Abstract
Plastomes of Fabaceae show both significant structural and size variation; however, most published plastomes are from subfamily Papilionoideae and only a few are from the other two subfamilies. In order to address the plastome structural and size variation of subfamily Mimosoideae, we integrated 11 newly sequenced plastomes from representing genera with three previously published ones. Each mimosoid plastome presented a typical quadripartite structure and contained 111 unique genes. Their inverted repeats (IRs) experienced multiple expansion/contraction; a ca. 13-kb IR expansion into small single copy (SSC) was detected in plastomes of a clade formed by tribe Ingeae and Acacia sensu stricto (s.s.), and a ca. 1.7-kb IR expansion into and a ca. 1.9-kb contraction out of large single copy (LSC) were found in Pithecellobium flexicaule and Acacia dealbata, respectively. Linear regression analysis showed decreased synonymous substitution rates of genes relocating from SSC into IR. A loss of both introns of clpP occurred in A. dealbata and Faidherbia albida, and a duplicated clpP copy was detected in A. dealbata. Furthermore, a 421-bp inversion that containing rps18 was found in A. dealbata. The size of mimosoid plastomes was found significantly affected by a IR-SC boundary shift, and also associated with repeat content. Plastome coding and noncoding regions with variable sequence divergence may supply valuable markers for molecular evolutionary and phylogenetic studies at different taxonomic levels. Plastid phylogenomics well resolved relationships among sampled mimosoid species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Having two copies of inverted repeat (IR) separated by the large and small single copy (LSC and SSC) regions is a typical feature of most plastomes (Jansen and Ruhlman 2012). The plastome size of land plants is typically 120–160 kb in length (Bock 2007) but varies considerably among some photosynthetic seed plant plastomes (Jansen and Ruhlman 2012). An extreme example comes from the plastome of the family Geraniaceae, which ranges from 116,935 bp in Erodium carvifolium to 217,942 bp in Pelargonium × hortorum (Blazier et al. 2011; Guisinger et al. 2011). The expansion/contraction or loss of an IR is one of the most evident causes of plastome size variation (Jansen and Ruhlman 2012). The IR size of land plants is typically 15–30 kb (Zhu et al. 2016) but ranges from 10 to 76 kb considering some extreme examples (Palmer 1991). Small IR expansions and contractions (<100 bp) are very frequent in angiosperm plastomes (Goulding et al. 1996), whereas large IR expansions (>1000 bp) occur less, and large contractions are even rare (Raubeson and Jansen 2004). Large IR expansions were reported from plastomes of Pelargonium × hortorum, Berberidaceae, Ranunculaceae, and some other plants (Chumley et al. 2006; Downie and Jansen 2015; Goulding et al. 1996; Hoot and Palmer 1994; Kim and Jansen 1994; Luo et al. 2016; Palmer et al. 1987a; Plunkett and Downie 2000; Shen et al. 1982). Large IR contractions were reported in the fern genus Osmunda (Stein et al. 1986) and Ophioglossum (Grewe et al. 2013) and some members from Myoporaceae, Loganiaceae, and Apiaceae (Plunkett and Downie 2000). Extremely, plastomes of some plants have contracted most or even lost IR, as observed in conifers (Guo et al. 2014; Wu and Chaw 2014), the inverted-repeat-lacking clade (IRLC) legumes (Doyle et al. 1996; Palmer et al. 1987b), some species of Erodium (Guisinger et al. 2011), and some broomrapes (Blazier et al. 2016; Wicke et al. 2013). In addition, repeat content has been also considered as an important contributor to plastome size variation (Dugas et al. 2015; Jo et al. 2011). The size of four published mimosoid plastomes varies significantly according to their IR length. Inga leiocalycina with expanded IR has the largest plastome (175,489 bp) among studied legumes (Dugas et al. 2015). Tandem repeat proportion was also inferred to be associated with mimosoid plastome size (Dugas et al. 2015). The size variation of mimosoid plastomes should be systematically addressed by including more representative taxa.
