Abstract
Growth, ionic responses, and expression of candidate genes to salinity stress were examined in two perennial ryegrass accessions differing in salinity tolerance. The salinity tolerant (PI265349) and sensitive accessions (PI231595) were subjected to 75-mM NaCl for 14 days in a growth chamber. Across two accessions, salinity stress increased shoot dry weight and concentrations of malondialdehyde (MDA) and Na+ in the shoots and roots, but decreased shoot Ca2+ and root K+ concentrations. Salinity stress also increased root expressions of SOS1, PIP1, and TIP1. Plant height and chlorophyll content were unaffected by salinity stress in the tolerant accession but significantly decreased in the sensitive accession. Shoot MDA content did not change in the tolerant accession but increased in the sensitive accession. A more dramatic increase in Na+ was found in the roots of the sensitive accession. Relative to the control, salinity stress reduced expression of SOS1, NHX1, PIP1, and TIP1 in the shoots but increased expression of these genes in the roots of the tolerant accession. Expression levels of SOS1 increased in the roots and expression of NHX1 increased in the shoots but decreased in the roots of the sensitive accession under salinity stress. A decline in PIP1 expression in the shoots and dramatic increases in TIP expression in both shoots and roots were found in the sensitive accession under salinity stress. The results suggested maintenance of plant growth and leaf chlorophyll content, lesser Na+ accumulation in the roots, and lower lipid peroxidation in the shoots which could be associated with salinity tolerance. The decreased expressions of SOS1, NHX1, and TIP1 in the shoots, and increased expressions of NHX1 and PIP1 in the roots might also be related to salinity tolerance in perennial ryegrass.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Plants response to salinity stress involve numerous growth, and physiological and biochemical alterations (Munns and Tester 2008). At the whole-plant level, plant growth is sensitive to salinity stress. Salinity reduced length of shoots and roots, leaf dry weight, leaf and root water content, and plant transpiration (Vysotskaya et al. 2010; Ruiz-Carrasco et al. 2011; Tang et al. 2013a). However, salinity effects on plant growth varied largely among or within a perennial grass species (Dai et al. 2008; Wang et al. 2011; Uddin et al. 2012; Tang et al. 2013a). Variations of plant growth under salinity stress depend on plant species, genotypes, and intensity and duration of a stress.
Salinity stress increases Na+ level in the plants. The decrease in K+ concentration under salinity stress often occurs along with Na+ accumulation, which is considered an imbalance of ion homeostasis. The tolerant genotypes of grass species showed low level of Na+, high level of K+, and high K+/Na+ ratio (Krishnan and Brown 2009; Hu et al. 2012b), but the results were not consistent. Salinity stress also interferes with uptake and accumulation of other nutrients such as Ca2+ and Mg2+, and reductions in Ca2+ and Mg2+ concentration were greater in the salinity-sensitive cultivar of pea (Pisum sativum L.) (Ahmad et al. 2008). The addition of Ca2+ considerably increased the uptake and transport capacity for K+ over Na+, suggesting that Ca2+ could regulate K+/Na+ homeostasis (Wu and Wang 2012). However, the effects of Ca2+ on the plants also varied with genotypes and NaCl concentrations (Cabot et al. 2009). Collectively, plant tolerance to salinity stress is associated with ionic stress tolerance (Munns and Tester 2008). Molecular mechanisms of ion transport may be essential for maintenance of ion hemostasis that contributes to salinity tolerance.
At the molecular level, candidate genes for ion accumulation and water movement are of interest in illustrating mechanisms of salinity tolerance. The salt overly sensitive (SOS1) gene, encoding a Na+/H+ antiporter in plasma membrane, can play an important role in excluding Na+ and controlling long-distance transport of Na+ from the roots to shoots in Arabidopsis thaliana (Shi et al. 2002). Salinity stress increased the expression level of SOS1, which might be associated with salinity tolerance (Gao et al. 2016). Expressions of CqSOS1 were differentially regulated in the shoots and roots as well as between the tolerant and sensitive genotypes of quinoa (Chenopodium quinoa Wild.) (Ruiz-Carrasco et al. 2011). Moreover, overexpression of SOS1 enhanced salinity tolerance of the transgenic plants (Gao et al. 2016). NHX1, a vacuolar Na+/H+ antiporter, controls Na+ movement across the tonoplast membrane into the vacuole (Munns and Tester 2008). Expression patterns of ZjNHX1 varied with tissue types in zoysia grass (Zoysia japonica Steud.) under salinity stress (Du et al. 2010). Expression of CqNHX1 also differed in shoots, roots, and genotypes (Ruiz-Carrasco et al. 2011). Overexpression of NHX1 improved salinity tolerance in some plant species (Banjara et al. 2012; Sahoo et al. 2016). However, overexpression of AtNHX1 alone did not improve salinity tolerance in transgenic Arabidopsis (Yang et al. 2009). The results indicate that gene coordination for controlling ion homeostasis can be vital for plant salinity tolerance.
