Abstract
Deep neural network methods have led to impressive breakthroughs in the medical image field. Most of them focus on single-modal data, while diagnoses in clinical practice are usually determined based on multi-modal data, especially for tumor diseases. In this paper, we intend to find a way to effectively fuse radiology images and pathology images for the diagnosis of gliomas. To this end, we propose a collaborative attention network (CA-Net), which consists of three attention-based feature fusion modules, multi-instance attention, cross attention, and attention fusion. We first take an individual network for each modality to extract the original features. Multi-instance attention combines different informative patches in the pathology image to form a holistic pathology feature. Cross attention interacts between the two modalities and enhances single modality features by exploring complementary information from the other modality. The cross attention matrixes imply the feature reliability, so they are further utilized to obtain a coefficient for each modality to linearly fuse the enhanced features as the final representation in the attention fusion module. The three attention modules are collaborative to discover a comprehensive representation. Our result on the CPM-RadPath outperforms other fusion methods by a large margin, which demonstrates the effectiveness of the proposed method.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
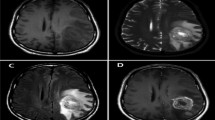
Gliomas are the most common primary intracranial tumors, accounting for 40% to 50% of all cranial tumors. World Health Organization (WHO) grading system grade the gliomas from 1 (least malignant and best prognosis) to 4 (most malignant and worst prognosis). According to the pathological malignancy of the tumor cells, brain gliomas are also divided into low-grade gliomas (including astrocytoma, oligodendroglioma) and high-grade gliomas (glioblastoma). Magnetic resonance imaging (MRI) is the common examination method for gliomas, which is mainly used to identify low-grade gliomas and high-grade gliomas. Due to the limitation of MRI in the identification of astrocytoma and oligodendroglioma, pathology images are also used. Hence, the diagnosis of gliomas in clinical practice is based on multiple modalities of medical images, which requires the doctors to have a rich experience. Computer aided diagnosis (CAD) systems are in demand to facilitate the diagnosis process.
Convolutional neural network (CNN) is the most widely used deep learning model to learn complex discriminative features of images and various architectures of CNN have been proposed, such as VGG16 [1], ResNet [2], and Densenet [3]. These networks achieve human-level performance on many tasks in the natural image field. Moreover, deep learning methods also bring significant progress in the medical field. For instance, the U-Net [4] architecture was proposed for the segmentation of neuronal structures and performed well on a variety of biomedical segmentation tasks. However, most models only focus on single modality data, such as X-ray images [5], CT images [6], or MRI images [7].
In order to obtain more information for better decision, learning methods on multi-modal data has been a growing trend. Incorporating visual information on many speech tasks has achieved great gains, such as speech enhancement [8], speech separation [9, 10]. Pretraining on vision and language data quickly become a popular task after the advent of BERT [11]. In the medical image field, multi-modal data refers to the images taken by different inspection methods and non-image data [36]. Although there are some public multi-modal datasets like BraTs [12, 37,38,39], CHAOS [13], CPM-RadPath [14, 40], the methods of fusing the multi-modal data are still deficient. To the best of our knowledge, most fusion methods on medical images are limited to direct fusion by concatenating or linear weighting at the input-level [15,16,17], feature-level [18,19,20, 28], or decision-level [21,22,23]. Pandya et al. [24] introduced a multi-channel MRI embedding strategy to improve the result of deep learning-based tumor segmentation models. This method linearly fused four modalities at the input-level. Neubauer et al. [18] improved the performance of tumor delineation by merging the features of MRI and PET/CT data after two modality-specific encoders. Kamnitsas et al. [22] trained three networks separately and averaged the confidence of each network as the final result.
MRI images and pathology images are the most common inspection methods for gliomas diagnoses. CPM-Radpath [14, 40] provided both modalities to evaluate the performance of computer-aided systems. This task is difficult as the two modalities are totally different. MRI images are 3D scanning data of the brain, while pathology images are 2D microscopy data of the sliced tissue. Ma et al. [25] fused the final results of the two modalities by logistic regression. Xue et al. [26] proposed a dual path model and fused the features before the last fully connected layer directly. However, due to the great difference between the two modalities, the relation between them is quite complicated and it can not be captured by these simple fusion methods. In this work, we adopt the powerful modeling capability of the attention mechanism and propose a collaborative attention network (CA-Net). It consists of three attention based feature fusion modules. Multi-instance attention combines different pathology patch features. Cross attention implicitly captures the relation between the two modalities and enhances both features by the complementary information from the other modality. Attention fusion fuses the two features according to the reliability of each feature, which is computed based on the learned cross attention matrixes, and obtain the final feature representation.
