Abstract
We report the isolation of Pi1, a gene conferring broad-spectrum resistance to rice blast (Magnaporthe oryzae). Using loss- and gain-of-function approaches, we demonstrate that Pi1 is an allele at the Pik locus. Like other alleles at this locus, Pi1 consists of two genes. A functional nucleotide polymorphism (FNP) was identified that allows differentiation of Pi1 from other Pik alleles and other non-Pik genes. A extensive germplasm survey using this FNP reveals that Pi1 is a rare allele in germplasm collections and one that has conferred durable resistance to a broad spectrum of pathogen isolates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The deployment of host resistance genes (R genes) is a cost-effective and environmentally sustainable means of controlling plant diseases (Khush and Jena 2009; Liu et al. 2010). Therefore, many crop breeding programs have focused on incorporating such alleles into elite cultivars (cvs). However, the majority of R genes do not provide long-term resistance, since most pathogens have sufficient genetic flexibility to eventually overcome them (Krattinger et al. 2009; Lee et al. 2009; Skamnioti and Gurr 2009; Liu et al. 2010). To ensure its longevity, the ideal R gene must combine a broad spectrum of resistance with durability. Of the several broad-spectrum R genes which have been characterized in some detail to date, perhaps the most renowned are the barley gene mlo (Büschges et al. 1997), and the rice genes Xa21 (Wang et al. 1996) and Pi9 (Qu et al. 2006). However, the underlying mechanisms of broad-spectrum resistance are still not well understood.
To keep pace with the diversifying pathogens, multiple alleles at a particular R locus are prevalent when they provide a principal fitness advantage by conferring effective disease resistance to the corresponding pathogens (Holub 2001). The basis of the race specificity expressed by R gene alleles can be conveniently studied by a comparative sequence analysis. A good example is the flax L locus, where Ellis et al. (1999) showed that the 13 known alleles shared >90 % sequence identity and that differences in the LRR domain and N-terminal region were associated with race specificity. A similar analysis of the wheat Pm3 gene suggested that specificity was associated with a single residue in the C-terminal LRR motif (Brunner et al. 2010). Three of the five alleles at the Pik locus of rice (Pik, Pik-p and Pik-m) have been isolated and characterized (Ashikawa et al. 2008; Yuan et al. 2011; Zhai et al. 2011), and their functionality is determined by a pair of closely linked CC-NBS-LRR genes, which contain functional nucleotide polymorphisms (FNPs) in either their N-terminal regions and/or their LRR domains (Zhai et al. 2011).
The conventional identification of alleles based on phenotype is limited by the number of pathogen isolates available. However, for R genes which have been isolated, it is now possible to replace phenotypes with DNA polymorphisms as the basis for defining alleles. The identification of FNPs, which include single nucleotide polymorphisms (SNPs) and insertion/deletions (InDels), which are responsible for allele specificity is particularly useful in the context of molecular breeding programs (Hausner et al. 1999; Tommasini et al. 2006; Shomura et al. 2008; Asano et al. 2011; Yuan et al. 2011; Zhai et al. 2011).
In rice, alleles at the Pik locus confer resistance to rice blast (incited by Magnaporthe oryzae). The Pi1 R gene, derived from the durably resistant West African cv. LAC23 (Mackill and Bonman 1992; Inukai et al. 1994), has displayed a high level of durable resistance against blast (Hittalmani et al. 2000; Fuentes et al. 2008; Yang et al. 2008; Tacconi et al. 2010) and was hypothesized to be an allele at the Pik locus. The objectives of the present study were to (i) isolate Pi1 by the in silico map-based cloning technique (Yuan et al. 2011; Zhai et al. 2011), (ii) characterize its effectiveness against a range of isolates and (iii) determine its frequency in a range of germplasm accessions. Pi1 proved to be a novel Pik allele and was distinguishable from other Pik alleles and R genes at other loci, conferring blast resistance by an FNP we identified.
