Abstract
Epigenetic modulations are crucial for the regulation of chromatin structures and genomic organizations, which are key factors determining gene expressions and cellular functions. Mechanical cues have also been shown to play important roles in modulating gene expressions and cellular functions. While there have been significant advances in our understanding of mechanotransduction in the nucleus, there is a lack of knowledge on the molecular details by which mechanical cues affect epigenetic and chromatin regulations to determine genetic outcomes. In this chapter we first introduce the current understanding on epigenetic regulations, particularly on histone modifications and DNA methylations. This is followed by the epigenetic regulations related to mechanobiology in the nucleus. We then introduce the development of genetically encoded molecular biosensors and the principles based on fluorescence proteins (FPs) and fluorescence resonance energy transfer (FRET) for the visualization of dynamic epigenetic regulations in single cells. Lastly, we present examples of the application of biosensors to visualize mechanotransduction events occurring in the nucleus in live cells. The single cell imaging of nuclear mechanotransduction can shed new lights on the molecular mechanisms regulating physiological and pathophysiological processes in living cells under different mechanical environments.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Live cell imaging
- Fluorescence proteins (FPs)
- Fluorescence resonance energy transfer (FRET)
- Mechanobiology
- Mechanotransduction
1 Introduction
Epigenetics refers to molecular variations in cells that change gene expression without altering DNA sequence, typically through changes in DNA modifications and chromatin proteins, e.g., DNA methylations and histone posttranslational modifications that alter DNA accessibility for the activation or silencing of transcription. It has been well documented that epigenetics and its modulation can govern genomic regulation and ultimately cell fates (Li et al. 2012). In particular, epigenetics has been shown to mediate the cellular functions in response to the changing mechanical cues in the cell microenvironment (Downing et al. 2013; Tan et al. 2014). However, it remains largely unclear how epigenetics is regulated in space and time in relation to the genomics upon receiving the mechanical cues. Most current studies utilize the DNA sequencing , Chip-on-Chip, and Chip-Seq technologies to investigate epigenetic regulation (Li et al. 2012). However, these assays in general require the signals to be averaged from a large number of lysed cells obtained at a given time. As such, the dynamic nature of epigenetic signals crucial for cellular functions in single cells, such as cell cycle control , can be masked by the noise engendered from the cell–cell heterogeneity, particularly in non-synchronized cells with relatively varied cellular processes. Therefore, there is an urgent need for the development of new imaging technologies and detection methods in order to elucidate the spatiotemporal landscape and epigenetic regulation.
2 Epigenetic Regulation and Mechanobiology in the Nucleus
2.1 Chromatin
Chromatins are highly ordered nuclear structures that contain DNAs, histones, and other chromosomal proteins. A fundamental subunit of chromatin in eukaryotes is the nucleosome, which consists of approximately 147 base pairs (bp) of DNA associated with a complex of eight core histones (two copies each of histone H2A, H2B, H3, and H4). The wrapping of genomic DNA sequence in 1.6 turns around the octamer results in a five- to tenfold compaction of DNA (Kornberg 1974; Kornberg and Thomas 1974; Black et al. 2012). The compact DNA is only partially accessible to regulatory proteins, but it can become more available if there is a conformational alteration of the nucleosome, or if the DNA is partly unwound from the histones. The histone tails protrude from the core complex and are readily accessible. Enzymes can chemically modify these tails to create docking motifs and recruit various chromatin modulators for the regulation of nucleosome remodeling and DNA unwinding, which can have profound effects on the chromatin complex and genomic regulation (Felsenfeld and Groudine 2003). Since chromatin has a very compact organization and most DNA sequences are structurally inaccessible at rest, it is in general unfavorable for transcription. It has been shown that enzymes recognizing DNA sequences have easier access to nucleosomes associated with active genes (Weintraub and Groudine 1976), suggesting a selective unfolding of the compact structure to activate gene transcription (Felsenfeld and Groudine 2003).
Chromatins and nucleosomes are subject to modifications to potentially alter their structures and regulate gene activity. There are three general ways in which chromatin structure can be altered by nucleosome modification. First, utilizing ATP, nucleosome remodeling can be directly induced by chromatin remodeling complexes designed specifically for the task (Becker and Horz 2002). Second, covalent modifications of histones can occur in the histone tails to recruit chromatin remodeling proteins (Zhang and Reinberg 2001). Third, histone variants may replace one or more of the core histones (Ahmad and Henikoff 2002; Redon et al. 2002; Smith 2002; Felsenfeld and Groudine 2003).
2.2 Chromatin and Histone Posttranslational Modifications (PTM)
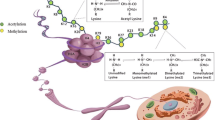
Since the pioneering studies by Allfrey in the early 1960s, posttranslational modifications on histones have been well documented (Allfrey et al. 1964). A large number of different histone posttranslational modifications (PTMs) have been reported and categorized (Table 7.1 and Fig. 7.1) (Luger et al. 1997; Cota et al. 2013). Among all the PTMs, histone acetylation, methylation, and phosphorylation are the most prevalent forms. In fact, all histones can be modified at different sites by acetylation, methylation, and phosphorylation. The determination of the high-resolution X-ray structure of the nucleosome in 1997 led to the insights into how these histone PTMs could affect chromatin structure (Luger et al. 1997). The resolved structure indicates that the highly basic histone amino (N)-terminal tails can extend from their own nucleosome and make contact with adjacent nucleosomes. It is likely that the modification of these tails can affect inter-nucleosomal interactions and thus affect the structure of the whole chromatin (Bannister and Kouzarides 2011).
Nucleosome with histone posttranslational modifications (Cota et al. 2013)
Histone acetylation was first reported in 1964 (Allfrey et al. 1964). The acetylation on lysine sites is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases (HATs) and histone deacetylase (HDACs) (Fig. 7.2) (Xhemalce et al. 2011). The HATs utilize acetyl CoA as a cofactor and catalyze the transfer of an acetyl group to the lysine side chains. This action neutralizes the lysine’s positive charge, and has the potential to weaken the interactions between histones and DNA for chromatin remodeling (Bannister and Kouzarides 2011).