Previous studies revealed significant structural variation among legume plastomes. Most have been found in plastomes of subfamily Papilionoideae. A 50-kb inversion in the LSC region occurs in plastomes of most papilionoids except a few early diverged lineages (Doyle et al. 1996). A further 78-kb inversion including the 50-kb segment occurs in Vigna radiata and Phaseolus vulgaris (Palmer et al. 1988), and a further 36-kb inversion within the 50-kb segment occurs in Lupinus luteus (Martin et al. 2014). Plastomes of the IRLC have lost their IRs (Doyle et al. 1996; Palmer et al. 1987b; Wojciechowski et al. 2000). Significant plastome structural rearrangements have been detected in many species of the IRLC. For example, the plastomes of the “core Trifolium” undergo extensive genomic reconfiguration including many gene duplications, losses, and order changes attributable to inversions (Cai et al. 2008; Sveinsson and Cronk 2014); plastome of Cicer arietinum exhibit many gene and intron losses (Jansen et al. 2008); plastomes of Astragalus species present several gene losses and large inversions (Choi et al. 2015; Lei et al. 2016). However, only limited studies focused on the other two subfamilies. Within Mimosoideae, a ca. 13-kb IR-SSC boundary shift was detected in the plastomes of Acacia and Inga and a loss of clpP intron 1 in the plastome of Inga (Dugas et al. 2015). Five plastomes of Caesalpinioideae are available in GenBank (National Center for Biotechnology Information), but relevant results have not been published. More samples should be included in comparative analyses to detect new plastome structural variation in these two subfamilies.
Mimosoideae is the second largest subfamily of Fabaceae and is characterized by regular flowers that often have increased stamen numbers (Käss and Wink 1996). It is comprised of 83 genera and approximately 3270 species in four recognized tribes of Acacieae, Ingeae, Mimoseae, and Mimozygantheae (Lewis et al. 2005; LPWG 2013). Mimosoids have a pantropical distribution and form ecologically abundant elements in all major tropical biomes (Lewis et al. 2005; LPWG 2013). Many mimosoid species are important horticultural plants (species of Albizia and Calliandra), tropical feeding crops (Leucaena leucocephala), timbers (species of Acacia, Anadenanthera, and Prosopis), glues, and food thickeners (species of Acacia) (Lewis et al. 2005).
For such an economically and ecologically important group, the classification and phylogenetic relationships of Mimosoideae remain unclear. The monophyly of the subfamily is strongly supported by previous molecular studies (LPWG 2013). Nevertheless, all traditionally recognized tribes (excluding the monotypic Mimozygantheae) are supported to be non-monophyletic by a few plastid and nuclear gene regions, and relationships among many major lineages remain unresolved (LPWG 2013). Phylogenomics applies genomic data to reconstruct the evolutionary history of organisms (Delsuc et al. 2005; Eisen 1998; Eisen and Fraser 2003) and has been applied to tackle rapidly radiating clades (Barrett et al. 2014; Bewick et al. 2012; Jian et al. 2008; Ruhfel et al. 2014; Wickett et al. 2014; Zhou et al. 2012). A plastid phylogenomic approach has been successfully applied to resolve many enigmatic relationships within angiosperms and across all green plants (Jansen et al. 2007; Moore et al. 2007; Moore et al. 2010; Xi et al. 2012; Zhong et al. 2010). To fully resolve phylogenetic relationships within this subfamily, more variable plastid gene fragments and even whole plastomes should be applied in the analysis.
In this study, we integrated 11 newly sequenced plastomes representing three of four traditionally recognized mimosoid tribes with three mimosoid plastomes available in GenBank. The main objectives of this study were (1) to reveal plastome structural and size variation in mimosoids, (2) to identify synonymous substitution rate shifts of genes relocated into or out of the IR, (3) to explore sequence divergence of plastome regions for further evolutionary and systematic study of mimosoids, and (4) to test the applicability of plastid phylogenomics in resolving phylogenetic relationships of mimosoids and trace the evolutionary pattern of IR expansion/contraction.
Materials and methods
Taxa sampling
Fresh leaves and silica-gel dried materials were sampled from 11 species representing 11 genera of three tribes of the subfamily Mimosoideae and one species (Cadellia pentastylis) of Surianaceae (unpublished). The voucher specimens for the ten fresh sampled plants collected from China and Australia were deposited at the Herbarium of Kunming Institute of Botany (KUN). The two silica-gel-dried materials from South Africa were obtained from the University of Johannesburg (JRAU). An Online Resource shows the details of collecting information (see Online Resource 1). Other three mimosoid plastomes obtained from GenBank were also included in analyses. We also included 27 other legume plastomes available in GenBank (Table 1) as closely related outgroups and the newly sequenced C. pentastylis as an outgroup of Fabaceae to reconstruct phylogenetic relationships and to infer the evolutionary pattern of IR expansion/contraction.
