Abstract
Blast is one of the devastating diseases of rice which cause significant yield losses. Blast is mainly managed by the use of fungicides. However, this approach is not eco-friendly and causes health hazards. Basmati is the specialty rice of India with great export potential. However, the importing nations have increased the stringency in maximum permissible limits of pesticide residues in the grain, which has led to rejection of several Basmati rice consignments. Therefore, developing genetic resistance is one of the most pragmatic approaches to address this issue. More than 100 blast resistance genes have been identified which can be effectively deployed into the high-yielding rice varieties through marker-assisted backcross breeding. In India the major blast resistance genes, namely Pi9, Pi2, Pi54, Pita, and Pi1, are widely used to develop blast resistance in popular rice varieties. Until recently, blast-resistant varieties developed in India are Pusa Basmati 1637, Pusa Basmati 1609, Pusa 1612, and Pusa Samba 1850. Effective adoption of these varieties would significantly reduce the use of chemical pesticides, thereby producing consumer-safe rice with eco-friendly approach.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
2.1 Introduction
Rice is the important staple food crop of the world (Seck et al. 2012) which suffices 76% of the calorific needs of South Asia (Fitzgerald et al. 2008). Globally, India stands at first position with 44 mha area under rice cultivation with a production of ~116 million tons of milled rice during 2017–2018 (www.agricoop.gov.in). Basmati is a specialty rice which is mainly grown in the Indo-Gangetic plains of India which include states of Punjab, Haryana, Delhi, Uttarakhand, Himachal Pradesh, Jammu and Kathua districts of Jammu and Kashmir, and parts of western Uttar Pradesh. Basmati rice is mainly grown in an area of 1.5 mha with a production of 5.16 mt during 2018–2019. The annual forex earning due to export of Basmati rice is Rs. 32,806 crores (www.apeda.gov.in). Basmati rice possesses unique quality traits such as extra-long slender grain, exceptionally long-cooked grain, and soft and fluffy texture of cooked rice with strong and appealing aroma (Singh 2000). Major Basmati rice varieties under cultivation are Pusa Basmati 1, Pusa Basmati 1121, Pusa Basmati 6, Pusa Basmati 1509, etc. However, most of these popular rice varieties are highly susceptible to various biotic stresses, of which rice blast is one of the most devastating diseases of rice which cause significant yield losses.
2.2 Pathogen
Rice blast caused by a fungi Magnaporthe oryzae (Hebert) (Couch and Kohn 2002) [anamorph Pyricularia grisea (Cooke) Saccardo] is considered one of the most devastating diseases of rice causing significant yield losses. Annually, 10–30% of yield losses occur due to rice blast (Talbot 2003). M. oryzae is a hemi-biotroph which infects and grows in living plant cells but kills the infected cells once spread onto the adjacent cells, although it grows biotrophically in rice roots (Wilson and Talbot 2009). The infection begins when a three-celled conidium lands on a host leaf and anchors to the leaf cuticle. M. oryzae causes the disease in a cyclic developmental process. It initially attaches to the host surface through its conidia after which the spore germination takes place which extends into a germ tube. The germ tube undergoes hooking and swelling at its tip which further differentiates into a unicellular, dome-shaped infection structure called the appressorium. In 6–10 h, the appressoria differentiates, making its cell wall melanized to restrict permeability allowing only water molecules to pass through (Howard et al. 1991a). The cytoplasm then flows from the conidia into the appressorium and as the disease progresses, the latter matures and the former gets empty and collapses (Braun and Howard 1994). Glycerol accumulation engenders cellular turgor as high as 8 MPa which breaches the plant cell (Howard et al. 1991b). Once inside the cell, M. oryzae forms bulbous invasive hypha which develops a specialized structure called biotrophic interfacial complex responsible for secreting virulence factors. The disease spreads through the plasmodesmata to the neighboring cells. At this stage, the small oval chlorotic lesions begin to appear which further turn necrotic and then coalesce.
2.3 Host Range
M. oryzae is known to infect a wide variety of monocotyledonous plants such as rice (Oryza sativa), pearl millet (Pennisetum glaucum), foxtail millet (Setaria italica), wheat (Triticum aestivum), and barley (Hordeum vulgare L) but not the dicotyledonous plants (Schulze-Lefert and Panstruga 2011). The fungus can attack several other species of grasses including annual, perennial ryegrass and turfgrasses. It is also known to infect Arabidopsis via a mechanism distinct from that in rice (Park et al. 2009).
2.4 Symptoms and Damage
The blast pathogen attacks all aerial parts including leaf blades, leaf sheaths, rachis, nodes, panicles, and glumes. Blast occurs mainly during the vegetative stage of the crop cycle, although the major injury occurs during its reproductive stage called neck blast. The epidemics occur generally at maximum tillering followed by a sharp decline and a very low disease severity from flowering until the end of the crop cycle. Yield losses associated to neck blast are much higher than yield losses associated to leaf blast in the tropical rice lowlands of Asia (Ghatak et al. 2013).
