Abstract
Ralstonia solanacearum species complex is the most destructive and economically important bacterial pathogen of many plant species around the world. It is a particular constraint to the production of the solanaceous vegetables such as tomato, chilli pepper and African eggplant in West Africa. Its broad host range, ability to survive in the soil for long periods and ability to sustain latent infection, make it difficult to control. Identification, characterization and mapping the distribution of the causal Ralstonia spp. strains are necessary to help design effective control strategies. Wilted tomato, African eggplant and pepper plant samples were collected from fields in different regions of Mali. The causal bacterial strains were isolated from the samples and all were identified as Ralstonia pseudosolanacearum. Multiplex PCR using four phylotype specific primer pairs identified the presence of phylotype I strains distributed across all the vegetable production areas whereas phylotype III strains were limited to areas of Sikasso, Koulikoro and Segou to the south and east of Bamako city. Phylogenetic analysis of part of the conserved endoglucanase virulence gene sequences revealed four sequevars with most phylotype I strains identified as sequevars 46 and 31 with only a few as sequevar 14 and one sequevar 18. The phylotype III strains were all in the sequevar 23–48 group. Pathogenicity testing of a selected subset of the strains isolated in Mali showed them all to be pathogenic to the susceptible tomato cultivar Roma. To our knowledge, this is the first study on the molecular diversity within the Ralstonia spp. complex in Mali. This information is important for developing wilt management strategies including breeding for resistance and specifying quarantine restrictions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial wilt (BW) caused by soil-borne bacteria of the Ralstonia solanacearum species complex (RSSC) occurs throughout the tropics, sub-tropics and warm temperate regions of the World. More than 250 plant species in more than 50 botanical families can be affected by bacterial wilt (Wicker et al. 2007; Prior et al. 2016) and the economically important solanaceous crop species potato, tomato, eggplant and chili pepper are particularly susceptible. BW disease is characterized by a sudden wilt of the entire plant, and when the end of a wilted stem cut near the collar is immersed in clean water, a slimy bacterial oose streams into the water. Yield losses due to bacterial wilt vary according to the host, cultivar, climate, soil type, cropping system, and bacterial strain causing the disease (Lebeau et al. 2011; Yuliar et al. 2015; Jiang et al. 2017). BW has caused yield loss of up to 90% in tomato and potato, and up to 100% in banana (Yuliar et al. 2015). Jiang et al. (2017) reported disease incidence of up to 80% on tomato. Similarly, Adebayo (2011) reported 70% loss in tomato yield and that tomato production was impossible in some areas of Nigeria because of BW. The disease is complex and difficult to control (Horita et al. 2014) mainly because the bacteria sometimes can survive in the soil for many years, depending on soil type, moisture content, amendments and cultural practices, it can survive in drain water, it is genetically diverse, it has a wide host range infecting many different host species, and it has the potential for latent infection (Yuliar et al. 2015; Lin et al. 2014; Stevens et al. 2018). Consequently, BW caused by members of the RSSC is considered one of the world’s most destructive bacterial plant diseases (Yuliar et al. 2015; Mansfield et al. 2012).
The plant pathogenic species of Ralstonia are phenotypically and genotypically diverse and are considered as a species complex (Ralstonia solanacearum species complex; RSSC) (Peeters et al. 2013; Prior et al. 2016). Strains of the members of the RSSC were initially sub divided into five “races” based on host ranges, and into five “biovars” based on carbohydrate utilization (Fegan and Prior 2005; Hayward and Pegg 2013). However, at the beginning of the millennium using sequence analysis of the internal transcribed spacer (ITS) region of the 16S–23S rRNA gene (Fegan et al. 1998), the members of the RSSC were divided into four phylogenetic groups. These “phylotypes” I to IV corresponded to the geographical origins of strains from Asia, America, Africa and Indonesia respectively (Prior and Fegan 2005). The phylotypes were further separated into “sequevars” based on the partial endoglucanase (egl) gene sequences (Fegan and Prior 2005). In 2014, the members of the RSSC underwent further taxonomic revision with phylotype I from Asia and phylotype III from Africa being reclassified as R. pseudosolanacearum, phylotype II remaining as R. solanacearum and phylotype IV from Indonesia and Australia shifting to R. syzygii (Safni et al. 2014). Subsequently, this taxonomic revision was supported using genomic and proteomic studies (Prior et al. 2016).
