Abstract
Gliomas are the most common primary malignant tumors of the brain caused by glial cell canceration of the brain and spinal cord. Its incidence accounts for the vast majority of intracranial tumors and has the characteristics of high incidence, high recurrence rate, high mortality, and low cure rate. Gliomas are graded into I to IV by the World Health Organization (WHO) and the treatment is highly dependent on the grade. Diagnosis and classification of brain tumors are traditionally done by pathologists, who examine tissue sections fixed on glass slides under a light microscope. This process is time-consuming and labor-intensive and does not necessarily lead to perfectly accurate results. The computer-aided method has the potential to improve tumor classification process. In this paper, we proposed two convolutional neural networks based models to predict the grade of gliomas from both radiology and pathology data. (1) 2D ResNet-based model for pathology whole slide image classification. (2) 3D DenseNet-based model for multimodal MRI images classification. Finally, we achieve first place in CPM-RadPath-2019 [1] challenge using these methods for the tasks of classifying lower grade astrocytoma (grade II or III), oligodendroglioma (grade II or III) and glioblastoma (grade IV).
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Brain tumors are one of the common diseases in the central nervous system, which are classified into primary and secondary tumors. Primary brain tumors originate from brain cells, while secondary tumors metastasize from other organs to the brain. Glioma is the first of all types of tumors in the brain and has the highest incidence of intracranial malignant tumors. The World Health Organization (WHO) grades I-IV for gliomas based on malignant behavior for clinical purposes. Preoperative glioma grading is critical for prognosis prediction and treatment planning. Grade I has the highest survival rate and Grade IV has the lowest.
The current standard diagnosis and grading of brain tumors are done by pathologists who test Hematoxylin and Eosin (H&E) staining tissue sections fixed on glass slides under an optical microscope after resection or biopsy. The cell of these subtypes has distinct features that pathologists use for grade confirmation. Some of the gliomas have the mixed above-mentioned features so it is difficult to distinguish accurately. The whole confirmation process is usually time-consuming, invasive, prone to sampling error and user interpretation. The previous study has shown that CNN can be helpful for glioma classification using pathology images [7]. CNN has shown its ability to work well with large labeled datasets of computer vision tasks, such as ImageNet. But it is not easy for medical imaging fields because of the limitations of data acquisition. Besides, especially for WSI images, it is a difficult point because the size is too large, such as 40k * 40k pixels.
Magnetic Resonance Imaging (MRI) is the standard medical imaging technique for brain tumor diagnosis in clinical practice. Usually, several complimentary 3D MRI modalities are obtained - such as Fluid Attenuation Inversion Recover (FLAIR), T1, T1 with contrast agent (T1c), and T2. Many works have been done on trying to grade gliomas using MRI images by radiomics feature-based machine learning methods [4] and CNN based deep learning approaches [5, 6, 13, 14] since MRI is relatively safe and non-invasive. However, since the golden standard for glioma classification is based on pathological information, it is difficult to predict the subtype of the glioma using only the MRI data.
With the development of machine learning, especially deep learning and computing ability, the automatic diagnosis technology based on computer vision has been applied in many fields, and has made great success in judging the types of diseases and the segmentation of lesions. It is natural to use deep learning to combine pathological and MRI images to predict gliomas subtypes.
In this work, we present a deep learning-based method for the glioma classification of (1) lower grade astrocytoma, IDH-mutant (Grade II or III), (2) oligodendroglioma, IDH-mutant, 1p/19q codeleted (Grade II or III), and (3) glioblastoma and diffuse astrocytic glioma with molecular features of glioblastoma, IDH-wildtype (Grade IV). We try to utilize both pathology and MRI data to train the model for a better classification result. Our model won first place in the MICCAI 2019 Computational Precision Medicine: Radiology-Pathology Challenge.
2 Dataset and Method
In this section, we describe the CPM-RadPath-2019 dataset and our method for preprocessing and prediction.
2.1 Dataset
The CPM-RadPath-2019 training dataset consists of a total of 221 paired radiology scans and digitized histopathology images. Three subtypes which need to be classified are glioblastoma, oligodendroglioma, and astrocytoma with the number of 133, 54 and 34. The data was provided in two formats: *.tiff for pathology images and *.nii.gz for MRI images with modalities of FLAIR, T1, T1c, and T2. In addition, we also get age information of the patient in the format of days (Fig. 1). The size of the validation and testing set is 35 and 73 respectively.