The diagram of histone acetylation regulation (Cota et al. 2013)
Histone phosphorylation takes place on residues including serine, threonine, and tyrosine, predominantly in the N-terminal histone tails (Xhemalce et al. 2011). Like histone acetylation, the phosphorylation of histones is also highly dynamic. The phosphorylation level is controlled by kinases and phosphatases that add and remove the phosphate group, respectively (Oki et al. 2007). Histone kinases transfer a phosphate group from ATP to the hydroxyl group of the target amino acid side chain, adding negative charge to the histone to affect the chromatin structure. However, it is unclear how the kinases are accurately recruited to the phosphorylation sites on histones at chromatins. Even less information is available regarding the roles of histone phosphatases, although it is clear that a high level of phosphatase activity exists in the nucleus to direct a rapid turnover of histone dephosphorylations . For example, the proteins phosphatase 1 (PP1) can rapidly neutralize the action of Aurora B kinase, which causes genome-wide phosphorylation events at H3S10 during mitosis (Fig. 7.3) (Goto et al. 2002; Sugiyama et al. 2002; Bannister and Kouzarides 2011).
The phosphorylation and dephosphorylation of H3S10 by Aurora kinase and PP1, respectively (Oki et al. 2007)
Unlike acetylation and phosphorylation , histone methylation does not alter the charge of histone proteins. Lysines and arginines are the main targets for histone methylation , with lysines being mono-, di-, or tri-methylated (Fig. 7.4) whereas arginines being mono-, or symmetrically or asymmetrically di-methylated (Bannister and Kouzarides 2005; Bedford and Clarke 2009; Lan and Shi 2009; Ng et al. 2009; Bannister and Kouzarides 2011; Xhemalce et al. 2011). The methylation level is controlled by methyl-transferase and demethylase that can add and remove the methyl group, respectively.
The diagram of different lysine methylations (Rea et al. 2000)
Among the different methyl-transferases , SUV39H1 is the first identified histone lysine methyl-transferase (HKMT) , which targets H3K9 for tri-methylation (Bannister and Kouzarides 2005). Numerous HKMTs have since been identified, with the vast majority of them methylating lysines at the N-terminal tails of histone proteins. Except the Dot1 enzyme that methylates H3K79 in the histone globular core, HKMTs that methylate N-terminal lysines contain a so-called SET domain harboring the enzymatic activity, which can specifically catalyze the transfer of a methyl group from S-adenosylmethionine (SAM) to a lysine’s ε-amino group. The resulted outcome can be either activation or repression of corresponding gene transcriptions (Martinez-Balbas et al. 1995; Bannister and Kouzarides 2011).
Current information suggests that some lysine methylation sites on histones are associated with transcriptional activation , e.g., H3K4, H3K36, and H3K79, with H3K4me and H3K36me implicated in transcriptional elongation. Other lysine methylation sites are connected to the repression of transcription, e.g., H3K9, H3K27, and H4K20. Methylation at H3K9 is implicated in the silencing of euchromatic genes, as well as forming silent heterochromatin such as pericentric heterochromatin surrounding centromeres. Gene repression related to the H3K9 methylation may involve the recruitment of methyl-transferase and heterochromatin protein 1 (HP1) to the promoter region of repressed genes. It is interesting that H3K9me3 and HP1γ are enriched in the coding region of active genes (Martinez-Balbas et al. 1995). It is possible that H3K9 methylation represents active signals within the coding regions, while promoting gene repression at the promoter sites (Kouzarides 2007).
Via histone posttranslational modifications, chromatin can undergo dynamic structural alterations to control the activation or repression of specific genes, e.g., during the cell cycle processes or in response to mechanical environmental cues. Indeed, the distinct H3 and H4 tail modifications may act sequentially or in combination to effect unique biological outcomes, constituting the so-called histone code (Strahl and Allis 2000). For example, H3K14 acetylation is required in addition to H3S10 phosphorylation to repel HP1 during G2/M transition in a cell cycle (Mateescu et al. 2004), with a minor influence of H3S10 phosphorylation observed on the HP1 binding to the tri-methylated H3K9 (Vermeulen et al. 2010). H3S10 phosphorylation can also lead to the recruitment of histone deacetylase HST2 for the deacetylation of H4K16 and its interaction with H2A for chromatin condensation (Wilkins et al. 2014). However, it remains unclear on how this hierarchy of multiple modifications extends (depicted as “higher-order combinations”), or how distinct combinatorial sets are established or maintained in the localized regions of the chromatin structure .
2.3 Cell Cycle and Histone Modifications
In eukaryotic cells , chromosomes occupy relatively large and intertwined regions in the interphase nucleus. As such, the packing and unpacking of chromosomes into discrete units are key tasks during cell division. The mechanics of chromatin organization is also important during cell division. For example, centromeres , the distinct chromatin regions of chromosome on which kinetochores form during mitosis, allow kinetochore formation and attachment to the mitotic spindle, which mechanically moves chromosomes into daughter cells. It has been shown that the large displacement of transcription factors from chromatin and general transcription are suppressed during mitosis (Gottesfeld and Forbes 1997; Egli et al. 2008). Since the defining features of a specific cell lineage can be maintained epigenetically during cell proliferation, mechanisms must also exist to ensure a “memory” of transcriptional program is inherited through mitosis (Egli et al. 2008; Moazed 2011; Sarkies and Sale 2012), which usually involves chromatin remodeling (Wang and Higgins 2013).