Bacterial wilt is widespread in Sub-Saharan African countries (Toukam et al. 2009; Adebayo 2011; N’Guessan et al. 2012; Sikirou et al. 2015, 2017) where it poses a major threat to the production of solanaceous and other crops. In Mali, BW is a major constraint and many farmers have abandoned tomato production because of high incidences of BW and the absence of effective and sustainable methods to manage it (Horita et al. 2014). In order to start devising sound management strategies for BW in different locations it is necessary to identify which member(s) of the RSSC strain-types are prevalent in each area. Phylotype I, II and III strains were reported in some countries of Sub-Saharan Africa (Toukam et al. 2009; N’Guessan et al. 2012; Shutt et al. 2018), but previously in Mali only race one, biovar three strains had been reported (Thera et al. 2010) and detailed analysis using reliable and phylogenetically meaningful molecular techniques (Fegan and Prior 2005) had not been done. Thus, the aims of this study were to confirm the pathogenicity of selected BW strains and to assess the prevalence, distribution and genetic diversity (phylotype groups and sequevars) of members of the RSSC strains in Mali.
Materials and methods
Survey and sample collection
The major vegetable growing regions of Mali (Bamako, Sikasso, Korlikoro, Segoue and Kayes) were surveyed for incidence of plants with suspected bacterial wilt and samples of wilted plants were collected from farmers’ fields with appropriate permissions both during the 2017 humid growing season (August–October) and in the 2018 dry-cool season (January–March). Sampling sites were selected based on the availability of vegetable production areas with a minimum distance of approximately 10–15 km between sites. At each site/field up to five plants of tomato (Solanum lycopersicum), African eggplant (Solanum aethiopicum) and/or chilli pepper (Capsicum spp.) showing symptoms of BW were cut at collar (just above soil) level and a stem piece of approximately 10 cm long from each was collected and preserved in a cool box with ice and transported to the laboratory. Upon arrival in the laboratory, the bacterial streaming (ooze) test was performed on each sample. Samples, which did not show streaming of a milky mass of bacterial cells into the clean water were discarded from further study. Twelve diseased samples comprising amaranth (Amaranthus spp.), tomato, African eggplant and basil (Ocimum basilicum) collected from Benin Republic in October 2017 were included in this study.
Isolation of R. solanacearum species complex bacteria and transfer to FTA™ cards
Samples confirmed as BW infected in the streaming test were surface sterilized by spraying with 70% ethanol and drying on soft tissue paper. To preserve samples directly from the infected stem for molecular analysis 0.5 cm sections of stem were cut and transferred into a 1.5 ml microfuge tube containing 150 μl 70% ethanol. Without macerating, this was allowed to stand for 20 min so that the bacteria would stream out of the stem section into the ethanol. Then, 100 μl the bacterial cell suspension were transferred onto FTA™ cards (Whatman), which were then dried for two hours in a laminar flow cabinet (Burlakoti et al. 2019) and stored in a desiccator until further use.
For preparation of pure cultures from the infected stem samples, subsamples of internal stem tissue (about 8 mm3) were prepared by paring away the epidermal tissue with a sterilized scalpel. One subsample of each sample was separately macerated by chopping using sterile scalpel in about 250 μl of sterile distilled water. Macerates were streaked onto 2,3,5-Triphenyltetrazolium chloride (TZC) agar medium supplemented with 1% yeast extract (Burlakoti et al. 2019). Plates were incubated at room temperature (25–28 °C) and fluidal, irregularly round, white colonies with a pink centre typical of Ralstonia were sub-cultured on fresh TZC medium to produce pure cultures for further study.
Pure cultures of each isolate were plated onto TZC media and incubated for three days at room temperature. To prepare samples from pure cultures to send to the World Vegetable Center (WorldVeg), Taiwan for molecular analysis, a small mass of bacterial colony was transferred from the TZC plates using a wire loop into a 1.5 mL microfuge tube containing 200 μl of 70% ethanol. After mixing the bacterial cells with the ethanol, 100 μl of the bacterial suspension were transferred onto an FTA™ card and the cards were dried in a laminar flow-hood for 2 h as above. Once dried, the FTA™ cards with bound bacterial cells were kept in a desiccator at room temperature until transported to WorldVeg (Taiwan) for molecular analysis.
Species and phylotype identification
Species and phylotype identification were performed on DNA from the samples captured on FTA™ cards. Two-millimetre diameter disks cut from the sample-loaded area of the FTA card using a Harris Uni-Core™ − 2.00 punch were placed into separate 1.5 ml microfuge tubes and washed using purification reagent and 1x TE buffer following the manufacturer’s instruction (Whatman™ FTA™ Card Technology). The cleaned and dried discs were transferred into new PCR tubes and species were detected using RSSC specific primer pairs AU759f and AU760r (Table 1). PCR was performed in a total volume of 25 μl mixture reaction comprising 2.5 μl of 10X PCR buffer with 15 mM MgCl2, 0.5 μl of 2.5 mM dNTPs, 1 μl of 10 μM of each primers, 0.2 μl of 5 U/μl Taq DNA polymerase, one 2 mm disc of FTA card (equivalent to 1.0 μl) and 18.8 μl of sterile deionized water. All PCR reagents were purchased from PROtech Technology Enter Prise Co. Ltd., Taiwan. DNA amplifications were conducted in a Bio-Rad DNAEngine ® Peltier Thermal Cycler with an initial cycle of 94 °C for 3 min, 53 °C for 1 min and 72 °C for 1.5 min, then followed by 30 cycles of 94 °C for 18 s, 60 °C for 18 s and 72 °C for 18 s, with a final extension step of 72 °C for 5 min and holding at 4 °C.
