368
Brutus A, Villard C, Durand A, Tahir T, Furniss C, Puigserver A, Juge N, Giardina T (2004) The inhibition specificity of recombinant Penicillium funiculosum xylanase B towards wheat proteinaceous inhibitors. Biochim Biophys Acta 1701(1–2):121–128
Brutus A, Reca IB, Herga S, Mattei B, Puigserver A, Chaix JC, Juge V, Bellincampi D, Giardina T (2005) A family 11 xylanase from the pathogen Botrytis cinerea is inhibited by plant endoxylanase inhibitors XIP-I and TAXI-I. Biochem Biophys Res Commun 337(1):160–166
Buchert J, Teleman A, Harjunpa V, Tenkanen M, Viikari L, Vuorinen T (1995) Effect of cooking and bleaching on the structure of xylan in conventional pine kraft pulp. Tappi J 78(11):125–130
Butt MS, Nadeem MT, Ahmad Z, Sultan MT (2008) Xylanases and their applications in baking industry. Food Technol Biotechnol 46(1):22–31
Cannio R, Di Prizito N, Rossi M, Morana A (2004) A xylan-degrading strain of Sulfolobus solfataricus: isolation and characterization of the xylanase activity. Extremophiles 8(2):117–124
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The carbohydrate-active enzymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res 37:D233–D238
Cao Y, Qiao J, Li Y, Lu W (2007) De novo synthesis, constitutive expression of Aspergillus sulphureus beta-xylanase gene in Pichia pastoris and partial enzymic characterization. Appl Microbiol Biotechnol 76(3):579–585
Carmona EC, Brochetto-braga MR, Pizzirani-kleiner AA, Jorge JA (1998) Purification and biochemical characterization of an endoxylanase from Aspergillus versicolor. FEMS Microbiol Lett 166:311–315
Carmona EC, Fialho MB, Buchgnani EB, Coelho GD, Brocheto-Braga MR, Jorge JA (2005) Production, purification and characterization of a minor form of xylanase from Aspergillus versicolor. Process Biochem 40(1):359–364
Cereghino JL, Cregg JM (2000) Heterologous protein expression in the methylotrophic yeast Pichia pastoris. FEMS Microbiol Rev 24(1):45–66
Cervera-Tison MC, Andre-Leroux G, Lafond M, Georis J, Juge N, Berrin J-G (2009) Molecular determinants of substrate and inhibitor specificities of the Penicillium griseofulvum family 11 xylanases. Biochim Biophys Acta 1794:438–445
Charnock SJ, Lakey JH, Virden R, Hughes N, Sinnott ML, Hazlewood GP, Pickersgill R, Gilbert HJ (1997) Key residues in subsite F play a critical role in the activity of Pseudomonas fluorescens subspecies cellulosa xylanase A against xylooligosaccharides but not against highly polymeric substrates such as xylan. J Biol Chem 272(5):2942–2951
Chauhan K, Trivedi U, Patel KC (2007) Statistical screening of medium components by Plackett–Burman design for lactic acid production by Lactobacillus sp. KCP01 using date juice. Bioresour Technol 98:98–103
Chauthaiwale VM, Deshpande VV (1992) Molecular cloning and expression of the xylanase gene from Chainia in Escherichia coli. FEMS Microbiol Lett 99(2–3):265–270
Chavez R, Schachter K, Navarro C, Peirano A, Bull P, Eyzaguirre J (2004) The acetyl xylan esterase II gene from Penicillium purpurogenum is differentially expressed in several carbon sources, and tightly regulated by pH. Biol Res 37(1):107–113
Chutani P and Sharma KK (2015) Biochemical evaluation of xylanases from various filamentous fungi and their application for the deinking of ozone treated newspaper pulp. Carbohydrate Polymers 127:54–63
Cheng YF, Yang CH, Liu WH (2005) Cloning and expression of Thermobifida xylanase gene in the methylotrophic yeast Pichia pastoris. Enz Microb Technol 37(5):541–546
Chiku K, Uzawa J, Seki H, Amachi S, Fujii T, Shinoyama H (2008) Characterization of a novel polyphenol-specific oligoxyloside transfer reaction by a family 11 xylanase from Bacillus sp. KT12. Biosci Biotechnol Biochem 72(9):2285–2293
Chipeta ZA, Du-Preez JC, Christopher L (2008) Effect of cultivation pH and agitation rate on growth and xylanase production by Aspergillus oryzae in spent sulphite liquor. J Ind Microbiol Biotechnol 35:587–594
Chithra M, Muralikrishna G (2008) Characterization of purified xylanase from finger millet (Eleusine coracana-Indaf 15) malt. Eur Food Res Technol 227(2):587–597
Claassen PAM, van Lier JB, Lopez Contreras AM, van Niel EJW, Sijtsma L, Stams AJM, de Vries SS, Weusthuis RA (1999) Utilisation of biomass for the supply of energy carriers. Appl Microbiol Biotechnol 52(1999):741–755
Clarkson K, Siika-aho M, Tenkanen M, Bower BS, Penttilä ME, Saloheimo MLA (2001) Trichoderma reesei xylanase. Patent WO 0149859
Cleemput G, Hessing M, van Oort M, Deconynck M, Delcour JA (1997) Purification and characterization of a [beta]-d-xylosidase and an endo-xylanase from wheat flour. Plant Physiol 113(2):377–386
Collins T, Gerday C, Feller G (2005a) Xylanases, xylanases families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
Collins T, De Vos D, Hoyoux A, Savvides SN, Gerday C, Van Beeumen J, Feller G (2005b) Study of the active site residues of a glycoside hydrolase family 8 xylanase. J Mol Biol 354(2):425–435
Connerton I, Cummings N, Harris GW, Debeire P, Breton C (1999) A single domain thermophilic xylanase can bind insoluble xylan: evidence for surface aromatic clusters. Biochim Biophys Acta 1433:110–121
Copa-Patino JL, Broda YG, Broda P (1993) Production and initial characterization of the xylan-degrading system of Phanerochaete chrysosporium. Appl Microbiol Biotechnol 40:69–76
Coughlan MP, Hazlewood GP (1993) Beta-1,4-d-xylan-degrading enzyme-systems-biochemistry, molecular-biology and applications. Biotechnol Appl Biochem 4:259–289
Couturier M, Haon M, Coutinho PM, Henrissat B, Lesage-Meessen L, Berrin JG (2010) Podospora anserina hemicellulases potentiate the Trichoderma reesei secretome for saccharification of lignocellulosic biomass. Appl Environ Microbiol 77:237–246
Crepin VF, Faulds CB, Connerton IF (2004) Functional classification of the microbial feruloyl esterases. Appl Microbiol Biotechnol 63:647–652
Croes E, Gebruers K, Carpentier S, Swennen R, Robben J, Laukens K, Witters E, Delcour JA, Courtin CM (2009) A quantitative portrait of three xylanase inhibiting protein families in different wheat cultivars using 2D-DIGE and multivariate statistical tools. J Proteomics 72(3):484–500
Csiszár E, Urbánszki K, Szakás G (2001) Biotreatment of desized cotton fabric by commercial cellulase and xylanase enzymes. J Mol Catal B Enzym 11:1065–1072
