Abstract
This study aimed to compare various responses of two Egyptian rice accessions bred for high yields, Sakha108 and Giza177, to salt stress at the seedling stage. Twenty-eight-day-old seedlings of two cultivars were grown in a hydroponic medium under control conditions (no NaCl) and salt stress (75 mM NaCl) for 12 days. Growth (dry weight), Na+ and K+ concentrations, enzymatic and non-enzymatic antioxidants, malondialdehyde (MDA), hydrogen peroxide (H2O2), and expression of Na+ and K+ transport-coding genes were recorded. Sakha108’s growth (18% rise from control) was significantly higher than Giza177’s. Both cultivars accumulated similar amounts of Na+ in the leaves and sheaths, however, Sakha108 had higher Na+ concentrations in the roots than Giza177 (13.3% higher). Root K+ concentration dropped dramatically (~ 2-fold reduction) in Giza177 roots while remaining unchanged in Sakha108 roots. The concentrations of H2O2 (root) and MDA (leaf and root) were higher in Giza177 than in Sakha108, although the difference was not statistically significant. Proline and total flavonoid (TF) contents in Sakha108 roots were greater than those in Giza177 roots. The expression of OsHKT1;5 and OsHKT2;1 genes declined in both cultivars, whereas expressions of OsSOS1, OsNHX1, and OsHAK7 were induced in Sakha108 but, except for OsHAK7, were repressed in Giza177. Combined, these findings suggest that Sakha108 is more resistant to salt stress than Giza177 is, and that this variation in tolerance may have its origins in the root systems of the two cultivars. Thus, these adaptive traits in the root of Sakha 108 could be explored for engineering stress tolerance in susceptible but high-yielding rice cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Rice is one of the most essential staples worldwide (Bandumula 2017). Several research institutes have developed high-yielding and high-quality rice cultivars, leading to a significant increase in production to meet the growing population’s demand. To address the challenges of adverse environmental conditions and climate change that are decreasing crop yields worldwide, new cultivars are being generated (Fang et al. 2021). Screening high-yielding cultivars for resistance to changing conditions is crucial for sustainable crop production. Failing to do so will hinder the development of these cultivars. The present study aims to fill the gap in knowledge regarding new and important rice cultivars that are bred.
Salinity is among the most important environmental stressors affecting crops globally (Munns and Tester 2008). It is particularly problematic in desert areas (including Egypt), where rainfall is limited, and the water requirements of plants are met via surface irrigation (Shahid et al. 2018). This method of field irrigation in the absence of rainfall often leads to water evaporation and the gradual build-up of salts in the irrigation water on the surface of the soil, leading to soil salinization (Shahid et al. 2018). Thus, it is necessary to screen emerging cultivars of crop plants, including rice, for salt stress tolerance to identify and disseminate planting material that will produce the expected yield even in salt-affected lands. Salinity affects plant growth and development by obstructing various physiological processes; for instance, salinity causes a more negative external solute potential than that of the root cells, which impacts soil water absorption, which further causes closed stomata and altered transpiration, gas exchange, photosynthesis, and ultimately biomass production (Munns and Tester 2008). This inability to absorb water is often associated with mineral deficiency as plants depend on the water uptake route for nourishment (Chuamnakthong et al. 2019; Sriskantharajah et al. 2020). In addition, uncontrolled Na+ accumulation in leaves often compromises cellular processes including enzyme deactivation as it replaces K+ on the active sites of these enzymes (Wang and Wu 2013). Furthermore, stress often generates reactive oxygen species (ROS) that inflict damage on many cellular components including cell membranes, proteins, and DNA, leading to cell death in extreme situations. Thus, to thrive under saline conditions, plants must be able to avert one or more of these aforementioned conditions.
These adaptive mechanisms include the synthesis and accumulation of osmolytes, such as proline to balance the osmotic potential required for sustaining water and mineral uptake from the soil by the roots. Plants would also have to be able to counter excess ROS production and accumulation by raising their antioxidant defense system, which is enzymatic (for example, catalase (CAT) and ascorbate peroxidase, (APX) and non-enzymatic (e.g., flavonoids, phenolics, proline, and glycine betaine) (Gill and Tuteja 2010). These antioxidants scavenge ROS, thereby protecting cellular structures and maintaining cellular homeostasis. Furthermore, to limit excess Na+ uptake and translocation to photosynthetic tissues, the induction of the expression of Na+ excluding (salt overly sensitive 1 (SOS1), retrieving (high-affinity K+ transporter, HKT1;5), and sequestering (Na+/H+ exchanger, NHX1) genes is required, whereas the expression of those involved in Na+ uptake and xylem Na+ loading (HKT2;1, and SOS1, respectively) should be repressed (Assaha et al. 2017b; Britto and Kronzucker 2015; Cotsaftis et al. 2012; Munns et al. 2012; Nieves-Cordones et al. 2016a). Moreover, one of the leading salt stress tolerance mechanisms is maintaining a low cytosolic Na+/K+ ratio (Anschutz et al. 2014; Himabindu et al. 2016), implying that for plants to tolerate salt stress, K+ uptake (via K+ transporters including the high-affinity K+ uptake transporters, AKT1 and HAK7) should be favored over Na+. Failure to do this will create an imbalance that will significantly affect cellular metabolism, as K+ is a co-factor of many enzymes. The replacement of K+ by Na+ may lead to impaired functions (Anschutz et al. 2014). Therefore, the regulation of Na+ transport in the root and the maintenance of healthy metabolic functions in the root will determine the plant’s overall health, as the root will ensure water and mineral uptake on which plants depend for proper growth, development, and yield.
Rice is a staple in Egypt, and its cultivation is carried out on 20% of the cultivated land (~ 0.65 million ha). This places Egypt as the top rice producer in Africa and the Near East region (Elbasiouny and Elbehiry 2020). The major area of rice cultivation is the Nile Delta region in the North of Egypt, where intrusion of saline water into agricultural lands and groundwater is common. The frequent and uncontrolled use of low-quality water (saline water) for irrigation is leading to soil salinization, which is a major challenge to rice cultivation in Egypt, claiming 35% of agricultural lands (Elbasiouny and Elbehiry 2020; Kotb et al. 2000; Zidan and Dawoud 2013). This poor irrigation practice is expected to reduce rice yields by 25% by 2025 (Elbasiouny and Elbehiry 2020). The quest for high-yielding rice cultivars in Egypt has been on the rise, and the rice cultivars, Giza 177 and Sakha 108, are among the new hybrids being developed at the Sakha Research Center of Field Crops, Kafrelsheikh, Egypt. Giza 177 has been shown to be sensitive to salt stress based on the field and greenhouse experiments (Abdallah et al. 2016; Alharbi et al. 2022; Zayed et al. 2014). The sensitivity has been attributed to high oxidative stress damage, high Na+ accumulation in the leaf blade, reduced photosynthetic pigment concentration, and low antioxidant enzyme activity (Alharbi et al. 2022). In addition, transcriptomic analysis of Giza 177 plants under salt stress revealed a downregulation of genes involved in oxidative stress defense, as well as of peroxidase genes (Shehab et al. 2022), implying that oxidative stress damage may be the primary cause of salt stress susceptibility in this plant. However, other stress adaptive pathways, such as Na+ and K+ transport and transporter regulators, have not been established for the cultivar. Sakha 108, on its part, is a high-yielding cultivar that is equally sensitive to salt stress, based on long-term exposure to salt stress in the field (Negm et al. 2023); however, the physiological and molecular basis for this sensitivity has not been elucidated yet, especially in the early development stage. Therefore, the present research was conducted to evaluate the impacts of salt stress on the two cultivars. This will help identify the possible causes behind salt tolerance or sensitivity in these rice cultivars, guiding breeding and disseminating genetic resources that will be both high-yielding and tolerant to adverse environmental conditions.