2.2 Preprocessing
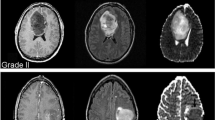
Pathology. For all patients, each one has a pathological image, respectively (Fig. 2). In each picture, only a small part of it is where the cells are stained, and a large part of it is a white background. Besides, there are different degrees of damage in some pictures, such as some strange color areas (blue or green). So we need to find an effective way to extract meaningful information. Pathology images of this dataset are compressed tiff format files which are very big after decompression (up to about 30 GB). Due to the limitations of the resources, it is not feasible to process the whole image. So we use openslide [8] to extract patches of 512 * 512 pixel with a stride of 512 in both directions from the whole slide images (see Fig. 3). During the extraction, we set several constraints to prevent sampling to data that we are not expecting like backgrounds and damaged images.
-
1)
100 < mean of the patch < 220.
-
2)
Standard deviation of the patch > 100.
-
3)
Convert the patch from RGB to HSV and the mean of channel 0 of it > 135 with the standard deviation > 25.
The iteration process will be stopped if two thousand patches are extracted or exceeded the limits of maximum time. The number of patches of each label is 108744, 49067 and 30318 with the order of G, A, and O. Although it may not be very accurate, we set the labels of all extracted patches to the labels of the entire WSI image.
Because of the different conditions of staining slice, using color normalization can make the result of the image better and extract more effective information. Now there are many robust color normalization methods for preprocessing the image [12]. In our model, for the sake of simplicity, we transform the original RGB images into grayscale images. The entire preprocessing process of pathological images can be seen in Fig. 4. All these extracted patches will be used as the input of the neural network for the tumor classification.
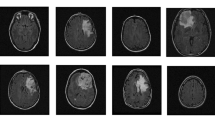
Magnetic Resonance Images. In general, for magnetic resonance images, extracting valid features requires information on the tumor mask, and then models are established by these radiomics features to predict the desired outcome. However, since the data set does not provide information about the mask, we do not intend to use additional information. Although the MR image of this dataset is very similar to the BraTS dataset [2], we have tried to train a segmentation network using data from BraTS. Four raw modalities data are used for the convolutional neural network to extract the 3D texture information. We use SimpleITK to convert the MRI images to numpy arrays and normalize them by subtracting the mean and scaling it to unit variance independently by each modality. The volume size of the input data for the convolutional neural network is 4 * 155 * 240 * 240. The sample MRI images can be seen in Figs. 5 and 6.
2.3 Convolutional Neural Network
Pathology. The extracted patches of the original WSI are grayscale *.png files. We use well-known ResNet34 and ResNet50 [9] deep learning network architectures for this classification task. The results of patch based classification on cross-validation set can be seen in Table 1. The size of the training and cross-validation set is 150504 and 37625 respectively.
ImageNet statistics are used for the normalization of the input images. The batch size is 24 due to the limitation of the GPU memory. The cross-validation set is obtained by randomly splitting the training set by 0.2. Adam optimizer is employed with the initial learning rate \(1e-4\) and weight decay \(1e-5\). The loss function is cross entropy loss. Extensive data augmentations are used including random crop, rotation, zoom, translation and color change.
Magnetic Resonance Images. 3D DenseNet is employed to explore the MRI volumes’ capabilities of the classification of gliomas because two-dimensional networks may lose some spatial contextual information and dense connections work better with small datasets [10]. The input and output layers are modified to meet our needs. Five-fold cross validation is employed to get a more robust model. The average results obtained could be found in Table 2. Besides, the confusion matrix can be seen in Fig. 7.
The batch size is 2 because of GPU memory efficiency. Mirroring, rotations, scaling, cropping, and color augmentation are used for data augmentation. The learning rate, loss function, and optimizer are set to be the same as the ResNet model described above. All models are implemented on PyTorch platform [11] and use NVIDIA Tesla V100 16GB GPU.
In the training stage, we train two models separately. In the inference stage, we make a simple regression model get the final result according to the outputs of the two models. Test time augmentation is performed at both the convolutional neural network models for a more stable prediction. In general, the results produced by the combined use of the outputs of the two models are more robust and accurate. The whole process can be seen in Fig. 8.
3 Results and Discussion
In this section, we will briefly present some of the best evaluation results and discuss the methods we have experimented with.
As you can see in Table 3, the results are obtained from the online evaluation tool. Above mentioned approaches are evaluated on the validation set. Several metrics are available from online evaluation to estimate the performance of our models.
Among the metrics, balanced accuracy is defined as the average recall obtained in each class. Kappa coefficients represent the proportion of error reduction between classifications and completely random classifications. \(p_o\) is the sum of the number of samples for each correct classification divided by the total number of samples. \(p_e\) is the expected agreement when both annotators assign labels randomly. The F1 score can be interpreted as a weighted average of the precision and recall, where an F1 score reaches its best value at 1 and the worst score at 0. [3] TP, TN, FN, FP stand for true positive, true negative, false negative, and false positive respectively.
