Abstract
Potato virus Y (PVY) causes one of the most serious and widespread diseases in North America. In recent years, the virus has become increasingly difficult to control. Durable dominant genes for resistance to PVY exist in potato germplasm and provide an effective control strategy. This paper describes a Solanum chacoense clone (M19) that is homozygous for a PVY resistance gene. The gene is linked to a previously published marker for Rychc found in Solanum chacoense. M19 was crossed with a diploid S. tuberosum clone to produce an adapted clone carrying the resistance gene. This hybrid clone is named M20. M20 tuberizes in the field, producing round tubers with white skin and flesh and moderate size. M20 is resistant to PVYO, PVYN:O, and PVYNTN. Both M19 and M20 are female and male fertile, so they are being released as sources of PVY resistance for breeding programs.
Resumen
El Virus Y de la Papa (PVY) causa una de las enfermedades más serias y de amplia distribución en Norteamérica. En años recientes, el virus ha aumentado la dificultad para su control. Los genes durables dominantes para resistencia al PVY existen en el germoplasma de papa y proporcionan una estrategia de control efectivo. Este artículo describe un clon (M19) de Solanum chacoense que es homozigótico para un gen de resistencia al PVY. El gen esta ligado a un marcador previamente publicado para Rychc encontrado en Solanum chacoense. Se cruzó M19 con un clon diploide de S. tuberosum para producir un clon adaptado llevando el gen de resistencia. A este clon híbrido se le llama M20. Este clon tuberiza en el campo, produciendo tubérculos redondos con piel y pulpa blancas, de tamaño moderado. M20 es resistente al PVYO, PVYN:O y PVYNTN. Ambos M19 y M20 son fértiles como hembra y macho, de manera que están siendo liberados como fuentes de Resistencia al PVY para los programas de mejoramiento.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Potato virus Y (PVY) is one of the most significant potato pathogens worldwide (German 2001; Scholthof et al. 2011). In recent decades, it has emerged as one of the most common and serious potato pathogens in the United States (Karasev and Gray 2013). Infection of potato tubers and foliage with PVY leads to yield losses and reductions in tuber quality. Yield losses of up to 80% have been reported (German 2001; Kopp et al. 2015).
PVY is a member of the genus Potyvirus and exists as a single-stranded, positive-sense RNA (Berger et al. 2005). The host range of PVY includes several solanaceous plants, including tomato, potato, pepper and tobacco (German 2001). PVY is transmitted across generations through infected tubers. During the growing season, non-persistent aphid transmission is responsible for spread of the pathogen. PVY can also be mechanically carried from plant to plant through foliar contact or through contaminated equipment during seed cutting (Quenouille et al. 2013).
Until the beginning of this century, seed certification programs effectively controlled PVY in North America. Then, several events converged, leading to PVY becoming a major potato pathogen. First, several new related cultivars, including Russet Norkotah, Shepody, and Silverton Russet, became some of the most popular cultivars and were planted on hundreds of thousands of acres. These highly susceptible cultivars develop mild or no symptoms when infected with PVY (Hane and Hamm 1999; Nolte et al. 2004), so growers are unable to rogue infected plants or easily identify fields that serve as regional inoculum sources. Seed inspectors rely mainly on visual symptoms when certifying seed lots, so it became more common for seed lots containing higher levels of PVY to pass inspection. Second, new recombinant PVY strains (PVYN:O, PVYN-Wi, and PVYNTN) which produce milder symptoms than the ordinary strain, PVYO, are now predominant in North America (Karasev and Gray 2013; MacKenzie et al. 2018). Of these strains, PVYNTN is the most troubling since it often produces necrotic tuber symptoms in some popular cultivars, such as Yukon Gold. PVYN:O and PVYN-Wi can also produce necrotic tuber symptoms, but fewer isolates of these strains are associated with these symptoms compared to PVYNTN isolates. Finally, the introduction of the soybean aphid into the northern U.S. in the early 2000’s brought a new PVY vector that arrived in potato fields in large numbers (Gray et al. 2010).