2.5 Blast Prevention and Management
Rice blast hampers crop yield massively. Various cultural practices including broad-spectrum seed treatment, split application of nitrogen, and burning-contaminated straw and stubbles are employed to prevent the disease. Chemical measures are widely used by spraying prescribed doses of fungicides such as edifenphos, dithiocarbamate, benomyl, carbendazim, tricyclazole, and pyroquilon (Kumar et al. 2013). However, the cultural practices are not sufficient and usage of chemical pesticides is not an eco-friendly and bio-safe measure to counteract the pathogen. Therefore, developing genetic resistance is the most potent and durable approach to manage rice blast.
2.6 Genes Governing Blast Resistance
Rice blast patho-system follows the gene-for-gene model of host pathogen interaction (Flor 1956), according to which the key to disease resistance lies in the recognition of avirulence proteins by the plant resistance gene products (Silue et al. 1992). More than 100 blast resistance genes and more than 500 QTLs governing blast resistance have been identified (Ashkani et al. 2015). Many blast R-genes have been clustered on chromosomes 6 and 11 (Tanweer et al. 2015). Further, 36 blast resistance genes, viz., Pib, Pit, Pita, Pita2, Pi1, Pi2, Pi5, Pi9, pi21, Pi25, Pi35, Pi36, Pi37, Pi50, Pi54, Pi56, Pi63, Pi64, Pizt, Pik, Pikm, Pikp, Pikh, Pike, Piks, Pigm, Pid2, Pid3, Pid3A4, Pish, Pb1, Pia, Pii, Pi-CO39, Pi54of, and Pi54rh, have been characterized (Table 2.1; Wang et al. 2017; Meng et al. 2020). The first blast resistance gene to be cloned was Pib in Japan (Wang et al. 1999), while the first blast resistance gene to be cloned in India was Pi54 (Sharma et al. 2005).
Correspondingly, >40 Avr genes have been identified using map-based cloning strategy (Feng et al. 2007); among these ten Avr genes, namely AvrPita (Orbach et al. 2000), ACE1 (Böhnert et al. 2004), AvrPia, AvrPii, AvrPik (Yoshida et al. 2009), AvrPiz-t (Li et al. 2009), Avr1-CO39 (Zheng et al. 2011), AvrPib (Zhang et al. 2015), AvrPi9 (Wu et al. 2015), and AvrPi54 (Ray et al. 2016), have been characterized with their corresponding R-genes.
2.7 Development of Differential Varieties
A differential set consisting of 24 monogenic lines with resistance genes Pia, Pib, Pik, Pikh, Pikm, Pikp, Piks, Pish, Pit, Pita, Pita2, Piz, Piz5, Pizt, Pi1, Pii, Pi3, Pi5 (t), Pi7 (t), Pi9, Pi11 (t), Pi12 (t), Pi19 (t), and Pi20 (t) in the genetic background of a highly susceptible, japonica rice variety LTH was developed to monitor and predict the evolution of new forms of blast races (Tsunematsu et al. 2000; Fukuta et al. 2004). Additionally, another set of 27 monogenic NILs carrying various blast resistance genes in the genetic background of CO39 was developed (Telebanco-Yanoria et al. 2011).
2.8 Molecular Mechanism Underlying Blast Resistance via R-Genes
Molecular markers have become the most potent solution for overcoming blast. As the pathogen enters the host, specific avirulence protein (Avr) secreted by M. oryzae is recognized by the respective resistance (R) gene present in the resistant variety. The interaction leads to the activation of effector-triggered immunity (ETI) provoking hypersensitive reaction, thereby curbing the advancement of the pathogen cycle. The effector proteins secreted by the pathogen ignite the infection process to hijack the host machinery, which in turn are recognized by the R-gene to execute the defense mechanism pathway in the resistant varieties (Jain et al. 2017). Another defense mechanism involves the recognition of pathogen-associated molecular patterns (PAMP) by the plant pattern recognition receptors (PRR) activating pathogen-triggered immunity (PTI) (Dangl et al. 2013). However, the former mechanism imparts comparatively stronger and faster defense activation (Dong et al. 2013). The blast-resistant Pi genes encode the nucleotide-binding site-leucine-rich repeat (NBS-LRR) proteins and mostly impart ETI. The Pi genes are reported mostly to be constitutively expressed except Pi5-1 (Lee et al. 2009), pi21 (Fukuoka et al. 2009), Pi63 (Xu et al. 2014), and Pb1 (Hayashi et al. 2010).
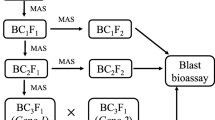
2.9 Marker-Assisted Backcross Breeding (MABB)
MABB is the promising approach to utilize molecular markers for incorporating target gene(s) into the elite crop variety without hampering its genetic background. A typical MABB comprises foreground, recombinant, and background selection. MABB requires the markers based on/linked to the target gene for foreground selection (Tanksley 1983), flanking markers for recombinant selection (Collard and Mackill 2008) and markers polymorphic between the recurrent and donor parents uniformly spanning across the genome for background selection (Hospital and Charcosset 1997). With the availability of molecular markers linked to the blast resistance genes as well as genome-wide markers, MABB has been effectively utilized to improve rice varieties for blast resistance.