2 Materials and Methods
2.1 Plant Material, Growth Conditions, and NaCl Treatment
The two rice cultivars (Sakha 108 and Giza 177) were obtained from the Sakha Research Center of Field Crops in Kafrelsheikh, Egypt. These genotypes were selected based on a preliminary screening experiment. In the experiment, several commercial Egyptian rice cultivars including Sakha (102, 103, 105, 106, 107 and 108) and Giza (177, 178, 181 and 182) accessions under increasing levels of salt stress (0, 25, 50, 75, and 100 mM of NaCl) in a hydroponic culture at the seedling stage (21-day-old seedlings) for two weeks. Then the most salt tolerant (Sakha 108) and the most salt sensitive (Giza 177) ones (under 75 mM of NaCl) were selected for further comparative analysis. The choice was based on the phenotype of seedling, dry weights, and Na+ and K+ contents. Thereafter, the seeds of the two cultivars were pretreated and then germinated as described earlier (Mekawy et al. 2015, 2018). After germination, uniformly germinated seeds were transferred to a nylon mesh suspended over 20 L of a half-strength Kimura B nutrient solution, whose composition was previously described (Mekawy et al. 2018). At day 28, the seedlings were transferred to either control (Kimura B nutrient solution) or salinity (Kimura B nutrient solution + 75 mM NaCl) conditions for 12 days. The salt concentration was attained by dissolving the required amounts of NaCl salt in the growth medium. Each treatment tank had 16 seedlings, eight from each cultivar. To avoid osmotic shock, the treatments were applied incrementally in the order of 25 mM per day up to the final concentration (75 mM NaCl). Every 2 d, the solutions (pH 5.0–6.0) were replaced. Seedlings were raised and experimentally treated in a greenhouse at Hiroshima University, Japan, under natural conditions. The experiment was designed to assess salt stress effects in the two cultivars at the seedling stage.
2.2 Growth Measurements
At the end of salinity treatment, 4 plants of each cultivar (representing 4 replicates) were taken out of the growth media and divided into roots, leaf sheaths, and leaf blades, and split into two sets of samples. One set of plants was oven-dried at 70 °C for three days, and the dry weights (DW) were recorded. The other set was maintained at − 80 °C for physiological analyses.
2.3 Determination of Na + and K + Concentrations
The concentrations of Na+ and K+ ions in the three parts of the plants were measured using a flame photometer (ANA-135; Tokyo Photoelectric, Tokyo, Japan) following a previously described method (Kushizaki 1968). The dried samples (0.23–0.56 g of root, 0.67–1.52 g of sheath and 0.6–1.2 g of leaf tissues) were soaked in 1 N HCl overnight, and the concentrations were determined from different doses of Na+ and K+ concentrations to establish standard curves.
2.4 Determination of Free Proline, Total Phenolics (TP), and Total Flavonoid (TF) Concentrations
Proline concentration was measured as a previously described method (Bates et al. 1973) with some modifications. Proline was extracted from 500 mg of fresh leaf or root tissues in 10 mL of 3% sulfosalicylic acid. To 2 mL of the extract was added 2 mL of ninhydrin reagent containing glacial acetic acid and incubated at 100 °C for 2/3 h. Then, the resultant reaction mixture was rapidly cooled in an ice-water bath. For chromophore development, 4 mL of toluene was added to the cooled samples and vigorously mixed. Finally, the chromophore layer was separated, and its absorbance measured at 520 nm. Total phenolic compounds were extracted and measured according to previously described methods (Assaha et al. 2017a). Briefly, 1 mL of extract and gallic acid were mixed with Na2CO3 and Folin-Ciocalteu reagent and incubated for 2 h. Then, the absorbance was read at 765 nm. Gallic acid was used as the standard, and the concentration of TP was expressed as gallic acid equivalents. The total flavonoids were extracted and measured according to the previously described method (Farajzadeh Memari-Tabrizi et al. 2021). Briefly, 0.5 mL of extract was mixed with 1.5 mL ethanol and 3 mL of a reaction mixture comAposed of 10% AlCl3 and 1 M potassium acetate. This mixture was then incubated at 25 oC for 30 min, and the absorbance was read at 415 nm. TF was expressed as quercetin equivalents.
2.5 Lipid Peroxidation (MDA) and Hydrogen Peroxide (H2O2) Determination
The thiobarbituric acid (TBA) reaction was employed to determine the malondialdehyde (MDA) concentration. One hundred (100) mg of fresh leaf and root tissues were ground in liquid nitrogen and MDA was extracted using extraction buffer (10 mM HEPES, pH 7, 15% trichloroacetic acid (TCA), 0.375% thiobarbituric acid, 0.25 N HCl, 0.04% butylated hydroxyl toluene (BHT), and 2% ethanol. The mixture was incubated at 95 oC and then centrifuged. The absorbance of the supernatant was read at 535 and 600 nm, and the MDA concentration was calculated using the extinction coefficient (155 mM− 1 cm− 1) as previously described (Assaha et al. 2015). The H2O2 extraction and measurement were performed as previously described (Mekawy et al. 2018).
2.6 Antioxidant Enzyme Activities
To measure the activities of CAT (EC 1.11.1.6) and APX (EC 1.11.1.11), enzyme extraction was performed using fresh leaf and root samples (0.5 g) according to previously described methods (Mekawy et al. 2018; Takagi and Yamada 2013). For CAT activity, 1 mL of assay mixture was composed of 5% enzyme extract, 50 mM potassium phosphate buffer (pH 7.0), and 10 mM H2O2. The consumption of H2O2 was monitored at 240 nm, and CAT activity was expressed as mmol H2O2 consumed per minute. For the APX activity, one mL of assay mixture comprised 10% enzyme extract, 25 mM phosphate buffer (pH 7.0), 0.25 mM ascorbic acid, 0.1 mM EDTA, and 0.1 mM H2O2. The ascorbate oxidation was followed at 290 nm, and the activity was calculated using 2.8 mM− 1cm− 1 as the extinction coefficient. A unit of APX was determined to be 1 µM of ascorbate oxidized per minute. The protein concentration in the enzyme extract was measured using a protein assay kit (Bio-Rad DC, California, USA) with bovine serum albumin as standard, according to the manufacturer’s instructions.