Salinity causes osmotic stress and interferes in plant–water relations. Aquaporins regulate water movement across the plant plasma membrane (e.g., PIP) and vacuolar membrane (e.g., TIP) (Maurel et al. 2008). The salinity tolerant wild barley (Hordeum spontaneumin L.) had a fast and persistent down-regulation of the HtPIP2;4 gene, while the sensitive line showed a delayed down-regulation of the gene, consistent with inhibited transpiration and hydraulic conductivity caused by salinity (Vysotskaya et al. 2010). In cucumber (Cucumis sativus L.), the decreased hydraulic conductivity under salinity stress was related to a down-regulation of CsPIP1;2 and CsPIP2;4, while the alterations of CsPIP expression were consistent with partial recovery of hydraulic conductivity of the roots after salinity treatment (Qian et al. 2015). The transcript levels of FrPIP2;1 were suppressed in red fescue (Festuca rubra L.) subjected to 500 mM NaCl stress, but PIP2;1 expression level was correlated with salinity tolerance (Diédhiou et al. 2009). In addition, overexpression of aquaporin genes such as PIP and TIP increased salinity tolerance of plants (Hu et al. 2012b; Zhou et al. 2014; Wang et al. 2015). It appears that aquaporins play a role in salinity tolerance by mediating water transport of seedlings, although the regulatory mechanisms could vary with species, stress durations, and specific organs.
Perennial ryegrass (Lolium perenne L.) is a commonly used perennial grass for turf, forage, and soil stabilization. This species exhibited large natural variation of salinity responses (Tang et al. 2013b). However, whole-plant and molecular mechanisms of salinity tolerance are not well understood in this species, especially under low salinity stress. Thus, the experiment was designed to study growth, ionic responses, and expression of candidate genes to salinity stress in two perennial ryegrass accessions differing in salinity tolerance. The results would contribute to a better understanding of whole-plant physiological and molecular mechanisms of salinity tolerance in perennial grass species under suboptimal growing conditions.
Materials and methods
Plant materials
The salinity tolerant (PI265349) and sensitive accessions (PI231595) of perennial ryegrass were seeded in plastic pots (4-cm diameter, 9-cm deep) containing sand in a greenhouse at Purdue University, USA, on 7 April 2014. Plants were cut twice a week to a height of 5–6 cm and irrigated three times a week or whenever needed using a half-strength Hoagland solution (pH ≈ 6.5, electrical conductivity ≈ 1.2 dS m−1). All the pots were well drained after irrigation. Grasses were grown at temperatures of 21/16 °C (day/night) with a photosynthetically active radiation (PAR) of 500 µmol m−2 s−1. On 23 June 2014, pots were placed in a growth chamber with temperatures of 20/15 °C (day/night) and PAR of 500 µmol m−2 s−1. The grasses were irrigated daily with a half-strength Hoagland solution and mowed twice a week for 7 days in the growth chamber before initiation of salinity stress.
Salinity treatment
Plants were irrigated daily with a half-strength Hoagland solution (non-stress control) or with 75-mM NaCl in a half-strength Hoagland solution (EC ≈ 8.5 dS m−1). Salt concentration was increased gradually by 25 mM every day. Plants were harvested after 14 days of 75-mM NaCl when two genotypes of perennial ryegrass visually showed differences in growth and color under salinity stress. Plants were not cut during salinity treatment.
Measurements
Plant height (HT) was determined from the soil surface to the top of leaf blade. Shoot tissues were weighed and dried at 80 °C for 3 days, and then, the dry weight was measured. For chlorophyll (Chl) extraction, approximately 50 mg of randomly selected fresh leaves were soaked in 15-mL dimethyl sulfoxide in dark for 48 h. Subsequently, the absorbance of Chl extract was read at 665, 649, and 480 nm, and Chl was calculated according to the method of Wellburn (1994). Roots were harvested, and the fresh and dried tissues were saved for analyses.