2.9.1 MABB for Developing Blast Resistance in India
The success of MABB has been depicted by various research programs utilizing the strategy with modified approaches. Distinct combinations of genes/QTLs have been introgressed into the background of popular cultivated varieties to overcome the virulent pathogenic spectrum of blast prevalent in a region.
Hittalmani et al. (2000) pyramided three blast resistance genes, i.e., Pi2, Pi1, and Pita, through MAS and concluded that Pi2 exhibits higher level of resistance owing to the broader resistance spectrum of this gene. However, this study did not yield a commercial product. A combination of constitutively expressed gene Pi2 from a donor parent C101A51 and a pathogen-induced gene Pi54 from a donor parent Tetep was used to develop resistance in the genetic background of a PRR78, a restorer parent of a popular Basmati quality rice hybrid Pusa RH10, through marker-assisted simultaneous but stepwise method of backcross breeding (Singh et al. 2012a, b, 2013). The recurrent parent genome recovery in the developed NILs ranged from 78.33 to 89.01%. Further, the cooking quality and agronomic performance of these introgression lines were at par to the recurrent parent PRR78. Also, in another study the blast resistance genes Piz5 and Pi1 were incorporated into the genetic background of PRR78 by employing marker-assisted foreground selection using SSR markers AP5659-5 and RM5926, respectively. These lines were forwarded up to BC3F5 and the recovery of RPG ranged from 85.4 to 92.1% in the derived lines.
The first attempt to develop blast resistance in Basmati rice varieties was made by Khanna et al. (2015a, b). Pusa Basmati 1, a widely cultivated popular Basmati rice variety, was found highly susceptible to blast disease. Therefore, seven blast resistance genes, namely Pi1, Pi54, Pita, Pi2, Pi9, Pi5, and Pib, were incorporated into the genetic background of Pusa Basmati 1 through MABB using the gene-based/linked markers RM224, RM206, YL87/155, AP4007, AP5659-5, C1454, and RM208, respectively. A cafeteria of 36 NILs comprising 14 monogenic, 16 two-gene pyramids, and 6 three-gene pyramids was developed. Among the genes tested, Pi9 was found to be most effective followed by Pi2 and Pita. Therefore, one of the Pusa Basmati 1 NILs carrying blast resistance gene Pi9 was released for commercial cultivation. Additionally, the blast resistance genes Pi2 and Pi54 were incorporated into the genetic background of most popular Basmati rice varieties Pusa Basmati 1121 and Pusa Basmati 6 (Ellur et al. 2016). In an effort to generate multiple biotic stress tolerance Singh et al. (2012a, b) carried out introgression of blast resistance gene Pi54 along with a major QTL qSBR11-1 for sheath blight using foreground and background selection in improved Pusa Basmati 1. The linkage drag was reduced up to 4.25 mb and 1.80 mb around Pi54 and qSBR11-1, respectively, with RPG recovery of up to 89.50%.
Similarly, MABB was effectively used to incorporate blast resistance genes Pi54, Pi1, and Pita in the genetic background of a short-grain aromatic rice variety Mushk Budji. The gene-linked markers, namely Pi54-MAS, RM224, and YL87/155, were used in foreground selection to incorporate the genes Pi54, Pi1, and Pita, respectively; STS markers helped to reduce linkage drag around the genes Pi54, Pi1, and Pita to 2.74, 4.60, and 2.03 Mb, respectively; and 50K SNP chip analysis for background analysis revealed more than 92% genome similarity to recurrent parent (Khan et al. 2018).
2.9.2 Blast-Resistant Varieties Developed Through MABB in India
Pusa 1612
It is the near-isogenic line carrying blast resistance genes Pi2 and Pi54 from donor parent C101A51 and Tetep, respectively, in the genetic background of high-yielding aromatic rice variety Pusa Sugandh 5 through marker-assisted simultaneous but stepwise method of backcross breeding. Pusa 1612 was released for commercial cultivation for the Basmati-growing regions of the country (Table 2.2).
Pusa Basmati 1637
The blast resistance gene Pi9 was transferred from the donor parent IRBL9-W into the genetic background of Pusa Basmati 1 through MABB. The gene-linked marker AP5659-5 was used in foreground selection of blast resistance gene Pi9 and background selection using 104 genome-wide polymorphic SSR markers revealed the recurrent parent genome recovery of 96.6% with the linkage drag of <1.5 mb around the gene Pi9. This variety has been released for commercial cultivation in the Basmati-growing regions of the country (Table 2.2).
