Abstract
Purpose of Review
Although extensive research has been conducted on microbial resilience, numerous unanswered questions persist. In this study, we highlight impactful research that elucidates the diverse mechanisms underlying the resilience of the gut microbiota against pathogen colonization and its implications on gut health and disease.
Recent Findings
The increasing importance of gut microbiota resistance in the context of pathogenic infections has been extensively reported. The establishment of a homeostatic microbiome-host interaction, facilitated by intricate mechanisms originating from both the microbiota and the host, plays a crucial role in fostering resilience. However, pathogens have evolved several evasion strategies that can disrupt healthy microbiota composition, trigger environmental alterations, and induce inflammation, thereby potentially exacerbating inflammatory diseases in the gut.
Summary
In this review, we aim to highlight the significance of different resilience mechanisms during intestinal infections and their potential for modulation to develop new interventions that can effectively ameliorate Inflammatory Bowel Disease (IBD).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The discovery of the gut microbiota dates back to the pre-1900 era, when a plethora of microorganisms, which are abundantly present in the human body, were identified [1]. While microbiota and microbiome are terms that are often used interchangeably, they exhibit certain differences. Microbiota is comprised of living microorganisms that exist within a particular environment, such as the gut microbiota. On the other hand, the microbiome encompasses the collection of genomes in the environment, including microorganisms and metabolites. Consequently, the microbiome has a broader scope than the microbiota [2].
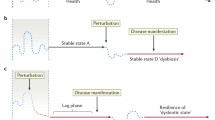
The microbiome is a complex ecosystem with diverse and variable compositions. The main phyla present in healthy humans are Firmicutes and Bacteroidetes, which represent 90% of the microbiota [3]. The extensive diversity observed in microbiomes complicates the task of defining a precise criterion for a natural or healthy microbiota. However, in the definitions, certain factors, such as increased diversity, symbiotic interactions with the host, stability, and resilience, which execute immune and metabolic functions, can be considered in the definitions [4]. The human microbiota is consistently influenced by the host and multiple external factors, including diet, exercise levels, medicine, antibiotics, genetics, immunity, and the intestinal barrier, which have the potential to perturb this ecosystem [5, 6]. Subsequently, the microbial ecosystem may or may not be able to revert to its original state; this phenomenon is referred to as resilience. The concept of resilience was recently proposed through a model that integrated tests and challenges to determine whether the microbiota comprises resilience, which was assessed by its capability to resist and recover from stress [7].
The development of chronic diseases can plausibly initiate a decline in the capacity of the microbiota to effectively resist challenges or promptly and fully recover to a state of homeostasis. This can potentially lead to a new state of dysbiosis [7]. Dysbiosis occurs when there is a critical involvement of the microbiota in the development of diseases [8]. Under certain conditions, patients treated with antibiotics exhibit a diminished ability to restore the microbiota to its baseline levels, which can contribute to the development of diseases [9]. Nevertheless, a highly resilient microbiota possesses the capacity to regain a healthy state [10]. Hryckowian et al. described diet as an intervention that increases microbiota diversity and resilience [11]. Thus, the maintenance of the microbiota in a resilient state may substantially contribute to homeostasis and health. In this review, we focus on resident microbiomes and the importance of microbiota resilience in health and disease management.
Inherent Mechanisms of Microbial Resilience
The gut microbiota is a collection of bacteria present in the intestinal tract. They exhibit distinct spatial organization based on the location, including the lumen, mucus layer, and crypts, which determine their interaction levels with each other and with the host [12].
Spatial and Nutritional Competition
The disparity in location and microbial density are features that are influenced by oxygen levels, nutrient availability, pH, and immune factors, and these variables have the potential to affect the gut microbial composition and spatial location [13]. Bacterial distribution within the lumen is not uniform; in the small intestine, for instance, the transit is faster, and and the sugar availability favors Proteobacteria and Lactobacillales expansion, which are rapidly dividing facultative anaerobes [14]. Considering the spatial structure of the colon microbiota, the phyla Firmicutes and Proteobacteria are the most predominant in the crypts and mucosa, while Actinobacteria and Bacteroides occur at lower levels [15]. Additionally, the microbiota in the small and large intestines rely on different nutrients for their developmental functions. In the large intestine, bacteria utilize nutrients that have not been absorbed in the upper gastrointestinal tract [16]. Furthermore, several studies have shown the nutritional competition between commensal and pathogenic bacteria, for instance, indigenous Escherichia coli has been reported to compete for amino acids and other nutrients with enterohemorrhagic E. coli, which causes morbidity [17,18,19]. Studies exploring competition for nutrients exhibited by the gut microbiota, as a mechanism to combat pathogens, are still relatively recent; however, it is evident that this competition is related to groups of commensal bacteria that are directly related to these pathogens in terms of their metabolism. For instance, in the case of mice infected with Citrobacter rodentium, the infection can be controlled when the mice are colonized with E. coli, which engages in nutritional competition [20].
Inhibitory Metabolites
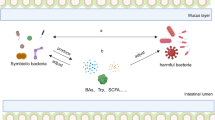
Compounds in the diet are metabolized by digestive enzymes in the upper gastrointestinal tract. However, some of these metabolites remain intact and are processed and metabolized by the gut microbiota [21]. The gut microbiota may interact either with the host directly, through metabolites provided by the bacteria, or by the transformation of diet-derived substrates, which are small molecules representing products of microbial metabolism [22]. These metabolites may affect the host (e.g., short-chain fatty acids (SCFA) and vitamins), other commensal bacteria (e.g., bacteriocins), and pathogens (e.g., lactic acid and hydrogen peroxide) [23]. These bacterial products can act as growth inhibitors for competitors. Additionally, the antibiotic effect exhibited by lactic acid-producing bacteria is attributed to the production of lactic acid through glucose fermentation. This process leads to alterations in the pH, which in turn acts as a protective mechanism against potential pathogen invasions, ultimately leading to oxidative stress and cell death [24]. The production of hydrogen peroxide (H2O2) by lactic acid bacteria, particularly within the Lactobacillus clade, is a commonly observed phenomenon, especially when exposed to aerobic conditions. This metabolic trait enables them to generate substantial amounts of H2O2 [25]. Additionally, the presence of both lactic acid and hydrogen peroxide not only influences the composition of microbial communities but also induces heightened elimination of Salmonella typhimurium by inducing DNA damage [26]. Similar to lactic acid, bacteriocins, which are proteins that induce cell death by altering membrane potential, exhibit a synergistic effect on the microbiome and cause its disruption [27]. These findings demonstrate the direct impact of microbiota metabolites on microbial resilience, acting as controllers and inhibitors of other pathogenic bacteria.
Host-Associated Mechanisms of Resilience
Besides the inherent colonization resistance displayed by microbiota components, there are pivotal interactions with the host that can limit the invasion and proliferation of pathogens. In this section, we will discuss the role of the physical barrier (e.g., mucous layer and epithelial cells) along with the oxygen gradient and components of the immune system, in the modulation of microbial resilience in the gut.
