Abstract
The essentiality of 14 mineral elements so far have been reported in plant nutrition. Eight of these elements were known as micronutrients due to their lower concentrations in plants (usually ≤100 mg/kg/dw). However, it is still challenging to mention an exact number of plant micronutrients since some elements have not been strictly proposed yet either as essential or beneficial. Micronutrients participate in very diverse metabolic processes, including from the primary and secondary metabolism to the cell defense, and from the signal transduction to the gene regulation, energy metabolism, and hormone perception. Thus, the attempt to understand the molecular mechanism(s) behind their transport has great importance in terms of basic and applied plant sciences. Moreover, their deficiency or toxicity also caused serious disease symptoms in plants, even plant destruction if not treated, and many people around the world suffer from the plant-based dietary deficiencies or metal toxicities. In this sense, shedding some light on this issue, the 13 mineral elements (Fe, B, Cu, Mn, Mo, Si, Zn, Ni, Cl, Se, Na, Al, and Co), required by plants at trace amounts, has been reviewed with the primary focus on the transport proteins (transporters/channels) in plant roots. So, providing the compiled but extensive information about the structural and functional roles of micronutrient transport genes/proteins in plant roots.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Plants are usually reported to need about 14 essential mineral elements for healthy growth and development [1]. From these elements, eight such as boron (B), chlorine (Cl), copper (Cu), iron (Fe), manganese (Mn), molybdenum (Mo), nickel (Ni), and zinc (Zn) are known as micronutrients since they are found in plants at lower concentrations (usually ≤100 mg/kg/dry weight (dw)) [1]. However, it is still challenging to mention an exact number of the micronutrients since some elements have not been strictly proposed yet either as essential or beneficial. For example, some authors report silicon (Si) in the micronutrient list while others mention it as a beneficial element [1–3]. Further molecular and physiological studies are thus required whether to include or exclude these controversial elements in the micronutrient list. Micronutrients participate in a number of metabolic processes, including primary and secondary metabolism, cell defense, signal transduction, energy metabolism, hormone perception, and gene regulation [1–4]. Taking into account that the very diverse group of physiological events in plant metabolism are directly or somewhat associated with the mineral elements, elucidating the molecular mechanism behind the element uptake and transport appears to have great significance in terms of basic and applied plant sciences [4, 5]. In addition, the element deficiencies or toxicities seriously affect the plant life cycle by causing the various symptoms, even plant death if not treated, and millions of people around the world also suffer from the plant-based element deficiencies or toxicities [6–8]. Regarding the essentiality and/or beneficial effects of the micronutrients in plants, this study has attempted to review 13 mineral elements such as iron (Fe), boron (B), copper (Cu), manganese (Mn), molybdenum (Mo), silicon (Si), zinc (Zn), nickel (Ni), chlorine (Cl), selenium (Se), sodium (Na), aluminum (Al), and cobalt (Co) with an emphasis on the micronutrient transporters or channels in plant roots. In this context, the very diverse group of root uptake micronutrient transporters or channels from various protein families was compiled and provided an easy-access information about their structural and functional roles in plants.
Iron (Fe)
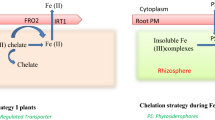
Fe is one of the important essential micronutrients in plants and also crucial for the plant growth and development [9, 10]. It participates in a number of biological processes, including respiration, photosynthesis, hormone production, nitrogen fixation, and chlorophyll and DNA syntheses. In addition, it functions as structural element in iron-sulfur clusters, heme cofactor, and other iron-binding sites [11–13]. Plants require about 10−9–10−4 M of Fe concentration to maintain the normal growth and development. Its deficiency causes the interveinal chlorosis in plant leaves and reduces the crop productivity; if not treated, even it could lead to plant death [14]. The Fe3+ form is abundant in nature but it is not directly available by plants at normal physiological conditions due to its less solubility [15]. Thus, iron transport necessitates either its reduction to Fe2+ or its chelation at root rhizosphere [16]. Moreover, Fe is a suitable element for redox reactions due to its chemical properties. However, its large quantities in free-state could also lead the generation of reactive oxygen species (ROS) [5]. So, its uptake, mobilization, utilization, and partitioning in plants require a tight regulation at cellular and molecular levels [11, 12, 17]. Regarding Fe uptake, two different strategies have been proposed for graminaceous and non-graminaceous plants [18]. In graminaceous plants, its uptake depends on the secretion of phytosiderophores (MAs) which are synthesized by S-adenosyl-L-methionine (SAM) pathway with three consecutive enzymatic reactions such as nicotianamine synthase (NAS), nicotianamine aminotransferase (NAAT), and deoxymugineic acid synthase (DMAS) [19–21]. The secreted MAs in rhizosphere solubilized the Fe3+ and formed an Fe3+-MA complex followed by its uptake into the root cells by Yellow Stripe 1-like (YSL) and Yellow Stripe 1 (YS1) transporters [22–24]. In non-graminaceous plants, Fe chelates are reduced at the root surface and the resulting ferrous ions (Fe2+) are taken into the cells through the plasma membrane. The phenolic compounds and protons, secreted into rhizosphere, also accompany this process to increase the ferric ions (Fe3+) solubility [12, 25]. Besides, some plants like rice demonstrate both types of mechanisms such as Fe3+-DMA uptake by OsYSL15 transporter as graminaceous plants and uptake by ferrous transporter OsIRT1 as non-graminaceous plants [22, 23]. For cytosolic Fe transport, several transporter families such as iron-regulated transporter (IRT)-like protein (ZIP), zinc-regulated transporter (ZRT), natural resistance-associated macrophage protein (NRAMP), and oligo peptide transporters (OPTs) have been reported so far [26]. ZIP family is a type of broad range metal transporters carrying the Fe2+, Zn2+, Mn2+, and Cd2+ [26, 27]. This family proteins are 309–476 residues long with eight putative TMDs and extracellular N- and C-terminals [27]. They contain a variable cytosolic region between TMD3 and 4 with a putative His-rich metal-binding domain [27]. TMD4 also harbors the most conserved part of the ZIP proteins with an amphipathic helix including a full conserved His residue. It has been also reported that this His amino acid and a semi polar adjacent residue could form an intramembranous metal-binding site [28]. As for the NRAMP family, they also contain metal transporters with broad subtract range such as Fe2+, Mn2+, Cu2+, Zn2+, Cd2+, Ni2+, Al3+, and Co2+ [29–31]. These family members have 10–12 putative TMDs with a consensus transport residue between TMD8 and 9 [32]. Members of the OPT family transport the metal ions such as Fe, Zn, Mn, Ni, Cd, and Cu, and amino acid-containing compounds and their derivatives [33, 34]. Thinking that most iron transporters have broad subtract range for different metals, therefore, plants could have developed a sophisticated iron homoeostasis maintaining the physiological iron limits.
Zinc (Zn)
Like Fe, Zn is also a crucial micronutrient in the plant life cycle due to its essentiality [35]. As a catalytic element, it functions in more than 300 enzymes, including alkaline phosphatase, alcohol dehydrogenase, carbonic anhydrase, and Cu-Zn superoxide dismutase. It also plays a structural role in stabilization of many proteins such as Zn cluster, Zn finger, and RING finger domains/motifs [36]. Its plant deficiency could cause the chlorosis, the formation of necrotic spots, and could reduce the plant growth [37]. From soil, it is taken as divalent cations (Zn2+) which also determine its physiological role since it is neither further reduced nor oxidized intracellularly [5, 36]. Zn homeostasis in plants is mainly maintained by the coordinated regulation of ZIP family uptake transporters [27, 38; refer to Fe (iron)]. In model organism Arabidopsis thaliana, seven ZIP family genes ZIP1–4 and IRT1–3 were characterized as functional Zn transporters with different affinities [6, 38–40]. ZIP1–3 genes were able to complement the Zn uptake in yeast and they were also overexpressed in Zn-deficient roots, while ZIP4 was expressed in roots and shoots of the Zn-deficient plants and also referred a role in intracellular Zn transport [6]. Although IRT1 is a high affinity iron transporter in roots, its overexpression was also able to accumulate the high levels of Zn and Cd in Fe-deficient Arabidopsis plants [41, 42]. IRT2 was reported to be expressed under Fe deficiency in Arabidopsis as well as could transport the Zn and Fe in yeast [42]. A. thaliana and Arabidopsis halleri IRT3 gene functionally complemented the Zn and Fe uptake in yeast, suggesting a Zn/Fe transport activity [39]. Moreover, Zn transporter homologs have been also reported in many plant species [43–47] but most are still waiting for their experimental characterization. Furthermore, taking into the account that reported Zn transporters have broad range affinity as well as have cross-talks with other metals, therefore it seems more challenging to propose a single specialized Zn transporter.