Pusa Basmati 1609
It is the blast-resistant Basmati rice variety possessing resistance genes Pi2 and Pi54. This variety was developed by intercrossing two breeding lines Pusa 1602 and Pusa 1603 carrying blast resistance genes Pi2 and Pi54, respectively, in the genetic background of PRR78. The markers AP5930 and RM206 linked to the genes Pi2 and Pi54 were used in foreground selection. This variety has been released for commercial cultivation in the Basmati-growing regions of the country (Table 2.2).
Pusa Samba 1850
It is a MAS-derived near-isogenic line of the mega-rice variety “BPT 5204” (Samba Mahsuri) possessing three blast genes, namely Pi54, Pi1, and Pita. These genes have been transferred from a doubled haploid line, DHMASQ164-2b. In a MABB program, foreground selection was carried out using the gene-based/linked markers Pi54MAS (Pi54), YL87/155 (Pita), YL87/183 (Pita), and RM1233 (Pi1); stringent phenotypic selection was carried out for agro-morphological, grain, and cooking qualities; and background selection was carried out using 43 polymorphic markers with genome-wide coverage. The recurrent parent genome recovery in Pusa Samba 1850 was estimated to be 86.05%. This variety has been released for the states of Chhattisgarh and Odisha (Table 2.2).
2.10 Conclusion and Future Prospects
With the identification of several blast resistance genes, marker-assisted backcross breeding has become one of the most successful approaches to develop blast resistance in the popular rice varieties. Till date, four blast-resistant rice varieties have been developed through marker-assisted selection and released for commercial cultivation. Incorporating blast resistance genes should become a routine breeding program. Adoption of blast-resistant rice varieties would significantly reduce the use of fungicides and thereby produce consumer-safe produce. Blast disease caused by M. oryzae is considered as one of the notorious diseases with population dynamics varying from location to location. Therefore, effectiveness of various genes should be tested across locations before deploying the blast resistance genes. The deployment plan needs to be developed to overcome the blast disease as well as reduce the evolution and spread of new races.
References
Ashikawa I, Hayashi N, Yamane H, Kanamori H, Wu J, Matsumoto T, Ono K, Yano M. Two adjacent nucleotide-binding site-leucine-rich repeat class genes are required to confer Pikm-specific rice blast resistance. Genetics. 2008;180:2267–76.
Ashkani S, Yusop MR, Shabanimofrad M, Azadi A, Ghasemzadeh A, Azizi P, Latif MA. Allele mining strategies: principles and utilisation for blast resistance genes in rice (Oryza sativa L.). Curr Issues Mol Biol. 2015;17:57–74.
Böhnert HU, Fudal I, Dioh W, Tharreau D, Notteghem JL, Lebrun MH. A putative polyketide synthase/peptide synthetase from Magnaporthe grisea signals pathogen attack to resistant rice. Plant Cell. 2004;16:2499–513.
Braun EJ, Howard RJ. Adhesion of fungal spores and germlings to host plant surfaces. Protoplasma. 1994;181(1–4):202–12.
Bryan GT, Wu KS, Farrall L, Jia Y, Hershey HP, McAdams SA, Faulk KN, Donaldson GK, Tarchini R, Valent B. A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta. Plant Cell. 2000;12:2033–46.
Cesari S, Thilliez G, Ribot C, Chalvon V, Michel C, Jauneau A, Rivas S, Alaux L, Kanzaki H, Okuyama Y, Fournier EJ, Tharreau D, Terauchi R, Kroja T. The rice resistance protein pair RGA4/RGA5 recognizes the Magnaporthe oryzae effectors AVR-Pia and AVR1-CO39 by direct binding. Plant Cell. 2013;25:1463–81.
Chen X, Shang J, Chen D, Lei C, Zou Y, Zhai W, Liu G, Xu J, Ling Z, Cao G, Ma B, Wang Y, Zhao X, Li S, Zhu L. A B-lectin receptor kinase gene conferring rice blast resistance. Plant J. 2006;46:794–804.
Chen J, Shi YF, Liu W, Chai R, Fu Y, Zhuang J, Wu J. A Pid3 allele from rice cultivar Gumei2 confers resistance to Magnaporthe oryzae. J Genet Genomics. 2011;38:209–16.
Chen J, Peng P, Tian J, He Y, Zhang L, Liu Z, Yin D, Zhang Z. Pike, a rice blast resistance allele consisting of two adjacent NBS-LRR genes, was identified as a novel allele at the Pik locus. Mol Breed. 2015;35:117.
Collard BCY, Mackill DJ. Marker-assisted selection: an approach for precision plant breeding in the 21st century. Philos Trans R Soc B Rev. 2008;363:557–72.
Couch BC, Kohn LM. A multilocus gene genealogy concordant with host preference indicates segregation of a new species, Magnaporthe oryzae, from M. grisea. Mycologia. 2002;94:683–93.
Dangl JL, Horvath DM, Staskawicz BJ. Pivoting the plant immune system from dissection to deployment. Science. 2013;341:746–51.
Das A, Soubam D, Singh PK, Thakur S, Singh NK, Sharma TR. A novel blast resistance gene, Pi54rh cloned from wild species of rice, Oryza rhizomatis confers broad spectrum resistance to Magnaporthe oryzae. Funct Integr Genomics. 2012;12:215–28.