Oxygen Gradient
Since bacteria consistently sense the surrounding environment, the available oxygen becomes a key signal and energy source for their metabolism and fitness. Enteric bacterial composition and resilience are specifically related to the gut oxygen gradient since their metabolic activity can reduce free oxygen in the lumen, leading to the prevalence of an anaerobic environment [28]. For instance, the SCFA butyrate produced by Clostridia can boost aerobic respiration in epithelial cells by enhancing the beta-oxidation pathway and causing the lowering of the oxygen concentration [29]. The oxygen levels increased after Clostridia depletion, which can enable S. typhimurium proliferation [30]. In addition, it was demonstrated that, depending on the oxygen levels, Enterobacteriaceae can contribute to colonization resilience by competing with S. entereditis [28, 31]. Complementary, the resistance is lost when the capacity to respire oxygen under micro-aerophilic conditions is genetically ablated in E. coli [31]. Similarly, Mucispirillum schaedler, another microbiota component, competes for nitrate in the absence of oxygen, limiting the proliferation of E. coli and S. typhimurium which depend on nitrate metabolism to succeed during gut inflammation [32]. Hence, the capacity of the symbiont microbiota to generate a hypoxic environment in the intestine largely restricts the virulence potential and invasion of pathogenic players.
Mucus Layer
The intestinal barrier consists of an inner and an outer layer layers, the epithelial barrier, and the immune cells. Goblet cells secrete mucus, which consists of a combination of highly glycosylated proteins, with mucin 2 (MUC2) being the most abundant and crucial component for mucus layer organization in the colon [33]. Its importance was highlighted when studies demonstrated that mice lacking MUC2 spontaneously developed colitis [34], showed a predisposition to inflammation-dependent colorectal cancer development [35], and heightened susceptibility to Citrobacter rodentium and Listeria monocytogenes [36, 37, 38•]. These findings provide a plausible explanation for the development of inflammatory diseases in the absence or dysfunction of the mucus layer, which typically plays a crucial role in preventing pathogen-driven inflammation. Complementarily, microbiota components have been reported to play a fundamental role in mucus production. For instance, germ-free (GF) mice exhibit a thinner mucus layer [39] and the administration of a fiber-free diet to gnotobiotic mice hosting a simple microbiota set leads to higher C rodentium proliferation and epithelial invasion [40•]. The diet also exerts a direct effect on the mucus layer thickness. Gnotobiotic mice fed with a Western diet low in microbiota-accessible carbohydrates (MACs), fostered the growth of mucus-degrading bacteria, including Akkermansia muciniphila and Bacteroides caccae. As a result, these bacteria consumed the outer mucus layer, thereby reducing the space between the microbiota community and epithelial cells. Intrinsically, the host increased the MUC2 expression, however, it was not sufficient for avoiding pathogen invasion [41]. Only with Bifidobacterium longum administration could the outer mucus layer damage be reversed, possibly because of its potential capacity to stimulate mucus production [42]. Together, this evidence suggests an intrinsic balance between the functional state of the mucus layer and the microbiota composition.
The Epithelial Cells
Intestinal epithelial cells (IECs), which include goblet cells, as well as enteroendocrine and Paneth cells, possess specialized functions aimed at preserving the digestive and barrier functions of the epithelium [43]. The enteroendocrine population acts as a fundamental network between the central and enteric neuroendocrine systems by releasing a wide range of hormone regulators. As mentioned above, goblet cells secrete mucins, whereas Paneth cells are responsible for antimicrobial protein (AMPs) production, constituting a physical and biochemical barrier that prevents microbial contact with the epithelial surface [43, 44]. The most relevant and recognized AMPs produced by enterocytes are the C-type lectin regenerating islet-derived protein IIIγ (RegIIIγ), found throughout the small intestine and colon. Moreover, Paneth cells have been reported to secrete additional proteins such as defensins, lysozyme, and cathelicidins in the crypts of the small intestine [43, 45]. By disrupting the conserved features of microbial biology, AMPs allow the modulation of commensal and pathogenic bacteria in the intestinal tract. Microbiota components possess the capacity to regulate the production of AMPs, thus exerting an influence on microbial abundance [46] and playing a pivotal role in colonization resistance. A study demonstrated that microbiota depletion by antibiotics administration in mice caused a decrease in the RegIIIγ expression, which was restored after stimulation with a synthetic ligand for Toll-like receptor 7 (TLR7) [47]. This directly impacted the host’s capacity to control vancomycin-resistant enterococci (VRE). Likewise, other studies have demonstrated that Nucleotide Binding Oligomerization Domain Containing 1 (NOD1) and NOD2 signaling through receptor-interacting serine–threonine-protein kinase 2 (RIPK2) can limit C. rodentium expansion and colonization by stimulation of RegIIIγ production during the early stages of infection [48]. In addition, microbiota-derived peptidoglycans stimulate NOD2 and lead to crypt expression, which facilitates L. monocytogenes clearance in mice [49]. Epithelial cells also act as sentinels through the expression of pattern-recognition receptors (PRR), including members of the Toll-like receptor (TLR) [50], NOD-like receptor (NOD) [51], and Retinoic acid-inducible gene I (RIG-I)-like receptor (RLR) [52]. Through signaling cascades that culminate in the production of several mediators, these receptors are pivotal for maintaining homeostasis between the host immune system and symbionts, recognizing pathogens, and initiating host defense and inflammation. The crosstalk between sensing molecules, the immune system, and microbiota resilience is discussed in detail in the next section.
Immune System Messengers
The intestinal microbiota also plays an essential role in orchestrating host immunity during homeostasis and disease. Cytokine signaling plays a vital role in host-microbiota interactions and serves to restrict pathogen invasion (Fig. 1). The primary producers of Interleukin (IL)-22 are lymphocytes, such as T helper (Th) 1, Th17, Th22, CD8+ T cells, γδ T cells, natural killer cells (NK), and type 3 innate lymphoid (ILC3) cells [53]. This cytokine is expressed in response to proinflammatory cytokines, such as IL-1β, IL-6, Tumor Necrosis Factor (TNF)-α, and mainly IL-23, produced by myeloid cells upon perception of microbial signals [54]. It has been reported that IL-22 production by Th17 cells is triggered by the presence of segmented filamentous bacteria (SFB) and promotes lower susceptibility to C. rodentium infection [55]. Additionally, colonization of GF mice with human microbiota components promotes IL-22 production, enhancing host glycosylation and the consequent growth of Phascolarctobacterium species, which in turn compete with C. difficile for succinate in the gut [56••]. Several studies have reported the role of IL-22 in promoting intestinal barrier function and altering the composition of the gut microbiota [55, 57,58,59], however, it has also been shown that this cytokine may favor the proliferation of some pathogens, such as S. tryphimurium, over bacterial symbionts [60]. Notably, microbes can manipulate the host to achieve a competitive advantage within the intestinal microbiome community. Bacteroides fragilis can act via TLR2 activation on T helper (Th) cells to establish symbiosis with the host, which was proved by the fact that the TLR2 deletion in CD4 + T cells drives an antimicrobial response that limits B. fragilis colonization [61]. Gut microbiota also enhances the priming of macrophages and IL-1β production to immediately respond to S. tryphimurium and P. aeruginosa infections, assisting neutrophil recruitment and further pathogen clearance [62]. A recent study showed that a mouse commensal E. coli isolate protected mice from C. rodentium infection and dextran sulfate sodium (DSS)-induced colitis by expansion IL-1B producing CX3CR1 + mononuclear phagocytes and IL-22-secreting type 3 innate lymphoid cells (ILC3) [63••]. A similar human commensal E. coli isolate also protected mice from infection and colitis [63••], revealing a surprising role for microbiota-mediated IL-1β secretion to endorse intestinal barrier repair.