2.7 Gene Expression Analysis
Collection of materials for gene expression analysis (leaves and roots) was carried out in the morning of the sampling day under natural conditions in the greenhouse (60–70% humidity and 25–30/20–25 oC day/night temperature). Total RNA was extracted from the roots and shoots of control and stressed plants using a total RNA Extraction Kit (Plant) (RBC Bioscience, SciTrove, Japan) according to the manufacturer’s instructions. One µg of the extracted total RNA was reverse transcribed to cDNA using the ReverTra Ace qPCR RT kit (Toyobo, Osaka, Japan). Primer design and specificity and quantitative real-time polymerase chain reaction (qRT-PCR) were conducted as previously described (Mekawy et al. 2015, 2018), using NCBI tools and Thunderbird SYBR qPCR Mix (Toyobo) and the ABI Step One Plus system (Applied Biosystems), respectively. The expression levels of the Oryza sativa tonoplast Na+/H+ exchanger (OsNHX1), high-affinity K+ transporters OsHKT1;5, OsHKT2;1, OsHAK7, OsAKT1, and salt overly sensitive 1 OsSOS1 genes were calculated using the 2−ΔΔCT method (Livak and Schmittgen 2001), using the OsUBQ5 gene as reference. Table 1 contains all the primer sequences used in these analyses. The melting curve analysis was used to verify the PCR products.
2.8 Statistical Analyses
Analysis of variance (ANOVA) tests were run using SPSS statistical software package version 21 (IBM Inc., USA). The Duncan’s multiple range test was used to separate means (n = 4 for physiological parameters and n = 3 for gene expression analysis) at α = 0.05.
3 Results
3.1 Effect of Salt Stress on the Growth (Biomass Accumulation) in Giza 177 and Sakha 108 at the Seedling Stage
After subjecting 28-day-old seedlings of Giza 177 and Sakha 108 to salt stress (75 mM NaCl) for 12 days, the dry mass of the leaves stems (sheaths), and roots were measured and are presented in Table 2. In Giza 177, the DW of leaves significantly decreased (23% reduction) under salt stress when compared with control plants, whereas the stress did not significantly alter the stem and root DWs. In Sakha 108 on the contrary, while salt stress significantly increased the stem and root DW (33 and 20%, respectively), it did not significantly affect the DW of leaves. In terms of total DW, salt stress reduced the biomass of Giza 177 (12%), although this reduction was not statistically significant, whereas the stress significantly increased the biomass of Sakha 108 (18% increase). Aside from changes in dry mass, the salt concentration and duration of the treatment did not induce any visible sign of salt stress injury on the plants.
3.2 Effect of Salt Stress on Na+ and K+ Concentration and Na+/K+ Ratio in Different Organs of Giza 177 and Sakha 108 at the Seedling Stage
The effects of salt stress on Na+ and K+ concentrations and Na+/K+ ratio are presented in Fig. 1. The Na+ concentration was lower in the leaves of both cultivars under salt stress, with no significant difference between them. The concentration was higher in the roots and sheaths, with a significantly higher concentration in the roots of Sakha 108 (13% higher) than in the roots of Giza 177. The concentration in the sheaths was not significantly different between the two cultivars. With regard to K+ (Fig. 1B), the concentration in root significantly decreased (1.9-fold reduction) in root Giza 177 but remained unaltered in Sakha 108, compared to the control. The K+ concentration in the sheath significantly decreased for both cultivars, although this reduction was more pronounced in Giza 177 (2-fold reduction) than in Sakha 108 (1.6-fold reduction). The K+ concentration in the leaves remained unchanged in Sakha 108 but decreased in Giza 177, although this decrease was not statistically significant. In terms of the Na+/K+ ratio (Fig. 1C), the sheaths and leaves of both cultivars had statistically similar ratios, whereas in the root, although the ratio markedly increased for both cultivars, that of Giza 177 was significantly higher (2.4-fold increase) than that of Sakha 108 (1.9-fold increase).
3.3 Effect of Salt Stress on the Concentration of Free Proline, TF, and TP of Leaves and Roots of Giza 177 and Sakha 108 at the Seedling Stage
The effects of NaCl on proline, total phenolics, and total flavonoid concentration in the leaves and roots of Giza 177 and Sakha 108 are shown in Fig. 2. The proline concentration significantly increased in the leaves and roots of both cultivars (Fig. 2A, B); however, this concentration increased more in the leaves of Giza 177 (3.1-fold) than in Sakha 108 (2-fold), whereas in the root, the concentration was higher in Sakha 108 (4-fold) than in Giza 177 (2.3-fold), compared with the control. The concentration of TF in the leaves of Sakha 108 significantly increased under salt stress (42% increase), while that of Giza 177 did not significantly alter (13% increase), compared with controls (Fig. 2C, D). The TP concentrations in the leaves and roots of both cultivars remained statistically unaltered compared with the control (Fig. 2E, F).
Effect of salt stress on the concentrations of proline (A and B), total flavonoids (TF, C and D), and total phenolics (TP, E and F) in the leaf blades and roots, respectively of Giza 177 and Sakha 108. Bars represent means ± SE (n = 4), and bars with the same letter are not statistically different (p ≤ 0.05)
3.4 Effect of Salt Stress on the Concentration of MDA and H2O2 in Leaves and Roots of Giza 177 and Sakha 108 at the Seedling Stage
The effects of salt stress on MDA and H2O2 concentrations are shown in Fig. 3. In the leaves, the MDA concentration significantly increased under salt stress in both cultivars, with Giza 177 exhibiting a higher concentration (33% increase) than Sakha 108 (28%), compared with the control. However, in the roots, Sakha 108 had a higher concentration (18% increase) than Giza 177 (6%), compared with the control, although the increase in Giza 177 was not statistically significant (Fig. 3B). Whereas H2O2 concentration was not statistically different in the leaves of both cultivars (Fig. 3C), it increased in the roots of Giza 177 (8% increase) but was not significantly altered in Sakha 108 (3% increase), compared with control (Fig. 3D).