Identification of phylotypes was performed using a multiplex PCR, combining four phylotype-specific primer pairs (Table 1). Amplification was performed in a 25 μl total reaction volume with 2.5 μl of 10X PCR buffer with 15 mM MgCl2, 2.0 μl of 2.5 mM dNTPs, 1.0 μl of all primers (10 μM) except Nmult23:AF which was 2.0 μl, 0.2 μl of 5 U/μl Taq DNA polymerase and two discs of FTA cards equivalent to a 2.0 μl. All PCR reagents were purchased from PROtech Technology EnterPrise Co. Ltd., Taiwan. The PCRs were performed in a Bio-Rad DNAEngine ® Peltier Thermal Cycler with an initial denaturation step at 96 °C for 5 min, followed by 35 cycles of denaturation at 94 °C for 15 s, annealing at 62 °C for 30 s and extension at 72 °C for 30 s, with a final extension step at 72 °C for 10 min and holding of 4 °C. PCR products were separated alongside a 100 bp DNA ladder for size comparison by electrophoresis in 1.5% agarose (aMResco®) gels. The gels were stained with ethidium bromide and amplicons were visualized and scored under UV illumination.
PCR amplification and sequencing of endoglucanase gene
In order to assess the genetic diversity and phylogeny of the strains the partial endoglucanase gene of selected isolates were amplified using the primer pair Endo-F/Endo-R (Table 1). A 25 μl reaction volume contained 2.5 μl of 10X PCR buffer with 15 mM MgCl2, 2.5 μl of 2.5 mM dNTPs, 2.0 μl of 10 μM of each primer, 0.2 μl of 5 U/μl Taq DNA polymerase and one disc of FTA card equal to 1.0 μl. The PCR condition comprised initial denaturation at 96 °C for 5 min followed by 30 cycles of denaturation at 95 °C for 1 min, annealing at 70 °C for 1 min and 72 °C for 2 min with the final extension at 72 °C for 10 min and holding at 4 °C. The PCR products were sequenced by Genomics BioSci & Tech Co. Ltd. Taiwan. Reference strains for phylotype I (PSS4 and PSS2016) and for phylotype II (PSS1632) from Taiwan were included in the partial egl gene sequencing for comparison. Sequences were edited using MEGA X (Kumar et al. 2018) and aligned using the online MAFFT 7 program (http://mafft.cbrc.jp/alignment/server) (Katoh and Standley 2013). Reference partial egl sequences from elsewhere in Africa and Asia in NCBI database (Supplementary Table 2) were included in the phylogenetic analysis which was performed using the Molecular Evolutionary Genetics Analysis - MEGA X application of the Neighbour-Joining method with bootstrap analysis of 1000 replicates. The evolutionary distances were computed using the Jukes-Cantor method and are in the units of the number of base substitutions per site.
Pathogenicity test
A set of 10 pure cultures of Ralstonia selected based on being from samples from diverse locations and host plants were tested for their pathogenicity on the commonly cultivated and susceptible tomato cultivar ‘Roma’, originally developed by the USDA Agricultural Research Services. Three-week-old seedlings were transferred into 12 cm diameter plastic pots containing sterile soil and compost (2:1 ratio) and grown-on in the greenhouse. Bacterial suspension of each of the pure cultures grown on TZC medium were prepared in sterile distilled water and the suspension was adjusted to OD600 = 0.3 using spectrophotometer resulting in a concentration of about 108 cfu/ml (Lin et al. 2014). At the four to six true leaves stage, the lateral roots of each seedling were damaged from one side of the pot using a sharp scalpel at 1–2 cm from the stem. Ten seedlings were used per strain and each seedling was drenched with 25 mL of bacterial suspension. Each plant was treated as a replicate in a completely randomized design experimental layout. Sterile distilled water was used as a negative control treatment. The study was conducted in the greenhouse using natural light and with an average temperature of 32 °C. Observations of wilting symptom development were taken every day starting from the seventh day after inoculation for three weeks. Disease severity was scored using the zero-to-five rating scale of Winstead and Kelman (1952) where 0 = no symptoms, 1 = 1 leaf partially wilted, 2 = 2 or 3 leaves wilted, 3 = all except the top 2 or 3 leaves wilted, 4 = all leaves wilted and 5 = dead. The strain was considered as pathogenic when at least one of the ten inoculated plants showed typical bacterial wilt symptoms. Percentage disease index (% DI) was calculated based on the scores at 28 days after inoculation using the formula
where: ni = number of plant with the particular disease score, vi = disease score, V = the highest disease score and N = the number of plants observed. After the final collection of disease data, all the inoculated plants were cut and tested for latent infection using the bacterial streaming test.