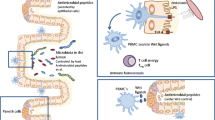
Host-associated mechanisms of resilience. Several host features facilitate the diversity of mechanisms that cover the microbiota’s resilience against pathogen invasions, including the low level of oxygen in the mucosa as a result of an interaction between commensals and epithelial cells; secretion of mucins by goblet cells compounding the mucus layer; secretion of AMPs and defensins by Paneth cells; cytokine production by innate immune cells that can activate and induce the differentiation of different types of lymphocytes, which together with ILCs produce key cytokines that control the exacerbated proliferation of microbiome components; secretion of IgA by plasma cells, which can also help controlling the expansion of prominent species in a balanced microbiota. AMPs: Antimicrobial peptides; ILCs: Innate lymphoid cells; IgA: Immunoglobulin A
Similarly, butyrate-producing bacteria can limit pathogen colonization by regulating tight junction protein expression via IL-10 signaling. IL-10 is secreted by several immune cells, the major producers of which are dendritic cells, macrophages, T cells, and ILC2 cells [64]. IL-10 can also promote intestinal Treg differentiation and maintenance, which is essential for preventing microbial-derived inflammation in the gut by controlling excessive effector T cell responses [65]. Studies have shown that diverse groups of symbionts can promote the expansion of Tregs from naïve T cells in the gut, thereby protecting mice from the inflammation caused by pathogenic invasion. For example, B. fragilis [66], Bifidobacterium infantis [67], and defined sets of bacteria including Firmicutes induce Treg expansion and reinforce intestinal barriers through several mechanisms [68]. Various studies have demonstrated differences in the microbiota composition between wild-type (WT) and IL-10-lacking mice. Using GF mice, Maharshak et al. [69] showed that IL-10−/− mice exhibited a decrease in microbiota complexity over time, however, this was not observed in wild-type littermates. Hence, E. coli is enhanced over time in IL-10−/− mice, converging with spontaneous inflammation and the initiation of colitis [69]. In summary, these studies highlighted the mechanisms employed by microbiota to promote cytokine production by immune components to maintain their colonization and resilience while facing pathogen invasion.
In addition to cytokines, the production of immunoglobulins is influenced by the gut microbiota, which can indirectly affect colonization fitness. Secretory immunoglobulin A (SIgA), produced by plasma cells, is the most prevalent isotype in the human intestinal lumen and is involved in the prevention of infections and maintenance of symbiont homeostasis. Fadlallah et al. [70] showed that co-dependent associations between commensals are disturbed in patients who exhibit deficiency in SIgA, which provides evidence for the participation of SIgA in the interactions between microbiota components [70]. Similarly, in mice, the binding of highly glycosylated SIgA to Bacteroides altered microbial metabolism and led to an indirect expansion of Clostridiales, culminating in impaired development of DSS colitis [71]. SIgA also induces the surveillance of B. fragilis in monocolonized mice [72] while promoting mutualistic associations between the host and possible pathogenic fungal symbionts by targeting and restraining virulent morphotypes. These evidence indicate the relevance of SIgA in modulating pathogen colonization and fostering ecological relationships between microbiota components [73].
Disruption of Microbiota Resilience by Pathogens
Despite multiple strategies developed through host-microbiota interactions, colonization resistance can be disrupted by pathogens. Through virulence programs, pathogens subvert the homeostasis of the healthy microbiota, impair barrier function, and trigger inflammation. The importance of pathogen virulence in combating microbiota resilience is emphasized by the integration of several environmental signals that dictate when and where to express virulence genes. For instance, the enhanced virulence of C. rodentium is essential for intestinal colonization in conventionally reared mice, but not in GF mice [20]. In addition, the proliferation of pathogens in diseases directly reflects the pathogen-microbiota interaction, since alterations in the physiological environment, such as oxygen levels and metabolic and nutritional settings, favor the growth of invading microbes over the commensals [74, 75]. The evasion and virulence strategies of pathogens and their consequences for health and disease are discussed in the next section.
Pathogen Evasion Strategies
As symbiont components, pathogens can successfully colonize the host through a virulence setting that modulates spatial niche construction and promotes their growth over the microbiota. Facultative anaerobic pathogens, including S. typhimurium, C. rodentium, and other Enterobacteriaceae, can create a physiologically challenging environment for symbiotic bacteria by modulating oxygen levels in the large intestine. As the majority of the microbes in the colon are fermenters and highly oxygen-sensitive, virulent programs that induce an increase in oxygen levels lead to a reduction in these components, facilitating pathogen invasion and growth [76]. For example, expression of the toxin type III secretion system (T3SS) by C. rodentium leads to crypt hyperplasia in mice, decreasing overall oxygen consumption through changes in epithelial metabolism [77]. Consequently, this increased oxygenation of the mucosa drives aerobic C. rodentium proliferation in the colon, favoring infection. It was recently reported that T3SS-mediated intimate attachment also enables the oxidation of hydrogen peroxide by C. rodentium, which is produced by epithelial NADPH oxidase (NOX1) even before crypt hyperplasia [78••]. As mentioned previously, one of the inherent strategies of the microbiota to resist pathogen colonization is its capacity, together with the epithelium, for rapid nutrient consumption. However, some pathogens can circumvent this obstacle by evading microbiota resilience and colonizing the host. For example, enteropathogenic E. coli (EPEC) can obtain nutrients from infected host cells via host nutrient extraction (HNE). Through an inner membrane complex, EPEC can protrude the structure of membranous nanotubes directly into host cells and draw on their nutrients [79]. Conversely, other pathogens have evolved the capacity to use alternative nutrient sources, such as C. rodentium, which can use diet-derived metabolites produced by commensal bacteria for initial proliferation and T3SS-driven inflammation [80]. Triggering inflammation is also a widely used strategy employed by pathogens [81, 82]. For example, S. typhimurium uses the Spi1 T3SS to induce the expression of inflammatory signals in the ileum and cecum of mice. Through the host’s generation of reactive oxygen and nitrogen species, S. typhimurium utilizes the byproducts, such as tetrathionate, nitrate, and oxidized sugars for respiration and proliferation [83, 84]. Together, this evidence exemplifies several specialized strategies to overcome the colonization resistance dictated by the microbiota (Fig. 2).