3.5 Effect of Salt Stress on Antioxidant Enzyme Activity in Leaves and Roots of Giza 177 and Sakha 108 at the Seedling Stage
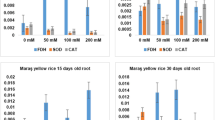
The effects of salt stress on CAT and APX activities are shown in Fig. 4. CAT activity in the leaves of both cultivars did not change significantly under salt stress compared to that in the controls (Fig. 4A). However, in the root, the CAT activity markedly decreased in Giza 177 (50% reduction) but significantly increased in Sakha 108 (36% increase), when compared with controls (Fig. 4B). APX activity, on the other hand, was significantly elevated in the leaves of both cultivars, although the activity was higher in Sakha 108 (36%) than in Giza 177 (20% increase), compared with controls (Fig. 4C). In the root (Fig. 4D), APX activity remained statistically unaltered in Giza 177, but the activity significantly increased in Sakha 108 (56% increase) compared to the control.
3.6 Effect of Salt Stress on the Expression of Na+ and K+ Transport Genes in the Leaves and Roots of Giza 177 and Sakha 108 at the Seedling Stage
The expression of the Na+ or K+ transport genes OsNHX1, OsHAK7, and OsAKT1 in the leaves and OsHKT1;5, OsHKT2;1, OsNHX1, OsSOS1, OsHAK7, and OsAKT1 in the roots of Giza 177 and Sakha 108, are presented in Figs. 5 and 6, respectively. In the leaves, the expression of OsHAK7 and OsNHX1 was upregulated in Giza 177 (1.7 and 1.2-fold, respectively) but was downregulated in Sakha 108 (Fig. 5A, C). However, OsAKT1 was downregulated in the leaves of both cultivars (Fig. 5B). In the roots, the expression of OsHKT1;5, OsHKT2;1, and OsAKT1 was downregulated in both cultivars (Fig. 6A, B, E). Whereas OsSOS1 and OsNHX1 were downregulated in Giza 177, they were induced in Sakha 108 (1.8- and 1.1-fold increase, respectively) (Fig. 6C, D). OsHAK7 was more markedly induced in Sakha 108 (1.7-fold) than in Giza 177 (1.1-fold) (Fig. 6F).
Effect of salt stress on the expression of Oryza sativa high-affinity K+transporter 1;5 (OsHKT1;5, A), Oryza sativa high-affinity K+transporter 2;1 (OsHKT2;1, B), Oryza sativa salt overly sensitive 1 (OsSOS1C), Oryza sativa Na+/H+exchanger 1 (OsNHX1, D), Oryza sativa Arabidopsis K+transporter 1 (OsAKT1, E), and Oryza sativa high-affinity K+7 (OsHAK7, F) genes in the roots of Giza 177 and Sakha 108. Bars represent means ± SE (n = 3, p ≤ 0.05)
4 Discussion
Soil salinization due to increasing salinity threatens global food security as it reduces crop yields. Therefore, studying crop plants’ adaptive responses to this stress and finding ways to mitigate it is a priority in agriculture. In this light, the responses of two Egyptian rice cultivars (Giza 177 and Sakha 108), bred for high yield, to salt stress were evaluated to obtain information that will guide breeders in the dissemination of improved plants that can withstand adverse environmental conditions with fewer yield penalties. Thus, the two cultivars were subjected to salt stress of 75 mM NaCl in a hydroponic culture medium for 12 days. The biomass (dry weight) of both cultivars was analyzed and revealed that although stress negatively affected the growth of Giza 177, it promoted that of Sakha 108, especially the mass of sheaths and roots. This clearly shows that Sakha 108 is more tolerant to salt stress than Giza 177. Enhanced plant growth by salt stress is known in halophytes and tolerant glycophytes, including the tolerant rice cultivar Pokkali. In a pot experiment to evaluate the relative tolerance of 3 rice cultivars, including Pokkali, subjected to salt stress (50, 100, and 150 mM NaCl) for two weeks, it was found that the shoot biomass of Pokkali was not affected by 50, and 100 mM NaCl stress, and tended to increase, although not significantly (Pattanagul and Thitisaksakul 2008). In another pot experiment, Pokkali subjected to 5, 10, and 15 dS m− 1 salt stress, showed reduced shoot (13, 52, and 67% reduction) and root (50, 82.89, and 88.15% reduction) dry weight (DW) (Ologundudu et al. 2014). However, in the present study, the growth of Sakha 108 significantly increased under 75 mM NaCl in a hydroponic culture. This contrasts with our observation of Egyptian Yasmine in a previous study (Mekawy et al. 2015), which, although appeared more tolerant than Sakha 102, had reduced growth (leaf biomass) under salt stress, like the pattern in Giza 177. This enhanced growth under salt stress is a trait seen mostly in halophytes (salt-loving plants); for example, the halophyte, ice plant, showed significant increases in biomass when exposed to 100 mM NaCl (Farzana et al. 2023). This suggests that salt stress stimulates growth in Sakha 108 at the seedling stage, contrary to the tolerant rice cultivars.
Furthermore, the tolerance of Sakha 108 contrasts previous reports of its susceptibility to salt stress (Negm et al. 2023). This difference is attributed to prolonged exposure of the cultivar to salt stress in the field till harvesting, without considering the short-term effects of salt stress as we have done here in a hydroponic system. Therefore, it will be essential to investigate, through a time course experiment, the tolerance of this cultivar, to establish its limit of tolerance through the growth cycle. Nonetheless, this enhanced tolerance, as revealed by the unusually significant increase in biomass, should normally place Sakha 108 in the class of tolerant rice cultivars, and a possibility of superior tolerance to the well-known tolerant cultivars (Nona Bokra and Pokkali) may be expected. Therefore, it would be interesting if further studies are conducted to investigate its growth alongside these established tolerant cultivars, and to understand more of the underlying mechanisms that confer salt stress tolerance, relevant for enhanced rice production in adverse environments.
To thrive in saline environments, Na+ exclusion from the leaf blade is one of the main adaptive strategies used by glycophytes (salt-sensitive plants), including rice (Assaha et al. 2017b; Munns and Tester 2008). In this strategy, regulatory mechanisms restrict the uptake and translocation of Na+ to the leaf blade. These include Na+ efflux from cells to the external environment of the root epidermal cells via SOS1 (salt overly sensitive 1), the absorption of Na+ from the xylem via SOS1 (Olías et al. 2009), HKT1;5 (high-affinity K+ transporter 1;5) and HKT1;4 (Cotsaftis et al. 2012), the compartmentation of Na+ into vacuoles of root cells, the recirculation of Na+ via the phloem for SOS1-mediated efflux (Fujimaki et al. 2015), and the repression of Na+ influx transporter genes (e.g. OsHKT2;1) (Mekawy et al. 2018). In this study, the two cultivars demonstrated the ability to exclude sodium ions (Na+) from the leaf blade. The mechanisms responsible for this exclusion likely occurred in the roots and sheaths, where the majority of absorbed sodium ions accumulated. In rice, OsHKT1;5 and OsHKT1;4 (root and sheath, respectively) proteins are implicated in Na+ extraction from the xylem for storage in the root and sheath (Cotsaftis et al. 2012), thus limiting the buildup of Na+ in the leaf blade. From the Na+ accumulation pattern in the present study, these two proteins should be implicated, although this was not reflected in the expression of OsHKT1;5 in the roots of both cultivars, as gene expression does not always correlate with protein abundance (Buccitelli and Selbach 2020). This HKT1;5 expression in the present study contrasts the observation in another Egyptian rice cultivar, Egyptian Yasmine, which was strongly induced and coincided with reduced Na+ delivery to the leaf blade of the cultivar and hence more tolerance over Sakha 102 (Mekawy et al. 2015). In addition, the repression of OsHKT2;1 would also indicate regulated Na+ influx in the roots of both cultivars, as has been observed in the rice cultivars Koshihikari and Egyptian Yasmine (Mekawy et al. 2018).
