Abstract
Production of bulk chemicals from renewable biomass has been proved to be sustainable and environmentally friendly. Escherichia coli is the most commonly used host strain for constructing cell factories for production of bulk chemicals since it has clear physiological and genetic characteristics, grows fast in minimal salts medium, uses a wide range of substrates, and can be genetically modified easily. With the development of metabolic engineering, systems biology, and synthetic biology, a technology platform has been established to construct E. coli cell factories for bulk chemicals production. In this chapter, we will introduce this technology platform, as well as E. coli cell factories successfully constructed for production of organic acids and alcohols.
Graphical Abstract

Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Bulk chemicals are produced at high volume and relatively low cost, which can be either directly used or used as platform chemicals for production of derivatives in the chemical industry [1, 2]. The annual production volumes of bulk chemicals are usually in the range of 1–100 million tons, and their selling prices are less than 2,000 dollars per ton [2]. Traditionally, bulk chemicals are predominantly produced from non-renewable fossil resources via the petrochemical routes [1, 3]. Because of the decrease of global storage, fluctuations of petroleum prices, trade imbalances, and political considerations, the cost for production of bulk chemicals through petrochemical process is increasing [1, 4–10]. In addition, petrochemical processes always consume a lot of energy and cause serious environmental pollutions [9–15].
Production of bulk chemicals from biomass resources by microbial cell factories is an alternative route, which is renewable and environment friendly compared to petrochemical route [11, 16–18]. However, only a few bulk chemicals can be produced by microbial cell factories. On the other hand, although some bulk chemicals can be synthesized by engineered microorganism, the producing capabilities, including titer, yield, productivity, and physiological characteristics, are not good enough to compete with petrochemical routes. Thus, it is important to expand the product range, as well as to improve producing capabilities of cell factories to decrease producing cost for commercialization.
The rapid development of metabolic engineering, systems biology, and synthetic biology has facilitated construction of microbial cell factories for producing bulk chemicals [1, 16, 19, 20]. Escherichia coli is the most commonly used host strain for cell factories construction since it has clear physiological and genetic characteristics and can be genetically modified easily [3, 21]. E. coli also grows fast in minimal salt medium and can utilize both hexose and pentose in the biomass. Currently, bio-based bulk chemicals produced by E. coli include not only those formed by E. coli native metabolic pathways such as lactate [22–25] and succinate [26–29], but also those produced by heterologous pathways or totally new synthetic pathways, such as 3-hydroxypropionic acid [30–33], 1,3-propanediol [2], isobutanol [13, 34–36], butanol [13], 1,4-butanediol [37], and alkanes [38, 39]. In this review, we will summarize the technology platforms for construction and optimization of E. coli cell factories, as well as representative cases of constructing E. coli cell factories for production of organic acids and alcohols.
2 Technology Platform for Construction of E. coli Cell Factories
With the developments of systems biology and synthetic biology, a technology platform has been established for the construction of E. coli cell factories for bulk chemicals production. This platform includes (1) design of the optimal synthetic pathway, (2) construction of the synthetic pathway, (3) optimization of the synthetic pathway, (4) optimization of the producing capability at the whole cell level, and (5) characterization of the genetic mechanisms (Fig. 1). An initial cell factory can be obtained after the first four steps. The genetic mechanisms identified for high production can be used to further improve the producing capabilities to construct the next-generation cell factories.
2.1 Design of the Optimal Synthetic Pathway
E. coli produces mixed acids during the glucose metabolism. In order to produce the target compound, other competitive pathways need to be inactivated. Enough energy supply is necessary to maintain cell growth and metabolism, while appropriate reducing equivalent supply is required to keep redox balanced, especially under anaerobic conditions. On the other hand, most synthetic pathways that convert the designated substrate to target product do not exist in E. coli, and sometimes are even not present in nature. Thus, designing the novel synthetic pathway is very important for construction of cell factories. With the development of bioinformatics tools, several genome-scale metabolic network models have been reconstructed for E. coli [40–43]. These models can help design the optimal synthetic pathway, discover new engineering target to improve production of target compound, and predict the cellular phenotypes [41]. There have been several reviews describing the developments of metabolic network models and their applications in constructing E. coli cell factories [44–49].
With the help of metabolic network models, the optimal synthetic pathway can be designed based on modification of the native metabolic pathways (such as succinate or d-lactate), integration of exogenous reactions by software predication (such as 1,4-Butanediol [12]), genome mining (such as alkanes [20]), and modification of the natural pathway to catalyze unnatural reactions (such as higher-chain alcohols [50, 51]). Different tools used for predication of novel biosynthetic pathways have been reported [52–55]. These tools can not only propose candidate pathways but also supply ranking of the pathways based on different factors employed in the process (such as thermodynamics [56]) to reduce the numbers of pathways to be a reasonable scope for experimental validation [16].
2.2 Construction of the Synthetic Pathway
Gene resources and DNA assembly method are two key factors for construction of synthetic pathway. It is desirable to obtain genes in an easy, quick, and inexpensive way, and to assemble different genes into a complete synthetic pathway with efficient and standard methods.
PCR has been commonly used to obtain target genes. However, original cells having the target genes need to be collected first, which is time-consuming. In addition, many heterologous genes cannot be expressed and translated efficiently in the host strain. With the rapid development of high-throughput chemical synthesis of DNA, achieving gene resources has become independent on original cells. It is also possible to optimize the transcriptional efficiency of target genes by codon optimization. The Church’s group developed a microchip-based technology for synthesis of genes and reduced the error rate by ninefold [57]. Using this technology, all 21 genes encoding the proteins of E. coli 30S ribosomal subunit were synthesized and the translation efficiency in vitro was optimized through alteration of codon bias. An on-chip gene synthesis technology, including ink-jet printing, isothermal oligonucleotide amplification, and parallel gene assembly, was integrated on a single microchip [58]. By using a mismatch-specific endonuclease for error correction, the error rate was reduced to about 0.19 errors per kb.
Standardized DNA assembly is another limiting technology for construction of the synthetic pathway. Several BioBrick assembly standards [59] have been developed. The DNA unit is flanked by standardized sequences, and the assembly can be achieved by a simple and standardized restriction/ligation method [60, 61]. However, the BioBrick approach still has several disadvantages, such as the remaining of 6-bp scars resulting from each binary BioBrick assembly and the limitation of rearrangement of every intermediate part [62]. Several new technologies, such as sequence- and ligation-independent cloning (SLIC), Gibson isothermal assembly, and circular polymerase extension cloning (CPEC), have been designed, which can supply standardized, scarless, sequence-independent, and multi-part DNA assembly [62–65]. All these technologies are dependent on the 5′ homology sequence flanked at the two ends of DNA part. The biological characteristics and mechanisms for these methods have been reviewed [61].
All methods mentioned above are carried out in vitro. The Zhao group at the University of Illinois at Urbana-Champaign developed a DNA assembly method in vivo [66, 67]. DNA parts with homologous sequences were transformed into Saccharomyces cerevisiae and assembled based on the high homologous recombination efficiency of the yeast. The assembled DNA devices were then extracted and transformed into E. coli for evaluation or expression. This method is efficient and independent on enzymes in vitro.
DNA integration into chromosome has also been developed to construct a genetically stable strain for industrial production. Homologous recombination based on the λ Red recombinase has been developed [68–70]. The one-step homologous recombination method uses an antibiotic marker for selection, which is flanked by two FLP recognition target (FRT) fragments. The antibiotic marker can be removed by the FLP recombinase, facilitating multiple rounds of genetic engineering. This method can integrate or delete genes quickly but leave a 68-bp FRT scar on chromosome each time, and repeated use of this system has the potential to result in large unintended chromosomal deletions [68, 69, 71]. To facilitate sequential gene manipulations, a two-stage recombination strategy was developed, which was based on the sensitivity of E. coli to sucrose when levansucrase (sacB) is expressed in cell. In the first recombination, the target chromosomal genes are replaced by a DNA cassette containing an antibiotic marker and the sacB gene. In the second recombination step, the antibiotic marker and the sacB gene are removed by selection for the resistance to sucrose [69, 72–74].
2.3 Optimization of the Synthetic Pathway
The native and new constructed synthetic pathways are always not efficient. Some enzymes might have low activities and become the rate-limiting steps for the whole pathway. Some toxic intermediates might accumulate in the cell, thus leading to decreased cell growth and flux imbalances. In order to improve the producing capability, several strategies have been developed to optimize the synthetic pathway in three levels.
2.3.1 Optimization of the Synthetic Pathway in a Single Gene Level
Gene overexpression was commonly used to increase the activities of rate-limiting enzymes among the synthetic pathway for the purpose of modulating metabolic fluxes. However, this simple overexpression strategy rarely reached the optimal transcript level and appeared to be unsuccessful in most cases to improve producing capability [75]. Promoter library has been developed as a solution to this all-or-nothing expression strategy, which could provide variable promoters with a wide range of strength for fine-tuning of gene expression [76–80]. Two methods have been developed for creating promoter library. One type was obtained by keeping the conserved −35 and −10 sequences intact and randomizing the surrounding nucleotides [78, 79]. The other type was obtained by mutating the sequence of an existing promoter using error-prone PCR [76, 77].
However, plasmid-based gene expression has several disadvantages for the engineering of genetically stable strains [69]. Plasmid maintenance is a metabolic burden on the host cell, especially for high-copy number plasmids [81], and only few natural unit-copy plasmids have the desirable genetic stability [82]. In addition, only low-copy number plasmids have replication that is timed with the cell cycle, and thus, it is difficult to maintain a consistent copy number in all cells [82]. It is thus desirable to integrate the target genes into chromosome followed by fine-tuning of their expression. With the aid of Red recombination technology [68], promoter libraries were recently constructed directly in the chromosome [83–85], which might be more suitable for modulation of gene expression directly in the chromosome.
Different promoters with varied strengths can be used to control gene transcription precisely to obtain a specific cellular phenotype. One example is the divergence of biomass yield by modulation of phosphoenolpyruvate carboxylase (ppc) gene transcription level. When the wild-type promoter of ppc was replaced by promoters with varied strengths, there was a positive correlation between the ppc transcription level and the biomass yield when the ppc transcription level was within a certain range. Excessive ppc transcription level led to decreased biomass yield. The promoter library facilitated identifying the optimum transcription level of ppc for biomass yield. Another example is the use of promoter library to investigate the relationship between succinate production and PPC or phosphoenolpyruvate carboxykinase (PCK) activity. There was a positive correlation between PCK activity and succinate production. In contrast, there was a positive correlation between PPC activity and succinate production only when PPC activity was within a certain range. Excessive PPC activity decreased the rates of both cell growth and succinate formation [29]. In contrast, plasmid overexpression of ppc gene always led to increased succinate production [86], which would mislead our understanding of the relationship between succinate production and PPC activity.
2.3.2 Optimization of the Synthetic Pathway in Multiple Gene Levels
The efficiencies of most synthetic pathways are always not limited by a single rate-limiting reaction [87]. A more broadly accepted opinion is to realize the coordinated expression of multiple genes involved in the synthetic pathway to increase the overall metabolic flux.
