Abstract
RNA sequencing technology has revealed a vast number of small RNAs (sRNAs) involved in plant development, stress management, and crop improvement. With improvements in molecular biology techniques and bioinformatic tools, sRNAs and their regulations have taken a mainstream position in various research fields, including plant science. Moreover, recent findings emphasize the significance of ncRNAs and sRNAs in spliceosome machinery. Non-coding RNAs in plants consist of ribosomal RNAs (rRNAs), transfer RNAs (tRNAs), small nucleolar RNAs (snoRNAs), small nuclear RNAs (snRNAs), microRNAs (miRNAs) (19–25 nt), short interfering RNAs (21–23 nt), and long non-coding RNAs (lncRNAs) (over 200 nt). The miRNAs and siRNAs have similar sizes but originate from structurally different RNA molecules and follow distinct biogenesis pathways with different modes of action. Uncovering their impact on plant developmental patterns and stress regulatory mechanisms assumed paramount importance in plant stress biology, specifically projects focused on crop improvement. Plants constantly face and adapt to environmental stresses that hinder their growth and development. Adverse effects of abiotic stresses, such as drought, heat, cold, and salinity, on plant productivity are well documented. With the onset of environmental stresses, certain sRNAs and ncRNAs take up prominent roles in cellular homeostasis and coordinate the stress responses. This review explores the functions of sRNAs and their interactions during plant development and stress regulation. The theme of this paper becomes pertinent given the challenge of developing crops for future environments.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Spectacular developments in the RNA world in the past few decades have contributed significantly to a better understanding of genomics and transcriptomics. Among them, the implications of non-coding and small RNAs assumed great importance in plant development, specifically tissue patterning, stress regulatory mechanism, and crop improvement, under rapidly changing climatic conditions [17, 92]. The non-coding RNAs (ncRNAs) include ribosomal RNA (rRNA), transfer RNA (tRNA), small nuclear RNA (snRNA) in spliceosome machinery, and regulatory non-coding RNAs. Where regulatory non-coding RNAs are categorized into micro RNAs (miRNAs, 19-25nt), small interfering RNAs (siRNAs, 21–23nt), and long non-coding RNA (lncRNAs, > 200 nt) in plants [7, 20, 66, 74]. Although miRNA and siRNA have somewhat similar sizes, they originate from two different precursor structures and follow different biogenesis pathways with varied modes of action [8, 99]. Long non-coding RNAs act as a source of miRNAs and also function as regulatory factors, whereas Phased small interfering RNAs (phasiRNAs) are produced from lncRNAs under the regulation of miRNA [10]. In the recent past, the study of small RNAs, especially their primary regulations, gained popularity due to their varied roles reported from diverse research fields.
In 1998, Fire et al. reported a breakthrough process of sequence-specific endogenous gene silencing from Caenorhabditis elegans under the act of long double-stranded RNAs (dsRNAs), subsequently popularized as RNA interference (RNAi) or RNA silencing [26, 39]. The process of RNAi involves siRNAs and miRNAs, which is an outcome of double-strand RNA cleavage effected by DICER (ribonuclease) or a Dicer-like protein (DCL) [62]. Later, such miRNAs were also found and reported from Arabidopsis [62, 88, 111]. Till now, various types of ncRNAs have been reported in plants like Arabidopsis [83], sugar beet [157], rice [155], wheat [23], tomato [12, 138], brassica [140], soybean [45], sorghum [127], ginseng [12, 15], pigeon pea [92], and Brachypodium [166]. Under high salinity and drought conditions, Arabidopsis Transacting Short Interfering RNA-Auxin Response Factor (tasiRNA-ARF) played a critical role in supporting the normal floral morphology in Arabidopsis [56]. Regulation of flower formation in pigeon pea is carried out by the interplay between Csa-lncRNA_1231 and Csa-miRNA-156b through the expression of SPL-12 (SQUAMOSA PROMPTER BINDING PROTEIN-LIKE 12) [34]. Moreover, under salt stress, lncRNA973 upregulated its expression in cotton, and lncRNA 973-overexpressing Arabidopsis showed remarkable salt tolerance capacity [161]. In rice, the knockdown of a gene OsDWARF4 through the RNAi promoted the development of shorter plants with enhanced photosynthetic capacity in lower leaves due to the erect leaf architecture of the RNAi plants [38]. Thus, small RNAs play a pivotal role in plant development, stress management, and crop improvement.
With the development of newer molecular biology techniques and bioinformatic tools, the involvement of ncRNAs or sRNAs in plant development and stress regulation has gained prominence. Sometimes, many identical ncRNAs play a pivotal role in plant development and stress tolerance [76]. Recently, it was found that the genetic transformation of ncRNAs can bring about phenotypic change [82, 92]. Finding out the ncRNA or sRNA-mediated developmental pattern and its interaction with stress regulatory mechanisms are essential to understanding plant stress biology and developmental strategies for crop improvement [98].
This review aims to understand the functional roles of sRNAs in the following three aspects: (1) plant development process (tissue patterning to lateral organ morphology); (2) use of RNAi technology in crop improvement to acquire agronomically desirable traits; and (3) understanding the sRNA-mediated stress regulatory network to deploy them judiciously to develop climate-resilient crops.
Non-coding RNA and their classification
Out of 90% of eukaryotic transcribed RNAs, only 2% is translated into protein products [113, 119]. The rest of the untranslated transcribed RNAs represent ncRNAs, which are treated as transcriptional noise with poorly conserved sequences [113]. Later, high-throughput sequencing technology revealed that ncRNAs play crucial roles in transcription, translation, and epigenetic regulation [25, 117, 118, 134].
Based on molecular structure, plant ncRNAs have two types- linear ncRNAs and circular ncRNAs. Linear ncRNAs are further classified into housekeeping ncRNAs and regulatory ncRNAs. Fundamentally, the housekeeping ncRNAs are of four types, namely, tRNAs, snRNAs, rRNAs, and small nucleolar RNAs (sno RNAs). Basically, they possess cellular and ribosomal functions. On the other hand, based on molecular function, regulatory ncRNAs are broadly categorized into two types- sRNAs and long ncRNAs (lncRNAs) [70]. In plants, sRNAs include miRNA, heterochromatic small interfering RNAs (hc-siRNAs), natural antisense transcript-derived small interfering RNAs (nat-siRNAs), repeat-associated siRNAs (rasi-RNAs) and transacting siRNAs (tasiRNAs) [32, 63, 92]. On the basis of genomic location of their protein-coding genes, long non-coding RNAs are further divided into long intergenic ncRNAs, antisense ncRNAs (ancRNAs), intron ncRNAs, and sense ncRNAs (sncRNAs) (Fig. 1). All lncRNAs control transcriptional activity as cis or trans regulators [92, 138].
Small RNA: miRNA and siRNA
MicroRNAs and siRNAs are nearly similar in size but have different precursor structures, modes of action, and biogenesis [8, 62]. Developmentally, miRNA and siRNA are derived from long RNA precursors via the action of dicer-like ribonucleases [13, 50, 51, 57, 62, 71] and are involved in various gene repression mechanisms through ribonucleoprotein silencing complexes [62].
Biogenesis of miRNA
Plant miRNAs (ncRNA molecules) consist of 19–25 nt and are considered under the class of small regulatory RNAs [59]. Intergenic genomic regions are the typical location of miRNA genes; however, sometimes, antisense or sense orientation RNA within a protein-coding gene intron are the sources of miRNAs [36]. Due to their clustered nature, miRNAs are often transcribed as long polycistronic RNAs [36].
In the plant nucleus, RNA polymerase II transcribes MIR genes into pri-miRNA (long primary transcripts) [10, 59]. They are capped at the 5′-end with modification and polyadenylated at the 3′-end [36, 109]. An RNase III-like enzyme, known as DICER-LIKE (DCL 1), cleaves the transcripts (pri-miRNAs) in two steps to form a hairpin-loop structure and a mature miRNA duplex (miRNA/miRNA*) in the nucleus [10, 110, 114]. The 3´-end of this duplex is methylated by the conserved HUA enhancer1 (S-adenosyl-L-methionine dependent RNA methyl transferase) [104]. After duplex formation, it forms a complex with RNA–induced silencing complex (RISC) and AGO1 [5] (see box 1) protein. A part of the RISC complex helps to unwind the miRNA duplex [4]. A small RNA-degrading nuclease eschews one strand of the miRNA duplex. When the miRNA maturation process is over, its production is regulated either by site-specific cleavage of the miRNAs (requires high homology between miRNA and targeted mRNA) or by translational repression of miRNA (that stops the downstream expression of targeted product) (Fig. 2) [36, 127].
