Abstract
Purpose
Surra is an economically important livestock disease in many low- and middle-income countries, including those of Northern Africa. The disease is caused by the biting fly-transmitted subspecies Trypanosoma brucei evansi, which is very closely related to the tsetse-transmitted subspecies T. b. brucei and the sexually transmitted subspecies T. b. equiperdum. At least two phylogenetically distinct groups of T. b. evansi can be distinguished, called type A and type B. These evolved from T. b. brucei independently. The close relationships between the T. brucei subspecies and the multiple evolutionary origins of T. b. evansi pose diagnostic challenges.
Methods
Here we use previously established and newly developed PCR assays based on nuclear and mitochondrial genetic markers to type the causative agent of recent trypanosome infections of camels in Southern Algeria.
Results/conclusion
We confirm that these infections have been caused by T. b. evansi type A. We also report a newly designed PCR assay specific for T. b. evansi type A that we expect will be of diagnostic use for the community.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The single cellular parasite Trypanosoma brucei evansi belongs to the subgenus Trypanozoon that also comprises T. b. brucei and T. b. equiperdum [1, 2] (the taxonomical status of T. b. evansi is controversial [1, 3, 4]; for this study, we will be referring to it as a subspecies of T. brucei). Trypanosoma b. evansi is the most widely distributed of the pathogenic animal trypanosomes, affecting a large number of wild and domesticated animal species in Asia, Africa and Latin America [5, 6]. In Europe, it is present in the Canary Islands, from where recent sporadic incursions into the French and Spanish mainland have occurred [5, 7, 8]. Trypanosoma b. evansi causes a trypanosomosis called “surra” in many countries [8,9,10]. It is an acute, chronic or subclinical disease that is very often fatal in camels, horses and dogs, but can also seriously affect cattle and buffaloes. Other animals, including wildlife, are also susceptible.
In affected countries, surra is an economically important disease, which causes high mortality, reduced milk and meat production, poor carcass quality, reduced reproductive performance, and decreased draft power and manure production [9]. Haematophagous flies of the genera Tabanus and Stomoxys are particularly relevant for transmitting the infection from host to host, acting as mechanical vectors without parasite development in the insect [9]. This is a key difference to T. b. brucei, where transmission is dependent on cyclic development in the tsetse fly [11]. Indeed, it is this mechanical transmission that has allowed the parasite to move beyond the tsetse fly region and out of Africa [12]. In South and Central America, T. b. evansi can also be transmitted by vampire bats (Desmodus rotundus), which act as both vectors and reservoirs [9].
Another key difference to T. b. brucei is that T. b. evansi strains are either ‘dyskinetoplastic’ or ‘akinetoplastic’, i.e., they have either dysfunctional kinetoplast DNA (the mitochondrial DNA network in these organisms) or lack it entirely. Where kDNA is present, T. b. evansi strains typically lack maxicircles—the equivalent of mitochondrial DNA in other eukaryotes—and are characterized by minicircle sequence homogeneity [13, 14]. By contrast, T. b. brucei contains hundreds of different minicircle classes [15]. Trypanosoma b. evansi is therefore incapable of mitochondrial gene expression, and a compensatory mutation in the nuclearly encoded subunit γ of the F1FO ATP synthase is necessary to enable viability [16]. Based on the minicircle class that dominates the kDNA networks, T. b. evansi can be divided into types A and B [1, 13, 17]. Indeed, this difference can be exploited for polymerase chain reaction (PCR)-based diagnostics and molecular characterization of the parasite. PCR-based assays that target Trypanozoon-specific satellite DNA or ribosomal DNA are regarded as the most sensitive for diagnosis or characterization of surra infections [18,19,20], while for genotyping T. b. evansi and/or to distinguish between T. b. evansi types A and B, PCR assays targeting type-specific variant surface glycoprotein genes, mitochondrial minicircles and maxicircles, microsatellite markers and the F1-ATP synthase γ subunit gene are being used [4, 17, 21,22,23].