Results
Collection of samples
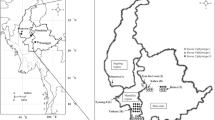
A total of 360 diseased samples from tomato, pepper and African eggplant were collected from vegetable production areas of Bamako Capital District and Cercles of four Regions of south and southwest Mali (Fig. 1). The incidence of BW was up to 70% in Bledougou (tomato and eggplant) and the disease killed all of the tomato plants in a net-house in Bamako. Ten samples comprising four from amaranth, two from tomato and two each from African eggplant and basil from the International Institute of Tropical Agriculture, IITA/WorldVeg fields and one tomato sample from farmers field in Benin Republic were included in the study. All the samples from Benin and 257 (71.4%) of those from Mali tested positive to the bacterial cell steaming test (Table 2) and subsamples of each were captured directly on separate FTA™ cards. Pure cultures with Ralstonia-like morphology and colouring on TZC medium were isolated from 114 of the samples from Mali, and bacterial suspension in 70% ethanol from each strain was captured on a separate FTA™ card and sent to WorldVeg, Taiwan for molecular analysis.
Phylotype analysis
The multiplex PCR which distinguishes between the four geographically linked monophyletic phylotypes within the members of the RSSC (Fegan and Prior 2005) was performed on DNA from 100 FTA™ card-captured samples from Mali and 11 samples from Benin. The multiplex PCR revealed that 93 (85%) of the strains were of phylotype I and 18 strains were of phylotype III (Table 3 and SpT. 1). As Safni et al. (2014) placed both phylotype I and phylotype III strains as R. pseudosolanacearum, this means that only strains of R. pseudosolanacearum were detected in the Mali samples. Although phylotype I strains were distributed across all the vegetable growing regions surveyed, the phylotype III (African origin) strains were far less dispersed with all having been collected from the area to the east and south of Bamako (Fig. 1). Both phylotypes III and I were identified from tomato, African eggplant and pepper in Mali. All the eleven strains analysed from Benin were phylotype I (Asian origin) and originated from tomato, African eggplant, basil and amaranth plants. None of the strains from Mali or Benin belonged to either the phylotype II (R. solanacearum; American origin) or phylotype IV (R. syzygii; Indonesian-Australia origin).
Sequevar analysis
Partial egl gene nucleotide sequences were obtained from 52 and five of the strains collected in Mali and Benin respectively. These 57 sequences were deposited in GenBank under accession numbers MN990211 to MN990267. Of the 57 sequences, 44 represented phylotype I strains and 13 were phylotype III strains (Table 3). The phylogenetic analysis of the partial egl gene sequences (687 positions in the final data set) in comparison to reference sequences from NCBI GenBank showed (Fig. 2) that all of the phylotype III strains were sequevar 48 (3 strains) or very closely related to and between sequevars 48 and 23 (10 strains). Of the 44 phylotype I strains sequences; 18 were sequevar 31, 19 were sequevar 46, one was sequevar 18 (very close to sequevar 46) and two were identical to the strain recently reported from amaranth in Benin (Sikirou et al. 2019) and were closely similar to sequevar 14. One of the strains from Benin was identified as sequevar 31, one was sequevar 14 and three were identical to the Benin Amaranth sample close to sequevar 14. The phylogenetic analysis confirmed the species/phylotype specific multiplex PCR result that all the strains from Mali and Benin were R. pseudosolanacearum, and no R. solanacearum (phylotype II) or R. syzygii (phylotype IV) strains were detected (Fig. 2).