Deng Y, Zhai K, Xie Z, Yang D, Zhu X, Liu J, Wang X, Qin P, Yang Y, Zhang G, Li Q, Zhang J, Wu S, Milazzo J, Mao B, Wang E, Xie H, Tharreau D, He Z. Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance. Science. 2017;355:962–5.
Devanna NB, Vijayan J, Sharma TR. The blast resistance gene Pi54of cloned from Oryza officinalis interacts with Avr-Pi54 through its novel non-LRR domains. PLoS One. 2014;9:e104840.
Dong TG, Ho BT, Yoder-Himes DR, Mekalanos JJ. Identification of T6SS-dependent effector and immunity proteins by Tn-seq in Vibrio cholera. Proc Natl Acad Sci U S A. 2013;110(7):2623–8.
Ellur RK, Khanna A, Yadav A, Pathania S, Rajashekara H, Singh VK, Krishnan SG, Bhowmick PK, Nagarajan M, Vinod KK, Prakash G. Improvement of Basmati rice varieties for resistance to blast and bacterial blight diseases using marker assisted backcross breeding. Plant Sci. 2016;242:330–41.
Feng S, Wang L, Ma J, Lin F, Pan Q. Genetic and physical mapping of AvrPi7, a novel avirulence gene of Magnaporthe oryzae using physical position-ready markers. Chin Sci Bull. 2007;52:903–11.
Fitzgerald MA, Sackville Hamilton NR, Calingacion MN, Verhoeven HA, Butardo VM. Is there a second fragrance gene in rice? Plant Biotechnol J. 2008;6:416–23.
Flor HH. The complementary genetic systems in flax rust. Adv Genet. 1956;8(29):54.
Fukuoka S, Saka N, Koga H, Ono K, Shimizu T, Ebana K, Hayashi N, Takahashi A, Hirochika H, Okuno K. Loss of function of a proline-containing protein confers durable disease resistance in rice. Science. 2009;325:998–1001.
Fukuoka S, Yamamoto SI, Mizobuchi R, Yamanouchi U, Ono K, Kitazawa N, Yasuda N, Fujita Y, Nguyen TT, Koizumi S, Sugimoto K, Matsumoto T, Yano M. Multiple functional polymorphisms in a single disease resistance gene in rice enhance durable resistance to blast. Sci Rep. 2014;4:1–7.
Fukuta Y, Araki E, Telebanco-Yanoria MJ, Ebron L, Mercado-Escueta D, Takai T, Khush GS. Identification of blast resistance gene, Pish in rice (Oryza sativa L.). In: Proc. Plant and Animal Genome XII Conference, San Diego, CA, 10–14 January. 2004. p. 401.
Ghatak A, Willocquet L, Savary S, Kumar J. Variability in aggressiveness of rice blast (Magnaporthe oryzae) isolates originating from rice leaves and necks: a case of pathogen specialization. PLoS One. 2013;8(6):e66180.
Hayashi K, Yoshida H. Refunctionalization of the ancient rice blast disease resistance gene Pit by the recruitment of a retrotransposon as a promoter. Plant J. 2009;57:413–25.
Hayashi N, Inoue H, Kato T, Funao T, Shirota M, Shimizu T, Kanamori H, Yamane H, Hayano-Saito Y, Matsumoto T. Durable panicle blast-resistance gene Pb1 encodes an atypical CC-NBS-LRR protein and was generated by acquiring a promoter through local genome duplication. Plant J. 2010;64:498–510.
Hittalmani S, Parco A, Mew TV, Zeigler RS, Huang N. Fine mapping and DNA marker-assisted pyramiding of the three major genes for blast resistance in rice. Theor Appl Genet. 2000;100(7):1121–8.
Hospital F, Charcosset A. Marker assisted introgression of quantitative trait loci. Genetics. 1997;147:1469–85.
Howard RJ, Bourett TM, Ferrari MA. In: Mendgen K, Lesemann DE, editors. Infection by Magnaporthe grisea: an in vitro analysis. Berlin: Springer; 1991a. p. 251–64.
Howard RJ, Ferrari MA, Roach DH, Money NP. Penetration of hard substrates by a fungus employing enormous turgor pressures. Proc Natl Acad Sci. 1991b;88:11281–4.
Hua L, Wu J, Chen C, Wu W, He X, Lin F, Wang L, Ashikawa I, Matsumoto T, Wang L, Pan Q. The isolation of Pi1, an allele at the Pik locus which confers broad spectrum resistance to rice blast. Theor Appl Genet. 2012;125:1047–55.
Jain P, Singh PK, Kapoor R, Khanna A, Solanke AU, Krishnan SG, Singh AK, Sharma V, Sharma TR. Understanding host-pathogen interactions with expression profiling of NILs carrying rice-blast resistance Pi9 gene. Front Plant Sci. 2017;8:9.