Despite the ability to exclude Na+ from the leaf blades of both cultivars, Sakha 108 exhibited significantly higher Na+ in the roots than Giza 177. For proper metabolic functions of root cells, Na+ has to be extruded via plasma membrane-bound SOS1 or sequestered into vacuoles via the tonoplast-bound NHX1 (Na+/H+ exchanger 1) for osmotic adjustment (Munns et al. 2020). To ascertain the involvement of these two Na+ transport systems in salt stress adaptation in the two cultivars, the expression of OsNHX1 and OsSOS1 in the roots of both cultivars were analyzed, and the results revealed that both genes were repressed in the roots of Giza 177 but were upregulated in the roots of Sakha 108. This implies that Giza 177 cannot effectively sequester Na+ into vacuoles, and in the absence of SOS1 function, excess Na+ accumulation in the cytosol can occur, damaging many cellular processes (Zhang et al. 2018). This vacuolar Na+ sequestration would be effective in Sakha 108, thereby preserving cell integrity by improving the water uptake and sustaining the cultivar’s growth under salt-stress conditions.
Similarly, the tolerant Egyptian Yasmine cultivar showed upregulated SOS1 expression in the root, which is essential for Na+ extrusion; however, it happened in the absence of NHX1 expression (Mekawy et al. 2015), indicating that Sakha 108 in the present study is more equipped to deal with excess Na+ in the root than Egyptian Yasmine. Furthermore, this Egyptian Yasmine was tested under a lower NaCl concentration (50 mM) compared to Sakha 108 (75 mM) in the present study. Moreover, Egyptian Yasmin accumulated more Na+ in the leaf blade (15 mg/g DW) than the two cultivars (~ 4 mg/g DW) in the present study, showing that the roots of Sakha 108 have more enhanced mechanisms to regulate Na+ delivery to the aerial parts of the plants. One of such mechanisms is vacuolar Na+ sequestration which may be operating in the root of Sakha 108 resulting from enhanced root NHX1 expression, which will reduce Na+ delivery to the shoot, as well as serve in osmotic adjustment to maintain water uptake. These must have contributed to enhanced root growth in Sakha108 when compared with that of Giza177 which had repressed root NHX1 and consequently unaltered root growth.
When plants can control the uptake and accumulation of Na+, the uptake, translocation, and accumulation of K+ for enhanced metabolic functions is assured. K+ is the most abundant macro-element in plants accounting for up to 10% of plant dry mass (Ahmad and Maathuis 2014). It is involved in multiple plant metabolic processes including guard cell volume regulation, enzyme activation, germination, osmoregulation, nyctinasty movement of leaves, stomatal regulation, loading and unloading of sugars in the phloem, as a counterion for nitrate translocation, membrane potential regulation, cytosolic pH regulation, and intracellular protein trafficking (Anschutz et al. 2014; Assaha et al. 2017b; Himabindu et al. 2016). Uncontrolled uptake and accumulation of Na+ often replace K+ functions in plants, thereby compromising them, including replacing K+ in enzyme-binding sites, leading to their deactivation (Nieves-Cordones et al. 2016b). Hence, maintaining a high cytosolic K+/Na+ ratio has been identified as a key salt stress tolerance strategy (Anschutz et al. 2014; Himabindu et al. 2016), which can be reflected in the tissue Na+/K+ ratios. To determine the effect of salt stress on K+ homeostasis, the K+ concentration in plant parts, as well as the expression of genes coding K+ transporters [OsHAK7 (Oryza sativa high-affinity K+ transporter) and OsAKT1 (Oryza sativa Arabidopsis K+ transporter)] in leaves and roots were measured. The results showed that although leaf K+ concentration was not altered in Sakha 108, it declined in Giza 177. Surprisingly, a sharp decline in root K+ of Giza (50% reduction) compared to the unaltered concentration in Sakha 108 was observed. This implies that Na+ uptake in the roots of Giza 177 is favored over K+ uptake. OsAKT1 was repressed in the roots of both cultivars and was almost silenced in Giza 177, whereas OsHAK7 was only induced in the roots of Sakha 108, implying that salt stress compromised the K+ uptake transport system in favor of Na+ uptake. It has been shown that many HAKs are permeable to Na+ (Takahashi et al. 2007; Wang et al. 2015) and so under conditions of high salinity will favor Na+ over K+ transport indicating that this may be the case in Giza 177.
HAKs are high-affinity K+/H+ transporters that require the activity of H+-ATPase to function properly. The activity of this transporter is inhibited because excess Na+ often depolarizes the plasma membrane to values higher than the equilibrium potential of K+. This depolarization then leads to the activation of outward rectifier K+ channels, consequently resulting in K+ efflux (Bacha et al. 2015; Pottosin and Dobrovinskaya 2014). In addition, SOS1 is integral to the uptake of K+ as it helps stimulate H+-ATPase to pump out more protons, thus creating a pmf that is used to energize Na+ efflux from the cytosol, which would otherwise impair AKT1 via plasma membrane depolarization (Horie et al. 2012; Mansour 2014). Therefore, the low root K+ and consequently higher Na+/K+ ratio in Giza 177 is the result of a defect in Na+ efflux via SOS1 (repressed) and sequestration via NHX1 (repressed), leading to excess cytosolic Na+ that induces plasma membrane depolarization thereby impairing the function of K+ uptake transporters, and hence, impaired K+ uptake. The low K+ could also result from ROS-induced K+ efflux (Shahzad et al. 2022), as H2O2 increased in the roots of Giza 177 without an antioxidant defense system. In fact, (Liu et al. 2019) found markedly higher H2O2-induced K+ efflux in the root of the salt sensitive rice cultivar Koshihikari, when compared with the tolerant Reizig cultivar. This indicates that K+ efflux may significantly contribute to salt stress susceptibility in sensitive rice cultivars including Giza 177. Thus, Sakha 108 had more control over Na+ and K+ uptake and accumulation than Giza 177, which probably affected water uptake and transport to the leaves for normal growth, the reason for which there is marked leaf mass reduction in the cultivar. This clearly shows that the overall performance of both cultivars depended on Na+ and K+ homeostatic mechanisms in the root. Their defects lead to poor growth, as in Giza177, while their enhancement leads to vigorous growth as in Sakha 108.