Dry shoot and root samples were ground into a fine powder with liquid nitrogen. For extraction of ions, we followed the method described by Tang et al. (2013a). Briefly, a 50-mg fine powder was mixed with 3 mL of 18 M H2SO4 in a digestion tube and then heated at 200 °C for 30 min. After the tubes cooled down to the room temperature, a 3-mL 30% H2O2 was added and tubes were heated again for 30 min until the mixture became transparent and clear. Distilled water was added to the extraction to bring the volume to 50 mL. A 2-mL aliquot of extraction was diluted to 15 mL with distilled water. The Na+, K+, Ca2+, and Mg2+ concentrations were determined using a plasma atomic emission spectrometer (ICP 9820; Shimadzu, Columbia, MD, USA).
The malondialdehyde (MDA) is indicator of lipid peroxidation (Dhindsa et al. 1981). We measured MDA concentration according to the method described by Liu and Jiang (2015). Approximately 50 mg frozen leaf or root tissues were ground into fine powder with liquid nitrogen. A 1 mL of extraction buffer (pH ≈ 7.8) containing 50-mM potassium phosphate, 1% polyvinylpyrrolidone, and 1 mM ethylenediaminetetraacetic acid was added to the powder. After centrifuging at 15,000×g for 30 min at 4 °C, a 0.5 mL of the supernatant was mixed with a 2 mL of trichloroacetic acid (20%) containing thiobarbituric acid (5%). The tube was heated at 95 °C for 30 min, immediately cooled in ice, and centrifuged at 10,000×g for 10 min. The MDA concentration was calculated using the absorbance at 532 and 600 nm, and an extinction coefficient of 155 mm−1cm−1 (Health and Packer 1968).
Gene expression was examined using the real-time quantitative reverse transcription polymerase chain reaction (qRT-PCR) procedure. Briefly, total RNA was extracted with a Direct-zol™ RNA MiniPrep Kit (Zymo Research Corporation, Irvine, CA, USA). About 0.5–1 µg of extracted RNA was mixed with an iScript™ cDNA Synthesis Kit (Bio-Rad, Hercules, CA, USA) to make cDNA under 25 °C for 5 min, 42 °C for 30 min, and 85 °C for 5 min. Quantitative RT-PCR was performed using a CFX96 Touch™ Real-Time PCR System with an iTaq™ Universal SYBR® Green Kit (Bio-Rad, Hercules, CA, USA), with a reaction at 95 °C for 3 min followed by 40 amplification cycles of 10 s at 95 °C and 1 min at 60 °C. All primers were designed using Primer Primer 5 (Lalitha 2004) from perennial ryegrass sequences as follows: for SOS1 (AY987046.1), forward 5′-ATTATTACAGAATCGGTGGT-3′ and reverse 5′-AATCCTCGCAATGACTAG-3′; for NHX1 (AY987047.1), forward 5′-CCCAATGAAATCAGTCGC-3′, and reverse 5′-CCAAATAATAACTTGCTGCCTA-3′; for PIP1 (genomic sequence), forward 5′-TGTGCTCCCTGTTGTAGATGA-3′ and reverse 5′-CCACTGCCTATTGGGTTCG-3′; for TIP1 (genomic sequence), forward 5′-TGAGGCAGGTTTATTAGTTTC-3′ and reverse 5′-CAGCAGATGATCGGTTTG-3′. Perennial ryegrass elongation factor 1-alpha (EU168438), forward 5′-GGCTGATTGTGCTGTGCTTA-3′, and reverse 5′-CTCACTCCAAGGGTGAAAGC-3′ were used as a housekeeping gene for internal control. The relative gene expression level between the control and salinity stress treatment was calculated using the method of 2−ΔΔCT (Livak and Schmittgen 2001), with three biological and three technical replicates for each treatment.
Experimental design and data analysis
The experimental was a split plot design, with main plot of salinity treatment (control and 75 mM NaCl) and subplot of accessions. Three pots representing three replicates were randomly assigned into each treatment. Analysis of variance was conducted to indicate effects of accessions, treatments, and accessions × treatments using SAS (version 9.1; SAS Institute, Cary, NC, USA). Least square means were tested in the mixed model at significance level of 0.05.