Neighbor-joining tree based on the partial endoglucanase (egl) gene sequences of strains from Mali, Benin and reference members of the RSSC sequences from NCBI GenBank (see supplementary Table 2). In the code such as ‘ML001 Tom’, ML001 stands for strain code and Tom for the host crop tomato. Numbers at the end of GenBank numbers are sequevar numbers. The number at each node is the bootstrap percentage value (1000 replications). The scale bar represents one nucleotide substitution per 100 nucleotides
Pathogenicity test
All the ten strains tested (phylotype I and III) were pathogenic and caused disease on tomato cv. Roma seedlings whereas the control plants inoculated with sterile distilled water remained healthy throughout the observation period (Table 4). Strains ML3 and ML87 produced wilting symptoms by the fifth day after inoculation. Although all the strains were pathogenic, they differed in aggressiveness as indicated by the % DI at 28 days post inoculation. The highest % DI was recorded for isolate ML3 (80%) followed by ML87 (72%). Disease index (DI) of strain ML34 was 18% with average infection score of 0.9 on the 28th day after inoculation. Plants inoculated with sterile distilled water remained healthy throughout the period of the experiment.
Discussion
This study confirmed that bacterial members of the RSSC cause bacterial wilt in tomato, capsicum peppers and African eggplant in Mali (and also in amaranth and basil in Benin) and disease incidences can be up to 80% in some farmer’s fields in Mali. Testing using specific PCR primers revealed that in Mali R. pseudosolanacearum (both phylotype I and III strains) could be detected whereas only phylotype I strains were detected among the few samples tested from Benin. No R. solanacearum (= phylotype II) strains were detected in any of the wilted samples from Mali or Benin. This is similar to Thera et al. (2010) who detected no Race 3 biovar 2 (=phylotype II) strains in Mali, but different compared to other African countries such as Cote d’Ivoire, Cameroon, Ethiopia, Ghana, Kenya, Madagascar, South Africa, and Uganda (Toukam et al. 2009; N’Guessan et al. 2012; Subedi et al. 2013; Abdurahman et al. 2017; Shutt et al. 2018; Abdurahman et al. 2019) where phylotype II has been detected at varying incidence, more usually associated with brown rot of potato (Solanum tuberosum) and production at higher elevations and/or in cooler, wetter conditions (Toukam et al. 2009; N’Guessan et al. 2012; Abdurahman et al. 2017). Phylotype II is believed to have originated in the Americas and is generally considered to have been spread to Europe, Africa and Asia in seed tubers of potato. As there are few sites suitable for potato production in Mali, it is a relatively minor crop here and there has been very little import or local exchange of seed tubers. This, with the fact that no samples were collected from potato or potato-growing areas may explain why no phylotype II strains were detected. Similarly, phylotype IV (= R. syzigii) strains originated in Indonesia-Australia and are generally associated with banana and other humid tropical crops which are not grown in and have not been imported to Mali.
Phylotype I (Asian origin) strains were the most prevalent and widely distributed throughout the vegetable cultivation areas surveyed in Mali whereas phylotype III strains (African origin) were detected relatively rarely (only 15% of samples tested) in this study and all were collected from Sikasso or Segou regions in an area south and east of Bamako Capital except one which was collected in Baguineda. All of the phylotype III strains included in the phylogenetic sequevar identification study were identified as sequevar 48 (3 strains) or between sequevar 48 and sequevar 23 (10 strains). Strains of this sequevar group previously have been identified from samples from Burkina Faso, Cote d’Ivoire, Nigeria and Zambia, and closely related sequevars have been identified from Guinea and Cameroon (Wicker et al. 2007). Since phylotype III strains are considered to have an African origin (Fegan and Prior 2005), it might be expected that they might be more widely dispersed and more diverse across Africa. However, a lack of diversity and patchy distribution of phylotype III strains as identified in Mali in this study has also been reported for several other African countries including Cote d’Ivoire (~1.8%, Phylotype III; N’Guessan et al. 2012), Cameroon (18.2% phylotype III; Toukam et al. 2009), Madagascar (18% phylotype III; Ravelomanantsoa et al. 2018) and South Africa (no phylotype III; Shutt et al. 2018). On the other hand, only phylotype III strains have been reported from Burkina Faso and Guinea.
Of the phylotype I strains identified from Mali, the highest proportion were sequevar 46 (19 strains), closely followed by sequevar 31 (18 strains). Strains of these sequevars have been identified from many areas both in and outside Africa (e.g. Madagascar, Reunion Island, Uganda, South Africa, Trinidad and Tobago). Sequevar 14 and close to 14 strains predominated among the Benin samples identified and a couple also were identified from Mali. This group of sequevars appears less commonly identified from countries in Africa, but have regularly been identified from elsewhere including China and Trinidad (Ramsubhag et al. 2012; Liu et al. 2016). Together, these observations suggests that there have been multiple relatively recent introductions of phylotype I strains into Mali (and Benin). However, sequevar identification is based on sequence of only part of the endoglucanase gene and more precise identification and differentiation of strains through a multilocus-phylogeny and network analysis approach, such as used by Lin et al. (2014) or Ravelomanantsoa et al. (2018) with strains from all around the world would be required to more accurately predict the likely source of these introductions.