Chloroplast DNA extraction and sequencing
Two different methods were used to obtain plastomes in accordance with the availability of fresh leaf material. For the four species with more than 50 g fresh leaves collected from the Kunming Botanical Garden (KBG) and Xishuangbanna Tropical Botanical Garden (XTBG), chloroplast DNA (cpDNA) was extracted using the protocol described in Zhang et al. (2011). For the remaining seven species obtained from Australia and South Africa, total genomic DNA was firstly isolated with a modified cetyl trimethyl ammonium bromide (CTAB) method (Doyle 1987), in which 4% CTAB with approximately 1% polyvinyl polypyrrolidone (PVP) and 0.2% dl-dithiothreitol (DTT) was included (Yang et al. 2014); cpDNA was subsequently amplified using long-range polymerase chain reaction (PCR) with 15 universal primer pairs and methods described in Zhang et al. (2016). The isolated and amplified cpDNA was fragmented to construct short-insert (500 bp) libraries following the manufacturer’s manual (Illumina). Paired-end (PE) sequencing was performed on the Illumina Hiseq 2500 instrument at Beijing Genomics Institute (BGI, Shenzhen, Guangdong, China) and Hiseq 2000 instrument at Plant Germplasm and Genomics Center (Kunming Institute of Botany, Chinese Academy of Sciences, Kunming, China).
Plastome assembly and annotation
Quality control checks on the short reads were performed using NGS QC Tool Kit (Patel and Jain 2012) with default parameters. High-quality short reads were de novo assembled into contigs using CLC Genomics Workbench version 8.5.1 (evaluation version, CLC Bio), with a k-mer of 63 and a minimum contig length of 1000 bp. Subsequently, all contigs were aligned with reference plastomes (plastomes of subfamily Mimosoideae; Dugas et al. 2015; Williams et al. 2015) using Basic Local Alignment Search Tool (BLAST; Altschul et al. 1990) with default search parameters. Then, we determined the proper orders of the aligned contigs according to the reference plastomes, and the gaps between the de novo assembled contigs were filled with consensus sequences of raw reads mapped to the reference plastomes. At this point, the target contigs were assembled into complete plastomes. Finally, we designed primer pairs (see Online Resource 2) to verify the four junctions between the single copy (SC) and IR regions of the plastomes. PCRs were performed in 25.5 μL volume using 12.5 μL Taq DNA polymerase, 0.5 μL each of forward and reverse primers (dissolved in 10× ddH2O), and 1 μL (30 ng/μL) template DNA under the following conditions: 3 min at 94 °C, followed by 35 cycles of 50 s at 94 °C, 2 min at 50 °C, and 1 min at 72 °C, and then a final 8-min extension at 72 °C. PCR products were sequenced at the Kunming Sequencing Department of Biosune Biotechnology Limited Company (Shanghai, China). Sanger sequences and assembled plastomes were aligned using Geneious version 8.1.4 (Kearse et al. 2012) to determine if there were any differences.
Annotation of the plastomes was performed using DOGMA (Wyman et al. 2004), coupled with manual correction of start and stop codons and intron/exon boundaries. The online tRNAscan-SE service (Schattner et al. 2005) was used to further determine tRNA genes. The final complete plastomes were deposited into GenBank with accession numbers KX852435–KX852445 (see Online Resource 1). OrganellarGenomeDRAW (Lohse et al. 2013) was applied to draw physical maps for all newly sequenced plastomes, and an Online Resource shows physical maps of ten species (see Online Resources 3 and 4). To detect the number of mapped paired-end reads and the depth of coverage, paired reads were remapped to the assembled genome using Bowtie 2 (Langmead and Salzberg 2012) implemented in Geneious version 8.1.4.