In northern Africa, the first cases of trypanosomosis were officially reported from Algeria, Mauritania, Morocco and Tunisia at the beginning of the last century [24,25,26,27]. In-depth epidemiological studies began at the end of the 1980s and showed that camel trypanosomosis could be considered as a dominant disease, with variable prevalence rates depending on the year, the sampling period and the provinces or wilayate (districts) surveyed [28,29,30,31,32,33,34]. A recent epidemiological study in southern Algeria carried out on 1056 dromedary camels revealed overall prevalence rates of 2.4% by Giemsa-stained thin smear (GST), 32.4% by card agglutination test for trypanosomosis (CATT/T. evansi), 23.1% by enzyme-linked immunosorbent assay (ELISA/VSG RoTat 1.2), 21.0% by immune trypanolysis (TL) and 11.2% by PCR (RoTat 1.2 PCR) [35].
Here, we present a genotyping analysis for six of the camels from the previous study [35], based on sequencing of minicircle DNA and of the F1FO ATP synthase subunit γ gene, and confirm the pathogen as T. b. evansi type A. Furthermore, we present a novel PCR assay based on primers with improved specificity for minicircle type A that will be useful for typing of surra infections.
Materials and Methods
All PCR primers are listed in Table 1. All trypanosome isolates or strains are listed in Table 2. Trypanosoma b. evansi and T. b. equiperdum reference strains were kind gifts from Kirsten Gillingwater, Swiss Tropical Institute [36] and from Philippe Büscher and Nick Van Reet, ITM Antwerp.
Growth of T. b. evansi and T. b. equiperdum Reference Strains and DNA Isolation
Trypanosome reference strains were grown in MFI mice and purified from blood using DEAE cellulose as described [40]. DNA extraction was performed using the QIAamp 250 mini blood kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions.
Preparation of FTA Punches
The Harris Uni-Core punch tool (Merck, Darmstadt, Germany) and cutting mat were prepared by soaking in 2% (w/v) sodium hypochlorite solution for 10 min, followed by three washes with ddH2O, soaking in 70% (v/v) ethanol for 5 min and air drying. Punches from FTA cards were washed three times with 200 µl FTA Purification Reagent (GE Healthcare) for 5 min each and twice with 200 µl TE buffer (10 mM Tris–HCl, 0.1 mM EDTA, pH 8.0) for 5 min each. Punches were dried at 50 °C for 15 min and added directly to PCR reaction tubes.
PCR Assays
All PCR assays were performed in 25 µl volumes and used FTA card punches or trypanosome genomic DNA (1–5 ng) as indicated (negative controls included additional H2O instead). Assays for all targets, with exception of the full-length F1FO ATP synthase subunit γ (Tb927.10.180), used the following reagents:
Reagent | Volume |
|---|---|
5 × GoTaq PCR buffer (Promega) | 5 µl |
MgCl2 (25 mM) | 2 µl |
dNTPs (10 mM) | 0.5 µl |
GoTaq G2 Hot Start (Promega) | 0.125 µl |
Specific primers, their volumes, and PCR cycling conditions were as follows:
Target | Primers (10 µM) | Volume (µl) | Cycling conditions |
|---|---|---|---|
F1FO ATP synthase subunit γ (Tb927.10.180), 511-bp fragment | #1, #2 | 1 | 95 °C 5 min 35x (95 °C 30 s, 55 °C 30 s, 72 °C 1 min) 72 °C 10 min |
Duplex assay minicircle type A (novel)/F1FO ATP synthase subunit γ 511-bp fragment | #3, #4, #5, #6 | 1.25 | 95 °C 5 min 40 × (95 °C 30 s, 51 °C 30 s, 72 °C 1 min) 72 °C 10 min |
Minicircle type A (novel) | #5, #6 | 2.5 | 95 °C 5 min 40 × (95 °C 30 s, 51 °C 30 s, 72 °C 1 min) 72 °C 10 min |
Minicircle type A (ref [17]) | MiniA, MiniB | 2.5 | 95 °C 5 min 40 × (95 °C 30 s, 51 °C 30 s, 72 °C 1 min) 72 °C 10 min |
Maxicircle gene A6 | #7, #8 | 1 | 95 °C 5 min 35 × (95 °C 30 s, 55 °C 30 s, 72 °C 1 min) 72 °C 10 min |
Maxicircle gene ND4 | #9, #10 | 1 | 95 °C 5 min 40 × (95 °C 30 s, 54 °C 30 s, 72 °C 1 min) 72 °C 10 min |
PCR reagents for the full-length F1FO ATP synthase subunit γ (Tb927.10.180) gene, including flanking regions, were as follows (25 µl total):
Reagent | Volume |
|---|---|
5 × Phusion PCR buffer (New England Biolabs) | 5 µl |
Primers #3 and #4 (10 µM) | 1.25 µl |
dNTPs (10 mM) | 0.5 µl |
Hot Start Phusion (New England Biolabs) | 0.25 µl |
PCR cycling conditions for the full-length gene were as follows: 98 °C 30 s, 40 cycles (98 °C 10 s, 60 °C 30 s, 72 °C 1 min), 72 °C 10 min.
