Abstract
Mechanical force regulates a variety of cellular functions through inducing modulations in nuclear chromatin structures and epigenetic landscapes (outside in). The epigenetic modifications, in turn, regulate gene expressions and affect phenotypic outcomes, including cytoskeletal organization and cell–cell/cell–ECM interactions (inside out). While there have been significant advances in the understanding of mechanotransduction in the nucleus, there is still a lack of knowledge on the potential mechanisms through which mechanical cues affect epigenetic and chromatin regulations to determine genetic outcomes. This review firstly focuses on the current understanding of epigenetic regulations and then summarizes how mechanotransduction and epigenetic modification couple together to regulate molecular and cellular functions, eventually causing functional phenotype changes e.g., diseases. Lastly, we introduce related technologies for mechanistic studies, particularly fluorescence resonance energy transfer (FRET) biosensors for the visualization of dynamic epigenetic regulations in single living cells, as well as the applications of FRET biosensors to visualize mechanotransduction events occurring in the nucleus. These studies could provide new insights into epigenetics in regulating the physiological and pathological processes in living cells under different mechanical environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Epigenetics, changing gene expression without altering the DNA sequence, plays an essential role in governing genomic regulation and ultimately cell fates (Li et al. 2012). It has been well documented that external mechanical force can be transmitted from the cell membrane through the cytoplasm to the nucleus, which can induce modulations in nuclear chromatin structures and epigenetic landscapes to impact gene transcription (Irianto et al. 2016; Kelkhoff et al. 2016; Peng et al. 2016; Wang et al. 2016). However, it is still largely unclear how epigenetics is regulated spatiotemporally in relation to genomics. Due to cell–cell heterogeneity, conventional ChIP-Seq technology is insufficient for measuring the dynamic nature of epigenetic signals crucial for cellular function in single cells (Li et al. 2012). Genetically encoded biosensors based on fluorescent proteins (FPs) and fluorescence resonance energy transfer (FRET) could be powerful tools for single live cell imaging to elucidate the mechano-chemical mechanisms underlying the adaptive epigenetic response and cell fate to understand nuclear mechanotransduction.
Epigenetic regulation and mechanobiology in the nucleus
Chromatin
Chromatins are highly ordered nuclear structures that contain DNA, histones, and other chromosomal proteins. The fundamental unit of chromatin is the nucleosome, which is wrapped by 147 base pairs of genomic DNAs. The nucleosome consists of two copies of the four core histones (H): H2A, H2B, H3, and H4, which are tied together by the linker histone H1. Each histone within the nucleosome contains a flexible N-terminus tail (Black et al. 2012; Kornberg 1974; Kornberg and Thomas 1974).
Epigenetic regulation of the chromatin is often achieved through the modulation of nucleosomes. It is the ensemble of mechanisms of concurrent chromatin modification that modulates global manners without affecting the DNA sequence itself. Such epigenetic regulations mainly include DNA methylation (Laird 2003), histone post-translational modifications (PTMs) (Jenuwein 2001), chromatin remodeling (Zhang and Reinberg 2001), and non-coding RNAs (ncRNAs); all these regulations exert their various functions in a coordinated manner (Allis and Jenuwein 2016). DNA methylation occurs on nucleotide cytosine next to guanine (CpG) to regulate the transition of chromatin states. During DNA methylation, DNA methyltransferases (DNMTs) aid the covalent addition of a methyl group from S-adenosylmethionine to the 5′ position of cytosine (Yu et al. 2011). DNA methylation is known to affect the formation of constitutive heterochromatin. Histone PTMs are also important for chromatin geometry and gene expression (Inaba et al. 2015). Histone tails can combine and undergo various PTMs, allowing regulatory proteins to access the genome dynamically (Önder et al. 2015). In addition to histone PTMs, chromatin remodeling can be achieved by ATP-dependent chromatin-remodeling complexes, during which ATP hydrolysis is used to alter the histone–DNA interactions. (Lusser and Kadonaga 2003). On the other hand, nucleosome positioning also contributes to epigenetic regulations. It has been found that nucleosomes assemble in heterogeneous groups of varying sizes interspersed with nucleosome-depleted regions. RNA polymerase II preferentially associates with smaller clutches, while linker histone H1 and heterochromatin are more enriched in larger ones, suggesting that transcription can be activated more preferentially in smaller clutches (Ricci et al. 2015). This review mainly focuses on histone post-translational modifications.
Chromatin and histone PTMs
The first histone modification reported was histone acetylation in the early 1960s (Allfrey et al. 1964). Since then, more than 100 different histone PTMs have been identified and categorized (Abranches et al. 2013; Cota et al. 2013; Goss 2014; Kouzarides 2007; Luger et al. 1997; Tan et al. 2011) (Table 1). Currently, six major PTMs are known to occur on histone tails in cells: acetylation, methylation, phosphorylation, adenosine diphosphate (ADP)-ribosylation, SUMOylation, and ubiquitylation, each with their own distinct functions and regulatory mechanisms.
Histone PTMs are important for the activation state of both promoters and enhancers. Histones are primarily modified on the lysine and arginine residues on histone tails of the H3 and H4 subunits. Histone PTMs have been shown to play central roles in regulating gene expression by modulating chromatin assembly and by coordinating the recruitment of specific readers or other chromatin-modifying or chromatin-remodeling enzymes (Strahl and Allis 2000). Distinct histone modifications are dynamically catalyzed or erased by specific enzymes in response to environmental challenges, while new types of biologically important PTMs, their modifiers, and their readers are continuing to be identified (Andrews et al. 2016).
Histone acetylation is the first reported PTM to be associated with gene activation and has since been identified as the most frequent PTM (Allfrey et al. 1964). Histone acetylation and deacetylation are observed on conserved lysines of the N-terminal tails of all four core histones and are conducted by histone acetyltransferases (HATs) and histone deacetylase (HDACs), respectively. The HATs catalyze the acetylation of specific lysines within histones by transferring an acetyl moiety from the cofactor acetyl-CoA to the lysine, which neutralizes the lysine’s positive charge and potentially weakens the interactions between histones and DNA for chromatin remodeling, a process important to gene expression (Fig. 1). HDACs, on the other hand, catalyze the reverse reaction and are also generally associated with gene repression (Bannister and Kouzarides 2011; Lee and Grant 2019). Histone acetylation has a significant effect on gene expressions. Researchers recently discovered that some cancer cells with increased acetylation levels on H3K18 and H3K27 share the characteristics of truncated acetyltransferase genes EP300 and CREBBP, suggesting a functional consequence of malfunction in epigenetic factors (Ghandi et al. 2019).
Schematic representation of a nucleosome. A nucleosome functions as the fundamental packing unit of chromatin. Different PTMs (mainly acetylation and methylation) at core histones and the processes of these modifications are also shown (He et al. 2018)
Histone methylation mainly occurs in lysines and arginines. Lysine residues can be mono-, di-, or tri-methylated, whereas arginines can be mono-methylated or symmetrically or asymmetrically di-methylated (Fig. 1) (Han et al. 2019). The different levels of methylation greatly increase the complexity of histone modification and regulation of gene expressions (Bannister and Kouzarides 2005, 2011; Bedford and Clarke 2009; Ng et al. 2009; Xhemalce et al. 2011; Han et al. 2019). Histone lysine methylations have been found on a range of lysine residues in various histones, including K4, K9, K27, K36, and K79 residues in histone H3, K20 in histone H4, K59 in the globular domain of histone H4 (Dillon et al. 2005), and K26 in histone H1B (Cai et al. 2010). The methylation level is controlled by enzymes called histone methyltransferases (HMTs) and histone demethylases (HDMs) that possess strong substrate specificity. Histone methylation does not alter the charge of histone proteins, unlike the case with histone acetylation and phosphorylation. Transcriptional activities can be regulated by the methylation of histone tails which function as a recognition motif to recruit effector proteins to local chromatin regions (Cloos et al. 2008). Thus, histone lysine methylation can be associated with either activation or repression of transcription, depending on the functions of the effectors.
Histone phosphorylation is another highly dynamic mark that takes place on serines, threonines, and tyrosines, predominantly in the N-terminal histone tails (Xhemalce et al. 2011). The phosphorylation level is controlled by kinases and phosphatases that add and remove the phosphate group, respectively (Oki et al. 2007). Histone kinases catalyze the addition of a phosphate group from ATP to the hydroxyl group of the target amino acid side chain, which increases the negative charge of histone to affect the chromatin structure (Fig. 2). The specific histone phosphorylation modifications are regulated during different stages of the cell cycle, involving different sets of kinases and phosphatases. For instance, the proteins phosphatase 1 (PP1) can rapidly neutralize the action of Aurora B kinase, which causes a large-scale genome-wide phosphorylation events at H3S10ph and H3S28ph during mitosis (Goto et al. 2002; Sugiyama et al. 2002). However, it is still unclear how the kinases are accurately recruited to the phosphorylation sites on histones at chromatins. Even less is known regarding the roles of histone phosphatases, although it is clear that a high level of phosphatase activity exists in the nucleus to direct a rapid turnover of histone dephosphorylations (Bannister and Kouzarides 2011).