Besides shoot Na+ exclusion, another strategy employed by plants to combat salt stress is deploying the antioxidant defense system (antioxidant enzymes and non-enzymes) to combat a surge in ROS (reactive oxygen species) generated via the oxidation of oxygen molecules by electrons that have escaped from cellular electron transport chains. These ROS damage several cellular components, including plasma membranes and nucleic acids (Gill and Tuteja 2010). Hence for plants to survive salt stress, they must be able to bring ROS (e.g., H2O2) levels within tolerable ranges, as observed by the level of MDA (malondialdehyde) concentration (indicator of lipid peroxidation). Thus, the activity of APX and CAT, which are the most effective in H2O2 detoxification (Gill and Tuteja 2010), and the non-enzyme antioxidants proline, TP and TF, is important. The present study revealed that the H2O2 (in roots) and MDA (in roots and leaves) concentrations of Giza 177 were higher than those of Sakha 108 under salt stress, although the difference was not statistically significant, indicating that the ability to control H2O2 accumulation in Giza 177 is less effective than in Sakha 108. This is evident in the activity of APX and CAT in the roots of both plants; whereas the activity of both enzymes was enhanced in Sakha 108, the activities either declined (CAT) or remained unaltered (APX) in Giza 177. This observation is similar to that of Liu et al. (2019), who found significantly reduced CAT and APX activities in the root of the sensitive rice cultivar Koshihikari, when compared with the tolerant cultivar, suggesting that weakened antioxidant defense leading to noxious levels of ROS, is a contributing factor to salt stress susceptibility in sensitive rice cultivars. Similarly, the concentrations of proline and TF significantly increased in Sakha 108, whereas they either increased (proline, lower than in Sakha) or significantly declined (TF) in Giza 177. However, TP remained unaltered in either cultivar, indicating that it is not an important stress tolerance factor in both cultivars. Thus, Sakha 108 is better equipped than Giza 177 to combat the oxidative stress induced by salt stress, and the root antioxidant system is probably central to this stress defense.
Furthermore, enhanced proline levels not only protect against oxidative stress damage but also suggest control of potassium ion (K+) efflux. Proline has been shown to reduce hydroxyl radical (OH•)-induced K+ efflux from roots (Demidchik et al. 2010). This implies that, due to its higher concentration, Sakha 108 exerts more influence on this phenomenon than Giza 177. Proline is an osmolyte that participates in osmotic adjustment to enable water uptake under salt stress (Hmidi et al. 2018), although, in some plants, proline accumulation is only a symptom of stress and not a defense strategy (Mansour and Ali 2017). As such, its synthesis and accumulation can lead to growth reduction as energy for growth is diverted to organic solute synthesis (Munns et al. 2020). This is probably the case for Giza 177, which had a higher proline concentration in the leaves. The enhanced accumulation of proline in the roots of Sakha 108, coupled with the possibility of vacuolar Na+ compartmentation, is essential to counter the osmotic potential difference with the soil, thus favoring water uptake for better growth over Giza 177. The accumulation of proline, especially in the early stages of growth, has been shown to be positively correlated with salt stress tolerance in both cultivated and wild rice cultivars (Chutipaijit et al. 2011; Nguyen et al. 2021), like what is here observed in Sakha 108. However, the contrary is noticed with Giza 177, suggesting that proline accumulation in this cultivar is simply a response (symptom)to the stress rather than a defense strategy.
5 Conclusions
In conclusion, Sakha 108 is more tolerant to salt stress than Giza 177 at the seedling stage based on dry biomass, as salt stress significantly increased the growth of Sakha 108 but did not significantly alter that of Giza 177. Although the two cultivars display similar leaf blade Na+ exclusion abilities (~ 5 mg/g dry weight, compared to > 20 mg/g dry weight in root or sheath) as tolerance strategy, the differences between them are associated with distinct responses occurring in the root including (1) the ability to regulate the uptake and accumulation of Na+ in the root, which is absent in Giza 177 following downregulation of Na+ efflux (repressed Oryza sativa salt overly sensitive 1, OsSOS1) and vacuolar compartmentation (repressed Oryza sativa Na+/H+exchanger 1, OsNHX1); (2) the ability to maintain K+ uptake, which is defective in Giza 177 owing to sharp decline in K+ concentration (2-fold reduction from control levels) following downregulation of Oryza sativa high-affinity K+7 (OsHAK7), as well as possible impairment of Arabidopsis K+ transporter 1 (AKT1) due to excess Na+ accumulation, which in the absence of an efflux transporter (SOS1), will depolarize membranes leading to K+ efflux instead of influx; (3) oxidative stress is more prominent in Giza 177 than Sakha 108, following more reduced enzyme and non-enzyme antioxidants; and (4) osmotic adjustment in Sakha 108 following higher proline accumulation and possible vacuolar Na+ sequestration, which is possibly absent in Giza 177. Hence, Giza 177 in addition to being susceptible at the reproductive stage (reduced yield) as mentioned earlier, is equally susceptible to salt stress at the seedling stage, whereas Sakha 108 as earlier mentioned is susceptible at the reproductive stage under prolonged exposure, but as reported here is tolerant to salt stress at the seedling stage. The tolerance mechanism in Sakha 108 could therefore be exploited to induce stress tolerance in other important but susceptible rice cultivars.