Repeat analysis
Tandem repeat composition and distribution were identified in 14 mimosoid plastomes using the Tandem Repeat Finder (TRF) web interface (Benson 1999) with default parameters. Repeats with overlapped regions were treated as one for counting repeat size and number. REPuter (Kurtz et al. 2001) was hired to identify dispersed repeats, including forward, reverse, and palindromic repeats. Dispersed repeats at a minimum size of 30 bp were detected using a Hamming distance of 3, corresponding to a sequence identity of over 90%. The IRA of each plastome was removed before the dispersed repeat detection, then the location of repeats in IRA was manually found based on those detected in IRB. Because REPuter overestimates the number of repetitive elements in a given sequence by recognizing nested or overlapping repeats within a given region containing multiple repeats (Curci et al. 2015; Downie and Jansen 2015), all overlapping repeats were manually identified and then adjusted or excluded based on the following criteria: repeats would be excluded when nested wholly within another one or nested in or overlapped mostly with tandem repeats that were found by TRF; if motifs of repeats within a certain region were all recognized as being repetitive with those of another region, only the first motifs within each of the two regions would have remained. Correlation between repeat proportion (total repeat length/plastome size) and plastome size was analyzed using R v.3.3.2. (R Development Core Team 2016). To compare results in the same dimensions, we divided the sampled mimosoids into IR-expanded species and IR-canonical species based on their IRs. Acacia dealbata and Pithecellobium flexicaule were excluded from the analysis of IR-expanded species for additional large IR-LSC junction shifts in their plastomes (see “Results”). We also counted the repeat number, measured the repeat length, and calculated their proportion to each region of LSC, SSC, and IR.
Sequence divergences
Alignment and comparison of all 14 sampled mimosoid plastomes were performed using mVISTA (Frazer et al. 2004) in LAGAN mode. Pithecellobium flexicaule was used as reference because of its largest genome size and most gene content in IR.
To identify sequence divergence in mimosoid plastomes, 111 coding (including protein coding genes, tRNAs, and rRNAs) and 128 noncoding regions (including intergenic spacers and introns) from 14 plastome sequences were extracted and then aligned using MEGA6 (Tamura et al. 2013). The 421-bp rps18-containing inversion in A. dealbata (see “Results”) was manually reversed (boundaries of this inversion were delimited by comparing the plastome of this species to that of Acacia podalyriifolia, an unpublished plastome). The duplicated clpP copy locating between psbZ and trnG GCC of A. dealbata was removed from alignment. Comparisons of extracted coding and noncoding regions including numbers of constant, parsimony uninformative sites, parsimony informative sites (PIS), and range of sequence divergence in pairwise comparisons (uncorrected “p” distance) were conducted by using PAUP* 4.0a147 (Swofford 2002) with treatment of “gap” states as “missing data.”
Synonymous substitution rate shifts of relocated genes
There are 14 protein-coding genes in mimosoid plastomes relocated into or out of IR (see “Results”). Nucleotide synonymous substitution rates were estimated for ten genes (ndhA, ndhD, ndhE, ndhG, ndhH, ndhI, psaC, rpl2, rps3, and rps15); genes ndhF, rpl23, rps19, and ycf1 across IR-SC boundaries were excluded from analyses following Zhu et al. (2016). Sequence alignment for each group of species (out-IR and in-IR) was performed by using MUSCLE (Edgar 2004). Adenanthera microsperma was used as reference for estimating the synonymous substitution rates in each species by using CodeML in PAML v4.8 (Yang 2007) with the options of runmode = −2, seqtype =1, CodonFreq =2. Then, average values of synonymous substitution rates for each data matrix were obtained. A significance test was conducted by using the cor.test function in R v3.3.2 (R Development Core Team 2016).
Phylogenetic analysis and the evolutionary pattern of IR expansion/contraction
All 77 protein-coding genes were extracted and aligned using MAFFT version 7 (Katoh and Standley 2013) implemented in Geneious version 8.1.4 with default parameters. These matrices were then concatenated to a final data matrix (deposited at the TreeBASE repository, http://purl.org/phylo/treebase/phylows/study/TB2:S20385). The maximum likelihood (ML) phylogeny was reconstructed using RAxML version 8.1.1179 (Stamatakis 2014) at the XSEDE Teragrid of the CIPRES science Gateway (Miller et al. 2010), including tree robustness assessment using 1000 replicates of rapid bootstrap with the GTRGAMMA substitution model.
Fitch parsimony (FP, a character optimization method) was performed using Mesquite version 3.10 (build 765; Maddison and Maddison 2016) to trace the evolutionary pattern of the IR expansion/contraction in legumes. The IR was treated as one character and five character states were coded (0, IR-lacked; 1, IR-canonical; 2, IR-expanded into SSC; 3, IR-expanded into LSC; 4, IR-contracted out of LSC). The state scores for each sampled species were listed in Online Resource 5. The newly reconstructed ML phylogeny was applied for tracing the ancestral states of IR.