In order to replicate the DNA sequence and pass it on in the genome, chromatin undergoes dynamic cycles of compaction and unpacking during cell cycle procession. Previous studies on replication timing have demonstrated that distinct chromatin replications happen at different times during S phases (Kennedy et al. 2000; Dimitrova and Berezney 2002). Euchromatic regions in general replicate early in the interior of the nucleus, followed by the replication of heterochromatin regions associated with the nuclear lamina. During this process, the local changes of chromatin microenvironment could dictate cell cycle and proliferation rate by regulating genes directly. The requirement of KMT4/Dot1 for the efficient entry into S phase is an example of transcriptional control of cell cycle genes through H3K79 di-methylation (Schulze et al. 2009). The chromatin microenvironment can also affect cell cycle indirectly by changing the replication timing. For example, KMT6/EZH2 , by regulating epigenetic H3K27me3 levels, can target transcriptional regulations of cell cycle proteins Cyclin D1, E1, and A2 (Bracken et al. 2003; Black and Whetstine 2011). The overexpression of the H3K9me3 demethylase KDM4A results in a faster progression through S phase, possibly due to the better chromatin accessibility, increase in replication forks, and altered replication timing at heterochromatin regions (Black et al. 2012). Similarly, the loss of KDM4A in MDA-MB-231 breast cancer cells leads to a G1/S arrest and decreased proliferation rates (Black et al. 2010). Consistent with these results, H3K9me3 levels are reduced in S phase, accompanied by an increase in H3K9me1/2 levels (Fig. 7.5) (O’Sullivan et al. 2010). Although the complete molecular mechanisms remain to be elucidated, these results suggest that H3K9me3 and other histone epigenetic marks may serve as conserved determinants of replication timing to regulate cell cycle processes. Therefore, the modulation of cell cycle genes can be a key determinant in cell cycle, which can be regulated by a dynamic balance between histone methyl-transferases and demethylases.
Histone methylations, as well as KMTs and KDMs, are dynamically regulated during the cell cycle (Black et al. 2012)
2.4 The Role of H3K9me3 and Chromatin Condensation in Cell Mitosis
In the interphase nucleus, chromatin is highly ordered for the storage of the genetic material (Campos and Reinberg 2009). Chromosome condensation occurs during mitosis to facilitate the separation of sister chromatids and the maintenance of genomic stability. It was demonstrated that histone modification within the nucleosome can trigger the structural changes for chromosome condensation (Trojer and Reinberg 2007; Duan et al. 2008). In fact, H3K9me3 was identified as a marker of heterochromatin and epigenetic silencing regions, which typically are located close to the centromeres (Schotta et al. 2004; Stewart et al. 2005). Accordingly, H3K9me3 is dynamically regulated in a cell cycle-dependent manner (McManus et al. 2006; Duan et al. 2008; Park et al. 2011). H3K9me3 increased rapidly in G2 to reach a maximum, followed by a quick decline (McManus et al. 2006). Loss of H3K9me3 is correlated with mitotic defects such as failure of chromosomes alignment, as shown in mouse embryonic fibroblasts (MEFs) following the elimination of histone-lysine-N-methyl transferases (SUV39h1/h2−/− cells) (Peters et al. 2001; McManus et al. 2006). Both SUV39h1 and SUV39h2 are primary enzymes catalyzing the tri-methylation of H3K9 (Rea et al. 2000; Heit et al. 2009). In these SUV39h1/2 knockout cells , the mitotic checkpoint is active and kinetochore proteins localize properly. However, pericentric chromatin in these cells is less condensed. Mis-segregation of chromosomes is also observed in cells treated with the methylation inhibitor adenosine dialdehyde (AdOx) , which affects the tri-methylation of both H3K9 and H4K20. The reduced integrity of pericentric heterochromatin might be responsible for the loss of tension at the centromere and activation of the spindle assembly checkpoint in AdOx-treated cells, which leads to the mitotic defects. Therefore, an increase in H3K9me3 at late G2 phase and early mitosis may be needed to stabilize the pericentric heterochromatin so that the centromeres and kinetochores have a rigid structure needed for proper tension transmission to the inner centromere (Heit et al. 2009). However, this result based on immunostaining is in contrast to the more recent results of genome-wide dynamic change of H3K9me3 in the overall cell cycle (Fig. 7.5). A recent study further revealed an association between chromatin and a lysine demethylase KDM4C during mitosis, which is accompanied by a decrease in the mitotic levels of H3K9me3 (Kupershmit et al. 2014).
Epigenetic modifications including histone methylation at different residues are early events that can guide gene regulation networks and expressions. The histone methylations and their combinations can serve as codes to determine the overall gene expressions and phenotypic outcomes. For example, mounting evidence suggests a direct role of histone H3K9 methylation as a histone marker positively correlating with DNA methylation and participating in repressive heterochromatin formation in tumorigenesis. In the cell cycle, H3K9me3 is important for HP1γ recruitment to regulate gene expression, chromatin packaging, and heterochromatin formation. However, the dynamics of H3K9me3 remains controversial, mainly due to the lack of the appropriate tool for H3K9me3 detection in living cells.
2.5 Epigenetics in Mechanobiology
Increasing evidence suggests that epigenetic modulations in DNA methylation and histone modification play a crucial role in regulating gene expression and cellular functions under different mechanical environments. Early studies have demonstrated that shear stress can cause the chromatin remodeling on both histones H3 and H4 to result in the eNOS gene expression modulation at the transcriptional level (Illi et al. 2003; Fish et al. 2005). Recently, hemodynamic force-induced histone modifications in cardiovascular systems have been extensively studied (Chen et al. 2013), mainly on histone acetylation/deacetylation (HAT/HDAC) . Three main groups of HDACs sensitive to hemodynamic force were characterized: class I (HDAC-1/2/3 and HDAC-8), class II (HDAC-4/5/6/7 and HDAC-9/10), and class III sirtuins (SIRT). Laminar flow was shown to increase the activity of class I HDACs and induce the association between HDAC1 and p53, leading to the deacetylation of p53 at Lys-320 and Lys-373 in ECs for the cell cycle arrest (Zeng et al. 2003). Oscillatory flow was also shown to modulate cell proliferation by increasing the activity of class I HDAC 1/2/3, but not class II HDAC4/7 (Lee et al. 2012). Both Class II and III HDACs have been demonstrated to play a critical role in the shear stress-induced eNOS expression (Illi et al. 2008; Chen et al. 2010), with Class III HDACs also playing a protective role in atherosclerosis (Stein and Matter 2011). Besides the modification of histones, DNA hypermethylation was shown to result in atheroprone gene expression in human umbilical vein endothelial cells (HUVECs) and rat carotid arteries as well as hematopoietic cell development (Lund et al. 2004; Zhou et al. 2014). Kim et al. further demonstrated that both the global DNA methylation profiles in atherosclerotic tissues and the DNA methylation patterns of estrogen receptor-β are correlated to the atherosclerotic development (Post et al. 1999).