Biogenesis of siRNA
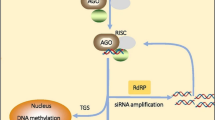
Given differences in function and biogenesis, the siRNAs are categorized into trans-acting siRNAs (tasiRNA), natural antisense siRNAs (nat-siRNAs), and heterochromatic siRNAs (hc-siRNA) [134]. TasiRNAs share a common regulatory mechanism with miRNAs. However, hc-siRNAs act in the cis region of their source. The nat-siRNAs are classified into cis and trans configurations [134, 167]. Long dsRNAs or short hairpin RNAs (snRNA) start the process of PTGS (Post Transcriptional Gene Silencing) or RNAi as they complement the target gene sequence. The RISC containing AGO, DCL enzyme, and dsRNA act as main precursor molecules for the siRNA synthesis [167]. The dicer enzyme truncates the dsRNA to a shorter form with 5´ phosphorylated end and 3′ overhangs(2nt) [62]. The siRNA-induced silencing complex (siRISC) degrades the sense strand of siRNA. After degradation, the siRISC complex binds to an antisense strand of siRNA, along with AGO and other effector proteins, to degrade the target mRNA [62, 79] (Fig. 3).
Formation of tasiRNAs depends on SUPPRESSOR OF GENE SILENCING 3 (SGS3), RNA-dependent PNA polymerase 6 (RDR6) and DCL4 [18, 43, 116]. In Arabidopsis, four types of DCLs are found, of which DCL4 and DCL2 regulate siRNA biogenesis (20–22 nt) [19]. DCL1 is involved in the processing of miRNA (21 nt long) [10], whereas DCL3 assists in the formation of heterochromatic siRNAs (hcsiRNAs, 24 nt long repeats) [149]. DCL1 and DCL2 contribute to nat-siRNA biogenesis along with SGS3 and RDR6 [18]. Once produced, siRNAs with some DNA and histone-modifying proteins, like the cytosine methyltransferase and chromomethylase 3, form a silent (transcriptionally inactive) chromatin state.
Movement of small RNAs
Cell-to-cell communication is the basis of multicellularity and tissue organization. To maintain cytoplasmic connection, plant cells allow intercellular interaction via plasmodesmata, i.e., ‘symplasm’ [95]. In plants, along with metabolites, peptides, proteins, and hormones, some RNAs (messenger and small RNAs) are also translocated from cell to cell via the symplast (by plasmodesmata and nanoscale channels of cell wall) [132]. Both long and short-distance RNA movements are reported in different plant organs. Moreover, some recent studies reported that RNA interchanges between various plants and colonizing organisms (microbes or parasites) take place via exosomes or selected vesicles [95]. More than a decade ago, it was observed that transgene silencing initiated in a single leaf or root and subsequently transmits systematically throughout the whole plant. Hence, it is evident that small RNA can traverse short (plasmodesmata) and long distances (vasculature) [29, 97].
An experiment with wild-type young shoots and rootstock (deficient in dcl2/3/4 siRNA biogenesis) demonstrated that endogenous siRNAs (23–24 nt) can travel from shoot to root and control the PTGS process [73, 100]. The miRNA can also travel through phloem assisted by various favorable physiological cues. Under sulfate, phosphate, and copper deficiencies, some miRNA species (miR395, miR398, and miR399) increase their level remarkably in phloem sap. In Lotus japonicus, miR211 traveled from shoot to root to control the availability of the symbiosis suppressor TOO MUCH LOVE (TML) to regulate healthy roots [44, 133]. Such mobility of small RNA species is really intriguing, particularly their roles in multiple stress responses, which might have a correlation with the magnitude of plant stress levels.
Role of small RNAs in the plant development
Plant development is an ‘umbrella term,’ which denotes the growth and differentiation of a plant during its entire lifespan, i.e., from germination to senescence. Some sRNAs play a central role in the developmental pathway through their participation in different signaling mechanisms (Table 1).
Meristem development
Shoot apical meristem (SAM), a specially organized structure, is the predominant meristem in a plant. The dividing cells of SAM are evolved into plant leaves [92]. In SAM, the development of meristematic cells are regulated by the STM-WUS-CLV pathway [41]. Here, various miRNAs interplay to regulate the development of SAM. In the L1 (protoderm) layer of the SAM, miR394 controls the expression of WUSCHEL (WUS), and later on, it moves through the organizing center (OC) of the SAM to inhibit the expression of LCR or LEAF CURLING RESPONSIVENESS (LCR) [66]—a direct inhibitor of WUS [124]. The process of SAM-related development of shoot apex is carried out by regulation of class III HOMEODOMAIN LEUCINE ZIPPER (HD-ZIP III), PHABULOSA (PHB)/ATHB14, PHAVOLUTA (PHV)/ATHB9, INTERFASCICULAR FIBERLESS/REVOLUTA (IFL1/REV), INCURVATA4/CORONA/ATHB15, and ATHB8 specifically targeted by both miR165 and miR166 species of miRNAs [168] (Fig. 4a).
Role of miRNA in the plant development. a Regulation of meristem identity. The expression of WUSCHEL(WUS) is confined to the organizing center solely through two parallel mechanisms, namely the CLV pathway and the miR394-mediated regulation. b Regulation of leaf shape, polarity and Shoot Apical Meristem. The adaxial and abaxial surfaces are determined by HD-ZIP III. [ab-abaxial, ad-adaxial] Positive regulation → Negative regulation 
Development of leaf
A set of meristematic cells in the peripheral area of the shoot apical meristem (SAM) are responsible for the development of leaves [96]. The leaf polarity (adaxial or upper surface and abaxial or lower surface) is determined by two HD-ZIP III genes, PHABULOSA (PHB) and PHAVOLUTA (PHV) [96] (Fig. 4b). The expression of HD-ZIP III is negatively regulated by miRNA165/166, which is expressed abaxially [64]. A maize mutant (rolled leaf 1, rdl1) (a Zea mays HD ZIP III homolog) and a gain-of-function Arabidopsis mutant (phb-1d) showed abnormality in the development of leaf polarity due to a mutation at the miR165/166 binding site of RDL1 and PHB [60, 103]. Other miRNAs also play an important role in the leaf lamina development. In Arabidopsis, leaf serration occurs due to auxin and CUC2 interaction [136] (Fig. 4b). The miR164 controls the CUC2 and performs a pivotal role in the formation of compound leaves [16]. In tomato and Arabidopsis, the mutation in the miR164 binding site forms a smooth leaf margin [12, 104]. The TCP gene exerts an indirect regulatory influence on the expression of CUC2 by inhibiting miR164 [67]. The formation of leaf shape is governed by GROWTH-REGULATING FACTORs (GRFs)—a cohort of transcription factors closely linked to cell division and cell elongation. Overexpression and loss of function study of GRF 1, 2, and 3 showed the development of large and small leaves [55, 65]. The miR396 regulates the expression of GRF genes. In the distal region of developing leaf blades of Arabidopsis, miR396 regulates the activity of GRF and helps to continue cell proliferation in the proximal region [121]. In the developing leaf of a mature plant, the amount of GRF is diminished, and the miR396 amount is increased [121]. Notably, the regulations of bidirectional, acropetal, and basipetal growth of different leaves are directed through the miR396-GRF pathway [48]. In Arabidopsis leaves, miR393 controls the expression of the TIR1/AFB2 (auxin receptor) involved in auxin-related development [122]. In rice, the formation of the enlarged flag leaf is correlated with overexpression of miR393, which targets related TIR1 homolog during auxin signaling [14, 80].
Flower development and patterning
Apart from various networks of different types of genes, many of the sRNAs are also engaged in floral patterning and development. However, floral transition, i.e., shifting from the vegetative to the reproductive phase, happens under the cooperation of some miRNAs and their target genes [92] (Figs. 5 and 6). In Arabidopsis, the acceleration of flowering is achieved through translational inhibition of the floral homeotic gene APETALA2 (AP2) by the overexpression of miR172 in shoot apical meristem (SAM) [26, 58, 141] (Fig. 5). The miR172 is a crucial factor in flower patterning, accumulates at the center of flower primordia where the class C gene AGAMOUS (AG) is expressed. This accumulation of miR172 leads to AP2 suppression, thereby inhibiting the co-expression of the A and C homeotic genes, delimiting the petal and stamen development [164]. Furthermore, AP2 functions to downregulate miR172, establishing a vital negative feedback loop that is essential for the precise determination of organ identity [27, 153]. The intricacy of this network is further exemplified by a recent investigation, which has revealed that TARGET OF EAT 3 (TOE3) plays a decisive role during the signaling interplay between miR156 and miR172 [61, 142]. Flowers with optimum organ size and preserved flower meristem identity were correlated with overexpressed transcripts of TOE3 (resistant to miR172) [51, 61]. The study offers further substantiation regarding the involvement of SPL3, which facilitates the activation of both TOE3 and its repressor miR172. This interaction among these three components (SPL3, TOE3, and miR172) establishes a negative feedback loop, which effectively governs the precise localization of TOE3 [61]. SPL9, a potent transcriptional activator of miR172, exerts precise and direct regulation over the expression of AP1, FUL, AGL24, and SOC1 through its binding to their individual promoters [135] (Fig. 6). The BLIND (BL) in Petunia and the FISTULATA (FIS) in Antirrhinum encode miR169, which regulate the expression of AG homologs [24, 92]. Thus, both miR172 and miR169 decisively regulate the expression of the ABC gene. Remarkably, the presence of a detectable miR172-binding site can be observed in barley AP2 homolog CLEISTOGAMY1 (CLY1) [94].