Histone phosphorylation regulation model. The main phosphorylated residues of histones are shown, with the corresponding kinases and phosphatases. Residues in red indicate phosphorylation during mitosis; in green during DNA transcription; in blue during DNA damage; and in light purple during spindle assembly checkpoint (SAC). Histone H2AX is colored in light green (A), H2A in dark green (B), H2B in purple, H3 in blue, H4 in pink, and CENP-A in light blue (C). Phosphatases involved are highlighted in orange. RM: Repo-man; PP1, PP2A, PP5, and PP6: protein phosphatase 1, 2A, 5, 6, respectively. ATM: ataxia-telangiectasia mutated; ATR: ataxia-telangiectasia and Rad3-related protein; VRK: vaccinia-related protein kinases; JAK: Janus kinase; PKC: protein kinase C; Dlk: death-associated protein (DAP)-like kinase; PRK: phosphoribulokinase; CKII: casein kinase II (Gil and Vagnarelli 2019)
Among the different HMTs, SUV39H1 is the first identified histone lysine methyltransferase (HKMT) which targets H3K9 for tri-methylation (Bannister and Kouzarides 2005). Many pieces of evidence suggest a direct role of histone H3K9 methylation as a histone marker, which is positively correlated with DNA methylation and participated in repressive heterochromatin formation in tumorigenesis. In the cell cycle, H3K9me3 is also important for HP1 recruitment to regulate gene expression, chromatin packaging, and heterochromatin formation (Dormann et al. 2006). However, the dynamics of H3K9me3 remains controversial, mainly due to the lack of the appropriate tool for H3K9me3 detection in living cells.
The interactions between different histone chemical modifications show that the same modification of different histone residues can modify the occurrence of other modifications. This process is also regulated by other histone residues. On the other hand, different modifications of the same amino acid residue on histones may be synergistic or antagonistic at the same time. The interaction between different modifications of the same histone is called cis-effect, and the interaction between modifications of different histones is called trans-effect.
Epigenetic regulations in relation to the cell cycle
The cell cycle consists of a chain of inter-connected events, during which DNA is accurately duplicated, and chromosomes are segregated into two genetically identical daughter cells. These events outline the two major phases of the cell cycle: the synthetic phase (S-phase), where DNA replication takes place, and the mitotic phase (M-phase), characterized by chromosomal segregation followed by cellular division. These phases are separated by two gap phases, G1- and G2-phase, respectively, during which cellular growth and checkpoint controls occur. The process of a cell cycle is controlled by chromatin modifiers. Locally, chromatin modifiers control the expression of individual genes; globally, they control chromatin condensation and chromosome segregation (Fig. 3). It is reported that the histone tail modifications on H3 and H4 are required for the normal progression of the cell cycle (Morgan et al. 1991). Indeed, many histone modifications are of fundamental importance across cell types (Ernst et al. 2011).
Histone dynamics across the cell cycle. Histone marks and chromatin-associated proteins are summarized by up- (red) and down- (blue) regulation during each phase of the cell cycle. The time courses of these histone marks within the cell cycle are also represented in the graph (Kheir and Lund 2010)
The S-phase is a crucial stage of the cell cycle where histones and DNA modifications must be deposited correctly to the newly synthesized chromatin. Global methylation levels of the histone H3 and histone H4 tails have been detected to increase in late G1- and S-phase (Bonenfant et al. 2007). H3K79 di-methylation by KMT4/Dot1 is required for efficient entry into the S-phase, which is an example of transcriptional control of cell cycle genes (Schulze et al. 2009). The overexpression of KDM4A, an H3K9me3 demethylase, results in a faster progression through S-phase, possibly due to the better chromatin accessibility increase in replication forks and alteration of replication timing at heterochromatin regions (Black et al. 2012). Conversely, the loss of KDM4A in MDA-MB-231 breast cancer cells leads to a G1/S arrest and decreased proliferation rates (Black et al. 2010). Chromatin microenvironment can also affect the cell cycle indirectly by changing the replication timing. For example, KMT6/EZH2, by regulating epigenetic H3K27me3 levels, can target transcriptional regulations of cell cycle proteins Cyclin D1, E1, and A2 (Black and Whetstine 2011; Bracken et al. 2003). H3K9Ac, which peaks during G1-phase, has been reported to remain high during S-phase together with H4K16Ac (McManus and Hendzel 2006; Rice 2002). In contrast, H4K20 and H3K79 methylations were not detected on newly deposited histones until later time points in the cell cycle (Scharf et al. 2009).
The G2-phase of the cell cycle is a short phase marked by significant protein synthesis. Histone PTMs often maintain across the G2-phase and the following M-phase. During mitosis, cell growth is halted, and cellular energy is focused on the division into two daughter cells. A large number of transcription factors are displaced from the chromatin and general transcription is also suppressed (Egli et al. 2008; Gottesfeld and Forbes 1997). Histone modifications are important for chromatin condensation and chromosomal segregation during this phase in several cell types (Liu et al. 2017; Stephens et al. 2018). Histone phosphorylation is upregulated during M-phase and linked to chromatin condensation, such as H3T3p, H3T11p, H3.3S31p, H2AS1p, and H4S1p (Bonenfant et al. 2007; Kang et al. 2007). Transcription is believed to be turned off during mitosis, which is in accordance with observed global deacetylation of histones. A dramatic decrease in acetylation has been reported on histones H3K9, H3K18, H3K23, H4K5, H4K8, H4K12, and H4K16 during this stage (Bonenfant et al. 2007; McManus and Hendzel 2006; Rice 2002). Lysine deacetylation has also been observed on histone H2A and H2B, in particular, H2AK5, H2BK12, H2BK15, and H2BK20. In addition to its role in transcriptional repression, lysine deacetylation is thought to ensure a correct packaging of nucleosomes into metaphase chromosomes (Bonenfant et al. 2007). This explanation is further supported by the observed destabilization of the chromatin fiber and the decondensation of the chromatin upon the induction of multiple events of acetylation on the histone tails (Kruhlak et al. 2001). Shogren-Knaak et al. showed that H4K16Ac, in particular, is responsible for chromatin decondensation (Shogren-Knaak 2006). In addition, SirT2, an HDAC specific for H4K16, has been shown to be highly expressed and associated with condensed chromatin exclusively during mitosis (Vaquero 2006).
The low levels of transcriptional activity during the mitotic phase is also evident from the histone methylation patterns. Lys20 methylation on histone H4 is generally increased, and its high level is correlated with the deacetylation of H4K5, H4K8, and H4K12 and, in particular, H4K16 observed during M-phase (Bonenfant et al. 2007; Rice 2002). Other notable methylation dynamics during G2/M-phase occur on H3K17, H3K79, and H4R3 (Bonenfant et al. 2007; Pesavento et al. 2008; Rice 2002). H3K9me3 was identified as a marker of heterochromatin and epigenetic silencing regions, which typically are located close to the centromeres (Schotta et al. 2004; Stewart et al. 2005). Accordingly, H3K9me3 is dynamically regulated in a cell cycle-dependent manner (Duan et al. 2008; McManus et al. 2006; Park et al. 2011). H3K9me3 is increased rapidly in G2 to reach a maximum, followed by a quick decline (McManus et al. 2006). It is also proposed that an increase in H3K9me3 at the late G2-phase and early mitosis may be needed to stabilize the pericentric heterochromatin so that the centromeres and kinetochores have a rigid structure required for proper tension transmission to the inner centromere (Heit et al. 2009). However, this hypothesized scenario is in contrast to the results of the genome-wise dynamic change of H3K9me3 in the overall cell cycle (Black et al. 2012). Another recent study further revealed an association between chromatin and a lysine demethylase KDM4C during mitosis, which is accompanied by a decrease in the mitotic levels of H3K9me3 (Kupershmit et al. 2014). A most recent study further proved that the histone methylations and their combinations could serve as codes to determine the overall gene expressions and phenotypic outcomes. In particular, the coordinated regulation of H3S10p and H3K9me3 may facilitate the timely dissolution of heterochromatin-like structures and allow accessibility by chromatin-remodeling complex for an efficient chromatin reorganization during mitosis (Peng et al. 2018).