The most important PVY control strategies are seed certification and host plant resistance. Since seed certification has become less effective at controlling PVY and since PVY strains that produce necrotic tuber symptoms are more prevalent in North America, interest in breeding for resistance has increased. Unlike most potato diseases, major dominant resistance genes for PVY exist in cultivated and wild potato germplasm. Extreme resistance, which protects against all PVY strains, has been identified in the wild potato species S. chacoense, S. hougasii, S. stoloniferum and the S. tuberosum Andigenum Group (Cockerham 1943; Munoz et al. 1975; Sato et al. 2006). The Rysto and Ryadg genes, from S. stoloniferum and S. tuberosum Andigenum, respectively, have been used extensively in breeding programs for decades in Europe and, more recently, in North America. These resistance genes have been remarkably durable (Quenouille et al. 2013). The third resistance gene, Rychc from S. chacoense is used less frequently by potato breeders in North America and Europe. Rychc has, however, been introduced into the Japanese cultivars Sakurafubuki (Murakami et al. 1995) and Konafubuki (Asama et al. 1982) and the germplasm release Saikai 35 (Mori et al. 2012). No linkage drag to inferior agronomic traits has been found when introgressing Rychc (Murakami et al. 1995). The three resistance genes (Rysto, Ryadg, and Rychc) have been mapped to chromosomes XII (Flis et al. 2005; Song et al. 2005), XI (Hamalainen et al. 1997), and IX (Hosaka et al. 2001; Sato et al. 2006), respectively.
Molecular markers for Rysto and Ryadg have been used by North American breeding programs when introgressing those genes (Ottoman et al. 2007; Vales et al. 2010; Whitworth et al. 2009). PVY resistant clones carrying Rysto have been reported to be male sterile, limiting their use to female parents only in breeding programs (Fulladolsa et al. 2015; Song and Schwarzfischer 2008; Vales et al. 2010). This is a problem because male sterility is very common in breeding programs and the number of male parents is typically limited. Ryadg is not associated with male sterility (Vales et al. 2010). We have generated and previously published two TaqMan® SNP markers and two SCAR markers linked to Rychc. All four markers are tightly linked to Rychc and are efficient for selecting resistant individuals, with less than 5.2% recombination (Fulladolsa et al. 2017).
In this paper, we present two PVY resistant clones for use in breeding. The first, M19, is a S. chacoense clone homozygous for the dominant resistance gene. Consequently, when used in breeding, all offspring will be PVY resistant. The second clone, M20, is heterozygous, but is more adapted. It is a hybrid between the dihaploid US-W4 and M19.
Materials and Methods
Germplasm
An evaluation of a diverse set of wild potato relatives identified several S. chacoense accessions with PVY resistance (Cai et al. 2011). Within these accessions, clone 39–7 (M19) from accession 275138 was selected as a source of PVY resistance because crosses using this clone revealed that it is homozygous for PVY resistance. M19 was crossed as a male to the S. tuberosum dihaploid US-W4 (De Jong and Rowe 1971), and clone XD3 (M20) was selected from the resulting family based on its high degree of fertility and self-compatibility.
PVY Resistance Screening
In a greenhouse trial in Wisconsin, three M20 plants were mechanically inoculated with each strain, and three plants were mock-inoculated. Three weeks post-inoculation, leaf samples were analyzed using DAS-ELISA. In field trials in Idaho, M20 plants were inoculated with PVYO, PVYN:O, and PVYNTN both mechanically and by aphids during the field season, across three replications for 2 years (2013 and 2014). Similar trials were carried out in New York, in a greenhouse in 2013 and the field in 2014 (Table 1). In 2013, tubers from the New York 2013 trial were planted into the field adjacent to healthy tubers. Infection due to natural aphid inoculation was recorded.
Fertility
Pollen from greenhouse-grown plants was collected, stained with 1% acetocarmine and evaluated using a compound microscope at 100x. M19 and M20 were used as parents in greenhouse crosses between 2011 and 2017 (Table 2). M19 was crossed as a female to three diploid parents, as a male to one diploid parent and it was also self-pollinated. M20 was crossed as a female to two diploid parents, as a male to 17 diploid parents, and it was self-pollinated.