The commensal microbiota colonization resistance is constantly challenged by external factors such as pathogens, which in turn develop strategies to modulate environmental factors, including oxygen levels and nutrient disposition. They also trigger inflammation, leading to physiological changes that impair the commensal’s growth. However, a healthy microbiota can resist and recover from these attacks, as well as restore its original composition. In certain individuals, plausibly due to genetic alterations, the microbiota cannot be restored and the resilience is disrupted, establishing dysbiosis and leading to the development and worsening of the disease
Microbiota Resilience Disruption
In addition to pathogen infection, other external stress factors such as severe dietary changes, antibiotics, and other medications can perturb the balanced ecosystem achieved by healthy microbiota. Consequently, the gut microbiome may return to its original shape. A resilient microbiota can return to its baseline equilibrium employing the various strategies discussed earlier, whereas a non-resilient microbiota is molded to acquire a new composition (Fig. 2). It is assumed that a healthy individual has a resilient microbiota that can rapidly return to its steady state after exposure to unavoidable environmental stressors [7, 10, 85]. An impaired ability to resist these challenges facilitates dysbiosis, which is associated with several chronic diseases. Specifically, in the gut, inflammatory bowel diseases (IBD), including Crohn’s disease and ulcerative colitis, are intimately associated with the disruption of healthy microbiota. Although several factors, including genetics, can be associated with IBD pathogenesis, the composition of the gut microbiome is thought to be an essential determinant of host susceptibility to IBD, since it can contribute to aberrant immune responses through multiple mechanisms d [86, 87]. Similarly, the transfer of IBD-associated microbiota into healthy mice induces intestinal inflammation [88], while microbiota depletion by antibiotic administration enhances intestinal inflammation in IBD-susceptible mouse models. However, the colonization of pathobionts, such as adherent-invasive E. coli (AIEC) [89••], Enterococcus faecium [90], enterotoxigenic Bacteroides ragilis [91], and multi-drug resistant Klebsiella pneumoniae (Kp) [92•] is related to IBD worsening in genetically susceptible individuals. Therefore, interventions that improve microbiome resilience are indispensable for disease amelioration. Understanding the features of resilient microbiota will assist in the conception of interventions aimed at enhancing resilience. In this way, dietary fibers can be a contributing factor in increasing microbiota diversity and, consequently, a more resilient one. Mice fed a fiber-enriched diet that was challenged with antibiotics and C. difficile returned to the pre-challenged composition, whereas mice fed a low-fiber diet did not [11], indicating that fibers have a direct effect on improving microbiota resilience in mice. Hence, the administration of probiotics containing species with anti-inflammatory features, such as Lactobacillus rhamnosus [93], or those that can improve the gut barrier, such as Lactobacillus plantarum [94], has been reported to be potential intervention strategies that can enhance microbiota resilience. Meanwhile, probiotic and dietary interventions that aim to boost certain species need to be carefully administered to avoid targeted components from becoming more prominent leading to a negative impact on the microbiome diversity [7]. Recently, Federici et al. [95••] demonstrated that IBD-associated Kp strains aggravate intestinal inflammation in colitis-prone, germ-free, and colonized mice, which is reversed using a lytic five-phage combination targeting Kp. They also assessed the proof-of-concept of Kp-targeting phages in an artificial human gut and healthy volunteers, demonstrating a feasible oral administration therapy that improves microbial resistance to pathobionts expansion [95••]. Together, these studies highlight the importance of colonization-resistant microbiota in health and disease and the emergence of combined therapies focused on reestablishing resilience.
Conclusion
In this review, we summarize the main strategies used by symbiotic bacteria to remain resilient against pathogen attacks throughout a host’s life. The inherent ecological and physiological characteristics of symbionts has the ability to confer a high degree of equilibrium between all microbiota components. However, the modulation of the host’s features, such as the mucosal barrier and immune system has been crucial for a closer host-microbiota relationship built through thousands of years of evolution. Its importance for human health has been highlighted in several studies showing the mechanisms underlying the disturbance of microbial resilience. However, a complete understanding of the role of microbiota resilience in health and disease remains a contentious subject, as experimental validation of models and further investigation in humans are required. In summary, as greater insights are acquired regarding microbiota resilience, innovative integrated interventions may be applied in the treatment of several microbiota-associated diseases.
Data Availability
N/A.
Abbreviations
- IBD :
-
Inflammatory bowel disease
- SCFA :
-
Short-chain fatty acids
- MUC2 :
-
Mucin 2
- GF :
-
Germ-free
- MACs :
-
Microbiota-accessible carbohydrates
- IECs :
-
intestinal epithelial cells
- RegIIIγ :
-
Regenerating islet-derived protein IIIγ
- AMPs :
-
Antimicrobial proteins
- VRE :
-
Vancomycin-resistant enterococcus
- RIPK2 :
-
Receptor-interacting serine–threonine-protein kinase 2
- TLR :
-
Toll-like receptor
- NOD :
-
NOD-like receptor
- RLR :
-
RIG-I like receptor
- PRR :
-
Pattern-recognition receptors
- SFB :
-
Segmented filamentous bacteria
- IL :
-
Interleukin
- Th :
-
T helper
- CX3CR1 :
-
C-X3-C Motif Chemokine Receptor 1
- ILCs :
-
Innate lymphoid cells
- DSS :
-
Dextran sulfate sodium
- WT :
-
Wild-type
- SIgA :
-
Secretory immunoglobulin A
- T3SS :
-
The toxin type III secretion system
- NOX1 :
-
NADPH oxidase
- EPEC:
-
Enteropathogenic E. coli
- HNE :
-
Host nutrient extraction
- AIEC :
-
Adherent-invasive E. coli
- Kp :
-
Klebsiella pneumoniae
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Hou K, Wu ZX, Chen XY, Wang JQ, Zhang D, Xiao C, et al. Microbiota in health and diseases. Signal Transduct Target Ther. 2022;7(1):135. https://doi.org/10.1038/s41392-022-00974-4.
Berg G, Rybakova D, Fischer D, Cernava T, Verges MC, Charles T, et al. Microbiome definition re-visited: old concepts and new challenges. Microbiome. 2020;8(1):103. https://doi.org/10.1186/s40168-020-00875-0.
Rinninella E, Raoul P, Cintoni M, Franceschi F, Miggiano GAD, Gasbarrini A, et al. What is the Healthy Gut Microbiota Composition? A Changing Ecosystem across Age, Environment, Diet, and Diseases. Microorganisms. 2019;7(1). https://doi.org/10.3390/microorganisms7010014.
Thursby E, Juge N. Introduction to the human gut microbiota. Biochem J. 2017;474(11):1823–36. https://doi.org/10.1042/BCJ20160510.
Cho I, Yamanishi S, Cox L, Methe BA, Zavadil J, Li K, et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature. 2012;488(7413):621–6. https://doi.org/10.1038/nature11400.
Magzal F, Shochat T, Haimov I, Tamir S, Asraf K, Tuchner-Arieli M, et al. Increased physical activity improves gut microbiota composition and reduces short-chain fatty acid concentrations in older adults with insomnia. Sci Rep. 2022;12(1):2265. https://doi.org/10.1038/s41598-022-05099-w.