Epigenetic regulation in mechanobiology
Cells in our body are constantly exposed to a spectrum of mechanical forces, such as blood flow-induced fluid shear stress, compression, stretches, differential tissue rigidity, and strain. Those forces modulate gene expressions and cellular functions to influence and govern physiology and pathophysiology in health and disease. Epigenetic modifications play crucial roles in these gene expressions to not only respond to the external mechanical cues but also to allow cells to adapt to differential mechanical environments (Chen et al. 2013; Miroshnikova et al. 2017). Here, we reviewed current understanding of how mechanical and epigenetic modifications couple together to regulate molecular and cellular functions.
One current understanding is that mechanical stress directly induces chromatin remodeling through epigenetic modifications to alter gene expressions and cellular functions. For example, shear stress (SS) is defined as the frictional force generated by blood flow in the endothelium. Endothelial cells (ECs) sense the changes of SS and activate intracellular signaling pathways, leading to the transcription of specific genes. Those genes include mechanosensory complexes of platelet-endothelial cell adhesion molecule 1 (PECAM-1) which directly transmits mechanical force, vascular endothelial cadherin (VE-cadherin) which functions as an adaptor, and vascular endothelial growth factor receptor 2 (VEGFR2) which activates phosphatidylinositol-3-OH kinase (Tzima et al. 2005). The expression of those important genes is regulated by the modification of chromatin structure, which depends on the activity of histone modification enzymes, including histone acetyltransferase (HAT) and histone deacetylase (HDAC) (Jenuwein 2001). HDACs are sensitive to hemodynamic forces and are mainly divided into three groups: Class I (HDAC-1/2/3 and HDAC-8), Class II (HDAC-4/5/6/7 and HDAC-9/10), and Class III sirtuins (SIRT) (Chen et al. 2013). Among the HDACs, it is worth noting that Class III HDACs play a protective role in atherosclerosis (Stein and Matter 2011). Besides, eNOS plays a key role in vascular wall homeostasis and regulation of vasomotor tone. SS can also remodel chromatin structure particularly on histone H3 and H4 by altering the state of enzymes like phosphatidylinositol 3-kinase and HDAC (Class II and III), resulting in modulating the endothelial nitric oxide synthase (eNOS) gene expression at the transcriptional level (Chen et al. 2010; Fish et al. 2005; Illi et al. 2003, 2008). Further research indicates that SS can increase H3K27ac level to open up chromatin and subsequently activate the inositol 1,4,5-trisphosphate receptor 3 (ITPR3)–eNOS axis in Ecs via krüppel-like factor 4(KLF4)-regulated ITPR3 transcription, which is a principal mechanosensitive cue of atheroma protective flow (He et al. 2019). Coupling shear stress and epigenetic modifications together to regulate vascular marker gene expression eventually can affect phenotypes (Gomez et al. 2015). For example, mechano-induced DNA demethylase ten-eleven translocation-2 (TET2) expression can modulate vascular tone by guiding the phenotype of vascular smooth muscle cells (VSMCs) between contractile and expansible (Liu et al. 2013). The blood pressure (BP) can, subsequently, be changed by the phenotype of vascular VSMCs (Brozovich et al. 2016). A trans-ancestry genome-wide study shows 12 genetic loci influencing blood pressure and implicates a role for DNA methylation, with four of them (IGFBP3, KCNK3, PDE3A and PRDM6) related to the vascular smooth muscle function (Kato et al. 2015; Liu et al. 2015). An array of cardiovascular diseases associated with epigenetic modifications caused by mechanotransduction are listed in Table 2 (Bauer and Martin 2017; Haberland et al. 2009; Koentges et al. 2016; Kumar et al. 2013; Lee et al. 2012; Liu et al. 2013; Nakamura and Sadoshima 2018; Ohtani et al. 2011; Oka et al. 2011; Shao et al. 2017; Shpargel et al. 2014; Zhang et al. 2011; Zheng et al. 2011; Zhu et al. 2016; Zhuang et al. 2017).
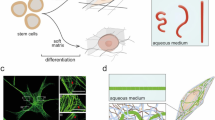
Another current understanding is that mechanical force mediated by nuclear lamina can indirectly organize chromosome through epigenetic modifications to regulate gene expression. Lamin A/C is a protein network that connects the linker of nucleoskeleton and cytoskeleton (LINC) complex to chromosomes. As a mechanosensory structure, it can respond to mechanical forces and regulate gene expressions through epigenetic modifications (Chowdhury et al. 2010; DuFort et al. 2011). Previous reports show that stem cell differentiation into bone on stiff matrix and into fat on soft matrix can be enhanced by the high and low lamin-A levels corresponding to high and low H3K9m3 levels on chromosomes, respectively (Swift et al. 2013). This result is also confirmed in melanoma cells. People found that soft fibrin matrices indeed promote H3K9 demethylation and increase Sox2 expression and self-renewal, while stiff fibrin has an opposite effect (Tan et al. 2014). A recent study demonstrated how mechanical force stretches chromatin to affect transcription. A green fluorescent protein (GFP)-tagged bacterial chromosome dihydrofolate reductase (DHFR) and an Arg-Gly-Asp-coated magnetic bead were used to demonstrate that local stresses can be propagated from the tensed actin cytoskeleton to the LINC complex via integrins and then stretch chromatin and upregulate transcription via lamina-chromatin interaction (Tajik et al. 2016). So, mutations in lamins can result in a wide array of pathologies, collectively referred to as laminopathies. Such diseases include Emery-Dreifuss muscular dystrophy, limb-girdle muscular dystrophy, dilated cardiomyopathy, familial partial lipodystrophy, Charcot-Marie-Tooth, and the accelerated aging disorder Hutchinson-Gilford progeria syndrome (HGPS) (Schreiber and Kennedy 2013). In fact, HGPS is caused by progerin which is a truncated version of the lamin-A protein. However, progerin remains permanently farnesylated and carboxymethylated since it lacks the second site for endoproteolytic cleavage. The accumulation of progerin leads to altered H3K27me3 through the downregulation of enhancer of zeste homolog 2 gene (EZH2) and disrupts heterochromatin–lamina interactions, which eventually lead to cellular and organismal decline (McCord et al. 2013). As a result, the senescence in HGPS may be delayed if the post-translational modifications of progerin are changed. Further research also showed that, blocking farnesylation of progerin (Gonzalo et al. 2017), via prenylation inhibitors (Moiseeva et al. 2016; Varela et al. 2008) and isoprenyl cysteine carboxyl methyltransferase inhibitors (Ibrahim et al. 2013), can lead to effective therapeutic treatment of HGPS. Another reported disease connected to lamin-A is Dunnigan-type lipodystrophy, namely, familial partial lipodystrophy type 2 (FPLD2), which can be caused by the R482W mutation on lamin-A, preventing lamin-A’s binding at MIR335 locus. This locus determines the aberrant transcription of the anti-adipogenic miR-335 with subsequent inhibition of adipogenic differentiation (Oldenburg et al. 2017). Lamina-associated domains (LADs) are redistributed for LMNA mutations in association with markedly altered CpG methylation and gene expression. As such, LADs can contribute to the pathogenesis of dilated cardiomyopathy (DCM) through genomic alterations and epigenetic regulation (Cheedipudi et al. 2019).
FRET imaging of epigenetic regulation in single living cells
Mechanical cues can serve as potent regulators of epigenetics, gene expression, and cell fate. In order to bridge mechanical regulation with epigenetic regulation at molecular levels, people have developed technologies to study the dynamic manipulation and regulation. For example, Guohong Li’s group has reported that the tetranucleosomes-on-a-string appears as a stable secondary structure during hierarchical organization of chromatin fibers using single-molecule magnetic tweezers and force spectroscopy. This stability is enhanced by histone chaperone facilitates chromatin transcription (FACT) in vitro (Li et al. 2016). They further reported that FACT displays dual functions in destabilizing the nucleosome and maintaining the original histones and nucleosome integrity at the single-nucleosome level (Chen et al. 2018). As we mentioned in “Epigenetic regulation in mechanobiology” section, Ning Wang’s group used 3D magnetic twisting cytometry to apply local stresses on the cell surface via an Arg-Gly-Asp-coated magnetic bead to deform chromatin as well as force-induced gene expression in a living cell (Tajik et al. 2016). Cheng Zhu’s group developed a DNA-origami tension probe (DOTP) that maps the piconewton forces generated by living cells at the nanometer-length scale (Dutta et al. 2018). However, the molecular pathways and mechanisms for mechanical and epigenetic coupled regulations remain unclear, requiring further studies using more advanced technologies like FRET epigenetic biosensors.