Damiano VB, Bocchini DA, Gomes E, Silva RD (2003) Application of crude xylanase from Bacillus licheniformis 77-2 to the bleaching of Eucalyptus kraft pulp. World J Microbiol Biotechnol 19:139–144
Dautova M, Roze E, Overmars H, de Graaff L, Schots A, Helder J, Goverse A, Bakker J, Smant G (2006) A symbiont-independent endo-1,4-beta-xylanase from the plant-parasitic nematode Meloidogyne incognita. Mol Plant Microbe Interact 19(5):521–529
De Lemos Esteves F, Gouders T, Lamotte-Brasseur J, Rigali S, Frere JM (2005) Improving the alkalophilic performances of the Xyl1 xylanase from Streptomyces sp. S38: structural comparison and mutational analysis. Protein Sci 14(2):292–302
De Vos D, Collins T, Nerinckx W, Savvides SN, Claeyssens M, Gerday C, Feller G, Van Beeumen J (2006) Oligosaccharide binding in family 8 glycosidases: crystal structures of active-site mutants of the beta-1,4-xylanase pXyl from Pseudoaltermonas haloplanktis TAH3a in complex with substrate and product. Biochemistry 45(15):4797–4807
De Vries RP, Visser J, de Graaff LH (1999) CreA modulates the XlnR-induced expression on xylose of Aspergillus niger genes involved in xylan degradation. Res Microbiol 150(4):281–285
De Vries RP, Kester HC, Poulsen CH, Benen JA, Visser J (2000) Synergy between enzymes from Aspergillus involved in the degradation of plant cell wall polysaccharides. Carbohydr Res 327(4):401–410
Dean JFD, Gross KC, Anderson JD (1991) Ethylene biosynthesis-inducing xylanase. 3. Product characterization. Plant Physiol 96(2):571–576
Debyser W, Peumans WJ, Van Damme EJM, Delcour JA (1999) Triticum aestivum xylanase inhibitor (TAXI), a new class of enzyme inhibitor affecting breadmaking performance. J Cereal Sci 30(1):39–43
Debyser W, Derdelinckx G, Delcour JA (1997) Arabinoxylan solubilization and inhibition of the barley malt xylanolytic system by wheat during mashing with wheat whole meal adjunct: evidence for a new class of enzyme inhibitors in wheat. Am Soc Brew Chem 55:153–156
Degefu Y, Lohtander K, Paulin L (2004) Expression patterns and phylogenetic analysis of two xylanase genes (htxyl1 and htxyl2) from Helminthosporium turcicum, the cause of northern leaf blight of maize. Biochimie 86(2):83–90
Dey D, Hinge J, Shendye A, Rao M (1992) Purification and properties of extracellular endoxylanases from alkalophilic thermophilic Bacillus sp. Can J Microbiol 38:436–442
Dheeran P, Nandhagopal N, Kumar S, Jaiswal YK, Adhikari DK (2012) A novel thermostable xylanase of Paenibacillus macerans IIPSP3 isolated from the termite gut. J Ind Microbiol Biotechnol 39(6):851–860
Dhillon A, Gupta JK, Khanna S (2000) Enhanced production, purification and characterisation of a novel cellulase-poor thermostable, alkalitolerant xylanase from Bacillus circulans AB 16. Process Biochem 35(8):849–856
Dhiman SS, Sharma J, Battan B (2008) Industrial applications and future prospects of microbial xylanases: a review. BioResources 3(4):1377–1402
Dijkerman R, Ledeboer J, Op den Camp HJM, Prins RA, van der Drift C (1997) The anaerobic fungus Neocallimastix sp. strain L2: growth and production of (Hemi) cellulolytic enzymes on a range of carbohydrate substrates. Curr Microbiol 34(2):91–96
Dornez E, Croes E, Gebruers K, De Coninck B, Cammue BPA, Delcour JA, Courtin CM (2010a) Accumulated evidence substantiates a role for three classes of wheat xylanase inhibitors in plant defense. Crit Rev Plant Sci 29(4):244–264
Dornez E, Croes E, Gebruers K, Carpentier S, Swennen R, Laukens K, Witters E, Urban M, Delcour JA, Courtin CM (2010b) 2-D DIGE reveals changes in wheat xylanase inhibitor protein families due to Fusarium graminearum DeltaTri5 infection and grain development. Proteomics 10(12):2303–2319
Durand A, Hughes R, Roussel A, Flatman R, Henrissat B, Juge N (2005) Emergence of a subfamily of xylanase inhibitors within glycoside hydrolase family 18. FEBS J 272(7):1745–1755
Ebanks R, Dupont M, Shareck F, Morosoli R, Kluepfel D, Dupont C (2000) Development of an Escherichia coli expression system and thermostability screening assay for libraries of mutant xylanase. J Ind Microbiol Biotechnol 25(6):310–314
El Enshasy H, Hellmuth K, Rinas U (1999) Fungal morphology in submerged cultures and its relation to glucose oxidase excretion by recombinant Aspergillus niger. Appl Biochem Biotechnol 81:1–11
El Enshasy H, Kleine J, Rinas U (2006) Agitation effects on morphology and protein productive fractions of filamentous and pelleted growth forms of recombinant Aspergillus niger. Process Biochem 41:2103–2112
El Enshasy H (2007) Filamentous fungal culture—process characteristics, products, and applications. In: Yang ST (ed) Bioprocessing for value-added products from renewable resources. Elsevier Press, Amsterdam. ISBN-10: 0-444-52114-3
Emami K, Hack E (2001) Characterisation of a xylanase gene from Cochliobolus sativus and its expression. Mycol Res 105:352–359
Esteban R, Villanueva JR, Villa TG (1982) β- d-xylanases of Bacillus circulans WL-12. Can J Microbiol 28:733–739
Fang HY, Chang SM, Lan CH, Fang TJ (2008) Purification and characterization of a xylanase from Aspergillus carneus M34 and its potential use in photoprotectant preparation. Process Biochem 43(1):49–55
Farliahati MR, Mohamad R, Puspaningsih NNT, Ariff AB (2009) Kinetics of xylanase fermentation by recombinant Escherichia coli DH5α in shake flask culture. Am J Biochem Biotechnol 5(3):110–118
Farliahati MR, Ramanan RN, Mohamad R, Puspaningsih NNTM, Ariff AB (2010) Enhanced production of xylanase by recombinant Escherichia coli DH5 alpha through optimization of medium composition using response surface methodology. Ann Microbiol 60(2):279–285
Fernández-Espinar M, Pinaga F, Degraaff L, Visser J, Ramon D, Valles S (1994) Purification, characterization and regulation of the synthesis of an Aspergillus nidulans acidic xylanase. Appl Microbiol Biotechnol 42(4):555–562
Fierens E, Gebruers K, Voet AR, De Maeyer M, Courtin CM, Delcour JA (2009) Biochemical and structural characterization of TLXI, the Triticum aestivum L. thaumatin-like xylanase inhibitor. J Enzyme Inhib Med Chem 24(3):646–654
Fierens E, Rombouts S, Gebruers K, Goesaert H, Brijs K, Beaugrand J, Volckaert G, Van Campenhout S, Proost P, Courtin CM, Delcour JA (2007) TLXI, a novel type of xylanase inhibitor from wheat (Triticum aestivum) belonging to the thaumatin family. Biochem J 403(3):583–591