Dogra SK, Dore J, Damak S. Gut Microbiota Resilience: Definition, Link to Health and Strategies for Intervention. Front Microbiol. 2020;11:572921. https://doi.org/10.3389/fmicb.2020.572921.
Wilkins LJ, Monga M, Miller AW. Defining Dysbiosis for a Cluster of Chronic Diseases. Sci Rep. 2019;9(1):12918. https://doi.org/10.1038/s41598-019-49452-y.
Dethlefsen L, Relman DA. Incomplete recovery and individualized responses of the human distal gut microbiota to repeated antibiotic perturbation. Proc Natl Acad Sci U S A. 2011;108(Suppl 1):4554–61. https://doi.org/10.1073/pnas.1000087107.
Sommer F, Anderson JM, Bharti R, Raes J, Rosenstiel P. The resilience of the intestinal microbiota influences health and disease. Nat Rev Microbiol. 2017;15(10):630–8. https://doi.org/10.1038/nrmicro.2017.58.
Hryckowian AJ, Van Treuren W, Smits SA, Davis NM, Gardner JO, Bouley DM, et al. Microbiota-accessible carbohydrates suppress Clostridium difficile infection in a murine model. Nat Microbiol. 2018;3(6):662–9. https://doi.org/10.1038/s41564-018-0150-6.
Nguyen J, Pepin DM, Tropini C. Cause or effect? The spatial organization of pathogens and the gut microbiota in disease. Microbes Infect. 2021;23(6–7):104815. https://doi.org/10.1016/j.micinf.2021.104815.
Tropini C, Earle KA, Huang KC, Sonnenburg JL. The Gut Microbiome: Connecting Spatial Organization to Function. Cell Host Microbe. 2017;21(4):433–42. https://doi.org/10.1016/j.chom.2017.03.010.
Gu S, Chen D, Zhang JN, Lv X, Wang K, Duan LP, et al. Bacterial community mapping of the mouse gastrointestinal tract. PLoS ONE. 2013;8(10):e74957. https://doi.org/10.1371/journal.pone.0074957.
Saffarian A, Mulet C, Regnault B, Amiot A, Tran-Van-Nhieu J, Ravel J, et al. Crypt- and Mucosa-Associated Core Microbiotas in Humans and Their Alteration in Colon Cancer Patients. mBio. 2019;10(4). https://doi.org/10.1128/mBio.01315-19.
Rowland I, Gibson G, Heinken A, Scott K, Swann J, Thiele I, et al. Gut microbiota functions: metabolism of nutrients and other food components. Eur J Nutr. 2018;57(1):1–24. https://doi.org/10.1007/s00394-017-1445-8.
Momose Y, Hirayama K, Itoh K. Competition for proline between indigenous Escherichia coli and E. coli O157:H7 in gnotobiotic mice associated with infant intestinal microbiota and its contribution to the colonization resistance against E. coli O157:H7. Antonie Van Leeuwenhoek. 2008;94(2):165–71. https://doi.org/10.1007/s10482-008-9222-6.
Fabich AJ, Jones SA, Chowdhury FZ, Cernosek A, Anderson A, Smalley D, et al. Comparison of carbon nutrition for pathogenic and commensal Escherichia coli strains in the mouse intestine. Infect Immun. 2008;76(3):1143–52. https://doi.org/10.1128/IAI.01386-07.
Leatham MP, Banerjee S, Autieri SM, Mercado-Lubo R, Conway T, Cohen PS. Precolonized human commensal Escherichia coli strains serve as a barrier to E. coli O157:H7 growth in the streptomycin-treated mouse intestine. Infect Immun. 2009;77(7):2876–86. https://doi.org/10.1128/IAI.00059-09.
Kamada N, Kim YG, Sham HP, Vallance BA, Puente JL, Martens EC, et al. Regulated virulence controls the ability of a pathogen to compete with the gut microbiota. Science. 2012;336(6086):1325–9. https://doi.org/10.1126/science.1222195.
Scott KP, Gratz SW, Sheridan PO, Flint HJ, Duncan SH. The influence of diet on the gut microbiota. Pharmacol Res. 2013;69(1):52–60. https://doi.org/10.1016/j.phrs.2012.10.020.
Lamichhane S, Sen P, Dickens AM, Oresic M, Bertram HC. Gut metabolome meets microbiome: A methodological perspective to understand the relationship between host and microbe. Methods. 2018;149:3–12. https://doi.org/10.1016/j.ymeth.2018.04.029.
Ruan W, Engevik MA, Spinler JK, Versalovic J. Healthy Human Gastrointestinal Microbiome: Composition and Function After a Decade of Exploration. Dig Dis Sci. 2020;65(3):695–705. https://doi.org/10.1007/s10620-020-06118-4.
Alakomi HL, Skytta E, Saarela M, Mattila-Sandholm T, Latva-Kala K, Helander IM. Lactic acid permeabilizes gram-negative bacteria by disrupting the outer membrane. Appl Environ Microbiol. 2000;66(5):2001–5. https://doi.org/10.1128/AEM.66.5.2001-2005.2000.
Knaus UG, Hertzberger R, Pircalabioru GG, Yousefi SP, Branco Dos Santos F. Pathogen control at the intestinal mucosa - H(2)O(2) to the rescue. Gut Microbes. 2017;8(1):67–74. https://doi.org/10.1080/19490976.2017.1279378.
Atassi F, Servin AL. Individual and co-operative roles of lactic acid and hydrogen peroxide in the killing activity of enteric strain Lactobacillus johnsonii NCC933 and vaginal strain Lactobacillus gasseri KS120.1 against enteric, uropathogenic and vaginosis-associated pathogens. FEMS Microbiol Lett. 2010;304(1):29–38. https://doi.org/10.1111/j.1574-6968.2009.01887.x.
Okuda K, Zendo T, Sugimoto S, Iwase T, Tajima A, Yamada S, et al. Effects of bacteriocins on methicillin-resistant Staphylococcus aureus biofilm. Antimicrob Agents Chemother. 2013;57(11):5572–9. https://doi.org/10.1128/AAC.00888-13.
Marteyn B, Scorza FB, Sansonetti PJ, Tang C. Breathing life into pathogens: the influence of oxygen on bacterial virulence and host responses in the gastrointestinal tract. Cell Microbiol. 2011;13(2):171–6. https://doi.org/10.1111/j.1462-5822.2010.01549.x.
Kelly CJ, Zheng L, Campbell EL, Saeedi B, Scholz CC, Bayless AJ, et al. Crosstalk between microbiota-derived short-chain fatty acids and intestinal epithelial HIF augments tissue barrier function. Cell Host Microbe. 2015;17(5):662–71. https://doi.org/10.1016/j.chom.2015.03.005.
Rivera-Chavez F, Zhang LF, Faber F, Lopez CA, Byndloss MX, Olsan EE, et al. Depletion of butyrate-producing Clostridia from the gut microbiota drives an aerobic luminal expansion of Salmonella. Cell Host Microbe. 2016;19(4):443–54. https://doi.org/10.1016/j.chom.2016.03.004.