Furthermore, it has been demonstrated that miR172 functions as a mobile element capable of diffusing over considerable distances in potato [102]. In potato tuberization, it is possible to augment the process (long-distance travel) through grafting with miR172 expression lines [102]. Additionally, this finding serves as an additional justification for the potential application of small RNAs to achieve targeted biotechnological objectives. The above findings validate the long-distance mobility of miRNA in plants during signal transmission.
After the successful establishment of floral patterns in an organized manner, floral primordia also form a complex structure and lead to the development of a complete flower under the cooperative action of several miRNAs and their target genes. Crosstalk between auxin and some miRNAs, such as ARF,6 ARF8, and miR167, contribute to the formation of male flowers [143]. Additionally, the expression of TIR1 and AUXIN SIGNALLING F-BOX (AFG) genes are regulated by miR393 during flower development [112]. Moreover, anther and silique fruit formation are determined by miR159 via MYB33 [1, 4]. Double-mutant study of miR159 binding site (miR159a and miR159b) in MYB33 showed impaired fertility, development of small siliques, retarded anther formation, and development of smaller seeds [1, 4, 120].
Besides being important regulators of floral organ identity, miR172 (responsible for the adult phase) and miR156 (responsible for the juvenile phase) are also associated with the vegetative to reproductive phase transition. Delay and induction of flowering were found to be controlled by overexpression of miR156 and miR172 [6, 148]. In monocot and dicot plants, AP2-LIKE genes [6] inhibit flowering by convergent action of siRNA [106] with other repressors during long-day conditions [72]. AP2-LIKE transcript is destructed by miR172 (promoted by the SPL family) [154], and the miR156 blocks the SPL activity in the juvenile phase [144], ultimately inhibiting the flowering. Contrarily, in adult plants, miR156 transcripts decrease and initiate the biogenesis of SPL protein to activate miR172 transcripts and hasten flowering. Therefore, the miR172-miR156 complex, along with their respective target genes, perhaps evolved to control flower patterning and phase transition (Figs. 5 and 6). The anatomical (cellular) transition from vegetative to reproductive phase, adaxial to abaxial form, and sepal to petal transformation are crucially linked to the participation of certain sRNAs.
The sRNAs perform diverse functions ranging from regulation of essential determinants of meristem identity, leaf polarity, and flowering process throughout the course of plant development. Mutations during transcription and processing of small RNAs produce pleiotropic effects across plant developmental stages, right from plant dimensions, leaf and flower morphology, as well as the timing of flowering. This underscores the significance of microRNAs in coordinating various phases of plant development.
Role of small RNAs in abiotic stress management
Plants, in their entire life period, constantly struggle to adjust to various environmental stresses that adversely affect their natural growth and development. Unfavorable conditions, especially abiotic stresses (drought, heat, cold, salinity), are well documented that severely impact plant productivity [162, 168]. In response to such abiotic stress conditions, a diverse array of sRNA and ncRNAs differentially express as a part of adaptive strategies (Table 2).
Heat stress
In the last few decades, there has been mounting evidence suggesting the loss of crop productivity due to drastic climate change, particularly due to heat stress [85]. By the end of the twentieth century, the loss of global wheat and maize productivity is expected to reach 5.5% and 3.8%, respectively, due to a 0.13° C temperature rise [169]. Plants are equipped with metabolic machinery to combat heat stress. There are several factors, like heat shock proteins (HSPs), heat shock transcription factor (HSFs), ROS scavenging enzymes, and small RNAs, that work cooperatively to offer thermotolerance [106]. Indeed, several sRNAs are found to be associated with heat stress mitigation and thereby offer thermotolerance. Several miRNA families (mi156/miR172, miR159/miR319, miR160/miR393, and miR398) function at the center of the gene regulatory network that play critical roles in coordinating heat stress tolerance by adaptive strategies towards thermal stress [169] (Fig. 7). The SQUAMOSA PROMOTER-BINDING PROTEIN-LIKE (SPL), the target of miR156, together regulate the phase transition (juvenile to adult and vegetative to reproductive) [91]. The miR156 also controls the high temperature-induced stress conditions in plants [103]. Under heat stress (37°–44°), miR156 isoform overaccumulates and regulates the expression of heat stress-responsive genes in Arabidopsis. The miR156 was reported to be responsible for heat stress memory, and it helps plants retain thermotolerance even after returning to favorable growing temperatures [69]. In this context, it is interesting to note that plants exposed to previous heat stress conditions retain the experience as “heat stress memory,” which they recall during subsequent exposures to heat stress. Heat stress memory can be increased through the inhibition of SPL2 by miR156. This entire phenomenon is controlled by the HSFA2 cascade, which is associated with HEAT STRESS-ASSOCIATED32 (HSA32) and ROF1 [126]. On the other side, miR172 targets the AP2 family genes (AP2, TOE 1, 2, and 3; SCHNARCHZAPFEN or SNZ; and SCHLAFMUTZE or SMZ) and acts as a positive regulator of phase transition in plant [91]. Under optimal growth conditions, the development of flowers takes place in a thermosensory manner by the action of miR172 [70]. Hence, the miR172-AP2 module plays a critical role in heat stress response [115].
The miR159 and miR319 regulate the male fertility and leaf growth [2]. Some members of GIBBERELLIC ACID MYB (GAMYB) are regulated by miR159 and support flower formation and male fertility. Under heat stress, miRNA levels lower while CsGAMYB1 and CsMYB29 (cucumber) levels increase, which underlines their involvement in heat stress response [77]. On the other hand, in Gossypium hirsutum, under extreme heat conditions, overexpressed miR160 blocks ARF10 and ARF17, thereby making the plant susceptible to heat stress, resulting in indehiscent anthers, which reduces crop productivity [35, 40].
The CSD, a significant ROS-scavenging enzyme, influences the expression of SODs (like Fe-SOD, Cu/Zn- SOD, and Mn-SOD) under heat stress. In Arabidopsis, in response to heat stress, the miR398 gets upregulated to suppress its target genes (CSD1, CSD2, and Copper Chaperone for SOD) [46].
A small RNA species, Osa-miR5144, in rice, is also associated with heat stress adaptation. This RNA manipulates the expression of OsPDIL1;1 (protein disulfide isomerase) responsible for protein disulfide bond formation [147]. MicroRNA400 is the primary transcript that undergoes heat stress responsive alternative splicing of the intron, causing lower availability of mature miR400. Therefore, in a heat stress resistance plant, miR400 acts as a negative regulator [151]. In light of crop productivity, it is noteworthy that a host of heat shock proteins, HSFs, miRNAs, hormones, and even alternative splicing collectively offer effective thermal protection to the crop plant (Fig. 7).
Cold stress
Cold stress poses significant challenges to the growth and development of plants, as it affects their physiology and metabolism by altering their cellular homeostasis [92]. Here, the reliable ABA-independent pathway rescues plants during cold stress by activating ICE or Inducer of CBF expression. This pathway further leads to the activation of CBF/DREB1, a downstream transcription factor, by ICE. Eventually, CBF/DREB1 attaches to CRT (C-repeat elements) or LTRE (Low Temperature-Response Element), resulting in the upregulation of COR or cold-responsive genes [47, 68]. Among diverse miRNAs, Osa-miR319b upregulates under cold stress in rice. However, the overexpression of Osa-miR319b simultaneously downregulates Ospcf6 and OsTCP21, upregulates cold-stress responsive genes, including Bdreb1/CBF (Dehydration Responsive Element Binding Protein/C-repeat Binding Factor), DREB2A, and TPP1/2 [7]. Additionally, OsSPL3, which is targeted by OsmiR156, exerts a positive control on OsWRKY71. Contrarily, both OsMYB2 and OsMYB3R-2 are negatively regulated by OsWRKY71. Consequently, this regulation leads the downstream activation of various stress-responsive genes such as OsLEA3, OsDREB24, and OsCTP1 [165]. In the case of wheat, both tasiRNA-ARF and miR167 play a critical role in governing the auxin-signaling pathway and facilitate the development of the cold stress response [130] (Fig. 8).