In the pathogenicity test, all the four phylotype III and six phylotype I strains were pathogenic on tomato cultivar Roma. However, they achieved different disease severities with DIs in the range 18–80% by 28 days post inoculation. Although there was indication that for both phylotypes the strains isolated from tomato were more aggressive on tomato than the strains isolated from African eggplant. Extensive testing of more strains on both tomato and African eggplant and other potential host species would be required to confirm if there are significant differences in pathogenicity (aggression) of the strains to different hosts. As the only previous report from Mali was of detecting only Biovar III race 1 strains (Thera et al. 2010), there is no historical data on the distribution and diversity of sequevars, it is not possible to predict how recent the introduction of phylotype I strains was, and since in this study there were phylotype I strains isolated from the same areas as phylotype III strains and the two phylotypes had similar ranges of aggression on tomato, it is not possible from this study to determine if the introduction of phylotype I strains is, or has, displaced the presumed indigenous phylotype III strains. It would be useful to determine the prevalence of phylotype I strains in neighbouring Guinea and Burkina Faso now since previously only phylotype III strains were identified in these countries (N’Guessan et al. 2012; Ravelomanantsoa et al. 2018).
Varying degrees of pathogenicity/aggression and host range and the ability to adapt to new hosts and agro-ecologies are among the reasons that members of the RSSC are some of the most destructive plant pathogenic bacteria infecting many plant species, including solanaceous crops, around the world (Mansfield et al. 2012). Better understanding of the species complex including its pathogenicity, phylotype, sequevar and distribution of the pathogen across a region are important for effective control measures to be put in place. This study has characterized the population of R. pseudosolanacearum on tomato and African eggplant in Mali using molecular tools and provided better insights for breeders, seed industries and policy makers. It is clear that breeding and selection of better adapted cultivars of tomato and African eggplant (and other crop species such as Capsicum peppers, amaranth and basil) for Mali or West Africa more generally, should include screening with both phylotype I and phylotype III strains and screening in multiple different agro-environments. Plant inspection, Ralstonia detection and quarantine measures for Mali (and other countries of West Africa) should be strengthened to reduce the likelihood of new and different strains of Ralstonia being introduced since these could overcome resistance in cultivars selected against the current locally predominant strains. The quarantine measures should also extend to potato seed tubers to prevent introduction or spread of R. solanacearum, particularly the virulent phylotype IIB brown rot strains (Carmeille et al. 2006), if there is any plan to expand and/or intensify potato production in suitable areas of the country. Phytosanitary and quarantine procedures should also apply to movement of plant materials and pathogen cultures for scientific or other purposes, and where appropriate exchange of DNA extract as suspension or captured on FTA ™ cards (Burlakoti et al. 2019) is the recommended practice.
References
Abdurahman A, Griffin D, Elphinstone J, Struik PC, Schulz S, Schulte-Geldermann E, Sharma K (2017) Molecular characterization of Ralstonia solanacearum strains from Ethiopia and tracing potential source of bacterial wilt disease outbreak in seed potatoes. Plant Pathol 66:826–834. https://doi.org/10.1111/ppa.12661
Abdurahman A, Parker ML, Kreuze J, Elphinstone JG, Struik PC, Kigundu A, Arengo E, Sharma K (2019) Molecular epidemiology of Ralstonia solanacearum species complex strains causing bacterial wilt of potato in Uganda. Phytopathology 109(11):1922–1931. https://doi.org/10.1094/PHYTO-12-18-0476-R
Adebayo OS (2011) Control of bacterial wilt disease of tomato: A review of research efforts in Nigeria. Proceedings of the 3rd International symposium on Tomato Disease. Acta Hortic 914:35–37. https://doi.org/10.17660/ActaHortic.2011.914.2