Litvak Y, Mon KKZ, Nguyen H, Chanthavixay G, Liou M, Velazquez EM, et al. Commensal Enterobacteriaceae Protect against Salmonella Colonization through Oxygen Competition. Cell Host Microbez. 2019;25(1):128-39 e5. https://doi.org/10.1016/j.chom.2018.12.003.
Herp S, Brugiroux S, Garzetti D, Ring D, Jochum LM, Beutler M, et al. Mucispirillum schaedleri Antagonizes Salmonella Virulence to Protect Mice against Colitis. Cell Host Microbe. 2019;25(5):681–94 e8. https://doi.org/10.1016/j.chom.2019.03.004.
Hansson GC, Johansson ME. The inner of the two Muc2 mucin-dependent mucus layers in colon is devoid of bacteria. Gut Microbes. 2010;1(1):51–4. https://doi.org/10.4161/gmic.1.1.10470.
Van der Sluis M, De Koning BA, De Bruijn AC, Velcich A, Meijerink JP, Van Goudoever JB, et al. Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection. Gastroenterology. 2006;131(1):117–29. https://doi.org/10.1053/j.gastro.2006.04.020.
Velcich A, Yang W, Heyer J, Fragale A, Nicholas C, Viani S, et al. Colorectal cancer in mice genetically deficient in the mucin Muc2. Science. 2002;295(5560):1726–9. https://doi.org/10.1126/science.1069094.
Bergstrom KS, Kissoon-Singh V, Gibson DL, Ma C, Montero M, Sham HP, et al. Muc2 protects against lethal infectious colitis by disassociating pathogenic and commensal bacteria from the colonic mucosa. PLoS Pathog. 2010;6(5):e1000902. https://doi.org/10.1371/journal.ppat.1000902.
Zarepour M, Bhullar K, Montero M, Ma C, Huang T, Velcich A, et al. The mucin Muc2 limits pathogen burdens and epithelial barrier dysfunction during Salmonella enterica serovar Typhimurium colitis. Infect Immun. 2013;81(10):3672–83. https://doi.org/10.1128/IAI.00854-13.
Zhang T, Sasabe J, Hullahalli K, Sit B, Waldor MK. Increased Listeria monocytogenes Dissemination and Altered Population Dynamics in Muc2-Deficient Mice. Infect Immun. 2021;89(4). https://doi.org/10.1128/IAI.00667-20. This work shows that Muc2 mucin plays a key role in controlling L. monocytogenes colonization, dissemination, and population dynamics.
Johansson ME, Jakobsson HE, Holmen-Larsson J, Schutte A, Ermund A, Rodriguez-Pineiro AM, et al. Normalization of Host Intestinal Mucus Layers Requires Long-Term Microbial Colonization. Cell Host Microbe. 2015;18(5):582–92. https://doi.org/10.1016/j.chom.2015.10.007.
Neumann M, Steimle A, Grant ET, Wolter M, Parrish A, Willieme S, et al. Deprivation of dietary fiber in specific-pathogen-free mice promotes susceptibility to the intestinal mucosal pathogen Citrobacter rodentium. Gut Microbes. 2021;13(1):1966263. https://doi.org/10.1080/19490976.2021.1966263. This study indicates that absence of fibers in Western-style diets impairs the mucus layer thickness and leads to disruption of epithelial barrier through changes in microbiome composition, facilitating the infection of enteropathogens.
Desai MS, Seekatz AM, Koropatkin NM, Kamada N, Hickey CA, Wolter M, et al. A Dietary Fiber-Deprived Gut Microbiota Degrades the Colonic Mucus Barrier and Enhances Pathogen Susceptibility. Cell. 2016;167(5):1339-53 e21. https://doi.org/10.1016/j.cell.2016.10.043.
Schroeder BO, Birchenough GMH, Stahlman M, Arike L, Johansson MEV, Hansson GC, et al. Bifidobacteria or Fiber Protects against Diet-Induced Microbiota-Mediated Colonic Mucus Deterioration. Cell Host Microbe. 2018;23(1):27-40 e7. https://doi.org/10.1016/j.chom.2017.11.004.
Peterson LW, Artis D. Intestinal epithelial cells: regulators of barrier function and immune homeostasis. Nat Rev Immunol. 2014;14(3):141–53. https://doi.org/10.1038/nri3608.
Kim YS, Ho SB. Intestinal goblet cells and mucins in health and disease: recent insights and progress. Curr Gastroenterol Rep. 2010;12(5):319–30. https://doi.org/10.1007/s11894-010-0131-2.
Bevins CL, Salzman NH. Paneth cells, antimicrobial peptides and maintenance of intestinal homeostasis. Nat Rev Microbiol. 2011;9(5):356–68. https://doi.org/10.1038/nrmicro2546.
Bosch TCG, Zasloff M. Antimicrobial Peptides-or How Our Ancestors Learned to Control the Microbiome. mBio. 2021;12(5):e0184721. https://doi.org/10.1128/mBio.01847-21.
Abt MC, Buffie CG, Susac B, Becattini S, Carter RA, Leiner I, et al. TLR-7 activation enhances IL-22-mediated colonization resistance against vancomycin-resistant enterococcus. Sci Transl Med. 2016;8(327):327ra25. https://doi.org/10.1126/scitranslmed.aad6663.
Waldschmitt N, Kitamoto S, Secher T, Zacharioudaki V, Boulard O, Floquet E, et al. The regenerating family member 3 beta instigates IL-17A-mediated neutrophil recruitment downstream of NOD1/2 signalling for controlling colonisation resistance independently of microbiota community structure. Gut. 2019;68(7):1190–9. https://doi.org/10.1136/gutjnl-2018-316757.
Kobayashi KS, Chamaillard M, Ogura Y, Henegariu O, Inohara N, Nunez G, et al. Nod2-dependent regulation of innate and adaptive immunity in the intestinal tract. Science. 2005;307(5710):731–4. https://doi.org/10.1126/science.1104911.
Abreu MT. Toll-like receptor signalling in the intestinal epithelium: how bacterial recognition shapes intestinal function. Nat Rev Immunol. 2010;10(2):131–44. https://doi.org/10.1038/nri2707.
Elinav E, Henao-Mejia J, Flavell RA. Integrative inflammasome activity in the regulation of intestinal mucosal immune responses. Mucosal Immunol. 2013;6(1):4–13. https://doi.org/10.1038/mi.2012.115.
Broquet AH, Hirata Y, McAllister CS, Kagnoff MF. RIG-I/MDA5/MAVS are required to signal a protective IFN response in rotavirus-infected intestinal epithelium. J Immunol. 2011;186(3):1618–26. https://doi.org/10.4049/jimmunol.1002862.
Sabat R, Ouyang W, Wolk K. Therapeutic opportunities of the IL-22-IL-22R1 system. Nat Rev Drug Discov. 2014;13(1):21–38. https://doi.org/10.1038/nrd4176.