Drought stress
Drought stress or water deficit condition poses a lethal threat to crop survival. It is interesting to note that changes in small RNA population correlate with severe dehydration, extreme drought, or even water deficit conditions [128]. The following case studies highlight these issues. Arabidopsis displayed upregulation of miR393, miR397b, and miR402 and downregulation of miR319c and miR389a in response to dehydration stress [128]. Rice seedlings subjected to drought stress upregulation of 16 miRNAs and downregulation of 17 miRNAs [158]. Maize leaves showed upregulation of 8 miRNAs and downregulation of 13 miRNAs, whereas root displayed upregulation of 7 miRNAs and downregulation of another seven miRNAs under drought stress conditions [87]. Notably, miR171, 156, 160, 162, and 164 species were found to downregulate the expression of their respective target genes during drought stress, which indicates their valuable roles. The intriguing study by Shen et al. [123] highlights that Mdm-miR160 moves from the scion to the rootstock, contributing to drought adaptation and root formation in apple. The MIR160 plays a vital role in promoting the activation of MdARF17 through the formation of a positive feedback loop, and it specifically binds to the promoter region of MdHLY1 [123]. In Arabidopsis, the miR165/166-mediated regulation of HD ZIP IIIs facilitates ABA signaling, which is crucial for drought adaptation. Moreover, HD-ZIP IIIs activate ARF, the target of miR16. This cooperative interaction between miR160 and miR165/166 enables plants to respond to and tolerate drought stress [86]. On the other hand, the ABA-dependent response to drought stress involves the miR159-MYB and miR169-NFYA components [86]. In Arabidopsis, miR159 modulates the cleavage and regulation of MYB33 and MYB101 during the germination phase, which assists in responding to drought stress and the accumulation of ABA [121]. Interestingly, the ABA-dependent pathway suppresses the expression of miR169 in the face of impending drought stress [92]. Indeed, the sensitivity to drought stress in Arabidopsis came to light through the study of miR169a expression and knockout experiments involving nfya5. However, the overexpression of NFYA5 promotes better adaptability to drought stress [78]. Similar patterns can be observed in Brassica napus and soybean, where the targets of miR169, namely NF-γAδ, AtNFYA1, and AtNFYA5, inhibit the production of stress-induced genes and exhibit a negative response to drought conditions [75, 154] (Fig. 9).
Salinity
Salinity or salt stress, akin to other abiotic stresses, impedes the natural growth and reproductive processes of plants. In this discourse, we shall delve into the significance of plant miRNAs and their respective target genes in alleviating the effects of salinity stress. In peanuts, similar to drought stress, the miR160 and its target gene ARF exhibit heightened expression in response to salt stress. In saline conditions, the ARF18 pathway, triggered by the overexpression of miR160, facilitates seedling growth and neutralizes the impact of ROS [131]. Furthermore, certain plants demonstrate distinct reactions or promotion effects toward salt tolerance by different or identical miRNAs that belong to the same family. For instance, in rice and Arabidopsis, the overexpression of Osa-miR393 results in diminished salt tolerance [42, 47, 146], whereas overexpression of the resistant form, MTIR1, of miR393 leads to enhanced salt tolerance [28]. In cotton, both miR414c and its target gene GhFSD1 (Iron Superoxide dismutase 1) contribute to salt tolerance. The overexpression study of miR414c illustrates a reduction in GhFSD1 expression, making it susceptible to salinity stress by regulating ROS metabolism [139] (Fig. 10).
Impact of small RNAs for the establishment of climate-resilient crops
Recent reports indicate that agro-climatic systems are becoming increasingly vulnerable to warming climatic conditions and changing precipitation patterns, with an expected impact on food security [53]. Various environmental stresses, like drought, soil contamination, rising temperatures, UV radiation, and atmospheric CO2 and O3 levels, pose a challenge to developing climate-resilient crops to meet the growing demand for food and feed. In the post-genomic era, biotechnological advancements provided a foundation for sustainable agriculture and increased crop flexibility in the face of climate change. To tackle these environmental challenges, breeding methods are combined with modern biotechnological tools to design stress-tolerant crops while maintaining high yields. The miRNAs, which play a crucial role in plant stress tolerance and environmental adaptation, promise immense potential for crop improvement [21]. The utilization of modern computational tools, databases, and high-throughput sequencing has contributed significantly to identifying numerous mRNAs that respond to single or multiple stress exposures [25]. Despite recent advancements in understanding the effects of plant miRNAs on growth, development, and stress responses, it is still limited for developing resilient crops that can thrive in challenging climatic conditions.
Exciting advancements have been made in the utilization of sRNAs to improve agronomic traits and bolster resistance to challenging environments. These small molecules play a crucial role in orchestrating the regulation of target genes to combat stress conditions [160]. Thanks to recent breakthroughs in high-throughput sequencing technology and bioinformatics, characterizing small RNAs and their corresponding targets has become more accessible than ever before. The techniques of overexpression and knockdown are gaining popularity as powerful tools to introduce functional improvements and to investigate the loss of specific functions in transgenic plants [37]. Till now, a group of transgenic plants with altered miRNAs has been designed to understand their response to abiotic stresses.
In Arabidopsis thaliana, the overexpression vector containing pre-miR408 was utilized to generate transformants exhibiting superior qualities, as compared to control plants, such as the absence of chlorosis, wilting, and drying. The study on overexpression of miR408 demonstrated reduced expression of the target gene Plantacyanin, whereas genes Rd17 and Rd29, along with DREB1/2A (a transcription factor of Rd17 and Rd29), revealed enhancement in transgenic plants, which mediates a positive response to drought stress [49]. The overexpression of miR408 in Cicer arientinum improved drought tolerance. The transgenic plant Agrostis stolonifera, harbouring the pre-miR393a of O. sativa, displayed stronger resilience to salinity (at 250 mM for 10 days) and heat (at 40 °C Day/35 °C night, 13 days), suggesting miR393 would be an excellent candidate to confer multi-stress tolerance [163]. The utilization of miR172 facilitated the development of transgenic plants with enhanced adaptability under high salt and water deficit conditions [67]. The overexpression of Glycine max-miR172 in transgenic Arabidopsis led to an increase in germination rate, greater number of green cotyledons, and reduced leaf water loss. Glyma o1g39520 (AP2/ERF), identified as the predicted target of this miRNA in Arabidopsis, acted as a regulatory component responsible for improved tolerance capacity [77]. Overexpression of miRNA genes helps to achieve cold tolerance in O. sativa [113]. Seedlings that were genetically modified to overexpress Osa-miR319b were subjected to 4 °C for 7 days, followed by a recovery period of 10 days. It was observed that the transgenic plants showed a better recovery rate (up to 90%) compared to the mere 30% recovery in wild-type plants. In support of these findings, gene expression profiles of two predicted targets, OsPCF6 and OsTCP21, were carried out, which indicated that these genes were inhibited by low temperatures, with wild-type plants displaying notable induction [137]. Another experiment conducted by Zhang et al. [158] involved the overexpression of miR396b from Poncirus trifoliata in Citrus lemon to enhance cold tolerance. Exposure of four-monthold transgenic and wild-type plants to cold (− 2 °C) for 12 h resulted in a higher accumulation of polyamines, reduced level of ROS, decreased ACO transcripts, and lowered ethylene content. These findings can be hypothesized as potential factors contributing to the successful recovery and growth of these plants under cold stress.
Currently, strategies involving the overexpression of small RNAs are being explored globally to develop climate-resilient plants. However, there is a growing trend for the knock-out approach, specifically through the short tandem target mimic (STTM) method, to achieve the same purpose. This approach results in the loss-of-function of identified miRNA, which acts as a negative regulator. Zhang et al. [159] demonstrated that miR166 knock-out lines (STTM166) showed reduced stomatal conductance and transpiration rate, thereby ensuring better photosynthetic efficiency. Additionally, the miRNA's major target, OsHB4, was identified as a potential factor in improving drought tolerance. The study concluded that manipulating miRNA can lead to developmental changes, such as leaf rolling and reduction in xylem diameter, which mimic the response of plants under water deficit conditions. Various studies are being conducted to deploy miRNAs in engineering crops with the aim to withstand multiple stress conditions. The emerging global research trends in miRNA underline its emerging significance in developing climate-resilient crops through targeted overexpression of concerned genes and their miRNAs.
Conclusion
Significant progress has been made in the study of plant sRNAs in recent years, establishing them as essential regulators of almost every aspect of the plant’s life, including growth, development, and sensitivity to environmental stimuli. Recombinant-DNA-based crop engineering generally targets specific genes and proteins of agro-economic traits to improve crop productivity with stress tolerance. This research trend is supported by the availability of databases and molecular tools dedicated to miRNA-based research. Scientists have discovered a large number of miRNAs that are crucial for plants’ responses to a variety of environmental challenges through the use of sRNAs and ‘degradome sequencing’ techniques and in silico biology. The ability of these miRNAs to target particular genes has been effectively predicted and increasingly explored.
Recent research suggests that plant miRNAs can be valuable targets for genetic engineering. The aim is to enhance stress tolerance in crop plants by manipulating miRNA expression through methods like overexpression or knock-out techniques, depending on their regulatory role. Despite limitations and the need for more research, early attempts in miRNA-based genetic engineering demonstrate their immense potential for developing climate-resilient crops. Furthermore, plant miRNAs are also being acknowledged as useful tools for genome editing, although further investigation is needed in this area.
Understanding the complexity of plant sRNAs and their biogenesis and action is still a significant challenge. It's important to acknowledge that the plant genome has evolved to contain a sophisticated and precise regulatory system through natural selection. A biased explanation of gene expression regulation based solely on structure, function, and mechanism is insufficient. Various components such as functional proteins, transcription factors, ncRNAs, small peptides, and epigenetic modification interact to create intricate networks that finely regulate gene expression in plant development and stress responses. Therefore, future research on sRNAs in plants should prioritize functional analysis and the development of transgenic crops that would be robust and proof of climatic fluctuations and exhibit consistent expression of desired agro-economic traits.