Results and discussion
Analysis of variance
Across the two cultivars, significant treatment effects were found for shoot dry weight (DW), concentrations of shoot and root malondialdehyde (MDA) and Na+, root K+ concentration, shoot Ca2+ concentration, expression of root SOS1, and expressions of PIP1 and TIP1 in the shoots and roots (Table 1).
The cultivar effects were observed for shoot chlorophyll content (Chl), plant height (HT), shoot and root DW, shoot MDA, shoot K+, root Na+, shoot SOS1, and expressions of NHX1, PIP1, and TIP1 in the shoots and roots (Table 1). Significant treatment by cultivar interactions was shown for Chl, HT, shoot MDA, root Na+, shoot SOS1, and NHX1, PIP1, and TIP1 in the shoots and roots (Table 1).
Treatment effects on traits across two cultivars
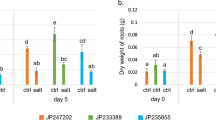
Shoot DW, MDA, and Na+ concentrations increased 33.1, 19.6%, and sevenfold, and shoot Ca2+ was reduced 21.7% under 75-mM NaCl compared with the control, respectively (Table 2). In addition, for shoots, salinity stress significantly increased transcript level of PIP1 but decreased expression of TIP1 (Table 2). Root MDA concentration increased 8.9% and root Na+ concentration dramatically increased 11.8-fold, but root K+ decreased 41.9% under salinity stress, compared to the control (Table 2). Expressions of root SOS1, PIP1, and TIP1 increased under salinity stress, especially for TIP1 (Table 2).
Cultivar differences in traits across treatments
Across the control and salinity treatments, the tolerant accession of PI265349 had lower Chl and shoot MDA but higher shoot DW and K+ than that of the sensitive accession of PI231595 (Table 3). The tolerant accession also had lower expression levels of SOS1, NHX1, and TIP1 and higher level of PIP1 than the sensitive accession (Table 3). In the roots, higher DW and lower Na+ were observed in the tolerant accession than in the sensitive accession (Table 3). In addition, higher expression levels of NHX1 and PIP1 and lower level of TIP1 were found in the tolerant accession than in the sensitive accession (Table 3).
Growth and physiological responses of individual cultivars to treatments
Plant height (HT) and Chl content were unaffected by 75-mM NaCl in the tolerant accession of PI265349, but significantly decreased 15.9 and 23.6% for the sensitive accession of PI231959, respectively, compared to the non-salinity control (Fig. 1). Salinity stress did not change MDA content in the shoots of the tolerant accession but increased 36.8% in the shoots of the sensitive accession (Fig. 1).
Plant height (HT), leaf chlorophyll content (Chl), shoot malondialdehyde (MDA), and root Na+ concentrations as affected by 14 days of 75-mM NaCl in the salinity tolerant accession of PI265349 and the sensitive accession of PI231595. Comparisons were made between the control (0-mM NaCl, open column) and 75-mM NaCl (shade column) for each accession. *p < 0.05
A reduction in HT in the sensitive accession suggested that plant growth of the sensitive accession of perennial ryegrass was inhibited by a relatively low NaCl concentration. However, the reduced HT was not consistent increase in SDW in the sensitive accession, suggesting that alterations of dry weight might not always be associated with salinity tolerance. A reduced or increased dry weight was found in plant species exposed to salinity stress including perennial grasses (Hu et al. 2012b; Uddin et al. 2012; Tang et al. 2013a; Yin et al. 2017). Under high salinity stress (~ 300-mM NaCl), HT was positively correlated with leaf dry weight among 56 perennial ryegrass accessions (Tang et al. 2013b). Genotypes, stress intensity, and duration may all cause variations of plant growth in different studies in perennial grasses.
Leaf Chl decreased or remained unchanged in perennial ryegrass under salinity stress, depending on salt concentration and plant materials (Hu et al. 2011; Tang et al. 2013a; Yin et al. 2017). Leaf Chl were unaffected in perennial ryegrass cultivar of Quick start II under 85- and 170-mM NaCl, but deceased under 255-mM NaCl (Hu et al. 2011). The decrease in Chl content in PI231595 indicated its sensitivity to salinity stress, which was consistent with its reduced HT and increased MDA in the shoots. MDA level is an indicator of lipid peroxidation. Increased leaf and root MDA content were observed in perennial grass and other plant species under salinity stress (Hu et al. 2011, 2012a; Chen and Ye 2014; Yin et al. 2017). The salinity tolerant perennial ryegrass cultivar accumulated lesser amount of MDA compared to the sensitive cultivar under 12 days of 250-mM NaCl (Hu et al. 2012a). The unchanged level of MDA in the tolerant accession and increased MDA in the sensitive accession in this study indicated that the degree of lipid peroxidation induced by salinity was more severe in the salinity-sensitive accession of perennial ryegrass.