Keir M, Yi Y, Lu T, Ghilardi N. The role of IL-22 in intestinal health and disease. J Exp Med. 2020;217(3):e20192195. https://doi.org/10.1084/jem.20192195.
Ivanov II, Atarashi K, Manel N, Brodie EL, Shima T, Karaoz U, et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell. 2009;139(3):485–98. https://doi.org/10.1016/j.cell.2009.09.033.
Nagao-Kitamoto H, Leslie JL, Kitamoto S, Jin C, Thomsson KA, Gillilland MG 3rd, et al. Interleukin-22-mediated host glycosylation prevents Clostridioides difficile infection by modulating the metabolic activity of the gut microbiota. Nat Med. 2020;26(4):608–17. https://doi.org/10.1038/s41591-020-0764-0. The authors demonstrated that IL-22, induced by colonization of the gut microbiota, is crucial for the prevention of C. difficile infection in human microbiota-associated (HMA) mice.
Zheng Y, Valdez PA, Danilenko DM, Hu Y, Sa SM, Gong Q, et al. Interleukin-22 mediates early host defense against attaching and effacing bacterial pathogens. Nat Med. 2008;14(3):282–9. https://doi.org/10.1038/nm1720.
Chen J, Waddell A, Lin YD, Cantorna MT. Dysbiosis caused by vitamin D receptor deficiency confers colonization resistance to Citrobacter rodentium through modulation of innate lymphoid cells. Mucosal Immunol. 2015;8(3):618–26. https://doi.org/10.1038/mi.2014.94.
Neil JA, Matsuzawa-Ishimoto Y, Kernbauer-Holzl E, Schuster SL, Sota S, Venzon M, et al. IFN-I and IL-22 mediate protective effects of intestinal viral infection. Nat Microbiol. 2019;4(10):1737–49. https://doi.org/10.1038/s41564-019-0470-1.
Behnsen J, Jellbauer S, Wong CP, Edwards RA, George MD, Ouyang W, et al. The cytokine IL-22 promotes pathogen colonization by suppressing related commensal bacteria. Immunity. 2014;40(2):262–73. https://doi.org/10.1016/j.immuni.2014.01.003.
Round JL, Lee SM, Li J, Tran G, Jabri B, Chatila TA, et al. The Toll-like receptor 2 pathway establishes colonization by a commensal of the human microbiota. Science. 2011;332(6032):974–7. https://doi.org/10.1126/science.1206095.
Franchi L, Kamada N, Nakamura Y, Burberry A, Kuffa P, Suzuki S, et al. NLRC4-driven production of IL-1beta discriminates between pathogenic and commensal bacteria and promotes host intestinal defense. Nat Immunol. 2012;13(5):449–56. https://doi.org/10.1038/ni.2263.
Wu WH, Kim M, Chang LC, Assie A, Saldana-Morales FB, Zegarra-Ruiz DF, et al. Interleukin-1beta secretion induced by mucosa-associated gut commensal bacteria promotes intestinal barrier repair. Gut Microbes. 2022;14(1):2014772. https://doi.org/10.1080/19490976.2021.2014772. This study demonstrated that a commensal E. coli isolate could induce IL-1b production by mononuclear phagocytes and IL-22 by ILC3 cells, improving the epithelial barrier repair after C. rodentium infection.
Kidess E, Kleerebezem M, Brugman S. Colonizing Microbes, IL-10 and IL-22: Keeping the Peace at the Mucosal Surface. Front Microbiol. 2021;12:729053. https://doi.org/10.3389/fmicb.2021.729053.
Josefowicz SZ, Niec RE, Kim HY, Treuting P, Chinen T, Zheng Y, et al. Extrathymically generated regulatory T cells control mucosal TH2 inflammation. Nature. 2012;482(7385):395–9. https://doi.org/10.1038/nature10772.
Round JL, Mazmanian SK. Inducible Foxp3+ regulatory T-cell development by a commensal bacterium of the intestinal microbiota. Proc Natl Acad Sci U S A. 2010;107(27):12204–9. https://doi.org/10.1073/pnas.0909122107.
O’Mahony C, Scully P, O’Mahony D, Murphy S, O’Brien F, Lyons A, et al. Commensal-induced regulatory T cells mediate protection against pathogen-stimulated NF-kappaB activation. PLoS Pathog. 2008;4(8):e1000112. https://doi.org/10.1371/journal.ppat.1000112.
Geuking MB, Cahenzli J, Lawson MA, Ng DC, Slack E, Hapfelmeier S, et al. Intestinal bacterial colonization induces mutualistic regulatory T cell responses. Immunity. 2011;34(5):794–806. https://doi.org/10.1016/j.immuni.2011.03.021.
Maharshak N, Packey CD, Ellermann M, Manick S, Siddle JP, Huh EY, et al. Altered enteric microbiota ecology in interleukin 10-deficient mice during development and progression of intestinal inflammation. Gut Microbes. 2013;4(4):316–24. https://doi.org/10.4161/gmic.25486.
Fadlallah J, El Kafsi H, Sterlin D, Juste C, Parizot C, Dorgham K, et al. Microbial ecology perturbation in human IgA deficiency. Sci Transl Med. 2018;10(439). https://doi.org/10.1126/scitranslmed.aan1217.
Nakajima A, Vogelzang A, Maruya M, Miyajima M, Murata M, Son A, et al. IgA regulates the composition and metabolic function of gut microbiota by promoting symbiosis between bacteria. J Exp Med. 2018;215(8):2019–34. https://doi.org/10.1084/jem.20180427.
Donaldson GP, Ladinsky MS, Yu KB, Sanders JG, Yoo BB, Chou WC, et al. Gut microbiota utilize immunoglobulin A for mucosal colonization. Science. 2018;360(6390):795–800. https://doi.org/10.1126/science.aaq0926.
Ost KS, O’Meara TR, Stephens WZ, Chiaro T, Zhou H, Penman J, et al. Adaptive immunity induces mutualism between commensal eukaryotes. Nature. 2021;596(7870):114–8. https://doi.org/10.1038/s41586-021-03722-w.
Ducarmon QR, Zwittink RD, Hornung BVH, van Schaik W, Young VB, Kuijper EJ. Gut Microbiota and Colonization Resistance against Bacterial Enteric Infection. Microbiol Mol Biol Rev. 2019;83(3). https://doi.org/10.1128/MMBR.00007-19.
Khan I, Bai Y, Zha L, Ullah N, Ullah H, Shah SRH, et al. Mechanism of the Gut Microbiota Colonization Resistance and Enteric Pathogen Infection. Front Cell Infect Microbiol. 2021;11:716299. https://doi.org/10.3389/fcimb.2021.716299.
Caballero-Flores G, Pickard JM, Nunez G. Microbiota-mediated colonization resistance: mechanisms and regulation. Nat Rev Microbiol. 2023;21(6):347–60. https://doi.org/10.1038/s41579-022-00833-.
Lopez CA, Miller BM, Rivera-Chavez F, Velazquez EM, Byndloss MX, Chavez-Arroyo A, et al. Virulence factors enhance Citrobacter rodentium expansion through aerobic respiration. Science. 2016;353(6305):1249–53. https://doi.org/10.1126/science.aag3042.