Materials and methods
Blast resistance characterization
Standard carriers of Pi1 (cv. C101LAC), Pik (cv. Kusabue), Pik-p (cv. K60), Pik-m (cv. Tusyuake) and Pi21 (cv. C101A5) and a universal susceptible cv. (CO39) were challenged with 1,292 blast isolates collected over 19 cropping seasons (between 1998 and 2008) in Guangdong Province (GD), and 480 isolates originating from Fujian (FJ, 40 isolates), Hunan (HN, 40 isolates), Yunnan (YN, 43 isolates), Guizhou (GZ, 60 isolates), Sichuan (SC, 66 isolates), Liaoning (LN, 108 isolates), Jilin (JL, 60 isolates) and Heilongjiang (HLJ, 63 isolates) provinces, China. The methods for inoculation and resistance evaluation followed Pan et al. (2003).
Genetic mapping of Pi1 and candidate gene identification
A set of F2 progeny generated from the cross cv. C101LAC (Pi1) × cv. K1 was screened for reaction to inoculation with blast isolate EHL0490. As Pi1 was suspected to be an allele of Pik (Inukai et al. 1994), DNA markers known to map within the Pik-m/Pik-p region (Li et al. 2007; Wang et al. 2009) were used to genotype the F2 progeny. The gene annotation tools RiceGAAS (http://ricegaas.dna.affrc.go.jp) and FGENSH (http://www.softberry.com) were then used to identify Pi1 candidates within the 277-kb segment of cv. Nipponbare defined by the closest flanking markers (CRG11-7 and K28). The DNA sequence surrounding Pik has been described as the functional “K type” (cvs. Tsuyuake, K60, and Kusabue) or the nonfunctional “N type” (cv. Nipponbare) (Ashikawa et al. 2008; Yuan et al. 2011; Zhai et al. 2011). Therefore, the four candidates for Pi1 within the critical insertion/deletion region were validated by a presence/absence analysis with “K” and “N” haplotype specific primer sets designed based on the BAC clone Ts18H12 of cv. Tsuyuake and the reference sequence of cv. Nipponbare, respectively. The association between SNPs within each candidate for Pi1- and Pi1-mediated resistance was examined by SNP genotyping a set of Pi1 carrier and non-carrier cvs.
Loss- and gain-of-function tests
Conventional F1 hybrids were made between the Pik allelic RNAi stock kh4i-16 (Yuan et al. 2011) and cv. C101LAC (Pi1), and between kh4i-11 (an alternative RNAi Pik knock-out line) and the Pia carrier cv. Aichi Asahi (Pia is not an allele of Pik), which were inoculated with blast isolate CHL381 (avirulent on both cvs. C101LAC and Aichi Asahi). A correlation between susceptibility and the presence of the RNAi construct was established using a PCR assay based on primers targeting the transgene. RT-PCR was used to verify that susceptibility was correlated with a lack of Pi1 expression. The RT-PCR reference gene was a fragment of the rice Actin gene. To test for gain of function, two of the candidates for Pi1 (Pi1-5C and Pi1-6C) were combined to form the sequence Pi1-56C. Based on the strategy used for Pik-p (Yuan et al. 2011), the Pi1-56C sequence was then divided into four fragments, each of which was independently amplified using Phusion™ High-Fidelity DNA Polymerase (NEB, Beijing, China) and introduced into either pMD20-T (TaKaRa, Dalian, China) or pCAMBIA 1300Asc I. The four fragments were fused in the appropriate orientation and order in pCAMBIA1300Asc I (details given in Fig. S2), which was then transformed into the blast susceptible cvs. Q1063, Longjing24 (LJ) and Nipponbare, by Agrobacterium-mediated transformation (Yuan et al. 2011; Zhai et al. 2011). As a control, two less-promising candidates for Pi1 (Pi1-3C and Pi1-4C) were similarly coupled to form Pi1-34, and this construct was also transformed into the susceptible cv. Q1063 in the same way. The resulting T1 plants were challenged with the Pi1 avirulent blast isolates CHL346 (for cvs. Q1063 and Nipponbare) or CHL357 (cv. LJ). T2 progeny segregating in a Mendelian fashion corresponding to the isolates were used to establish the relationship between resistance and the presence of the transgene.