Fierens K, Geudens N, Brijs K, Courtin CM, Gebruers K, Robben J, Van Campenhout S, Volckaert G, Delcour JA (2004) High-level expression, purification, and characterization of recombinant wheat xylanase inhibitor TAXI-I secreted by the yeast Pichia pastoris. Protein Expr Purif 37(1):39–46
Flatman R, McLauchlan WR, Juge N, Furniss C, Berrin JG, Hughes RK, Manzanares P, Ladbury JE, O’Brien R, Williamson G (2002) Interactions defining the specificity between fungal xylanases and the xylanase-inhibiting protein XIP-I from wheat. Biochem J 365(Pt 3):773–781
Frederix SA, Courtin CM, Delcour JA (2004) Substrate selectivity and inhibitor sensitivity affect xylanase functionality in wheat flour gluten–starch separation. J Cer Sci 40:41–49
Fushinobu S, Ito K, Konno M, Wakagi T, Matsuzawa H (1998) Crystallographic and mutational analyses of an extremely acidophilic and acid-stable xylanase: biased distribution of acidic residues and importance of Asp37 for catalysis at low pH. Protein Eng 11(12):1121–1128
Gao J, Zhang HJ, Yu SH, Wu SG, Yoon I, Quigley J, Gao YP, Qi GH (2008) Effects of yeast culture in broiler diets on performance and immunomodulatory functions. Poult Sci 87:1377–1384
Garai D, Kumar V (2012) Response surface optimization for xylanase with high volumetric productivity by indigenous alkali tolerant Aspergillus candidus under submerged cultivation. Biotechnology 3:127–136
Garg AP, Roberts JC, McCarthy AJ (1998) Bleach boosting effect of cellulase-free xylanase of Streptomyces thermoviolaceus and its comparison with two commercial enzyme preparations on Birchwood kraft pulp. Enzyme Microb Technol 22:594–598
Gebruers K, Brijs K, Courtin CM, Fierens K, Goesaert H, Raedschelders G, Robben J, Sorensen JF, Van Campenhout S, Delcour JA (2004) Properties of TAXI-type endoxylanase inhibitors. Biochim Biophys Acta 1696:213–221
Gebruers K, Beaugrand J, Croes E, Dornez E, Courtin CM, Delcour JA (2008) Quantification of wheat TAXI and XIP type xylanase inhibitors: a comparison of analytical techniques. Cer Chem 85(5):586–590
Georlette D, Blaise V, Collins T, D’Amico S (2004) Some like it cold: biocatalysis at low temperatures. FEMS Microbiol Rev 28:25–42
Gessesse A, Mamo G (1998) Purification and characterization of an alkaline xylanase from alkaliphilic Micrococcus sp AR-135. J Ind Microbiol Biotechnol 20(3–4):210–214
Ghosh M, Nanda G (1994) Purification and some properties of a xylanase from Aspergillus sydowii MG49. Appl Environ Microbiol 60(12):4620–4623
Goesaert H, Gebruers K, Courtin CM, Delcour JA (2005) Purification and characterization of a XIP-type endoxylanase inhibitor from rice (Oryza sativa). J Enz Inhib 20:95–101
Georis J, Giannotta F, De Buyl E, Granier B, Frère J (2000) Purification and properties of three endo-β-1,4-xylanases produced by Streptomyces sp. strain S38 which differ in their ability to enhance the bleaching of kraft pulps. Enzyme Microb Technol 26:178–186
Gomes DJ, Gomes J, Steiner W (1994) Factors influencing the induction of endo-xylanase by Thermoascus aurantiacus. J Biotechnol 33(1):87–94
Gomez LD, Steele-King CG, McQueen-Mason SJ (2008) Sustainable liquid biofuels from biomass: the writing’s on the walls. New Phytol 178(3):473–485
Gruber K, Klintschar G, Hayn M, Schlacher V, Steiner W, Kratky C (1998) Thermophilic xylanase from Thermomyces lanuginosus: high-resolution X-ray structure and modeling studies. Biochemistry 37(39):13475–13485
Guo B, Chen XL, Sun CY, Zhou BC, Zhang YZ (2009) Gene cloning, expression and characterization of a new cold-active and salt-tolerant endo-beta-1,4-xylanase from marine Glaciecola mesophila KMM 241. Appl Microbiol Biotechnol 84(6):1107–1115
Gupta S, Bhushan B, Hoondal GS (2000) Isolation, purification and characterization of xylanase from Staphylococcus sp. SG-13 and its application in biobleaching of kraft pulp. J Appl Microbiol 88:325–334
Gupta U, Kar R (2009) Xylanase production by a thermos-tolerant Bacillus species under solid-state and submerged fermentation. Braz Arch Biol Technol 52:1363–1371
Hakulinen N, Tenkanen M, Rouvinen J (1998) Crystallization and preliminary X-ray diffraction studies of the catalytic core of acetyl xylan esterase from Trichoderma reesei. Acta Crystallogr D Biol Crystallogr 54(Pt 3):430–443
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293(3):781–788
Hessing JGM, van Rotterdam C, Verbakel JMA, Roza M, Maat J, van Gorcom RFM, van den Hondel CAMJJ (1994) Isolation and characterization of a 1,4-b-endoxylanase gene of A. awamori. Curr Genet 26:228–232
Himmel ME, Ding SY, Johnson DK, Adney WS, Nimlos MR, Brady JW, Foust TD (2007) Biomass recalcitrance: engineering plants and enzymes for biofuels production. Science 315(5813):804–807
Honda Y and Kitaoka M (2004) A family 8 glycoside hydrolase from Bacillus halodurans C-125 (BH2105) is a reducing end xylose-releasing exo-oligoxylanase. J Biol Chem 279:55097–55103
Hu YJ, Smith DC, Cheng KJ, Foresberg CW (1991) Cloning of a xylanase gene from Fibrobacter succinogenes 135 and its expression in Escherichia coli. Can J Microbiol 37(7):554–561
Huang J, Wang G, Xiao L (2006) Cloning, sequencing and expression of the xylanase gene from a Bacillus subtilis strain B10 in Escherichia coli, Bioresour Technol 97:802–808
Humphry DR, GeorgeA Black GW, Cummings SP (2001) Flavobacterium frigidarium sp. an aerobic, psychrophilic, xylanolytic and laminarinolytic bacterium from Antarctica. Int J Syst Evol Microbiol 51:1235–1243
Hung KS, Liu SM, Fang TY, Tzou WS, Lin FP, Sun KH, Tang SJ (2011) Characterization of a salt-tolerant xylanase from Thermoanaerobacterium saccharolyticum NTOU1. Biotechnol Lett 33(7):1441–1447
Igawa T, Tokai T, Kudo T, Yamaguchi I, Kimura M (2005) A wheat xylanase inhibitor gene, Xip-I, but not Taxi-I, is significantly induced by biotic and abiotic signals that trigger plant defense. Biosci Biotechnol Biochem 69(5):1058–1063
Ingelbrecht JA, Moers K, Abecassis J, Rouau X, Delcour JA (2001) Influence of arabinoxylans and endoxylanases on pasta processing and quality. Production of high-quality pasta with increased levels of soluble fiber. Cereal Chem 78:721–729
