Abstract
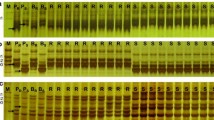
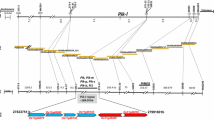
Rice blast, caused by the fungal pathogen Magnaporthe oryzae, is one of the most devastating crop diseases worldwide. The avirulence gene corresponding to rice blast resistance gene Pi7 in field isolate CHL346 was inherited as a single gene, designated AvrPi7, in a segregating population consisting of 189 ascospore progenies derived from a cross between field isolates CHL346 and CHL42. In order to determine the chromosomal location of the AvrPi7 locus, a total of 121 simple sequence repeat (SSR) markers were developed based on the whole-genome sequence of reference isolate 70-15 of M. oryzae. Linkage analysis of the locus with these SSR markers showed that eight SSR markers on chromosome 1 were linked to the locus, among which the closest flanking markers MS1-9 and MS1-15 were 3.2 and 16.4 cM from the locus, respectively. For fine mapping, additional PCR-based makers including eight SSR markers and three candidate avirulence gene (CAG) markers were developed in the region flanking both markers. The AvrPi7 locus was genetically delimited within a 1.6-cM region flanked by markers MS1-21 and MS1-22, and co-segregated with the marker CAG2. To construct a physical map of the AvrPi7 locus, molecular markers linked to the Avr gene were mapped on the supercontigs of the reference isolate 70-15 through bioinformation analysis (BIA). Consequently, the AvrPi7 locus was delimited to a 75-kb interval flanked by markers MS1-21 and MS1-22 based on the reference sequence. Merodiploids observed in this study are also discussed.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ou S H. Rice Diseases. 2nd ed. Surrey, England: Commonwealth Mycological Institute, 1985. 109–201
Dean R A, Talbot N J, Ebbole D J, et al. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature, 2005, 434: 980–986
Luo C X, Yin L F, Koyanagi S, et al. Genetic mapping and chromosomal assignment of Magnaporthe oryzae avrirulence genes AvrPik, AvrPiz, and AvrPiz-t controlling cultivar specificity on rice. Phypathology, 2005, 95: 640–647
Flor H. Current status of the gene-for-gene concept. Annu Rev Phytopathol, 1971, 9: 275–296
Herbert T T. The perfect stage of Pyricularia grisea. Phytopathology, 1971, 61: 83–87
Notteghem J L, Tharreau D, Silue D, et al. Present knowledge of rice resistance genetics and strategies for Magnaporthe grisea pathogenicity and avirulence gene analysis. In: Zeigler R S, Leong S A, Teng P S, eds. Rice Blast Disease. Walliingford: CAB International, 1994. 155–165
Leung H, Williams P H. Genetic analyses of electrophoretic enzyme variants, mating type, and hermaphroditism in Pyricularia oryzae Cavara. Can J Genet Cytol, 1985, 27: 697–704
Leung H, Borromeo E S, Bernardo M A, et al. Genetic analysis of virulence in the rice blast fungus Magnaporthe grisea. Phytopathology, 1988, 78: 1227–1233
Farman M L, Leong S A. Chromosome walking to AVR1-CO39 avirulence gene of Magnaporthe grisea discrepancy between the physical and genetic maps. Genetics, 1998, 150: 1049–1058
Smith J R, Leong S A. Mapping of Magnaporthe grisea locus affecting rice (Oryza sativa) cultivar specifity. Theor Appl Genet, 1994, 88: 901–908
Lau G W, Chao C T, Ellingboe A H. Interaction of genes controlling avirulence/virulence of Magnaporthe grisea on rice cultivar Katy. Phytopathology, 1993, 83: 375–382
Silué D, Tharreau D, Notteghem J L. Identification of Magnaporthe grisea avirulence genes to seven rice cultivars. Phytopathology, 1992, 82: 1462–1467
Dioh W, Tharreau D, Notteghem J L, et al. Mapping of avirulence genes in the rive blast fungus, Magnaporthe grisea, with RFLP and RAPD markers. Mol Plant Microbe Interact, 2000, 13: 217–227