2 Method
Based on pathology images and MRI images, our task is to identify the subtypes of gliomas. The pipeline of the proposed CA-Net is shown in Fig. 1, including five parts, two feature extractors of pathological images and MRI images, three collaborative attention-based feature fusion modules, i.e. Multi-Instance Attention (MIA), Cross Attention (CA), Attention Fusion (AF).
2.1 Features Extraction
The resolution of pathological images is around \(100000 \times 100000\), which is too huge for computation devices to process. A typical solution is extracting patches from the whole slide image. We exclude the white background regions and crop patches sized \(256 \times 256\) without overlap. Then we filter out the patches that have low entropy. The extracted patches are then fed to a Densenet [3] structure network which consists of four stages and the number of dense blocks in each stage is 4, 8, 12, and 24.
The MRI images of each patient contain four types of scans, including T1, T2, T1-CE, and Flair. In order to reduce the useless information, extraction of the lesion is first performed by a U-Net structured lesion segmentation model with 23 layers, which is pre-trained on BraTS2019 [12, 37,38,39]. Lesion regions are then cropped and resized to \(128 \times 128 \times 128\). The four types of scans are concatenated to form a 4D tensor. The feature extractor is a 3D-Densenet [3], which consists of four stages and the number of dense blocks in each stage is 4, 8, 12, and 12.
Both the pathology image and MRI image feature extractors are trained with a cross-entropy loss. Since the pathological images are only annotated with image labels, we have no label for each patch. Thereby, we directly assign the whole image label to the sampled patches, as most studies [27] do.
2.2 Multi-instance Attention
There are multiple patches and multiple features in each pathology image, which is unbalanced when fusing with the radiology feature. So we should combine the features of all the patches to form a holistic feature, which is similar to the setting in multi-instance learning (MIL). The extracted patch is regarded as an instance and we shall build a bag feature to represent the pathology image. To this end, we propose a multi-instance attention module, as illustrated in Fig. 2.
For the convenience of parallel training, we only sample a fixed number (500 in this paper) of instances for training and inference. All the sampled instances with a feature size of \(c \times 8 \times 8\) are sent to a global average pooling (GAP) layer, result in a feature size of \(c \times 1\). c is the channel number. Then the attention coefficient is computed by Eq. 1.
\(g_j\) is the feature of the jth instance after GAP. M is the number of instances. \(w \in {R^{M \times 1}},v \in {R^{M \times c}}\) are the parameters of two fully connected layers. Tanh is employed as the activation function. The learned attention coefficients are further utilized to accumulate all the instances’ features and get the bag-level feature.
2.3 Cross Attention
Pathology features and radiology features have plenty of complementary information. Previous feature fusion methods including concatenation and linear fusion can not effectively explore the relation between the two modalities. In this work, we propose a cross-attention module to deeply learn their relations, which is illustrated in Fig. 3.
Attention is a popular mechanism in deep learning models, especially after the introduction of self-attention [29]. The most frequently used attention is scaled dot-product attention, which computes the relation by the dot product of the feature vector. The dot-product attention implies that similar features have a close relation. However, in our task, the features come from two totally different modalities, therefore, it’s not a valid way to adopt the dot-product attention. We adopt additive attention [30] to explore the relationship between different modalities, which is formulated as follows:
The pathology feature size is \(c \times 8 \times 8\) and the radiology feature size is \(c \times 4 \times 4 \times 4\). Both of them are reshaped to \(c \times 64\) before sent to the attention module. c is the channel number, i.e. feature length. Attention is computed at every position. \(q_i\) is the query feature from one modality and \(k_j\) is the key feature from the other modality. N is the number of positions (64 in our setting). A shared multi-layer perceptron (MLP) followed by a softmax normalization is employed to learn their relation. Note that \(q_i\) and \(k_j\) are concatenated before sent to the MLP, which means \(e_{ij}\) will be different when the modality of the query feature changes. Then the complementary feature from the other modality can be obtained by a simple linearly weighted summation. The complementary feature \(g_i\) is added to the original query feature \(q_i\) to enhance the feature of each modality, obtaining \(F_p\) and \(F_r\).