Mechanical signals are also important regulators of epigenetics in guiding stem cell behavior and cell/tissue differentiation. Recent study has demonstrated that cues from the mechanical microenvironment can alter the DNA methylation in promoter regions of osteogenic genes in bone marrow mesenchymal stem cells (bMSCs) to control the osteogenic cell fate (Arnsdorf et al. 2010). Biophysical cues such as microgroove topography were also shown to increase the histone H3 acetylation in somatic fibroblasts to promote cell reprogramming (Downing et al. 2013). Mechanical matrix stiffness was further shown to affect structural protein lamin-A and guide cell differentiations (Swift et al. 2013). In fact, stem cell differentiation into fat cells on soft matrices and into bone cells on stiff matrices can be enhanced by the low and high lamin-A levels, respectively (Swift et al. 2013). These observations are consistent with the subcellular location and function of lamin-A proteins, lying inside the nuclear envelope and interacting with both chromatins and the cytoskeleton to transmit mechanical signals into the nucleus for the regulation of epigenetics and gene expression. Recent observation revealed that a high level of H3K9 tri-methylation is correlated with the peripheral localization of chromosomes proximal to Lamins, with the inhibition of H3K9 tri-methylation leading to the relocation of chromosomes (Bian et al. 2013). Consistently, Lamin-A/C-deficient (Lmna−/−) and LmnaN195K/N195K mutant cells have impaired nuclear morphology and downstream signaling of the mechanosensitive transcription factor MKL1 (Ho et al. 2013). Moreover, a single point mutation (G608G) in lamin-A gene (LMNA) caused aging-associated nuclear defects including disorganization of nuclear lamina and loss of heterochromatin (Scaffidi and Misteli 2005; Scaffidi and Misteli 2006; Dechat et al. 2008; Pegoraro et al. 2009). This critical role of lamin A in regulating epigenetics and gene expression can also be seen in Hutchinson–Gilford progeria syndrome (HGPS) fibroblasts , which showed abnormal nuclear morphology and lamina, loss of heterochromatin markers H3K9me3, HP1α, and HDAC1 (Liu et al. 2011). Therefore, lamin A, by mechanically connecting cytoskeleton outside and chromatin inside the nucleus, may serve as a mechanotransducer to relay mechanical signals into nucleus to regulate epigenetic modifications and gene transcriptions for the determination of cell fates .
3 FRET Imaging of Epigenetic Regulations in Single Cells
3.1 FRET Imaging
Recent advances in genetic technologies have allowed the sequencing of the whole human genome, providing invaluable information toward our molecular understanding of human pathophysiological regulations. However, genomic sequences lack spatial and temporal information of the target molecules, as well as information on post-genomic modifications, which directly link to biological regulations in health and disease. Therefore, imaging technologies are crucial for the direct determinations of the spatiotemporal characteristics of different functional molecules in cells and organisms (Wang et al. 2008; Wang and Wang 2009). Parallel progresses in the developments of molecular imaging probes and microscopic technologies have allowed the dynamic monitoring of molecular localization, activities, and interactions in real time in live cells. In this aspect, the fluorescence proteins (FPs ) and fluorescence resonance energy transfer (FRET) technology have become timely developments. FPs such as the green-colored GFP have allowed a variety of revolutionary discoveries, but the limitation of the GFP-tagging approach is that only the location of the molecules can be monitored. In contrast, FRET technology, which utilizes FPs to provide ratiometric readout on molecular activities and interactions, has been increasingly powerful in live cell imaging (Wang et al. 2008; Wang and Wang 2009).
The concept of FRET is based on the interactions of a pair of FPs . When two FPs are very close in distance, with the emission spectrum of the donor FP overlapping the excitation spectrum of the acceptor FP, energy transfer occurs if these two FPs are in favorable orientations (Clegg 1996; Clegg 2005; Wang and Wang 2009). The FRET efficiency between the donor and acceptor FPs of a FRET pair is mainly dependent on three factors: The first is the overlapping area between the emission spectrum of the donor and the excitation spectrum of the acceptor (Fig. 7.6a), with a larger area providing a higher efficiency. The second factor is the distance between the donor and acceptor. The FRET efficiency is inversely proportional to the 6th power of the distance between donor and acceptor. Hence, a slight modification of this distance can markedly affect the FRET signals. Typically, sufficient FRET between FPs occurs only when this distance is within 10 nm (Fig. 7.6b). The second factor is the relative orientation between donor and acceptor. It has been reported that FRET can change dramatically by changing the orientation while maintaining the distance between the two FPs (Nagai et al. 2004; Giepmans et al. 2006). Because FRET can provide such a high sensitivity in detecting a change in the distance or orientation between the donor and acceptor FPs , it has been widely applied to designing biosensors for the measurement of molecular signals with high precision (Piston and Kremers 2007; Wang et al. 2008; Wang and Wang 2009).