FRET imaging
Understanding the molecular basis of physiological regulations is crucial to understanding the pathways of pathological events to find treatments. However, many detection methods like genomic sequencing lack spatial and temporal resolutions (Wang and Wang 2009). Furthermore, those methods typically provide little information on epigenetic modifications. As a result, an imaging method that can simultaneously provide both spatial and temporal information is much needed. Fluorescence resonance energy transfer (FRET) emerged as a recent advance in molecular detection, offering high spatiotemporal resolutions (Wang et al. 2008).
FRET technique uses a pair of fluorescence proteins (FP) to detect molecular signals in organisms. The first discovered and isolated FP is the GFP from aequorin (Shimomura et al. 1962), after which numerous other natural and engineered FPs were discovered (Tsien 2005). Among those FPs, cyan-colored ECFP and yellow-colored EYFP and its various derivatives are believed to be the best FRET pairs (Miyawaki et al. 1997). There are three main types of FRET biosensors. In the first type, a donor FP and an acceptor FP are pulled by two interacting sensing molecules (Fig. 4A). When activated, the two sensing molecules bind together to change the distance and orientation of FPs, resulting in a FRET signal change. In the second type, only one sensing molecule is present (Fig. 4B). This sensing molecule can change conformation when activated, causing FRET change. FRET biosensors with two interacting molecules fused separately with the donor and acceptor FPs are also possible (Fig. 4C). The interaction of the two sensing domains can bring two FPs closer in the distance, resulting in a signal which measures this interaction (Wang et al. 2008). Three main factors influencing the efficiency of FRET imaging are the distance between FPs, the relative orientation between the donor and acceptor FPs, and the excitation/emission spectrum of FPs. High FRET efficiency requires close distance, correct orientation, and overlapping between emission wavelengths of donor protein and excitation wavelengths of acceptor protein (Clegg 1995; Wang and Wang 2009).
Three designs of FRET biosensors. A Intramolecular FRET biosensor with two interacting sensing domains. The two sensing domains interact with each other upon activation, resulting in FRET. B Intramolecular FRET biosensor with one target sensing domain. The sensing domain changes conformation upon activation, bringing the two FPs into close distance and correct orientation. C Intermolecular FRET biosensor. One molecule is fused to donor FP, and the other to the acceptor FP. Upon activation, two molecules interact with each other to induce FRET changes
One application of a FRET biosensor is to monitor kinase activity in living cells. For example, biosensors using FRET can monitor the mechanical activation of Src (Wang et al. 2005). This biosensor consists of an N-terminal ECFP, an SH2 domain from Src kinase, a flexible linker, a substrate-binding domain derived from p130cas which is sensitive to Src phosphorylation, and a C-terminal citrine (a variant of YFP) (Wang et al. 2005, 2008; Wang and Wang 2009). Recently, improved FRET biosensors with enhanced specificity are being developed. For example, a biosensor that monitors the activation of Fyn, a member of the Src family, is reported. This biosensor used a similar design with previously reported Src biosensor and was altered only in the substrate peptide derived from p34cdc2. In vitro kinase assays suggest that this biosensor has a clear preference of the activation of Fyn over other Src family kinases like Src, Yes, and Abl (Ouyang et al. 2019). Other enzymatic activities besides Src family kinases can also be detected using FRET biosensors. For example, using a FRET biosensor, Pan et al. reported that stronger EphA4 activation might occur in non-raft regions than raft regions on the plasma membrane (Pan et al. 2019).
FRET-based epigenetic biosensors
Recently, FRET-based biosensors capable of monitoring histone epigenetic modifications have been developed to study the effect of epigenetic events on cell fate. Lin and Ting first developed a histone H3S28 phosphorylation biosensor (Lin and Ting 2004), and then, Lin et al. reported a histone H3K9me3 and H3K27me3 biosensor which can visualize the histone methylation both in vitro and in single living cells (Lin et al. 2004). Later, Chu et al. developed another FRET-based and centromere-targeted H3K9me3 biosensor to visualize the methylation dynamics during chromosome segregation (Chu et al. 2012). As for acetylation, Sasaki et al. developed an H4K5 and H4K8 biosensor, followed by an H4K12 biosensor developed by Ito et al. (Ito et al. 2011; Sasaki et al. 2009). Recent studies further reported the monitoring of H3K9ac and H3K14ac by FRET biosensors in 2016 (Nakaoka et al. 2016; Sasaki and Yoshida 2016).
FRET epigenetic biosensors have many applications. For example, by using an H3K9me3 FRET biosensor, it was reported that a low level of H3K9me3 is present in tumor-repopulating cells that are not sensitive to matrix stiffness and applied forces (Tan et al. 2014). Using an H3K9 tri-methylation biosensor together with an H3S10 phosphorylation biosensor in the same cell, Peng et al. found an anticorrelation between H3K9me3 and H3S10p during cell cycles. Further studies revealed that this coordinated regulation might allow increased access of remodeling complexes to the chromatin in preparation of the global reorganization of chromatin during mitosis (Fig. 5) (Peng et al. 2018). To study the dynamic epigenetics in mechano-regulation, epigenetic FRET biosensors can be introduced into endogenous genome to create stable cell lines by using CRISPR/Cas9, such as EC stable cell line. Combining with liveFISH labeling technology (Wang et al. 2019), the transcription and epigenetic dynamics at specific genomic loci in EC stable cell lines can be visualized in real time under shear stress stimulation. As such, epigenetic visualization will provide more mechanistic understanding on mechano-epigenetic regulation in gene expression at single living cell levels.
The designed activation mechanism of H3K9 tri-methylation FRET biosensor. The H3K9me3 tri-methylation biosensor contains a full-length histone H3, an ECFP (donor), a flexible EV linker (120aa), a heterochromatin protein 1 (HP1), and a YPet (acceptor). At rest state, the H3K9me3 biosensor has an open conformation with low FRET. When the histone H3 lysine 9 is methylated by an upstream methyltransferase, such as SUV39H1, HP1 binds to the tri-methylated H3K9, causing a strong FRET. The excitation of ECFP at 433 nm then results in the emission from YPet at 527 nm
Even though several FRET biosensors have been developed to study epigenetic events, the actual usage of the FRET epigenetic biosensor is still limited. The sensitivity of most epigenetic biosensors is typically low comparing to other types of chemical or biological imaging probes (Sasaki et al. 2009; Tan et al. 2014). Some strategies to optimize the FRET biosensor are proposed. For example, peptide scaffold (e.g., monobody)-based directed evolution will be a promising technology to increase the specificity and sensitivity of the FRET biosensors (Limsakul et al. 2018). Tools other than the FRET biosensor are also helpful in understanding the relationship between epigenetics and mechanobiology. For example, methods such as fluorescence correlation spectroscopy (FCS), fluorescence recovery after photobleaching (FRAP), and split fluorescence-activating and absorption shifting tag (splitFAST) are used to monitor protein dynamics (Liu and Irudayaraj 2019; Tebo and Gautier 2019). Should more FRET biosensors with increased specificity and sensitivity become available in the future, mechanobiological studies using FRET biosensors will thrive.
Summary and perspectives
In general, there are multiple concurrent epigenetic modifications involved in cellular processes. These modifications synergistically cooperate to govern cellular functions at different subcellular compartments, e.g., in the nucleus. As such, developing more FRET epigenetic biosensors for the visualization and quantification of histone and other epigenetic modifications is in great need. Simultaneous imaging of two or more histone modifications in the same live cell, together with the associated traction force map, could allow us to generate a spatiotemporal landscape of multiplex histone orchestration in association with the corresponding mechanical stress distribution. Moreover, by combining the CRISPR/dCas9 labeling system, these biosensors should allow the more precise monitoring of spatiotemporal epigenetics at specific genomic loci in response to mechanical force. In summary, the single-cell study of epigenetic regulation should not only advance our precise understanding of the dynamic coordination of epigenetics and genomic regulations, but also identify new therapeutic targets in response to mechanical cues in diseases.