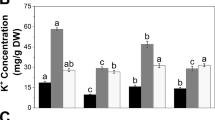
Salinity increased root Na+ concentrations for both accessions (Fig. 1). The Na+ concentration of the roots increased 6.3-fold in the tolerant accession and 18.6-fold in the sensitive accession under salinity stress, compared with the control (Fig. 1). The low uptake of Na+ and a higher K+/Na+ ratio could be associated with salinity tolerance (Thalji and Shalaldeh 2007; Azadi et al. 2011; Hu et al. 2011; Tang et al. 2013a, b), but the results are not often consistent (Munns and James 2003; Yin et al. 2017). Compared with the sensitive accession, higher K+ accumulation in the shoot and lower Na+ accumulation in the roots were found in the tolerant accession (Table 3; Fig. 1), which may contribute to its better salinity tolerance.
Accumulation of Na+ in the soil decreased Ca2+ uptake and transport (Hu and Schmidhalter 1997). Meanwhile, Ca2+ highly competes with Mg2+ at the binding sites of root plasma membranes, which can interfere with both Mg2+ and Ca2+ uptake under salinity stress (Grattan and Grieve 1998). The reduced Ca2+ in the shoots under salinity stress could be due to a high level of Na+ accumulation (Table 2). The reductions in leaf Ca2+ and Mg2+ concentrations were found in two citrus stocks in response to increasing NaCl concentrations but not in other stocks (Ruiz et al. 1997). Collectively, the results suggested that the salinity effect of Ca2+ and Mg2+ varied with genotypes, but a relatively lower Na+ accumulation and higher levels of Ca2+ and K+/Na+ could contribute to salinity tolerance in perennial ryegrass.
Gene expression under salinity
The gene expression patterns were consistent in the shoots and roots in the tolerant accession of PI2653496, but not in the sensitive accession of PI231595 (Figs. 2, 3). Relative to the non-stress control, the transcript abundance of SOS1 and NHX1 decreased in the shoots, but increased in the roots of the tolerant accession under 75-mM NaCl (Fig. 2). For the sensitive accession, the transcript levels of SOS1 in the shoots were unaffected by salinity stress, but was reduced in the roots, while expression of NHX1 increased in the shoots and decreased in the roots, relative to the control (Fig. 2). Expression patterns of PIP1 and TIP1 were similar to that of SOS1 and NHX1 in the roots of the tolerant accession exposed to 75-mM NaCl (Fig. 3). A decline in PIP1 expression was found in the shoots but not in the roots, while dramatic increases in TIP1 expressions were noted in both shoot and root tissues of the sensitive accession under salinity stress (Fig. 3).
Gene expression of SOS1 and NHX1 in the shoots and roots as affected by 14 days of 75-mM NaCl in the salinity tolerant accession of PI265349 and the sensitive accession of PI231595. Comparisons were made between the control (0-mM NaCl, open column) and 75-mM NaCl (shade column) for each accession. Data were normalized in comparison with their respective control. *p < 0.05
Gene expression of PIP1 and TIP1 in the shoots and roots as affected by 14 days of 75-mM NaCl in the salinity tolerant accession of PI265349 and the sensitive accession of PI231595. Comparisons were made between the control (0-mM NaCl, open column) and 75-mM NaCl (shade column) for each accession. Data were normalized in comparison with their respective control. *p < 0.05
The transport and compartment of Na+ and K+ between different tissues and subcellular locations may allow plants to adjust to a saline environment (Silva et al. 2015). Expression of SOS1 gene encoding Na+/H+ antiporter in plasma membrane and NHX1 encoding vascular membrane Na+/H+ antiporter may confer salinity tolerance (Wu et al. 2005; Gao et al. 2016). Variations in Na+ accumulation in the shoots were associated with the expression level of AtSOS1 in different Arabidopsis ecotypes (Jha et al. 2010). Higher expression levels of SOS1 in the roots of both perennial ryegrass accessions were consistent with accumulation of Na+ in the roots under salinity stress (Fig. 1). However, down-regulation of SOS1 and NHX1 in the shoots of the tolerant accession was not consistent with shoot Na+ accumulation under salinity. Overall, our results in perennial ryegrass supported that expressions of SOS1 and NHX1 had tissue-specific patterns and were regulated differentially in the shoots and roots as well as between the tolerant and sensitive genotypes (Du et al. 2010; Ruiz-Carrasco et al. 2011). In addition, allelic variation of NHX1 was strongly associated with leaf K+/Na+ ratio in natural populations of perennial ryegrass (Tang et al. 2013b). In this study, reduced root K+/Na+ along with the lesser expressed level of NHX1 in the roots of the sensitive accession might suggest a role of this gene in salinity tolerance. However, other genes involved in K+ transport could also be important for illustrating ion homeostasis under salinity stress.