2n Gamete Production
To evaluate 2n gamete production and the potential for the two clones to be used in tetraploid breeding programs, they were crossed as males and females to tetraploid cultivars (Table 3). Pollen was also observed microscopically for the presence diploid gametophytes.
Verticillium Wilt Resistance
M20 was included in the 2012 National Verticillium wilt resistance trial, planted on May 8 on a Verticillium-infested field at the Hancock, WI, Agricultural Experiment Station. Three replications of five-hill units were planted on a field that was inoculated with V. dahliae in 2006 and has been maintained as a Verticillium wilt screening plot. On August 29, vines were killed and on September 4, stems were collected and allowed to air dry at room temperature. All main stems from a plot were ground in a Wiley mill and 50 mg per plot were plated on selective medium. After 2 weeks of incubation in the dark at room temperature, plates were evaluated microscopically for the presence of V. dahliae colonies. On September 11, each plot was harvested with a single row digger, and tubers were picked up by hand and weighed. Tubers were then stored at 6 °C until January 7, 2013, when they were fried and visually scored for chip color (1 = light, 10 = dark; 4 or lower is acceptable).
Glycoalkaloids
Glycoalkaloid analyses were carried out by Dr. Brian Perkins (University of Maine) on one set of M19 tubers and two sets of M20 tubers. Alpha-solanine and alpha-chaconine were quantified by reverse-phase liquid chromatography (RP-LC). Approximately 2 g of powdered, lyophilized sample was added to an acidified ion pairing solution (0.02 M 1-heptanesulfonic acid sodium salt, monohydrate in 2% (v/v) acetic acid) and extracted for 3 min with a Polytron tissue homogenizer. The resulting extract was centrifuged to pellet tissue debris and a 10 mL aliquot of the supernatant was passed through a methanol activated tC-18 SPE (Waters Corp, Milford, MA, Cat # WAT036810) cartridge followed by an acetonitrile-water (20:80, v/v) wash. After vacuum drying, the sample was eluted from the cartridge with a tetrahydrofuran-wateracetonitrile (50:30:20, v/v) solution and 20 μl was injected into the LC system, equipped with a diode array detector (DAD) for analysis. Separation was accomplished on a C-6 analytical column, with a buffered (pH 3.5) mobile phase.
Dr. David Douches (Michigan State University) also determined glycoalkaloid content of a set of M20 tubers. A mixture of 50 mg of ground tissue and 1 ml of extraction solution (methanol/water/glacial acetic acid in a ratio 49:49:2, v/v/v) was homogenized by vortexing. Samples were incubated at 60 °C for 30 min and then centrifuged at 14000 rpm for 1 min. The supernatant was filtered using Corning Costar® Spin-X centrifuge tube filter equipped with a 0.22 μm pore size nylon membrane (Sigma-Aldrich, Inc.). The final extract was diluted 50 fold and complemented with 0.5 μM telmisartan (internal standard for quantification). Targeted glycoalkaloid quantification was conducted at the Mass Spectrometry and Metabolomics Facility at Michigan State University. The most common glycoalkaloids (α-chaconine and α-solanine) were separated using a reverse phase Supelco® Ascentis Express C-18 column (50 × 2.1 mm, 2.7 μm particle size) and analyzed by liquid chromatography/mass spectrometry (LC/MS) using a Waters (Milford, MA) ACQUITY® system and TQ Detector interfaced to a Waters Acquity binary solvent manager and 2777c autosampler. Steroidal glycoalkaloids were eluted using a binary gradient system with solvent A (0.1% formic acid in water) and solvent B (100% acetonitrile) at a flow rate of 0.3 ml/min and starting conditions of 10% B. The gradient elution was 0–2 min, 10% to 100% B; 2–3 min, hold at 100% B; 3.01 min return to 10% B and hold to 4 min. Standard solutions of α-chaconine and α-solanine at different concentrations (0.625 to 40 μl) with 0.5 μM internal standard were injected (10 μM) to obtain the calibration curve. Targeted detection was used to quantify α-chaconine and α-solanine glycoalkaloids. Peak areas and calibration curves were generated using the Quanlynx tool in Masslynx (Waters).