Sequence analysis
The entire coding sequences of Pi1-5C and Pi1-6C were amplified using the primer sets KP3ORF F/R and KP4ORF F/R, respectively (Table S1). Both 5′ and 3′-rapid amplification of cDNA ends was obtained using a GeneRacer™-kit (Invitrogen, http://www.invitrogen.com), following the same strategy used for Pik-p (Fig. S2; Yuan et al. 2011). The intron–exon structure of both genes was deduced by comparing the resulting cDNA sequences with their respective gDNA sequence. Further sequence comparisons were performed by pairwise BLAST (http://www.ncbi.nlm.nih.gov/BLAST/bl2seq/bl2.html), and the similarity between predicted peptide sequences by BLASTP. The prediction of the CC domain was based on COILS (http://www.ch.embnet.org/software/COILS_form.html) and Paircoil2 (http://groups.csail.mit.edu/cb/paircoil2/) analysis. The peptide sequences of the various Pik allele products were aligned using Multalin software (http://bioinfo.genotoul.fr/multalin/multalin.html).
SNP genotyping
A germplasm array consisting of 636 rice accessions (198 wild rice accessions, 180 indica landraces, 91 japonica landraces, 86 indica cultivars and 81 japonica cultivars) was genotyped for N and K haplotypes (Table S2), following the method described by Zhai et al. (2011). Accessions having the K type of genome were subjected to genotyping using the Pi1-specific FNP marker through dCAPS analysis. The haplotypes of the four Pik alleles (Pi1, Pik, Pik-p and Pik-m) were further subjected to allele distribution analysis.
Results
The blast resistance spectrum and durability of Pi1
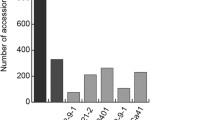
The differential reactions to the various blast isolates showed that Pi1 is distinct from other Pik alleles (Pik, Pik-p and Pik-m), as well as from R genes at the Pia and Pi2 loci. Pi1 gave a consistent level of resistance in GD throughout the period 1998–2008, suggesting that it is durable (Fig. 1a). The gene also conferred resistance to a broad range of blast isolates from across China, although it was ineffective against some blast isolates collected from the JL Province (Fig. 1b).
Frequency of resistance among sets of blast isolates shown by three near isogenic lines (cvs. CO39, C101A51 and C101LAC) and three cvs carrying different Pik alleles. a A set of pathogen populations collected over 19 growing seasons in Guangdong Province. Codes 98E and 99L indicate populations collected, respectively, in the early season (March–July) of 1998 and the late season (July–November) of 1999, etc. b Populations collected from Fujian (FJ), Hunan (HN), Yunan (YN), Guizhou (GZ), Sichuan (SC), Jiangsu (JS), Liaoning (LN), Jilin (JL), Heilongjiang (HLJ) provinces in China
Genetic and physical maps of the Pi1 locus
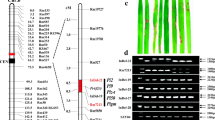
With respect to the reaction to inoculation with blast isolate EHL0490 (avirulent on cv. C101LAC, virulent on cv. K1), the C101LAC/K1 F2 progeny segregated at a ratio 489 resistant to 161 susceptible, consistent with the presence of a single gene for resistance (χ 2c = 0.01, P < 0.90). When the markers for the Pik-m/Pik-p/Pik region (Table S1) were applied to the 161 blast susceptible segregants, five individuals were shown to be recombinant with respect to blast reaction and marker K37, one with respect to K28 and none with respect to K33 (Fig. 2). Of the three polymorphic CRG (Candidate Resistance Gene) markers identified from the equivalent region in cv. Nipponbare, CRG11-7 identified three recombinants, while both CRG11-5 and CRG11-6 identified none (Fig. 2). We concluded that the location of Pi1 was confined to a region delimited by CRG11-7 and K28, equivalent to a 277-kb stretch of chromosome 11, overlapping with the location of the Pik locus (Fig. 2).