References
Achard P, et al. Modulation of floral development by a gibberellin-regulated microRNA. Development. 2004;131:3357–65.
Ahmed W, Xia Y, Zhang H, Li R, Bai G, Siddique KHM, et al. Identification of conserved and novel miRNAs responsive to heat stress in flowering Chinese cabbage using high-throughput sequencing. Sci Rep. 2019;9:14922. https://doi.org/10.1038/s41598-019-51443-y.
Akdogan G, Tufekci ED, Uranbey S, Unver T. miRNA-based drought regulation in wheat. Funct Integr Genomics. 2016;16(3):221–33.
Allen RS, et al. Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc Natl Acad Sci U S A. 2007;104:16371–6.
Arribas-Hernández L, Marchais A, Poulsen C, Haase B, Hauptmann J, Benes V, et al. The slicer activity of ARGONAUTE1 is required specifically for the phasing, not production, of trans-acting short interfering RNAs in Arabidopsis. Plant Cell. 2016;28:1563–80. https://doi.org/10.1105/tpc.16.00121.
Aukerman MJ, Sakai H. Regulation of flowering time and floral organ identity by a microRNA and its APETALA2- like target genes. Plant Cell. 2003;15:2730–41.
Aung K, Lin SI, Wu CC, Huang YT, Su CL, Chiou TJ. pho2, a phosphate over accumulator, is caused by a nonsense mutation in a microRNA399 target gene. Plant Physiol. 2006;141:1000–11.
Axtell MJ. Classification and comparison of small RNAs from plants. Annu Rev PlantBiol. 2013;64:137–59. https://doi.org/10.1146/annurev-arplant-050312120043.
Barakat A, Sriram A, Park J, Zhebentyayeva T, Main D, Abbott A. Genome wide identification of chilling responsive microRNAs in Prunus persica. BMC Genom. 2012;13:1–11.
Bartel DP. MicroRNAs: genomics, biogenesis mechanism, and function. Cell. 2004;116:281–97. https://doi.org/10.1016/S0092-8674(04)00045-5.
Barrera-Figueroa BE, Gao L, Wu Z, Zhou X, Zhu J, Jin H, Liu R, Zhu J-K. High throughput sequencing reveals novel and abiotic stress-regulated microRNAs in the inflorescences of rice. BMC Plant Biol. 2012;12:1–11.
Berger Y, et al. The NAC-domain transcription factor GOBLET specifies leaflet boundaries in compound tomato leaves. Development. 2009;136(823–832):49.
Bernstein E, Caudy AA, Hammond SM. Hannon GJ Rolefor a bidentate ribonuclease in the initiation step of RNA interference. Nature. 2001;409:363–6. https://doi.org/10.1038/35053110.
Bian H, Xie Y, Guo F, Han N, Ma S, Zeng Z, Wang J, Yang Y, Zhu M. Distinctive expression patterns and roles of the miRNA393/TIR1 homolog module in regulating flag leaf inclination and primary and crown root growth in rice (Oryza sativa). New Phytol. 2012;196:149–61.
Bian XB, Yu PC, Dong L, Zhao Y, Yang H, Han YZ, Zhang LX. Regulatory role of non-coding RNA in ginseng rusty root symptom tissue. Sci Rep. 2021;11:9211.
Blein T, et al. A conserved molecular framework for compound leaf development. Science. 2008;322:1835–9.
Bhutia KL, Khanna VK, Meetei TNG, Bhutia ND. Effects of climate change on growth and development of chilli. Agrotechnology. 2018;7(2):1–4.
Borsani O, Zhu J, Verslues PE, Sunkar R, Zhu JK. Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell. 2005;123:1279–91.
Bouche N, Lauressergues D, Gasciolli V, Vaucheret H. Anantagonistic function for Arabidopsis DCL2 in development and a newfunc- tion for DCL4 in generating viral siRNAs. EMBO J. 2006;25:3347–56. https://doi.org/10.1038/sj.emboj.7601217.
Brant E, Budak H. Plant small non-coding RNAs and their roles in biotic stresses. Front Plant Sci. 2018;9:1038. https://doi.org/10.3389/fpls.2018.01038.
Budak H, Zhang B. MicroRNAs in model and complex organisms. Funct Integr Genomics. 2017;17:121–4. https://doi.org/10.1007/s10142-017-0544-1.
Busch BL, et al. Shoot branching and leaf dissection in tomato are regulated by homologous gene modules. Plant Cell. 2011;23:3595–609.
Cagirici HB, Alptekin B, Budak H. RNA sequencing and co-expressed long non-coding RNA in modern and wild wheats. Sci Rep. 2017;7:10670.
Cartolano M, et al. A conserved microRNA module exerts homeotic control over Petunia hybrida and Antirrhinum majus floral organ identity. Nat Genet. 2007;39:901–5.
Cech TR, Steitz JA. The noncoding RNA revolution-trashing old rules to forge new ones. Cell. 2014;157:77–94.
Chen X. Small RNAs and their roles in plant development. BioloAnnu Rev Cell Dev Biol. 2009;25:21–44.
Chen X. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science. 2004;303:2022–5.
Chen Z, Hu L, Han N, Hu J, Yang Y, Xiang T, Zhang X, Wang L. Overexpression of a miR393-resistant form of transport inhibitor response protein 1 (mTIR1) enhances salt tolerance by increased osmoregulation and Na+ exclusion in Arabidopsis thaliana. Plant Cell Physiol. 2015;56:73–83.
Chitwood DH, Timmermans MCP. Small RNAs are on the move. Nature. 2010;467:415–9.
Chuck G, et al. The maize tasselseed4 microRNA controls sex determination and meristem cell fate by targeting Tasselseed6/indeterminate spikelet1. Nat Genet. 2007;39:1517–21.
Chuck G, et al. The maize SBP-box transcription factor encoded by tasselsheath4 regulates bract development and the establishment of meristem boundaries. Development. 2010;137:1243–50.
Contreras-Cubas C, Palomar M, Arteaga-Vazquez M, Reyes JL, Covarrubias AA. Non-coding RNAs in the plant response to abiotic stress. Planta. 2012;236:943–58.
D’Ario M, Griffiths-Jones S, Kim M. Small RNAs: big impact on plant development. Trends Plant Sci. 2017;22(12):1056.
Das A, Saxena S, Kumar K, Tribhuvan KU, Singh NK, Gaikwad K. Non-coding RNAs having strong positive interaction with mRNAs reveal their regulatory nature during flowering in a wild relative of pigeonpea (Cajanus scarabaeoides). Mol Biol Rep. 2020;47:3305–17.
Ding Y, Ma Y, Liu N, Xu J, Hu Q, Li Y, et al. microRNAs involved in auxin signalling modulate male sterility under high-temperature stress in cotton (Gossypium hirsutum). Plant J. 2017;91:977–94. https://doi.org/10.1111/tpj.13620.
Djami-Tchatchou AT, Sanan-Mishra N, Ntushelo K, Dubery IA. Functional roles of microRNAs in agronomically important plants—potential as targets for crop improvement and protection. Front Plant Sci. 2017;8:378. https://doi.org/10.3389/fpls.2017.00378.
Esmaeli F, Shiran B, Fallahi H, Mirakhorli N, Budak H, Martínez-Gómez P. In silico search and biological validation of microRNAs related to drought response in peach and almond. Funct Integr Genomics. 2017;17:189–201.
Feldmann KA. Steroid regulation improves crop yield. Nat Biotechnol. 2006;4:46–7. https://doi.org/10.1038/nbt0106-46.
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391:806–11.
Gahlaut V, Baranwal VK, Khurana P. miRNomes involved in imparting thermotolerance to crop plants. 3 Biotech. 2018;8:497. https://doi.org/10.1007/s13205-018-1521-7.
Gaillochet C, Lohmann JU. The never-ending story: from pluripotency to plant developmental plasticity. Development. 2015;142:2237–49.
Gao P, Bai X, Yang L, Lv D, Pan X, Li Y, Cai H, Ji W, Chen Q, Zhu Y. osa-MIR393: a salinity- and alkaline stress-related microRNA gene. Mol Biol Rep. 2011;38:237–42.
Gasciolli V, Mallory AC, Bartel DP, Vaucheret H. Partially redundant functions of Arabidopsis DICER-like enzymes and a role for DCL4 in producing trans-acting siRNAs. Curr Biol. 2005;15:1494–500.
Gautrat P, Laffont C, Frugier F. Compact root architecture 2 promotes root competence for nodulation through the miR2111 systemic effector. Curr Biol. 2020;30:1339–45.
Golicz AA, Singh MB, Bhalla PL. The long intergenic noncoding RNA (LincRNA) landscape of the soybean genome. Plant Physiol. 2018;176:2133–47.
Guan Q, Lu X, Zeng H, Zhang Y, Zhu J. Heat stress induction of miR398 triggers a regulatory loop that is critical for thermotolerance in Arabidopsis. Plant J. 2013;74:840–51. https://doi.org/10.1111/tpj.12169.