2.4 Attention Fusion
The last step is to fuse the features from the two modalities. Although the enhanced feature of each modality has contained the information of both modalities, we believe that the representational ability, i.e. reliabilities, of them are still different. An easy solution is to learn an adaptive linear coefficient for each modality. But this will bring in extra parameters, which will lead to overfitting. We notice that the attention matrix in the cross-attention module refers to the relation between two modalities. Thereby, we attempt to explore the reliability according to the attention matrix. Actually, when \(e_{ij}\) in Eq. 2 is bigger, it means the query feature \(q_i\) is more dependent on the key feature \(k_j\), implying that the query feature is less reliable. Although the query feature is enhanced by the cross attention module, the complementary feature is scaled by a normalized coefficient \({\alpha _{ij}}\) for the sake of stable training. Hence, the enhanced feature still does not contain sufficient complementary information. Thus we can infer the feature reliability according to \(e_{ij}\). We compute the reliability as in Eq. 5.
\(\sigma \) is a measure function, which is sigmoid in this work. The final feature representation is obtained by Eq. 6.
\(F_p\) and \(F_r\) are the enhanced pathology feature and radiology feature. \(r_p\) and \(r_r\) are the corresponding reliabilities calculated by Eq. 5 when taking pathology features and radiology features as the query feature, respectively. The higher the reliability is, the higher the weight is.
The final feature representation is sent to the classifier to be classified into three subtypes of gliomas. The loss function is cross entropy. The three attention based feature modules are jointly trained, while the feature extractors of the two modalities are trained independently.
3 Results
3.1 Experiment Setup
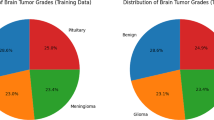
Dataset. CPM-RadPath [14, 40] consists of 221 paired radiology images and histopathology images for training. Since we can not obtain the validation data and test data, we only utilized its training data for experiments. Due to the limited number of images in medical tasks, all the experiments were evaluated by 3-folder cross-validation. The MRI images of each patient contain four types of scans, Flair, T1, T1-Ce, and T2. Due to the differences in the staining process of slices, pathology images have a big variance in color, we converted the RGB pathology images into gray images. CPM-RadPath aims to distinguish between three subtypes of brain tumors, namely astrocytoma, oligodendroglioma, and glioblastoma. The number of each subtype is shown in Table 1.
Implementation Details. Feature extractors of pathology images and radiology images were trained with a batch size of 400 and 20 respectively, and the number of feature channel was set to 64. Xavier initialization was adopted in all the models. Parameters were optimized by SGD [31], and the weight decay and momentum were set as 1e−4 and 0.95 respectively. The learning rate was initially set to 0.001 and was divided by 10 at 50% and 75% of the total training epochs. All the models were trained based on MXNet [32] for 200 epochs on a TeslaV100 GPU. For the pathology images, the same augmentation methods as the study [35] were used, including random brightness and contrast, random saturation and hue, flip, and rotation. Random crop and flip were adopted as data augmentation for the radiology images.
The feature extractors of the two modalities were first trained with a cross-entropy loss. Then we frozen the feature extractors and jointly trained the three attention modules.
3.2 Results of Gliomas Classification
The same evaluation metrics of the CPM-RadPath challenge [14, 40] were employed to evaluate the effectiveness of the proposed method in this paper.
Results on a Single Modality. The dataset consists of pathology images and radiology images (MRI). We first evaluated the performance on single modality data. Results are displayed in Table 2. Compared with the pathology image, the results of the radiology image are much worse. The reason is that astrocytoma and oligodendroglioma only have a slight difference in radiology images, so it is difficult for models to learn a discriminative feature. And that is also why we need pathology images in this task.
When evaluated on the pathology images, we compared our multi-instance attention with another common feature fusion method, max-out [33]. Max-out selects the biggest value among all the extracted patches as the output for each feature element. We do not use concatenation because the patch number is too much, i.e. 500, leading to a higher feature length, which is hard to fuse with the radiology feature. Compared with max-out, our multi-instance attention achieved higher performance, indicating that different patches have different importance and our attention mechanism can effectively incorporate all the patches.
Results on Multiple Modalities. Then we evaluated our methods on the multiple modality data. Since the training of feature extraction and feature fusion are independent, we directly used the output feature of the single modality model as the input feature of the fusion stage. Particularly, the pathology feature refers to the feature obtained by our proposed multi-instance attention. We compared our methods with other feature fusion methods and the results are displayed in Table 3. Simply concatenating the features is treated as the baseline. Xue et al. [26] fused the two features by a learned linear weight, while Ma et al. [25] fused the scores of each modality by logistic regression. We reimplemented them on the proposed framework.
As pathology features and radiology features focus on different characteristics of gliomas, simple concatenation can not capture the relation between the two modalities. So when we concatenated pathology features and radiology features, the results got even worse compared with the single pathology feature. Linear feature fusion and score fusion introduce extra parameters to capture the relation between the two modalities, thus they got an improvement and were higher than every single modality. The results show that the two modalities are complementary and can benefit from each other.