(a) The excitation and emission spectra of a typical FRET pair, with cyan fluorescent protein (CFP) as the donor and yellow fluorescent protein (YFP) as the acceptor. The broken lines represent the excitation spectra and the solid lines the emission spectra of CFP and YFP. The spectra curves of CFP and YFP are color-coded with cyan and yellow, respectively. The shaded red area represents the overlap between the CFP emission and the YFP excitation. (b) The cartoon shows that the FRET efficiency between a typical FRET pair, enhanced CFP (ECFP ) as the donor and enhanced YFP (EYFP ) as the acceptor, is mainly dependent on the distance and the relative orientation between the donor and acceptor (Wang and Wang 2009)
3.2 FRET Biosensors
Based on the principle of FRET, different kinds of biosensors have been developed for biological applications. For example, hybridized sensors consisting of polypeptides and protein domains recognizing active target molecules have been applied to detect the small GTPases RhoA and Cdc42 (Nalbant et al. 2004; Pertz and Hahn 2004; Hodgson et al. 2008). However, genetically encoded FRET biosensors are particularly appealing for live cell imaging , because these biosensors can be conveniently introduced into cells for targeting subcellular compartments to continuously monitor local molecular signals (Wang et al. 2008; Wang and Wang 2009). For the development of genetically encoded FRET biosensors, researchers have originally chosen the enhanced and blue-colored EBFP and green-colored EGFP as the FRET donor and acceptor pair (Romoser et al. 1997; Mahajan et al. 1998). Although the acceptor EGFP is quite bright, the donor EBFP is dim and poor in photo-stability, hence hampering further usage of the EBFP/EGFP FRET pair. Later, it was discovered that the enhanced cyan-colored ECFP and yellow-colored EYFP , including the yellow-colored variants of EYFP such as Venus and Citrine, provide excellent FRET pairs (Miyawaki et al. 1997; Griesbeck et al. 2001; Itoh et al. 2002; Nagai et al. 2002; Yoshizaki et al. 2003). Therefore, these FRET pairs have become increasingly popular for the biosensor development. In fact, ECFP and a YFP variant, YPet, have been shown to serve as a high-efficiency FRET pair for a variety of biosensors (Ouyang et al. 2008). At this stage, it seems that the brightness of ECFP is still a limiting factor for cell imaging. Recently, new variants for EBFP and ECFP have been developed, including Azurite (Mena et al. 2006), EBFP2 (Pedelacq et al. 2006), Cerulean (Rizzo et al. 2004), and mTFP1 (Ai et al. 2006). All these newly developed FP variants display significantly enhanced fluorescent properties, such as brightness and photo-stability, and they provide promising candidates for further improvement of FRET biosensors (Wang and Wang 2009).
A single-molecule FRET biosensor typically consists of a FRET pair of FPs and two intramolecular domains capable of interacting with each other. For example, a FRET-based biosensor capable of detecting Src kinase activation has been developed consisting of an N-terminal ECFP , a SH2 domain derived from Src kinase, a flexible linker, a substrate peptide derived from p130cas and specifically sensitive to Src phosphorylation, and a C-terminal Citrine (EYFP ) (Wang et al. 2005; Wang et al. 2008; Wang and Wang 2009). This Src FRET biosensor has been applied to monitor Src activity continuously in live cells (Wang et al. 2005; Wang et al. 2008; Wang and Wang 2009). Alternatively, the donor and acceptor FPs can also be fused with two different target molecules, with the distance between donor and acceptor representing the interaction or separation of two target molecules (Zaccolo et al. 2000; Ai et al. 2006; Schulze et al. 2009; Wang and Wang 2009). Comparing these design approaches, the single-molecule FRET biosensors have two advantages: (1) the intramolecular interaction between the biosensor domains is robust and resistant to the interference caused by the interactions between the endogenous target molecules and the biosensor domains; (2) the acceptor/donor ratio will not be influenced by the relative expression levels of donors and acceptors in a single cell. Therefore, the single-molecule FRET biosensors are quite popular for monitoring intracellular signals (Wang and Wang 2009).
3.3 FRET-Based Epigenetic Biosensor
FRET-based biosensors have been recently developed to study the dynamics of histone epigenetics at single living cell level, and to investigate the mechanism of epigenetic regulations on cell fates. The Ting group first developed FRET-based histone methylation reporters to visualize H3K9me3 and H3K27me3 in vitro and in single living cells (Lin et al. 2004). To visualize the spatial patterns of aurora B kinase in anaphase, Fuller et al. developed a strategy using FRET-based sensors to report quantitative changes in substrate phosphorylation in living cells (Fuller et al. 2008). Sasaki et al. developed a FRET-based histone acetylation biosensor, which monitored the dynamic fluctuation of histone H4 acetylation levels during mitosis (Sasaki et al. 2009). Later, Chu et al. developed another FRET-based and centromere-targeted H3K9me3 biosensor to visualize the methylation dynamics during chromosome segregation (Chu et al. 2012). However, the usage of current FRET-based epigenetic biosensors are relatively limited, possibly due to: (1) the substrates of those epigenetic biosensors are in general short histone peptides (N-terminal tail), which cannot be incorporated correctly into target nucleosomes. Therefore, the specificity of the resulting biosensors is not high; (2) the sensitivity of most current epigenetic biosensors is still awaiting further improvement (Sasaki et al. 2009) (Tan et al. 2014). As such, some physiologically important epigenetic signals with moderate magnitude may not be readily detectable. These limitations have led to the scarce application of epigenetic FRET biosensors in mechanobiology. An H3K9me3 FRET biosensor was applied to reveal a low level of H3K9me3 in tumor-repopulating cells (TRCs), which is unresponsive to matrix stiffness or applied forces (Tan et al. 2014). It is apparent that more FRET biosensors will be available to monitor epigenetic modulations, and this may promote the study of epigenetic regulations in mechanobiology .
4 Conclusions and Perspectives
Given the crucial role of epigenetics in genomic regulations and cellular fate determinations, there is a great need to monitor the epigenetic regulation in space and time. FRET biosensors should provide powerful tools in elucidating these spatiotemporal landscapes of epigenetic regulations, particularly in single live cells. The results should advance our precise understanding on the dynamic coordination of epigenetics and genomic regulations at different chromatin locations inside the nucleus, which should exceed the capability of traditional bulky assays based on the averaged extraction of a large number of cells.