Salinity stress influences plant water status. Aquaporins could affect salinity tolerance by mediating water transport across plant vacuolar and plasma membranes. Up- and down-regulations of PIP1 and TIP1 found in the shoots and roots of both accessions supported the differential expression patterns of PIP and TIP identified in other plant species under salinity stress (Diédhiou et al. 2009; Vysotskaya et al. 2010; Xu et al. 2013; Hove et al. 2015; Qian et al. 2015). The lowered PIP activity contributed to decreased water uptake capacity in the roots grown under salinity stress, but up-regulation of the PIP1 gene enhanced salinity tolerance in transgenic Nicotiana tabacum L. (Hu et al. 2012c). Our results also suggested that increased expression of PIP1 and TIP1 in the roots of the tolerant accession could contribute to salinity tolerance of this accession. Expression patterns of PIP1 and TIP1 were not consistent in the sensitive accession, suggesting differential roles of these genes in salinity response in the sensitive genotype of perennial ryegrass. In wheat (Triticum aestivum L.), the TIP2;2 gene was down-regulated by salinity, and heterologous expression of TaTIP2;2 compromised salinity and drought tolerance of Arabidopsis thaliana (Xu et al. 2013). The dramatic increases in TIP1 in both shoots and roots of the sensitive accession indicated that TIP1 might be a negative regulator of salinity tolerance in perennial ryegrass. This deserves further investigation.
In summary, maintenance of HT and Chl and absence of accumulation of MDA in the shoots were associated with salinity tolerance. Lesser increases of Na+ in the roots contributed to salinity tolerance. The decreased expressions of SOS1, NHX1, and TIP1 in the shoots and increased expressions of NHX1 and PIP1 in the roots might also be related to salinity tolerance in perennial ryegrass.
References
Ahmad P, John R, Sarwat M, Umar S (2008) Responses of proline, lipid peroxidation and antioxidative enzymes in two varieties of Pisum sativum L. under salt stress. Intl J Plant Production 2:353–366
Azadi A, Hervan EM, Mohammadi SA, Moradi F, Nakhoda B, Vahabzade M (2011) Screening of recombinant inbred lines for salinity tolerance in bread wheat (Triticum aestivum L.). Afri J Biotech 10:12875–12881
Banjara M, Zhu L, Shen G, Payton P, Zhang H (2012) Expression of an Arabidopsis sodium/proton antiporter gene (AtNHX1) in peanut to improve salt tolerance. Plant Biotechnol Rep 6:59–67
Cabot C, Sibole JV, Barceló J, Poschenrieder C (2009) Sodiu–-calcium interactions with growth, water, and photosynthetic parameters in salt-treated beans. J Plant Nutr Soil Sci 172:637–643
Chen Y, Ye Y (2014) Effects of salinity and nutrient addition on mangrove Excoecaria agallocha. PLoS One 9:e93337
Dai J, Schlossberg MJ, Huff DR (2008) Salinity tolerance of 33 greens-type Poa annua experimental lines. Crop Sci 48:1187–1192
Dhindsa RS, Plumb-Dhindsa P, Thorpe TA (1981) Leaf senescence: correlated with increased leaves of membrane permeability and lipid peroxidation and decreased levels of superoxide dismutase and catalase. J Exp Bot 32:93–101
Diédhiou CJ, Popova OV, Golldack D (2009) Transcript profiling of the salt-tolerant Festuca rubra ssp. litoralis reveals a regulatory network controlling salt acclimatization. J Plant Physiol 166:697–711
Du Y, Hei Q, Liu Y, Zhang H, Xu K, Xia T (2010) Isolation and characterization of a putative vacuolar Na+/H+ antiporter gene from Zoysia japonica L. J Plant Biol 53:251–258
Gao J, Sun J, Cao P, Ren L, Liu C, Chen S, Chen F, Jiang J (2016) Variation in tissue Na+ content and the activity of SOS1 genes among two species and two related genera of Chrysanthemum. BMC Plant Biol 16:98