Boron (B)
Another important essential element involving in the healthy plant growth and development is boron [48]. It involves in many physiological processes, including the ammonium and nitrogen assimilation [49, 50], cytoskeleton polymerization [51], cellular signaling [52], regulation of the membrane potential and permeability [53, 54], control of the cell wall porosity and tensile strength [55, 56], and changes of the phenolic compounds [57]. B-deficiency forms one of the most widespread micronutrient deficiencies in the world and causes the significant losses in crop yield and quality [48]. It affects the vegetative and reproductive growth in plants; inhibits the cell expansion, reduces the plant fertility, and causes the meristem death [5]. In soil solution, B is present as boric acid (B(OH)3) or borate but boric acid is the most accepted form by plants since it is abundant in soil at optimum pH (5.5–7.5). Based on the B availability, three different mechanisms have been proposed for boric acid uptake; (i) passive diffusion by plasma membrane, (ii) facilitated transport by MIP proteins, and (iii) active transport by BOR transporters [58]. In case of B availability, boric acid is taken up via passive diffusion and facilitated transport, and under B deficiency, BOR transporters mainly involve in the boron uptake [59]. In model organism A. thaliana, B uptake was achieved by two transport molecules NIP5;1 and BOR1 [60, 61]. NIP5;1 is a boric acid channel protein from major intrinsic protein (MIP) family. It is localized to epidermal cell and root cap plasma membranes with a soil-facing polarity and significantly upregulated under B-deficient conditions [60, 61]. This family (MIP/aquaporin) members are characterized with six putative TMDs and an NPA (Asn-Pro-Ala/Ser/Val) motif [62]. BOR1 transporter was first characterized in B-deficient Arabidopsis roots [60]. Arabidopsis BOR1 gene encoded a protein of 704 amino acid residues including 10 putative TMDs. Besides, Arabidopsis AtBOR2–7 sequences also demonstrated the similarity to AtBOR1 [60]. Additionally, under B-deficient conditions, rice NIP3;1 channels and BOR1 transporters participated in B uptake [63, 64]. They also showed high homology to Arabidopsis NIP5;1 boric acid channels and BOR1 transporters, respectively [65]. Grapevine VvBOR1 gene, which is an AtBOR1 ortholog, encoded a polypeptide of 720 amino acid residues in Arabidopsis root pericycle cells [66]. In addition, functional BOR1 transporter homologs were also reported in some other species, including baker’s yeast [60], Brassica napus [67], and Eucalyptus [68]. Moreover, maize mutants tls1 and rte encoded the NIP3;1 and boron transporter proteins, homologous to AtNIP5;1 and AtBOR1, respectively, with some vegetative and reproductive defects like B deficiency [69–71]. Furthermore, boron transporter genes were reported to have great importance from agricultural perspectives to alleviate the B-deficiency symptoms and enhance the B tolerance.
Copper (Cu)
Cu stands another crucial element in plant micronutrient list [72]. It participates in a number of metabolic processes such as mitochondrial respiration, hormone signaling, photosynthetic electron transport, cell wall metabolism, and superoxide scavenging. It also functions as structural component in variety of enzymes, including cytochrome c oxidase, laccase, amino oxidase, plastocyanin and polyphenol oxidases, and Cu/Zn superoxide dismutase [5, 73]. Cu deficiency mainly affects the reproductive organs and younger leaves while its toxicity causes the necrosis, chlorosis, leaf discoloration and stunting, inhibits the root growth, and incites the ROS generation [5, 72]. Thus, plants could have acquired a sophisticated Cu network to maintain its uptake, mobilization, utilization, and storage [74]. In model species A. thaliana, Cu is transported into the cytosol by six high affinity transporters COPT1–6 while it is effluxed by P-type ATPases such as HMA1, HMA5, PAA1, PAA2, and RAN1. In addition, its intracellular distribution occurs with the help of metallochaperones such as CCS1, ATX1, and CCH [75–78]. COPT transporters are the members of CTR protein family with three putative TMDs and they include an extracellular N-terminal and a cytosolic C-terminal region [79–81]. With the exception of AtCOPT4, all Arabidopsis COPT proteins contain the His-rich residues and variable number of Met-rich motifs in their N-terminal regions. Yeast studies have demonstrated that Met-rich motifs in CTR family involve in Cu translocation into the cytosol by selectively sequestering copper ions from environment and stabilizing them with thioester groups in Met residues [80, 82, 83]. Green algae CTR1 and 2 transporters contained the six Met- and Cys-rich copper binding motifs [84]. In Arabidopsis, a Met residue before TMD1 (around 20 amino acids) and an Mx3M motif in TMD2 were essential in copper transport [85]. In yeast ctr1 mutants, a conserved Gx3G motif in TMD3 functioned as glycine zipper in helix packing and trimer assembly, and involved in delivery of transporters to their targets [86]. Mx3Mx12Gx3G motif was also highly conserved in CTR proteins [85]. In addition, many CTR sequences contained the Cys-rich (CxC) motif in their C-terminal regions. Yeast CTR1 studies demonstrated that this motifs (CxC) could involve in regulation of intracellular copper concentrations as well as could function in copper delivery to cytosolic metallochaperones [87]. Moreover, transcriptional activation of copper deficiency responsive genes, including COPT1 and 2 in Arabidopsis, is regulated by SPL7 transcription factor. This transcription factor also modulates a group of Cu-microRNAs such as miR397, 398, 408, and 857, causing the degradation of messenger RNAs (mRNAs) encoding the non-essential cuproproteins [88, 89]. All these mechanisms suggest the dynamic regulation of intracellular copper concentrations in response to plant physiological demand.
Molybdenum (Mo)
In essential plant micronutrient list, Mo is another important element with crucial metabolic functions [1]. It is used in synthesis of molybdenum cofactor (Moco), which forms the active sites of molybdoenzymes such as sulfite oxidase (SOX), aldehyde oxidase (AO), xanthine dehydrogenase (XDH), and nitrate reductase (NR) [90]. These enzymes participate in many substantial metabolic processes, including the phytohormone biosynthesis, purine metabolism, sulfite detoxification, and nitrate assimilation [91]. Thus, its deficiency or toxicity hinders the plant growth and development, and decreases the agricultural productivity [92]. Mo is present in soil at various forms but only molybdate (MoO4 2−), a dissolved form of the molybdenum, is taken up by the plants [93]. In model plant A. thaliana, MOT1 (previously SULTR5;2) is a high-affinity molybdate transporter in plasma membrane or endomembranes [94, 95]. Another Arabidopsis protein reported was the MOT2 in vacuoles; its accumulation in leaves and decrease in seed as well as the increased MOT2 activity in senescing leaves in mot2-deficient plants suggested that MOT2 may export the molybdate from vacuole into the cytosol [96]. In addition, MOT1 and 2 homologs have been also identified as in silico in a number of plant species [97]. Previously, MOT1 and 2 proteins were included into the sulfate transporters under SLC26A/SulP transporter family but they were separated from them with the absence of a C-terminal STAS (sulfate transporter and anti-sigma factor antagonist) domain [98]. SLC26A/SulP family members were reported to have 10–12 putative TMDs with STAS domain [99]. However, recently, Mo transporters were included into the molybdate transporter family (Pfam: PF16983) [100]. Molybdate (MoO4 2−) and sulfate (SO4 2−) show high-degree similarity due to their double negative charges and tetrahedral structures. Thus, sulfate transporters are reported to facilitate the molybdate uptake and distribution [96]. Green algae CrMOT1 included nine putative TMDs, and residues PXPVQPMKX(I/L)(A/G)AXA between TMD1 and 2, and residues FGXMPXCHG(S/A)GGLAXQ(Y/H)XFG(A/G)RXG between TMD6 and 7 showed the significant conservation in plants, algae, fungi, and bacteria [101]. Overall, further molecular and physiological studies are required to more elaborately elucidate the molybdenum transporters from the sulfate transporters.
Silicon (Si)
Among the plant micronutrients, Si is usually regarded as beneficial element [102]. It improves the canopy photosynthesis, reduces the transpiration loss, and increases the plant resistance against various biotic/abiotic stresses such as cold, drought, salinity, temperature, and various bacterial and fungal diseases [102–105]. However, beneficial effects of the silicon are related with the accumulation capacity of the plants; effects are clearer in high Si accumulators while less in low accumulating plants [106]. Silicon is taken up and transported to the shoots as silicic acid (Si(OH)4) and there it is converted to hydrated amorphous silica and stored on cell the walls, which forms the silica cuticle and silica cellulose double layers on stem, leaf and hull surfaces [104, 107]. Si transporter Lsi1 (NIP2;1) was first characterized in the root plasma membrane of the rice [108]. This transporter is a member of nodulin 26-like intrinsic proteins (NIPs) from plant aquaporins, which carry the water and small uncharged solutes such as silicic and boric acids, glycerol, and ammonia [109]. Substrate preference in NIP members were associated with two pore forming sites; (i) two conserved NPA (Asn-Pro-Ala) motifs and (ii) an aromatic/arginine (ar/R) region [110]. Adjacent Asn residues in two NPA motifs constitute the conserved NPA site. Besides, these residues are reported to make H bond with the molecules to be transported and function as proton exclusion [111, 112]. ar/R region is composed of four residues from helix 2, helix 5, loops E1 and E2, and forms a selectivity filter for substrate molecules [113, 114]. It plays a barrier role, regulates the transport rate, and makes van der Waals and H bonds [115, 116]. In Si transport, only Gly-Ser-Gly-Arg (GSGR) residues are reported to form the correct specificity filter with high conservancy in all known silicon transporters [114]. Two silicon transporters, Lsi1 and Lsi2 were reported to involve in efficient silicon uptake in rice; knockout of either transporter resulted in the decreased silicon uptake [108, 117, 118]. Besides, these transporter (Lsi1 and 2) homologs were also identified in some silicon accumulators including maize, barley, and pumpkin [119–122]. Moreover, plants absent with Lsi1 or Lsi2 homologs were either low accumulators or inefficiently accumulated the silicon [108, 117]. Although many studies so far have reported the beneficial effects of silicon in plant stress tolerance; however, its accumulation by different plants still waits to be elucidated. Besides, up to date, silicon transporters were mainly characterized in monocots therefore we have limited knowledge about many dicot species.