References
Abranches E, Bekman E, Henrique D (2013) Generation and characterization of a novel mouse embryonic stem cell line with a dynamic reporter of nanog expression. PLoS ONE 8:e59928
Allfrey VG, Faulkner R, Mirsky AE (1964) Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci USA 51:786–794
Allis CD, Jenuwein T (2016) The molecular hallmarks of epigenetic control. Nat Rev Genet 17:487–500
Andrews FH, Strahl BD, Kutateladze TG (2016) Insights into newly discovered marks and readers of epigenetic information. Nat Chem Biol 12:662–668
Bannister AJ, Kouzarides T (2011) Regulation of chromatin by histone modifications. Cell Res 21:381–395
Bannister AJ, Kouzarides T (2005) Reversing histone methylation. Nature 436:1103–1106
Bauer AJ, Martin KA (2017) Coordinating regulation of gene expression in cardiovascular disease: interactions between chromatin modifiers and transcription factors. Front Cardiovasc Med. https://doi.org/10.3389/fcvm.2017.00019
Bedford MT, Clarke SG (2009) Protein arginine methylation in mammals: who, what, and why. Mol Cell 33:1–13
Black JC, Whetstine JR (2011) Chromatin landscape: methylation beyond transcription. Epigenetics 6:9–15
Black JC, Allen A, van Rechem C, Forbes E, Longworth M, Tschöp K, Rinehart C, Quiton J, Walsh R, Smallwood A, Dyson NJ, Whetstine JR (2010) Conserved antagonism between JMJD2A/KDM4A and HP1γ during cell cycle progression. Mol Cell 40:736–748
Black JC, van Rechem C, Whetstine JR (2012) Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol Cell 48:491–507
Bonenfant D, Towbin H, Coulot M, Schindler P, Mueller DR, van Oostrum J (2007) Analysis of dynamic changes in post-translational modifications of human histones during cell cycle by mass spectrometry. Mol Cell Proteomics 6:1917–1932
Bracken AP, Pasini D, Capra M, Prosperini E, Colli E, Helin K (2003) EZH2 is downstream of the pRB-E2F pathway, essential for proliferation and amplified in cancer. EMBO J 22:5323–5335
Brozovich FV, Nicholson CJ, Degen CV, Gao YZ, Aggarwal M, Morgan KG (2016) Mechanisms of vascular smooth muscle contraction and the basis for pharmacologic treatment of smooth muscle disorders. Pharmacol Rev 68:476–532
Cai Y, Jin J, Swanson SK, Cole MD, Choi SH, Florens L, Washburn MP, Conaway JW, Conaway RC (2010) Subunit composition and substrate specificity of a MOF-containing histone acetyltransferase distinct from the male-specific lethal (MSL) complex. J Biol Chem 285:4268–4272
Cheedipudi SM, Matkovich SJ, Coarfa C, Hu X, Robertson MJ, Sweet M, Taylor M, Mestroni L, Cleveland J, Willerson JT, Gurha P, Marian AJ (2019) Genomic reorganization of lamin-associated domains in cardiac myocytes is associated with differential gene expression and DNA methylation in human dilated cardiomyopathy. Circ Res 124:1198–1213
Chen Z, Peng I-C, Cui X, Li YS, Chien S, Shyy JY (2010) Shear stress, SIRT1, and vascular homeostasis. Proc Natl Acad Sci USA 107:10268–10273
Chen L-J, Wei S-Y, Chiu J-J (2013) Mechanical regulation of epigenetics in vascular biology and pathobiology. J Cell Mol Med 17:437–448
Chen P, Dong L, Hu M, Wang YZ, Xiao X, Zhao Z, Yan J, Wang PY, Reinberg D, Li M, Li W, Li G (2018) Functions of FACT in breaking the nucleosome and maintaining its integrity at the single-nucleosome level. Mol Cell 71(2):284–293
Chowdhury F, Na S, Li D, Poh YC, Tanaka TS, Wang F, Wang N (2010) Material properties of the cell dictate stress-induced spreading and differentiation in embryonic stem cells. Nat Mater 9:82–88
Chu L, Zhu T, Liu X, Yu R, Bacanamwo M, Dou Z, Chu Y, Zou H, Gibbons GH, Wang D, Ding X, Yao X (2012) SUV39H1 orchestrates temporal dynamics of centromeric methylation essential for faithful chromosome segregation in mitosis. J Mol Cell Biol 4:331–340
Clegg RM (1995) Fluorescence resonance energy transfer. Curr Opin Biotechnol 6:103–110
Cloos PAC, Christensen J, Agger K, Helin K (2008) Erasing the methyl mark: histone demethylases at the center of cellular differentiation and disease. Genes Dev 22:1115–1140
Cota P, Shafa M, Rancourt ED (2013) Stem cells and epigenetic reprogramming. In: Bhartiya D (ed) Pluripotent stem cells. InTech, London
Dillon SC, Zhang X, Trievel RC, Cheng X (2005) The SET-domain protein superfamily: protein lysine methyltransferases. Genome Biol 6:227. https://doi.org/10.1186/gb-2005-6-8-227
Dormann HL, Tseng BS, Allis CD, Funabiki H, Fischle W (2006) Dynamic regulation of effector protein binding to histone modifications: the biology of HP1 switching. Cell Cycle 5:2842–2851
Duan Q, Chen H, Costa M, Dai W (2008) Phosphorylation of H3S10 blocks the access of H3K9 by specific antibodies and histone methyltransferase. Implication in regulating chromatin dynamics and epigenetic inheritance during mitosis. J Biol Chem 283:33585–33590
DuFort CC, Paszek MJ, Weaver VM (2011) Balancing forces: architectural control of mechanotransduction. Nat Rev Mol Cell Biol 12:308–319
Dutta PK, Zhang Y, Blanchard AT, Ge C, Rushdi M, Weiss K, Zhu C, Ke Y, Salaita K (2018) Programmable multivalent DNA-origami tension probes for reporting cellular traction forces. Nano Lett 18:4803–4811
Egli D, Birkhoff G, Eggan K (2008) Mediators of reprogramming: transcription factors and transitions through mitosis. Nat Rev Mol Cell Biol 9:505–516
Ernst J, Kheradpour P, Mikkelsen TS, Shoresh N, Ward LD, Epstein CB, Zhang X, Wang L, Issner R, Coyne M, Ku M, Durham T, Kellis M, Bernstein BE (2011) Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473:43–49
Fish JE, Matouk CC, Rachlis A, Lin S, Tai SC, D'Abreo C, Marsden PA (2005) The expression of endothelial nitric-oxide synthase is controlled by a cell-specific histone code. J Biol Chem 280:24824–24838
Ghandi M, Huang FW, Jané-Valbuena J, Kryukov GV, Lo CC, McDonald ER 3rd, Barretina J, Gelfand ET, Bielski CM, Li H, Hu K, Andreev-Drakhlin AY, Kim J, Hess JM, Haas BJ, Aguet F, Weir BA, Rothberg MV, Paolella BR, Lawrence MS, Akbani R, Lu Y, Tiv HL, Gokhale PC, de Weck A, Mansour AA, Oh C, Shih J, Hadi K, Rosen Y, Bistline J, Venkatesan K, Reddy A, Sonkin D, Liu M, Lehar J, Korn JM, Porter DA, Jones MD, Golji J, Caponigro G, Taylor JE, Dunning CM, Creech AL, Warren AC, McFarland JM, Zamanighomi M, Kauffmann A, Stransky N, Imielinski M, Maruvka YE, Cherniack AD, Tsherniak A, Vazquez F, Jaffe JD, Lane AA, Weinstock DM, Johannessen CM, Morrissey MP, Stegmeier F, Schlegel R, Hahn WC, Getz G, Mills GB, Boehm JS, Golub TR, Garraway LA, Sellers WR (2019) Next-generation characterization of the cancer cell line encyclopedia. Nature 569:503–508
Gil RS, Vagnarelli P (2019) Protein phosphatases in chromatin structure and function. Biochim Biophys Acta BBA 1866:90–101
Gomez D, Swiatlowska P, Owens GK (2015) Epigenetic control of smooth muscle cell identity and lineage memory. Arterioscler Thromb Vasc Biol 35:2508–2516
Gonzalo S, Kreienkamp R, Askjaer P (2017) Hutchinson-Gilford Progeria Syndrome: a premature aging disease caused by LMNA gene mutations. Ageing Res Rev 33:18–29
Goss HM (2014) Histone modifications across the cell cycle in undifferentiated and differentiating mouse embryonic stem cells. PhD, University of Birmingham
Goto H, Yasui Y, Nigg EA, Inagaki M (2002) Aurora-B phosphorylates Histone H3 at serine28 with regard to the mitotic chromosome condensation. Genes Cells Devoted Mol Cell Mech 7:11–17