Ito K, Ikemasu T, Ishikawa T (1992) Cloning and sequencing of the xynA gene encoding xylanase A of Aspergillus kawachii. Biosci Biotechnol Biochem 56(6):906–912
Jeong MY, Kim S, Yun CW, Choi YJ, Cho SG (2007) Engineering a de novo internal disulfide bridge to improve the thermal stability of xylanase from Bacillus stearothermophilus No. 236. J Biotechnol 127(2):300–309
Joo JC, Pack SP, Kim YH, Yoo YJ (2011) Thermostabilization of Bacillus circulans xylanase: computational optimization of unstable residues based on thermal fluctuation analysis. J Biotechnol 151(1):56–65
Joshi MD, Hedberg A, McIntosh LP (1997) Complete measurement of the pKa values of the carboxyl and imidazole groups in Bacillus circulans xylanase. Protein Sci 6(12):2667–2670
Jun H, Bing Y, Zhang K, Ding X, Daiwen C (2008) Expression of a Trichoderma reesei Β-xylanase gene in Escherichia coli and activity of the enzyme on fiber-bound substrates. Protein Expr Purif 67:1–6
Kapoor M, Nair LM, Kuhad RC (2008) Cost-effective xylanase production from free and immobilized Bacillus pumilus strain MK001 and its application in saccharification of Prosopis juliflora. Biochem Eng J 38:88–97
Karlsson EN, BartonekRoxa E, Holst O (1997) Cloning and sequence of a thermostable multidomain xylanase from the bacterium Rhodothermus marinus. Biophys Acta Gene Struct Expr 1353(2):118–124
Karlsson EN, Bartonek Roxa E, Holst O (1998) Evidence for substrate binding of a recombinant thermostable xylanase originating from Rhodothermus marinus. FEMS Microbiol Lett 168(1):1–7
Karlsson EN, Holst O, Tocaj A (1999) Efficient production of truncated thermostable xylanases from Rhodothermus marinus in Escherichia coli fed-batch cultures. J Biosci Bioeng 87:598–606
Katapodis P, Christakopoulos P (2008) Enzymatic production of feruloyl xylo-oligosaccharides from corn cobs by a family 10 xylanase from Thermoascus aurantiacus. LWT Food Sci Technol 41(7):1239–1243
Khandeparkar R, Bhosle NB (2006) Purification and characterization of thermoalkalophilic xylanase isolated from the Enterobacter sp. MTCC 5112. Res Microbiol 157(4):315–325
Khasin A, Alchanati I, Shoham Y (1993) Purification and characterization of a thermostable xylanase from Bacillus stearothermophilus T-6. Appl Environ Microbiol 59(6):1725–1730
Kimura T, Kitamoto N, Kito Y, Karita S, Sakka K, Ohmiya K (1998) Molecular cloning of xylanase gene xynG1 from Aspergillus oryzae KBN 616, a shoyu koji mold, and analysis of its expression. J Ferm Bioeng 85(1):10–16
Kimura T, Ito J, Kawano A, Makino T, Kondo H, Karita S, Sakka K, Ohmiya K (2000) Purification, characterization, and molecular cloning of acidophilic xylanase from Penicillium sp. 40. Biosci Biotechnol Biochem 64(6):1230–1237
Kitamoto N, Yoshino S, Ohmiya K, Tsukagoshi N (1999) Purification and characterization of the over expressed Aspergillus oryzae xylanase, XynF1. Biosci Biotechnol Biochem 63(10):1791–1794
Ko EP, Akatsuka H, Moriyama H, Shinmyo A, Hata Y, Katsube Y, Urabe I, Okada H (1992) Site-directed mutagenesis at aspartate and glutamate residues of xylanase from Bacillus pumilus. Biochem J 288(Pt 1):117–121
Ko CH, Tsai CH, Tu J, Yang BY, Hsieh DL, Jane WN, Shih TL (2011) Identification of Paenibacillus sp. 2S-6 and application of its xylanase on biobleaching. Int Biodeterior Biodegrad 65:334–339
Kohli U, Nigam P, Singh D, Chaudhary K (2001) Thermostable, alkalophilic and cellulase free xylanase production by Thermoactinomyces thalophilus subgroup C. Enzyme Microb Technol 28(7–8):606–610
Kolenova K, Vrsanska M, Biely P (2006) Mode of action of endo-beta-1,4-xylanases of families 10 and 11 on acidic xylooligosaccharides. J Biotechnol 121:338–345
Kormelink FJ, Gruppen H, Vietor RJ, Voragen AG (1993) Mode of action of the xylan-degrading enzymes from Aspergillus awamori on alkali-extractable cereal arabinoxylans. Carbohydr Res 249(2):355–367
Krengel U, Dijkstra BW (1996) Three-dimensional structure of Endo-1,4-beta-xylanase I from Aspergillus niger: molecular basis for its low pH optimum. J Mol Biol 263(1):70–78
Kudo T, Ohkoshi A, Horikoshi K (1985) Molecular cloning and expression of a xylanase gene of alkalophilic Aeromonas sp. no. 212 in Escherichia coli. J Gen Microbiol 131(10):2825–2830
Kui H, Luo H, Shi P, Bai Y, Yuan T, Wang Y, Yang P, Dong S, Yao B (2010) Gene cloning, expression, and characterization of a thermostable xylanase from Nesterenkonia xinjiangensis CCTCC AA001025. Appl Biochem Biotechnol 162(4):953–965
Kulkarni N, Lakshmikumaran M, Rao M (1999) Xylanase II from an alkaliphilic thermophilic Bacillus with a distinctly different structure from other xylanases: evolutionary relationship to alkaliphilic xylanases. Biochem Biophys Res Commun 263(3):640–645
Kumar KS, Manimaran A, Permaul K, Singh S (2009) Production of β-xylanase by a Thermomyces lanuginosus MC 134 mutant on corn cobs and its application in biobleaching of bagasse pulp. J Biosci Bioeng 107(5):494–498
Kumar PR, Eswaramoorthy S, Vithayathil PJ, Viswamitra MA (2000) The tertiary structure at 1.59 Å resolution and the proposed amino acid sequence of a family-11 xylanase from the thermophilic fungus Paecilomyces varioti Bainier. J Mol Biol 295(3):581–593
Kumar V, Satyanarayana T (2011) Applicability of thermo-alkali-stable and cellulase-free xylanase from a novel thermo-halo-alkaliphilic Bacillus haloduransin producing xylooligosaccharides. Biotechnol Lett 33:2279–2285
La Grange DC, Claeyssens M, Pretorius IS, Van Zyl WH (2000) Coexpression of the Bacillus pumilus beta-xylosidase (xynB) gene with the Trichoderma reesei beta xylanase 2 (xyn2) gene in the yeast Saccharomyces cerevisiae. Appl Microbiol Biotechnol 54(2):195–200
Lagaert S, Van Campenhout S, Pollet A, Bourgois TM, Delcour JA, Courtin CM, Volckaert G (2007) Recombinant expression and characterization of a reducing-end xylose-releasing exo-oligoxylanase from Bifidobacterium adolescentis. Appl Environ Microbiol 73:5374–5377
Lama L, Calandrelli V, Gambacorta A, Nicolaus B (2004) Purification and characterization of thermostable xylanase and beta-xylosidase by the thermophilic bacterium Bacillus thermantarcticus. Res Microbiol 155(4):283–289
Lamed R, Setter E, Bayer EA (1983) Characterization of a cellulose-binding, cellulase-containing complex in Clostridium thermocellum. J Bacteriol 156(2):828–836