Results
Plastome organization
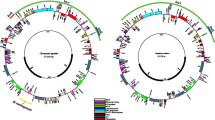
All newly sequenced mimosoid plastomes displayed the typical quadripartite structure, conserved gene content, and a similar gene order. The mean coverage of the 11 newly sequenced mimosoid plastomes ranged from 1011.8× (Pararchidendron pruinosum) to 2521.6× (Parkia javanica). The overall GC content was relatively conserved and around 36% (Table 2). Each plastome contained 111 different genes, including 77 protein-coding, 30 tRNA, and 4 ribosomal RNA (rRNA) genes (Fig. 1, Table 3 and Online Resources 3 and 4). Eighteen genes (six of which are tRNAs) contained introns. The protein-coding genes rps12 and ycf3 contained two introns. The rps12 gene was spliced into two transcriptions, with exon 1 in LSC and exons 2 and 3 in IR. The clpP gene also had introns in our sampled species except Faidherbia albida and Acacia dealbata, each of which lacked both two introns in their clpP. In addition, another duplicated clpP copy was found between psbZ and trnG GCC in A. dealbata. A 421-bp inversion containing rps18 was found in A. dealbata.
Gene maps of the plastome of Acacia dealbata. Genes are indicated by boxes on the inside (clockwise transcription) and outside (counterclockwise transcription) of the outermost circle. The inner circle identifies the major structural components of the plastome (LSC, IR, and SSC). Genes belonging to different functional groups are color-coded. The dashed area in the inner circle indicates the GC content of the plastome
IR expansion and contraction in mimosoid plastomes
Within the sampled mimosoids, six species of Adenanthera, Dichrostachys, Leucaena, Parkia, Piptadenia, and Prosopis belonging to tribe Mimoseae had canonical IRs (ranging from 26,007 bp in P. javanica to 26,142 bp in Dichrostachys cinerea; Table 2) which contain 16 complete genes (including five protein-coding genes, seven tRNAs, and all four rRNAs) and parts of the 5′ end of ycf1 (692–779 bp) and rps19 (91–105 bp) (Fig. 2). The remaining eight species belonging to tribe Ingeae and Acacia s.s. had much longer IRs ranging from 38,254 bp (A. dealbata) to 41,503 bp (Pithecellobium flexicaule), because all of them experienced ca. 13-kb IR expansion into SSC. Accordingly, their IRs included nine more protein-coding genes, with the IRB/SSC junction (JSB) occurring within ndhF, resulting in the duplication of the 3′ end (6–239 bp) of this gene in IRs, and the SSC/IRA junction (JSA) occurring between ccsA and ndhD in these eight species except F. albida, in which the last nucleotide of the stop codon of ccsA was included in IR. The LSC/IRB junction (JLB) of most of the aforementioned eight (IR-expanded) species was also located in rps19, making 98–109 bp of the 5′ end of this gene run into IR. However, a ca. 1.7-kb IR expansion into LSC was detected in P. flexicaule, making its JLB occur between rpl16 and rps3 and two more genes (rps3 and rps19) in IR. In contrast, a ca. 1.9-kb IR contraction out of LSC was found in A. dealbata, making its JLB occur within rpl23, and hence rpl2, rps19, and 199 bp of the 3′ end of rpl23 ran into LSC. Accordingly, the length of LSC within sampled mimosoid plastomes showed relatively moderate variation and ranged from 88,577 bp (Adenanthera microsperma) to 93,690 bp (Leucaena trichandra), whereas the length of SSC varied significantly, ranging from 4573 bp (Archidendron lucyi) to 18,941 bp (Piptadenia communis).
Repeat analyses
Tandem repeats were detected in 54 (A. microsperma)–96 (L. trichandra) sites among IR-canonical species and in 64 (Albizia odoratissima)–104 (P. flexicaule) sites among IR-expanded species (a detailed list of repeat numbers and lengths is shown in Online Resource 6). The total length of tandem repeats ranged from 2902 (A. microsperma) to 7018 bp (P. flexicaule), and the proportion ranged from 1.82% (A. microsperma) to 3.92% (A. dealbata and P. flexicaule). After excluding the overlapped repeats detected by REPuter and TRF and taking one IR copy into account, 11 (A. odoratissima)–57 (L. trichandra) pairs of dispersed repeats were identified: 12 (A. microsperma)–57 (L. trichandra) within IR-canonical species and 11 (A. odoratissima)–45 (P. flexicaule) within IR-expanded species. In the majority of these species, the most dispersed repeats were forward, then palindromic and the least reverse. The length and proportion of the dispersed repeats ranged from 813 and 0.46% (A. odoratissima) to 3222 and 1.85% (A. dealbata). Correlation (p < 0.05) was found between both tandem and dispersed repeat proportion and plastome size in both IR-expanded species and IR-canonical species (Table 4). The LSC regions of all studied mimosoid plastomes possessed the highest proportion of both tandem and dispersed repeats (Fig. 3 and Online Resource 6). Within IR-canonical species, the SSCs contained a higher repeat proportion than IRs. Within IR-expanded species, SSC had a higher repeat proportion than IR in some species, while less in other species.