Gottesfeld JM, Forbes DJ (1997) Mitotic repression of the transcriptional machinery. Trends Biochem Sci 22:197–202
Haberland M, Montgomery RL, Olson EN (2009) The many roles of histone deacetylases in development and physiology: implications for disease and therapy. Nat Rev Genet 10:32–42
Han D, Huang M, Wang T, Li Z, Chen Y, Liu C, Lei Z, Chu X (2019) Lysine methylation of transcription factors in cancer. Cell Death Dis 10:290. https://doi.org/10.1038/s41419-019-1524-2
He H, Hu Z, Xiao H, Zhou F, Yang B (2018) The tale of histone modifications and its role in multiple sclerosis. Hum Genomics 12:31. https://doi.org/10.1186/s40246-018-0163-5
He M, Huang T-S, Li S, Hong HC, Chen Z, Martin M, Zhou X, Huang HY, Su SH, Zhang J, Wang WT, Kang J, Huang HD, Zhang J, Chien S, Shyy JY (2019) Atheroprotective flow upregulates ITPR3 (inositol 1,4,5-trisphosphate receptor 3) in vascular endothelium via KLF4 (Krüppel-like factor 4)-mediated histone modifications. Arterioscler Thromb Vasc Biol 39:902–914
Heit R, Rattner JB, Chan GKT, Hendzel MJ (2009) G2 histone methylation is required for the proper segregation of chromosomes. J Cell Sci 122:2957–2968
Ibrahim MX, Sayin VI, Akula MK, Liu M, Fong LG, Young SG, Bergo MO (2013) Targeting isoprenylcysteine methylation ameliorates disease in a mouse model of progeria. Science 340:1330–1333
Illi B, Nanni S, Scopece A, Farsetti A, Biglioli P, Capogrossi MC, Gaetano C (2003) Shear stress-mediated chromatin remodeling provides molecular basis for flow-dependent regulation of gene expression. Circ Res 93:155–161
Illi B, Russo Claudio D, Colussi C, Rosati J, Pallaoro M, Spallotta F, Rotili D, Valente S, Ragone G, Martelli F, Biglioli P, Steinkuhler C, Gallinari P, Mai A, Capogrossi MC, Gaetano C (2008) Nitric oxide modulates chromatin folding in human endothelial cells via protein phosphatase 2A activation and class II histone deacetylases nuclear shuttling. Circ Res 102:51–58
Inaba H, Tsukagoshi A, Kida S (2015) PARP-1 activity is required for the reconsolidation and extinction of contextual fear memory. Mol Brain 8(1):63. https://doi.org/10.1186/s13041-015-0153-7
Irianto J, Ivanovska IL, Swift J, Discher DE (2016) The nuclear lamina: from mechanosensing in differentiation to cancer cell migration. In: Chien S, Engler AJ, Wang PY (eds) Molecular and cellular mechanobiology. Springer, New York, pp 175–195
Ito T, Umehara T, Sasaki K, Nakamura Y, Nishino N, Terada T, Shirouzu M, Padmanabhan B, Yokoyama S, Ito A, Yoshida M (2011) Real-time imaging of histone H4K12-specific acetylation determines the modes of action of histone deacetylase and bromodomain inhibitors. Chem Biol 18:495–507
Jenuwein T (2001) Translating the histone code. Science 293:1074–1080
Kang T-H, Park D-Y, Choi YH, Kim KJ, Yoon HS, Kim KT (2007) Mitotic histone H3 phosphorylation by vaccinia-related kinase 1 in mammalian cells. Mol Cell Biol 27:8533–8546
Kato N, Loh M, Takeuchi F, Verweij N, Wang X, Zhang W, Kelly TN, Saleheen D, Lehne B, Leach IM, Drong AW, Abbott J, Wahl S, Tan ST, Scott WR, Campanella G, , Chadeau-Hyam M, Afzal U, Ahluwalia TS, Bonder MJ, Chen P, Dehghan A, Edwards TL, Esko T, Go MJ, Harris SE, Hartiala J, Kasela S, Kasturiratne A, Khor CC, Kleber ME, Li H, Yu Mok Z, Nakatochi M, Sapari NS, Saxena R, Stewart AFR, Stolk L, Tabara Y, Teh AL, Wu Y, Wu JY, Zhang Y, Aits I, Da Silva Couto Alves A, Das S, Dorajoo R, Hopewell JC, Kim YK, Koivula RW, Luan J, Lyytikäinen LP, Nguyen QN, Pereira MA, Postmus I, Raitakari OT, Scannell Bryan M, Scott RA, Sorice R, Tragante V, Traglia M, White J, Yamamoto K, Zhang Y, Adair LS, Ahmed A, Akiyama K, Asif R, Aung T, Barroso I, Bjonnes A, Braun TR, Cai H, Chang LC, Chen CH, Cheng CY, Chong YS, Collins R, Courtney R, Davies G, Delgado G, Do LD, Doevendans PA, Gansevoort RT, Gao YT, Grammer TB, Grarup N, Grewal J, Gu D, Wander GS, Hartikainen AL, Hazen SL, He J, Heng CK, Hixson JE, Hofman A, Hsu C, Huang W, Husemoen LLN, Hwang JY, Ichihara S, Igase M, Isono M, Justesen JM, Katsuya T, Kibriya MG, Kim YJ, Kishimoto M, Koh WP, Kohara K, Kumari M, Kwek K, Lee NR, Lee J, Liao J, Lieb W, Liewald DCM, Matsubara T, Matsushita Y, Meitinger T, Mihailov E, Milani L, Mills R, Mononen N, Müller-Nurasyid M, Nabika T, Nakashima E, Ng HK, Nikus K, Nutile T, Ohkubo T, Ohnaka K, Parish S, Paternoster L, Peng H, Peters A, Pham ST, Pinidiyapathirage MJ, Rahman M, Rakugi H, Rolandsson O, Ann Rozario M, Ruggiero D, Sala CF, Sarju R, Shimokawa K, Snieder H, Sparsø T, Spiering W, Starr JM, Stott DJ, Stram DO, Sugiyama T, Szymczak S, Tang WHW, Tong L, Trompet S, Turjanmaa V, Ueshima H, Uitterlinden AG, Umemura S, Vaarasmaki M, van Dam RM, van Gilst WH, van Veldhuisen DJ, Viikari JS, Waldenberger M, Wang Y, Wang A, Wilson R, Wong TY, Xiang YB, Yamaguchi S, Ye X, Young RD, Young TL, Yuan JM, Zhou X, Asselbergs FW, Ciullo M, Clarke R, Deloukas P, Franke A, Franks PW, Franks S, Friedlander Y, Gross MD, Guo Z, Hansen T, Jarvelin MR, Jørgensen T, Jukema JW, Kähönen M, Kajio H, Kivimaki M, Lee JY, Lehtimäki T, Linneberg A, Miki T, Pedersen O, Samani NJ, Sørensen TIA, Takayanagi R, Toniolo D, BIOS-consortium, CARDIo GRAMplusCD, LifeLines Cohort Study, InterAct Consortium, Ahsan H, Allayee H, Chen YT, Danesh J, Deary IJ, Franco OH, Franke L, Heijman BT, Holbrook JD, Isaacs A, Kim BJ, Lin X, Liu J, März W, Metspalu A, Mohlke KL, Sanghera DK, Shu XO, van Meurs JBJ, Vithana E, Wickremasinghe AR, Wijmenga C, Wolffenbuttel BHW, Yokota M, Zheng W, Zhu D, Vineis P, Kyrtopoulos SA, Kleinjans JCS, McCarthy MI, Soong R, Gieger C, Scott J, Teo YY, He J, Elliott P, Tai ES, van der Harst P, Kooner JS, Chambers JC (2015) Trans-ancestry genome-wide association study identifies 12 genetic loci influencing blood pressure and implicates a role for DNA methylation. Nat Genet 47:1282–1293
Kelkhoff D, Downing T, Li S (2016) Mechanotransduction to epigenetic remodeling. In: Chien S, Engler AJ, Wang PY (eds) Molecular and cellular mechanobiology. Springer, New York, pp 163–173
Kheir TB, Lund AH (2010) Epigenetic dynamics across the cell cycle. Essays Biochem 48:107–120
Koentges C, Bode C, Bugger H (2016) SIRT3 in cardiac physiology and disease. Front Cardiovasc Med. https://doi.org/10.3389/fcvm.2016.00038
Kornberg RD (1974) Chromatin structure: a repeating unit of histones and DNA. Science 184:868–871
Kornberg RD, Thomas JO (1974) Chromatin structure; oligomers of the histones. Science 184:865–868
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705
Kruhlak MJ, Hendzel MJ, Fischle W, Bertos NR, Hameed S, Yang XJ, Verdin E, Bazett-Jones DP (2001) Regulation of global acetylation in mitosis through loss of histone acetyltransferases and deacetylases from chromatin. J Biol Chem 276:38307–38319
Kumar A, Kumar S, Vikram A, Hoffman TA, Naqvi A, Lewarchik CM, Kim YR, Irani K (2013) Histone and DNA methylation-mediated epigenetic downregulation of endothelial Kruppel-like factor 2 by low-density lipoprotein cholesterol. Arterioscler Thromb Vasc Biol 33:1936–1942
Kupershmit I, Khoury-Haddad H, Awwad SW, Guttmann-Raviv N, Ayoub N (2014) KDM4C (GASC1) lysine demethylase is associated with mitotic chromatin and regulates chromosome segregation during mitosis. Nucleic Acids Res 42:6168–6182
Laird PW (2003) The power and the promise of DNA methylation markers. Nat Rev Cancer 3:253–266