Lappalainen A, Siika-aho M, Kalkkinen N, Fagerstrom R, Tenkanen M (2000) Endoxylanase II from Trichoderma reesei has several isoforms with different isoelectric points. Biotechnol Appl Biochem 31:61e68
Larson SB, Day J, Barba de la Rosa AP, Keen NT, McPherson A (2003) First crystallographic structure of a xylanase from glycoside hydrolase family 5: implications for catalysis. Biochemistry 42(28):8411–8422
Lee J (1997) Biological conversion of lignocellulosic biomass to ethanol. J Biotechnol 56(1):1–24
Lee JM, Hu Y, Zhu H, Cheng KJ, Krell PJ, Forsberg CW (1993) Cloning of a xylanase gene from the ruminal fungus Neocallimastix patriciarum 27 and its expression in Escherichia coli. Can J Microbiol 39(1):134–139
Lee TH, Lim PO, Lee YE (2007) Cloning, characterization, and expression of xylanase A gene from Paenibacillus sp. DG-22 in Escherichia coli. J Microbiol Biotechnol 17(1):29–36
Levasseur A, Asther M, Record E (2005) Overproduction and characterization of xylanase B from Aspergillus niger. Can J Microbiol 51(2):177–183
Li XL, Ljungdahl LG (1996) Expression of Aureobasidium pullulans xynA in, and secretion of the xylanase from Saccharomyces cerevisiae. Appl Environ Microbiol 62(1):209–213
Li JH, Wang LS (2011) Why substituting the asparagine at position 35 in Bacillus circulans xylanase with an aspartic acid remarkably improves the enzymatic catalytic activity? A quantum chemistry-based calculation study. Polym Degrad Stab 96(5):1009–1014
Li C, Knierim B, Manisseri C, Arora R, Scheller HV, Auer M, Vogel KP, Simmons BA, Singh S (2010) Comparison of dilute acid and ionic liquid pretreatment of switchgrass: biomass recalcitrance, delignification and enzymatic saccharification. Bioresour Technol 101(13):4900–4906
Li XL, Skory CD, Ximenes EA, Jordan DB, Dien BS, Hughes SR, Cotta MA (2007) Expression of an AT-rich xylanase gene from the anaerobic fungus Orpinomyces sp. strain PC-2 in and secretion of the heterologous enzyme by Hypocrea jecorina. Appl Microbiol Biotechnol 74:1264–1275
Liu JR, Duan CH, Zhao X, Tzen JT, Cheng KJ, Pai CK (2008) Cloning of a rumen fungal xylanase gene and purification of the recombinant enzyme via artificial oil bodies. Appl Microbiol Biotechnol 79(2):225–233
Liu MQ, Weng XY, Sun JY (2006) Expression of recombinant Aspergillus niger xylanase A in Pichia pastoris and its action on xylan. Protein Expr Purif 48(2):292–299
Lo YC, Lu WC, Chen CY, Chen WM, Chang JS (2010) Characterization and high-level production of xylanase from an indigenous cellulolytic bacterium Acinetobacter junii F6-02 from southern Taiwan soil. Biochem Eng J 53(1):77–84
Lundgren KR, Bergkvist L, Hogman S, Joves H, Eriksson G, Bartfai T, Vanderlaan J, Rosenberg E, Shoham Y (1994) TCF mill trial on softwood pulp with korsnas thermostable and alkaline stable xylanase T6. Fems Microbiology Reviews 13(2–3):365–368
Luthi E, Love DR, McAnulty J, Wallace C, Caughey PA, Saul D, Bergquist PL (1990) Cloning, sequence analysis, and expression of genes encoding xylan-degrading enzymes from the thermophile Caldocellum saccharolyticum. Appl Environ Microbiol 56(4):1017–1024
Lynd LR, Cushman JH, Nichols RJ, Wyman CE (1991) Fuel ethanol from cellulosic biomass. Science 251(4999):1318–1323
Mach-Aigner A, Pucher ME, Mach RL (2010) d-Xylose as a repressor or inducer of xylanase expression in Hypocrea jecorina (Trichoderma reesei). Appl Environ Microbiol 76:1770–1776
Mamo G, Hatti-Kaul R, Mattiasson B (2006) A thermostable alkaline active endo-b-1-4-xylanase from Bacillus halodurans S7: Purification and characterization. Enzyme Microb Technol 39:1492–1498
Mamo G, Hatti-Kaul R, Mattiasson B (2007) Fusion of carbohydrate binding modules from Thermotoga neapolitana with a family 10 xylanase from Bacillus halodurans S7. Extremophiles 11:169–177
Mannarelli BM, Evans S, Lee D (1990) Cloning, sequencing, and expression of a xylanase gene from the anaerobic ruminal bacterium Butyrivibrio fibrisolvens. J Bacteriol 172(8):4247–4254
Margeot A, Hahn-Hagerdal B, Edlund M, Slade R, Monot F (2009) New improvements for lignocellulosic ethanol. Curr Opin Biotechnol 20:372–380
Marrone L, McAllister KA, Clarke AJ (2000) Characterization of function and activity of domains A, B and C of xylanase C from Fibrobacter succinogenes S85. Protein Eng 13(8):593–601
McCarthy AA, Morris DD, Bergquist PL, Baker EN (2000) Structure of XynB, a highly thermostable beta 1,4-xylanase from Dictyoglomus thermophilum Rt46B.1, at 1.8 Å resolution. Acta Crystallogr D56:1367–1375
Mirande C, Mosoni P, Bera-Maillet C, Bernalier-Donadille A, Forano E (2010) Characterization of Xyn10A, a highly active xylanase from the human gut bacterium Bacteroides xylanisolvens XB1A. Appl Microbiol Biotechnol 87(6):2097–2105
Moreau A, Durand S, Morosoli R (1992) Secretion of a Cryptococcus albidus xylanase in Saccharomyces cerevisiae. Gene 116(1):109–113
Morris DD, Gibbs MD, Bergquist PL (1996) Cloning of a family G xylanase gene (XYNB) from the extremely thermophilic bacterium Dictyoglomus thermophilum and action of the gene product on kraft pulp. Am Chem Soc 211:234
Muilu J, Torronen A, Perakyla M, Rouvinen J (1998) Functional conformational changes of endo-1,4-xylanase II from Trichoderma reesei: a molecular dynamics study. Proteins 31:434–444
Mullai P, Fathima NSA, Rene ER (2010) Statistical analysis of main and interaction effects to optimize xylanase production under submerged cultivation conditions. J Agric Sci 2(1):144–153
Nakamura S, Wakabayashi K, Nakai R, Aono R, Horikoshi K (1993) Production of alkaline xylanase by a newly isolated alkaliphilic Bacillus sp strain 41 m-1. World J Microbiol Biotechnol 9(2):221–224
Narang S, Satyanarayana T (2001) Thermostable alpha-amylase production by an extreme thermophile Bacillus thermooleovorans. Lett Appl Microbiol 32:31–35
Nascimento RP, Coelho RRR, Marques S, Alves L, Girio FM, Bon EPS, Amaral-Collaco MT (2002) Production and partial characterisation of xylanase from Streptomyces sp strain AMT-3 isolated from Brazilian cerrado soil. Enzyme Microb Technol 31(4):549–555
Natesh R, Bhanumoorthy P, Vithayathil PJ, Sekar K, Ramakumar S, Viswamitra MA (1999) Crystal structure at 1.8 angstrom resolution and proposed amino acid sequence of a thermostable xylanase from Thermoascus aurantiacus. J Mol Biol 288(5):999–1012