Cloning and Sequencing
All PCR products were cleaned up using the PCR Clean-Up kit from Macherey–Nagel (Dueren, Germany). Sequencing was either direct, using the same primers that had been used for the PCR reaction, or after cloning into pCR-Blunt (Invitrogen; for Phusion PCR products) or into pGEM-T easy (Promega; for GoTaq PCR products), following the manufacturer’s instructions. Cloned products were sequenced using Sanger technology (Edinburgh Genomics or MRC Sequencing Service, Dundee) and standard M13 forward and reverse primers.
Phylogenetic Analysis
A phylogenetic tree was constructed with IQ-TREE [41], using a maximum likelihood model with HKY + G substitution.
Results and Discussion
PCR assays for TbATPase subunit γ confirm infection with T. b. evansi type A
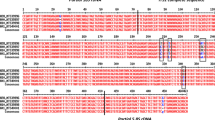
To confirm the diagnosis of a T. b. evansi infection in camels from 5 different Algerian regions (Table 2) [35], we amplified by PCR a 511 bp fragment of subunit γ of the mitochondrial F1FO ATP synthase (systematic TriTrypDB ID Tb927.10.180). In the T. b. evansi types identified so far, this gene contains adaptive mutations that are differentially diagnostic for types A and B [1, 4, 21]. Punches from FTA cards containing DNA purified from blood samples from cases 1 to 6 were washed and placed in reaction tubes, together with PCR reagents and primers #1 and #2 (Table 1). Initial reactions were carried out with (non-proof-reading) Taq polymerase because of its robust performance. Total cellular DNA from a T. b. brucei strain served as positive control. Reactions for all six cases showed a single amplicon of the expected size, suggesting infection with a Trypanozoon (Fig. 1A). To identify the type of T. b. evansi, we next amplified the entire ATP synthase γ gene with primers #3 and #4 and a proof-reading polymerase, followed by cloning and sequencing. Sequence analysis confirmed presence of a heterozygous A281del mutation in the ATP synthase γ protein for all cases (Fig. 1B), providing conclusive evidence for infection with T. b. evansi type A [1]. These results are consistent with the previously reported RoTat1.2-positive PCR results for these isolates [35]. RoTat1.2 is a VSG gene that, when present, is generally considered as being diagnostic for T. b. evansi type A [5]. These results are also consistent with the fact that the only other type of T. b. evansi currently known, type B, has so far only been reported from countries in East Africa, namely Kenya and Ethiopia [5, 17, 21].
Detection by PCR of ATP synthase subunit γ sequences diagnostic of T. b. evansi type A. A PCR assay for detection of a 511-bp fragment of ATP synthase subunit γ (Tb927.10.180). Aliquots of completed PCR reactions (15 μl) were fractionated by electrophoresis on an agarose gel containing ethidium bromide. Images were captured using a UV light box. Lanes 1, 19: New England Biolabs 100-bp ladder (kbp: kilobasepairs); lanes 3–8: Algerian camel cases 1–6; lanes 9, 18: PCR reactions with water instead of samples; lanes 10–17: varying amounts of total cellular DNA from T. b. brucei strain EATRO 1125 AnTat1.1 90:13. B Sequencing of ATP synthase γ sequences. Top, trace files of direct sequencing (from the 5′ end) of PCR amplicons from cases 1, 3 and 4. Bottom, representative sequences obtained after cloning of PCR amplicons. Sequencing of cloned amplicons confirmed that T. b. evansi strains responsible for infections 1, 2, 5 and 6 are heterozygous for deletion of amino acid alanine 281 (A281del). All cloned sequences obtained for case 3 were wild-type, and no cloned sequences were obtained for case 4, but direct sequencing of PCR amplicons confirmed heterozygosity for A281del for those cases as well
Development of a Novel PCR Assay Specific for Minicircle Type A
A defining characteristic of T. b. evansi type A is that (unless the strain is akinetoplastic [1]) its kDNA is dominated by, or consists entirely of, thousands of copies of a particular class of minicircle [13]. A PCR assay for this minicircle class developed by Njiru and colleagues [17] uses primers (‘MiniA’ and ‘MiniB’) derived from a region semi-conserved among all minicircle classes (Supplementary Figure S1) and can therefore result in false positive reactions [21]. We therefore aimed to develop a PCR assay that is highly specific for type A minicircles. Alignment of type A minicircles available in GenBank identified several regions of perfect conservation outside of the universally conserved region. Based on this information, we designed primer sequences that are