Khan GH, Shikari AB, Vaishnavi R, Najeeb S, Padder BA, Bhat ZA, Parray GA, Bhat MA, Kumar R, Singh NK. Marker-assisted introgression of three dominant blast resistance genes into an aromatic rice cultivar Mushk Budji. Sci Rep. 2018;8(1):1–3.
Khanna A, Sharma V, Ellur RK, Shikari AB, Gopala Krishnan S, Singh UD, Prakash G, Sharma TR, Rathour R, Variar M, Prashanthi SK, Nagarajan M, Vinod KK, Bhowmick PK, Singh NK, Prabhu KV, Singh BD, Singh AK. Development and evaluation of near-isogenic lines for major blast resistance gene(s) in Basmati rice. Theor Appl Genet. 2015a;128(7):1243–59.
Khanna A, Sharma V, Ellur RK, Shikari AB, Krishnan SG, Singh UD, Prakash G, Sharma TR, Rathour R, Variar M, Prashanthi SK. Marker assisted pyramiding of major blast resistance genes Pi9 and Pita in the genetic background of an elite Basmati rice variety, Pusa Basmati 1. Indian J Genet. 2015b;75(4):417–25.
Kumar MKP, Gowda DKS, Moudgal R, Kumar NK, Gowda KTP, Vishwanath K. Impact on fungicides on rice production in India. In: Nita M, editor. Fungicides-showcases of integrated plant disease management from around the world. InTech; 2013. ISBN: 978-953-51-1130-6. https://doi.org/10.5772/51009. http://www.intechopen.com/books/fungicides-showcases-of-integrated-plant-disease-managementfrom-around-the-world/impact-of-fungicides-on-rice-production-in-india. Accessed 18 Aug 2014.
Lee S, Song M, Seo Y, Kim S, Ko S, Cao P, Suh J, Yi G, Roh J, Lee S, An G, Hahn TR, Wang GL, Ronald P, Jeon JS. Rice Pi5-mediated resistance to Magnaporthe oryzae requires the presence of two coiled-coil-nucleotide-binding-leucine-rich repeat genes. Genetics. 2009;181:1627–38.
Li W, Wang B, Wu J, Lu G, Hu Y, Zhang X, Zhang Z, Zhao Q, Feng Q, Zhang H, Wang Z, Wang G, Han B, Wang Z, Zhou B. The Magnaporthe oryzae avirulence gene AvrPiz-t encodes a predicted secreted protein that triggers the immunity in rice mediated by the blast resistance gene Piz-t. Mol Plant Microbe Interact. 2009;22:411–20.
Lin F, Chen S, Que Z, Wang L, Liu X, Pan Q. The blast resistance gene Pi37 encodes a nucleotide binding site leucine-rich repeat protein and is a member of a resistance gene cluster on rice chromosome 1. Genetics. 2007;177:1871–80.
Liu X, Lin F, Wang L, Pan Q. The in silico map-based cloning of Pi36, a rice coiled-coil-nucleotide-binding site leucine-rich repeat gene that confers race-specific resistance to the blast fungus. Genetics. 2007;176:2541–9.
Liu Y, Liu B, Zhu X, Yang J, Bordeos A, Wang G, Leach JE, Leung H. Fine-mapping and molecular marker development for Pi56(t), a NBS-LRR gene conferring broad-spectrum resistance to Magnaporthe oryzae in rice. Theor Appl Genet. 2013;126:985–98.
Lü Q, Xu X, Shang J, Jiang G, Pang Z, Zhou Z, Wang J, Liu Y, Li T, Li X, Xu J, Cheng Z, Zhao X, Li S, Zhu L. Functional analysis of Pid3-A4, an ortholog of rice blast resistance gene Pid3 revealed by allele mining in common wild rice. Phytopathology. 2013;103:594–9.
Ma J, Lei C, Xu X, Hao K, Wang J, Cheng Z, Ma X, Ma J, Zhou K, Zhang X, Guo X, Wu F, Lin Q, Wang C, Zhai H, Wang H, Wan J. Pi64, encoding a novel CC-NBS-LRR protein, confers resistance to leaf and neck blast in rice. Mol Plant-Microbe Interact. 2015;28:558–68.
Meng X, Xiao G, Telebanco-Yanoria MJ, Siazon PM, Padilla J, Opulencia R, Bigirimana J, Habarugira G, Wu J, Li M, Wang B. The broad-spectrum rice blast resistance (R) gene Pita2 encodes a novel R protein unique from Pita. Rice. 2020;13(1):1–5.
Okuyama Y, Kanzaki H, Abe A, Yoshida K, Tamiru M, Saitoh H, Fujibe T, Matsumura H, Shenton M, Galam DC, Undan J, Ito A, Sone T, Terauchi R. A multifaceted genomics approach allows the isolation of the rice Pia-blast resistance gene consisting of two adjacent NBS-LRR protein genes. Plant J. 2011;66:467–79.
Orbach MJ, Farrall L, Sweigard JA, Chumley FG, Valent B. A telomeric avirulence gene determines efficacy for the rice blast resistance gene Pi-ta. Plant Cell. 2000;12:2019–32.