Ogasawara W, Shida Y, Furukawa T, Shimada R, Nakagawa S, Kawamura M, Yagyu T, Kosuge A, Xu J, Nogawa M, Okada H, Morikawa Y (2006) Cloning, functional expression and promoter analysis of xylanase III gene from Trichoderma reesei. Appl Microbiol Biotechnol 72(5):995–1003
Ohta K, Moriyama S, Tanaka H, Shige T, Akimoto H (2001) Purification and characterization of an acidophilic xylanase from Aureobasidium pullulans var. melanigenum and sequence analysis of the encoding gene. J Biosci Bioeng 92(3):262–270
Okeke BC, Obi SKC (1994) Lignocellulose and sugar compositions of some agro-waste materials. Bioresour Technol 47(3):283–284
Olfa E, Mondher M, Issam S, Ferid L, Nejib NM (2007) Induction, properties and application of xylanase activity from Sclerotinia sclerotiorum S2 fungus. J Food Chem 31:96–107
Paës G, Berrin J-G, Beaugrand J (2012) GH11 xylanases: structure/function/properties relationships and applications. Biotechnol Adv 30:564–592
Paës G, O’Donohue MJJ (2006) Engineering increased thermostability in the thermostable GH-11 xylanase from Thermobacillus xylanilyticus. J Biotechnol 125:338–350
Payan F, Leone P, Porciero S, Furniss C, Tahir T, Williamson G, Durand A, Manzanares P, Gilbert HJ, Juge N, Roussel A (2004) The dual nature of the wheat xylanase protein inhibitor XIP-I: structural basis for the inhibition of family 10 and family 11 xylanases. J Biol Chem 279(34):36029–36037
Payan F, Flatman R, Porciero S, Williamson G, Juge N, Roussel A (2003) Structural analysis of xylanase inhibitor protein I (XIP-I), a proteinaceous xylanase inhibitor from wheat (Triticum aestivum var soisson). Biochem J 372(Pt 2):399–405
Petrescu I, Lamotte-Brasseur J, Chessa JP, Ntarima P, Claeyssens M, Devreese B, Marino G, Gerday C (2000) Xylanase from the psychrophilic yeast Cryptococcus adeliae. Extremophiles 4(3):137–144
Pokhrel S, Joo JC, Yoo YJ (2013) Shifting the optimum pH of Bacillus circuulans xylanase towards acidic side by introducing arginine. Biotechnol Bioprocess Eng 18:35–42
Polizeli MLTM, Rizzatti ACS, Monti R, Terenzi HF, Jorge JA, Amorim DS (2005) Xylanases from fungi: properties and industrial applications. Appl Microbiol Biotechnol 67:577–591
Pollet A, Schoepe J, Dornez E, Strelkov SV, Delcour JA, Courtin CM (2010) Functional analysis of glycosidehydrolase family 8 xylanases shows narrow but distinct substrate specificities and biotechnological potential. Appl Microbiol Biotechnol 87:2125–2135
Prade RA (1996) Xylanases: from biology to biotechnology. Biotechnol Genet Eng Rev 13(1):101–132
Puchkaev AV, Koo LS, Ortiz de Montellano PR (2003) Aromatic stacking as a determinant of the thermal stability of CYP119 from Sulfolobus solfataricus. Arch Biochem Biophys 409:52–58
Puls J, Tenkanen M, Korte HE, Poutanen K (1991) Products of hydrolysis of beechwood acetyl-4-O-methylglucuronoxylan by a xylanase and an acetyl xylan esterase. Enzyme Microb Technol 13(6):483–486
Qureshy AF, Khan LA, Khanna S (2002) Cloning, regulation and purification of cellulase-free xylanase from Bacillus circulans Teri-42. Ind J Microbiol 42:35–41
Raedschelders G, Debefve C, Goesaert H, Delcour JA, Volckaert G, Van Campenhout S (2004) Molecular identification and chromosomal localization of genes encoding Triticum aestivum xylanase inhibitor I-like proteins in cereals. Theor Appl Genet 109(1):112–121
Ratto M, Poutanen K, Viikari L (1992) Production of xylanolytic enzymes by an alkalitolernt Bacillus circulans stain. Appl Microbiol Biotechnol 37:470–473
Rizzatti ACS, Jorge JA, Terenzi HF, Rechia CGV, Polizeli MLTM (2001) Purification and properties of a thermostable extracellular β-xylosidase produced by a thermotolerant Aspergillus phoenicis. J Ind Microbiol Biotechnol 26:156–160
Rose SH, van Zyl WH (2002) Constitutive expression of the Trichoderma reesei beta-1,4-xylanase gene (xyn2) and the beta-1,4-endoglucanase gene (egl) in Aspergillus niger in molasses and defined glucose media. Appl Microbiol Biotechnol 58(4):461–468
Rouau X (1993) Investigations into the effects of an enzyme preparation for baking on what flour dough pentosans. J Cereal Sic 18:145–157
Rouau X, Daviet S, Tahir T, Cherel B, Saulnier L (2006) Effect of the proteinaceous wheat xylanase inhibitor XIP-I on the performances of an Aspergillus niger xylanase in bread making. J Sci Food Agric 86(11):1604–1609
Ruanglek V, Sriprang R, Ratanaphan N, Tirawongsaroj P, Chantasigh D, Tanapongpipat S, Pootanakit K, Eurwilaichitr L (2007) Cloning, expression, characterization, and high cell-density production of recombinant endo-1,4-beta-xylanase from Aspergillus niger in Pichia pastoris. Enzyme Microb Technol 41(1–2):19–25
Saha BC (2003) Hemicellulose bioconversion. J Ind Microbiol Biotechnol 30(5):279–291
Sakka K, Maeda Y, Hakamada Y, Takahashi N, Shimada K (1991) Purification and some properties of xylanase from Clostridium stercorarium strain Hx-1. Agric Biol Chem 55(1):247–248
Salles BC, Cunha RB, Fontes W, Sousa MV, Filho EXF (2000) Purification and characterization of a new xylanase from Acrophialophora nainiana. J Biotechnol 81(2–3):199–204
Salles BC, Te’o VS, Gibbs MD, Bergquist PL, Filho EX, Ximenes EA, Nevalainen KM (2007) Identification of two novel xylanase-encoding genes (xyn5 and xyn6) from Acrophialophora nainiana and heterologous expression of xyn6 in Trichoderma reesei. Biotechnol Lett 29(8):1195–1201
Sanghi A, Garg N, Kuhar K, Kuhad RC, Gupta VK (2009) Enhanced production of cellulase-free xylanase by alkalophilic Bacillus subtilis ASH and its application in biobleaching of kraft pulp. BioResources 4:1109–1129
Sandhu JS, Kennedy JF (1984) Molecular cloning of Bacillus polymyxa (1–4)-P-D-xylanase gene in Escherichia coli. Enzyme Microb Technol 6:271–274
Sapre MP, Jha H, Patil MB (2005) Purification and characterization of a thermoalkalophilic xylanase from Bacillus sp. World J Microbiol Biotechnol 21(5):649–654
Schlacher A, Holzmann K, Hayn M, Steiner W, Schwab H (1996) Cloning and characterization of the gene for the thermostable xylanase XynA from Thermomyces lanuginosus. J Biotechnol 49(1–3):211–218
Schulze E (1891) Amphibia Europaea, annual report and treatises society of natural scientists in Magdeburg 1890:163–178
Shallom D, Shoham Y (2003) Microbial hemicellulases. Curr Opin Microbiol 6(3):219–228