Manganese (Mn)
Mn is considered as an essential element in the list of major plant micronutrients [8, 123]. It functions in biosynthesis of the lipids, lignins, and carbohydrates, and involves in photosystem II (PSII) as well as it plays a structural role as cofactor in many enzymes. Its deficiency or toxicity hinders the plant growth, reduce the biomass, and causes the tissue necrosis, and interveinal chlorosis [8, 124–126]. Several transporter families, with broad range substrate affinities have been reported in influx and efflux of Mn. The NRAMP, ZIP, and YSL families were in Mn influx, and cation exchanger (CAX), cation calcium exchanger (CCX), P-type ATPases, and vacuolar iron transporter (VIT), and cation diffusion facilitator/metal tolerance protein (CDF/MTP) families were in Mn efflux [126]. Many studies have also reported the crucial roles of NRAMP, ZIP, and YSL families in Mn uptake. In NRAMP family, NRAMP1 is a high-affinity Mn transporter in Arabidopsis root plasma membrane [127, 128]. Arabidopsis NRAMP3 and 4 are reported to complement the Mn/Fe deficiency in yeast [129, 130]. NRAMP3 and 4 proteins are shown to involve in the Mn/Fe remobilization from vacuole [129]. Rice NRAMP3 differentially transports the Mn depending on the environmental conditions [131] while rice NRAMP5 is an essential Mn root uptake transporter from the soil [132]. In the ZIP family, Arabidopsis high-affinity iron transporter IRT1 was also to have low affinity for Mn [133]. Besides, Arabidopsis ZIP1, 2, 5–7, and 9 were reported to restore the Mn uptake in yeast mutants [134]. In YSL family, rice YSL2 and 6, and Arabidopsis YSL4 and 6 were implicated in Mn homeostasis [135–137]. Moreover, Ca2+-permeable channels were also reported to involve in the Mn transport in Arabidopsis and maize roots [138, 139]. NRAMP genes encode the highly hydrophobic membrane proteins with 10–12 putative TMDs possessing a consensus transport residue between TMD8 and 9 [29, 32]. Arabidopsis NRMAP1 and 2 transporters, respectively, are 532 and 530 residue proteins with 12 putative TMDs and a consensus transport residue between TMD8 and 9 [140]. Despite the reports of several transporter families in Mn uptake in some plants, we still have limited knowledge in many other plant species.
Nickel (Ni)
Ni has been proposed as essential plant micronutrient. It involves in the structure of many enzymes, including glyoxalases (family I), ureases, methyl-CoM reductase, superoxide dismutases, peptide deformylases, and some hydrogenases [141–143]. It plays crucial roles in ureolysis, methane biogenesis, acitogenesis, and hydrogen metabolism as well as in maintaining the cellular redox state, stress tolerance/defense, and optimum nitrogen use efficiency (NUE) [143–148]. Its deficiency was reported to cause the accumulation of urea and necrotic lesions in plant leaves [143], while toxicity reduced the plant growth and photosynthesis, induced the oxidative stress, inhibited the nitrogen metabolism, and enzymatic and mitotic activities, and interfered with other metals uptake [149–152]. Thus, nickel seems to play substantial role/s in plant growth and development. Nickel uptake could be achieved by active transport or passive diffusion depending on the plant species, soil pH, nickel form and concentration, and availability of other metals [153–155]. Solubilized nickel compounds could be transported by various cation transport systems such as Fe2+, Mg2+, Cu2+, and Zn2+, by chelators such as citric acid, histidine (His), and nicotianamine (NA) as well as with various other proteins such as permeases, metallothionein (MT), YS1-like (YSLs) and metallochaperones [151–156].
Chlorine (Cl−)
Cl, as being an essential micronutrient, involves in regulation of the pH and turgor pressure, and cytoplasmic enzyme activities, helps to stabilize the membrane potential, and plays a cofactor role in photosynthesis [5, 157–160]. Its deficiency causes the reduced leaf growth and wilting resulted with chlorosis and necrosis, and causes the stunted roots and reduced fruit size. However, its deficiency rarely occurs in normal conditions because the chlorine concentrations in soil are usually high and plants only require the trace amounts [160]. Besides, higher concentrations are reported to be toxic to plants, particularly many economically valuable cereals, vegetables, and fruit crops are susceptible to chlorine toxicity (4–7 mg/g/dw for sensitive species and 15–50 mg/g/dw for tolerant plants) [159]. In plants, several transporter families have been reported to participate in anion transport thereby in chlorine transport. These anion transporters include the slow anion channel associated protein (SLAC1), aluminum activated malate channels (ALMT), anion/H+ antiporters, ATP-binding cassette (ABC) transporter family, CLC anion channels, cation-coupled Cl− (CCC), voltage-dependent anion channels (VDAC), NRT (NOD)s nitrate and peptide transporter family, and MscS-like mechanosensitive channels [161]. Moreover, some other channels such as stretch-activated anion channels, slowly activated anion channels (S-type) and rapidly activated anion channels (R-type) were also shown to involve in chlorine ion efflux in plasma membranes of guard cells [160]. Furthermore, chlorine homeostasis in plants was also reported to be closely related with the salt tolerance [162–166].
Selenium (Se)
Se is regarded as a beneficial element for plants but non-essential. It involves in antioxidant and ROS regulation, heavy metal uptake and transport inhibition, photosynthetic system improvement, and construction of chloroplast components and cell membrane [167]. Besides, it also helps plants to alleviate the abiotic stress factors such as cold [168], drought [169], temperature [170], desiccation [171], salinity [169], heavy metals [172], water [173], senescence [174], and UV [175]. Its low concentrations improve the plant growth and development, and alleviate the stress factors, while higher concentrations could be toxic to plants [167]. For example, 1 mg/kg H2SeO4 addition to soil showed the toxic effect in ryegrass [174]. However, application of 1 mg/L Se with foliar or fruit spray was reported to improve the fruit quality in pears and peaches [176]. Besides, Se levels (up to 5 mg/L) were beneficial for some Se accumulators such as Spirulina platensis, red seaweed, and Pteris vittata [172, 177, 178]. Thus, plants demonstrate the variations for its accumulation and effects. Its concentration was reported to usually vary of 0.01–2.0 mg/kg in most soils [179]. Se is mainly present as selenate in alkaline and well-oxidized soils, as selenite in well-drained mineral soils, and as selenide in strongly reduced soils [180]. Plants could transport the selenate by sulfate transporters since both (sulfate and selenate) demonstrate the chemical similarity [181]. Sulfate selectivity over selenate is low under high sulfate availability. In addition, inducible sulfate transporters have higher selectivity than constitutive sulfate transporters for sulfate over selenate [182]. So, it is of great importance to characterize the sulfate transporter homologs particularly in Se hyperaccumulators [183]. Besides, plant selenite uptake is less known. Some studies suggested that its uptake is not metabolically associated [184]. Additionally, phosphate and carbonyl cyanide m-chlorophenyl hydrazone (CCCP) are reported to hinder the selenite uptake in wheat while P-deficiency improves it [185].
Sodium (Na)
Na significance is usually associated with the soil salinity (mainly with NaCl) because excessive Na+ concentrations are toxic to plants [186]. In woody plants citrus and grapevine, Na+ accumulated in roots and stems, and Cl− in shoots are harmful to plants [187]. In many higher plants, Na+ was also reported to cause an ion-specific damage [188]. However, some C4 plants and dicotyledonous halophytes were also reported to show considerable growth in response to high Na+ treatments [189]. Some specific/non-specific channels and transporters such as voltage-independent channels (VICs), nonselective cation channels (NSCCs), low-affinity cation transporter 1 (LCT1), KUP/HAK/KT, HKT1, HKT2, AKT1, and CCCs were suggested to involve in Na+ uptake in plants [186, 190]. Electrophysiological studies demonstrated that Na+ flows through NSCC/VIC channels in the root cortical cell plasma membranes [191–193]. It was therefore suggested that salt stress in plants can be alleviated by manipulation of NSCCs channels [193]. Despite the implication of LCT1 in Na+ influx, it was mainly reported to involve in Ca+ acquisition and Cd2+ toxicity recovery [194]. HKT1 via K+-independent way and HKT2 via K+-dependent way were involved in Na+ transport [195]. KUP/HAK/KT and AKT1 transporters involved in low and high affinity K+ uptake, and were relatedly sensitive to Na+ [191, 196, 197]. Cation-Cl–cotransporters (CCCs) participated in Na+, K+, and Cl− ions uptake in plants [198]. In light of these studies, a number of specific/non-specific transporters and channels appeared to involve in Na+ uptake in plants.
Aluminum (Al)
No experimental studies have been available so far showing the essentiality of Al in plants [199]. But its toxicity was demonstrated to be a major problem for many crops, particularly those growing in the acidic soils [200–202]. However, Al also stimulated the plant growth in Al accumulators [203, 204]. For example, tea plants responded by growth stimulation to the Al accumulation in shoots [205, 206]. Several reports have been proposed about the Al-induced growth mechanisms in plants. For example, Al-associated growth stimulation in tea plants could have derived from the improvement of latent iron toxicity [207]. In a different study, amelioration of proton toxicity was regarded as reason of Al-induced root elongation in proton-sensitive plants [208, 209]. In some other studies, improved antioxidant defense system was attributed as the reason of Al-related growth simulation in plants [199, 204, 210]. Thus, further molecular and physiological studies are required with more Al accumulator plants to demonstrate whether Al has any beneficial effects in plants.
Cobalt (Co)
Co is not regarded as essential element but reported as beneficial for plants. Its plant concentrations have importance because of its essentiality in animal nutrition; it is a vitamin B12 component required by all animals [211]. Normal Co concentrations in plants were as low as 0.1–10 μg/g/dw [212]. Thus, its higher concentrations were toxic to plants, severely interfering with the metabolic functions [213–215]. It hinders the plant growth, photosynthesis, and seed germination [216] as well as could cause the neurotoxicity in animals and memory deficit in humans [217]. Co transport in plants is physiologically regulated at a species-specific way. Co as divalent cation (Co2+) could be transported into the cells by various broad range transporters thereby its homeostasis in plants are regulated by different metabolic pathways [211]. Besides, it is reported to be distributed in plant body via organic complexes due to its low mobility [212, 218, 219]. However, its precise transport mechanism from soil to root and to plant organs still awaits for further elucidation. In addition, soil properties significantly affect the Co bioavailability by plants [220, 221] therefore eliminating soil properties in Co studies could lead to the incorrect reports of toxicity thresholds [222]. So, cobalt being a heavy metal as well as a micronutrient, its elaborative control is required by plants for normal plant cycle.