A technology for tuning the expression of multiple genes by employing post-transcriptional mechanisms was developed by the Keasling group at the University of California, Berkeley [88]. Libraries of tunable intergenic regions (TIGRs) consisting of several control elements composed by mRNA secondary structures, RNase cleavage sites, and the RBS sequence were constructed and used to differentially change the processes of transcription termination, mRNA stability, and translation initiation [87, 88]. When using this strategy to balance expression of three genes in an operon encoding a heterologous mevalonate biosynthetic pathway, a sevenfold increase of mevalonate production was achieved. Another technology for fine-tuning pathway flux was developed by the same group in 2009 [89]. Synthetic protein scaffolds was built to spatially recruit metabolic enzymes in a specific manner to increase the valid concentration of metabolic intermediates and avoid their accumulation to toxic level. Also, the production levels can be optimized by balancing relative quantities of individual enzymes via changing the number of interaction-domain repeats that locate different enzymes to the synthetic complex. Using this technology, a 77-fold improvement was achieved for the mevalonate biosynthesis.
The Church group at Harvard Medical School developed a powerful tool termed multiple automated genome engineering (MAGE), which can modify many genes in the E. coli genome in parallel. This technique was used to optimize the 1-deoxy-d-xylulose-5-phosphate (DXP) synthetic pathway in E. coli for improving lycopene production. Twenty-four genes in the DXP pathway were modified simultaneously, and over 4.3 billion variants were created per day. E. coli variants with more than fivefold increase in lycopene production were isolated within 3 days [90, 91].
Instead of repeating multiple rounds of gene knockout, synthetic regulatory small RNAs (sRNAs) were designed to finely control gene expression in E. coli by the Lee group at Korea Advanced Institute of Science and Technology. Customized synthetic sRNAs were consisted by a scaffold and a target-binding sequence. With utilization of the plasmid-based synthetic sRNA system, one can study the effects of multiple knockdowns on the cell’s producing capability in a high-throughput way and simultaneously screen target genes in different E. coli strains [92].
2.3.3 Optimization of the Synthetic Pathway Using the Sensor–Regulator System
To precisely control and regulate the heterologous pathway expression due to the change of environment or intracellular conditions, an efficient strategy is to use the sensor–regulator system which can respond to a particular intermediate and stimulate the desired cellular response to enable the cell to efficiently use the cellular resources and improve the producing capability while decrease the accumulation of toxic metabolite [93]. Malonyl-CoA is the rate-limiting precursor involved in the synthetic pathway of several value-added pharmaceuticals and biofuels. By incorporating the trans-regulatory protein FapR and the cis-regulatory element fapO of Bacillus subtilis, a hybrid promoter–regulator system was constructed and could respond to a wide range of intracellular malonyl-CoA concentrations in E. coli [93]. In another study, the Liao’s group designed and engineered a regulatory circuit by recruiting and altering the Ntr regulon which is a global regulatory system to control the pathway expression for lycopene synthesis in E. coli. The artificially engineered regulon controlled the gene expression in the lycopene synthetic pathway by sensing the concentration of acetyl phosphate which is the glycolytic pathway hallmark metabolite [94]. Recently, a dynamic sensor–regulator system (DSRS) was developed to dynamically regulate the gene expression in biodiesel biosynthetic pathway by responding to the key intermediate fatty acyl-CoA in E. coli. Using this strategy, the fatty acid ethyl ester production was increased threefold compared to that of using constitutive promoters in E. coli [95].
2.4 Optimization of Producing Capability at the Whole Cell Level
After optimization of the synthetic pathway, the producing capability of the engineered cell, such as titer, yield, productivity, and physiological characteristics, might still be not good enough for industrial application. The desired cell phenotypes may be affected by factors which are not directly related to the synthetic pathway [96]. In order to obtain an efficient cell factory, the producing capability needs to be optimized further at the whole cell level.
Metabolic evolution was developed by the Ingram group at the University of Florida and has been demonstrated to be an excellent strategy for strain improvement [69]. Synthesis of target product is designed to be the only fermentation pathway to oxidize NADH under anaerobic condition. The cell growth of the engineered cell is coupled with the synthesis of target product, since this is the only way to regenerate NAD+ for continuous glycolysis to provide ATP for cell growth. This technology has been used widely to improve the production of several bulk chemicals by E. coli cell factories, such as d-lactate [97–99], succinate [100–102], and ethanol [103]. In addition, this technology can also be used to improve cell’s physiological characteristics, especially tolerance to toxic metabolites or high concentration of target products [104, 105].
Global transcription machinery engineering, or termed gTME, was developed by the Stephanopoulos group at Massachusetts Institute of Technology and has been proved to be a powerful strategy for optimization of a desired phenotype at system level [106–109]. For optimization of a desired phenotype at systems level, gTME has been used to improve ethanol tolerance, lycopene production, and simultaneous tolerance to sodium dodecyl sulfate (SDS) and ethanol. To realize these purposes, one of the components of global cellular transcription machinery (specifically rpoD encoding the σ70) in E. coli was engineered to globally perturb the transcriptome to help unlock complex phenotypes [109].
Aside from technologies mentioned above, some other tools, such as genome shuffling [110–113] and trackable multiplex recombineering (TRMR) [114], have been designed and used to optimize a target pathway at system level. Great improvement for the chemical production properties has been achieved based on these strategies. It should be noted that high-throughput screening methods are required for the efficient selection.
2.5 Characterization of the Genetic Mechanism
Although metabolic evolution or other global perturbation methods are efficient for improving the producing capability of the engineered cell, the genetic backgrounds of strains obtained by these strategies often remain unclear. Characterization of the genetic mechanisms relative to the improved producing capability is very important. The fast accumulation of omics data, including genomics, transcriptomics, proteomics, metabolomics, and fluxomics, has provided foundation for the understanding of the genetic mechanisms in depth [115–119], which is crucial for further round of engineering to obtain the next-generation cell factories.
Up to now, many E. coli cell factories with abilities of producing different bulk chemicals have been constructed and some have been applied in industrial scale. The bio-based bulk chemicals produced by E. coli cell factories mainly include organic acids and alcohols, which will be described in detail in the following chapters.
3 Organic Acids
Organic acids have received attractive attentions for their increasing utilization in food industry and great potential as platform chemicals for the manufacture of biodegradable polymers [120, 121]. As an alternative of petroleum-based production, microbial production of organic acids from renewable biomass has been accepted as a feasible process. E. coli has been widely engineered to produce organic acids, such as acetate, lactate, pyruvate, 3-hydroxypropionate, succinate, malate, fumarate, glucaric acid, and muconic acid (Table 1).
3.1 d-lactate
As a specialty chemical, d-lactate is widely applied in the food and pharmaceutical industry. A potential huge market for d-lactate is to be combined with l-lactate to produce polylactic acid (PLA), an increasingly attractive biodegradable plastic. The commercial success of PLA will greatly depend on the production cost of the monomers [69]. Wild-type E. coli can produce d-lactate in its mixed acid fermentation process (Fig. 2). However, the productivity is low and several undesirable metabolites are produced at the same time. To realize the production of d-lactate in an efficient way, it is necessary to reengineer the metabolic network of E. coli.
d-lactate-producing strains were engineered from E. coli W3110 by the Ingram’s group by inactivating the competitive fermentation pathways, including fumarate reductase (frdABCD), alcohol/aldehyde dehydrogenase (adhE), and pyruvate formate lyase (pflB). A further deletion of the acetate kinase gene (ackA) increased the cell mass and lactate productivity. D-lactate production yield of these strains approached the theoretical maximum yield (2 mol/mol glucose) using mineral salts medium [122]. For expanding the substrate range, a cluster of sucrose utilization genes which were characterized and cloned from E. coli KO11 were introduced, resulting in production of over 500 mM D-lactate from sucrose [123]. However, these biocatalysts were unable to ferment glucose or sucrose with concentration of up to 10 % completely. Inspired by the construction of ethanol producing strain, a derivative of E. coli B was selected as the starting strain for d-lactate production. Based on the growth-based selection (Fig. 3), metabolic evolution was carried out to improve strain performance. The resulting strain SZ194 produced 1.22 M d-lactate with a yield of 1.9 mol/mol using mineral salts medium. The production capability was comparable with lactic acid bacteria [98].
In another study, E. coli strain B0013 was engineered for d-lactate production by deletion of acetate kinase and phosphotransacetylase (ackA-pta), phosphoenolpyruvate synthase (pps), pflB, FAD-binding d-lactate dehydrogenase (dld), pyruvate oxidase (poxB), and adhE and frd genes. The resulting strain, B0013-070, produced 125 g/L d-lactate [25]. Replacing the ldhA promoter with the λ p R and p L promoter in strain B0013-070 led to a thermocontrollable strain B0013-070B in which the LDH activity was twofold higher than the parent strain B0013-070 at 42 °C. When the culture temperature reached to 33 °C, the genetic switch would be turned off and strain B0013-070B produced 10 % more biomass under aerobic conditions than stain B0013-070 with trace d-lactate produced. This modification reduced the growth inhibition which was caused by oxygen insufficiency in large-scale fermentation process [23].
3.2 3-Hydroxypropionate
3-Hydroxypropionate (3HP), a non-chiral carboxylic acid, has received much attention for its potential applications to produce biodegradable polymer by itself or with other compounds [124, 125]. Additionally, 3HP was an important C3 platform chemical and can be used for the production of various commercially valuable chemicals, such as 1,3-propanediol, acrylic acid, and malonic acid [126]. 3HP has been identified as a metabolic intermediate naturally present in several microorganisms [127–134]. More than a dozen of pathways for 3HP biosynthesis have been proposed based on the natural metabolic pathways or in silico design [135–137]. However, a little fraction of the pathways have been evaluated. The Park’s group developed a recombinant E. coli strain producing 3HP from glucose involved malonyl-CoA as an intermediate. In this strain, a mcr gene encoding the NADPH-dependent malonyl-CoA reductase (MCR) of Chloroflexus aurantiacus DSM 635 was introduced into E. coli. The recombinant strain produced 0.064 g/L 3HP when cultivated aerobically for 24 h using glucose as the sole carbon source. To improve the 3HP production, the gene cluster accADBCb encoding the acetyl-CoA carboxylase and biotinilase of E. coli K-12 were overexpressed and this resulted in a twofold improvement in 3HP production. Further genetic modification was carried out to express the gene pntAB encoding the membrane-bound transhydrogenase to convert the NADH to NADPH which increased 3HP titer to 0.193 g/L [33].