For pathological data, we have used directly extracted pictures and binarized and denoised pictures after extraction. The first method can obtain above 95% accuracy easily by using a convolutional neural network, while the second approach is stuck on about 75% accuracy. As for the patch extraction method, I believe there could be some better constraints other than what we have mentioned above. The speed and the rate of successful sampling of pre-set value need to be further improved. Besides, the further annotation of pathological whole slide images could be a help for our method.
For MRI data, in addition to the methods mentioned above, we also try to use the segmentation tumors regions as the input of the neural network. The segmentation network is trained by BraTS2018 challenge. However, we do not use the tumor segmentation information because we do not want to use external datasets. 2D network has also experimented and the results are worse than the 3D network as we can infer.
Overall, our experiments show the power of the convolutional neural network on the task of glioma classification. Through pathological and MRI data, our model could support the diagnosis and treatment planning of glioma for the pathologists and radiologist. Effective combination of the computer-aided and manual method can improve efficiency of the remedy process.
4 Conclusion
In this paper, we explore the potential of computer-aided method’s ability to the diagnosis and grading of glioma by developing a CNN based ensemble model from pathology and radiology data. Our method achieves first place for astrocytoma, oligodendroglioma, and glioblastoma classification problem from CPM-RadPath-2019. The results suggest that our model could be useful for improving the accuracy of glioma grading. The proposed model could be further developed in the future.
References
Computational precision medicine: Radiology-pathology challenge on brain tumor classification 2019, CBICA. https://www.med.upenn.edu/cbica/cpm2019.html
Multimodal brain tumor segmentation challenge 2018. CBICA. https://www.med.upenn.edu/sbia/brats2018/evaluation.html
Buitinck, L., et al.: API design for machine learning software: experiences from the SCIKIT-learn project. In: ECML PKDD Workshop: Languages for Data Mining and Machine Learning, pp. 108–122 (2013)
Chen, Q., Wang, L., Wang, L., Deng, Z., Zhang, J., Zhu, Y.: Glioma grade predictions using scattering wavelet transform-based radiomics (2019)
Citak-Er, F., Firat, Z., Kovanlikaya, I., Ture, U., Ozturk-Isik, E.: Machine-learning in grading of gliomas based on multi-parametric magnetic resonance imaging at 3T. Comput. Biol. Med. 99, 154–160 (2018)
Decuyper, M., Van Holen, R.: Fully automatic binary glioma grading based on pre-therapy MRI using 3D convolutional neural networks (2019)
Ertosun, M.G., Rubin, D.L.: Automated grading of gliomas using deep learning in digital pathology images: a modular approach with ensemble of convolutional neural networks. In: AMIA Annual Symposium Proceedings, vol. 2015, p. 1899. American Medical Informatics Association (2015)
Goode, A., Gilbert, B., Harkes, J., Jukic, D., Satyanarayanan, M.: Openslide: a vendor-neutral software foundation for digital pathology. J. Pathol. Inform. 4, 27 (2013)
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. CoRR abs/1512.03385 (2015). http://arxiv.org/abs/1512.03385
Huang, G., Liu, Z., Weinberger, K.Q.: Densely connected convolutional networks. CoRR abs/1608.06993 (2016). http://arxiv.org/abs/1608.06993
Paszke, A., et al.: Automatic differentiation in pytorch (2017)
Tellez, D., et al.: Quantifying the effects of data augmentation and stain color normalization in convolutional neural networks for computational pathology. CoRR abs/1902.06543 (2019). http://arxiv.org/abs/1902.06543
Wang, X., et al.: Machine learning models for multiparametric glioma grading with quantitative result interpretations. Front. Neurosci. 12, 1046 (2018)
Yang, Y., et al.: Glioma grading on conventional MR images: a deep learning study with transfer learning. Front. Neurosci. 12, 804 (2018)
Acknowledgements
This work was supported in part by Shenzhen Key Basic Science Program (JCYJ20170413162213765 and JCYJ20180507182437217), the Shenzhen Key Laboratory Program (ZDSYS201707271637577), the NSFC-Shenzhen Union Program (U1613221), and the National Key Research and Development Program (2017YFC0110903).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Ma, X., Jia, F. (2020). Brain Tumor Classification with Multimodal MR and Pathology Images. In: Crimi, A., Bakas, S. (eds) Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries. BrainLes 2019. Lecture Notes in Computer Science(), vol 11993. Springer, Cham. https://doi.org/10.1007/978-3-030-46643-5_34
Download citation
DOI: https://doi.org/10.1007/978-3-030-46643-5_34
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-46642-8
Online ISBN: 978-3-030-46643-5
eBook Packages: Computer ScienceComputer Science (R0)