Results and Discussion
PVY Resistance
M20 showed resistance to three PVY strains, PVYO, PVYN:O, and PVYNTN. In the greenhouse trial, no virus was detected using DAS-ELISA. M20 plants did not exhibit any typical foliar symptoms in greenhouse or field trials and analysis of plant samples using DAS-ELISA (Clark and Adams 1977) did not detect any virus antigen. Furthermore, tubers harvested from inoculated plants did not generate PVY-infected daughter plants (Table 1).
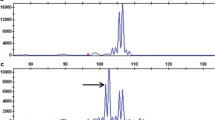
Molecular markers 38–530 (Hosaka et al. 2001), Ry186 (Mori et al. 2011), and Ry364 (Mori et al. 2012), linked to Rychc were used to genotype M19. A 587 bp fragment associated with Ry186 was detected, but markers 38–530 and Ry364 were not present. This suggests that the PVY resistance gene in M19 could be Rychc. In a subsequent manuscript, we will describe PVY resistance markers in this germplasm.
Fertility
Pollen stainability of M19 and M20 was 95% and 90%, respectively. Both M19 and M20 are male and female fertile, as shown by pollination data (Table 2). They are both self-compatible, but M19 produced only a few seeds following a large number of pollinations. M20 is much more self-fertile, producing an abundance of spontaneous berries in the greenhouse. Two M20 plants in the greenhouse produced 292 and 248 berries, containing a total of 4750 and 4000 seeds, respectively. High levels of self-compatibility in M20 are expected, since US-W4 is male fertile (De Jong and Rowe 1971).
2n Gamete Production
Microscopic evaluations of M19 and M20 pollen did not reveal evidence of 2n pollen production. This observation supported the pollination data. A few seeds were produced following large numbers of pollinations of M20 onto emasculated tetraploid White Pearl flowers, indicating that 2n pollen is functioning at a low level (Table 3). Similarly, a low level of 2n eggs appears to be present in M20, as four seeds were generated from 110 pollinations.
Verticillium Wilt Resistance
Across three replications in the Verticillium wilt trial, M20 produced an average of 54 V. dahliae propagules per 50 mg of dried basal stem tissue. For comparison, propagule counts in susceptible Russet Norkotah and resistant Ranger Russet averaged 121 and 83, respectively. Consequently, M20 appears to have moderate resistance to Verticillium wilt. Additional trials are needed to confirm the presence of stable resistance.
M20 tubers exhibit high specific gravity and moderate resistance to cold-induced sweetening (Table 4). Yields were measured on small plots in a V. dahliae-infested field. Nevertheless, these data demonstrate that M20 produces moderate yields in the field (Fig. 1).
Glycoalkaloids
Glycoalkaloids were not detected in M19 tubers. Total glycoalkaloids ranged from 4.53 to 5.8 mg/100 g FW in M20, which is well below the acceptable level of 20 mg/100 g FW. Since it is common for wild potatoes to produce high levels of glycoalkaloids, we did not expect to see the extremely low levels detected by the University of Maine lab. Tubers were then sent to another lab (Michigan State University), which confirmed the low levels in M20. M19 tubers are difficult to generate in quantity because wild species produce small tubers. Consequently, only M20 tubers were re-tested.
References
Asama, K., H. Ito, N. Murakami, and T. Itoh. 1982. New potato variety “Konafubuki”. Bull Hokkaido Pref Agr Exp Station 48: 75–84.
Berger, P.H., M.J. Adams, O.W. Barnett, A.A. Brunt, J. Hammond, et al. 2005. Family potyviridae. In Virus taxonomy: Classification and nomenclature of viruses. 8th report of the international committee on taxonomy of viruses, ed. C.M. Fauquet, M.A. Mayo, J. Maniloff, U. Desselberger, and L.A. Ball, 819–841. London: Elsevier Acad.