Shao W, Wiegel J (1995) Purification and characterization of two thermostable acetyl xylan esterases from Thermoanaerobacterium sp. strain JW/SL-YS485. Appl Environ Microbiol 61(2):729–733
Sharma A, Adahikari S, Styanarayana T (2007) Alkali-thermostable and cellulase-free xylanase production by an extreme thermophile Geobacillus thermoleovorans. World J Microbiol Biotechnol 23:483–490
Shatalov AA, Pereira AA (2007) Xylanase pre-treatment of giant reed organosolv pulps: direct bleaching effect and bleach boosting. Ind Crops Prod 25:248–256
Shibuya H, Kaneko S, Hayanshi K (2000) Enhancement of thermostability and hydrolytic activity of xylananse by random gene shuffling. Biochem J 349:651–656
Shrinivas D, Savitha G, Raviranjan K, Naik GR (2010) A highly thermostable alkaline cellulase-free xylanase from thermoalkalophilic Bacillus sp. JB 99 suitable for paper and pulp industry: purification and characterization. Appl Biochem Biotechnol 162(7):2049–2057
Sidhu G, Withers SG, Nguyen NT, McIntosh LP, Ziser L, Brayer GD (1999) Sugar ring distortion in the glycosyl-enzyme intermediate of a family G/11 xylanase. Biochemistry 38(17):5346–5354
Silva CHC, Puls J, de Sousa MV, Ferreira EX (1999) Purification and characterization of a low molecular weight xylanase from solid-state cultures of Aspergillus fumigatus Fresenius. Rev Microbiol 30(2):114–119
Simpson HD, Haufler UR, Daniel RM (1991) An extremely thermostable xylanase from the thermophilic bacterium Thermotoga. Biochem J 277(2):413–417
Simpson PJ, Bolam DN, Cooper A, Ciruela A, Hazlewood GP, Gilbert HJ, Williamson MP (1999) A family IIb xylan-binding domain has a similar secondary structure to a homologous family IIa cellulose-binding domain but different ligand specificity. Structure 7(1999):853–864
Singh S, Pillay B, Dilsook V, Prior BA (2000) Production and properties of hemicellulases by a Thermomyces lanuginosus strain. J Appl Microbiol 88:975–982
Sorensen JF, Sibbesen O (2006) Mapping of residues involved in the interaction between the Bacillus subtilis xylanase A and proteinaceous wheat xylanase inhibitors. Protein Eng Des Sel 19(5):205–210
Sriprang R, Asano K, Gobsuk J, Tanapongpipat S, Champreda V, Eurwilaichitr L (2006) Improvement of thermostability of fungal xylanase by using site-directed mutagenesis. J Biotechnol 126(4):454–462
Srivastava R, Srivastava AK (1993) Characterization of a bacterial xylanase resistant to repression by glucose and xylose. Biotechnol Lett 15(8):847–852
Sriyapai T, Somyoonsap P, Matsui K, Kawai F, Chansiri K (2011) Cloning of a thermostable xylanase from Actinomadura sp. S14 and its expression in Escherichia coli and Pichia pastoris. J Biosci Bioeng 111(5):528–536
St John FJ, Godwin DK, Preston JF, Pozharski E, Hurlbert JC (2009) Crystallization and crystallographic analysis of Bacillus subtilis xylanase C. Acta Crystallogr Sect F Struct Biol Crystallogr Commun 65(Pt 5):499–503
Strauss J, Mach RL, Zeilinger S, Hartler G, Stoffler G, Wolschek M, Kubicek CP (1995) Cre1, the carbon catabolite repressor protein from Trichoderma reesei. FEBS Lett 376:103–107
Stephens DE, Singh S, Permaul K (2009) Error-prone PCR of a fungal xylanase for improvement of its alkaline and thermal stability. FEMS Microbiol Lett 293(1):42–47
Subramaniyan S (2000) Studies on the production of bacterial xylanases. Ph.D. Thesis, Cochin University of Science and Technology, India
Subramaniyan S, Prema P (2002) Biotechnology of microbial xylanases: enzymology, molecular biology and application. Crit Rev Biotechnol 22:33–46
Sun JY, Liu MQ, Weng XY, Qian LC, Gu SH (2007) Expression of recombinant Thermomonospora fusca xylanase A in Pichia pastoris and xylooligosaccharides released from xylans by it. Food Chem 104(3):1055–1064
Sunna A, Prowe SG, Stoffregen T, Antranikian G (1997) Characterization of the xylanases from the new isolated thermophilic xylan-degrading Bacillus thermoleovorans strain K-3d and Bacillus flavothermus strain LB3A. FEMS Microbiol Lett 148(2):209–216
Suzuki T, Ibata K, Hatsu M, Takamizawa K, Kawai K (1997) Cloning and expression of a 58-kDa xylanase VI gene (xynD) of Aeromonas caviae ME-1 in Escherichia coli which is not categorized as a family F or family G xylanase. J Ferm Bioeng 84(1):86–89
Takahashi-Ando N, Inaba M, Ohsato S, Igawa T, Usami R, Kimura M (2007) Identification of multiple highly similar XIP-type xylanase inhibitor genes in hexaploid wheat. Biochem Biophys Res Commun 360(4):880–884
Tanaka H, Okuno T, Moriyama S, Muguruma M, Ohta K (2004) Acidophilic xylanase from Aureobasidium pullulans: efficient expression and secretion in Pichia pastoris and mutational analysis. J Biosci Bioeng 98(5):338–343
Taneja K, Gupta S, Kuhad RC (2002) Properties and application of a partially purified alkaline xylanase from an alkalophilic fungus Aspergillus nidulans KK-99. Bioresour Technol 85(1):39–42
Teng C, Jia H, Yan Q, Zhou P, Jiang Z (2011) High-level expression of extracellular secretion of a β-Xylosidase gene from Paecilomyces thermophila in Escherichia coli. Bioresour Technol 102:1822–1830
Tomme P, Gilkes NR, Miller RC, Warren AJ, Kilburn DG (1994) An internal cellulose-binding domain mediates adsorption of an engineered bifunctional xylanase/cellulase. Protein Eng 7(1):117–123
Torronen A, Harkki A, Rouvinen J (1994) 3-Dimensional structure of endo-1,4-beta-xylanase-II from Trichoderma reesei-2 conformational states in the active-site. EMBO J 13(11):2493–2501
Torronen A and Rouvinen J (1997) Structural and functional properties of low molecular weight endo-1,4-beta-xylanases. J Biotechnol 57:137–149
Trudel J, Grenier J, Potvin C, Asselin A (1998) Several thaumatin-like proteins bind to beta-1,3-glucans. Plant Physiology 118:1431–1438
Tsai CT, Huang CT (2008) Overexpression of the Neocallimastix frontalis xylanase gene in the methylotrophic yeasts Pichia pastoris and Pichia methanolica. Enzyme Microb Technol 42(6):459–465
Turunen O, Etuaho K, Fenel F, Vehmaanpera J, Wu XY, Rouvinen J, Leisola M (2001) A combination of weakly stabilizing mutations with a disulfide bridge in the alpha-helix region of Trichoderma reesei endo-1,4-beta-xylanase II increases the thermal stability through synergism. J Biotechnol 88(1):37–46
Valenzuela SV, Diaz P, Javier Pastor FI (2010) Recombinant expression of an alkali stable GH10 xylanase from Paenibacillus barcinonensis. J Agric Food Chem 58(8):4814–4818