Integrated genetic and physical maps of the Pik/Pi1 locus. a Genetic map of the Pi1 locus. The numbers in parenthesis are recombinants/gametes identified at the respective marker locus. b Contig map of the target region flanked by markers CRG11-7 and K28. c Physical map of the Pi1 locus in the Nipponbare type genome, where six candidate genes with NBS-LRR structure were predicted by the gene annotation program, RiceGAAS. d Physical map of of the Pi1 locus in the C101LAC type genome, where six candidate genes with the conserved R gene structures, i.e., Pi1-1C (NB-ARC), Pi1-2C (PK), Pi1-3C (NB-ARC), Pi1-4C (NBS), Pi1-5C (NBS-LRR), and Pi1-6C (NBS-LRR), were predicted based on the sequence of the BAC clones, Ts50A13 and Ts18H12 derived from cv Tsuyuake. A large InDel between Nipponbare and Tsuyuake/K60/C101LAC genomes was also indicated according to Ashikawa et al. (2008) (also see Fig. S1). The anchor markers were extracted from a Zhai et al. (2011); b Ashikawa et al. (2008); c Xu et al. (2008)
Candidate gene identification
The K- and N-haplotypes surrounding Pik differ by a large InDel (Ashikawa et al. 2008; Yuan et al. 2011; Zhai et al. 2011), so both the sequences of cv Nipponbare and BAC clones Ts50A13 and Ts18H12 from cv. Tsuyuake were used as a reference to annotate the candidate region (Fig. 2d). This produced a set of six genes having the expected R gene structure (Fig. 2c, d). The presence/absence analysis applied to genes Pi1-1 to Pi1-4 showed that Pi1-1N to Pi1-4N were all absent in cv. C101LAC, while Pi1-1C to Pi1-4C were all present (Figs. 2; S1). Thus, cv. C101LAC is the K-haplotype. A perfect association was established between SNP genotype and blast reaction among a set of Pi1 carriers and non-carriers for both Pi1-5C and Pi1-6C (Table 1), promoting these two genes above the four others as candidates for Pi1. Both are orthologs of Pik-m/Pik-p/Pik (Fig. 2d; Ashikawa et al. 2008; Yuan et al. 2011; Zhai et al. 2011). The Pik-m, Pik-p and Pik “alleles” are defined by specific sequences at two physically linked genes (Ashikawa et al. 2008; Yuan et al. 2011; Zhai et al. 2011). In the remainder of this report, we will follow the convention for describing alleles at Pik, where “allele” is used to define specific sequences at two genes. We hypothesize that Pi1 also comprises two genes: Pi1-5C and Pi1-6C. This hypothesis will be tested in further experiments.
Functional characterization of the candidate genes
The three C101LAC/Kh4i-16 F1 plants tested showed no resistance to the Pi1/Pia-avirulent blast isolate CHL381 (Figs. 3a; S3), while those tested from the Aichi Asahi/Kh4i-11 cross maintained resistance to the blast isolate CHL381 (Fig. 3a). Gene expression analysis revealed that the loss of Pi1 function reflected the lack of its expression (Fig. 3b). To confirm that Pi1 function required the presence of Pi1-56C, a binary construct harboring Pi1-56C and Pi1-34C was introduced into cvs. Q1063, LJ and Nipponbare (Table 2). Many of the transformants carrying the Pi1-56C transgene were resistant to infection by Pi1-avirulent blast isolates, unlike any of the Pi1-34C transformants (Table 2; Figs. S4–S6). An unambiguous association between resistance and the presence of the Pi1-56C transgene was established in later generations derived from the primary transformants (Fig. 4). Thus, it was clear that Pi1-56C, and not Pi1-34C, was required for Pi1 function. Pi1-56C homozygous progeny shared the same race specificity as the Pi1 standard cv. IRBL1-CL (data not shown).