Cai, X., D. Spooner, and S. Jansky. 2011. A test of taxonomic and biogeographic predictivity: Resistance to Potato virus Y in wild relatives of the cultivated potato. Phytopathology 101: 1074–1080.
Clark, M.F., and A.N. Adams. 1977. Characteristics of the microplate method of enzyme linked immunosorbent assay for the detection of plant viruses. Journal of General Virology 34: 475–483.
Cockerham, G. 1943. Potato breeding for virus resistance. Annals of Applied Biology 30: 105–108.
De Jong, H., and P.R. Rowe. 1971. Inbreeding in cultivated diploid potatoes. Potato Research 14: 74–83.
Flis, B., J. Hennig, D. Strzelczyk-Zyta, C. Gebhardt, and W. Marczewski. 2005. The Ry-fsto gene from Solanum stoloniferum for extreme resistant to Potato virus Y maps to chromosome XII and is diagnosed by PCR marker GP122 718 in PVY resistant potato cultivars. Molecular Breeding 15: 95–101.
Fulladolsa, A.C., F.M. Navarro, R. Kota, K. Severson, J.P. Palta, and A.O. Charkowski. 2015. Application of marker assisted selection for Potato virus Y resistance in the University of Wisconsin Potato Breeding Program. American Journal of Potato Research 92: 444–450.
Fulladolsa, A. C., Jansky, S. H., Smith, D. R., Abramczak, C. M., & Charkowski, A. O. 2017. Development and evaluation of four molecular markers tightly linked to the Potato virus Y resistance gene Ry chc in diploid potato populations. American Phytopathological Society 258-P.
German, T.L. 2001. Potato virus Y. In Compendium of potato diseases, ed. W.R. Stevenson, R. Loria, G.D. Franc, and D.P. Weingartner, 2nd ed., 69–71. Minnesota: The American Phytopathological Society Press.
Gray, S., S. De Boer, J. Lorenzen, A. Karasev, J. Whitworth, P. Nolte, R. Singh, A. Boucher, and H. Xu. 2010. Potato virus Y: An evolving concern for potato crops in the United States and Canada. Plant Disease 94: 1384–1397.
Hamalainen, J.H., K.N. Watanabe, J.P. Valkonen, A. Arihara, R.L. Plaisted, E. Pehu, L. Miller, and S.A. Slack. 1997. Mapping and marker-assisted selection for a gene for extreme resistance to Potato virus Y. Theoretical and Applied Genetics 94: 192–197.
Hane, D.C., and P.B. Hamm. 1999. Effects of seedborne Potato virus Y infection in two potato cultivars expressing mild disease symptoms. Plant Disease 83: 43–45.
Hosaka, K., Y. Hosaka, M. Mori, T. Maida, and H. Matsunaga. 2001. Detection of a simplex RAPD marker linked to resistance to Potato virus Y in a tetraploid potato. American Journal of Potato Research 78: 191–196.
Karasev, A.V., and S.M. Gray. 2013. Continuous and emerging challenges of Potato virus Y in potato. Annual Review of Phytopathology 51: 571–586.
Kopp, A., M. Kondrák, and Z. Bánfalvi. 2015. Review article: Molecular mechanisms of resistance to potato virus X and Y in potato. Acta Phytopathologica et Entomologica Hungarica 50: 151–160.
MacKenzie, T.D.B., J. Lavoie, X. Nie, and M. Singh. 2018. Differential spread of Potato virus Y (PVY) strains O, N:O and NTN in the field: Implications for the rise of recombinant PVY strains in New Brunswick, Canada. American Journal of Potato Research 95: 301–310.
Mori, K., Y. Sakamoto, N. Mukojima, S. Tamiya, T. Nakao, T. Ishii, and K. Hosaka. 2011. Development of a multiplex PCR method for simultaneous detection of diagnostic DNA markers of five disease and pest resistance genes in potato. Euphytica 180: 347–355.