Repeat proportion in different regions of mimosoid plastomes. a Tandem repeats. b Dispersed repeats. Species are ordered by plastome size from small to large. LSC large single copy, SSC, small single copy, IR inverted repeat. The yellow-dashed area refers to IR-conical species, while the gray-dashed area refers to IR-expanded species
Identification of sequence divergence
Comparison of sampled mimosoid plastomes using mVISTA revealed that noncoding regions were more diverged than coding regions. Generally, noncoding regions located in SCs were more divergent than those in IRs (Fig. 4).
Aligned length for each of 111 coding and 128 noncoding regions ranged from 9 bp (psbF-psbE) to 6957 bp (ycf2). The number of PIS ranged from 0 (for 45 loci) to 302 (trnS GCU-trnG UCC). The number of variable sites ranged from 0 (for 17 loci) to 948 (ycf1). The five regions with relatively high sequence divergence were ccsA-ndhD (3.56–24.80%), clpP (0.68–29.25%), psaA-ycf3 (0.83–28.95%), rps12-clpP (1.49–26.04%), and trnS GCU-trnG UCC (2.77–25.41%), while those with no sequence divergence were mostly tRNAs and rrn5. An Online Resource shows the detailed comparative results (see Online Resource 7). Percentages of variable sites and PIS in coding and noncoding regions are shown in Fig. 5. As to coding regions, accD, clpP, rpl32, rps3, and ycf1 had high percentage of variable sites and PIS, with clpP having an especially high PIS percentage (19.59%) comparing with the four other regions (all less than 5%) (Fig. 5a). The percentages of both variable sites and PIS of noncoding regions were much higher than those of coding regions. The top five variable regions were accD-psaI, ccsA-ndhD, psaA-ycf3, psbI-trnS GCU, and trnS GCU-trnG UCC. The top five PIS regions were accD-psaI, ccsA-ndhD, psbI-trnS GCU, rpl33-rps18, and trnS GCU-trnG UCC (Fig. 5b).
Synonymous substitution rate shifts of relocated genes
Nucleotide synonymous substitution rate shifts for genes relocated into and out of IR were shown in Online Resource 8. Most of these genes had higher synonymous substitution rates in SC than those in IR (Fig. 6). Linear regression analysis showed decreased synonymous substitution rates of genes relocating from SSC into IR (y = 0.8899x). However, the synonymous substitution rate of rps3 increased following translocation from LSC into IR.
Synonymous substitution rate shifts of relocated genes. Hollow circles refer to the substitution rates of ndhA, ndhD, ndhE, ndhG, ndhH, ndhI, psaC, rpl2, rps3, and rps15 in-IR (ordinate) and out-IR (abscissa). The red line shows the best-fit regression (constrained to pass through the origin); the black diagonal line refers to the y = x line
Phylogenetic relationships and evolutionary pattern of IR expansion/contraction in mimosoids
Phylogenetic analysis based on 77 protein coding genes well resolved relationships among sampled species from three subfamilies with high bootstrap supports (BS) (Fig. 7). Mimosoideae and Papilionoideae were strongly supported to be monophyletic (BS = 100) and nested within the paraphyletic Caesalpinioideae. Phylogenetic relationships among sampled mimosoids were resolved with high supports. The clade comprising Ingeae and Acacia s.s. (nested within Ingeae) was strongly supported (BS = 100). The tribe Mimoseae was supported to be paraphyletic.
The state IR-canonical was traced as plesiomorphy of Fabaceae (Fig. 8). The states IR-expanding into SSC and IR-lacked were synapomorphies of the Ingeae + Acacia s.s. clade and the IRLC, respectively. The state IR-expanded into LSC and IR-contracted out of LSC evolved separately in different species within the Ingeae + Acacia s.s. clade, which should be their autapomorphies.