Conclusion
Present study has extensively reviewed the 13 mineral elements (Fe, B, Cu, Mn, Mo, Si, Zn, Ni, Cl, Se, Na, Al, and Co) that are required by plants at trace amounts, with an emphasis on their transport proteins (transporters/channels) in plant roots. Although the essentiality of some micronutrients have been well established by many studies while some others have not been strictly proposed yet. Thus, it is still challenging to mention an exact number of the micronutrients in plants. Further molecular and physiological studies are needed to elucidate the roles of some elements in plant nutrition whether as essential or beneficial. In addition, transporter or channel proteins involved in the micronutrient transport usually demonstrate the broad range affinity for various other metals. Therefore, the cellular element uptake and its metabolism in plants are physiologically regulated depending on the plants demand, availability of other elements, physiological state of the plants, and other environmental conditions.
References
Barker, A. V., & Pilbeam, D. J. (Eds.). (2015). Handbook of plant nutrition. CRC press.
Mengel, K., Kosegarten, H., Kirkby, E. A., & Appel, T. (Eds.) (2001). Principles of plant nutrition. Springer Science and Business, Media.
Maathuis, F. J. M. (2013). Plant mineral nutrients: methods and protocols. New York: Humana Press.
Maathuis, F. J., & Diatloff, E. (2013). Roles and functions of plant mineral nutrients. Plant Mineral Nutrients: Methods and Protocols, 1–21.
Marschner, H. (1995). Functions of mineral nutrients: macronutrients. Mineral nutrition of higher plants, 2, Academic Press, London, 379–396.
Grotz, N., Fox, T., Connolly, E., Park, W., Guerinot, M. L., & Eide, D. (1998). Identification of a family of zinc transporter genes from Arabidopsis that respond to zinc deficiency. Proceedings of the National Academy of Sciences, 95, 7220–7224.
Underwood, E. (2012). Trace elements in human and animal nutrition 4e. Elsevier.
Marschner, P. (2012). Marschner’s mineral nutrition of higher plants. Boston, MA: Academic Press.
Tavsan, Z., & Kayali, H. A. (2013). The effect of iron and copper as an essential nutrient on mitochondrial electron transport system and lipid peroxidation in Trichoderma harzianum. Applied Biochemistry and Biotechnology, 170(7), 1665–1675.
Vatansever, R., Filiz, E., & Ozyigit, I. I. (2015). Genome-wide analysis of iron-regulated transporter 1 (IRT1) genes in plants. Horticulture, Environment, and Biotechnology, 56(4), 516–523.
Vert, G., Grotz, N., Dédaldéchamp, F., Gaymard, F., Guerinot, M. L., Briat, J. F., et al. (2002). IRT1, an Arabidopsis transporter essential for iron uptake from the soil and for plant growth. Plant Cell Online, 14, 1223–1233.
Kobayashi, T., & Nishizawa, N. K. (2012). Iron uptake, translocation, and regulation in higher plants. Annual Review of Plant Biology, 63, 131–152.
Tombuloğlu, H., Ablazov, A., & Filiz, E. (2016). Genome-wide analysis of response to low sulfur (LSU) genes in grass species and expression profiling of model grass species Brachypodium distachyon under S-deficiency. Turkish Journal of Biology. doi:10.3906/biy-1508-32.
López-Millán, A. F., Grusak, M. A., Abadía Bayona, A., & Abadía Bayona, J. (2013). Iron deficiency in plants: an insight from proteomic approaches. Frontiers in Plant Science, 4, 254.
Kim, S. A., & Guerinot, M. L. (2007). Mining iron: iron uptake and transport in plants. FEBS Letters, 581, 2273–2280.
Briat, J. F., & Lobr’eaux, S. (1997). Iron transport and storage in plants. Trends in Plant Sciences, 2, 187–193.
Briat, J. F., Dubos, C., & Gaymard, F. (2015). Iron nutrition, biomass production, and plant product quality. Trends in Plant Science, 20(1), 33–40.
Römheld, V., & Marschner, H. (1986). Evidence for a specific uptake system for iron phytosiderophore in roots of grasses. Plant Physiology, 80, 175–180.
Higuchi, K., Suzuki, K., Nakanishi, H., Yamaguchi, H., Nishizawa, N. K., & Mori, S. (1999). Cloning of nicotianamine synthase genes, novel genes involved in the biosynthesis of phytosiderophores. Plant Physiology, 119, 471–479.
Takahashi, M., Yamaguchi, H., Nakanishi, H., Shioiri, T., Nishizawa, N. K., & Mori, S. (1999). Cloning two genes for nicotianamine aminotransferase, a critical enzyme in iron acquisition (strategy II) in graminaceous plants. Plant Physiology, 121, 947–956.
Bashir, K., Inoue, H., Nagasaka, S., Takahashi, M., Nakanishi, H., Mori, S., et al. (2006). Cloning and characterization of deoxymugineic acid synthase genes from graminaceous plants. Journal of Biological Chemistry, 43, 32395–32402.
Inoue, H., Kobayashi, T., Nozoye, T., Takahashi, M., Kakei, Y., Suzuki, K., et al. (2009). Rice OsYSL15 is an iron-regulated iron (III)-deoxymugineic acid transporter expressed in the roots and is essential for iron uptake in early growth of the seedlings. Journal of Biological Chemistry, 284, 3470–3479.
Lee, S., Chiecko, J. C., Kim, S. A., Walker, E. L., Lee, Y., Guerinot, M. L., et al. (2009). Disruption of OsYSL15 leads to iron inefficiency in rice plants. Plant Physiology, 150, 786–800.
Nozoye, T., Nagasaka, S., Kobayashi, T., Takahashi, M., Sato, Y., Sato, M., et al. (2011). Phytosiderophore efflux transporters are crucial for iron acquisition in graminaceous plants. Journal of Biological Chemistry, 286, 5446–5454.
Thomine, S., & Vert, G. (2013). Iron transport in plants: better be safe than sorry. Current Opinion in Plant Biology, 16(3), 322–327.
Colangelo, E. P., & Guerinot, M. L. (2006). Put the metal to the petal: metal uptake and transport throughout plants. Current Opinion in Plant Biology, 9, 322–330.
Guerinot, M. L. (2000). The ZIP family of metal transporters. Biochimica et Biophysica Acta (BBA), 1465, 190–198.
Eng, B. H., Guerinot, M. L., Eide, D., & Saier Jr., M. H. (1998). Sequence analyses and phylogenetic characterization of the ZIP family of metal ion transport proteins. Journal of Membrane Biology, 166, 1–7.
Thomine, S., Wang, R., Ward, J. M., Crawford, N. M., & Schroeder, J. I. (2000). Cadmium and iron transport by members of a plant metal transporter family in Arabidopsis with homology to Nramp genes. Proceedings of the National Academy of Sciences, USA, 97, 4991–4996.
Nevo, Y., & Nelson, N. (2006). The NRAMP family of metal-iontransporters. Biochimica et Biophysica Acta (BBA), 1763, 609–620.
Xia, J., Yamaji, N., Kasai, T., & Ma, J. F. (2010). Plasma membrane-localized trans-Porter for aluminum in rice. Proceedings of the National Academy of Sciences, USA, 107, 18381–18385.
Cellier, M., Prive, G., Belouchi, A., Kwan, T., Rodrigues, V., Chia, W., et al. (1995). Nramp defines a family of membrane proteins. Proceedings of the National Academy of Sciences, USA, 92, 10089–10093.
Schaaf, G., Ludewig, U., Erenoglu, B. E., Mori, S., Kitahara, T., & Von Wiren, N. (2004). ZmYS1 functions as a proton-coupled symporter for phytosiderophore- and nicotianamine-chelated metals. Journal of Biological Chemistry, 279, 9091–9096.
Yen, M. R., Tseng, Y. H., & Saier Jr., M. H. (2001). Maize yellow Stripe1, an iron-phytosiderophore uptake transporter, is a member of the oligopeptide transporter (OPT) family. Microbiology, 147, 2881–2883.
Höreth, S., Detterbeck, A., Ahmadi, H., Pongrac, P., & Clemens, S. (2015). Molecular analysis of within-plant Zn mobility in a metal hyperaccumulator and a crop model system. In OF ABSTRACTS (p. 22).
Fox, T. C., & Guerinot, M. L. (1998). Molecular biology of cation transport in plants. Annual Review of Plant Biology, 49, 669–696.
Sadeghzadeh, B., & Rengel, Z. (2011). Zinc in soils and crop nutrition. The molecular and physiological basis of nutrient use efficiency in crops, John Wiley & Sons, Inc., pp. 335–375.
Maser, P., Thomine, S., Schroeder, J. I., Ward, J. M., Hirschi, K., Sze, H., et al. (2001). Phylogenetic relationships within cation transporter families of Arabidopsis. Plant Physiology, 126, 1646–1667.
Lin, Y. F., Liang, H. M., Yang, S. Y., Boch, A., Clemens, S., Chen, C. C., et al. (2009). Arabidopsis IRT3 is a zinc-regulated and plasma membrane localized zinc/iron transporter. New Phytologist, 182, 392–404.
Assunção, A. G., Herrero, E., Lin, Y. F., Huettel, B., Talukdar, S., Smaczniak, C., Immink, R. H., Eldik, Mv., Fiers, M., Schat, H. et al. (2010). Arabidopsis thaliana transcription factors bZIP19 and bZIP23 regulate the adaptation to zinc deficiency. Proceedings of the National Academy of Sciences of the United States of America, 10296–10301.
Connolly, E. L., Fett, J. P., & Guerinot, M. L. (2002). Expression of the irt1 metal transporter is controlled by metals at the levels of transcript and protein accumulation. Plant Cell, 14, 1347–1357.
Vert, G., Briat, J. F., & Curie, C. (2001). Arabidopsis irt2 gene encodes a root-periphery iron transporter. Plant Journal, 26, 181–189.
Moreau, S., Thomson, R. M., Kaiser, B. N., Trevaskis, B., Guerinot, M. L., Udvardi, M. K., et al. (2002). GmZIP1 encodes a symbiosis-specific zinc transporter in soybean. Journal of Biological Chemistry, 15, 4738–4746.
Ramesh, S. A., Shin, R., Eide, D. J., & Schachtman, D. P. (2003). Defferential metal selectivity and gene expression of two zinc transporters from rice. Plant Physiology, 133, 126–134.