Compared to producing 3HP from glucose, more studies have been focused on the production of 3HP from glycerol. By heterologous overexpression of the glycerol dehydratase (DhaB) from Klebsiella pneumobiae DSM 2026 and aldehyde dehydrogenase (AldH) from E. coli K-12 MG1655 in E. coli BL21 (DE3), a recombinant E. coli strain SH254 was obtained. When fermented aerobically in M9 minimal medium supplemented with glycerol as substrate in shake flask, this strain produced 0.58 g/L 3HP with a yield of 0.48 mol/mol glycerol [138]. Further optimization of the fermentation parameters, such as pH, IPTG concentration, aeration rate, and substrate concentration, led to production of 31 g/L 3HP in 72 h when a fed-batch fermentation process was used [139]. Though the titer of 3HP was improved by optimization of the fermentation parameters, several problems, including the imbalance between DhaB and AldH and instability of DhaB, were still not solved. To overcome these limitations, DhaB and AldH were overexpressed in two compatible plasmids with inducible expression systems and the glycerol dehydratase reactivase (GDR) was expressed at the same time. Then, by using α-ketoglutaric semialdehyde dehydrogenase (KGSADH) from A. brasilense to replace the AldH, a recombinant E. coli strain SH-BGK1 was constructed which produced 38.7 g/L 3HP aerobically using a fed-batch process [140]. Modulation of glycerol metabolism further increased 3HP titer to 57.3 g/L with a yield of 0.88 g/g glycerol [30]. In another work, based on in silico simulation, two genes tpiA (encoding triose phosphate isomerase) and zwf (encoding glucose 6-phosphate dehydrogenase) involved in the central metabolism and yqhD gene (encoding NADPH-dependent aldehyde reductase) involved in the biosynthetic pathway of the major by-product 1,3-propanediol were identified as the engineering targets to improve 3HP production from glycerol. Deletion of these three genes led to 7.4-fold increase of 3HP titer compared to the parent strain [141].
3.3 Succinate
Succinate can be produced by native E. coli as a minor product [56]. In order to produce succinate as the sole product, other competitive pathways need to be eliminated. Strain NZN111 is an engineered E. coli, which has lactate dehydrogenase (ldhA) and pflB inactivated [142–144]. This strain produces undetectable lactate and formate under anaerobic conditions [142, 143]. However, inactivation of these NADH-consuming pathways could also cause redox imbalance within cells, thus leading to decreased cell growth and glucose utilization [143]. Strain NZN111 consumed only 1.8 g/L glucose and produced 1.8 g/L succinate under anaerobic conditions for 44 h [145]. A mutant strain of NZN111 (strain AFP111) was isolated, which recovered cell growth and had increased succinate production under anaerobic conditions. A spontaneous mutation in ptsG gene [146–148] of NZN111, which encodes EIIBGlc subunit of phosphoenolpyruvate (PEP): carbohydrate phosphotransferase systems (PTS), was identified to be responsible for the increased cell growth and succinate production [143]. This ptsG mutation could enhance PEP precursor supply for succinate synthesis, as well as alleviating glucose repression to the expression of several genes which are crucial to the fermentation. Several genetic manipulations were performed to further improve succinate production of strain AFP111. For instance, overexpression of pyruvate carboxylase (PYC) gene of Rhizobium etli in AFP111 increased succinate titer and yield to 99.2 g/L (841 mM) and 1.1 g/g (1.68 mol/mol), respectively, under dual-phase conditions [149].
Under anaerobic conditions, 1 molecule glucose produces 2 molecules NADH through glycolysis, while the production of 1 molecule succinate requires two NADH through the reductive TCA pathway. The maximal succinate yield is only 1 mol/mol glucose, which is much less than the theoretical maximum yield (1.71 mol/mol) [150–153]. In comparison, NADH requirement for succinate synthesis decreases when glyoxylate shunt pathway is utilized. It was calculated that 1.25 molecules NADH was required to synthesize 1 molecule succinate [150]. Glyoxylate shunt pathway is composed of isocitrate lyase (encoded by aceA) and malate synthase (encoded by aceB) [154]. In the presence of glucose, aceBAK operon is strictly repressed by IclR regulator [155–157], and iclR deletion was proven to efficiently activate glyoxylate shunt pathway [155]. Sanchez et al. found that deletion of iclR in strain SBS550MG (∆adhE, ∆pflB, ∆ack-pta) harboring pyruvate carboxylase from Lactococcus lactis increased its succinate yield to 1.61 mol/mol [150]. Obtaining NADH through formate dehydrogenase is another strategy to improve succinate yield [158]. Blazer et al. reported that overexpression of heterologous NAD+-dependent formate dehydrogenase from Candida boidinii increased succinate yield to 1.74 mol/mol glucose [158]. External formate supplementation further resulted in 6 % increase in succinate yields [158].
Although plenty of successes were obtained in metabolic engineering of E. coli to improve succinate production as mentioned above [143, 145, 149, 150, 158, 159], there were several problems that remained to be improved. Many research groups have employed dual-phase fermentation for succinate production, i.e., aerobic growth phase followed by anaerobic fermentation phase [149, 150]. Part of the carbon source is converted to cell mass and carbon dioxide during the aerobic phase, which leads to decreased succinate yield. Supply of dissolved oxygen also increases the energy costs during industrial production. In addition, rich medium is frequently used for fermentation [149, 150], which would increase material costs and downstream purification costs. It is very important to use mineral salts medium and one-step anaerobic process for succinate production. By combining metabolic engineering to inactivate competitive fermentation pathways and metabolic evolution to improve cell growth and succinate production, a high-succinate-producing strain KJ073 was obtained by the Ingram’s group which produced 668 mM succinate with a yield of 1.2 mol/mol using mineral salts medium and one-step anaerobic process [160]. The genetic mechanisms for efficient succinate production of strain KJ073 were further identified [101]. PCK activity was increased due to a G-to-A transition at −64 position relative to the ATG start codon of pck, which increased the energy supply for cell growth and succinate production under anaerobic condition [101]. In addition, a frame-shift mutation in ptsI gene, which encodes the EI component of PTS system [161], was also found in KJ073, which increased PEP precursor supply for succinate production [101]. Reverse metabolic engineering was performed to verify the effects of these two core mutations. After increasing PCK activity and deleting ptsI gene in wild-type E. coli ATCC 8739, succinate titer and yield increased 3.7- and 4.6-fold compared with parent strain, respectively [162].
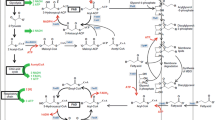
Besides inactivating competitive fermentation pathways, increasing energy supply, and increasing precursor supply, the fourth key factor for efficient succinate production is increasing reducing equivalent supply. As mentioned above, activating glyoxylate bypass and recruiting formate dehydrogenase could increase reducing equivalent supply [150, 158]. In addition, two reducing equivalent conserving pathways were identified recently, which could increase succinate yield [163]. By combining metabolic engineering and metabolic evolution, a high-succinate-producing strain HX024 was obtained (Fig. 4), which produced 813 mM succinate with a yield of 1.36 mol/mol using mineral salts medium and one-step anaerobic process [163]. Genetic mechanisms for high yield were then identified through genome sequencing and transcriptome and enzyme assay analysis. Pyruvate dehydrogenase (PDH) activity increased significantly, and sensitivity of PDH to NADH was eliminated by three mutations in LpdA, which is the E3 component of PDH [164–166]. On the other hand, pentose phosphate pathway (PPP) and transhydrogenase SthA [167–169] were activated. More carbon flux could go through the pentose phosphate pathway, thus leading to production of more reducing equivalent in the form of NADPH, which was then converted to NADH through soluble transhydrogenase for succinate production. Reverse metabolic engineering was further performed in the parent strain. Succinate yield increased from 1.12 to 1.5 mol/mol (88 % of theoretical maximum yield) by activating PDH, PPP, and SthA transhydrogenase in combination. It was suggested that the theoretical maximum succinate yield can also be obtained if 85.7 % of the carbon source goes through PPP, using both NADH and NADPH as the reducing equivalents [163]. The other benefit of using the PPP for succinate production is that only half exogenous CO2 is required, which could reduce the fermentation cost [163].
Genetic mechanisms for high-succinate production in E. coli strain HX024. Green square lines represent activated metabolic modules (including pentose phosphate, transhydrogenase, and pyruvate dehydrogenase) which are responsible for increased succinate yield. Pentose phosphate pathway was activated to increase the supply of NADPH, which was then converted to NADH through soluble transhydrogenase for succinate production. The sensitivity of pyruvate dehydrogenase to NADH inhibition was eliminated by the lpdA gene mutation. Pyruvate dehydrogenase activity increased under anaerobic conditions, which provided additional NADH for succinate production. Red oval lines represent activated metabolic modules (including glucose transport, carboxylation, reductive TCA, and succinate transport) which are responsible for increased succinate productivity. G6P glucose-6-phosphate, 6PGL gluconolactone-6-phosphate, 6PGC 6-phospho gluconate, Ru5P ribulose-5-phosphate, X5P xylurose-5-phosphate, R5P ribose-5-phosphate, S7P sedoheptulose-7-phosphate, E4P erythrose-4-phosphate, F6P fructose- 6-phosphate, FBP fructose-1,6-bisphosphate, GAP glyceraldehyde-3-phosphate, DHAP dihydroxyacetonephosphate, 1,3-BPG 1,3-bisphosphoglycerate, 3-PG 3-phosphoglycerate, 2-PG 2-phosphoglycerate, PEP phosphoenolpyruvate, ACP acetylphosphate, Ace acetate, DLAC D-lactate, FOR formate, ETH ethanol, OAA oxaloacetate, CIT citrate, ICIT isocitrate, GLO glyoxylate, MAL malate, FUM fumarate, SUC succinate, NAD + oxidized nicotinamide adeninedinucleotide, NADPH reducednicotinamide adenine dinucleotidephosphate, NADH reduced nicotinamide adeninedinucleotide, and NADP + oxidized nicotinamide adeninedinucleotide phosphate
3.4 Malate
Malate, together with fumarate and succinate, has been identified as one of the 12 most valuable bulk chemicals by the US Department of Energy [126]. It can be produced by several native microorganisms [170–174]. Since converting one molecule pyruvate to one molecule malate only requires one NADH, the theoretical maximum yield for malate production can be 2 mol/mol glucose. Starting from a succinate-producing strain KJ073, the Ingram’s group developed an engineered E. coli strain for l-malate production [175]. Inactivating fumarase isoenzymes could not convert the succinate-producing strain to produce malate, and the resulting strain still accumulated large amounts of succinate. Fumarate appears to be the immediate precursor for succinate production in a fumarase-negative background. By contrast, it was surprisingly found that inactivation of fumarate reductase alone could reforce the carbon flow into malate production. It was suggested that the thermodynamic equilibrium favors the hydration of fumarate to malate and E. coli might have a better malate-transporting capability than fumarate. Inactivation of fumarase and malic enzymes further improve malate production. Strain XZ-T658 was obtained which produced 163 mM malate with a yield of 1.0 mol/mol glucose. When using a two-stage process, 253 mM malate was produced within 72 h and the yield reached 1.42 mol/mol [175].
3.5 Fumarate
Production of fumarate using fermentative process has been studied a century ago [176, 177], and the focus has been concentrated on the Rhizopus strains [176, 178–188]. The best reported strain can produce 126 g/L fumarate with a yield of 0.97 g/g form glucose [189]. Recently, E. coli was also engineered by the Lee’s group for fumarate production under aerobic conditions [190]. The carbon flux was redirected through the glyoxylate shunt by deletion of the iclR gene, while the fumarate production was increased by the deletion of the fumA, fumB, and fumC genes. The engineered strain produced 1.45 g/L fumarate when glucose was used as the substrate. The ppc gene was then overexpressed, and the fumarate production increased to 4.09 g/L. To reach better performance, further genetic modifications were carried out including deletion of arcA (encoding ArcA transcriptional dual regulator) and ptsG genes to increase the oxidative TCA cycle flux, deletion of aspA (encoding aspartate ammonia-lyase) to decrease the degradation of fumarate, and replacement of the native promoter of galP by a strong trc promoter to promote the uptake of glucose. Strain CWF812 was obtained which produced 28.2 g/L fumarate with a yield of 0.389 g/g glucose when fermented in fed-batch for 63 h [190].