predicted to amplify a fragment of ~ 570 bp in a PCR assay (Supplementary Figure S1, primers #5 and #6). Alignment of the minicircle type A consensus to the closest match in the recently defined minicircle population of T. b. brucei EATRO 1125 [15], a minicircle that contains the same set of gRNA genes, suggests that primers #5 and #6 should be specific for T. b. evansi minicircle type A (Supplementary Figure S2). Indeed, when tested against a panel of type A and non-type A strains or isolates, the PCR assay was highly specific (Fig. 2A). Sequencing of the ~ 570 bp amplicons confirmed that they corresponded to the expected minicircle type A. The only unexpected result was absence of a ~ 570 bp amplicon for strain T. b. evansi CAN86/Brazil (Fig. 2A, lane 6). As a PCR reaction using the MiniA/MiniB primers also failed to produce an amplicon for this strain (data not shown), we suspect that this strain has spontaneously lost its kDNA. This phenomenon is not unusual in T. b. evansi and T. b. equiperdum [14, 42].
A specific PCR assay for minicircle type A. A PCR assay for detection of a ~ 570 bp fragment of minicircle type A (‘mini A’) in samples. In the same reactions (duplex PCR), primers #3 and #4 for amplification of a ~ 1.4-kb ATP synthase subunit γ amplicon (‘subunit γ’) were included as positive internal controls. Per reaction, 1–5 ng total DNA were used as template. Lane 1: Bioline 1-kbp ladder; lanes 2, 19: New England Biolabs 100-bp ladder; lanes 3, 18: empty; lane 4: control PCR reaction with water instead of total DNA; lane 5: control PCR reaction with mouse genomic DNA instead of total trypanosome DNA (several trypanosome strains/isolates were grown in mice); lanes 6–17: reactions with total trypanosome DNA. Trypanosome strains/isolates were as follows. 1 = T. b. evansi CAN86/Brazil; 2 = T. b. evansi Antat3/3 (akinetoplastic); 3 = T. b. evansi KETRI 2479; 4 = T. b. equiperdum BoTat1.1; 5 = T. b. equiperdum OVI; 6 = T. b. equiperdum Hamburg; 7 = T. b. evansi RoTat1.2; 8 = T. b. evansi Philippines; 9 = T. b. brucei Lister 427; 10 = T. b. brucei EATRO 1125 AnTat1.1; 11 = T. b. equiperdum American; 12 = T. b. equiperdum AnTat4.1. Strains/isolates previously identified as belonging to the type A group [1, 3] are indicated by an asterisk. Please note: (i) T. b. equiperdum in this group have been suggested to be misidentified or mislabelled T. b. evansi [3]; (ii) T. b. evansi AnTat3/3 (lane 7) is a type A strain [43], but the strain in our lab had spontaneously lost its kDNA [44]; (iii) T. b. evansi CAN86/Brazil is a type A strain [1, 3], but, like AnTat3/3, may have spontaneously lost its kDNA; (iv) amplification of minicircle type A in the same reaction appears to diminish the signal for subunit γ, perhaps by competing for nucleotides, this is particularly evident in lane 11. B Analysis of cases 1–6 using the PCR assay with primers #5/#6 (left panel) or primers MiniA/MiniB (right panel). Lane 1: New England Biolabs 100-bp ladder; lane 2: control PCR reaction with water instead of total DNA; lanes 3–8: FTA card punches from cases 1 to 6; lane 9: empty; lane 10: empty (left panel); lane 11 (left panel) / lane 10 (right panel): T. b. evansi RoTat1.2 (positive control)
Next, we used the new PCR assay to analyze samples from cases 1 to 6. For cases 1, 2, 3 and 6, we obtained a specific band of ~ 570 bp (Fig. 2B, left panel), and direct sequencing confirmed that the amplicons were the type A fragment (Supplementary Figure S3). We did not obtain a product for cases 4 and 5. The same result was obtained with primers MiniA/MiniB: strong amplification products of the expected size for cases 1, 2, 3 and 6, but no products for cases 4 and 5 (Fig. 2B, right panel). We conclude that, in all six cases, camels had been infected with T. b. evansi type A. In cases 4 and 5, the parasites may have become akinetoplastic, and our typing relies exclusively on the presence of the A281del mutation of ATP synthase subunit γ. A phylogenetic tree based on the ~ 570 bp minicircle type A amplicon shows a separation into two main branches that is supported by strong bootstrap values (Supplementary Fig. 4). The Algerian cases branch together with two other isolates from Africa, and also a single isolate from South America, whereas the other isolates, all from non-African countries, form a separate branch. It will be interesting to expand the phylogenetic analysis of type A T. b. evansi based on their minicircle sequence to include other isolates, perhaps using the entire ~ 1-kb sequence to further improve resolution and reliability.