Molecular characterization of hybrid F1 plants derived from the conventional crosses for loss-of-function analysis of Pi1. a The amplification of a fragment of the gus-linker sequence indicates the presence of the RNAi construct targeting Pik-p. b Expression analysis of Pi1 (lanes 1–5) or Pia (6–9). The lower panel shows the expression of the reference gene actin. Lane 1 cv. C101LAC (Pi1), lane 2 kh4i-16 (Pik-p-RNAi construct), lanes 3–5 C101LAC/kh4i-16 F1 plants, lane 6 cv. Aichi Asahi (Pia), lane 7 kh4i-11 (Pik-p-RNAi construct), lanes 8–9 Aichi Asahi/kh4i-11 F1 plants. M DNA ladder. Plants inoculated with the blast isolate CHL381 (avirulent on all Pik alleles including Pi1, as well as on Pia). The F1 plants in lanes 3–5 were susceptible (S), and those in lanes 8–9 were resistant
Co-segregation of resistance with the presence of Pi1-56C in segregating transgenic progeny. M DNA ladder, V vector, LJ cv Longjing 24, R resistant, MS moderately susceptible, S susceptible (also see Table 2). The codes of Pi1-56C/LJ-46 and Pi1-56C/LJ-48 indicate #46 and #48 T2 lines generated from transformation of Pi1-56C
The structure of Pi1
Pi1-5C encodes a 1,143 residue polypeptide and has the same intron/exon structure as Pikm-1 and Pik-1 and a very similar one to that of Pikp-1 (Fig. S7; Ashikawa et al. 2008; Yuan et al. 2011; Zhai et al. 2011). The Pi1-5C sequence shares, respectively, 99, 99 and 95 % peptide identity with those of Pikm-1, Pik-1 and Pikp-1. Pi1-6C encodes a 1,021 residue polypeptide and, apart from an extra intron in its 5′-UTR, has the same intron/exon structure as Pikm-2 and Pik-2. The 5′-UTR intron is also present in Pikp-2, but not in either Pik-2 or Pikm-2 (Fig. S8; Ashikawa et al. 2008; Yuan et al. 2011; Zhai et al. 2011). The Pi1-6C peptide sequence is 99 % identical with those of Pikm-2, Pik-2 and Pikp-2. At the nucleotide sequence level, Pi1 can be distinguished from other Pik alleles with respect to two non-synonymous SNPs, namely Pi1-5SNP in the Pi1-5C CC domain (T1-687G) and Pi1-6SNP in the Pi1-6C LRR domain (T2-2261A) (Figs. S7, S8). The Pi1 nucleotide sequence has been deposited in GenBank as accession HQ606329.
Distribution of Pi1
When the Pi1-5SNP and Pi1-6SNP SNP assays were applied to a large germplasm panel, it was apparent that only the latter was diagnostic for Pi1 (Fig. 5). The survey of wild rice accessions and indica and japonica landrace and cvs (Tables S2 and S3) showed that Pi1 was only present in two indica cvs (Tetep and C101LAC) and one japonica cv (IRBL1-CL, which was derived from cv. C101LAC). The presence of Pi1 only in the modern cultivar populations suggested that it probably evolved later than Pik-m, Pik-p or Pik (Table S3). Tetep was bred in SE Asia (Vietnam) and LAC23 (the Pi1 donor to C101LAC and IRBL1-CL) originated in W Africa (Liberia).
The Pi1 allele-specific FNP distinguishes it from other blast R genes including the five Pik alleles. Lane 1 cv. C101LAC (Pi1), Lane 2 cv. CO39 (susceptible parent of cv. C101LAC), lane 3 IRBL1-CL (Pi1), lane 4 cv. LTH, lane 5 cv. Tetep (Pi1), lane 6 IRBLks-S (Pik-s), lane 7 IRBLk-Ka (Pik), lane 8 IRBLkp-K60 (Pik), lane 9 IRBLkh-K3 (Pik-h), lane 10 IRBLKM-TS (Pik-m), lane 11 IRBLz-CA (Pi2), lane 12 IRBL7-M (Pi7), lane 13 IRBL9-W (Pi9). M DNA ladder
Discussion
We have shown that Pi1 is a novel and rare allele at the Pik locus. Pi1 is now the fourth cloned and characterized allele at this complex locus. The Pik locus has proven valuable to rice breeders concerned with blast resistance, as most alleles impart broad spectrum and durable resistance (Kiyosawa 1987; Ashikawa et al. 2008; Xu et al. 2008; Wang et al. 2009; Yuan et al. 2011; Zhai et al. 2011). Pi1 has been used in a number of rice breeding programs. For example, it was combined with Pi2 and Pita by IRRI (Hittalmani et al. 2000) for use in both the Philippines and India; by Fuentes et al. (2008) in Latin America; in Europe by Tacconi et al. (2010), and in China it is regarded as having great potential for achieving broad spectrum and durable resistance (Li et al. 2005; Yang et al. 2008; Hu et al. 2010; Zhang et al. 2010). Our results confirm that the Pi1 allele is effective against many, but not all, isolates collected from the broad range of cropping seasons and regions.