Lee CY, Grant PA (2019) Role of histone acetylation and acetyltransferases in gene regulation. In: McCullough S, Dolinoy D (eds) Toxicoepigenetics. Elsevier, Cambridge, pp 3–30
Lee S, Lee JW, Lee S-K (2012) UTX, a histone H3-lysine 27 demethylase, acts as a critical switch to activate the cardiac developmental program. Dev Cell 22:25–37
Li M, Liu G-H, Izpisua Belmonte JC (2012) Navigating the epigenetic landscape of pluripotent stem cells. Nat Rev Mol Cell Biol 13:524–535
Li W, Chen P, Yu J, Dong L, Liang D, Feng J, Yan J, Wang PY, Li Q, Zhang Z, Li M, Li G (2016) FACT remodels the tetranucleosomal unit of chromatin fibers for gene transcription. Mol Cell 64:120–133
Limsakul P, Peng Q, Wu Y, Allen ME, Liang J, Remacle AG, Lopez T, Ge X, Kay BK, Zhao H, Strongin AY, Yang XL, Lu S, Wang Y (2018) Directed evolution to engineer monobody for FRET biosensor assembly and imaging at live-cell surface. Cell Chem Biol 25:370–379
Lin C-W, Ting AY (2004) A genetically encoded fluorescent reporter of histone phosphorylation in living cells. Angew Chem Int Ed 43:2940–2943
Lin C-W, Jao CY, Ting AY (2004) Genetically encoded fluorescent reporters of histone methylation in living cells. J Am Chem Soc 126:5982–5983
Liu W, Irudayaraj J (2019) Understanding the dynamics and structure of epigenetic states with single molecule fluorescence microscopy. Curr Opin Biomed Eng. https://doi.org/10.1016/j.cobme.2019.08.010
Liu R, Jin Y, Tang WH, Qin L, Zhang X, Tellides G, Hwa J, Yu J, Martin KA (2013) Ten-eleven translocation-2 (TET2) is a master regulator of smooth muscle cell plasticity. Circulation 128:2047–2057
Liu R, Leslie KL, Martin KA (2015) Epigenetic regulation of smooth muscle cell plasticity. Biochim Biophys Acta 1849:448–453
Liu Y, Chen S, Wang S, Soares F, Fischer M, Meng F, Du Z, Lin C, Meyer C, DeCaprio JA, Brown M, Liu XS, He HH (2017) Transcriptional landscape of the human cell cycle. Proc Natl Acad Sci USA 114:3473–3478
Luger K, Mäder AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389:251–260
Lusser A, Kadonaga JT (2003) Chromatin remodeling by ATP-dependent molecular machines. BioEssays News Rev Mol Cell Dev Biol 25:1192–1200
McCord RP, Nazario-Toole A, Zhang H, Chines PS, Zhan Y, Erdos MR, Collins FS, Dekker J, Cao K (2013) Correlated alterations in genome organization, histone methylation, and DNA–lamin A/C interactions in Hutchinson-Gilford progeria syndrome. Genome Res 23:260–269
McManus KJ, Hendzel MJ (2006) The relationship between histone H3 phosphorylation and acetylation throughout the mammalian cell cycle. This paper is one of a selection of papers published in this Special Issue, entitled 27th International West Coast Chromatin and Chromosome Conference, and has undergone the Journal’s usual peer review process. Biochem Cell Biol 84:640–657
McManus KJ, Biron VL, Heit R, Underhill DA, Hendzel MJ (2006) Dynamic changes in histone H3 lysine 9 methylations: identification of a mitosis-specific function for dynamic methylation in chromosome congression and segregation. J Biol Chem 281:8888–8897
Miroshnikova YA, Nava MM, Wickström SA (2017) Emerging roles of mechanical forces in chromatin regulation. J Cell Sci 130:2243–2250
Miyawaki A, Llopis J, Heim R, McCaffery JM, Adams JA, Ikura M, Tsien RY (1997) Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388:882–887
Moiseeva O, Lopes-Paciencia S, Huot G, Lessard F, Ferbeyre G (2016) Permanent farnesylation of lamin A mutants linked to progeria impairs its phosphorylation at serine 22 during interphase. Aging 8:366–381
Morgan BA, Mittman BA, Smith MM (1991) The highly conserved N-terminal domains of histones H3 and H4 are required for normal cell cycle progression. Mol Cell Biol 11:4111–4120
Nakamura M, Sadoshima J (2018) Mechanisms of physiological and pathological cardiac hypertrophy. Nat Rev Cardiol 15:387–407
Nakaoka S, Sasaki K, Ito A, Nakao Y, Yoshida M (2016) A genetically encoded FRET probe to detect intranucleosomal histone H3K9 or H3K14 acetylation using BRD4, a BET family member. ACS Chem Biol 11:729–733
Ng SS, Yue WW, Oppermann U, Klose RJ (2009) Dynamic protein methylation in chromatin biology. Cell Mol Life Sci CMLS 66:407–422
Ohtani K, Vlachojannis GJ, Koyanagi M, Boeckel J-N, Urbich C, Farcas R, Bonig H, Marquez VE, Zeiher AM, Dimmeler S (2011) Epigenetic regulation of endothelial lineage committed genes in pro-angiogenic hematopoietic and endothelial progenitor cells. Circ Res 109:1219–1229
Oka S, Alcendor R, Zhai P, Park JY, Shao D, Cho J, Yamamoto T, Tian B, Sadoshima J (2011) PPARα-Sirt1 complex mediates cardiac hypertrophy and failure through suppression of the ERR transcriptional pathway. Cell Metab 14:598–611
Oki M, Aihara H, Ito T (2007) Role of histone phosphorylation in chromatin dynamics and its implications in diseases. Subcell Biochem 41:319–336
Oldenburg A, Briand N, Sørensen AL, Cahyani I, Shah A, Moskaug JØ, Collas P (2017) A lipodystrophy-causing lamin A mutant alters conformation and epigenetic regulation of the anti-adipogenic MIR335 locus. J Cell Biol 216:2731–2743
Önder Ö, Sidoli S, Carroll M, Garcia BA (2015) Progress in epigenetic histone modification analysis by mass spectrometry for clinical investigations. Expert Rev Proteomics 12:499–517
Ouyang M, Wan R, Qin Q, Peng Q, Wang P, Wu J, Allen M, Shi Y, Laub S, Deng L, Lu S, Wang Y (2019) Sensitive FRET biosensor reveals Fyn kinase regulation by submembrane localization. ACS Sens 4:76–86
Pan Y, Lu S, Lei L, Lamberto I, Wang Y, Pasquale EB, Wang Y (2019) Genetically encoded FRET biosensor for visualizing EphA4 activity in different compartments of the plasma membrane. ACS Sens 4:294–300
Park J-A, Kim A-J, Kang Y, Jung YJ, Kim HK, Kim KC (2011) Deacetylation and methylation at histone H3 lysine 9 (H3K9) coordinate chromosome condensation during cell cycle progression. Mol Cells 31:343–349
Peng Q, Cheng B, Lu S, Chien S, Wang Y (2016) Perspectives of FRET imaging to study epigenetics and mechanobiology in the nucleus. In: Chien S, Engler AJ, Wang PY (eds) Molecular and cellular mechanobiology. Springer, New York, pp 143–161
Peng Q, Lu S, Shi Y, Pan Y, Limsakul P, Chernov AV, Qiu J, Chai X, Shi Y, Wang P, Ji Y, Li YJ, Strongin AY, Verkhusha VV, Izpisua Belmonte JC, Ren B, Wang Y, Chien S, Wang Y (2018) Coordinated histone modifications and chromatin reorganization in a single cell revealed by FRET biosensors. Proc Natl Acad Sci USA 115:E11681–E11690
Pesavento JJ, Yang H, Kelleher NL, Mizzen CA (2008) Certain and progressive methylation of histone H4 at lysine 20 during the cell cycle. Mol Cell Biol 28:468–486
Ricci MA, Manzo C, García-Parajo MF, Lakadamyali M, Cosma MP (2015) Chromatin fibers are formed by heterogeneous groups of nucleosomes in vivo. Cell 160:1145–1158
Rice JC (2002) Mitotic-specific methylation of histone H4 Lys 20 follows increased PR-Set7 expression and its localization to mitotic chromosomes. Genes Dev 16:2225–2230
Sasaki K, Yoshida M (2016) The exploitation of FRET probes to track bromodomain/histone interactions in cells for bromodomain inhibitors. Drug Discov Today Technol 19:51–56
Sasaki K, Ito T, Nishino N, Khochbin S, Yoshida M (2009) Real-time imaging of histone H4 hyperacetylation in living cells. Proc Natl Acad Sci USA 106:16257–16262
Scharf AND, Barth TK, Imhof A (2009) Establishment of histone modifications after chromatin assembly. Nucleic Acids Res 37:5032–5040