PCR assays for maxicircle-encoded genes A6 and ND4 (primer pairs #7/#8 and #9/#10, respectively; Table 2) were negative (data not shown), consistent with the expected absence of the maxicircle in T. b. evansi [13, 14].
Conclusion
Based on nuclear and mitochondrial genetic markers, we have confirmed that the recently reported trypanosome infections in southern Algerian camels were caused by T. b. evansi type A, adding to an accumulating body of recent reports of surra infections in that country [45,46,47]. We also report a novel PCR assay based on careful sequence analysis of type A minicircles that we expect will be a useful tool for the community to diagnose T. b. evansi type A infections in livestock. Our data reported here suggest good specificity and sensitivity for type A strains and compatibility with samples prepared on FTA cards. Further studies should compare specificity and sensitivity with other assays, such as the recently reported recombinase polymerase amplification lateral flow assay for T. b. evansi [48].
References
Carnes J, Anupama A, Balmer O, Jackson A, Lewis M, Brown R et al (2015) Genome and phylogenetic analyses of Trypanosoma evansi reveal extensive similarity to T. brucei and multiple independent origins for dyskinetoplasty. PLoS Negl Trop Dis 9:e3404. https://doi.org/10.1371/journal.pntd.0003404
Cuypers B, Van den Broeck F, Van Reet N, Meehan CJ, Cauchard J, Wilkes JM et al (2017) Genome-wide SNP analysis reveals distinct origins of Trypanosoma evansi and Trypanosoma equiperdum. Genome Biol Evol 9:1990–1997. https://doi.org/10.1093/gbe/evx102
Claes F, Büscher P, Touratier L, Goddeeris BM (2005) Trypanosoma equiperdum: master of disguise or historical mistake? Trends Parasitol 21:316–321. https://doi.org/10.1016/j.pt.2005.05.010
Lai D-H, Hashimi H, Lun Z-R, Ayala FJ, Lukes J (2008) Adaptations of Trypanosoma brucei to gradual loss of kinetoplast DNA: Trypanosoma equiperdum and Trypanosoma evansi are petite mutants of T. brucei. Proc Natl Acad Sci USA 105:1999–2004. https://doi.org/10.1073/pnas.0711799105
Aregawi WG, Agga GE, Abdi RD, Büscher P (2019) Systematic review and meta-analysis on the global distribution, host range, and prevalence of Trypanosoma evansi. Parasit Vectors 12:67. https://doi.org/10.1186/s13071-019-3311-4
Morrison LJ, Vezza L, Rowan T, Hope JC (2016) Animal African trypanosomiasis: time to increase focus on clinically relevant parasite and host species. Trends Parasitol 32:599–607. https://doi.org/10.1016/j.pt.2016.04.012
Gutierrez C, Desquesnes M, Touratier L, Buscher P (2010) Trypanosoma evansi: recent outbreaks in Europe. Vet Parasitol 174:26–29. https://doi.org/10.1016/j.vetpar.2010.08.012
Desquesnes M, Holzmuller P, Lai DH, Dargantes A, Lun ZR, Jittaplapong S (2013) Trypanosoma evansi and surra: a review and perspectives on origin, history, distribution, taxonomy, morphology, hosts, and pathogenic effects. Biomed Res Int 2013:194176. https://doi.org/10.1155/2013/194176
Desquesnes M, Dargantes A, Lai D-H, Lun Z-R, Holzmuller P, Jittapalapong S (2013) Trypanosoma evansi and surra: a review and perspectives on transmission, epidemiology and control, impact, and zoonotic aspects. Biomed Res Int 2013:321237. https://doi.org/10.1155/2013/321237