Ishimaru, Y., Suzuki, M., Kobayashi, T., Takahashi, M., Nakanishi, H., Mori, S., et al. (2005). OsZIP4, a novel zinc-regulated zinc transporter in rice. Journal of Experimental Botany, 56, 3207–3214.
Lopez-Millan, A. F., Ellis, D. R., & Grusak, M. A. (2004). Identification and characterization of several new members of the zip family of metal ion transporters in Medicago truncatula. Plant Mol Biology, 54, 583–596.
Vatansever, R., Ozyigit, I. I., & Filiz, E. (2015). Comparative and phylogenetic analysis of zinc (Zn) transporter genes/proteins in plants. Turkish Journal of Biology. doi:10.3906/biy-1501-91.
Coudray, N., Zhang, Z., Clark, K. M., Ubarretxena, I., Beckstein, O., Dumont, M. E., et al. (2016). Structure of the borate transporter Bor1p by cryo-EM. Biophysical Journal, 110(3), 137–138.
Shen, Z. G., Liang, Y. C., & Shen, K. (1993). Effect of boron on the nitrate reductase-activity in oilseed rape plants. Journal of Plant Nutrition, 16, 1229–1239.
Camacho-Cristóbal, J. J., & González-Fontes, A. (2007). Boron deficiency decreases plasmalemma H+−ATPase expression and nitrate uptake, and promotes ammonium assimilation into asparagine in tobacco roots. Planta, 226, 443–451.
Bassil, E., Hu, H., & Brown, P. H. (2004). Use of phenylboronic acids to investigate boron function in plants. Possible role of boron in transvacuolar cytoplasmic strands and cell-to-wall adhesion. Plant Physiology, 136, 3383–3395.
Gonzalez-Fontes, A., Rexach, J., Navarro-Gochicoa, M. T., Herrera-Rodrıguez, M. B., Beato, V. M., Maldonado, J. M., et al. (2008). Is boron involved solely in structural roles in vascular plants? Plant Signaling and Behavior, 3, 24–26.
Ferrol, N., & Donaire, J. P. (1992). Effect of boron on plasma membrane proton extrusion and redox activity in sunflower cells. Plant Science, 86, 41–47.
Wang, Z. Y., Tang, Y. L., Zhang, F. S., & Wang, H. (1999). Effect of boron and low temperature on membrane integrity of cucumber leaves. Journal of Plant Nutrition, 22, 543–550.
Fleischer, A., O’Neill, M. A., & Ehwald, R. (1999). The pore size of nongraminaceous plant cell walls is rapidly decreased by borate ester cross-linking of the pectic polysaccharide rhamnogalacturonan II. Plant Physiology, 121, 829–838.
Ryden, P., Sugimoto-Shirasu, K., Smith, A. C., Findlay, K., Reiter, W. D., & McCann, M. C. (2003). Tensile properties of Arabidopsis cell walls depend on both a xyloglucan cross-linked microfibrillar network and rhamnogalacturonan II-borate complexes. Plant Physiology, 132, 1033–1040.
Camacho-Cristóbal, J. J., Anzelotti, D., & González-Fontes, A. (2002). Changes in phenolic metabolism of tobacco plants during short-term boron deficiency. Plant Physiology and Biochemistry, 40, 997–1002.
Tanaka, M., & Fujiwara, T. (2008). Physiological roles and transport mechanisms of boron: perspectives from plants. European Journal of Physiology, 456, 671–677.
Dannel, F., Pfeffer, H., & R¨omheld, V. (2002). Update on boron in higher plant-uptake, primary translocation and compartmentation. Plant Biology, 4, 193–204.
Takano, J., Noguchi, K., Yasumori, M., Kobayashi, M., Gajdos, Z., Miwa, K., et al. (2002). Arabidopsis boron transporter for xylem loading. Nature, 420, 337–340.
Takano, J., Wada, M., Ludewig, U., Schaaf, G., von Wir ´en, N., & Fujiwara, T. (2006). The Arabidopsis major intrinsic protein NIP5;1 is essential for efficient boron uptake and plant development under boron limitation. Plant Cell, 18, 1498–1509.
Kruse, E., Uehlein, N., & Kaldenhoff, R. (2006). The aquaporins. Genome Biology, 7, 206.
Nakagawa, Y., Hanaoka, H., Kobayashi, M., Miyoshi, K., Miwa, K., & Fujiwara, T. (2007). Cell-type specificity of the expression of OsBOR1, a rice efflux boron transporter gene, is regulated in response to boron availability for efficient boron uptake and xylem loading. Plant Cell, 19, 2624–2635.
Hanaoka, H., Uraguchi, S., Takano, J., Tanaka, M., & Fujiwara, T. (2014). OsNIP3; 1, a rice boric acid channel, regulates boron distribution and is essential for growth under boron-deficient conditions. Plant Journal, 78, 890–902.
Takano, J., Miwa, K., & Fujiwara, T. (2008). Boron transport mechanisms: collaboration of channels and transporters. Trends in Plant Sciences, 13, 451–457.
Pérez-Castro, R., Kasai, K., Gainza-Cortés, F., Ruiz-Lara, S., Casaretto, J. A., Peña-Cortés, H., et al. (2012). VvBOR1, the grapevine ortholog of AtBOR1, encodes an efflux boron transporter that is differentially expressed throughout reproductive development of Vitis vinifera L. Plant and Cell Physiology, 53(2), 485–494.
Sun, J., Shi, L., Zhang, C., & Xu, F. (2012). Cloning and characterization of boron transporters in Brassica napus. Molecular Biology Reports, 39, 1963–1973.
Domingues, D. S., Maximino Leite, S. M., & Cazerta Farro, A. P. (2005). Boron transport in eucalyptus. 2. Identification in silico of a putative boron transporter for xylem loading in eucalypt. Genetics and Molecular Biology, 28, 625–629.
Chatterjee, M., Tabi, Z., Galli, M., Malcomber, S., Buck, A., Muszynski, M., et al. (2014). The boron efflux transporter ROTTEN EAR is required for maize inflorescence development and fertility. Plant Cell, 26, 2962–2977.
Durbak, A. R., Phillips, K. A., Pike, S., O’Neill, M. A., Mares, J., Gallavotti, A., et al. (2014). Transport of boron by the tassel-less1 aquaporin is critical for vegetative and reproductive development in maize. Plant Cell, 26, 2978–2995.
Leonard, A., Holloway, B., Guo, M., Rupe, M., Yu, G., Beatty, M., & Li, B. (2014). tassel-less1 encodes a boron channel protein required for inflorescence development in maize. Plant Cell Physiology, pcu036.
Adrees, M., Ali, S., Rizwan, M., Ibrahim, M., Abbas, F., Farid, M., et al. (2015). The effect of excess copper on growth and physiology of important food crops: a review. Environmental Science and Pollution Research, 22(11), 8148–8162.
Ng, I. S., Zheng, X., Wang, N., Chen, B. Y., Zhang, X., & Lu, Y. (2014). Copper response of Proteus hauseri based on proteomic and genetic expression and cell morphology analyses. Applied Biochemistry and Biotechnology, 173(5), 1057–1072.
Clemens, S. (2001). Molecular mechanisms of plant metal tolerance and homeostasis. Planta, 212, 475–486.
Huffman, D. L., & O’Halloran, T. V. (2001). Function, structure, and mechanism of intracellular copper trafficking proteins. Annual Review of Biochemistry, 70, 677–701.
Burkhead, J. L., Gogolin, R. K. A., Abdel-Ghany, S. E., Cohu, C. M., & Pilon, M. (2009). Copper homeostasis. New Phytologist, 182, 799–816.
Pilon, M., Cohu, C. M., Ravet, K., Abdel-Ghany, S. E., & Gaymard, F. (2009). Essential transition metal homeostasis in plants. Current Opinion in Plant Biology, 12, 347–357.
Puig, S., & Peñarrubia, L. (2009). Placing metal micronutrients in context: transport and distribution in plants. Current Opinion in Plant Biology, 12, 299–306.
Eisses, J. F., & Kaplan, J. H. (2002). Molecular characterization of hCTR1, the human copper uptake protein. Journal of Biological Chemistry, 277, 29162–29171.
Puig, S., Lee, J., Lau, M., & Thiele, D. J. (2002). Biochemical and genetic analyses of yeast and human high affinity copper transporters suggest a conserved mechanism for copper uptake. Journal of Biological Chemistry, 277, 26021–26030.
Vatansever, R., Ozyigit, I. I., & Filiz, E. (2016). Genome-wide identification and comparative analysis of copper transporter genes in plants. Interdisciplinary Sciences: Computational Life Sciences, 1–14.
Jiang, J., Nadas, I. A., Kim, M. A., & Franz, K. J. (2005). A Mets motif peptide found in copper transport proteins selectively binds Cu (I) with methionine-only coordination. Inorganic Chemistry, 44, 9787–9794.
Beaudoin, J., Laliberté, J., & Labbé, S. (2006). Functional dissection of Ctr4 and Ctr5 amino-terminal regions reveals motifs with redundant roles in copper transport. Microbiology, 152, 209–222.
Page, M. D., Kropat, J., Hamel, P. P., & Merchant, S. S. (2009). Two Chlamydomonas CTR copper transporters with a novel cys-met motif are localized to the plasma membrane and function in copper assimilation. Plant Cell Online, 21, 928–943.
Peñarrubia, L., Andrés-Colás, N., Moreno, J., & Puig, S. (2010). Regulation of copper transport in Arabidopsis thaliana: a biochemical oscillator? Journal of Biological Inorganic Chemistry, 15, 29–36.
Aller, S., Eng, G., De, E. T., Feo, C. J., & Unger, V. M. (2004). Eukaryotic CTR copper uptake transporters require two faces of the third transmembrane domain for helix packing, oligomerization, and function. Journal of Biological Chemistry, 279, 53435–53441.
Wu, X., Sinani, D., Kim, H., & Lee, J. (2009). Copper transport activity of yeast Ctr1 is down-regulated via its C terminus in response to excess copper. Journal of Biological Chemistry, 284, 4112–4122.