Grattan S, Grieve C (1998) Salinity–mineral nutrient relations in horticultural crops. Sci Hort 78:127–157
Health RL, Packer L (1968) Photoperoxidation in isolated chloroplasts. I. Kinetics and stoichiometry of fatty acid peroxidation. Arch Biochem Biophys 125:189–198
Hove RM, Ziemann M, Bhave M (2015) Identification and expression analysis of the barley (Hordeum vulgare L.) aquaporin gene family. PLoS One 10:e0128025
Hu Y, Schmidhalter U (1997) Interactive effects of salinity and macronutrient level on wheat. II. Composition. J Plant Nutr 20:1169–1182
Hu T, Li H, Zhang X, Luo H, Fu J (2011) Toxic effect of NaCl on ion metabolism, antioxidative enzymes and gene expression of perennial ryegrass. Ecotoxicol Environ Saf 74:2050–2056
Hu L, Li H, Pang H, Fu J (2012a) Responses of antioxidant gene, protein and enzymes to salinity stress in two genotypes of perennial ryegrass (Lolium perenne) differing in salt tolerance. J Plant Physiol 169:146–156
Hu L, Huang Z, Liu S, Fu J (2012b) Growth response and gene expression in antioxidant-related enzymes in two bermudagrass genotypes differing in salt tolerance. J Amer Soc Hort Sci 137:134–143
Hu W, Yuan Q, Wang Y, Cai R, Deng X, Wang J, Zhou S, Chen M, Chen L, Huang C, Ma Z, Yang G, He G (2012c) Overexpression of a wheat aquaporin gene, TaAQP8, enhances salt stress tolerance in transgenic tobacco. Plant Cell Physiol 53:2127–2141
Jha D, Shirley N, Tester M, Roy J (2010) Variation in salinity tolerance and shoot sodium accumulation in Arabidopsis ecotypes linked to differences in the natural expression levels of transporters involved in sodium transport. Plant Cell Environ 33:793–804
Krishnan S, Brown RN (2009) Na+ and K+ accumulation in perennial ryegrass and red fescue accessions differing in salt tolerance. Intl Soc Turf Res J 11:817–827
Lalitha S (2004) Primer premier 5. Biotech Softw Internet Rep 1:270–272. https://doi.org/10.1089/152791600459894
Liu M, Jiang Y (2015) Genotypic variation in growth and metabolic responses of perennial ryegrass exposed to short-term waterlogging and submergence stress. Plant Physiol Biochem 95:57–64
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT C T method. Methods 25:402–408
Maurel C, Verdoucq L, Luu DT, Santoni V (2008) Plant aquaporins: membrane channels with multiple integrated functions. Ann Rev Plant Biol 59:595–624
Munns R, James RA (2003) Screening methods for salinity tolerance: a case study with tetraploid wheat. Plant Soil 253:201–218
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Qian Z-J, Song J-J, Chaumont F, Ye Q (2015) Differential responses of plasma membrane aquaporins in mediating water transport of cucumber seedlings under osmotic and salt stresses. Plant Cell Environ 38:461–473
Ruiz D, Martínez V, Cerdá A (1997) Citrus response to salinity: growth and nutrient uptake. Tree Physiol 17:141–150
Ruiz-Carrasco K, Antognoni F, Coulibaly AK, Lizardi S, Covarrubias A, Martínez EA, Molina-Montenegro MA, Biondi A, Zurita-Silva A (2011) Variation in salinity tolerance of four lowland genotypes of quinoa (Chenopodium quinoa Willd.) as assessed by growth, physiological traits, and sodium transporter gene expression. Plant Physiol Biochem 49:333–1341
Sahoo DP, Kumar S, Mishra S, Kobayashi Y, Panda SK, Sahoo L (2016) Enhanced salinity tolerance in transgenic mungbean overexpressing Arabidopsis antiporter (NHX1) gene. Mol Breed 36:144
Shi H, Quintero FJ, Pardo JM, Zhu J-K (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long distance Na+ transport in plants. Plant Cell 14:465–477
Silva EN, Silveira JAG, Rodrigues CRF, Viégas RA (2015) Physiological adjustment to salt stress in Jatropha curcas is associated with accumulation of salt ions, transport and selectivity of K+, osmotic adjustment and K+/Na+ homeostasis. Plant Biol 5:1023–1029