Health EPoA, Welfare, More S, Botner A, Butterworth A, Calistri P et al (2017) Assessment of listing and categorisation of animal diseases within the framework of the Animal Health Law (Regulation (EU) No 2016/429): Trypanosoma evansi infections (including Surra). EFSA J 15:e04892. https://doi.org/10.2903/j.efsa.2017.4892
Sharma R, Gluenz E, Peacock L, Gibson W, Gull K, Carrington M (2009) The heart of darkness: growth and form of Trypanosoma brucei in the tsetse fly. Trends Parasitol 25:517–524. https://doi.org/10.1016/j.pt.2009.08.001
Jensen RE, Simpson L, Englund PT (2008) What happens when Trypanosoma brucei leaves Africa. Trends Parasitol 24:428–431. https://doi.org/10.1016/j.pt.2008.06.007
Borst P, Fase-Fowler F, Gibson WC (1987) Kinetoplast DNA of Trypanosoma evansi. Mol Biochem Parasitol 23:31–38. https://doi.org/10.1016/0166-6851(87)90184-8
Schnaufer A, Domingo GJ, Stuart K (2002) Natural and induced dyskinetoplastic trypanosomatids: how to live without mitochondrial DNA. Int J Parasitol 32:1071–1084. https://doi.org/10.1016/s0020-7519(02)00020-6
Cooper S, Wadsworth ES, Ochsenreiter T, Ivens A, Savill NJ, Schnaufer A (2019) Assembly and annotation of the mitochondrial minicircle genome of a differentiation-competent strain of Trypanosoma brucei. Nucleic Acids Res 47:11304–11325. https://doi.org/10.1093/nar/gkz928
Dean S, Gould MK, Dewar CE, Schnaufer AC (2013) Single point mutations in ATP synthase compensate for mitochondrial genome loss in trypanosomes. Proc Natl Acad Sci USA 110:14741–14746. https://doi.org/10.1073/pnas.1305404110
Njiru ZK, Constantine CC, Masiga DK, Reid SA, Thompson RCA, Gibson WC (2006) Characterization of Trypanosoma evansi type B. Infect Genet Evol 6:292–300. https://doi.org/10.1016/j.meegid.2005.08.002
Gari FR, Ashenafi H, Tola A, Goddeeris BM, Claes F (2010) Comparative diagnosis of parasitological, serological, and molecular tests in dourine-suspected horses. Trop Anim Health Prod 42:1649–1654. https://doi.org/10.1007/s11250-010-9615-1
Masiga DK, Smyth AJ, Hayes P, Bromidge TJ, Gibson WC (1992) Sensitive detection of trypanosomes in tsetse flies by DNA amplification. Int J Parasitol 22:909–918. https://doi.org/10.1016/0020-7519(92)90047-o
Njiru ZK, Constantine CC, Guya S, Crowther J, Kiragu JM, Thompson RCA et al (2005) The use of ITS1 rDNA PCR in detecting pathogenic African trypanosomes. Parasitol Res 95:186–192. https://doi.org/10.1007/s00436-004-1267-5
Birhanu H, Gebrehiwot T, Goddeeris BM, Büscher P, Van Reet N (2016) New Trypanosoma evansi type B isolates from Ethiopian dromedary camels. PLoS Negl Trop Dis 10:e0004556. https://doi.org/10.1371/journal.pntd.0004556
Masiga DK, Gibson WC (1990) Specific probes for Trypanosoma (Trypanozoon) evansi based on kinetoplast DNA minicircles. Mol Biochem Parasitol 40:279–283. https://doi.org/10.1016/0166-6851(90)90049-r
Claes F, Radwanska M, Urakawa T, Majiwa PA, Goddeeris B, Büscher P (2004) Variable Surface Glycoprotein RoTat 1.2 PCR as a specific diagnostic tool for the detection of Trypanosoma evansi infections. Kinetoplastid Biol Dis 3:3. https://doi.org/10.1186/1475-9292-3-3
Curasson G (ed) (1943) Traité de Protozoologie Vétérinaire et Comparée; Tome I: Trypanosomes. Vigot Frères, Paris, pp 335–366
Hoare CA (1972) The trypanosomes of mammals: a zoological monograph. Blackwell Scientific Publications, Oxford
Sergent E, Donatien A (1921) De l’infection latente dans la trypanosomiase des dromadaires (le Debab). Archives des Instituts Pasteur de l'Afrique du Nord (Alger), XXIIe, pp 179–184