It is expected that, with the rapid development in computational molecular modeling and high-throughput screening of large mutant libraries, highly sensitive and specific FRET biosensors will be developed in a systematic and rapid fashion. These biosensors, together with DNA binding domains and genome targeting motifs, should allow the precise monitoring of spatiotemporal epigenetic regulations at specific locus-sites. Similar approaches should also lead to the development of novel FRET biosensors capable of monitoring the spatiotemporal landscapes of DNA methylation evolution during different cellular processes including mechanotransduction, i.e., how cells perceive the mechanical environmental cues and transmit them into the regulation signals of epigenome and genome. In summary, the single cell study of epigenetic regulations in mechanobiology is at its infant stage. FRET imaging integrated with rapidly progressing sequencing technologies should allow the revelation and construction of the spatiotemporal landscape of epigenetic regulations in response to mechanical cues in the near future.
References
Ahmad K, Henikoff S (2002) Histone H3 variants specify modes of chromatin assembly. Proc Natl Acad Sci U S A 99(Suppl 4):16477–16484
Ai HW, Henderson JN et al (2006) Directed evolution of a monomeric, bright and photostable version of Clavularia cyan fluorescent protein: structural characterization and applications in fluorescence imaging. Biochem J 400(3):531–540
Allfrey VG, Faulkner R et al (1964) Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci U S A 51:786–794
Arnsdorf EJ, Tummala P et al (2010) The epigenetic mechanism of mechanically induced osteogenic differentiation. J Biomech 43(15):2881–2886
Bannister AJ, Kouzarides T (2005) Reversing histone methylation. Nature 436(7054):1103–1106
Bannister AJ, Kouzarides T (2011) Regulation of chromatin by histone modifications. Cell Res 21(3):381–395
Becker PB, Horz W (2002) ATP-dependent nucleosome remodeling. Annu Rev Biochem 71:247–273
Bedford MT, Clarke SG (2009) Protein arginine methylation in mammals: who, what, and why. Mol Cell 33(1):1–13
Bian Q, Khanna N et al (2013) Beta-globin cis-elements determine differential nuclear targeting through epigenetic modifications. J Cell Biol 203(5):767–783
Black JC, Whetstine JR (2011) Chromatin landscape: methylation beyond transcription. Epigenetics 6(1):9–15
Black JC, Allen A et al (2010) Conserved antagonism between JMJD2A/KDM4A and HP1gamma during cell cycle progression. Mol Cell 40(5):736–748
Black JC, Van Rechem C et al (2012) Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol Cell 48(4):491–507
Bracken AP, Pasini D et al (2003) EZH2 is downstream of the pRB-E2F pathway, essential for proliferation and amplified in cancer. EMBO J 22(20):5323–5335
Campos EI, Reinberg D (2009) Histones: annotating chromatin. Annu Rev Genet 43:559–599
Chen Z, Peng IC et al (2010) Shear stress, SIRT1, and vascular homeostasis. Proc Natl Acad Sci U S A 107(22):10268–10273
Chen LJ, Wei SY et al (2013) Mechanical regulation of epigenetics in vascular biology and pathobiology. J Cell Mol Med 17(4):437–448
Chu L, Zhu T et al (2012) SUV39H1 orchestrates temporal dynamics of centromeric methylation essential for faithful chromosome segregation in mitosis. J Mol Cell Biol 4(5):331–340
Clegg RM (1996) Fluorescence resonance energy transfer, in fluorescence imaging spectroscopy and microscopy. Wiley, New York
Clegg RM (2005) Nuts and bolts of excitation energy migration and energy transfer, in chlorophyll a fluorescence: a signature of photosynthesis. Adv Photosynth Resp 19:83–105
Cota P, Shafa M, Rancourt DE (2013) Stem cells and epigenetic reprogramming. In book, Pluripotent Stem Cells. Ed. by Bhartiya D, Lenka N, InTech. doi: 10.5772/45917
Dechat T, Pfleghaar K et al (2008) Nuclear lamins: major factors in the structural organization and function of the nucleus and chromatin. Genes Dev 22(7):832–853
Dimitrova DS, Berezney R (2002) The spatio-temporal organization of DNA replication sites is identical in primary, immortalized and transformed mammalian cells. J Cell Sci 115(Pt 21):4037–4051
Downing TL, Soto J et al (2013) Biophysical regulation of epigenetic state and cell reprogramming. Nat Mater 12(12):1154–1162
Duan Q, Chen H et al (2008) Phosphorylation of H3S10 blocks the access of H3K9 by specific antibodies and histone methyltransferase. Implication in regulating chromatin dynamics and epigenetic inheritance during mitosis. J Biol Chem 283(48):33585–33590
Egli D, Birkhoff G et al (2008) Mediators of reprogramming: transcription factors and transitions through mitosis. Nat Rev Mol Cell Biol 9(7):505–516
Felsenfeld G, Groudine M (2003) Controlling the double helix. Nature 421(6921):448–453
Fish JE, Matouk CC et al (2005) The expression of endothelial nitric-oxide synthase is controlled by a cell-specific histone code. J Biol Chem 280(26):24824–24838
Fuller BG, Lampson MA et al (2008) Midzone activation of aurora B in anaphase produces an intracellular phosphorylation gradient. Nature 453(7198):1132–1136
Giepmans BN, Adams SR et al (2006) The fluorescent toolbox for assessing protein location and function. Science 312(5771):217–224
Goto H, Yasui Y et al (2002) Aurora-B phosphorylates Histone H3 at serine28 with regard to the mitotic chromosome condensation. Genes Cells 7(1):11–17
Gottesfeld JM, Forbes DJ (1997) Mitotic repression of the transcriptional machinery. Trends Biochem Sci 22(6):197–202