Discussion
This study revealed moderate variation in mimosoid plastomes that is consistent with and adds to what was found by Dugas et al. (2015). The previously reported ca. 13-kb IR-SSC boundary shift (Dugas et al. 2015) was detected in plastomes of all sampled species within the clade formed by Ingeae and Acacia s.s. and led to a major increase of plastome size. In addition, some structural variations were firstly found in this subfamily, including a 421-bp inversion containing rps18 and a duplicated copy of clpP in plastome of Acacia dealbata, and the loss of both introns of clpP in plastomes of A. dealbata and Faidherbia albida (Fig. 1 and Online Resources 3 and 4).
Structural and size variation of mimosoid plastomes
The shift of IR-SC boundaries played an important role in mimosoid plastome size variation (Dugas et al. 2015). The ca. 13-kb IR expansion into SSC resulted in significant larger legume plastomes in the Ingeae + Acacia s.s. clade. Plastome of Pithecellobium flexicaule experienced further IR expansion into LSC, which led to the largest known legume plastome (Fig. 2). In contrast, plastome of A. dealbata, which experienced IR contraction out of LSC, was the shortest within this clade.
Goulding et al. (1996) proposed a double-strand break (DSB) model that starts with a double-strand break followed by strand invasion and recombination to explain the IR expansion in Nicotiana acuminata. This model was applied by Wang et al. (2008) to explain IR expansions in many angiosperm lineages. Previous studies also suggested that regions with high content of short repeats or “poly A tract” are closely associated with the dynamics of IR-LSC junctions and expansions of IR (Dugas et al. 2015; Wang et al. 2008). In all eight mimosoid plastomes that exhibit ca. 13-kb IR expansion into SSC, the ca. 100-bp regions upstream of the IR-SSC junctions are extremely AT-rich (>90%) and have many poly A tracts and short repeats. Initiating with the DSBs and ending in the AT-rich or poly A-tract regions could be applied to explain the large IR expansion of mimosoid plastomes. Large IR contractions have been rarely reported; illegitimate recombination has been considered as the most plausible explanation (Blazier et al. 2016; Downie and Jansen 2015; Goulding et al. 1996).
The duplicated copy of clpP was found to be located between psbZ and trnG GCC in A. dealbata (Fig. 1). Gene duplications caused by IR are common in plastomes and is believed to be an important driving force in the evolution of genomes, leading to the creation of new genes and new gene functions (Xiong et al. 2009). Gene duplication is previously reported in multiple angiosperm lineages and most of them are tRNAs (Drabkova et al. 2004; Haberle et al. 2008; Hipkins et al. 1995; Koch et al. 2005; Lee et al. 2007; Lin et al. 2012; Vijverberg and Bachmann 1999). Duplication of protein-coding genes outside of the IR is rare, which have been reported in Pinus thunbergii (psaM; Wakasugi et al. 1994), Jasminum and Poaceae (rpl23; Lee et al. 2007; Xiong et al. 2009), Silene and Lychnis (clpP; Erixon and Oxelman 2008), Trachelium (psbJ; Haberle et al. 2008) and Pelargonium (many genes; Chumley et al. 2006). Haberle et al. (2008) supposed that duplicative transposition may be responsible for the three complete copies of psbJ occurring in the LSC region of Trachelium. Same mechanism could also be applied to explain the duplication of the clpP in A. dealbata. Any yet, the underlying mechanisms of gene duplication in SCs of plastome remain unclear (Lee et al. 2007).
Effect of repeats on mimosoid plastome
Plastome size variation were previously reported to be contributive to tandem (Dugas et al. 2015; Jo et al. 2011) and dispersed repeats (Cosner et al. 1997; Guisinger et al. 2011; Haberle et al. 2008; Ogihara et al. 1988; Weng et al. 2014). Congruent with Dugas et al. (2015), our analyses illustrated that both tandem and dispersed repeat contributed to plastome size of mimosoids (p < 0.05; Table 4), and LSC contains the highest repeat proportion and contributed most to plastome size (Fig. 3 and Online Resource 6). Repeats were previously inferred to associate with plastome structural variation (Cai et al. 2008; Cosner et al. 1997; Greiner et al. 2008). In this study, the plastome of A. dealbata possessed an extremely high repeat proportion, and presented the most structural variations. These together supposed that repeats may also affect both structure and size variation in mimosoid plastomes.