Miller BM, Liou MJ, Zhang LF, Nguyen H, Litvak Y, Schorr EM, et al. Anaerobic Respiration of NOX1-Derived Hydrogen Peroxide Licenses Bacterial Growth at the Colonic Surface. Cell Host Microbe. 2020;28(6):789-97 e5. https://doi.org/10.1016/j.chom.2020.10.009. The authors describes one of the several evasion strategies of C. rodentium in which host NOX1-derived H2O2 fuels its anaerobic growth during early infection, while the pathogen relies on aerobic respiration for its expansion in the inflamed gut.
Pal RR, Baidya AK, Mamou G, Bhattacharya S, Socol Y, Kobi S, et al. Pathogenic E coli Extracts Nutrients from Infected Host Cells Utilizing Injectisome Components. Cell. 2019;177(3):683-96 e18. https://doi.org/10.1016/j.cell.2019.02.022.
Jimenez AG, Ellermann M, Abbott W, Sperandio V. Diet-derived galacturonic acid regulates virulence and intestinal colonization in enterohaemorrhagic Escherichia coli and Citrobacter rodentium. Nat Microbiol. 2020;5(2):368–78. https://doi.org/10.1038/s41564-019-0641-0.
Lupp C, Robertson ML, Wickham ME, Sekirov I, Champion OL, Gaynor EC, et al. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe. 2007;2(3):204. https://doi.org/10.1016/j.chom.2007.08.002.
Stecher B, Robbiani R, Walker AW, Westendorf AM, Barthel M, Kremer M, et al. Salmonella enterica serovar typhimurium exploits inflammation to compete with the intestinal microbiota. PLoS Biol. 2007;5(10):2177–89. https://doi.org/10.1371/journal.pbio.0050244.
Lopez CA, Winter SE, Rivera-Chavez F, Xavier MN, Poon V, Nuccio SP, et al. Phage-mediated acquisition of a type III secreted effector protein boosts growth of salmonella by nitrate respiration. mBio. 2012;3(3). https://doi.org/10.1128/mBio.00143-12.
Faber F, Tran L, Byndloss MX, Lopez CA, Velazquez EM, Kerrinnes T, et al. Host-mediated sugar oxidation promotes post-antibiotic pathogen expansion. Nature. 2016;534(7609):697–9. https://doi.org/10.1038/nature18597.
Taroncher-Oldenburg G, Jones S, Blaser M, Bonneau R, Christey P, Clemente JC, et al. Translating microbiome futures. Nat Biotechnol. 2018;36(11):1037–42. https://doi.org/10.1038/nbt.4287.
Neurath MF. Host-microbiota interactions in inflammatory bowel disease. Nat Rev Gastroenterol Hepatol. 2020;17(2):76–7. https://doi.org/10.1038/s41575-019-0248-1.
Caruso R, Lo BC, Nunez G. Host-microbiota interactions in inflammatory bowel disease. Nat Rev Immunol. 2020;20(7):411–26. https://doi.org/10.1038/s41577-019-0268-7.
Schaubeck M, Clavel T, Calasan J, Lagkouvardos I, Haange SB, Jehmlich N, et al. Dysbiotic gut microbiota causes transmissible Crohn’s disease-like ileitis independent of failure in antimicrobial defence. Gut. 2016;65(2):225–37. https://doi.org/10.1136/gutjnl-2015-309333.
Elhenawy W, Hordienko S, Gould S, Oberc AM, Tsai CN, Hubbard TP, et al. High-throughput fitness screening and transcriptomics identify a role for a type IV secretion system in the pathogenesis of Crohn’s disease-associated Escherichia coli. Nat Commun. 2021;12(1):2032. https://doi.org/10.1038/s41467-021-22306-w. This study reveals the role of type IV secretion system in biofilm formation by adherent-invasive E. coli isolated from Crohn’s disease patients, which was essential in promoting its persistence in the gut.
Seishima J, Iida N, Kitamura K, Yutani M, Wang Z, Seki A, et al. Gut-derived Enterococcus faecium from ulcerative colitis patients promotes colitis in a genetically susceptible mouse host. Genome Biol. 2019;20(1):252. https://doi.org/10.1186/s13059-019-1879-9.
Zamani S, HesamShariati S, Zali MR, AsadzadehAghdaei H, SarabiAsiabar A, Bokaie S, et al. Detection of enterotoxigenic Bacteroides fragilis in patients with ulcerative colitis. Gut Pathog. 2017;9:53. https://doi.org/10.1186/s13099-017-0202-0.
Kitamoto S, Nagao-Kitamoto H, Jiao Y, Gillilland MG 3rd, Hayashi A, Imai J, et al. The Intermucosal Connection between the Mouth and Gut in Commensal Pathobiont-Driven Colitis. Cell. 2020;182(2):447-62 e14. https://doi.org/10.1016/j.cell.2020.05.048. This study reported that periodontitis leads to an expansion of oral pathobionts enabling its translocation into gut, where they activate inflammasomes of colonic mononuclear phagocytes and triggering intestinal inflammation.
Capurso L. Thirty Years of Lactobacillus rhamnosus GG: A Review. J Clin Gastroenterol. 2019;53(Suppl 1):S1–41. https://doi.org/10.1097/MCG.0000000000001170.
Karczewski J, Troost FJ, Konings I, Dekker J, Kleerebezem M, Brummer RJ, et al. Regulation of human epithelial tight junction proteins by Lactobacillus plantarum in vivo and protective effects on the epithelial barrier. Am J Physiol Gastrointest Liver Physiol. 2010;298(6):G851–9. https://doi.org/10.1152/ajpgi.00327.2009.
Federici S, Kredo-Russo S, Valdes-Mas R, Kviatcovsky D, Weinstock E, Matiuhin Y, et al. Targeted suppression of human IBD-associated gut microbiota commensals by phage consortia for treatment of intestinal inflammation. Cell. 2022;185(16):2879-98 e24. https://doi.org/10.1016/j.cell.2022.07.003. In this study the authors developed a feasible oral administration of a phage therapy targeting IBD-associated Klebsiella pneumoniae that contributes to amelioration of the disease.
Funding
MC, EMS and NOSC are supported by Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) with grant numbers 2018/24350–4, 2020/14388–4 and 2017/05264–7, respectively.
Author information
Authors and Affiliations
Contributions
MC: Writing—original draft, figure preparation; writing—review and editing.
EMS: Writing—original draft; writing—review and editing.
NOSC: Conceptualization, lead; funding acquisition, supporting; lead; writing—original draft, lead; writing—review and editing, lead.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Topical Collection on Microbe-Microbiome Interactions
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cipelli, M., da Silva, E.M. & Câmara, N.O.S. Gut Microbiota Resilience Mechanisms Against Pathogen Infection and its Role in Inflammatory Bowel Disease. Curr Clin Micro Rpt 10, 187–197 (2023). https://doi.org/10.1007/s40588-023-00207-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40588-023-00207-4