Guo J, Ren Y, Tang Z, Shi W, Zhou M. Characterization and expression profiling of the ICE-CBF-COR genes in wheat. PeerJ. 2019;7:81–90.
Gupta M, Das NU. Divergence in patterns of leaf growth polarity is associated with the expression divergence of miR396. Plant Cell. 2015;27:2785–99.
Hajyzadeh M, Turktas M, Khawar KM, Unver T. MiR408 overexpression causes increased drought tolerance in chickpea. Gene. 2015;555:186–93. https://doi.org/10.1016/j.gene.2014.11.002.
Hamilton AJ, Baulcombe DC. A novel species of small anti- sense RNA in post transcriptional gene silencing. Science. 1999;286:950–2. https://doi.org/10.1126/science.286.5441.950.
Hammond SM, Bernstein E, Beach D, Hannon GJ. AnRNA- directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature. 2000;404:293–6. https://doi.org/10.1038/35005107.
Hamza NB, Sharma N, Tripathi A, Sanan-Mishra N. MicroRNA expression profiles in response to drought stress in Sorghum bicolor. Gene Expr Patterns. 2016;20(2):88–98.
Hatfield JL, Antle J, Garrett KA, Izaurralde RC, Mader T, Marshall E, Nearing M, Philip Robertson G, Ziska L. Indicators of climate change in agricultural systems. Clim Chang. 2018. https://doi.org/10.1007/s10584-018-2222-2.
Hibara K, et al. Arabidopsis CUP-SHAPED COTYLE-DON3 regulates postembryonic shoot meristem and organ boundary formation. Plant Cell. 2006;18:2946–57.
Horiguchi G, et al. The transcription factor AtGRF5 and the transcription coactivator AN3 regulate cell proliferation in leaf primordia of Arabidopsis thaliana. Plant J. 2005;43(68–78):57.
Huang L, Dong H, Zhou D, Li M, Liu Y, Zhang F, Feng Y, Yu D, Li S, Cao J. Syatematic identification of long non-coding RNAs during pollen development and fertilization in Brassica rapa. Plant J. 2018;96:203–22.
Hutvagner G, McLachlan J, Pasquinelli AE, Balint E, Tuschl T, Zamore PD. A cellular function for the RNA-interference enzyme Dicerin the maturation of the let-7 small temporal RNA. Science. 2001;293:834–8. https://doi.org/10.1126/science.1062961.
Irish VF, Sussex IM. Function of the APETALA-1 gene during Arabidopsis floral development. Plant Cell. 1990;2:741–53.
Jones-Rhoades MW, Bartel DP, Bartel B. MicroRNAs and their regulatory roles in plants. Ann Rev Plant Biol. 2006;57:19–53. https://doi.org/10.1146/annurev.arplant.57.032905.105218.
Juarez MT, et al. MicroRNA-mediated repression of rolled leaf1 specifies maize leaf polarity. Nature. 2004;428:84–8.
Jung JH, et al. The miR172 target TOE3 represses AGAMOUS expression during Arabidopsis floral patterning. Plant Sci. 2014;215:29–38.
Kamthan A, Chaudhuri A, Kamthan M, Datta A. Small RNAs in plants: recent development and application for crop improvement. Front Plant Sci. 2015;6:208. https://doi.org/10.3389/fpls.2015.00208.
Khraiwesh B, Zhu JK, Zhu J. Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. Biochim Biophys Acta. 2012;1819:137–48.
Kidner CA, Martienssen RA. Spatially restricted microRNA directs leaf polarity through ARGONAUTE1. Nature. 2004;428:81–4.
Kim JH, et al. The AtGRF family of putative transcription factors is involved in leaf and cotyledon growth in Arabidopsis. Plant J. 2003;36:94–104.
Knauer S, et al. A protodermal miR394 signal defines a region of stem cell competence in the Arabidopsis shoot meri-stem. Dev Cell. 2013;24:125–32.
Koyama T, et al. TCP transcription factors regulate the activities of ASYMMETRIC LEAVES1 and miR164, as well as the auxin response, during differentiation of leaves in Arabidopsis. Plant Cell. 2010;22:3574–88.
Kumar R. Role of microRNAs in biotic and abiotic stress responses in crop plants. Appl Biochem Biotechnol. 2014;174:93–115.
Lämke J, Brzezinka K, Altmann S, Bäurle I. A hit-and-run heat shock factor governs sustained histone methylation and transcriptional stress memory. EMBO J. 2016;35:162–75. https://doi.org/10.15252/embj.201592593.
Lee H, Yoo SJ, Lee JH, Kim W, Yoo SK, Fitzgerald H, et al. Genetic framework for flowering-time regulation by ambient temperature-responsive miRNAs in Arabidopsis. Nucleic Acids Res. 2010;38:3081–93. https://doi.org/10.1093/nar/gkp1240.
Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic genelin-encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843–54. https://doi.org/10.1016/0092-8674(93)90529-Y.
Lee YS, et al. Rice miR172 induces flowering by suppressing OsIDS1 and SNB, two AP2 genes that negatively regulate expression of Ehd1 and florigens. Rice. 2014;7:31.
Lewsey MG, Hardcastle TJ, Melnyk CW, Molnar A, Valli A, Urich MA, Nery JR, Baulcombe DC, Ecker JR. Mobile small RNAs regulate genome-wide DNA methylation. Proc Natl Acad Sci U S A. 2016;113:E801–10.
Li DD, Qiao HL, Qiu WJ, Xu X, Liu TM, Jiang QL, et al. Identification and functional characterization of intermediate-size non-coding RNAs in maize. BMC Genom. 2018;19:1–11.
Li J, Duan YJ, Sun NL, Wang L, Feng SS, Fang YJ, Wang YP. The miR169n-NF-YA8 regulation module involved in drought resistance in Brassica napus L. Plant Sci. 2021;313:111062.
Li S, Castillo-Gonzalez C, Yu B, Zhang X. The functions of plant small RNAs in development and in stress responses. Plant J. 2017;90:654–70.
Li W, Wang T, Zhang Y, Li Y. Overexpression of soybean miR172c confers tolerance to water deficit and salt stress, but increases ABA sensitivity in transgenic Arabidopsis thaliana. J Exp Bot. 2016;67:175–94. https://doi.org/10.1093/jxb/erv450.
Li WX, Oono Y, Zhu JH, He XJ, Wu JM, Iida K, Lu XY, Cui XP, Jin HL, Zhu JK. The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and post transcriptionally to promote drought resistance. Plant Cell. 2008;20:2238–51.
Li W, Cui X, Meng Z, Huang X, Xie Q, Wu H, et al. Transcriptional regulation of Arabidopsis MIR168a and argonaute1 homeostasis in abscisic acid and abiotic stress responses. Plant Physiol. 2012;158:1279–92. https://doi.org/10.1104/pp.111.188789.
Li X, et al. Control of tillering in rice. Nature. 2003;422:618–21.
Lin JS, Kuo CC, et al. MicroRNA160 modulates plant development and heat shock protein gene expression to mediate heat tolerance in Arabidopsis. Front Plant Sci. 2018;9:68.
Liu DG, Mewalal R, Hu RB, Tuskan GA, Yang XH. New technologies accelerate the exploration of non-coding RNAs in horticultural plants. Hortic Res. 2017;4:17031.
Liu J, Jung C, Xu J, Wang H, Deng S, Bernad L, et al. Genome-wide analysis uncovers regulation of long intergenic noncoding RNAs in Arabidopsis. Plant Cell. 2012;24:4333–45.
Liu H, et al. OsmiR396d-regulated OsGRFs function in floral organogenesis in rice through binding to their targets OsJMJ706 and OsCR4. Plant Physiol. 2014;165:160–74.
Liu Q, Yan S, Yang T, Zhang S, Chen Y, Liu B. Small RNAs in regulating temperature stress response in plants. J Integr Plant Biol. 2017;59:774–91. https://doi.org/10.1111/jipb.12571.
Liu Q, Yang T, Yu T, Zhang S, Mao X, Zhao J, Wang X, Dong J, Liu B. Integrating small RNA sequencing with qtl mapping for identification of miRNAs and their target genes associated with heat tolerance at the flowering stage in rice. Front Plant Sci. 2017;8:43.
Liu X, Zhang X, Sun B, Hao L, Liu C, Zhang D, Tang H, Li C, Li Y, Shi Y, et al. Genome-wide identification and comparative analysis of drought-related microRNAs in two maize inbred lines with contrasting drought tolerance by deep sequencing. PLoS ONE. 2019;14:e0219176.
Llave C, Xie Z, Kasschau KD, Carrington JC. Cleavage of Scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science. 2002;297:2053–6. https://doi.org/10.1126/science.1076311.
Luan M, Xu M, Lu Y, Zhang Q, Zhang L, Zhang C, Fan Y, Lang Z, Wang L. Family-wide survey of miR169s and NF-Yas and their expression profiles response to abiotic stress in maize roots. PLoS ONE. 2014;9:91369.
Ma C, Burd S, Lers A. miR408 is involved in abiotic stress responses in Arabidopsis. Plant J. 2015;84:169–87.