Schotta G, Lachner M, Sarma K, Ebert A, Sengupta R, Reuter G, Reinberg D, Jenuwein T (2004) A silencing pathway to induce H3–K9 and H4–K20 trimethylation at constitutive heterochromatin. Genes Dev 18:1251–1262
Schreiber KH, Kennedy BK (2013) When lamins go bad: nuclear structure and disease. Cell 152:1365–1375
Schulze JM, Jackson J, Nakanishi S, Gardner JM, Hentrich T, Haug J, Johnston M, Jaspersen SL, Kobor MS, Shilatifard A (2009) Linking cell cycle to histone modifications: SBF and H2B monoubiquitination machinery and cell-cycle regulation of H3K79 dimethylation. Mol Cell 35:626–641
Shao M, Chen G, Lv F, Liu Y, Tian H, Tao R, Jiang R, Zhang W, Zhuo C (2017) LncRNA TINCR attenuates cardiac hypertrophy by epigenetically silencing CaMKII. Oncotarget 8:47565–47573
Shimomura O, Johnson FH, Saiga Y (1962) Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J Cell Comp Physiol 59:223–239
Shogren-Knaak M (2006) Histone H4–K16 acetylation controls chromatin structure and protein interactions. Science 311:844–847
Shpargel KB, Starmer J, Yee D, Pohlers M, Magnuson T (2014) KDM6 demethylase independent loss of histone H3 lysine 27 trimethylation during early embryonic development. PLoS Genet 10:e1004507. https://doi.org/10.1371/journal.pgen.1004507
Stein S, Matter CM (2011) Protective roles of SIRT1 in atherosclerosis. Cell Cycle 10:640–647
Stephens AD, Liu PZ, Banigan EJ, Almassalha LM, Backman V, Adam SA, Goldman RD, Marko JF (2018) Chromatin histone modifications and rigidity affect nuclear morphology independent of lamins. Mol Biol Cell 29:220–233
Stewart MD, Li J, Wong J (2005) Relationship between histone H3 lysine 9 methylation, transcription repression, and heterochromatin protein 1 recruitment. Mol Cell Biol 25:2525–2538
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403:41–45
Sugiyama K, Sugiura K, Hara T, Sugimoto K, Shima H, Honda K, Furukawa K, Yamashita S, Urano T (2002) Aurora-B associated protein phosphatases as negative regulators of kinase activation. Oncogene 21:3103–3111
Swift J, Ivanovska IL, Buxboim A, Harada T, Dingal PC, Pinter J, Pajerowski JD, Spinler KR, Shin JW, Tewari M, Rehfeldt F, Speicher DW, Discher DE (2013) Nuclear lamin-A scales with tissue stiffness and enhances matrix-directed differentiation. Science 341:1240104. https://doi.org/10.1126/science.1240104
Tajik A, Zhang Y, Wei F, Sun J, Jia Q, Zhou W, Singh R, Khanna N, Belmont AS, Wang N (2016) Transcription upregulation via force-induced direct stretching of chromatin. Nat Mater 15:1287–1296
Tan M, Luo H, Lee S, Jin F, Yang JS, Montellier E, Buchou T, Cheng Z, Rousseaux S, Rajagopal N, Lu Z, Ye Z, Zhu Q, Wysocka J, Ye Y, Khochbin S, Ren B, Zhao Y (2011) Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146:1016–1028
Tan Y, Tajik A, Chen J, Jia Q, Chowdhury F, Wang L, Chen J, Zhang S, Hong Y, Yi H, Wu DC, Zhang Y, Wei F, Poh YC, Seong J, Singh R, Lin LJ, Doğanay S, Li Y, Jia H, Ha T, Wang Y, Huang B, Wang N (2014) Matrix softness regulates plasticity of tumour-repopulating cells via H3K9 demethylation and Sox2 expression. Nat Commun 5:4619. https://doi.org/10.1038/ncomms5619
Tebo AG, Gautier A (2019) A split fluorescent reporter with rapid and reversible complementation. Nat Commun 10:1–8
Tsien RY (2005) Building and breeding molecules to spy on cells and tumors. FEBS Lett 579:927–932
Tzima E, Irani-Tehrani M, Kiosses WB, Dejana E, Schultz DA, Engelhardt B, Cao G, DeLisser H, Schwartz MA (2005) A mechanosensory complex that mediates the endothelial cell response to fluid shear stress. Nature 437:426–431
Vaquero A (2006) SirT2 is a histone deacetylase with preference for histone H4 Lys 16 during mitosis. Genes Dev 20:1256–1261
Varela I, Pereira S, Ugalde AP, Navarro CL, Suárez MF, Cau P, Cadiñanos J, Osorio FG, Foray N, Cobo J, de Carlos F, Lévy N, Freije JM, López-Otín C (2008) Combined treatment with statins and aminobisphosphonates extends longevity in a mouse model of human premature aging. Nat Med 14:767–772
Wang Y, Wang N (2009) FRET and mechanobiology. Integr Biol Quant Biosci Nano Macro 1:565–573
Wang Y, Botvinick EL, Zhao Y, Berns MW, Usami S, Tsien RY, Chien S (2005) Visualizing the mechanical activation of Src. Nature 434:1040. https://doi.org/10.1038/nature03469
Wang Y, Shyy JY-J, Chien S (2008) Fluorescence proteins, live-cell imaging, and mechanobiology: seeing is believing. Annu Rev Biomed Eng 10:1–38
Wang Y, Makhija E, Damodaran K, Shivashankar GV (2016) Role of cell geometry on nuclear mechanics, chromosome reorganization, and gene expression. In: Chien S, Engler AJ, Wang PY (eds) Molecular and cellular mechanobiology. Springer, New York, pp 197–216
Wang H, Nakamura M, Abbott TR, Zhao D, Luo K, Yu C, Nguyen CM, Lo A, Daley TP, La Russa M, Liu Y, Qi LS (2019) CRISPR-mediated live imaging of genome editing and transcription. Science 365:1301–1305
Xhemalce B, Dawson MA, Bannister AJ (2011) Histone modifications. In: Reviews in cell biology and molecular medicine. American Cancer Society, New York
Yu N-K, Baek SH, Kaang B-K (2011) DNA methylation-mediated control of learning and memory. Mol Brain 4:5. https://doi.org/10.1186/1756-6606-4-5
Zhang Y, Reinberg D (2001) Transcription regulation by histone methylation: interplay between different covalent modifications of the core histone tails. Genes Dev 15:2343–2360
Zhang Q-J, Chen H-Z, Wang L, Liu DP, Hill JA, Liu ZP (2011) The histone trimethyllysine demethylase JMJD2A promotes cardiac hypertrophy in response to hypertrophic stimuli in mice. J Clin Invest 121:2447–2456
Zheng B, Han M, Shu Y-N, Li YJ, Miao SB, Zhang XH, Shi HJ, Zhang T, Wen JK (2011) HDAC2 phosphorylation-dependent Klf5 deacetylation and RARα acetylation induced by RAR agonist switch the transcription regulatory programs of p21 in VSMCs. Cell Res 21:1487–1508
Zhu W-S, Tang C-M, Xiao Z, Zhu JN, Lin QX, Fu YH, Hu ZQ, Zhang Z, Yang M, Zheng XL, Wu SL, Shan ZX (2016) Targeting EZH1 and EZH2 contributes to the suppression of fibrosis-associated genes by miR-214-3p in cardiac myofibroblasts. Oncotarget 7:78331. https://doi.org/10.18632/oncotarget.13048
Zhuang J, Luan P, Li H, Wang K, Zhang P, Xu Y, Peng W (2017) The Yin-Yang dynamics of DNA methylation is the key regulator for smooth muscle cell phenotype switch and vascular remodeling. Arterioscler Thromb Vasc Biol 37:84–97
Acknowledgements
This work is supported by grants from the Scientific and Technological Innovation Programs of Higher Education Institutions in Shanxi (201802075), the Key Research and Development Program of Shanxi Province (201803D221013-4), and the Young Scientists Fund of the National Natural Science Foundation of China (81802740).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Yingxiao Wang is a scientific co-founder of Cell E&G Inc. and Acoustic Cell Therapy LLC. However, these financial interests do not affect the content of this review article. Shitian Li, Dingyi Yang, Li Gao, and Qin Peng declare that they have no conflict of interest.
Human and animal rights and informed consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, S., Yang, D., Gao, L. et al. Epigenetic regulation and mechanobiology. Biophys Rep 6, 33–48 (2020). https://doi.org/10.1007/s41048-020-00106-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41048-020-00106-x