Griesbeck O, Baird GS et al (2001) Reducing the environmental sensitivity of yellow fluorescent protein. Mechanism and applications. J Biol Chem 276(31):29188–29194
Heit R, Rattner JB et al (2009) G2 histone methylation is required for the proper segregation of chromosomes. J Cell Sci 122(Pt 16):2957–2968
Ho CY, Jaalouk DE et al (2013) Lamin A/C and emerin regulate MKL1-SRF activity by modulating actin dynamics. Nature 497(7450):507–511
Hodgson L, Pertz O et al (2008) Design and optimization of genetically encoded fluorescent biosensors: GTPase biosensors. Methods Cell Biol 85:63–81
Illi B, Nanni S et al (2003) Shear stress-mediated chromatin remodeling provides molecular basis for flow-dependent regulation of gene expression. Circ Res 93(2):155–161
Illi B, Dello Russo C et al (2008) Nitric oxide modulates chromatin folding in human endothelial cells via protein phosphatase 2A activation and class II histone deacetylases nuclear shuttling. Circ Res 102(1):51–58
Itoh RE, Kurokawa K et al (2002) Activation of rac and cdc42 video imaged by fluorescent resonance energy transfer-based single-molecule probes in the membrane of living cells. Mol Cell Biol 22(18):6582–6591
Kennedy BK, Barbie DA et al (2000) Nuclear organization of DNA replication in primary mammalian cells. Genes Dev 14(22):2855–2868
Kornberg RD (1974) Chromatin structure: a repeating unit of histones and DNA. Science 184(4139):868–871
Kornberg RD, Thomas JO (1974) Chromatin structure; oligomers of the histones. Science 184(4139):865–868
Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705
Kupershmit I, Khoury-Haddad H et al (2014) KDM4C (GASC1) lysine demethylase is associated with mitotic chromatin and regulates chromosome segregation during mitosis. Nucleic Acids Res 42(10):6168–6182
Lan F, Shi Y (2009) Epigenetic regulation: methylation of histone and non-histone proteins. Sci China C Life Sci 52(4):311–322
Lee DY, Lee CI et al (2012) Role of histone deacetylases in transcription factor regulation and cell cycle modulation in endothelial cells in response to disturbed flow. Proc Natl Acad Sci U S A 109(6):1967–1972
Li M, Liu GH et al (2012) Navigating the epigenetic landscape of pluripotent stem cells. Nat Rev Mol Cell Biol 13(8):524–535
Lin CW, Jao CY et al (2004) Genetically encoded fluorescent reporters of histone methylation in living cells. J Am Chem Soc 126(19):5982–5983
Liu GH, Barkho BZ et al (2011) Recapitulation of premature ageing with iPSCs from Hutchinson-Gilford progeria syndrome. Nature 472(7342):221–225
Luger K, Mader AW et al (1997) Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389(6648):251–260
Lund G, Andersson L et al (2004) DNA methylation polymorphisms precede any histological sign of atherosclerosis in mice lacking apolipoprotein E. J Biol Chem 279(28):29147–29154
Mahajan NP, Linder K et al (1998) Bcl-2 and Bax interactions in mitochondria probed with green fluorescent protein and fluorescence resonance energy transfer. Nat Biotechnol 16(6):547–552
Martinez-Balbas MA, Dey A et al (1995) Displacement of sequence-specific transcription factors from mitotic chromatin. Cell 83(1):29–38
Mateescu B, England P et al (2004) Tethering of HP1 proteins to chromatin is relieved by phosphoacetylation of histone H3. EMBO Rep 5(5):490–496
McManus KJ, Biron VL et al (2006) Dynamic changes in histone H3 lysine 9 methylations: identification of a mitosis-specific function for dynamic methylation in chromosome congression and segregation. J Biol Chem 281(13):8888–8897
Mena MA, Treynor TP et al (2006) Blue fluorescent proteins with enhanced brightness and photostability from a structurally targeted library. Nat Biotechnol 24(12):1569–1571
Miyawaki A, Llopis J et al (1997) Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388(6645):882–887
Moazed D (2011) Mechanisms for the inheritance of chromatin states. Cell 146(4):510–518
Nagai T, Ibata K et al (2002) A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat Biotechnol 20(1):87–90
Nagai T, Yamada S et al (2004) Expanded dynamic range of fluorescent indicators for Ca(2+) by circularly permuted yellow fluorescent proteins. Proc Natl Acad Sci U S A 101(29):10554–10559
Nalbant P, Hodgson L et al (2004) Activation of endogenous Cdc42 visualized in living cells. Science 305(5690):1615–1619
Ng SS, Yue WW et al (2009) Dynamic protein methylation in chromatin biology. Cell Mol Life Sci 66(3):407–422
O’Sullivan RJ, Kubicek S et al (2010) Reduced histone biosynthesis and chromatin changes arising from a damage signal at telomeres. Nat Struct Mol Biol 17(10):1218–1225
Oki M, Aihara H et al (2007) Role of histone phosphorylation in chromatin dynamics and its implications in diseases. Subcell Biochem 41:319–336
Ouyang M, Sun J et al (2008) Determination of hierarchical relationship of Src and Rac at subcellular locations with FRET biosensors. Proc Natl Acad Sci U S A 105(38):14353–14358
Park JA, Kim AJ et al (2011) Deacetylation and methylation at histone H3 lysine 9 (H3K9) coordinate chromosome condensation during cell cycle progression. Mol Cells 31(4):343–349
Pedelacq JD, Cabantous S et al (2006) Engineering and characterization of a superfolder green fluorescent protein. Nat Biotechnol 24(1):79–88
Pegoraro G, Kubben N et al (2009) Ageing-related chromatin defects through loss of the NURD complex. Nat Cell Biol 11(10):1261–1267
Pertz O, Hahn KM (2004) Designing biosensors for Rho family proteins—deciphering the dynamics of Rho family GTPase activation in living cells. J Cell Sci 117(Pt 8):1313–1318