Van Campenhout S, Pollet A, Bourgois TM, Rombouts S, Beaugrand J, Gebruers K, De Backer E, Courtin CM, Delcour JA, Volckaert G (2007) Unprocessed barley aleurone endo-beta-1,4-xylanase X-I is an active enzyme. Biochem Biophys Res Commun 356(3):799–804
Van Der Borght A, Goesaert H, Veraverbeke WS, Delcour JA (2005) Fractionation of wheat and wheat flour into starch and gluten: overview of the main processes and the factors involved. J Cereal Sci 41:221–237
Van Peij NN, Brinkmann J, Vrsanska M, Visser J, de Graaff LH (1997) Beta-xylosidase activity, encoded by xlnD, is essential for complete hydrolysis of xylan by Aspergillus niger but not for induction of the xylanolytic enzyme spectrum. Eur J Biochem 245(1):164–173
Van Petegem F, Collins T, Meuwis MA, Gerday C, Feller G, Van Beeumen J (2003) The structure of a cold-adapted family 8 xylanase at 1.3 Å resolution. Structural adaptations to cold and investigation of the active site. J Biol Chem 278(9):7531–7539
Vandeplas S, Dauphin RD, Thonart P, Thewis A, Beckers Y (2010) Effect of the bacterial or fungal origin of exogenous xylanases supplemented to a wheat-based diet on performance of broiler chickens and nutrient digestibility of the diet. Can J Anim Sci 90:221–228
Vardakou M, Flint J, Christakopoulos P, Lewis RJ, Gilbert HJ, Murray JW (2005) A family 10 Thermoascus aurantiacus xylanase utilizes arabinose decorations of xylan as significant substrate specificity determinants. J Mol Biol 352:1060–1067
Verjans P, Dornez E, Delcour JA, Courtin CM (2010) Selectivity for water-unextractable arabinoxylan and inhibition sensitivity govern the strong bread improving potential of an acidophilic GH11 Aureobasidium pullulans xylanase. Food Chem 123:331–337
Verma D, Satyanarayana T (2012) Cloning, expression and applicability of thermo-alkali-stable xylanase of Geobacillus thermoleovorans in generating xylooligosaccharides from agro-residues. Bioresour Technol 107:333–338
Wang JS, Bai YG, Yang PL, Shi PJ, Luo HY, Meng K, Huang HQ, Yin J, Yao B (2010) A new xylanase from thermoalkaline Anoxybacillus sp E2 with high activity and stability over a broad pH range. World J Microbiol Biotechnol 26(5):917–924
Wang Q, Xia T (2008) Enhancement of the activity and alkaline pH stability of Thermobifida fusca xylanase A by directed evolution. Biotechnol Lett 30(5):937–944
Whitehead TR, Hespell RB (1989) Cloning and expression in Escherichia coli of a xylanase gene from Bacteroides ruminicola 23. Appl Environ Microbiol 55(4):893–896
Winterhalter C, Liebl W (1995) Two extremely thermostable xylanases of the hyperthermophilic bacterium Thermotoga maritima MSB8. Appl Environ Microbiol 61:1810–1815
Wong KK, Tan LU, Saddler JN (1988) Multiplicity of beta-1,4-xylanase in microorganisms: functions and applications. Microbiol Rev 52(3):305–317
Wu SJ, Liu B, Zhang XB (2006) Characterization of a recombinant thermostable xylanase from deep-sea thermophilic Geobacillus sp MT-1 in East Pacific. Appl Microbiol Biotechnol 72(6):1210–1216
Xiong H, Turunen O, Pastinen O, Leisola M, von Weymarn N (2004) Improved xylanase production by Trichoderma reesei grown on L-arabinose and lactose or d-glucose mixtures. Appl Microbiol Biotechnol 64:353–358
Xiong H, Weymarn N, Turunen O, Leisola M, Pastinen O (2005) Xylanase production by Trichoderma reesei Rut C-30 grown on L-arabinose-rich plant hydrolysates. Bioresour Technol 96:753–759
Xue GP, Gobius KS, Orpin CG (1992) A novel polysaccharide hydrolase cDNA (celD) from Neocallimastix patriciarum encoding three multi-functional catalytic domains with high endoglucanase, cellobiohydrolase and xylanase activities. J Gen Microbiol 138(11):2397–2403
Yang HJ, Xie CY (2010) Assessment of fibrolytic activities of 18 commercial enzyme products and their abilities to degrade the cell wall fraction of corn stalks in in vitro enzymatic and ruminal batch cultures. Anim Feed Scie Technol 159:110–121
Yang JH, Park JY, Kim SH, Yoo YJ (2008) Shifting pH optimum of Bacillus circulans xylanase based on molecular modeling. J Biotechnol 133(3):294–300
Yang RC, MacKenzie CR, Narang SA (1988) Nucleotide sequence of a Bacillus circulans xylanase gene. Nucleic Acids Res 16(14B):7187
Yamura I, Koga T, Matsumoto T, Kato T (1997) Purification and some properties of endo-1,4-[5-d-xylanase from a fresh water mollusc, Pomacea insularus (de Ordigny). Biosci Biotechol Biochem 61(4):615–620
Yin LJ, Lin HH, Chiang YI, Jiang ST (2010) Bioproperties and purification of xylanase from Bacillus sp. YJ6. J Agric Food Chem 58(1):557–562
You C, Yuan H, Huang Q, Lu H (2010) Substrate molecule enhances the thermostability of a mutant of a family 11 xylanase from Neocallimastix patriciarum. African J Biotechnol 9:1288–1294
Yuan Q, Wang J, Zhang H, Qian Z (2005) Effect of temperature shift on production of xylanase by Aspergillus niger. Process Biochem 40:3255–3257
Zhao Y, Chany CJ, Sims PF, Sinnott ML (1997) Definition of the substrate specificity of the ‘sensing’ xylanase of Streptomyces cyaneus using xylooligosaccharide and cellooligosaccharide glycosides of 3,4-dinitrophenol. J Biotechnol 57(1–3):181–190
Zhou CY, Bai JY, Deng SS, Wang J, Zhu J, Wu MC, Wang W (2008a) Cloning of a xylanase gene from Aspergillus usamii and its expression in Escherichia coli. Bioresour Technol 99:831–838
Zhou CY, Li DF, Wu MC, Wang W (2008b) Optimized expression of an acid xylanase from Aspergillus usamii in Pichia pastoris and its biochemical characterization. World J Microbiol Biotechnol 24(8):1393–1401
Zhu H, Paradis FW, Krell PJ, Phillips JP, Forsberg CW (1994) Enzymatic specificities and modes of action of the two catalytic domains of the XynC xylanase from Fibrobacter succinogenes S85. J Bacteriol 176(13):3885–3894
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
El Enshasy, H.A., Kandiyil, S.K., Malek, R., Othman, N.Z. (2016). Microbial Xylanases: Sources, Types, and Their Applications. In: Gupta, V. (eds) Microbial Enzymes in Bioconversions of Biomass. Biofuel and Biorefinery Technologies, vol 3. Springer, Cham. https://doi.org/10.1007/978-3-319-43679-1_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-43679-1_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-43677-7
Online ISBN: 978-3-319-43679-1
eBook Packages: EnergyEnergy (R0)