Wang B H, Lu G D, Lin W M, et al. Genetic analysis and molecular marker of Avr-Pi1, Avr-Pi2 and Avr-Pi4a of Magnaporthe grisea. Acta Genet Sin (in Chinese), 2002, 29: 820–826
Li C Y, Luo C X, Li J B, et al. Mapping avirulence gene in the rice blast fungus Magnaporthe grisea. Sci Agr Sin (in Chinese), 2000, 33: 49–53
Lin F, Li J B, Li C Y, et al. Identification of two avirulence genes in the rice blast fungus Magnaporthe grisea using random amplified polymorphic DNA (RAPD) markers. Sci Agr Sin (in Chinese), 2002, 35: 1079–1084
Hayashi N, Li C Y, Li J R, Naito H. In vitro production on rice plants of perithecia of Magnaporthe grisea from Yunnan, China. Mycol Res, 1997, 101: 1308–1310
Valent B, Chumley F G. Avirulence genes and mechanisms of genetic instability in the rice blast fungus. In: Zeigler R, Leong S A, Teng P S, eds. Rice Blast Disease. Wallingford: CAB International, 1994. 111–134
Valent B, Chumley F G. Molecular genetic analysis of the rice blast fungus, Magnaporthe grisea. Annu Rev Phytopathol, 1991, 29: 443–467
Yasuda N, Fujita Y, Noguchi M. Identification of avirulence genes in the rice blast fungus corresponding to three resistance genes in Japanese differentials. J Gen Plant Pathol, 2004, 70: 202–206
Yasuda N, Noguchi M T, Fijita Y. Identification of an avirulence gene in the fungus Magnaporthe grisea corresponding to a resistance gene at the Pik locus. Phytopathology, 2005, 95: 768–772
Liu J F, Dong N, Hou Z J, et al. Identification of a RAPD markers linked with the locus in rice blast fungus conferring avirulence to rice cultivar Tsuyuke. Acta Phytopathol Sin (in Chinese), 2001, 31: 10–15
Orbach M J, Farrall L, Sweigard J A, et al. A telomeric avirulence gene determines efficacy for the rice blast resistance gene Pi-ta. Plant Cell, 2000, 12: 2019–2032
Böhnert H U, Fudal I, Dioh W, et al. A putative polyketide synthase/peptide synthetae from Magnaporthe grisea signals pathogen attack to resistance rice. Plant cell, 2004, 16: 2499–2513
Lauge R, De Wit P J G M. Fungal avirulence genes: Structure and possible functions. Fungal Genet Biol, 1998, 24: 285–297
Ma J H, Wang L, Feng S J, et al. Identification and fine mapping of AvrPi15, a novel avirulence gene of Magnaporthe grisea. Theor Appl Genet, 2006, 113: 875–883
Pan Q H, Hu Z D, Tanisaka T, et al. Fine mapping of the blast resistance gene Pi15, linked to Pii, on rice chromosome 9. Acta Bot Sin, 2003, 45: 871–877
Tsunematsu H, Yanoria M J T, Ebron L A, et al. Development of monogenic lines of rice for blast resistance. Breed Sci, 2000, 50: 229–234
Pan Q H, Wang L, Ikehashi H, et al. Identification of a new blast resistance gene in the indica rice cultivar Kasalath using Japanese differential cultivars and isozyme markers. Phytopathology, 1996, 86: 1071–1075
Farman M L, Eto Y, Nakao T, et al. Analysis of the structure of the AVR1-CO39 avirulence locus in virulent rice-infecting isolates of Magnaporthe grisea. Mol Plant Microbe Interact, 2002, 15: 6–16
Chen S, Wang L, Que Z Q, et al. Genetic and physical mapping of Pi37(t), a new gene conferring resistance to rice blast in the famous cultivar St.No.1. Theor Appl Genet, 2005, 111: 1563–1570
Liu X Q, Wang L, Chen S, et al. Genetic and physical mapping of Pi36(t), a new rice blast resistance gene located on rice chromosome 8. Mol Gen Genomics, 2005, 274: 394–401
Chen J W, Pang X F, Wang L, et al. Genetic analysis and fine mapping of a rice brown planthopper (Nilaparvata lugens Stål) resistance gene bph19(t). Mol Genet Genomics, 2006, 275: 321–329
Michelmore R W, Paran I, Kesseli R V. Identification of markers linked to disease resistance genes by bulked segregant analysis: A rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA, 1991, 88: 9828–9832