The linear fusion method is a simple linear combination of two features and there is no interaction between the two modalities. So we propose the cross attention module to interact between the two modalities and intend to enhance single modality features by digging complementary information from the other modality. The enhanced features are further fused by two linear weights which are derived from the attention matrix, i.e. attention fusion. As Table 3 shows, our results outperform other methods by a large margin. We also conducted an ablation experiment that replaced the attention fusion module with a concatenation operation. The performance is also higher than other methods, which further demonstrates that the cross-attention module can explore complementary information from each other and form a comprehensive feature representation.
4 Conclusion
In this paper, we propose a collaborative attention network to utilize multiple modality data for the diagnosis of gliomas. The network consists of three attention-based feature fusion modules. The multi-instance attention combines different patch features from the pathology images to construct a holistic pathology feature. Then the pathology feature and radiology feature are fused by the cross attention module. The final feature representation is obtained by the attention fusion module. Experimental results on CPM-RadPath demonstrate the effectiveness of the proposed method.
The proposed attention fusion module recovers the reliability of different features according to their cross-attention matrices. No additional parameters are introduced in this module and it can be implemented with one line of code. Thereby, it can be served as a plug-and-play module and used in other multi-feature fusion tasks.
References
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv: 1409.1556
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (ICCV), pp. 770–778 (2016)
Huang, G., Liu, Z., et al.: Densely connected convolutional networks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), pp. 2261–2269 (2016)
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Frid-Adar, M., Ben-Cohen, A., Amer, R., Greenspan, H.: Improving the segmentation of anatomical structures in chest radiographs using U-Net with an ImageNet pre-trained encoder. In: Stoyanov, D., et al. (eds.) RAMBO/BIA/TIA -2018. LNCS, vol. 11040, pp. 159–168. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-00946-5_17
Fan, D., Zhou, T., Ji, G., et al.: Inf-Net: automatic COVID-19 lung infection segmentation from CT images. IEEE Trans. Med. Imaging 39, 2626–2637 (2020)
Quande, L., Lequan, Y., Pheng, A.: MS-Net: multi-site network for improving prostate segmentation with heterogeneous MRI data. IEEE Trans. Med. Imaging 39(9), 2713–2724 (2020)
Gu, R., Zhang, S., Xu, Y., et al.: Multi-modal multi-channel target speech separation. IEEE J. Sel. Top. Signal Process. 14, 530–541 (2020)
Wang, W., Xing, C., Wang, D., et al.: A robust audio-visual speech enhancement model. In: IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), pp. 7529–7533 (2020)
Sadeghi, M., Alameda-Pineda, X.: Robust unsupervised audio-visual speech enhancement using a mixture of variational autoencoders. In: IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP) (2020)
Devlin, J., Lee, K., et al.: BERT: pre-training of deep bidirectional transformers for language understanding. In: Computation and Language (2018)
Kavur, A.E., Gezer, N.S., Bar, M.M., et al.: CHAOS challenge - combined (CT-MR) healthy abdominal organ segmentation. In: Image and Video Processing (2020)
CPM-RadPath. https://www.med.upenn.edu/cbica/cpm2020.html
Kamnitsas, K., Ledig, C., Newcombe, V.F.J., et al.: Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 36, 61–78 (2016)
Clèrigues, A., Valverde, S., Bernal, J., et al.: SUNet: a deep learning architecture for acute stroke lesion segmentation and outcome prediction in multimodal MRI. arXiv preprint arXiv: 1810.13304 (2018)
Shaoguo, C., Lei, M., Jingfeng, J., et al.: Automatic semantic segmentation of brain gliomas from MRI images using a deep cascaded neural network. J. Healthcare Eng. 1–14 (2018)
Neubauer, T., et al.: Soft tissue sarcoma co-segmentation in combined MRI and PET/CT data. In: Syeda-Mahmood, T., et al. (eds.) CLIP/ML-CDS -2020. LNCS, vol. 12445, pp. 97–105. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-60946-7_10
Chen, Yu., Chen, J., Wei, D., Li, Y., Zheng, Y.: OctopusNet: a deep learning segmentation network for multi-modal medical images. In: Li, Q., Leahy, R., Dong, B., Li, X. (eds.) MMMI 2019. LNCS, vol. 11977, pp. 17–25. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-37969-8_3