References
Abdallah MMS, Abdelgawad ZA, El-Bassiouny HMS (2016) Alleviation of the adverse effects of salinity stress using trehalose in two rice varieties. S Afr J Bot 103:275–282. https://doi.org/10.1016/j.sajb.2015.09.019
Ahmad I, Maathuis FJ (2014) Cellular and tissue distribution of potassium: physiological relevance, mechanisms and regulation. J Plant Physiol 171:708–714. https://doi.org/10.1016/j.jplph.2013.10.016
Alharbi K, Osman HS, Rashwan E, Hafez EM, Omara AE (2022) Stimulating the growth, anabolism, antioxidants, and yield of rice plants grown under salt stress by combined application of bacterial inoculants and nano-silicon. Plants 11:3431. https://doi.org/10.3390/plants11243431
Anschutz U, Becker D, Shabala S (2014) Going beyond nutrition: regulation of potassium homoeostasis as a common denominator of plant adaptive responses to environment. J Plant Physiol 171:670–687. https://doi.org/10.1016/j.jplph.2014.01.009
Assaha DVM, Liu LY, Mekawy AMM, Ueda A, Nagaoka T, Saneoka H (2015) Effect of salt stress on na accumulation, antioxidant enzyme activities and activity of cell wall peroxidase of Huckleberry (Solanum scabrum) and eggplant (Solanum melongena). Int J Agric Biol 17:1149–1156. https://doi.org/10.17957/Ijab/15.0052
Assaha DVM, Mekawy AMM, Liu L, Noori MS, Kokulan KS, Ueda A, Nagaoka T, Saneoka H (2017a) Na+ retention in the root is a key adaptive mechanism to low and high salinity in the glycophyte, Talinum paniculatum (Jacq.) Gaertn. (Portulacaceae). J Agron Crop Sci 203:56–67. https://doi.org/10.1111/jac.12184
Assaha DVM, Ueda A, Saneoka H, Al-Yahyai R, Yaish MW (2017b) The role of Na+ and K+ transporters in salt stress adaptation in glycophytes. Front Physiol 8:509. https://doi.org/10.3389/fphys.2017.00509
Bacha H, Rodenas R, Lopez-Gomez E, Garcia-Legaz MF, Nieves-Cordones M, Rivero RM, Martinez V, Botella MA, Rubio F (2015) High Ca2+ reverts the repression of high-affinity K+ uptake produced by Na+ in Solanum Lycopersycum L. (var. Microtom) plants. J Plant Physiol 180:72–79. https://doi.org/10.1016/j.jplph.2015.03.014
Bandumula N (2017) Rice production in Asia: Key to global food security. Proc Natl Acad Sci India Sect B Biol Sci 88:1323–1328. https://doi.org/10.1007/s40011-017-0867-7
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207. https://doi.org/10.1007/Bf00018060
Britto DT, Kronzucker HJ (2015) Sodium efflux in plant roots: what do we really know? J Plant Physiol 186–187:1–12. https://doi.org/10.1016/j.jplph.2015.08.002
Buccitelli C, Selbach M (2020) mRNAs, proteins and the emerging principles of gene expression control. Nat Rev Genet 21:630–644. https://doi.org/10.1038/s41576-020-0258-4
Chuamnakthong S, Nampei M, Ueda A (2019) Characterization of Na+ exclusion mechanism in rice under saline-alkaline stress conditions. Plant Sci 287:110171. https://doi.org/10.1016/j.plantsci.2019.110171
Chutipaijit S, Cha-um S, Sompornpailin K (2011) High contents of proline and anthocyanin increase protective response to salinity in Oryza sativa L. spp. indica. Aust J Crop Sci 5:1191–1198
Cotsaftis O, Plett D, Shirley N, Tester M, Hrmova M (2012) A two-staged model of Na+ exclusion in rice explained by 3D modeling of HKT transporters and alternative splicing. PLoS ONE 7:e39865. https://doi.org/10.1371/journal.pone.0039865
Demidchik V, Cuin TA, Svistunenko D, Smith SJ, Miller AJ, Shabala S, Sokolik A, Yurin V (2010) Arabidopsis root K+-efflux conductance activated by hydroxyl radicals: single-channel properties, genetic basis and involvement in stress-induced cell death. J Cell Sci 123:1468–1479. https://doi.org/10.1242/jcs.064352
Elbasiouny H, Elbehiry F (2020) Rice production in Egypt: The challenges of climate change and water deficiency. In: Ewis Omran, ES., Negm, A. (eds) Climate change impacts on agriculture and food security in Egypt. https://doi.org/10.1007/978-3-030-41629-4_14
Fang S, Hou X, Liang X (2021) Response mechanisms of plants under saline-alkali stress. Front Plant Sci 12:667458. https://doi.org/10.3389/fpls.2021.667458
Farajzadeh Memari-Tabrizi E, Yousefpour-Dokhanieh A, Babashpour-Asl M (2021) Foliar-applied silicon nanoparticles mitigate cadmium stress through physio-chemical changes to improve growth, antioxidant capacity, and essential oil profile of summer savory (Satureja hortensis L). Plant Physiol Biochem 165:71–79. https://doi.org/10.1016/j.plaphy.2021.04.040
Farzana T, Guo Q, Rahman MS, Rose TJ, Barkla BJ (2023) Salinity and nitrogen source affect productivity and nutritional value of edible halophytes. PLoS ONE 18:e0288547. https://doi.org/10.1371/journal.pone.0288547
Fujimaki S, Maruyama T, Suzui N, Kawachi N, Miwa E, Higuchi K (2015) Base to tip and long-distance transport of sodium in the root of Common Reed [Phragmites australis (Cav.) Trin. Ex Steud.] At steady state under constant high-salt conditions. Plant Cell Physiol 56:943–950. https://doi.org/10.1093/pcp/pcv021
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930. https://doi.org/10.1016/j.plaphy.2010.08.016
Himabindu Y, Chakradhar T, Reddy MC, Kanygin A, Redding KE, Chandrasekhar T (2016) Salt-tolerant genes from halophytes are potential key players of salt tolerance in glycophytes. Environ Exp Bot 124:39–63. https://doi.org/10.1016/j.envexpbot.2015.11.010
Hmidi D, Abdelly C, Athar HU, Ashraf M, Messedi D (2018) Effect of salinity on osmotic adjustment, proline accumulation and possible role of ornithine-delta-aminotransferase in proline biosynthesis in Cakile maritima. Physiol Mol Biol Plants 24:1017–1033. https://doi.org/10.1007/s12298-018-0601-9
Horie T, Karahara I, Katsuhara M (2012) Salinity tolerance mechanisms in glycophytes: an overview with the central focus on rice plants. Rice 5:11. https://doi.org/10.1186/1939-8433-5-11
Kotb THS, Watanabe T, Ogino Y, Tanji KK (2000) Soil salinization in the Nile Delta and related policy issues in Egypt. Agric Water Manag 43:239–261. https://doi.org/10.1016/S0378-3774(99)00052-9
Kushizaki M (1968) An extraction procedure of plant materials for the rapid determination of Mn, Cu, zn and mg by the atomic absorption analysis. J Sci Soil Manure 39:489–490
Liu J, Shabala S, Shabala L, Zhou M, Meinke H, Venkataraman G, Chen Z, Zeng F, Zhao Q (2019) Tissue-specific regulation of Na+ and K+ transporters explains genotypic differences in salinity stress tolerance in rice. Front Plant Sci 10:1361. https://doi.org/10.3389/fpls.2019.01361
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-∆∆CT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Mansour MM (2014) The plasma membrane transport systems and adaptation to salinity. J Plant Physiol 171:1787–1800. https://doi.org/10.1016/j.jplph.2014.08.016
Mansour MMF, Ali EF (2017) Evaluation of proline functions in saline conditions. Phytochem 140:52–68. https://doi.org/10.1016/j.phytochem.2017.04.016
Mekawy AM, Assaha DV, Yahagi H, Tada Y, Ueda A, Saneoka H (2015) Growth, physiological adaptation, and gene expression analysis of two Egyptian rice cultivars under salt stress. Plant Physiol Biochem 87:17–25. https://doi.org/10.1016/j.plaphy.2014.12.007
Mekawy AMM, Abdelaziz MN, Ueda A (2018) Apigenin pretreatment enhances growth and salinity tolerance of rice seedlings. Plant Physiol Biochem 130:94–104. https://doi.org/10.1016/j.plaphy.2018.06.036
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. https://doi.org/10.1146/annurev.arplant.59.032607.092911
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C, Byrt CS, Hare RA, Tyerman SD, Tester M, Plett D, Gilliham M (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol 30:360–364. https://doi.org/10.1038/nbt.2120
Munns R, Passioura JB, Colmer TD, Byrt CS (2020) Osmotic adjustment and energy limitations to plant growth in saline soil. New Phytol 225:1091–1096. https://doi.org/10.1111/nph.15862
Negm ME, Abo-Marzoka EA, Abdelhameed MM, Ghidan WF (2023) Inheritance of some biochemical, morpho-physiological and yield attributes trait of some rice genotypes under salinity condition. Egypt J Agric Res 101:552–568. https://doi.org/10.21608/ejar.2023.191201.1330
Nguyen HTT, Das Bhowmik S, Long H, Cheng Y, Mundree S, Hoang LTM (2021) Rapid accumulation of proline enhances salinity tolerance in Australian wild rice Oryza australiensis Domin. Plants 10:2044. https://doi.org/10.3390/plants10102044
Nieves-Cordones M, Martinez V, Benito B, Rubio F (2016a) Comparison between Arabidopsis and rice for main pathways of K+ and Na+ uptake by roots. Front Plant Sci 7:992. https://doi.org/10.3389/fpls.2016.00992
Nieves-Cordones M, Shiblawi A, Razzaq F, Sentenac H (2016b) Roles and transport of sodium and potassium in plants. The alkali metal ions: their role for life. Springer, pp 291–324. https://doi.org/10.1007/978-3-319-21756-7.