3.6 Glucaric Acid
d-Glucaric acid, a compound present in fruits, vegetables, and mammals, has been studied for therapeutic purpose [191–193], and it has potential applications for polymers [126]. The synthetic pathway for d-glucaric acid production from glucose is present naturally in mammals. However, this natural pathway is composed of more than 10 reactions and limits its construction in E. coli. To realize the production of d-glucaric acid, the Prather’s group designed a synthetic pathway by coexpression of ino1 encoding myoinositol-1-phosphate synthase from S. cerevisiae, miox encoding myoinositol oxygenase from mice, and udh encoding the urinate dehydrogenase from Pseudomonas syringae in E. coli. The resulting strain produced more than 1 g/L of glucaric acid using LB medium with 10 g/L glucose [194]. MIOX was identified as the rate-limiting step in the whole pathway, and its activity was strongly affected by the myoinositol concentration. To improve the flux for glucaric acid production, two strategies were carried out. Utilization of protein scaffold to colocalize the three heterologous enzymes in a designable complex resulted in fivefold improvement of glucaric acid titer [195]. On the other hand, protein fusion tags and directed evolution were used to improve MIOX activity, leading to the production of 4.85 g/L glucaric acid from 10.8 g/L myoinositol [196].
3.7 Muconic Acid
Muconic acid (MA) is an important unsaturated dicarboxylic acid and has great potential for the production of bioplastics [197–199]. It can also be used as the precursor for the synthesis of important bulk chemicals, such as adipic acid, terephthalic acid, and trimellitic acid [197]. Biosynthesis of muconic acid in E. coli has been studied from 1994 by the Frost’s group [200]. They described an artificial pathway for muconic acid biosynthesis from glucose by combining the shikimic acid pathway, which is natively present in E. coli for aromatic amino acid synthesis, with three heterologous enzymes including 3-dehydroshikimate (DHS) dehydratase, protocatechuic acid (PCA) decarboxylase, and catechol 1,2-dioxygenase (CDO). Inactivation of shikimate dehydrogenase to reduce the DHS consumption and overexpression of transketolase, 3-deoxy-d-arabinoheptulosonate 7-phosphate (DAHP) synthase, and 3-dehydroquinate (DHQ) synthase to increase the availability of DHS were further performed to improve muconic acid production. The resulting strain produced 2.4 g/L muconic acid in a batch fermentation. Deregulation of the feedback inhibition of shikimic acid pathway and overexpression of the critical genes increased muconic acid titer to 38.6 g/L [201]. Optimization of the fermentation process using fed-batch conditions further improved the titer to 59.2 g/L [202].
A novel artificial pathway for MA production in E. coli was established by integration of the native tryptophan biosynthetic pathway with a heterologous anthranilate degradation pathway [199]. In this pathway, anthranilate which is an intermediate involved in the native tryptophan biosynthetic pathway was transformed into MA sequentially by anthranilate 1,2-dioxygenase (ADO) from Pseudomonas aeruginosa and catechol 1,2-dioxygenase (CDO) from P. putida. The MA production was optimized by screening several enzyme candidates and improving the native tryptophan biosynthetic pathway. The resulting strain produced 389 mg/L muconic acid using the modified M9 minimal medium with a mixture carbon sources of glycerol and glucose [199].
Another novel MA synthetic pathway was designed via extending shikimate pathway by introducing the hybrid of a salicylic acid (SA) biosynthetic pathway with its partial degradation pathway [198]. A well-developed phenylalanine-producing strain was first engineered to produce SA by heterologous expression of the isochorismate synthase and isochorismate pyruvate lysate, leading to production of 1.2 g/L of SA. The SA was then converted into MA by introducing salicylate 1-monoxygenase and catechol 1,2-dioxygenase. Optimization of the whole pathway resulted in the production of MA up to 1.5 g/L after 48-h fermentation in shake flasks [198].
3.8 Adipic Acid
As the most important dicarboxylic acid, it is estimated that the market volume of adipic acid is about 2.6 million tons per year in global and an increase of 3–3.5 % will be expected annually [203, 204]. The primary use of adipic acid is as precursor for the production of polyamide nylon-6,6 [200, 203–205]. Traditionally, adipic acid is produced by chemical catalytic pathway in industrial large-scale processes using benzene, an important compound derived from non-renewable fossil resource, as the principal starting compound [200, 203]. To decrease the dependence on fossil feedstock, many efforts have been made in the past years to develop an alternative way to produce adipic acid from renewable biomass resources [11, 200, 201, 203, 205–211].
Although cis,cis-muconic acid can be converted to adipic acid by chemical hydrogenation [200, 201], it is still designed to construct cell factories for producing adipic acid directly through glucose fermentation [203, 205, 206, 208, 210]. Recently, the Zhong group at Shanghai Jiao Tong University constructed an artificial adipic acid synthetic pathway in E. coli [205]. Acetyl-CoA and succinyl-CoA were condensed to produce the C6 backbone 3-oxoadipyl-CoA, which was then converted to adipic acid sequentially via 3-hydroxyadipyl-CoA, 2,3-dehydroadipyl-CoA, and adipyl-CoA. The six enzymatic steps were catalyzed respectively by the β-ketoadipyl-CoA thiolase (PaaJ) from E. coli, 3-hydroxybutyryl-CoA dehydrogenase (Hbd) and crotonase (Crt) from Clostridium acetobutylicum, trans-enoyl-CoA reductase (Ter) from Euglena gracilis, and phosphate butyryltransferase (Ptb) and butyryl kinase (Buk1) from C. acetobutylicum. The constructed strain AA1 produced 31 μg/L adipic acid when fermented in minimal R/2 medium supplemented with 10 g/L glucose aerobically at 30°C for 120 h. The adipic acid titer increased to 120 μg/L when replacing Ter with butyryl-CoA dehydrogenase (Bcd) from C. acetobutylicum, replacing Hbd with 3-hydroxyacyl-CoA reductase (PaaH1) from Ralstonia eutropha, and replacing Crt with the putative enoyl-CoA hydratase (ECH) from R. eutropha H16. Supplies of acetyl-CoA and succinyl-CoA precursors were then increased to further improve adipic acid production, resulting in strain AA7 which produced 639 μg/L adipic acid which was about 20-fold higher than that of the starting strain AA1 [205].
4 Alcohols
E. coli cell factories have been constructed for production of a variety of alcohols, such as 1,3-propanediol [19, 212], 1-propanol [213], 1,2-propanediol [214, 215], isopropanol [216], n-butanol [217], isobutanol [13], 1,4-butanediol [12], and higher-chain alcohols [13, 51, 218, 219]. Some reviews have been focused on the elucidation of the bio-based production of alcohols using E. coli cell factories [2, 3, 217, 220], which will not be described here. This chapter will focus on recently developed cell factories for the production of higher-chain alcohols and 1,4-butanediol.
4.1 Higher-Chain Alcohols
Higher-chain alcohols are attractive biofuel targets because they exhibit higher energy density, lower hygroscopicity, lower vapor pressure, and compatibility with present transportation devices [218]. However, these compounds are not synthesized economically by native organisms [50]. Two different synthetic pathways have been created for the production of these compounds.
By the introduction of the broad-substrate-range 2-keto acid decarboxylases (KDCs) and alcohol dehydrogenases (ADHs) genes, the amino acid synthetic pathways can be redirected to produce higher-chain alcohols from 2-keto acids in E. coli [221]. Isobutanol is a representative example. A synthetic pathway for isobutanol production from glucose had been created by the Liao’s group through combining branched-chain amino acid synthetic pathway and Ehrlich pathway with 2-keto-isovalerate serving as a precursor [13]. Overexpression of the valine biosynthetic pathway (ilvIHCD) and the alcohol producing pathway (kivD from Lactococcus lactis and adh2 from S. cerevisiae) resulted in production of 1.7 g/L isobutanol. Competitive fermentation pathways, including adhE, ldhA, frdAB, fnr, and pta, were further deleted to increase the pyruvate supply for the isobutanol production. The resulted strain produced 2.2 g/L isobutanol with a yield of 0.21 g/g glucose. Further improvement was carried out by replacing the native ilvIH by the alsS gene from B. subtilis. AlsS has higher affinity for pyruvate than IlvIH, and the replacement increased isobutanol titer up to 3.7 g/L. With a deletion of the pflB gene, the isobutanol titer increased to 22 g/L under microaerobic conditions [13]. When fermented in a 1-L bioreactor instead of the shake flask with in situ isobutanol removal using gas stripping, the isobutanol production could reach a concentration of more than 50 g/L in 72 h [221].
NADPH is the reducing equivalent required for the production of isobutanol. Both keto acid reductoisomerase and alcohol dehydrogenase are NADPH dependent, and two equivalents of NADPH are required for the conversion of pyruvate to isobutanol. In contrast, the common reducing equivalent under anaerobic condition is NADH, which is produced through glycolysis [222]. In order to solve this cofactor imbalance problem, the cofactor specificity of keto acid reductoisomerase and alcohol dehydrogenase enzymes was changed from NADPH to NADH, and theoretical yield was obtained under anaerobic condition [222]. On the other hand, membrane-bound transhydrogenase PntAB and NAD kinase were activated in combination to increase the NADPH supply for improved isobutanol production [35]. Activating these two enzymes increased anaerobic isobutanol yield by 39 % to 0.92 mol/mol glucose [35].
Enzymes involved in the native L-leucine biosynthesis pathway were designed to catalyze the chain elongation, and various 2-keto acids were obtained. Diverse LeuA mutants were generated to suit the different size of the substrates [13, 16, 51, 217, 218]. These 2-keto acids were then successively converted to aldehyde and alcohols in turn by introducing KDCs and ADHs. By this pathway design, one carbon atom was added to the chain in each cycle and a new pool of alcohols can be produced.
A second pathway designed for higher-chain alcohol production was based on acetyl-CoA. Two carbon atoms were added to the chain in each cycle [217, 223–225]. Five reactions were involved in the carbon chain elongation reactions and the production of alcohols, which were catalyzed by the thiolase (AtoB/BktB), dehydrogenase (Hbd/PaaH1), dehydrase (Crt), reductase (Ter), and thioesterase (TesB), respectively. These enzymes were not specific to all compounds with different carbon number. It is critical to find more specific enzymes to increase the efficiency for certain products.