Mori, K., N. Mukojima, T. Nakao, S. Tamiy, Y. Sakamoto, N. Sohbaru, K. Hayashi, H. Watanuki, K. Nara, K. Yamazaki, T. Ishii, and K. Hosaka. 2012. Germplasm release: Saikai 35, a male and female fertile breeding line carrying Solanum phureja-derived cytoplasm and potato cyst nematode resistance (H1) and potato virus Y resistance (Ry chc) genes. American Journal of Potato Research 89: 63–72.
Munoz, R.J., R.L. Plaisted, and H.D. Thurston. 1975. Resistance to Potato virus Y in Solanum tuberosum ssp. andigena. American Potato Journal 52: 107–115.
Murakami, N., H. Matsunaga, K. Senda, Y. Okuyama, M. Iritani, K. Asama, Y. Mitsui, and K. Shimizu. 1995. A new potato variety “Konamuso” (=Sakaruafubuki)". Bull Hokkaido Pref Agr Exp Station 68: 1–16.
Nolte, P., J.L. Whitworth, M.K. Thornton, and C.S. McIntosh. 2004. Effect of seedborne Potato virus Y on performance of Russet Burbank, Russet Norkotah, and Shepody potato. Plant Disease 88: 248–252.
Ottoman, R.J., D. Hane, C. Brown, S. Yilma, A. Mosley, and M.I. Vales. 2007. Usefulness of molecular markers to screen for PVY resistance (Ry adg gene) in potato. American Journal of Potato Research 84: 108.
Quenouille, J., N. Vassilakos, and B. Moury. 2013. Potato virus Y: A major crop pathogen that has provided major insights into the evolution of viral pathogenicity. Molecular Plant Pathology 14: 439–452.
Sato, M., K. Nishikawa, K. Komura, and K. Hosaka. 2006. Potato virus Y resistance gene, Ry chc, mapped to the distal end of potato chromosome 9. Euphytica 149: 367–372.
Scholthof, K.-B.G., S. Adkins, H. Czosnek, P. Palukaitis, E. Jacquot, T. Hohn, B. Hohn, K. Saunders, T. Candresse, P. Ahlquist, C. Hemenway, and G.D. Foster. 2011. Top 10 plant viruses in molecular plant pathology. Molecular Plant Pathology 12: 938–954.
Song, Y.-S., and A. Schwarzfischer. 2008. Development of STS markers for selection of extreme resistance (Ry sto) to PVY and maternal pedigree analysis of extremely resistant cultivars. American Journal of Potato Research 85: 159–170.
Song, Y.-S., L. Hepting, G. Schweizer, L. Hartl, G. Wenzel, and A. Schwarzfischer. 2005. Mapping of extreme resistance to PVY (Ry sto) on chromosome XII using anther-culture-derived primary dihaploid potato lines. Theoretical and Applied Genetics 111: 879–887.
Vales, M.I., R.J. Ottoman, J.A. Ortega, S. Yilma, and E. Karaagac. 2010. Marker-assisted selection for PVY resistance in tetraploid potatoes. Acta Horticulturae 859: 409–416.
Whitworth, J.L., R.G. Novy, D.G. Hall, J.M. Crosslin, and C.R. Brown. 2009. Characterization of broad spectrum Potato Virus Y resistance in a Solanum tuberosum ssp. andigena -derived population and select breeding clones using molecular markers, grafting, and field inoculations. American Journal of Potato Research 86: 286–296.
Acknowledgements
This work is supported by SCRI grant number 73999-10921 from USDA National Institute of Food and Agriculture. Any opinions, findings, conclusions, or recommendations expressed in this publication are those of the author(s) and do not necessarily reflect the view of the U.S. Department of Agriculture. The authors thank Drs. Douches and Perkins for their assistance with glycoalkaloid analysis.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fulladolsa, A.C., Charkowski, A., Cai, X. et al. Germplasm with Resistance to Potato virus Y Derived from Solanum chacoense: Clones M19 (39–7) and M20 (XD3). Am. J. Potato Res. 96, 390–395 (2019). https://doi.org/10.1007/s12230-019-09719-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12230-019-09719-6