Tanksley S D, Ganal M W, Martin G B. chromosome landing: A paradigm for map-based gene cloning in plants with large genomes. Trends Genet, 1995, 11: 63–68
Joosten M H A J, Cozijnsen T J, De Wit P J G M. Host resistance to a fungal tomato pathogen lost by a single base-pair change in an avirulence gene. Nature, 1994, 367: 384–386
Sweigard J A, Carroll A M, Kang S, et al. Identification, cloning, and characterization of PWL2, a gene for host species specificity in the rice blast fungus. Plant Cell, 1995, 7: 1221–1233
Shan W X, Cao M, Leung D, et al. The Avr1b locus of Phytophthora sojae encodes an elicitor and a regulator required for avirulence on soybean plants carrying resistance gene Rps1b. Mol Plant Micobe Interact, 2004, 17: 394–403
Gout L, Fudal I, Kuhn M L, et al. Lost in the middle of nowhere: The AvrLm1 avirulence gene of the Dothideomycete Leptosphaeria maculans. Mol Microbiol, 2006, 60: 67–80
Schuler G D. Sequence mapping by electronic PCR. Genome Res, 1997, 7: 541–550
Davis C R, Kempainen R R, Srodfs M S, et al. Correlation of the physical and genetic maps of the centromere region of the right arm of linkage group III of Neurospora crassa. Genetics, 1994, 136, 1297–1306
Chauhan R S, Farman M L, Zhang H B, et al. Genetic and physical mapping of a rice blast resistance locus, Pi-CO39(t), that corresponds to the avirulence gene AVR1-CO39 of Magnaporthe grisea. Mol Gen Genomics, 2002, 26: 603–612
Clarke L, Carbon J. Isolation of the centromere-linked CDC10 gene by complementation in yeast. Proc Natl Acad Sci, USA, 1980, 77: 2173–2177
Wu W H, Wang L, Pan Q H. Comparison of genetic structures of two rice blast fungus populations originated from Jiangsu and Yunnan Provinces, China. Chin J Tropical Crops (in Chinese), 2006, 17: 95–100
Sun X, Yang Z, Wang S, et al. Identification of a 47-kb DNA fragment containing Xa4, a locus for bacterial blight resistance in rice. Theor Appl Genet, 2003, 106: 683–687
Nitta N, Farman M L, Leong S A. Genome organization of Magnaporthe grisea: Integration of genetic maps, clustering of transposable elements and identification of genome duplications and rearrangements. Theor Appl Genet, 1997, 95: 20–32
Hartl D L, Jones E W. Genetics: Analysis of Genes and Genomes. 5th ed. Boston: Jones and Bartlett Publishers, 2001. 393–443
Covert S F. Supernumerary chromosomes in filamentous fungi. Curr Genet, 1998, 33: 311–319
Orbach M J, Chumley F G, Valent B. Electrophoretic karyotypes of Magnaporthe grisea pathogens of diverse grasses. Mol Plant Microbe Interact, 1996, 9: 261–271
Talbot N J, Salch Y P, Ma M, et al. Karyotypic variation within clonal lineages of the rice blast fungus, Magnaporth grisea. Appl Environ Microbiol, 1993, 59: 585–593
Chuma I, Tosa Y, Taga M, et al. Meiotic behavior of a supernumerary chromosome in Magnaporthe oryzae. Curr Genetic, 2003, 43: 191–198
Enkerli J, Reed H, Briley A, et al. Physical map of a conditionally dispensable chromosome in Nectria haematococca mating population VI and location of chromosome breakpoints. Genetics, 2000, 155: 1083–1094
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the National Basic Research Program of China (Grant No. 2006CB100206), the Program for Changjiang Scholars and Innovation Research Team in University (Grant No. IRT0448), and the Natural Science Foundation of Guangdong Province (Grant Nos. 021006 and 039254)
About this article
Cite this article
Feng, S., Wang, L., Ma, J. et al. Genetic and physical mapping of AvrPi7, a novel avirulence gene of Magnaporthe oryzae using physical position-ready markers. CHINESE SCI BULL 52, 903–911 (2007). https://doi.org/10.1007/s11434-007-0125-0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s11434-007-0125-0