LaLonde, R., et al.: INN: inflated neural networks for IPMN diagnosis. In: Shen, D., et al. (eds.) MICCAI 2019. LNCS, vol. 11768, pp. 101–109. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-32254-0_12
Myronenko, A.: 3D MRI brain tumor segmentation using autoencoder regularization. In: Crimi, A., Bakas, S., Kuijf, H., Keyvan, F., Reyes, M., van Walsum, T. (eds.) BrainLes 2018. LNCS, vol. 11384, pp. 311–320. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-11726-9_28
Kamnitsas, K., et al.: Ensembles of multiple models and architectures for robust brain tumour segmentation. In: Crimi, A., Bakas, S., Kuijf, H., Menze, B., Reyes, M. (eds.) BrainLes 2017. LNCS, vol. 10670, pp. 450–462. Springer, Cham (2018). https://doi.org/10.1007/978-3-319-75238-9_38
Aygün, M., Yusuf, H., et al.: Multi modal convolutional neural networks for brain tumor segmentation. arXiv preprint arXiv: 1809.06191 (2018)
Apurva, P., Catherine, S., Nisargkumar, P.: Multi-channel MRI embedding: an effective strategy for enhancement of human brain whole tumor segmentation. arXiv preprint arXiv: 2009.06115 (2020)
Ma, X., Jia, F.: Brain tumor classification with multimodal MR and pathology images. In: Crimi, A., Bakas, S. (eds.) BrainLes 2019. LNCS, vol. 11993, pp. 343–352. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-46643-5_34
Xue, Y., et al.: Brain tumor classification with tumor segmentations and a dual path residual convolutional neural network from MRI and pathology images. In: Crimi, A., Bakas, S. (eds.) BrainLes 2019. LNCS, vol. 11993, pp. 360–367. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-46643-5_36
Patil, A., Tamboli, D., Meena, S., et al.: Breast cancer histopathology image classification and localization using multiple instance learning. In: IEEE International WIE Conference on Electrical and Computer Engineering (WIECON-ECE) (2019)
Yin, B., Cheng, H., Wang, F., Wang, Z.: Brain tumor classification based on MRI images and noise reduced pathology images. In: Crimi, A., Bakas, S. (eds.) BrainLes 2020. LNCS, vol. 12659, pp. 465–474. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-72087-2_41
Vaswani, A., Shazeer, N., Parmar, N., et al.: Attention is all you need. In: Neural Information Processing Systems, pp. 5998–6008 (2017)
Dzmitry, B., KyungHyun, C., Yoshua, B.: Neural machine translation by jointly learning to align and translate. In: 3rd International Conference on Learning Representations (ICLR) (2015)
Niu, F., Recht, B., Re, C., et al.: HOGWILD!: a lock-free approach to parallelizing stochastic gradient descent. In: Advances in Neural Information Processing Systems 24, pp. 693–701 (2011)
Chen, T., Li, M., Li, Y., et al.: MXNet: a flexible and efficient machine learning library for heterogeneous distributed systems. In: Statistics (2015)
Soumya, R., David, P.: Multiple instance regression. In: Proceedings of the Eighteenth International Conference on Machine Learning (2001)
Landis, J.R., Koch, G.G.: The measurement of observer agreement for categorical data. Biometrics 33(1), 159–174 (1977)
Liu, Y., Gadepalli, K., Norouzi, M., et al.: Detecting cancer metastases on gigapixel pathology images. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2017)
Chen, R.J., Lu, M.Y., Wang, J., et al.: Pathomic fusion: an integrated framework for fusing histopathology and genomic features for cancer diagnosis and prognosis. IEEE Trans. Med. Imaging 41, 757–770 (2020)
Bakas, S., Reyes, M., Jakab, A., et al.: Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the BRATS challenge. arXiv preprint arXiv: 1811.02629 (2018)
Bakas, S., Akbari, H., Sotiras, A., et al.: Advancing the cancer genome atlas glioma MRI collections with expert segmentation labels and radiomic features. Nat. Sci. Data 4, 170117 (2017)
Menze, B.H., Jakab, A., Bauer, S., et al.: The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans. Med. Imaging 34(10), 1993–2024 (2015)
Kurc, T., Bakas, S., Ren, X., et al.: Segmentation and classification in digital pathology for glioma research: challenges and deep learning approaches. Front. Neurosci. 14, 27 (2020)
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2020AAA0107900).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Yin, B., Cheng, H., Wang, F., Wang, Z. (2022). CA-Net: Collaborative Attention Network for Multi-modal Diagnosis of Gliomas. In: Crimi, A., Bakas, S. (eds) Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries. BrainLes 2021. Lecture Notes in Computer Science, vol 12962. Springer, Cham. https://doi.org/10.1007/978-3-031-08999-2_4
Download citation
DOI: https://doi.org/10.1007/978-3-031-08999-2_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-08998-5
Online ISBN: 978-3-031-08999-2
eBook Packages: Computer ScienceComputer Science (R0)