Peters AH, O’Carroll D et al (2001) Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107(3):323–337
Piston DW, Kremers GJ (2007) Fluorescent protein FRET: the good, the bad and the ugly. Trends Biochem Sci 32(9):407–414
Post WS, Goldschmidt-Clermont PJ et al (1999) Methylation of the estrogen receptor gene is associated with aging and atherosclerosis in the cardiovascular system. Cardiovasc Res 43(4):985–991
Rea S, Eisenhaber F et al (2000) Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406(6796):593–599
Redon C, Pilch D et al (2002) Histone H2A variants H2AX and H2AZ. Curr Opin Genet Dev 12(2):162–169
Rizzo MA, Springer GH et al (2004) An improved cyan fluorescent protein variant useful for FRET. Nat Biotechnol 22(4):445–449
Romoser VA, Hinkle PM et al (1997) Detection in living cells of Ca2+-dependent changes in the fluorescence emission of an indicator composed of two green fluorescent protein variants linked by a calmodulin-binding sequence. A new class of fluorescent indicators. J Biol Chem 272(20):13270–13274
Sarkies P, Sale JE (2012) Cellular epigenetic stability and cancer. Trends Genet 28(3):118–127
Sasaki K, Ito T et al (2009) Real-time imaging of histone H4 hyperacetylation in living cells. Proc Natl Acad Sci U S A 106(38):16257–16262
Scaffidi P, Misteli T (2005) Reversal of the cellular phenotype in the premature aging disease Hutchinson-Gilford progeria syndrome. Nat Med 11(4):440–445
Scaffidi P, Misteli T (2006) Lamin A-dependent nuclear defects in human aging. Science 312(5776):1059–1063
Schotta G, Lachner M et al (2004) A silencing pathway to induce H3-K9 and H4-K20 trimethylation at constitutive heterochromatin. Genes Dev 18(11):1251–1262
Schulze JM, Jackson J et al (2009) Linking cell cycle to histone modifications: SBF and H2B monoubiquitination machinery and cell-cycle regulation of H3K79 dimethylation. Mol Cell 35(5):626–641
Smith MM (2002) Centromeres and variant histones: what, where, when and why? Curr Opin Cell Biol 14(3):279–285
Stein S, Matter CM (2011) Protective roles of SIRT1 in atherosclerosis. Cell Cycle 10(4):640–647
Stewart MD, Li J et al (2005) Relationship between histone H3 lysine 9 methylation, transcription repression, and heterochromatin protein 1 recruitment. Mol Cell Biol 25(7):2525–2538
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403(6765):41–45
Sugiyama K, Sugiura K, Hara T, Sugimoto K, Shima H, Honda K, Furukawa K, Yamashita S, Urano T (2002) Aurora-B associated protein phosphatases as negative regulators of kinase activation. Oncogene 21:3103–3111
Swift J, Ivanovska IL et al (2013) Nuclear lamin-A scales with tissue stiffness and enhances matrix-directed differentiation. Science 341(6149):1240104
Tan Y, Tajik A et al (2014) Matrix softness regulates plasticity of tumour-repopulating cells via H3K9 demethylation and Sox2 expression. Nat Commun 5:4619
Trojer P, Reinberg D (2007) Facultative heterochromatin: is there a distinctive molecular signature? Mol Cell 28(1):1–13
Vermeulen M, Eberl HC et al (2010) Quantitative interaction proteomics and genome-wide profiling of epigenetic histone marks and their readers. Cell 142(6):967–980
Wang F, Higgins JM (2013) Histone modifications and mitosis: countermarks, landmarks, and bookmarks. Trends Cell Biol 23(4):175–184
Wang Y, Wang N (2009) FRET and mechanobiology. Integr Biol (Camb) 1(10):565–573
Wang Y, Botvinick EL et al (2005) Visualizing the mechanical activation of Src. Nature 434(7036):1040–1045
Wang Y, Shyy JY et al (2008) Fluorescence proteins, live-cell imaging, and mechanobiology: seeing is believing. Annu Rev Biomed Eng 10:1–38
Weintraub H, Groudine M (1976) Chromosomal subunits in active genes have an altered conformation. Science 193(4256):848–856
Wilkins BJ, Rall NA et al (2014) A cascade of histone modifications induces chromatin condensation in mitosis. Science 343(6166):77–80
Xhemalce B, Dawson MA et al (2011) Histone modifications. In: Meyers RA (ed) Encyclopedia of molecular cell biology and molecular medicine, epigenetic regulation and epigenomics, 2nd edn. Wiley, New York, pp 1–45
Yoshizaki H, Ohba Y et al (2003) Activity of Rho-family GTPases during cell division as visualized with FRET-based probes. J Cell Biol 162(2):223–232
Zaccolo M, De Giorgi F et al (2000) A genetically encoded, fluorescent indicator for cyclic AMP in living cells. Nat Cell Biol 2(1):25–29
Zeng L, Zhang Y et al (2003) The role of p53 deacetylation in p21Waf1 regulation by laminar flow. J Biol Chem 278(27):24594–24599
Zhang Y, Reinberg D (2001) Transcription regulation by histone methylation: interplay between different covalent modifications of the core histone tails. Genes Dev 15(18):2343–2360
Zhou J, Li YS et al (2014) Epigenetic mechanism in regulation of endothelial function by disturbed flow: induction of DNA hypermethylation by DNMT1. Cell Mol Bioeng 7(2):218–224
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 The American Physiological Society
About this chapter
Cite this chapter
Peng, Q., Cheng, B., Lu, S., Chien, S., Wang, Y. (2016). Perspectives of FRET Imaging to Study Epigenetics and Mechanobiology in the Nucleus. In: Chien, S., Engler, A., Wang, P. (eds) Molecular and Cellular Mechanobiology. Physiology in Health and Disease. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-5617-3_7
Download citation
DOI: https://doi.org/10.1007/978-1-4939-5617-3_7
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-5615-9
Online ISBN: 978-1-4939-5617-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)