Tang J, Camberato JJ, Yu X, Luo N, Bian S, Jiang Y (2013a) Growth response, carbohydrate and ion accumulation of diverse perennial ryegrass accessions to increasing salinity. Sci Hort 154:73–81
Tang J, Yu X, Luo N, Xiao F, Camberato JJ, Jiang Y (2013b) Natural variation of salinity response, population structure and candidate genes associated with salinity tolerance in perennial ryegrass accessions. Plant Cell Environ 36:2021–2033
Thalji T, Shalaldeh G (2007) Screening wheat and barley genotype for salinity resistance. J Agron 6:75–78
Uddin MK, Juraimi AS, Ismail MR, Hossain MA, Othman R, Rahim AA (2012) Physiological and growth responses of six turfgrass species relative to salinity tolerance. Sci World J 2012:905468. https://doi.org/10.1100/2012/905468
Vysotskaya L, Hedley PE, Sharipova G, Veselov D, Kudoyarova G, Morris J, Jones HG (2010) Effect of salinity on water relations of wild barley plants differing in salt tolerance. AoB PLANTS. https://doi.org/10.1093/aobpla/plq006
Wang S, Zhang Q, Watkins E (2011) Evaluation of salinity tolerance of prairie junegrass, a potential low-maintenance turfgrass species. HortScience 46:1038–1043
Wang L, Li Q, Lei Q, Feng C, Gao Y, Zheng X, Zhao Y, Wang Z, Kong J (2015) MZPIP2;1: an aquaporin involved in radial water movement in both water uptake and transportation, altered the drought and salt tolerance of transgenic Arabidopsis. PLoS One 10:e0142446
Wellburn AR (1994) The spectral determination of chlorophylls a and b, as well as total carotenoids, using various solvents with spectrophotometers of different resolution. J Plant Physiol 144:307–313
Wu G, Wang S (2012) Calcium regulates K+/Na+ homeostasis in rice (Oryza sativa L.) under saline conditions. Plant Soil Environ 58:121–127
Wu Y, Chen Q, Chen M, Chen J, Wang X (2005) Salt tolerant transgenic perennial ryegrass (Lolium perenne L.) obtained by Agrobacterium tumefaciens-mediated transformation of the vacuolar Na+/ H+ antiporter gene. Plant Sci 169:65–73
Xu C, Wang M, Zhou L, Quan T, Xia G (2013) Heterologous expression of the wheat aquaporin gene TaTIP2;2 compromises the abiotic stress tolerance of Arabidopsis thaliana. PLoS One 8:e79618
Yang Q, Chen Z-Z, Zhou X-F, Yin H-B, Li X, Xin X-F, Hong X-H, Zhu J-K, Gong Z (2009) Overexpression of SOS (Salt Overly Sensitive) genes increases salt tolerance in transgenic Arabidopsis. Mol Plant 2:22–31
Yin X, Zhang C, Song X, Jiang Y (2017) Interactive short-term effects of waterlogging and salinity stress on growth and carbohydrate, lipid peroxidation, and nutrients in two perennial ryegrass cultivars. J Am Soc Hort Sci 142:110–118
Zhou L, Wang C, Liu RF, Han Q, Vandeleur RK, Du J, Tyerman S, Shou H (2014) Constitutive overexpression of soybean plasma membrane intrinsic protein GmPIP1;6 confers salt tolerance. BMC Plant Biol 14:181
Acknowledgements
This work was supported by the China Scholarship Council (No. 201708430007).
Author information
Authors and Affiliations
Contributions
ML conducted the experiments, performed the statistical analysis of data and prepared the manuscript. XS performed ion analysis. YJ supervised the overall work and contributed to data interpretation and discussion as well as paper writing.
Corresponding author
Additional information
Communicated by S Abe.
Rights and permissions
About this article
Cite this article
Liu, M., Song, X. & Jiang, Y. Growth, ionic response, and gene expression of shoots and roots of perennial ryegrass under salinity stress. Acta Physiol Plant 40, 112 (2018). https://doi.org/10.1007/s11738-018-2687-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-018-2687-7