Olías R, Eljakaoui Z, Li J, De Morales PA, Marin-Manzano MC, Pardo JM, Belver A (2009) The plasma membrane Na/H antiporter SOS1 is essential for salt tolerance in tomato and affects the partitioning of na between plant organs. Plant Cell Environ 32:904–916. https://doi.org/10.1111/j.1365-3040.2009.01971.x
Ologundudu AF, Adelusi AA, Akinwale RO (2014) Effect of salt stress on germination and growth parameters of rice (Oryza sativa L). Not Sci Biol 6:237–243. https://doi.org/10.15835/nsb.6.2.9163
Pattanagul W, Thitisaksakul M (2008) Effect of salinity stress on growth and carbohydrate metabolism in three rice (Oryza sativa L.) cultivars differing in salinity tolerance. Indian J Exp Biol 46:736–742
Pottosin I, Dobrovinskaya O (2014) Non-selective cation channels in plasma and vacuolar membranes and their contribution to K+ transport. J Plant Physiol 171:732–742. https://doi.org/10.1016/j.jplph.2013.11.013
Shahid SA, Zaman M, Heng L (2018) Introduction to soil salinity, sodicity and diagnostics techniques. Guideline for salinity assessment, mitigation and adaptation using nuclear and related techniques. Springer, pp 1–42
Shahzad B, Yun P, Rasouli F, Shabala L, Zhou MX, Venkataraman G, Chen ZH, Shabala S (2022) Root K+ homeostasis and signalling as a determinant of salinity stress tolerance in cultivated and wild rice species. Environ Exp Bot 201:104944. https://doi.org/10.1016/j.envexpbot.2022.104944
Shehab M, Iovene M, Ciancio A, Colagiero M, Finetti-Sialer M (2022) Transcriptome analysis provides novel insights into salt stress response in two Egyptian rice varieties with different tolerance levels. Rice Sci 29:499–502. https://doi.org/10.1016/j.rsci.2022.09.001
Sriskantharajah K, Osumi S, Chuamnakthong S, Nampei M, Amas JC, Gregorio GB, Ueda A (2020) Contribution of two different Na+ transport systems to acquired salinity tolerance in rice. Plant Sci 297:110517. https://doi.org/10.1016/j.plantsci.2020.110517
Takagi H, Yamada S (2013) Roles of enzymes in anti-oxidative response system on three species of chenopodiaceous halophytes under NaCl-stress condition. Soil Sci Plant Nutr 59:603–611. https://doi.org/10.1080/00380768.2013.809600
Takahashi R, Nishio T, Ichizen N, Takano T (2007) High-affinity K+ transporter PhaHAK5 is expressed only in salt-sensitive reed plants and shows Na+ permeability under NaCl stress. Plant Cell Rep 26:1673–1679. https://doi.org/10.1007/s00299-007-0364-1
Wang Y, Wu WH (2013) Potassium transport and signaling in higher plants. Annu Rev Plant Biol 64:451–476. https://doi.org/10.1146/annurev-arplant-050312-120153
Wang Q, Guan C, Wang P, Lv ML, Ma Q, Wu GQ, Bao AK, Zhang JL, Wang SM (2015) AtHKT1;1 and AtHAK5 mediate low-affinity Na+ uptake in Arabidopsis thaliana under mild salt stress. Plant Growth Regul 75:615–623. https://doi.org/10.1007/s10725-014-9964-2
Zayed BA, Salem AE-aK, Ali OAM (2014) Physiological characterization of Egyptian salt tolerant rice varieties under different salinity levels. Life Sci J 11:1364–1372. https://doi.org/10.7537/marslsj111014.199
Zhang Y, Fang J, Wu X, Dong L (2018) Na+/K+ Balance and Transport Regulatory Mechanisms in Weedy and Cultivated Rice (Oryza sativa L.) under salt stress. BMC Plant Biol 18:375. https://doi.org/10.1186/s12870-018-1586-9
Zidan MS, Dawoud MA (2013) Agriculture use of marginal water in Egypt: Opportunities and challenges. In: Developments in Soil Salinity Assessment and Reclamation. pp 661–679. https://doi.org/10.1007/978-94-007-5684-7_43
Acknowledgements
This research was supported by JSPS KAKENHI Grant Numbers 20KK0129 to AU. AMMM extends his appreciation to the Academy of Scientific Research and Technology (ASRT, Egypt) for funding the Graduation Projects conducted at Laboratory of Plant Nutritional Physiology, Minia University, Egypt.
Funding
Open Access funding provided by Hiroshima University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare there are no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mekawy, A.M.M., Assaha, D.V., Li, J. et al. Differential Physiological and Molecular Processes in the Root May Underlie Contrasting Salt Tolerance in Two Egyptian Rice Cultivars at the Seedling Stage. J Soil Sci Plant Nutr 24, 3100–3114 (2024). https://doi.org/10.1007/s42729-024-01736-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42729-024-01736-7