Ma J, Zhao P, Liu S, Yang Q, Guo H. The control of developmental phase transitions by microRNAs and their targets in seed plants. Int J Mol Sci. 2020;21:1971. https://doi.org/10.3390/ijms21061971.
Ma X, Zhao F, Zhou B. The characters of non-coding RNAs and their biological roles in plant development and abiotic stress response. Int J Mol Sci. 2022;23:4124. https://doi.org/10.3390/ijms23084124.
Marin E, et al. miR390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregu-latory network quantitatively regulating lateral root growth. Plant Cell. 2010;22:1104–17.
Martin A, et al. Graft-transmissible induction of potato tuberization by the microRNA miR172. Development. 2009;136:2873–81.
Maizel A, Markmann K, Timmermans M, Wachter A. To move or not to move: roles and specificity of plant RNA mobility. Curr Opin Plant Biol. 2020;57:52–60.
McConnell JR, Barton MK. Leaf polarity and meri- stem formation in Arabidopsis. Development. 1998;125:2935–42.
Melnyk CW, Molnar A, Baulcombe DC. Intercellular and systemic movement of RNA silencing signals. EMBO J. 2011;30:3553–63.
Meng XX, Li AX, Yu B, Li SJ. Interplay between miRNAs and lncRNAs: mode of action and biological roles in plant development and stress adaptation. Comput Struct Biotechnol. 2021;19:2567–74.
Mette MF, Vander Winden J, Matzke M, Matzke AJ. Short RNAs can identify new candidate transposable element families in Arabidopsis. Plant Physiol. 2002;130:6–9. https://doi.org/10.1104/pp.007047.
Molnar A, Melnyk CW, Bassett A, Hardcastle TJ, Dunn R, Baulcombe DC. Small silencing RNAs in plants are mobile and direct epigenetic modification in recipient cells. Science. 2010;328:872–5.
Montgomery TA, et al. Specificity of ARGONAUTE7– miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell. 2008;133:128–41.
Nair SK, et al. Cleistogamous flowering in barley arises from the suppression of microRNA-guided HvAP2 mRNA cleav-age. Proc Natl Acad Sci U S A. 2010;107:490–5.
Nelson JM, et al. Expression of a mutant maize gene in the ventral leaf epidermis is sufficient to signal a switch of the leaf’s dorsoventral axis. Development. 2002;129:4581–9.
Nikovics K, et al. The balance between the MIR164A and CUC2 genes controls leaf margin serration in Arabidopsis. Plant Cell. 2006;18:2929–45.
Ochando I, et al. Alteration of the shoot radial pattern in Arabidopsis thaliana by a gain-of-function allele of the class III HD-Zip gene INCURVATA4. Int J Dev Biol. 2008;52:953–61.
Ohama N, Sato H, Shinozaki K, Yamaguchi-Shinozaki K. Transcriptional regulatory network of plant heat stress response. Trends Plant Sci. 2017;22:53–65. https://doi.org/10.1016/j.tplants.2016.08.015.
Ori N, et al. Regulation of LANCEOLATE by miR319 is required for compound-leaf development in tomato. Nat Genet. 2007;39:787–91.
Pan WJ, Tao JJ, Cheng T, Bian XH, Wei W, Zhang WK, Ma B, Chen SY, Zhang JS. Soybean miR172a improves salt tolerance and can function as a long-distance signal. Mol Plant. 2016;9:1337–40.
Pantaleo V, Szittya G, Moxon S, Miozzi L, Moulton V, Dalmay T, et al. Identification of grapevine microRNAs and their targets using high-throughput sequencing and degradome analysis. Plant J. 2010;62:960–76. https://doi.org/10.1111/j.0960-7412.2010.04208.x.
Papp I, Mette MF, Aufsatz W, Daxinger L, Schauer SE, Ray A, et al. Evidence for nuclear processing of plant micro RNA and short interfering RNA precursors. Plant Physiol. 2003;132:1382–90. https://doi.org/10.1104/pp.103.021980.
Park W, Li J, Song R, Messing J, Chen X. Carpelfactory, a dicer homolog, and HEN1, a novel protein, actin microRNA metabolism in Arabidopsis thaliana. Curr Biol. 2002;12:1484–95. https://doi.org/10.1016/S0960-9822(02)01017-5.
Parry G, et al. Complex regulation of the TIR1/AFB family of auxin receptors. Proc Natl Acad Sci U S A. 2009;106:22540–5.
Pauli A, Rinn JL, Schier AF. Non-coding RNAs as regulators of embryogenesis. Nat Rev Genet. 2011;12:136–49.
Peláez P, Trejo MS, Iñiguez LP, Estrada-Navarrete G, Covarrubias AA, Reyes JL, et al. Identification and characterization of microRNAs in Phaseolus vulgaris by high-throughput sequencing. BMC Genom. 2012;13:83. https://doi.org/10.1186/1471-2164-13-83.
Peng Y, Zhang X, Liu Y, Chen X. Exploring heat-response mechanisms of microRNAs based on microarray data of rice post-meiosis panicle. Int J Genomics. 2020;17:7582612. https://doi.org/10.1155/2020/7582612.
Peragine A, Yoshikawa M, Wu G, Albrecht HL, Poethig RS. SGS3 and SGS2/SDE1/RDR6 are required for juvenile development and the production of trans-acting siRNAs in Arabidopsis. Genes Dev. 2004;18:2368–79.
Peschansky VJ, Wahlestedt CWC. Non-coding RNAs as direct and indirect modulators of epigenetic regulation. Epigenetics. 2014;9:3–12.
Ponjavic J, Ponting CP, Lunter G. Functionality or transcriptional noise? Evidence for selection within long noncoding RNAs. Genome Res. 2007;17:556–65.
Rai MI, Alam M, Lightfoot DA, Gurha P, Afzal AJ. Classification and experimental identification of plant long non-coding RNAs. Genomics. 2019;111:997–1005.
Reyes JL, Chua NH. ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. Plant J. 2007;49:592–606.
Rodriguez RE, et al. Control of cell proliferation in Arabidopsis thaliana by microRNA miR396. Development. 2010;137:103–12.
Si-Ammour A, Windels D, Arn-Bouldoires E, Kutter C, Ailhas J, Meins F Jr, Vazquez F. miR393 and secondary siRNAs regulate expression of the TIR1/AFB2 auxin receptor clade and auxin-related development of Arabidopsis leaves. Plant Physiol. 2011;157:683–91.
Shen X, He J, Ping Y, Guo J, Hou N, Cao F, Li X, Geng D, Wang S, Chen P, et al. The positive feedback regulatory loop of miR160-Auxin Response Factor 17- HYPONASTIC LEAVES 1 mediates drought tolerance in apple trees. Plant Physiol. 2021;188:1686–708.
Song JB, et al. Regulation of leaf morphology by micro- RNA394 and its target leaf curling responsiveness. Plant Cell Physiol. 2012;53:1283–94.
Song JB, Gao S, Wang Y, Li BW, Zhang YL, Yang ZM. miR394 and its target gene LCR are involved in cold stress response in Arabidopsis. Plant Gene. 2016;5:56–64.
Stief A, Altmann S, Hoffmann K, Pant BD, Scheible WR, Bäurle I. Arabidopsis miR156 regulates tolerance to recurring environmental stress through SPL transcription factors. Plant Cell. 2014;26:1792–807. https://doi.org/10.1105/tpc.114.123851.
Sun X, Zheng HX, Li JL, Liu LN, Zhang XS, Sui N. Comparative transcriptome analysis reveals new lncRNAs responding to salt stress in sweet sorghum. Front Bioeng Biotechnol. 2020;8:331.
Sunkar R, Zhu JK. Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell. 2004;16:2001–19.
Swiezewski S, et al. Small RNA-mediated chromatin silencing directed to the 30 region of the Arabidopsis gene encoding the developmental regulator. FLC Proc Natl Acad Sci U S A. 2007;104:3633–8.
Tang Y, Du G, Xiang J, Hu C, Li X, Wang W, Zhu H, Qiao L, Zhao C, Wang J, et al. Genome-wide identification of auxin response factor (ARF) gene family and the miR160-ARF18-mediated response to salt stress in peanut (Arachis hypogaea L.). Genomics. 2022;114:171–84.
Tang Z, Zhang L, Xu C, Yuan S, Zhang F, Zheng Y, Zhao C. Uncovering small RNA-mediated responses to cold stress in a wheat thermosensitive genic male-sterile line by deep sequencing. Plant Physiol. 2012;159:721–38.
Tilsner J, Nicolas W, Rosado A, Bayer EM. Staying tight: plasmodesmal membrane contact sites and the control of cell-to-cell connectivity in plants. Annu Rev Plant Biol. 2016;67:337–64.
Tsikou D, Yan Z, Holt DB, Abel NB, Reid DE, Madsen LH, Bhasin H, Sexauer M, Stougaard J, Markmann K. Systemic control of legume susceptibility to rhizobial infection by a mobile microRNA. Science. 2018;362:233–6.