R genes are characteristically organized into clusters, and even as alleles at the particular locus (Bai et al. 2002; Wang et al. 2009). Before the development of marker technology, the effective characterization of R genes depended entirely on testing with a panel of race-specific pathogen isolates. This approach is problematic whenever no matching race is available which is virulent against a carrier of a novel allele. The advent of DNA markers has encouraged the implementation of the R gene pyramiding concept, which in principle avoids the need for inoculation (Huang et al. 1997; Hayashi et al. 2004). The requirement for this strategy is that markers for all genes in the pyramid are both diagnostic and robust (Tommasini et al. 2006; Hayashi et al. 2010; Perumalsamy et al. 2010). Ideally, each marker needs to be gene (or allele) specific, as is an FNP. The bread wheat R locus Pm3 provides an example of the development of FNPs (Tommasini et al. 2006), in which seven distinct alleles have been independently marked. Not only can these FNPs be used to search for novel alleles in large germplasm sets and as effective selection tools for resistance breeding, but they also have the potential to allow the parallel introduction of multiple alleles into a single individual plant via a transgenic approach. The current situation with the Pik locus is that four allele-specific FNPs (Pik, Pik-p, Pik-m, and Pi1) are now available (Fig. 5; Yuan et al. 2011; Zhai et al. 2011). These FNP markers can be widely used to stack the Pik alleles with other R genes in elite cvs via MAS and transgenic approaches.
Only 2 of 636 germplasm accessions surveyed carry Pi1. One accession (LAC23) is from W Africa and the other (Tetep) in SE Asia. The presence of Pi1 only in both modern cultivars suggests that it evolved relatively recently, unlike Pik-p which pre-dates the domestication of rice, and Pik and Pik-m, both of which evolved somewhat after rice domestication (Zhai et al. 2011). The geographic dispersal of Pi1 may trace to the introduction of rice to Africa in the sixteenth century (Linares 2002). Alternatively, active selection for blast resistance could easily have promoted the incorporation of the rare Pi1 into local cultivars in both SE Asia and W Africa (Gross et al. 2010). It is testimony to the effectiveness of plant breeding that such a valuable and potentially durable allele would occur at such low frequency in germplasm collections and at a higher frequency in contemporary breeding programs.
References
Asano K, Yamasaki M, Takuno S, Miura K, Katagiri S, Ito T, Doi K, Wu J, Ebana K, Matsumoto T, Innan H, Kitano H, Ashikari M, Matsuoka M (2011) Artificial selection for a green revolution gene during japonica rice domestication. Proc Natl Acad Sci 108:11034–11039
Ashikawa I, Hayashi N, Yamane H, Kanamori H, Wu J, Matsumoto T, Ono K, Yano M (2008) Two adjacent nucleotide-binding site-leucine rich repeat class genes are required to confer Pikm-specific rice blast resistance. Genetics 180:2267–2276
Bai J, Pennill LA, Ning J, Lee S, Ramalingam J, Webb C, Zhao B, Sun Q (2002) Diversity in nucleotide binding site-leucine-rich repeat genes in cereals. Genome Res 12:1871–1884
Brunner S, Hurni S, Streckeisen P, Mayr G, Albrecht M, Yahiaoui N, Keller B (2010) Intragenic allele pyramiding combines different specificities of wheat Pm3 resistance alleles. Plant J 64:433–445
Büschges R, Hollricher K, Panstruga R, Simons G, Wolter M, Frijters A, van Daelen R, van der Lee T, Diergaarde P, Groenendijk J, Töpsch S, Vos P, Salamini F, Schulzelefert P (1997) The barley mlo gene: a novel control element of plant pathogen resistance. Cell 88:695–705