Abdel-Ghany, S. E., & Pilon, M. (2008). MicroRNA-mediated systemic down-regulation of copper protein expression in response to low copper availability in Arabidopsis. Journal of Biological Chemistry, 283, 15932–15945.
Yamasaki, H., Hayashi, M., Fukazawa, M., Kobayashi, Y., & Shikanai, T. (2009). SQUAMOSA promoter binding protein–like7 is a central regulator for copper homeostasis in Arabidopsis. Plant Cell Online, 21, 347–361.
Kruse, T., Gehl, C., Geisler, M., Lehrke, M., Ringel, P., Hallier, S., et al. (2010). Identification and biochemical characterization of molybdenum cofactor-binding proteins from Arabidopsis thaliana. Journal of Biological Chemistry, 285, 6623–6635.
Mendel, R. R., & Leimkühler, S. (2015). The biosynthesis of the molybdenum cofactors. JBIC Journal of Biological Inorganic Chemistry, 20(2), 337–347.
Kaiser, B. N., Gridley, K. L., Brady, J. N., Phillips, T., & Tyerman, S. D. (2005). The role of molybdenum in agricultural plant production. Annual Botany, 96, 745–754.
Ide, Y., Kusano, M., Oikawa, A., Fukushima, A., Tomatsu, H., Saito, K., et al. (2011). Effects of molybdenum deficiency and defects in molybdate transporter MOT1 on transcript accumulation and nitrogen/Sulphur metabolism in Arabidopsis thaliana. Journal of Experimental Botany, 62, 1483–1497.
Hawkesford, M. J. (2003). Transporter gene families in plants: the sulphate transporter gene family-redundancy or specialization? Physiologia Plantarum, 117, 155–163.
Baxter, I., Muthukumar, B., Park, H. C., Buchner, P., Lahner, B., Danku, J., & Salt, D. E. (2008). Variation in molybdenum content across broadly distributed populations of Arabidopsis thaliana is controlled by a mitochondrial molybdenum transporter (MOT1). Plos Genetics, 4.
Bittner, F. (2014). Molybdenum metabolism in plants and crosstalk to iron. Frontier in Plant Sciences, 5.
Vatansever, R., Filiz, E., & Ozyigit, I. I. (2015). In silico identification and comparative analysis of molybdenum (Mo) transporter genes in plants. Brazilian Journal of Botany, 1–13.
Shibagaki, N., & Grossman, A. R. (2006). The role of the STAS domain in the function and biogenesis of a sulfate transporter as probed by random mutagenesis. Journal of Biological Chemistry, 281(32), 22964–22973.
Compton, E. L., Karinou, E., Naismith, J. H., Gabel, F., & Javelle, A. (2011). Low resolution structure of a bacterial SLC26 transporter reveals dimeric stoichiometry and mobile intracellular domains. Journal of Biological Chemistry, 286, 27058–27067.
Finn, R. D. (2012). Pfam: the protein families database. Encyclopedia of Genetics, Genomics, Proteomics and Bioinformatics.
Tejada-Jiménez, M., Llamas, Á., Sanz-Luque, E., Galván, A., & Fernández, E. (2007). A high- affinity molybdate transporter in eukaryotes. Proceedings of the National Academy of Sciences, 104, 20126–20130.
Zhu, Y., & Gong, H. (2014). Beneficial effects of silicon on salt and drought tolerance in plants. Agronomy for Sustainable Development, 34(2), 455–472.
Rodrigues, F. A., Resende, R. S., Dallagnol, L. J., & Datnoff, L. E. (2015). Silicon potentiates host defense mechanisms against infection by plant pathogens. In Silicon and Plant Diseases (pp. 109–138). Springer International Publishing.
Mitani, N., Yamaji, N., & Ma, J. F. (2008). Characterization of substrate specificity of a rice silicon transporter, Lsi1. Pflügers Archiv-European Journal of Physiology, 456(4), 679–686.
Zhang, C., Wang, L., Nie, Q., Zhang, W., & Zhang, F. (2008). Long-term effects of exogenous silicon on cadmium translocation and toxicity in rice (Oryza sativa L.). Environmental and Experimental Botany, 62, 300–307.
Ma, J. F. (2004). Role of silicon in enhancing the resistance of plants to biotic and abiotic stresses. Soil Science and Plant Nutrition, 50, 11–18.
Ashraf, M. A., Morshed, M. M., Ahammad, A. S., & Morshed, M. N. (2013). Computational study of silicon transporter protein in rice and wheat. International Journal of Computational Bioinformatics and In Silico Model, 2(4), 199–205.
Ma, J. F., Tamai, K., Yamaji, N., Mitani, N., Konishi, S., Katsuhara, M., et al. (2006). A silicon transporter in rice. Nature, 440, 688–691.
Bienert, G., & Chaumont, F. (2011). Plant aquaporins: roles in water homeostasis, nutrition, and signaling processes. In M. Geisler & K. Venema (Eds.), Transporters and pumps in plant signaling processes (pp. 3–36). Berlin: Springer-Verlag.
Wu, B., & Beitz, E. (2007). Aquaporins with selectivity for unconventional permeants. Cellular and Molecular Life Sciences, 64, 2413–2421.
Ilan, B., Tajkhorshid, E., Schulten, K., & Voth, G. A. (2004). The mechanism of proton exclusion in aquaporin channels. Proteins, 55, 223–228.
Forrest, K. L., & Bhave, M. (2007). Major intrinsic proteins (MIPs) in plants: a complex gene family with major impacts on plant phenotype. Functional Integrative Genomics, 7, 263–289.
Harries, W. E., Akhavan, D., Miercke, L. J., Khademi, S., & Stroud, R. M. (2004). The channel architecture of aquaporin 0 at a 2.2 A° resolution. Proceedings of the National Academy of Sciences, USA, 101, 14045–14050.
Mitani-Ueno, N., Yamaji, N., Zhao, F. J., & Ma, J. F. (2011). The aromatic/arginine selectivity filter of NIP aquaporins plays a critical role in substrate selectivity for silicon, boron, and arsenic. Journal of Experimental Botany, 62(12), 4391–4398.
Fu, D., Libson, A., Miercke, L. J., Weitzman, C., Nollert, P., Krucinski, J., et al. (2000). Structure of a glycerol-conducting channel and the basis for its selectivity. Science, 290, 481–486.
Sui, H., Han, B. G., Lee, J. K., Walian, P., & Jap, B. K. (2001). Structural basis of water-specific transport through the AQP1 water channel. Nature, 414, 872–878.
Ma, J. F., Yamaji, N., Mitani, N., Kazunori, T., Konishi, S., Fujiwara, T., et al. (2007). An efflux transporter of silicon in rice. Nature, 448, 209–212.
Ma, J. F. (2010). Si transporters in higher plant. In P. T. Jhon & P. G. Bienert (Eds.), MIPs and their role in the exchange of materials (pp. 99–109). Texas: Landes Bioscience.
Chiba, Y., Mitani, N., Yamaji, N., & Ma, J. F. (2009). HvLsi1 is a silicon influx transporter in barley. Plant Journal, 57, 810–818.
Mitani, N., Chiba, Y., Yamaji, N., & Ma, J. F. (2009). Identification and characterization of maize and barley Lsi2-like silicon efflux transporters reveals a distinct silicon uptake system from that in rice. Plant Cell, 21, 2133–2142.
Mitani, N., Yamaji, N., & Ma, J. F. (2009). Identification of maize silicon influx transporters. Plant and Cell Physiology, 50, 5–12.
Mitani, N., Yamaji, N., Ago, Y., Iwasaki, K., & Ma, J. F. (2011). Isolation and functional characterization of an influx silicon transporter in two pumpkin cultivars contrasting in silicon accumulation. Plant Journal, 66, 231–240.
Fernando, D. R., & Lynch, J. P. (2015). Manganese phytotoxicity: new light on an old problem. Annals of Botany, 116(3), 313–319.
Yin, C., Zhao, W., Zheng, L., Chen, L., Tan, Q., Shang, X., et al. (2014). High-level expression of a manganese superoxide dismutase (PoMn-SOD) from Pleurotus ostreatus in Pichia pastoris. Applied Biochemistry and Biotechnology, 174(1), 259–269.
Nickelsen, J., & Rengstl, B. (2013). Photosystem II assembly: from cyanobacteria to plants. Annual Review of Plant Biology, 64, 609–635.
Socha, A. L., & Guerinot, M. L. (2014). Mn-euvering manganese: the role of transporter gene family members in manganese uptake and mobilization in plants. Frontiers in Plant Sciences, 5, 106.
Cailliatte, R., Schikora, A., Briat, J. F., Mari, S., & Curie, C. (2010). High- affinity manganese uptake by the metal transporter NRAMP1 is essential for Arabidopsis growth in low manganese conditions. Plant Cell, 22, 904–917.
Vatansever, R., Filiz, E., & Ozyigit, I. I. (2016). In silico analysis of Mn transporters (NRAMP1) in various plant species. Molecular Biology Reports. doi:10.1007/s11033-016-3950-x.
Thomine, S., Lelievre, F., Debarbieux, E., Schroeder, J. I., & Barbier-Brygoo, H. (2003). AtNRAMP3, a multi specific vacuolar metal transporter involved in plant responses to iron deficiency. Plant Journal, 34, 685–695.
Lanquar, V., Ramos, M. S., Lelievre, F., Barbier-Brygoo, H., Krieger-Liszkay, A., Kramer, U., et al. (2010). Export of vacuolar manganese by AtNRAMP3 and AtNRAMP4 is required for optimal photosynthesis and growth under manganese deficiency. Plant Physiology, 152, 1986–1999.
Yamaji, N., Sasaki, A., Xia, J. X., Yokosho, K., & Ma, J. F. (2013). Anode-based switch for preferential distribution of manganese in rice. Nature Communications, 4, 2442.
Sasaki, A., Yamaji, N., Yokosho, K., & Ma, J. F. (2012). Nramp5 is a major transporter responsible for manganese and cadmium uptake in rice. Plant Cell Online, 24, 2155–2167.