Waititu JK, Zhang C, Liu J, Wang H. Plant non-coding RNAs: origin, biogenesis, mode of action and their roles in abiotic stress. Int J Mol Sci. 2020;21:8401. https://doi.org/10.3390/ijms21218401.
Wang JW, Czech B, Weigel D. miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell. 2009;138:738–49.
Wang Q, et al. Divide et impera: boundaries shape the plant body and initiate new meristems. New Phytol. 2016;209:485–98.
Wang S, Sun X, Hoshino Y, Yu Y, Jia B, Sun Z, Sun M, Duan X, Zhu Y. MicroRNA319 positively regulates cold tolerance by targeting OsPCF6 and OsTCP21 in Rice (Oryza sativa L.). PLoSOne. 2014;9:e91357. https://doi.org/10.1371/journal.pone.0091357.
Wang Y, Gao L, Li J, Zhu B, Zhu H, Luo Y, et al. Analysis of long-non-coding RNAs associated with ethylene in tomato. Gene. 2018;674:151–60.
Wang W, Liu D, Chen DD, Cheng YY, Zhang XP, Song LR, Hu MJ, Dong J, Shen FF. MicroRNA414c affects salt tolerance of cotton by regulating reactive oxygen species metabolism under salinity stress. RNA Biol. 2019;16:362–75.
Wei W, Li G, Jiang X, Wang Y, Ma Z, Niu Z, et al. Small RNA and degradome profiling involved in seed development and oil synthesis of Brassica napus. PLoS ONE. 2018;13:e0204998.
Wollmann H, et al. On reconciling the interactions between APETALA2, miR172 and AGAMOUS with the ABC model of flower development. Development. 2010;137:3633–42.
Wu G, et al. The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell. 2009;138:750–9.
Wu G, Poethig RS. Temporal regulation of shoot development in Arabidopsis thaliana by miR156 and its target SPL3. Development. 2006;133:3539–47.
Wu MF, et al. Arabidopsis microRNA167 controls pat-terns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development. 2006;133:4211–8.
Xia J, Zeng C, Chen Z, et al. Endogenous small-noncoding RNAs and their roles in chilling response and stress acclimation in Cassava. BMC Genomics. 2014;15:1–19.
Xia K, Wang R, Ou X, Fang Z, Tian C, Duan J, Wang Y, Zhang M. OsTIR1 and OsAFB2 downregulation via OsmiR393 overexpression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS ONE. 2012;7:e30039.
Xia K, Zeng X, Jiao Z, Li M, Xu W, Nong Q, Mo H, Cheng T, Zhang M. Formation of protein disulfide bonds catalyzed by OsPDIL1;1 is mediated by MicroRNA5144-3p in rice. Plant Cell Physiol. 2018;59:331–42.
Xie K, et al. Genomic organization, differential expres-sion, and interaction of SQUAMOSA promoter-binding-like tran-scription factors and microRNA156 in rice. Plant Physiol. 2006;14:280–93.
Xie Z, Allen E, Wilken A, Carrington JC. DICER-LIKE4functions in trans-acting small interfering RNA biogenesis and vegetative phase change in Arabidopsis thaliana. Proc Natl Acad Sci USA. 2005;102:12984–9. https://doi.org/10.1073/pnas.0506426102.
Xin M, Wang Y, Yao Y, Xie C, Peng H, Ni Z, Sun Q. Diverse set of microRNAs are responsive to powdery mildew infection and heat stress in wheat (Triticum aestivum L.). BMC Plant Biol. 2010;10:1–11.
Yan K, Liu P, Wu CA, Yang GD, Xu R, Guo QH, Huang JG, Zheng CC. Stress-induced alternative splicing provides mechanism for the regulation of microRNA processing in Arabidopsis thaliana. Mol Cell. 2012;48:521–31.
Yang C, Li D, Mao D, Liu X, Ji C, Li X, Zhao X, Cheng Z, Chen C, Zhu L. Overexpression of micro RNA 319 impacts leaf morphogenesis and leads to enhanced cold tolerance in rice (Oryza sativa L.). Plant Cell Environ. 2013;36:2207–18.
Yant L, et al. Orchestration of the floral transition and floral development in Arabidopsis by the bifunctional transcription factor APETALA2. Plant Cell. 2010;22:2156–70.
Yu YH, Ni ZY, Wang Y, Wan HN, Hu Z, Jiang QY, Sun XJ, Zhang H. Overexpression of soybean miR169c confers increased drought stress sensitivity in transgenic Arabidopsis thaliana. Plant Sci. 2019;285:68–78.
Yuan J, Li J, Yang Y, Tan C, Zhu Y, Hu L, et al. Stress-responsive regulation of long non-coding RNA polyadenylation in Oryza sativa. Plant J. 2018;93:814–27.
Yuan S, Li Z, Li D, Yuan N, Hu Q, Luo H. Constitutive expression of rice microRNA528 alters plant development and enhances tolerance to salinity stress and nitrogen starvation in creeping bent grass. Plant Physiol. 2015;169:576–93.
Zou CL, Wang YB, Wang B, Liu D, Liu L, Gai ZJ, Li CF. Long non-coding RNAs in the alkaline stress response in sugar beet (Beta vulgaris L.). BMC Plant Biol. 2020;20:227.
Zhang F, Luo X, Zhou Y, Xie J. Genome-wide identification of conserved microRNA and their response to drought stress in Dongxiang wild rice (Oryza rufipogon Gri.). Biotechnol Lett. 2016;38:711–21.
Zhang J, Zhang H, Srivastava AK, Pan BJ, Fang J, Shi H, Zhu JK. Knockdown of rice microRNA166 confers drought resistance by causing leaf rolling and altering stem xylem development. Plant Physiol. 2018;176:2082–94. https://doi.org/10.1104/pp.17.01432.
Zhang X, Wang W, Wang M, Zhan HY, Liu JH. The miR396b of Poncirus trifoliata functions in cold tolerance by regulating ACC oxidase gene expression and modulating ethylene–polyamine homeostasis. Plant Cell Physiol. 2016;57:1865–78. https://doi.org/10.1093/pcp/pcw108.
Zhang XP, Dong J, Deng FN, Wang W, Cheng YY, Song LR, Hu MJ, Shen J, Xu QJ, Shen FF. The long non-coding RNA lncRNA973 is involved in cotton response to salt stress. BMC Plant Biol. 2019;19:459.
Zhao J, He Q, Chen G, Wang L, Jin B. Regulation of non-coding RNAs in heat stress responses of plants. Front Plant Sci. 2016;7:1213. https://doi.org/10.3389/fpls.2016.01213.
Zhao J, Yuan S, Zhou YN, Li Z, Hu BFG, Liu H, Li S, Luo H. Transgenic creeping bent grass overexpressing Osa-miR393a exhibits altered plant development and improved multiple stress tolerance. Plant Biotechnol J. 2018. https://doi.org/10.1111/pbi.12960.
Zhao L, et al. miR172 regulates stem cell fate and defines the inner boundary of APETALA3 and PISTILLATA expression domain in Arabidopsis floral meristems. Plant J. 2007;51:840–9.
Zhou M, Tang W. MicroRNA156 amplifies transcription factor-associated cold stress tolerance in plant cells. Mol Genet Genom. 2019;294:379–93.
Zhdanov VP. Stochastic bursts in the kinetics of gene expression with regulation by long non-coding RNAs. JETP Lett. 2010;92:410–5.
Zheng B, Wang Z, Li S, Yu B, Liu JY, Chen X. Intergenic transcription by RNA polymerase II coordinates Pol IV and Pol V in siRNA-directed transcriptional gene silencing in Arabidopsis. Genes Dev. 2009;23:2850–60.
Zhu H, Hu F, Wang R, Zhou X, Sze SH, Liou LW, Barefoot A, Dickman M, Zhang X. Arabidopsis Argonaute10 specifically sequesters miR166/165 to regulate shoot apical meristem development. Cell. 2011;145:242–56.
Zuo ZF, He W, Li J, Mo B, Liu L. Small RNAs: the essential regulators in plant thermotolerance. Front Plant Sci. 2021;12:726762. https://doi.org/10.3389/fpls.2021.726762.
Acknowledgements
The authors express their gratitude to the Council of Scientific & Industrial Research (CSIR)-University Grant Commission (UGC), New Delhi, India for financial support in the form of Senior Research Fellowship to Miss. Rinku Mondal (2018-MAY-351826) and Mr. Adwaita Das (09/025(0250)/2018-EMR-I)
The authors would like to acknowledge Department of Botany, The University of Burdwan for research and infrastructure facilities.
Author information
Authors and Affiliations
Contributions
RM and AD wrote the manuscript. AB provided the critical comments and edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to disclose.
Additional information
Corresponding Editor: Sachin Rustagi; Reviewers: Anjana Rustagi, Neeraj Kumar Vasistha.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mondal, R., Das, A. & Bandyopadhyay, A. Implications of small RNAs in plant development, abiotic stress response and crop improvement in changing climate. Nucleus 66, 321–339 (2023). https://doi.org/10.1007/s13237-023-00447-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13237-023-00447-1