4.2 1,4-Butanediol
As one of the important C4 platform chemicals, 1,4-butanediol (BDO) owns a world market exceeding 1 million tons and is used widely in the manufacture of biopolymers, cosmetics, fine chemicals, and solvents [3, 19]. BDO is predominantly produced from crude oil and natural gas. No biosynthetic pathways have been reported in any natural organisms. The project of constructing an E. coli cell factory for BDO production was initiated by Genomatica [12]. All candidate pathways from E. coli central metabolites to BDO were elucidated based on the SimPheny Biopathway Predictor software. Rather than by known enzyme reactions, the transformation of functional groups by known chemistry was used as the basis for the Biopathway Predictor algorithm, and this gave a chance to identify novel enzyme activities or to engineer enzymes with specific activities to a particular substrate. As a result, 10,000 pathways for the BDO synthesis from common central metabolites were identified. Then, the proposed pathways were evaluated based on the different factors including maximum theoretical yield, pathway length, number of non-native steps, and thermodynamic feasibility. At last, two pathways for BDO biosynthesis, which involved 4-hydroxybutyrate as the intermediate, were proposed as the highest priority and tested in vivo. A pathway was proved to be potential for BDO production in which BDO was produced from succinate via six enzymatic reactions catalyzed by two E. coli native enzymes (succinyl-CoA synthetase and alcohol dehydrogenase) and four heterologous enzymes (CoA-dependent succinate semialdehyde dehydrogenase, 4-hydroxybutyrate dehydrogenase, 4-hydroxybutyryl-CoA transferase, and 4-hydroxybutyryl-CoA reductase). The engineered strain produced over 18 g/L BDO from glucose in 5 days [12]. It could also produce BDO from sucrose, xylose, and biomass-derived mixed sugar streams.
To increase the efficiency of the BDO biosynthetic pathway, butyraldehyde dehydrogenase (Bld) and butanol dehydrogenase from Clostridium saccharoperbutylacetonicum were selected and used for BDO production in E. coli. Furthermore, random mutagenesis and site-directed mutagenesis were carried out in turn to improve the activity of Bld. The resulted strain could produce BDO with the titer fourfold greater than those of strains expressing the wild-type Bld [37].
5 Perspectives
Production of bulk chemicals by E. coli cell factories from renewable biomass resources has been proved to be a sustainable and environment-friendly process to replace the petroleum-based process. Due to the rapid development of metabolic engineering, systems biology, and synthetic biology, great progress has been achieved and many successful E. coli cell factories have been constructed. However, there are still several challenges.
Only a few bulk chemicals can be produced biologically by microorganisms. Although several new synthetic pathways have been designed and created, most of the bulk chemicals still do not have biosynthetic pathways. The primary reason is that many chemical reactions do not have natural enzymes. Creation of new enzymes to catalyze the desired chemical reaction by integrating chemistry, protein rational design, and directed evolution will be in demand to full in the gaps existing in the novel biosynthetic pathway [16].
The created synthetic pathway is usually not efficient due to the low catalytic activities of some specific enzymes, especially those new enzymes. It is thus very important to improve the catalytic capabilities of these enzymes so that they are not rate limiting within the whole pathway. In addition, coordinated expression of multiple genes involved in the synthetic pathway is desirable so that there will be no metabolic imbalance problem such as exhaustion of precursor and accumulation of toxic intermediates [226]. Product yield is an important factor to realize low-cost production of bulk chemicals. On the one hand, redox balance is necessary to maintain anaerobic cell growth since the only way to consume the reducing equivalent is through the synthetic pathway of target compound. On the other hand, enough reducing equivalent is required to reach the theoretical maximum yield.
Finally, good physiological characteristics of cell factories are necessary for large-scale industrial production of bulk chemicals [227]. Tolerance to high osmolality and high concentration of target chemicals can increase the final titer and productivity. Tolerance to high temperature can reduce the energy cost and contamination problems. Tolerance to low pH can produce organic acid directly so that complex downstream purification process can be avoided to convert organic acid salt to organic acid. Since modifying single gene usually has no effect on improving physiological characteristics, global perturbation strategies together with high-throughput omics analysis are needed to improve these physiological characteristics and identify the genetic mechanisms so that bulk chemicals can be produced biologically in a cost-comparable way compared to petrochemical process.
References
Vickers C, Klein-Marcuschamer D, Krömer J (2012) Examining the feasibility of bulk commodity production in Escherichia coli. Biotechnol Lett 34:585–596
Zeng AP, Biebl H (2002) Bulk chemicals from biotechnology: the case of 1,3-propanediol production and the new trends. Adv Biochem Eng Biotechnol 74:239–259
Chen X, Zhou L, Tian K et al (2013) Metabolic engineering of Escherichia coli: a sustainable industrial platform for bio-based chemical production. Biotechnol Adv 31:1200–1223
Keasling JD (2010) Manufacturing molecules through metabolic engineering. Science 330:1355–1358
Taher H, Al-Zuhair S, Al-Marzouqi AH et al (2011) A review of enzymatic transesterification of microalgal oil-based biodiesel using supercritical technology. Enzyme Res 2011:468292
U.S. Department of Energy EIA (2012) Petroleum marketing monthly. Department of Energy, Washington, DC, DOE Publ. No. EIA-0380(20012/02)
U.S. Department of Energy EIA (2006) Annual energy outlook 2006 with projections to 2030. Department of Energy, DOE Publ. No. EIA-0383(2006), Washington, DC
U.S. Department of Energy EIA (2007) Annual energy outlook 2007 with projections to 2030. Departmentt of Energy, DOE/EIA-0383(2007), Washington, DC
Lee JW, Na D, Park JM et al (2012) Systems metabolic engineering of microorganisms for natural and non-natural chemicals. Nat Chem Biol 8:536–546
Demirbas A (2009) Political, economic and environmental impacts of biofuels: a review. 86. Appl Energy 86(1):S108–S117
Jang YS, Kim B, Shin JH et al (2012) Bio-based production of C2-C6 platform chemicals. Biotechnol Bioeng 109:2437–2459
Yim H, Haselbeck R, Niu W et al (2011) Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat Chem Biol 7:445–452
Atsumi S, Hanai T, Liao JC (2008) Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451:86–89
Jambunathan P, Zhang K (2014) Novel pathways and products from 2-keto acids. Curr Opin Biotechnol 29:1–7
Schiweck H, Bär A, Vogel R et al (2000) Sugar Alcohols, in Ullmann’s Encyclopedia of Industrial Chemistry. Wiley-VCH Verlag GmbH & Co, KGaA
Shin JH, Kim HU, Kim DI et al (2013) Production of bulk chemicals via novel metabolic pathways in microorganisms. Biotechnol Adv 31:925–935
Jarboe LR, Liu P, Kautharapu KB et al (2012) Optimization of enzyme parameters for fermentative production of biorenewable fuels and chemicals. Comput Struct Biotechnol J 3:e201210005
Stephanopoulos G (2012) Synthetic biology and metabolic engineering. ACS Synth Biol 1:514–525
Nielsen J, Fussenegger M, Keasling J et al (2014) Engineering synergy in biotechnology. Nat Chem Biol 10:319–322
Schirmer A, Rude MA, Li X et al (2010) Microbial biosynthesis of alkanes. Science 329:559–562
Yu C, Cao Y, Zou H et al (2011) Metabolic engineering of Escherichia coli for biotechnological production of high-value organic acids and alcohols. Appl Microbiol Biotechnol 89:573–583
Zhou L, Tian KM, Niu DD et al (2012) Improvement of D-lactate productivity in recombinant Escherichia coli by coupling production with growth. Biotechnol Lett 34:1123–1130
Zhou L, Niu DD, Tian KM et al (2012) Genetically switched D-lactate production in Escherichia coli. Metab Eng 14:560–568
Utrilla J, Licona-Cassani C, Marcellin E et al (2012) Engineering and adaptive evolution of Escherichia coli for D-lactate fermentation reveals GatC as a xylose transporter. Metab Eng 14:469–476
Zhou L, Zuo ZR, Chen XZ et al (2011) Evaluation of genetic manipulation strategies on D-lactate production by Escherichia coli. Curr Microbiol 62:981–989
Yang J, Wang Z, Zhu N et al (2014) Metabolic engineering of Escherichia coli and in silico comparing of carboxylation pathways for high succinate productivity under aerobic conditions. Microbiol Res 169:432–440
Jiang M, Chen X, Liang L et al (2014) Co-expression of phosphoenolpyruvate carboxykinase and nicotinic acid phosphoribosyltransferase for succinate production in engineered Escherichia coli. Enzyme Microb Technol 56:8–14
Tang J, Zhu X, Lu J et al (2013) Recruiting alternative glucose utilization pathways for improving succinate production. Appl Microbiol Biotechnol 97:2513–2520
Tan Z, Zhu X, Chen J et al (2013) Activating phosphoenolpyruvate carboxylase and phosphoenolpyruvate carboxykinase in combination for improvement of succinate production. Appl Environ Microbiol 79:4838–4844
Kim K, Kim SK, Park YC et al (2014) Enhanced production of 3-hydroxypropionic acid from glycerol by modulation of glycerol metabolism in recombinant Escherichia coli. Bioresour Technol 156:170–175
Jung WS, Kang JH, Chu HS et al (2014) Elevated production of 3-hydroxypropionic acid by metabolic engineering of the glycerol metabolism in Escherichia coli. Metab Eng 23:116–122
Kwak S, Park YC, Seo JH (2013) Biosynthesis of 3-hydroxypropionic acid from glycerol in recombinant Escherichia coli expressing Lactobacillus brevis dhaB and dhaR gene clusters and E. coli K-12 aldH. Bioresour Technol 135:432–439
Rathnasingh C, Raj SM, Lee Y et al (2012) Production of 3-hydroxypropionic acid via malonyl-CoA pathway using recombinant Escherichia coli strains. J Biotechnol 157:633–640
Desai SH, Rabinovitch-Deere CA, Tashiro Y et al (2014) Isobutanol production from cellobiose in Escherichia coli. Appl Microbiol Biotechnol 98:3727–3736
Shi A, Zhu X, Lu J et al (2013) Activating transhydrogenase and NAD kinase in combination for improving isobutanol production. Metab Eng 16:1–10