Yang, T. J., Perry, P. J., Ciani, S., Pandian, S., & Schmidt, W. (2008). Manganese deficiency alters the patterning and development of root hairs in Arabidopsis. Journal of Experimental Botany, 59, 3453–3464.
Milner, M. J., Seamon, J., Craft, E., & Kochian, L. V. (2013). Transport properties of members of the ZIP family in plants and their role in Zn and Mn homeostasis. Journal of Experimental Botany, 64, 369–381.
Koike, S., Inoue, H., Mizuno, D., Takahashi, M., Nakanishi, H., Mori, S., et al. (2004). OsYSL2 is a rice metal-nicotianamine transporter that is regulated by iron and expressed in the phloem. Plant Journal, 39, 415–424.
Sasaki, A., Yamaji, N., Xia, J., & Ma, J. F. (2011). OsYSL6 is involved in the detoxification of excess manganese in rice. Plant Physiology, 157, 1832–1840.
Conte, S. S., Chu, H. H., Rodriguez, D. C., Punshon, T., Vasques, K. A., Salt, D. E., et al. (2013). Arabidopsis thaliana yellow stripe1-like4 and yellowstripe1-like6 localize to internal cellular membranes and are involved in metal ion homeostasis. Frontiers in Plant. Sciences, 4, 283.
Marshall, J., Corzo, A., Leigh, R. A., & Sanders, D. (1994). Membrane potential dependent calcium transport in right-side-out plasma membrane vesicles from Zea mays L. Roots. Plant Journal, 5, 683–694.
Wymer, C. L., Bibikova, T. N., & Gilroy, S. (1997). Cytoplasmic free calcium distributions during the development of root hairs of Arabidopsis thaliana. Plant Journal, 12, 427–439.
Curie, C., Alonso, J., Le Jean, M., Ecker, J., & Briat, J. (2000). Involvement of NRAMP1 from Arabidopsis thaliana in iron transport. Biochemical Journal, 347, 749–755.
Fabiano, C. C., Tezotto, T., Favarin, J. L., Polacco, J. C., & Mazzafera, P. (2015). Essentiality of nickel in plants: a role in plant stresses. Frontiers in plant science, 6.
Küpper, H., & Kroneck, P. M. (2007). Nickel in the environment and its role in the metabolism of plants and cyanobacteria. Metal Ions in Life Sciences, 2, 31–62.
Polacco, J. C., Mazzafera, P., & Tezotto, T. (2013). Opinion–nickel and urease in plants: still many knowledge gaps. Plant Science, 199, 79–90.
Maier, T., Jacobi, A., Sauter, M., & Böck, A. (1993). The product of the hypB gene, which is required for nickel incorporation into hydrogenases, is a novel guanine nucleotide-binding protein. Journal of Bacteriology, 175(3), 630–635.
van der Lelie, D., Wilmotte, A., & Wuertz, S. (1994). Plasmids for heavy metal resistance in Alcaligenes eutrophus CH34: mechanisms and applications. FEMS Microbiology Reviews, 14, 405–414.
Ragsdale, S. W. (1998). Nickel biochemistry. Current Opinion in Chemical Biology, 2(2), 208–215.
Mulrooney, S. B., & Hausinger, R. P. (2003). Nickel uptake and utilization by microorganisms. FEMS Microbiology Reviews, 27(2–3), 239–261.
Yusuf, M., Fariduddin, Q., Hayat, S., & Ahmad, A. (2011). Nickel: an overview of uptake, essentiality and toxicity in plants. Bulletin of Environmental Contamination and Toxicology, 86(1), 1–17.
Rajendran, P., Ashokkumar, B., Muthukrishnan, J., & Gunasekaran, P. (2002). Toxicity assessment of nickel using Aspergillus niger and its removal from an industrial effluent. Applied Biochemistry and Biotechnology, 102(1–6), 201–206.
Molas, J. (2002). Changes of chloroplast ultrastructure and total chlorophyll concentration in cabbage leaves caused by excess of organic Ni II complexes. Environmental and Experimental Botany, 47, 115–126.
Chen, C., Huang, D., & Liu, J. (2009). Functions and toxicity of nickel in plants: recent advances and future prospects. Clean, 37, 304–313.
Gajewska, E., Wielanek, M., Bergier, K., & Skłodowska, M. (2009). Nickel induced depression of nitrogen assimilation in wheat roots. Acta Physiologiae Plantarum, 31, 1291–1300.
Dan, T. V., Krishnaraj, S., & Saxena, P. K. (2002). Cadmium and nickel uptake and accumulation in scented geranium (Pelargonielm sp. ‘Frensham’). Water. Air and Soil Pollution, 137, 355–364.
Vogel-Mikus, K., Drobne, D., & Regvar, M. (2005). Zn, Cd and Pb accumulation and arbuscular mycorrhizal colonization of pennycress Thlaspi Praecox Wulf. (Brassicaceae) from the vicinity of a lead mine and smelter in Slovenia. Environmental Pollution, 133, 233–242.
Seregin, I. V., & Kozhevnikova, A. D. (2006). Physiological role of nickel and its toxic effects on higher plants. Russian Journal of Plant Physiology, 53, 257–277.
Nishida, S., Tsuzuki, C., Kato, A., Aisu, A., Yoshida, J., & Mizuno, T. (2011). AtIRT1, the primary iron uptake transporter in the root, mediates excess nickel accumulation in Arabidopsis thaliana. Plant and Cell Physiology, 52(8), 1433–1442.
Baetz, U., Eisenach, C., Tohge, T., Martinoia, E., & De Angeli, A. (2016). Vacuolar chloride fluxes impact ion content and distribution during early salinity stress. Plant Physiology, 00183.
Teodoro, A. E., Zingarelli, L., & Lado, P. (1998). Early changes of Cl efflux band H+ extrusion induced by osmotic stress in Arabidopsis thaliana cells. Physiologia Plantarum, 102, 29–37.
Xu, G., Magen, H., Tarchitzky, J., & Kafkafi, U. (2000). Advances in chloride nutrition. Advances in Agronomy, 68, 96–150.
White, P. J., & Broadley, M. R. (2001). Chloride in soils and its uptake and movement within the plant: a review. Annals of Botany, 88, 967–988.
Teakle, N. L., & Tyerman, S. D. (2010). Mechanisms of Cl-transport contributing to salt tolerance. Plant, Cell & Environment, 33(4), 566–589.
Moya, J. L., Gomez-Cadenas, A., Primo-Millo, E., & Talon, M. (2003). Chloride absorption in salt-sensitive Carrizo citrange and salt tolerant Cleopatra mandarin citrus rootstocks is linked to water use. Journal of Experimental Botany, 54, 825–833.
Franklin, J. A., & Zwiazek, J. J. (2004). Ion uptake in Pinus banksiana treated with sodium chloride and sodium sulphate. Physiologia Plantarum, 120, 482–490.
Luo, Q., Bingjun, Y., & Liu, Y. (2005). Differential selectivity to chloride and sodium ions in seedlings of Glycine max and G. soja under NaCl stress. Journal of Plant Physiology, 162, 1003–1012.
Teakle, N. L., Real, D., & Colmer, T. D. (2006). Growth and ion relations in response to combined salinity and waterlogging in the perennial forage legumes Lotus corniculatus and Lotus tenuis. Plant and Soil, 289, 369–383.
Teakle, N., Flowers, T., Real, D., & Colmer, T. (2007). Lotus tenuis Tolerates the interactive effects of salinity and waterlogging by ‘excluding’ Na+ and Cl- from the xylem. Journal of Experimental Botany, 58, 2169–2180.
Pilon-Smits, E. A. (2015). Selenium in plants. In Progress in botany (pp. 93–107). Springer International Publishing.
Chu, J. Z., Yao, X. Q., & Zhang, Z. N. (2010). Responses of wheat seedlings to exogenous selenium supply under cold stress. Biological Trace Element Research, 136(3), 355–363.
Hasanuzzaman, M., & Fujita, M. (2011). Selenium pretreatment upregulates the antioxidant defense and methylglyoxal detoxification system and confers enhanced tolerance to drought stress in rapeseed seedlings. Biological Trace Element Research, 143, 1758–1776.
Djanaguiraman, M., Prasad, P. V. V., & Seppänen, M. (2010). Selenium protects sorghum leaves from oxidative damage under high temperature stress by enhancing antioxidant defense system. Plant Physiology and Biochemistry, 48(12), 999–1007.
Pukacka, S., Ratajczak, E., & Kalemba, E. (2011). The protective role of selenium in recalcitrant Acer saccharium L. Seeds subjected to desiccation. Journal of Plant Physiology, 168(3), 220–225.
Kumar, M., Bijo, A. J., Baghel, R. S., Reddy, C. R. K., & Jha, B. (2012). Selenium and Spermine alleviates cadmium induced toxicity in the red seaweed Gracilaria dura by regulating antioxidant system and DNA methylation. Plant Physiology and Biochemistry, 51, 129–138.
Wang, C. Q. (2011). Water-stress mitigation by selenium in Trifolium repens L. Journal of Plant Nutrition and Soil Science, 174(2), 276–282.
Hartikainen, H., Xue, T. L., & Piironen, V. (2000). Selenium as an anti-oxidant and prooxidant in ryegrass. Plant and Soil, 225, 193–200.
Yao, X., Chu, J., & Ba, C. (2010). Antioxidant responses of wheat seedlings to exogenous selenium supply under enhanced ultraviolet-B. Biological Trace Element Research, 136(1), 96–105.
Pezzarossa, B., Remorini, D., Gentile, M. L., & Massai, R. (2012). Effects of foliar and fruit addition of sodium selenate on selenium accumulation and fruit quality. Journal of the Science of Food and Agriculture, 92, 781–786.
Belokobylsky, A., Ginturi, E., Kuchava, N., Kirkesali, E., Mosulishvili, L., Frontasyeva, M., et al. (2004). Accumulation of selenium and chromium in the growth dynamics of Spirulina platensis. Journal of Radioanalytical and Nuclear Chemistry, 259(1), 65–68.