Park JY, Jin J, Lee YW, Kang S, Lee YH. Rice blast fungus (Magnaporthe oryzae) infects Arabidopsis via a mechanism distinct from that required for the infection of rice. Plant Physiol. 2009;149:474–86.
Qu SH, Liu GF, Zhou B, Bellizzi M, Zeng LR, Dai LY, Han B, Wang GL. The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rich repeat protein and is a member of a multigene family in rice. Genetics. 2006;172:1901–14.
Ray S, Singh PK, Gupta DK, Mahato AK, Sarkar C, Rathour R, Singh NK, Sharma TR. Analysis of Magnaporthe oryzae genome reveals a fungal effector, which is able to induce resistance response in transgenic rice line containing resistance gene, Pi54. Front Plant Sci. 2016;7:1140.
Schulze-Lefert P, Panstruga R. A molecular evolutionary concept connecting nonhost resistance, pathogen host range, and pathogen speciation. Trends Plant Sci. 2011;16:117–25.
Seck PA, Diagne A, Mohanty S, Wopereis MCS. Crops that feed the world. Food Sec. 2012;4:7–24.
Shang J, Tao Y, Chen X, Zou Y, Lei C, Wang J, Li X, Zhao X, Zhang M, Lu Z, Xu J, Cheng Z, Wan J, Zhu L. Identification of a new rice blast resistance gene, Pid3, by genome wide comparison of paired nucleotide-binding site-leucine-rich repeat genes and their pseudogene alleles between the two sequenced rice genomes. Genetics. 2009;182:1303–11.
Sharma TR, Madhav MS, Singh BK, Shanker P, Jana TK, Dalal V, Pandit A, Singh A, Gaikwad K, Upreti HC, Singh NK. High-resolution mapping, cloning and molecular characterization of the Pi-kh gene of rice, which confers resistance to Magnaporthe grisea. Mol Gen Genomics. 2005;274:569–78.
Sharma TR, Rai AK, Gupta SK, Singh NK. Broad-spectrum blast resistance gene Pi-kh cloned from rice line Tetep designated as Pi54. J Plant Biochem Biotechnol. 2010;19:87–9.
Silue D, Notteghem JL, Tharreau D. Evidence of a gene-for-gene relationship in the Oryza sativa-Magnaporthe oryzae pathosystem. Phytopathology. 1992;82:577–80.
Singh VP. In: Singh RK, Singh US, Khush GS, editors. Aromatic rices. New Delhi: Oxford and India Book House; 2000. p. 135–53.
Singh VK, Singh A, Singh SP, Ellur RK, Choudhary V, Sarkel S, Singh D, Gopala SK, Nagarajan M, Vinod KK, Singh UD, Rathore R, Prashanthi SK, Agrawal PK, Bhatt JC, Mohapatra T, Prabhu KV, Singh AK. Incorporation of blast resistance into “PRR78”, an elite Basmati rice restorer line through marker assisted backcross breeding. Field Crop Res. 2012a;128:8–16.
Singh A, Singh VK, Singh SP, Pandian RT, Ellur RK, Singh D, Bhowmick PK, Gopala Krishnan S, Nagarajan M, Vinod KK, Singh UD. Molecular breeding for the development of multiple disease resistance in Basmati rice. AoB Plants. 2012b;1:2012.
Singh VK, Singh A, Singh SP, Ellur RK, Singh D, Gopala Krishnan S, Bhowmick PK, Nagarajan M, Vinod KK, Singh UD, Mohapatra T, Prabhu KV, Singh AK. Marker-assisted simultaneous but stepwise backcross breeding for pyramiding blast resistance genes Pi2 and Pi54 into an elite Basmati rice restorer line PRR78. Plant Breed. 2013;132(5):486–95.
Su J, Wang W, Han J, Chen S, Wang C, Zeng L, Feng A, Yang J, Zhou B, Zhu X. Functional divergence of duplicated genes results in a novel blast resistance gene Pi50 at the Pi2/9 locus. Theor Appl Genet. 2015;128:2213–25.
Takagi H, Uemura A, Yaegashi H, Tamiru M, Abe A, Mitsuoka C, Utsushi H, Natsume S, Kanzaki H, Matsumura H, Saitoh H, Yoshida K, Cano LM, Kamoun S, Terauchi R. MutMap-Gap: whole-genome resequencing of mutant F2 progeny bulk combined with de novo assembly of gap regions identifies the rice blast resistance gene Pii. New Phytol. 2013;200:276–83.
Takahashi A, Hayashi N, Miyao A, Hirochika H. Unique features of the rice blast resistance Pish locus revealed by large scale retrotransposon-tagging. BMC Plant Biol. 2010;10:175.
Talbot NJ. On the trail of a cereal killer: exploring the biology of Magnaporthe grisea. Annu Rev Microbiol. 2003;57:177–202.