Sergent E, Sergent E (1905) El-Debab. Trypanosomiase des dromadaires de l’Afrique du Nord. Annales de l’Institut Pasteur d’Alger, Alger
Atarhouch T, Rami M, Bendahman MN, Dakkak A (2003) Camel trypanosomosis in Morocco 1: results of a first epidemiological survey. Vet Parasitol 111:277–286. https://doi.org/10.1016/s0304-4017(02)00382-5
Boushaki D (2007) Prévalence de la Trypanosomose Cameline en Algérie. M.Sc. thesis, École Nationale Supérieure Vétérinaire d’Alger.http://archive.ensv.dz:8080/jspui/handle/123456789/215. Accessed 29 Apr 2022
Dia ML, Barry Y, Ould Ahmed M, Claes F, Büscher P, Ba A (2009) Nouvelles données sur la trypanosomose cameline à T. evansi en Mauritanie. In: Proceedings 30th biennial conference of the International Scientific Council for Trypanosomiasis Research and Control (ISCTRC). Kampala, Uganda, pp 391–398. https://www.au-ibar.org/sites/default/files/2020-11/doc_20110922_isctrc_30thconference_proceedings_en.pdf. Accessed 29 Apr 2022
Dia ML, Boushaki D, Dakkak A, Atarhouch T, Rami T, Jemli MH et al (2007) Situation épidémiologique de la trypanosomiase cameline en Mauritanie, en Algérie, au Maroc et en Tunisie. In: Proceedings 29th biennial conference of the International Scientific Council for Trypanosomiasis Research and Control (ISCTRC). Luanda, Angola, pp 388–398. https://www.au-ibar.org/sites/default/files/2020-11/doc_20110929_isctrc_29thconference_proceedings_en.pdf. Accessed 29 Apr 2022
Dia ML, Diop C, Aminetou M, Jacquiet P, Thiam A (1997) Some factors affecting the prevalence of Trypanosoma evansi in camels in Mauritania. Vet Parasitol 72:111–120. https://doi.org/10.1016/S0304-4017(97)00054-X
Jemli MH, Megdiche F, Laaridi M, Bahri S, Kallel A, Mejri M (2000) Principales maladies du dromadaire en Tunisie. In: Proceedings Maladies parasitaires et infectieuses du dromadaire, Rabat, Marocco. Actes Éditions, pp 19–21
Rami M, Atarhouch T, Bendahman MN, Azlaf R, Kechna R, Dakkak A (2003) Camel trypanosomosis in Morocco. 2. A pilot disease control trial. Vet Parasitol 115:223–231. https://doi.org/10.1016/S0304-4017(03)00222-X
Boushaki D, Adel A, Dia ML, Büscher P, Madani H, Brihoum BA et al (2019) Epidemiological investigations on Trypanosoma evansi infection in dromedary camels in the South of Algeria. Heliyon 5:e02086. https://doi.org/10.1016/j.heliyon.2019.e02086
Gillingwater K, Büscher P, Brun R (2007) Establishment of a panel of reference Trypanosoma evansi and Trypanosoma equiperdum strains for drug screening. Vet Parasitol 148:114–121. https://doi.org/10.1016/j.vetpar.2007.05.020
Domingo GJ, Palazzo SS, Wang B, Pannicucci B, Salavati R, Stuart KD (2003) Dyskinetoplastic Trypanosoma brucei contains functional editing complexes. Eukaryotic Cell 2:569–577. https://doi.org/10.1128/EC.2.3.569-577.2003
Engstler M, Boshart M (2004) Cold shock and regulation of surface protein trafficking convey sensitization to inducers of stage differentiation in Trypanosoma brucei. Genes Dev 18:2798–2811. https://doi.org/10.1101/gad.323404
Wirtz E, Leal S, Ochatt C, Cross GAM (1999) A tightly regulated inducible expression system for conditional gene knock-outs and dominant-negative genetics in Trypanosoma brucei. Mol Biochem Parasitol 99:89–101. https://doi.org/10.1016/S0166-6851(99)00002-X
Dewar CE, MacGregor P, Cooper S, Gould MK, Matthews KR, Savill NJ et al (2018) Mitochondrial DNA is critical for longevity and metabolism of transmission stage Trypanosoma brucei. PLoS Pathog 14:e1007195. https://doi.org/10.1371/journal.ppat.1007195