Trinh CT, Li J, Blanch HW et al (2011) Redesigning Escherichia coli metabolism for anaerobic production of isobutanol. Appl Environ Microbiol 77:4894–4904
Hwang HJ, Park JH, Kim JH et al (2014) Engineering of a butyraldehyde dehydrogenase of Clostridium saccharoperbutylacetonicum to fit an engineered 1,4-butanediol pathway in Escherichia coli. Biotechnol Bioeng 111:1374–1384
Wang W, Lu X (2013) Microbial synthesis of alka(e)nes. Front Bioeng Biotechnol. doi: 10.3389/fbioe.2013.00010
Howard TP, Middelhaufe S, Moore K et al (2013) Synthesis of customized petroleum-replica fuel molecules by targeted modification of free fatty acid pools in Escherichia coli. Proc Natl Acad Sci U S A 110:7636–7641
Riemer SA, Rex R, Schomburg D (2013) A metabolite-centric view on flux distributions in genome-scale metabolic models. BMC Syst Biol 7:33
McCloskey D, Palsson BO, Feist AM (2013) Basic and applied uses of genome-scale metabolic network reconstructions of Escherichia coli. Mol Syst Biol 9:661
Xu Z, Sun X, Sun J (2013) Construction and analysis of the model of energy metabolism in E. coli. PLoS One 8:e55137
Khodayari A, Zomorrodi AR, Liao JC et al (2014) A kinetic model of Escherichia coli core metabolism satisfying multiple sets of mutant flux data. Metab Eng. doi:10.1016/j.ymben.2014.05.014
Zomorrodi AR, Lafontaine Rivera JG, Liao JC et al (2013) Optimization-driven identification of genetic perturbations accelerates the convergence of model parameters in ensemble modeling of metabolic networks. Biotechnol J 8:1090–1104
Xu Z, Zheng P, Sun J et al (2013) ReacKnock: identifying reaction deletion strategies for microbial strain optimization based on genome-scale metabolic network. PLoS One 8:e72150
Chang RL, Andrews K, Kim D et al (2013) Structural systems biology evaluation of metabolic thermotolerance in Escherichia coli. Science 340:1220–1223
Durot M, Bourguignon P-Y, Schachter V (2009) Genome-scale models of bacterial metabolism: reconstruction and applications. FEMS Microbiol Rev 33:164–190
O’Brien EJ, Lerman JA, Chang RL et al (2013) Genome-scale models of metabolism and gene expression extend and refine growth phenotype prediction. Mol Syst Biol 9:693
Cho BK, Kim D, Knight EM et al (2014) Genome-scale reconstruction of the sigma factor network in Escherichia coli: topology and functional states. BMC Biol 12:4
Atsumi S, Liao JC (2008) Metabolic engineering for advanced biofuels production from Escherichia coli. Curr Opin Biotechnol 19:414–419
Marcheschi RJ, Li H, Zhang K et al (2012) A synthetic recursive “+1” pathway for carbon chain elongation. ACS Chem Biol 7:689–697
Planson AG, Carbonell P, Grigoras I et al (2012) A retrosynthetic biology approach to therapeutics: from conception to delivery. Curr Opin Biotechnol 23:948–956
Rodrigo G, Carrera J, Prather KJ et al (2008) DESHARKY: automatic design of metabolic pathways for optimal cell growth. Bioinformatics 24:2554–2556
Hatzimanikatis V, Li C, Ionita JA et al (2005) Exploring the diversity of complex metabolic networks. Bioinformatics 21:1603–1609
Cho A, Yun H, Park JH et al (2010) Prediction of novel synthetic pathways for the production of desired chemicals. BMC Syst Biol 4:35
Mavrovouniotis M, Stephanopoulos G (1992) Synthesis of biochemical production routes. Comput Chem Eng 16:605–619
Tian J, Gong H, Sheng N et al (2004) Accurate multiplex gene synthesis from programmable DNA microchips. Nature 432:1050–1054
Quan J, Saaem I, Tang N et al (2011) Parallel on-chip gene synthesis and application to optimization of protein expression. Nat Biotechnol 29:449–452
Shetty RP, Endy D, Knight TF Jr (2008) Engineering BioBrick vectors from BioBrick parts. J Biol Eng 2:5
Xu P, Vansiri A, Bhan N et al (2012) ePathBrick: a synthetic biology platform for engineering metabolic pathways in E. coli. ACS Synth Biol 1:256–266
Ellis T, Adie T, Baldwin GS (2011) DNA assembly for synthetic biology: from parts to pathways and beyond. Integr Biol (Camb) 3:109–118
Hillson N (2011) DNA Assembly method standardization for synthetic biomolecular circuits and systems. In: Koeppl H, Setti G, di Bernardo M et al (eds) Design and analysis of biomolecular circuits. Springer, New York, pp 295–314
Quan J, Tian J (2014) Circular polymerase extension cloning. Methods Mol Biol 1116:103–117
Li MZ, Elledge SJ (2007) Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nat Methods 4:251–256
Gibson DG, Young L, Chuang RY et al (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6:343–345
Shao Z, Zhao H (2009) DNA assembler, an in vivo genetic method for rapid construction of biochemical pathways. Nucleic Acids Res 37:e16
Du J, Yuan Y, Si T et al (2012) Customized optimization of metabolic pathways by combinatorial transcriptional engineering. Nucleic Acids Res 40:e142
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645
Jarboe LR, Zhang X, Wang X et al (2010) Metabolic engineering for production of biorenewable fuels and chemicals: contributions of synthetic biology. J Biomed Biotechnol 2010:761042
Gaj T, Sirk SJ, Barbas CF (2014) Expanding the scope of site-specific recombinases for genetic and metabolic engineering. Biotechnol Bioeng 111:1–15
Posfai G, Koob MD, Kirkpatrick HA et al (1997) Versatile insertion plasmids for targeted genome manipulations in bacteria: isolation, deletion, and rescue of the pathogenicity island LEE of the Escherichia coli O157:H7 genome. J Bacteriol 179:4426–4428
Jantama K, Zhang X, Moore JC et al (2008) Eliminating side products and increasing succinate yields in engineered strains of Escherichia coli C. Biotechnol Bioeng 101:881–893
Zhang X, Jantama K, Moore JC et al (2007) Production of L -alanine by metabolically engineered Escherichia coli. Appl Microbiol Biotechnol 77:355–366
Gay P, Le Coq D, Steinmetz M et al (1985) Positive selection procedure for entrapment of insertion sequence elements in gram-negative bacteria. J Bacteriol 164:918–921
Hammer K, Mijakovic I, Jensen PR (2006) Synthetic promoter libraries–tuning of gene expression. Trends Biotechnol 24:53–55
Nevoigt E, Kohnke J, Fischer CR et al (2006) Engineering of promoter replacement cassettes for fine-tuning of gene expression in Saccharomyces cerevisiae. Appl Environ Microbiol 72:5266–5273
Alper H, Fischer C, Nevoigt E et al (2005) Tuning genetic control through promoter engineering. Proc Natl Acad Sci USA 102:12678–12683
Solem C, Jensen PR (2002) Modulation of gene expression made easy. Appl Environ Microbiol 68:2397–2403
Jensen PR, Hammer K (1998) The sequence of spacers between the consensus sequences modulates the strength of prokaryotic promoters. Appl Environ Microbiol 64:82–87
Jensen PR, Hammer K (1998) Artificial promoters for metabolic optimization. Biotechnol Bioeng 58:191–195
de la Cueva-Mendez G, Pimentel B (2007) Gene and cell survival: lessons from prokaryotic plasmid R1. EMBO Rep 8:458–464
Keasling JD (2008) Synthetic biology for synthetic chemistry. ACS Chem Biol 3:64–76
Lu J, Tang J, Liu Y et al (2012) Combinatorial modulation of galP and glk gene expression for improved alternative glucose utilization. Appl Microbiol Biotechnol 93:2455–2462
De Mey M, Maertens J, Boogmans S et al (2010) Promoter knock-in: a novel rational method for the fine tuning of genes. BMC Biotechnol 10:26
Meynial-Salles I, Cervin MA, Soucaille P (2005) New tool for metabolic pathway engineering in Escherichia coli: one-step method to modulate expression of chromosomal genes. Appl Environ Microbiol 71:2140–2144
Millard CS, Chao YP, Liao JC et al (1996) Enhanced production of succinic acid by overexpression of phosphoenolpyruvate carboxylase in Escherichia coli. Appl Environ Microbiol 62:1808–1810
Santos CN, Stephanopoulos G (2008) Combinatorial engineering of microbes for optimizing cellular phenotype. Curr Opin Chem Biol 12:168–176
Pfleger BF, Pitera DJ, Smolke CD et al (2006) Combinatorial engineering of intergenic regions in operons tunes expression of multiple genes. Nat Biotechnol 24:1027–1032
Dueber JE, Wu GC, Malmirchegini GR et al (2009) Synthetic protein scaffolds provide modular control over metabolic flux. Nat Biotechnol 27:753–759
Cobb RE, Sun N, Zhao H (2013) Directed evolution as a powerful synthetic biology tool. Methods 60:81–90
Wang HH, Isaacs FJ, Carr PA et al (2009) Programming cells by multiplex genome engineering and accelerated evolution. Nature 460:894–898
Yoo SM, Na D, Lee SY (2013) Design and use of synthetic regulatory small RNAs to control gene expression in Escherichia coli. Nat Protoc 8:1694–1707
Xu P, Wang W, Li L et al (2014) Design and kinetic analysis of a hybrid promoter-regulator system for malonyl-CoA sensing in Escherichia coli. ACS Chem Biol 9:451–458
Farmer WR, Liao JC (2000) Improving lycopene production in Escherichia coli by engineering metabolic control. Nat Biotechnol 18:533–537
Zhang F, Carothers JM, Keasling JD (2012) Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat Biotechnol 30:354–359
Cobb RE, Si T, Zhao H (2012) Directed evolution: an evolving and enabling synthetic biology tool. Curr Opin Chem Biol 16:285–291
Zhou S, Yomano LP, Shanmugam KT et al (2005) Fermentation of 10 % (w/v) sugar to D: (-)-lactate by engineered Escherichia coli B. Biotechnol Lett 27:1891–1896
Zhou S, Shanmugam KT, Yomano LP et al (2006) Fermentation of 12 % (w/v) glucose to 1.2 M lactate by Escherichia coli strain SZ194 using mineral salts medium. Biotechnol Lett 28:663–670
Grabar TB, Zhou S, Shanmugam KT et al (2006) Methylglyoxal bypass identified as source of chiral contamination in l(+) and d(-)-lactate fermentations by recombinant Escherichia coli. Biotechnol Lett 28:1527–1535
Zhang X, Shanmugam KT, Ingram LO (2010) Fermentation of glycerol to succinate by metabolically engineered strains of Escherichia coli. Appl Environ Microbiol 76:2397–2401
Zhang X, Jantama K, Moore JC et al (2009) Metabolic evolution of energy-conserving pathways for succinate production in Escherichia coli. Proc Natl Acad Sci USA 106:20180–20185
Jiang M, Wan Q, Liu R et al (2014) Succinic acid production from corn stalk hydrolysate in an E. coli mutant generated by atmospheric and room-temperature plasmas and metabolic evolution strategies. J Ind Microbiol Biotechnol 41:115–123
Trinh CT, Srienc F (2009) Metabolic engineering of Escherichia coli for efficient conversion of glycerol to ethanol. Appl Environ Microbiol 75:6696–6705
Fernandez-Sandoval MT, Huerta-Beristain G, Trujillo-Martinez B et al (2012) Laboratory metabolic evolution improves acetate tolerance and growth on acetate of ethanologenic Escherichia coli under non-aerated conditions in glucose-mineral medium. Appl Microbiol Biotechnol 96:1291–1300