Tanksley SD. Molecular markers in plant breeding. Plant Mol Biol Rep. 1983;1:1–3.
Tanweer F, Rafii M, Sijam K, Rahim H, Ahmed F, Latif M. Current advance methods for the identification of blast resistance genes in rice. C R Biol. 2015;338:321–34.
Telebanco-Yanoria MJ, Koide Y, Fukuta Y, Imbe T, Tsunematsu H, Kato H, Ebron LA, Nguyet TMN, Kobayashi N. A set of near-isogenic lines of Indica-type rice variety CO39 as differential varieties for blast resistance. Mol Breed. 2011;27:357–73.
Tsunematsu H, Yanoria MJT, Ebron LA, Hayashi N, Ando I, Kato H, Imbe T, Khush GS. Development of monogenic lines of rice for blast resistance. Breed Sci. 2000;50:229–34.
Wang Z, Yano M, Yamanouchi U, Iwamoto M, Monna L, Hayasaka H, Katayose Y, Sasaki T. The Pib gene for rice blast resistance belongs to the nucleotide binding and leucine-rich repeat class of plant disease resistance genes. Plant J. 1999;19:55–64.
Wang B, Ebbole DJ, Wang Z. The arms race between Magnaporthe oryzae and rice: diversity and interaction of Avr and R genes. J Integr Agric. 2017;16(12):2746–60.
Wilson RA, Talbot NJ. Under pressure: investigating the biology of plant infection by Magnaporthe oryzae. Nat Rev Microbiol. 2009;7(3):185–95.
Wu J, Kou Y, Bao J, Li Y, Tang M, Zhu X, Ponaya A, Xiao G, Li J, Li C, Song MY, Cumagun CJR, Deng Q, Lu G, Jeon JS, Naqvi NI, Zhou B. Comparative genomics identifies the Magnaporthe oryzae avirulence effector AvrPi9 that triggers Pi9-mediated blast resistance in rice. New Phytol. 2015;206:1463–75.
Xu X, Hayashi N, Wang C, Fukuoka S, Kawasaki S, Takatsuji H, Jiang C. Rice blast resistance gene Pikahei-1(t), a member of a resistance gene cluster on chromosome 4, encodes a nucleotide-binding site and leucine-rich repeat protein. Mol Breed. 2014;34:691–700.
Yoshida K, Saitoh H, Fujisawa S, Kanzaki H, Matsumura H, Yoshida K, Tosa Y, Chuma I, Takano Y, Win J, Kamoun S, Terauchi R. Association genetics reveals three novel avirulence genes from the rice blast fungal pathogen Magnaporthe oryzae. Plant Cell. 2009;21:1573–91.
Yuan B, Zhai C, Wang WJ, Zeng XS, Xu XK, Hu HQ, Lin F, Wang L, Pan QH. The Pik-p resistance to Magnaporthe oryzae in rice is mediated by a pair of closely linked CC-NBS-LRR genes. Theor Appl Genet. 2011;122:1017–28.
Zhai C, Lin F, Dong ZQ, He XY, Yuan B, Zeng XS, Wang L, Pan QH. The isolation and characterization of Pik, a rice blast resistance gene which emerged after rice domestication. New Phytol. 2011;189:321–34.
Zhai C, Zhang Y, Yao N, Lin F, Liu Z, Dong Z, Wang L, Pan Q. Function and interaction of the coupled genes responsible for Pik-h encoded rice blast resistance. PLoS One. 2014;9:e98067.
Zhang S, Wang L, Wu W, He L, Yang X, Pan Q. Function and evolution of Magnaporthe oryzae avirulence gene AvrPib responding to the rice blast resistance gene Pib. Sci Rep. 2015;5:11642.
Zheng Y, Zheng W, Lin F, Zhang Y, Yi Y, Wang B, Lu G, Wang Z, Wu W. AVR1-CO39 is a predominant locus governing the broad avirulence of Magnaporthe oryzae 2539 on cultivated rice (Oryza sativa L.). Mol Plant-Microbe Interact. 2011;24:13–7.
Zhou B, Qu SH, Liu GF, Dolan M, Sakai H, Lu GD, Bellizzi M, Wang GL. The eight amino-acid differences within three leucine-rich repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to Magnaporthe grisea. Mol Plant-Microbe Interact. 2006;19:1216–28.
Zhu X, Chen S, Yang J, Zhou S, Zeng L, Han J, Su J, Wang L, Pan Q. The identification of Pi50(t), a new member of the rice blast resistance Pi2/Pi9 multigene family. Theor Appl Genet. 2012;124:1295–304.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Khanna, A. et al. (2021). Utilizing Host-Plant Resistance to Circumvent Blast Disease in Rice. In: Nayaka, S.C., Hosahatti, R., Prakash, G., Satyavathi, C.T., Sharma, R. (eds) Blast Disease of Cereal Crops. Fungal Biology. Springer, Cham. https://doi.org/10.1007/978-3-030-60585-8_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-60585-8_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-60584-1
Online ISBN: 978-3-030-60585-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)