Ellis JG, Lawrence GJ, Luck JE, Dodds PN (1999) Identification of regions in alleles of the flax rust resistance gene L that determine differences in gene-for-gene specificity. Plant Cell 11:495–506
Fuentes JL, Correa-Victoria FJ, Escobar F, Prado G, Aricapa G, Duque MC, Tohme J (2008) Identification of microsatellite markers linked to the blast resistance gene Pi-1(t) in rice. Euphytica 160:295–304
Gross BL, Steffen FT, Olsen KM (2010) The molecular basis of white pericarps in African domesticated rice: novel mutations at the Rc gene. Evol Biol 23:2747–2753
Hausner G, Rashid KY, Kenaschuk EO, Procunier JD (1999) The development of codominant PCR/RFLP based markers for the flax rust-resistance alleles at the L locus. Genome 42:1–8
Hayashi K, Hashimoto N, Daigen M, Ashikawa I (2004) Development of PCR-based SNP markers for rice blast resistance genes at the Piz locus. Theor Appl Genet 108:1212–1220
Hayashi K, Yasuda N, Fujita Y, Koizumi S, Yoshida H (2010) Identification of the blast resistance gene Pit in rice cultivars using functional markers. Theor Appl Genet 121:1357–1367
Hittalmani S, Parco A, Mew TV, Zeigler RS (2000) Fine mapping and DNA marker-assisted pyramiding of three major genes for blast resistance in rice. Theor Appl Genet 100:1121–1128
Holub EB (2001) The arms race is ancient history in Arabidopsis, the wildflower. Nat Rev Cenet 2:516–527
Hu J, Li X, Wu C (2010) Gene pyramiding to improve the resistance of rice hybrids to brown planthopper and blast disease using molecular marker-assisted selection. Mol Plant Breed 8:1180–1187 (in Chinese)
Huang N, Angeles ER, Domingo J, Magpantay G, Singh S, Zhang G, Kumaravadivel N, Bennett J, Khush GS (1997) Pyramiding of bacterial blight resistance genes in rice: marker-assisted selection using RFLP and PCR. Theor Appl Genet 95:313–320
Inukai T, Nelson RJ, Zeigler RS, Sarkarung S, Mackill DJ, Bonman JM, Takamure I, Kinoshita T (1994) Allelism of blast resistance genes in near-isogenic lines of rice. Phytopathology 84:1278–1283
Khush GS, Jena KK (2009) Current status and future prospects for research on blast resistance in rice (Oryza sative L.). In: Wang GL, Valent B (eds) Advances in genetics, genomics and control of rice blast disease. Springer, Dordrecht, pp 1–10
Kiyosawa S (1987) With genetic view on the mechanism of resistance and virulence. Jpn J Genet 41:89–92
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363
Lee S, Costanzo S, Jia Y, Olsen KM, Caicedo AL (2009) Evolutionary dynamics of the genomic region around the blast resistance gene Pi-ta in AA genome Oryza species. Genetics 183:1315–1325
Li J, Li C, Chen Y, Lei C, Ling Z (2005) Evaluation of twenty-two blast resistance genes in Yunnan using monogenetic rice lines. Acta Phytophylacica Sin 32:113–119 (in Chinese)
Li L, Wang L, Jing J, Li Z, Lin F, Pan Q (2007) The Pikm gene, conferring stable resistance to isolates of Magnaporthe oryzae was finely mapped in a crossover-cold region on rice chromosome 11. Mol Breed 20:179–188
Linares OF (2002) African rice (Oryza glaberrima): history and future potential. Proc Natl Acad Sci 99:16360–16365
Liu J, Wang X, Mitchell T, Hu Y, Liu X, Dai L, Wang GL (2010) Recent progress and understanding of the molecular mechanisms of the rice–Magnaporthe oryzae interaction. Mol Plant Pathol 11:419–427
Mackill DJ, Bonman LM (1992) Inheritance of blast resistance in near-isogenic lines of rice. Phytopathology 82:746–749
Pan Q, Hu Z, Tanisaka T, Wang L (2003) Fine mapping of the blast resistance gene Pi15, linked to Pii, on rice chromosome 9. Acta Bot Sin 45:871–877