Burlakoti RR, Hsu C-F, Chen J-R, Sheu Z-M, Bihon W, Kenyon L (2019) Capture of Ralstonia solanacearum species complex strains directly from plant tissue sampled on FTA cards for molecular characterization. J Plant Pathol 102:11–17. https://doi.org/10.1007/s42161-019-00361-z
Carmeille A, Prior P, Kodja H, Chiroleu F, Luisetti J, Besse P (2006) Evaluation of resistance to race 3, biovar 2 of Ralstonia solanacearum in tomato germplasm. J Phytopathol 154:398–402. https://doi.org/10.1111/j.1439-0434.2006.01112.x
Fegan M, Prior P (2005) How complex is the Ralstonia solanacearum species complex? Pages 449–461 In: Bacterial wilt disease and the Ralstonia solanacearum species complex. Allen C, Prior P and Hayward AC eds. Aps press. St. Paul, MN
Fegan M, Taghavi M, Sly LI, Hayward AC (1998) Phylogeny, diversity and molecular diagnositcs of Ralstonia Solanacearum. Pages 19-33 In: Bacterial wilt disease: molecular and ecological aspects. Prior P, Allen C, Elphinstone J, eds. Springer, Berlin
Hayward C, Pegg KG (2013) Bacterial wilt of ginger in Queensland: reappraisal of a disease outbreak. Australas Plant Path 42:235–239. https://doi.org/10.1007/s13313-012-0174-y
Horita M, Tsuchiya K, Suga Y, Yano K, Waki T, Kurose D, Furuya N (2014) Current classification of Ralstonia solanacearum and genetic diversity of the strains in Japan. J Gen Plant Pathol 80:455–465. https://doi.org/10.1007/s10327-014-0537-z
Jiang G, Wei Z, Xu J, Chen H, Zhang Y, She X, Macho AP, Ding W, Liao B (2017) Bacterial wilt in China: history, current status, and future perspectives. Front Plant Sci 8:1549. https://doi.org/10.3389/fpls.2017.01549
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549 https://www.megasoftware.net/
Lebeau A, Daunay M-C, Frary A, Palloix A, Wang JF, Dintinger J, Chiroleu F, Wicker E, Prior P (2011) Bacterial wilt resistance in tomato, pepper and eggplant: genetic resources respond to diverse strains in the Ralstonia solanacearum species complex. Phytopathology 101:154–165. https://doi.org/10.1094/PHYTO-02-10-0048
Lin CH, Tsai KC, Prior P, Wang JF (2014) Phylogenetic relationships and population structure of Ralstonia solanacearum isolated from diverse origins in Taiwan. Plant Pathol 63(6):1395–1403. https://doi.org/10.1111/ppa.12209
Liu Y, Wu D, Liu Q, Zhang S, Tang Y, Jiang G, Li S, Ding W (2016) The sequevar distribution of Ralstonia solanacearum in tobacco-growing zones of China is structured by elevation. Eur J Plant Pathol. https://doi.org/10.1007/s10658-016-1023-6
Mansfield J, Genin S, Magori S, Citovsky V, Sriariyanum M, Ronald P, Dow M, Verdier V, Beer SV, Machado MA, Toth I, Salmond G, Foster GD (2012) Top 10 plant pathogenic bacteria in molecular plant pathology. Mol Plant Pathol 13(6):614–629. https://doi.org/10.1111/j.1364-3703.2012.00804.x
N’Guessan CA, Abo K, Fondio L, Chiroleu F, Lebeau A, Poussier S, Wicker E, Koné D (2012) So near and yet so far: the specific case of Ralstonia solanacearum populations from Côte d’Ivoire in Africa. Phytopathology 102:733–740. https://doi.org/10.1094/PHYTO-11-11-0300
Opina N, Tavner F, Hollway G, Wang J, Li T, Maghirang R, Fegan M, Hayward AC, Krishnapillai V, Hong WF, Holloway BW, Timmis JN (1997) A novel method for development of species and strain-specific DNA probes and PCR primers for identifying Burkholderia solanacearum (formerly Pseudomonas solanacearum). Asia Pac J Mol Biol Biotechnol 5:19–30
Peeters N, Guidot A, Vailleau F, Valls M (2013) Ralstonia solanacearum, a widespread bacterial plant pathogen in the post-genomic era. Mol Plant Pathol 14(7):651–662. https://doi.org/10.1111/mpp.12038
Prior P, Fegan M (2005) Recent development in the phylogeny and classification of Ralstonia solanacearum. Acta Hortic 695:127–136. https://doi.org/10.17660/ActaHortic.2005.695.14
Prior P, Ailloud F, Dalsing BL, Remenant B, Sanchez B, Allen C (2016) Genomic and proteomic evidence supporting the division of the plant pathogen Ralstonia solanacearum into three species. BMC Genomics 17(1):90. https://doi.org/10.1186/s12864-016-2413-z
Ramsubhag A, Lawrence D, Cassie D, Fraser R, Umaharan P, Prior P, Wicker E (2012) Wide genetic diversity of Ralstonia solanacearum strains affecting tomato in Trinidad, West Indies. Plant Pathol. https://doi.org/10.1111/j.1365-3059.2011.02572.x