Zheng H, Wang X, Yomano LP et al (2013) Improving Escherichia coli FucO for furfural tolerance by saturation mutagenesis of individual amino acid positions. Appl Environ Microbiol 79:3202–3208
Liu H, Yan M, Lai C et al (2010) gTME for improved xylose fermentation of Saccharomyces cerevisiae. Appl Biochem Biotechnol 160:574–582
Tyo KE, Alper HS, Stephanopoulos GN (2007) Expanding the metabolic engineering toolbox: more options to engineer cells. Trends Biotechnol 25:132–137
Alper H, Stephanopoulos G (2007) Global transcription machinery engineering: a new approach for improving cellular phenotype. Metab Eng 9:258–267
Alper H, Moxley J, Nevoigt E et al (2006) Engineering yeast transcription machinery for improved ethanol tolerance and production. Science 314:1565–1568
Shi DJ, Wang CL, Wang KM (2009) Genome shuffling to improve thermotolerance, ethanol tolerance and ethanol productivity of Saccharomyces cerevisiae. J Ind Microbiol Biotechnol 36:139–147
Otte B, Grunwaldt E, Mahmoud O et al (2009) Genome shuffling in Clostridium diolis DSM 15410 for improved 1,3-propanediol production. Appl Environ Microbiol 75:7610–7616
Hida H, Yamada T, Yamada Y (2007) Genome shuffling of Streptomyces sp. U121 for improved production of hydroxycitric acid. Appl Microbiol Biotechnol 73:1387–1393
Zhang YX, Perry K, Vinci VA et al (2002) Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature 415:644–646
Warner JR, Reeder PJ, Karimpour-Fard A et al (2010) Rapid profiling of a microbial genome using mixtures of barcoded oligonucleotides. Nat Biotechnol 28:856–862
Royce LA, Boggess E, Fu Y et al (2014) Transcriptomic analysis of carboxylic acid challenge in Escherichia coli: beyond membrane damage. PLoS One 9:e89580
McCloskey D, Gangoiti JA, King ZA et al (2014) A model-driven quantitative metabolomics analysis of aerobic and anaerobic metabolism in E. coli K-12 MG1655 that is biochemically and thermodynamically consistent. Biotechnol Bioeng 111:803–815
Wiback SJ, Mahadevan R, Palsson BO (2004) Using metabolic flux data to further constrain the metabolic solution space and predict internal flux patterns: the Escherichia coli spectrum. Biotechnol Bioeng 86:317–331
Oliver DJ, Nikolau B, Wurtele ES (2002) Functional genomics: high-throughput mRNA, protein, and metabolite analyses. Metab Eng 4:98–106
Patterson SD, Aebersold RH (2003) Proteomics: the first decade and beyond. Nat Genet 33(Suppl):311–323
Alonso S, Rendueles M, Diaz M (2014) Microbial production of specialty organic acids from renewable and waste materials. Crit Rev Biotechnol . doi:10.3109/07388551.2014.904269
Tsao GT, Cao NJ, Du J et al (1999) Production of multifunctional organic acids from renewable resources. In: Tsao GT, Brainard AP, Bungay HR et al (eds) Recent progress in bioconversion of Lignocellulosics. Springer, Berlin, pp 243–280
Zhou S, Causey TB, Hasona A et al (2003) Production of optically pure D-lactic acid in mineral salts medium by metabolically engineered Escherichia coli W3110. Appl Environ Microbiol 69:399–407
Shukla VB, Zhou S, Yomano LP et al (2004) Production of D(-)-lactate from sucrose and molasses. Biotechnol Lett 26:689–693
Wang Q, Yang P, Liu C et al (2013) Biosynthesis of poly(3-hydroxypropionate) from glycerol by recombinant Escherichia coli. Bioresour Technol 131:548–551
Meng DC, Shi ZY, Wu LP et al (2012) Production and characterization of poly(3-hydroxypropionate-co-4-hydroxybutyrate) with fully controllable structures by recombinant Escherichia coli containing an engineered pathway. Metab Eng 14:317–324
Werpy T GPe (2004) Top value added chemicals from biomass. U.S. Department of Energy, Washington, DC. http://www1.eere.energy.gov/biomass/pdfs/35523.pdf
Holo H (1989) Chloroflexus aurantiacus secretes 3-hydroxypropionate, a possible intermediate in the assimilation of CO2 and acetate. Arch Microbiol 151:252–256
Strauss G, Eisenreich W, Bacher A et al (1992) 13C-NMR study of autotrophic CO2 fixation pathways in the sulfur-reducing Archaebacterium Thermoproteus neutrophilus and in the phototrophic Eubacterium Chloroflexus aurantiacus. Eur J Biochem 205:853–866
Hugler M, Huber H, Stetter KO et al (2003) Autotrophic CO2 fixation pathways in archaea (Crenarchaeota). Arch Microbiol 179:160–173
Berg IA, Kockelkorn D, Buckel W et al (2007) A 3-hydroxypropionate/4-hydroxybutyrate autotrophic carbon dioxide assimilation pathway in Archaea. Science 318:1782–1786
Ansede JH, Pellechia PJ, Yoch DC (1999) Metabolism of acrylate to beta-hydroxypropionate and its role in dimethylsulfoniopropionate lyase induction by a salt marsh sediment bacterium, Alcaligenes faecalis M3A. Appl Environ Microbiol 65:5075–5081
Loh KD, Gyaneshwar P, Papadimitriou EM et al (2006) A previously undescribed pathway for pyrimidine catabolism. Proc Natl Acad Sci U S A 103:5114–5119
Kim KS, Pelton JG, Inwood WB et al (2010) The Rut pathway for pyrimidine degradation: novel chemistry and toxicity problems. J Bacteriol 192:4089–4102
Andersen G, Bjornberg O, Polakova S et al (2008) A second pathway to degrade pyrimidine nucleic acid precursors in eukaryotes. J Mol Biol 380:656–666
Valdehuesa KN, Liu H, Nisola GM et al (2013) Recent advances in the metabolic engineering of microorganisms for the production of 3-hydroxypropionic acid as C3 platform chemical. Appl Microbiol Biotechnol 97:3309–3321
Kumar V, Ashok S, Park S (2013) Recent advances in biological production of 3-hydroxypropionic acid. Biotechnol Adv 31:945–961
Jiang X, Meng X, Xian M (2009) Biosynthetic pathways for 3-hydroxypropionic acid production. Appl Microbiol Biotechnol 82:995–1003
Raj S, Rathnasingh C, Jo J-E et al (2008) Production of 3-hydroxypropionic acid from glycerol by a novel recombinant Escherichia coli BL21 strain. Process Biochem 43:1440–1446
Mohan Raj S, Rathnasingh C, Jung WC et al (2009) Effect of process parameters on 3-hydroxypropionic acid production from glycerol using a recombinant Escherichia coli. Appl Microbiol Biotechnol 84:649–657
Rathnasingh C, Raj SM, Jo JE et al (2009) Development and evaluation of efficient recombinant Escherichia coli strains for the production of 3-hydroxypropionic acid from glycerol. Biotechnol Bioeng 104:729–739
Tokuyama K, Ohno S, Yoshikawa K et al (2014) Increased 3-hydroxypropionic acid production from glycerol, by modification of central metabolism in Escherichia coli. Microb Cell Fact 13:64
Bunch PK, Mat-Jan F, Lee N et al (1997) The ldhA gene encoding the fermentative lactate dehydrogenase of Escherichia coli. Microbiology 143(Pt 1):187–195
Chatterjee R, Millard CS, Champion K et al (2001) Mutation of the ptsG gene results in increased production of succinate in fermentation of glucose by Escherichia coli. Appl Environ Microbiol 67:148–154
Donnelly MI, Millard CS, Clark DP et al (1998) A novel fermentation pathway in an Escherichia coli mutant producing succinic acid, acetic acid, and ethanol. Appl Biochem Biotechnol 70–72:187–198
Stols L, Donnelly MI (1997) Production of succinic acid through overexpression of NAD(+)-dependent malic enzyme in an Escherichia coli mutant. Appl Environ Microbiol 63:2695–2701
Escalante A, Cervantes AS, Gosset G et al (2012) Current knowledge of the Escherichia coli phosphoenolpyruvate-carbohydrate phosphotransferase system: peculiarities of regulation and impact on growth and product formation. Appl Microbiol Biotechnol 94:1483–1494
Gabor E, Gohler AK, Kosfeld A et al (2011) The phosphoenolpyruvate-dependent glucose-phosphotransferase system from Escherichia coli K-12 as the center of a network regulating carbohydrate flux in the cell. Eur J Cell Biol 90:711–720
Postma PW, Lengeler JW, Jacobson GR (1993) Phosphoenolpyruvate:carbohydrate phosphotransferase systems of bacteria. Microbiol Rev 57:543–594
Vemuri GN, Eiteman MA, Altman E (2002) Succinate production in dual-phase Escherichia coli fermentations depends on the time of transition from aerobic to anaerobic conditions. J Ind Microbiol Biotechnol 28:325–332
Sanchez AM, Bennett GN, San KY (2005) Novel pathway engineering design of the anaerobic central metabolic pathway in Escherichia coli to increase succinate yield and productivity. Metab Eng 7:229–239
Cheng KK, Wang GY, Zeng J et al (2013) Improved succinate production by metabolic engineering. Biomed Res Int 2013:538790
Clark DP (1989) The fermentation pathways of Escherichia coli. FEMS Microbiol Rev 5:223–234
Shanmugam KT, Ingram LO (2008) Engineering biocatalysts for production of commodity chemicals. J Mol Microbiol Biotechnol 15:8–15
Maloy SR, Nunn WD (1982) Genetic regulation of the glyoxylate shunt in Escherichia coli K-12. J Bacteriol 149:173–180
Lorca GL, Ezersky A, Lunin VV et al (2007) Glyoxylate and pyruvate are antagonistic effectors of the Escherichia coli IclR transcriptional regulator. J Biol Chem 282:16476–16491
Yamamoto K, Ishihama A (2003) Two different modes of transcription repression of the Escherichia coli acetate operon by IclR. Mol Microbiol 47:183–194
Cozzone AJ, El-Mansi M (2005) Control of isocitrate dehydrogenase catalytic activity by protein phosphorylation in Escherichia coli. J Mol Microbiol Biotechnol 9:132–146
Balzer GJ, Thakker C, Bennett GN et al (2013) Metabolic engineering of Escherichia coli to minimize byproduct formate and improving succinate productivity through increasing NADH availability by heterologous expression of NAD(+)-dependent formate dehydrogenase. Metab Eng 20:1–8
Stols L, Kulkarni G, Harris BG et al (1997) Expression of Ascaris suum malic enzyme in a mutant Escherichia coli allows production of succinic acid from glucose. Appl Biochem Biotechnol 63–65:153–158
Jantama K, Haupt MJ, Svoronos SA et al (2008) Combining metabolic engineering and metabolic evolution to develop nonrecombinant strains of Escherichia coli C that produce succinate and malate. Biotechnol Bioeng 99:1140–1153
Gosset G (2005) Improvement of Escherichia coli production strains by modification of the phosphoenolpyruvate:sugar phosphotransferase system. Microb Cell Fact 4:14
Zhang X, Jantama K, Shanmugam KT et al (2009) Reengineering Escherichia coli for Succinate Production in Mineral Salts Medium. Appl Environ Microbiol 75:7807–7813
Zhu X, Tan Z, Xu H et al (2014) Metabolic evolution of two reducing equivalent-conserving pathways for high-yield succinate production in Escherichia coli. Metab Eng 24:87–96
Steiert PS, Stauffer LT, Stauffer GV (1990) The lpd gene product functions as the L protein in the Escherichia coli glycine cleavage enzyme system. J Bacteriol 172:6142–6144