Perumalsamy S, Bharani M, Sudha M, Nagarajan P, Arul L, Saraswathi R, Balasubramanian P, Ramalingam J (2010) Functional marker-assisted selection for bacterial leaf blight resistance genes in rice (Oryza sativa L.). Plant Breed 129:400–406
Qu S, Liu G, Zhou B, Bellizzi M, Zeng L, Dai L, Han B, Wang GL (2006) The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rice repeat protein and is a member of a multigene family in rice. Genetics 172:1901–1914
Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M (2008) Deletion in a gene associated with grain size increased yields during rice domestication. Nat Genet 40:1023–1028
Skamnioti P, Gurr SJ (2009) Against the grain: safeguarding rice from rice blast disease. Trend Biotechnol 27:141–150
Tacconi G, Baldassarre V, Lanzanova C, Faivre-Rampant O, Cavigiolo S, Urso S, Lupotto E, Valè G (2010) Polymorphism analysis of genomic regions associated with broad-spectrum effective blast resistance genes for marker development in rice. Mol Breed 26:595–617
Tommasini L, Yahiaoui N, Srichumpa P, Keller B (2006) Development of functional markers specific for seven Pm3 resistance alleles and their validation in the bread wheat gene pool. Theor Appl Genet 114:165–175
Wang GL, Song WY, Ruan DL, Sideris S, Ronald PC (1996) The closed gene, Xa21, confers resistance to multiple Xanthomonas oryzae pv. oryzae isolates in transgenic plants. Mol Plant Microbe Interact 9:850–855
Wang L, Xu X, Lin F, Pan Q (2009) Characterization of rice blast resistance genes in the Pik cluster and fine mapping of the Pik-p locus. Phytopathology 99:900–905
Xu X, Hayashi N, Wang CT, Kato H, Fujimura T, Kawasaki S (2008) Efficient authentic fine mapping of the rice blast resistance gene Pik-h in the Pik cluster, using new Pik-h-differentiating isolates. Mol Breed 22:289–299
Yang X, Zhu C, Ruan H, Du Y, Guan R, Chen F (2008) Pathogenic types of Magnaporthe grisea Barr. and resistance of some rice cultivars to the pathogens in Fujian province. J Fujian Agr Fore Uni 37:243–247 (in Chinese)
Yuan B, Zhai C, Wang W, Zeng X, Xu X, Hu H, Lin F, Wang L, Pan Q (2011) The Pik-p resistance to Magnaporthe oryzae in rice is mediated by a pair of closely linked CC-NBS-LRR genes. Theor Appl Genet 122:1017–1028
Zhai C, Lin F, Dong Z, He X, Yuan B, Zeng X, Wang L, Pan Q (2011) The isolation and characterization of Pik, a rice blast resistance gene which emerged after rice domestication. New Phytol 189:321–334
Zhang C, Ma J, Xiao J, Liu Y, Xin A, Ren Y (2010) The blast resistance of 24 monogenic rice lines to prevalence physiologic races of Heilongjiang and analysis of pathogenicity association. Chi Agr Sci Bull 26:233–237 (in Chinese)
Acknowledgments
We thank Z. Wang (Fujian Agricultural University), Z. Zhao (Hunan Province Agricultural Academy of Sciences), D. Lu (Sichuan Province Agricultural Academy Sciences), J. Yuan (Guizhou Province Agricultural Academy of Sciences), Z. Liu (Shenyang Agricultural University), X. Guo (Jilin Province Agricultural Academy of Sciences) and G. Zhang (Heilongjiang Province Agricultural Academy of Sciences) for providing blast isolates. This study was supported by the Chinese National Natural Science Foundation (U1131003), the National 973 project (2011CB1007) and the National Transgenic Research Projects (2009ZX08009-023B; 2011ZX08001-002).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by P. Hayes.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hua, L., Wu, J., Chen, C. et al. The isolation of Pi1, an allele at the Pik locus which confers broad spectrum resistance to rice blast. Theor Appl Genet 125, 1047–1055 (2012). https://doi.org/10.1007/s00122-012-1894-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-012-1894-7