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Suganuma K, Narantsatsral S, Battur B, Yamasaki S, Otgonsuren D, Musinguzi SP et al (2016) Isolation, cultivation and molecular characterization of a new Trypanosoma equiperdum strain in Mongolia. Parasit Vectors 9:481. https://doi.org/10.1186/s13071-016-1755-3
Songa EB, Paindavoine P, Wittouck E, Viseshakul N, Muldermans S, Steinert M et al (1990) Evidence for kinetoplast and nuclear DNA homogeneity in Trypanosoma evansi isolates. Mol Biochem Parasitol 43:167–179. https://doi.org/10.1016/0166-6851(90)90142-9
Schnaufer A, Clark-Walker GD, Steinberg AG, Stuart K (2005) The F1-ATP synthase complex in bloodstream stage trypanosomes has an unusual and essential function. EMBO J 24:4029–4040. https://doi.org/10.1038/sj.emboj.7600862
Boutellis A, Bellabidi M, Benaissa MH, Harrat Z, Brahmi K, Drali R, Kernif T (2021) New haplotypes of Trypanosoma evansi identified in dromedary camels from Algeria. Acta Parasitol 66:294–302. https://doi.org/10.1007/s11686-020-00316-w
Benfodil K, Büscher P, Abdelli A, Van Reet N, Mohamed-Herif A, Ansel S et al (2020) Comparison of serological and molecular tests for detection of Trypanosoma evansi in domestic animals from Ghardaïa district, South Algeria. Vet Parasitol 280:109089. https://doi.org/10.1016/j.vetpar.2020.109089
Benfodil K, Büscher P, Ansel S, Mohamed Cherif A, Abdelli A, Van Reet N et al (2020) Assessment of Trypanosoma evansi prevalence and associated risk factors by immune trypanolysis test in camels from Ghardaïa district, southern Algeria. Vet Parasitol Reg Stud Rep 22:100460. https://doi.org/10.1016/j.vprsr.2020.100460
Li Z, Pinto Torres JE, Goossens J, Stijlemans B, Sterckx YG-J, Magez S (2020) Development of a recombinase polymerase amplification lateral flow assay for the detection of active Trypanosoma evansi infections. PLoS Negl Trop Dis 14:e0008044. https://doi.org/10.1371/journal.pntd.0008044
Acknowledgements
We thank Kirsten Gillingwater (Swiss Tropical Institute), Philippe Büscher and Nick Van Reet (ITM Antwerp) for the kind gifts of T. b. evansi and T. b. equiperdum reference strains. This work was supported by the UK Medical Research Council (Grant MR/L019701/1 to A.S.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that no potential conflicts of interests exist.
Compliance with ethical standards
Authorisation to conduct the original survey [35] had been obtained from the Direction des Services Vétérinaires (DSV, Ministry of Agriculture, Rural Development and Fisheries). At each wilaya, the study had been authorized and supervised by the respective Inspection Vétérinaire de Wilaya (IVW Béchar, El Bayadh, Ouargla, and Tamanrasset), operating under the umbrella of the Direction des Services Vétérinaires. All experiments in Achim Schnaufer’s laboratory are carried out after local ethical approval at the University of Edinburgh by the School of Biological Sciences Ethics Committee (application “aschnauf-0002 Mitochondrial biology of trypanosomes”).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Boushaki, D., Wallis, J., Van den Broeck, F. et al. Molecular Analysis of Trypanosome Infections in Algerian Camels. Acta Parasit. 67, 1246–1253 (2022). https://doi.org/10.1007/s11686-022-00577-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11686-022-00577-7