Guest JR, Creaghan IT (1974) Further studies with lipoamide dehydrogenase mutants of Escherichia coli K12. J Gen Microbiol 81:237–245
Guest JR, Creaghan IT (1972) Lipoamide dehydrogenase mutants of Escherichia coli K 12. Biochem J 130:8
Kim YM, Cho HS, Jung GY et al (2011) Engineering the pentose phosphate pathway to improve hydrogen yield in recombinant Escherichia coli. Biotechnol Bioeng 108:2941–2946
Sauer U, Canonaco F, Heri S et al (2004) The soluble and membrane-bound transhydrogenases UdhA and PntAB have divergent functions in NADPH metabolism of Escherichia coli. J Biol Chem 279:6613–6619
Sauer U, Lasko DR, Fiaux J et al (1999) Metabolic flux ratio analysis of genetic and environmental modulations of Escherichia coli central carbon metabolism. J Bacteriol 181:6679–6688
Battat E, Peleg Y, Bercovitz A et al (1991) Optimization of L-malic acid production by Aspergillus flavus in a stirred fermentor. Biotechnol Bioeng 37:1108–1116
Peleg Y, Rokem JS, Goldberg I (1990) A simple plate-assay for the screening of L-malic acid producing microorganisms. FEMS Microbiol Lett 55:233–236
Pines O, Even-Ram S, Elnathan N et al (1996) The cytosolic pathway of L-malic acid synthesis in Saccharomyces cerevisiae: the role of fumarase. Appl Microbiol Biotechnol 46:393–399
Taing O, Taing K (2007) Production of malic and succinic acids by sugar-tolerant yeast Zygosaccharomyces rouxii. Eur Food Res Technol 224:343–347
Kawagoe M, Hyakumura K, Suye S-I et al (1997) Application of bubble column fermentors to submerged culture of Schizophyllum commune for production of L-malic acid. J Ferment Bioeng 84:333–336
Zhang X, Wang X, Shanmugam KT et al (2011) L-malate production by metabolically engineered Escherichia coli. Appl Environ Microbiol 77:427–434
Straathof AJ, van Gulik WM (2012) Production of fumaric Acid by fermentation. Subcell Biochem 64:225–240
Ehrlich F (1911) Über die bildung von fumarsäure durch schimmelpilze. Ber Dtsch Chem Ges 44:3737–3742
Wang G, Huang D, Qi H et al (2013) Rational medium optimization based on comparative metabolic profiling analysis to improve fumaric acid production. Bioresour Technol 137:1–8
Zhou Y, Nie K, Zhang X et al (2014) Production of fumaric acid from biodiesel-derived crude glycerol by Rhizopus arrhizus. Bioresour Technol 163:48–53
Gu S, Xu Q, Huang H et al (2014) Alternative respiration and fumaric acid production of Rhizopus oryzae. Appl Microbiol Biotechnol 98:5145–5152
Goldberg I, Steiglitz B (1985) Improved rate of fumaric acid production by Tweens and vegetable oils in rhizopus arrhizus. Biotechnol Bioeng 27:1067–1069
Zhou Y, Du J, Tsao GT (2002) Comparison of fumaric acid production by Rhizopus oryzae using different neutralizing agents. Bioprocess Biosyst Eng 25:179–181
Fu YQ, Li S, Chen Y et al (2010) Enhancement of fumaric acid production by Rhizopus oryzae using a two-stage dissolved oxygen control strategy. Appl Biochem Biotechnol 162:1031–1038
Xu Q, Li S, Fu Y et al (2010) Two-stage utilization of corn straw by Rhizopus oryzae for fumaric acid production. Bioresour Technol 101:6262–6264
Ding Y, Li S, Dou C et al (2011) Production of fumaric acid by Rhizopus oryzae: role of carbon-nitrogen ratio. Appl Biochem Biotechnol 164:1461–1467
Roa Engel CA, van Gulik WM, Marang L et al (2011) Development of a low pH fermentation strategy for fumaric acid production by Rhizopus oryzae. Enzyme Microb Technol 48:39–47
Zhang B, Skory CD, Yang ST (2012) Metabolic engineering of Rhizopus oryzae: effects of overexpressing pyc and pepc genes on fumaric acid biosynthesis from glucose. Metab Eng 14:512–520
Gu C, Zhou Y, Liu L et al (2013) Production of fumaric acid by immobilized Rhizopus arrhizus on net. Bioresour Technol 131:303–307
Ling LB, Ng TK (1989). US Patent 4,877,731. Google Patents
Song CW, Kim DI, Choi S et al (2013) Metabolic engineering of Escherichia coli for the production of fumaric acid. Biotechnol Bioeng 110:2025–2034
Singh J, Gupta KP (2003) Calcium glucarate prevents tumor formation in mouse skin. Biomed Environ Sci 16:9–16
Singh J, Gupta KP (2007) Induction of apoptosis by calcium D-glucarate in 7,12-dimethyl benz [a] anthracene-exposed mouse skin. J Environ Pathol Toxicol Oncol 26:63–73
Walaszek Z, Szemraj J, Hanausek M et al (1996) d-Glucaric acid content of various fruits and vegetables and cholesterol-lowering effects of dietary d-glucarate in the rat. Nutr Res 16:673–681
Moon TS, Yoon SH, Lanza AM et al (2009) Production of glucaric acid from a synthetic pathway in recombinant Escherichia coli. Appl Environ Microbiol 75:589–595
Moon TS, Dueber JE, Shiue E et al (2010) Use of modular, synthetic scaffolds for improved production of glucaric acid in engineered E. coli. Metab Eng 12:298–305
Shiue E, Prather KL (2014) Improving D-glucaric acid production from myo-inositol in E. coli by increasing MIOX stability and myo-inositol transport. Metab Eng 22:22–31
Xie NZ, Liang H, Huang RB et al (2014) Biotechnological production of muconic acid: current status and future prospects. Biotechnol Adv 32:615–622
Lin Y, Sun X, Yuan Q et al (2014) Extending shikimate pathway for the production of muconic acid and its precursor salicylic acid in Escherichia coli. Metab Eng 23:62–69
Sun X, Lin Y, Huang Q et al (2013) A novel muconic acid biosynthesis approach by shunting tryptophan biosynthesis via anthranilate. Appl Environ Microbiol 79:4024–4030
Draths KM, Frost JW (1994) Environmentally Compatible Synthesis of Adipic Acid from D-Glucose. J Am Chem Soc 116:399–400
Niu W, Draths KM, Frost JW (2002) Benzene-free synthesis of adipic acid. Biotechnol Prog 18:201–211
Bui V, Lau MK, MacRae D et al (2013) Methods for producing isomers of muconic acid and muconate salts, United States Patent application. US 2013/0030215 A1, Google Patents
Polen T, Spelberg M, Bott M (2013) Toward biotechnological production of adipic acid and precursors from biorenewables. J Biotechnol 167:75–84
Musser MT (2000) Adipic Acid, in Ullmann’s Encyclopedia of Industrial Chemistry. Wiley-VCH Verlag GmbH & Co, KGaA
Yu JL, Xia XX, Zhong JJ et al (2014) Direct biosynthesis of adipic acid from a synthetic pathway in recombinant Escherichia coli. Biotechnol Bioeng. doi:10.1002/bit.25293
Han L, Chen W, Yuan F et al (2013) Biosynthesis of adipic acid. Sheng Wu Gong Cheng Xue Bao 29:1374–1385
Picataggio S, Beardslee T (2012) Biological methods for preparing adipic acid. Google Patents US 8,241,879 B2
Parthasarathy A, Pierik AJ, Kahnt J et al (2011) Substrate specificity of 2-hydroxyglutaryl-CoA dehydratase from Clostridium symbiosum: toward a bio-based production of adipic acid. Biochemistry 50:3540–3550
Noack H, Georgiev V, Blomberg MR et al (2011) Theoretical insights into heme-catalyzed oxidation of cyclohexane to adipic acid. Inorg Chem 50:1194–1202
Burgard AP, Pharkya P, Osterhout RE (2010) Microorganisms for the production of adipic acid and other compounds. Google Patents US 2010/0330626 A1
Dugal M, Sankar G, Raja R et al (2000) Designing a Heterogeneous Catalyst for the Production of Adipic Acid by Aerial Oxidation of Cyclohexane. Angew Chem Int Ed Engl 39:2310–2313
Nakamura CE, Whited GM (2003) Metabolic engineering for the microbial production of 1,3-propanediol. Curr Opin Biotechnol 14:454–459
Jain R, Yan Y (2011) Dehydratase mediated 1-propanol production in metabolically engineered Escherichia coli. Microb Cell Fact 10:97
Altaras NE, Cameron DC (1999) Metabolic engineering of a 1,2-propanediol pathway in Escherichia coli. Appl Environ Microbiol 65:1180–1185
Altaras NE, Cameron DC (2000) Enhanced production of (R)-1,2-propanediol by metabolically engineered Escherichia coli. Biotechnol Prog 16:940–946
Soma Y, Inokuma K, Tanaka T et al (2012) Direct isopropanol production from cellobiose by engineered Escherichia coli using a synthetic pathway and a cell surface display system. J Biosci Bioeng 114:80–85
Lan EI, Liao JC (2013) Microbial synthesis of n-butanol, isobutanol, and other higher alcohols from diverse resources. Bioresour Technol 135:339–349
Zhang K, Sawaya MR, Eisenberg DS et al (2008) Expanding metabolism for biosynthesis of nonnatural alcohols. Proc Natl Acad Sci USA 105:20653–20658
Dhande YK, Xiong M, Zhang K (2012) Production of C5 carboxylic acids in engineered Escherichia coli. Process Biochem 47:1965–1971
Bhan N, Xu P, Koffas MA (2013) Pathway and protein engineering approaches to produce novel and commodity small molecules. Curr Opin Biotechnol 24:1137–1143
Baez A, Cho KM, Liao JC (2011) High-flux isobutanol production using engineered Escherichia coli: a bioreactor study with in situ product removal. Appl Microbiol Biotechnol 90:1681–1690
Bastian S, Liu X, Meyerowitz JT et al (2011) Engineered ketol-acid reductoisomerase and alcohol dehydrogenase enable anaerobic 2-methylpropan-1-ol production at theoretical yield in Escherichia coli. Metab Eng 13:345–352
Machado HB, Dekishima Y, Luo H et al (2012) A selection platform for carbon chain elongation using the CoA-dependent pathway to produce linear higher alcohols. Metab Eng 14:504–511
Tseng HC, Prather KL (2012) Controlled biosynthesis of odd-chain fuels and chemicals via engineered modular metabolic pathways. Proc Natl Acad Sci USA 109:17925–17930
Yu P, Tai Y-S, Woodruff AP et al (2012) Engineering artificial metabolic pathways for biosynthesis. Curr Opin Chem Eng 1:373–379
Pitera DJ, Paddon CJ, Newman JD et al (2007) Balancing a heterologous mevalonate pathway for improved isoprenoid production in Escherichia coli. Metab Eng 9:193–207
Chowdhury R, Sahu G, Das J (1996) Stress response in pathogenic bacteria. J Bioscience 21:149–160
Acknowledgments
This research was supported by grants from Tianjin Key Technology R&D program of Tianjin Municipal Science and Technology Commission (11ZCZDSY09100, 13ZCZDSY05300), the National Natural Science Foundation of China (31370136), and the Key Deployment Project of the Chinese Academy of Sciences (KGZD-EW-606; KSZD-EW-Z-016-2).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Liu, P., Zhu, X., Tan, Z., Zhang, X., Ma, Y. (2015). Construction of Escherichia Coli Cell Factories for Production of Organic Acids and Alcohols. In: Ye, Q., Bao, J., Zhong, JJ. (eds) Bioreactor Engineering Research and Industrial Applications I. Advances in Biochemical Engineering/Biotechnology, vol 155. Springer, Berlin, Heidelberg. https://doi.org/10.1007/10_2014_294
Download citation
DOI: https://doi.org/10.1007/10_2014_294
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-49159-1
Online ISBN: 978-3-662-49161-4
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)