Ravelomanantsoa S, Vernière C, Rieux A, Costet L, Chiroleu F, Arribat S, Cellier G, Pruvost O, Poussier S, Robène I, Guérin F, Prior P (2018) Molecular epidemiology of bacterial wilt in the Madagascar highlands caused by Andean (phylotype IIB-1) and African (phylotype III) Brown rot strains of the Ralstonia solanacearum species complex. Front Plant Sci 8:2258. https://doi.org/10.3389/fpls.2017.02258
Safni I, Cleenwerck I, De Vos P, Fegan M, Sly L, Kappler U (2014) Polyphasic taxonomic revision of the Ralstonia solanacearum species complex: proposal to emend the descriptions of Ralstonia solanacearum and Ralstonia syzygii and reclassify current R. syzygii strains as Ralstonia syzygii subsp. syzygii subsp. nov., R. solanacearum phylotype IV strains as Ralstonia syzygii subsp. indonesiensis subsp. nov., banana blood disease bacterium strains as Ralstonia syzygii subsp. celebesensis subsp. nov. and R. solanacearum phylotype I and III strains as Ralstonia pseudosolanacearum sp. nov. Int J Syst Evol Micr 64(9):3087–3103. https://doi.org/10.1099/ijs.0.066712-0
Shutt VM, Shin G, van der Waals GE, Goszczynska T, Coutinho TA (2018) Characterization of Ralstonia strains infecting tomato plants in South Africa. Crop Prot 112:56–62. https://doi.org/10.1016/j.cropro.2018.05.013
Sikirou R, Zocli B, Paret ML, Deberdt P, Coranson-Beaudu R, Huat J, Assogba-Komlan F, Dossoumou MEEA, Simon S, Wicker E (2015) First report of bacterial wilt of Gboma (Solanum macrocarpon) caused by Ralstonia solanacearum in Benin. Plant Dis 99(11):1640–1640. https://doi.org/10.1094/PDIS-02-15-0213-PDN
Sikirou R, Beed F, Ezin V, Hoteigni J, Miller SA (2017) Distribution, pathological and biochemical characterization of Ralstonia solanacearum in Benin. Ann Agric Sci 62(1):83–88. https://doi.org/10.1016/j.aoas.2017.05.003
Sikirou R, Dossoumou M-EEA, Zocli B, Afari-Sefa V, Honfoga J, Azoma K, Chen J-R, Paret ML, Bihon W (2019) First report of bacterial wilt of Amaranth (Amaranthus cruentus) caused by Ralstonia solanacearum in Benin. Plant Dis 103(3):578–578. https://doi.org/10.1094/pdis-07-18-1140-pdn
Stevens LH, van der Zouwen PS, van Tongeren CAM, Kastelein P, van der Wolf JM (2018) Survival of Ralstonia solanacearum and Ralstonia pseudosolanacearum in drain water. EPPO Bulletin 48(1):97–104. https://doi.org/10.1111/epp.12450
Subedi N, Gilbertson RL, Osei MK, Cornelius E, Miller SA (2013) First report of bacterial wilt caused by Ralstonia solanacearum in Ghana, West Africa. Plant Dis 98(6):840–840. https://doi.org/10.1094/PDIS-09-13-0963-PDN
Thera AT, Jacobsen BJ, Neher OT (2010) Bacterial wilt of Solanaceae caused by Ralstonia solanacearum race 1 Biovar 3 in Mali. Plant Dis 94(3):372–372. https://doi.org/10.1094/PDIS-94-3-0372B
Toukam GMS, Cellier G, Wicker E, Guilbaud C, Rm K, Allen C, Prior P (2009) Broad diversity of Ralstonia solanacearum strains in Cameroon. Plant Dis 93(11):1123–1130. https://doi.org/10.1094/PDIS-93-11-1123
Wicker E, Grassart L, Coranson-Beaudu R, Mian D, Guilbaud C, Fegan M, Prior P (2007) Ralstonia solanacearum strains from Martinique (French West Indies) exhibiting a new pathogenic potential. Appl Environ Microbiol 73(21):6790–6801. https://doi.org/10.1128/aem.00841-07
Winstead NN, Kelman A (1952) Inoculation techniques for evaluating to Pseudomonas solanacearum. Phytopathology 42:628–634
Yuliar, Nion YA, Toyota K (2015) Recent trends in control methods for bacterial wilt diseases caused by Ralstonia Solanacearum. Microbes Environ 30:1–11. https://doi.org/10.1264/jsme2.ME14144
Acknowledgments
Funding for this research was provided by the WorldVeg Innovation Fund program and the strategic long-term donors to the World Vegetable Center: Republic of China (Taiwan), UK aid from the UK government, United States Agency for International Development (USAID), Australian Center for International Agricultural Research (ACIAR), Germany, Thailand, Philippines, Korea and Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bihon, W., Chen, JR. & Kenyon, L. Identification and characterization of Ralstonia spp. causing bacterial wilt disease of vegetables in Mali. J Plant Pathol 102, 1029–1039 (2020). https://doi.org/10.1007/s42161-020-00631-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42161-020-00631-1