Feng, R. W., & Wei, C. Y. (2012). Antioxidative mechanisms on selenium accumulation in Pteris vittata L., a potential selenium phytoremediation plant. Plant, Soil and Environment, 58(3), 105–110.
Fordyce, F.M. (2005). Selenium deficiency and toxicity in the environment. In Essentials of Medical Geology (Selinus, O. et al., eds), Elsevier pp. 373–415.
Elrashidi, M. A., et al. (1987). Chemical-equilibria of seleniumin soils—a theoretical development. Soil Science, 144, 141–152.
Anderson, J. (1993). Selenium interactions in sulfur metabolism. In Sulfur nutrition and assimilation in higher plants: regulatory agricultural and environmental aspects (De Kok, I.S. et al., eds), SPB Academic, pp. 49–60.
White, P. J., Bowen, H. C., Parmaguru, P., Fritz, M., Spracklen, W. P., Spiby, R. E., et al. (2004). Interactions between selenium and sulphur nutrition in Arabidopsis thaliana. Journal of Experimental Botany, 55(404), 1927–1937.
Zhu, Y. G., Pilon-Smits, E. A., Zhao, F. J., Williams, P. N., & Meharg, A. A. (2009). Selenium in higher plants: understanding mechanisms for biofortification and phytoremediation. Trends in Plant Science, 14(8), 436–442.
Arvy, M. P. (1993). Selenate and selenite uptake and translocation in bean-plants (Phaseolus vulgaris. Journal of Experimental Botany, 44, 1083–1087.
Li, H. F., McGrath, S. P., & Zhao, F. J. (2008). Selenium uptake, translocation and speciation in wheat supplied with selenate or selenite. New Phytologist, 178(1), 92–102.
Zhang, J. L., Flowers, T. J., & Wang, S. M. (2010). Mechanisms of sodium uptake by roots of higher plants. Plant and Soil, 326(1–2), 45–60.
Flowers, T. J., & Yeo, A. R. (1988). Ion relation of salt tolerance. In: Baker DA, Hall JL (eds) Solute transport in plant cells and tissues. Longman Scientific and Technical, Harlow, pp 392–413.
Tester, M., & Davenport, R. (2003). Na+ tolerance and Na+ transport in higher plants. Annual Botany, 91, 503–527.
Flowers, T. J., & Colmer, T. D. (2008). Salinity tolerance in halophytes. New Phytologist, 179, 945–963.
Kronzucker, H. J., & Britto, D. T. (2011). Sodium transport in plants: a critical review. New Phytologist, 189(1), 54–81.
Amtmann, A., & Sanders, D. (1999). Mechanism of Na+ uptake by plant cells. Advances in Botanical Research, 29, 75–112.
Tyerman, S. D., & Skerrett, I. M. (1999). Root ion channels and salinity. Scientia Horticulturae, 78, 175–235.
White, P. J. (1999). The molecular mechanism of sodium influx to root cells. Trends in Plant Science, 4(7), 245–246.
Antosiewicz, D. M., & Hennig, J. (2004). Overexpression of LCT1 in tobacco enhances the protective action of calcium against cadmium toxicity. Environmental Pollution, 129, 237–245.
Liu, W. H., Schachtman, D. P., & Zhang, W. (2000). Partial deletion of a loop region in the high affinity K+ transporter HKT1 changes ionic permeability leading to increased salt tolerance. Journal of Biological Chemistry, 275, 27924–27932.
Blumwald, E., Aharon, G. S., & Apse, M. P. (2000). Sodium transport in plant cells. Biochim Biophys Acta-Biomembranes, 1465, 140–151.
?>Senn, M. E., Rubio, F., Banuelos, M. A., & Rodrıguez-Navarro, A. (2001). Comparative functional features of plant potassium HvHAK1 and HvHAK2 transporters. Journal of Biological Chemistry, 30, 44563–44569.
Colmenero-Flores, J. M., Martinez, G., Gamba, G., Vazquez, N., Iglesias, D. J., Brumos, J., et al. (2007). Identification and functional characterization of cation-chloride cotransporters in plants. Plant Journal, 50, 278–292.
Hajiboland, R., Bahrami Rad, S., Barceló, J., & Poschenrieder, C. (2013). Mechanisms of aluminum-induced growth stimulation in tea (Camellia sinensis). Journal of Plant Nutrition and Soil Science, 176(4), 616–625.
Barcelo, J., & Poschenrieder, C. (2002). Fast root growth responses, root exudates, and internal detoxification as clues to the mechanisms of aluminium toxicity and resistance: a review. Environmental and Experimental Botany, 48(1), 75–92.
Sade, H., Meriga, B., Surapu, V., Gadi, J., Sunita, M. S. L., Suravajhala, P., et al. (2016). Toxicity and tolerance of aluminum in plants: tailoring plants to suit to acid soils. Biometals, 29(2), 187–210.
Poschenrieder, C., Gunsé, B., Corrales, I., & Barceló, J. (2008). A glance into aluminum toxicity and resistance in plants. Science of the Total Environment, 400(1), 356–368.
Konishi, S. (1992). Promotive effects of aluminium on tea plant growth. Japan Agricultural. Research Quarterly, 26, 26–26.
Watanabe, T., & Osaki, M. (2002). Mechanisms of adaptation to high aluminum condition in native plant species growing in acid soils: a review. Communications in Soil Sciences and Plant Analysis, 33, 1247–1260.
Jansen, S., Watanabe, T., Caris, P., Geuten, K., Lens, F., Pyck, N., et al. (2004). The distribution and phylogeny of aluminium accumulating plants in the Ericales. Plant Biology, 6, 498–505.
Osawa, H., Ikeda, S., & Tange, T. (2013). The rapid accumulation of aluminum is ubiquitous in both the evergreen and deciduous leaves of Theaceae and Ternstroemiaceae plants over a wide pH range in acidic soils. Plant and Soil, 363(1–2), 49–59.
Hajiboland, R., Barceló, J., Poschenrieder, C., & Tolrà, R. (2013). Amelioration of iron toxicity: a mechanism for aluminum-induced growth stimulation in tea plants. Journal of Inorganic Biochemistry, 128, 183–187.
Kinraide, T. B. (1994). Use of a Gouy-Chapman-stern model for membrane-surface electrical potential to interpret some features of mineral rhizotoxicity. Plant Physiology, 106(4), 1583–1592.
Kidd, P. S., & Proctor, J. (2001). Why plants grow poorly on very acid soils: are ecologists missing the obvious? Journal of Experimental Botany, 52(357), 791–799.
Ghanati, F., Morita, A., & Yokota, H. (2005). Effects of aluminum on the growth of tea plant and activation of antioxidant system. Plant and Soil, 276(1–2), 133–141.
Komeda, H., Kobayashi, M., & Shimizu, S. (1997). A novel transporter involved in cobalt uptake. Proceedings of the National Academy of Sciences, 94, 36–41.
Bakkaus, E., Gouget, B., Gallien, J. P., Khodja, H., Carrot, F., Morel, J. L., et al. (2005). Concentration and distribution of cobalt in higher plants: the use of micro-PIXE spectroscopy. Nuclear Instruments and Methods in Physic Research, 231, 350.
Sree, K. S., Keresztes, Á., Mueller-Roeber, B., Brandt, R., Eberius, M., Fischer, W., et al. (2015). Phytotoxicity of cobalt ions on the duckweed Lemna minor—morphology, ion uptake, and starch accumulation. Chemosphere, 131, 149–156.
Cai, Z., Kastell, A., Speiser, C., & Smetanska, I. (2013). Enhanced resveratrol production in Vitis vinifera cell suspension cultures by heavy metals without loss of cell viability. Applied Biochemistry and Biotechnology, 171(2), 330–340.
Jayakumar, K., Jaleel, C. A., & Azooz, M. M. (2008). Impact of cobalt on germination and seedling growth of Eleusine coracana L. And Oryza sativa L. Under hydroponic culture. Global Journal of Molecular Sciences, 3(1), 18–20.
Wei, W., Wang, Y., Wei, Z. G., Zhao, H. Y., Li, H. X., & Hu, F. (2009). Roles of organic acids and nitrate in the long-distance transport of cobalt in xylem saps of Alyssum murale and Trifolium subterraneum. Biological Trace Element Research, 131(2), 165–176.
Karovic, O., Tonazzini, I., Rebola, N., Edström, E., Lövdahl, C., Fredholm, B. B., et al. (2007). Toxic effects of cobalt in primary cultures of mouse astrocytes similarities with hypoxia and role of HIF-1α. Biochemical Pharmacology, 73, 694–708.
Palit, S., Sharma, A., & Talukder, G. (1994). Effects of cobalt on plants. Botanical Review, 60, 149–181.
Barysas, D., Cesniene, T., Balciuniene, L., Vaitkuniene, V., & Rancelis, V. (2002). Genotoxicity of Co2+ in plants and other organisms. Biologija, 2, 58–63.
Kukier, U., Peters, C. A., Chaney, R. L., Angle, J. S., & Roseberg, R. J. (2004). The effect of pH on metal accumulation in two alyssum species. Journal of Environmental Quality, 33, 2090–2102.
Li, Z., McLaren, R. G., & Metherell, A. K. (2004). The availability of native and applied soil cobalt to ryegrass in relation to soil cobalt and manganese status and other soil properties. New Zealand Journal of Agricultural Research, 47, 33–43.
Li, H. F., Gray, C., Mico, C., Zhao, F. J., & McGrath, S. P. (2009). Phytotoxicity and bioavailability of cobalt to plants in a range of soils. Chemosphere, 75(7), 979–986.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vatansever, R., Ozyigit, I.I. & Filiz, E. Essential and Beneficial Trace Elements in Plants, and Their Transport in Roots: a Review. Appl Biochem Biotechnol 181, 464–482 (2017). https://doi.org/10.1007/s12010-016-2224-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-016-2224-3