Identification of plastome sequence divergence
Plastomes supply valuable markers for reconstructing phylogenetic relationships. Many fragments of coding regions, introns, and intergenic spacers, including atpB, atpB-rbcL, matK, ndhF, rbcL, rpl16, rps4-trnS, rps16, trnH-psbA, trnL-F, trnS-G, etc., have been used for phylogenetic reconstructions at various taxonomic levels (Gao et al. 2008; Hilu et al. 2008; Kim and Jansen 1995; Li 2008; Peterson et al. 2010; Wilson 2009). Some plastome regions, such as atpF-H, matK, psbK-I, rbcL, rpoB, rpoC1, trnH-psbA, etc., have been relied upon heavily for development of candidate markers for plant DNA barcoding (Chase et al. 2007; Dong et al. 2012; Hollingsworth et al. 2011; Kress et al. 2005; Newmaster et al. 2006). The noncoding trnH-psbA and coding matK have been applied to distinguish the sister species complex of pantropical Acacia (Newmaster and Ragupathy 2009). Mimosoid plastomes contain multiple variable regions. The noncoding regions located in SCs show higher divergence than those in IRs (Fig. 4), which is accordant with previous studies (reviewed by Zhu et al. 2016). Many of their moderately evolved gene loci could be applied to resolve deep and intergeneric relationships of this subfamily. Multiple fast-evolving loci such as psaA-ycf3, ccsA-ndhD, trnS GCU-trnG UCC, psbI-trnS GCU, and accD-psaI could be applied to resolve inter- or intraspecific relationships (Fig. 5 and Online Resource 7).
Substitution rate shift of relocated genes
Nucleotide substitution rates of plastome coding genes have been proved to be decreased after translocating from SC regions to IR in many plant lineages (Li et al. 2016; Lin et al. 2012; Perry and Wolfe 2002; Zhu et al. 2016). In this study, we again found a decrease of substitution rates for genes shifted into IR (Fig. 6). One exception is that rps3 had a higher substitution rate after relocation into IR. Relocation of rps3 to IR may occur too recent for a rate decreasing to a detectable level; a similar phenomenon was detected for ycf2 in Ginkgo biloba (Lin et al. 2012).
Phylogenetic analysis and evolutionary pattern of IR expansion/contraction
The deep relationships of mimosoids were poorly resolved by phylogenetic studies applying a few plastid markers (Bouchenak-Khelladi et al. 2010; Luckow et al. 2003; Luckow et al. 2000; Miller et al. 2003; Miller and Seigler 2012). Plastid phylogenomics has been proved to be efficient to resolve difficult relationships at family level like Orchidaceae (Givnish et al. 2015) and lower taxonomic level such as subfamilies Bambusoideae (Wysocki et al. 2015; Zhang et al. 2011) and Chloridoideae (Duvall et al. 2016) of Poaceae. In this study, plastid phylogenomics was proved to be an efficient way to resolve relationships of Mimosoideae and those of Fabaceae (Fig. 7), which could be applied to reconstruct a robust mimosoid or even legume phylogeny with systematic sampling.
Large structural rearrangements of plastome have been proved to have a strong phylogenetic signal and were applied to define monophyletic lineages in many plant groups (Downie and Palmer 1992; Raubeson and Jansen 2004). For instance, the IRLC of Papilionoideae lacks IR (Lavin et al. 1990; Wojciechowski et al. 2000), and the “Aegopodium group” and “Apium group” of Apiaceae are characterized by large IR expansion and contraction, respectively (Downie and Jansen 2015). In Mimosoideae, tribe Ingeae and Acacia s.s. (nested within Ingeae) formed a strongly supported clade, and “IR-expanded into SSC” was explored to be synapomorphy of the Ingeae + Acacia s.s. clade (Fig. 8). This clade could be named as “inverted-repeat-expanding clade (IREC)” comparing with the IRLC in Papilionoideae.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Barrett CF, Specht CD, Leebens-Mack J, Stevenson DW, Zomlefer WB, Davis JI (2014) Resolving ancient radiations: can complete plastid gene sets elucidate deep relationships among the tropical gingers (Zingiberales)? Ann Bot 113:119–133
Benson G (1999) Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27:573–580
Bewick AJ, Chain FJJ, Heled J, Evans BJ (2012) The pipid root. Syst Biol 61:913–926
Blazier J, Guisinger MM, Jansen RK (2011) Recent loss of plastid-encoded ndh genes within Erodium (Geraniaceae). Plant Mol Biol 76:263–272
Blazier JC, Jansen RK, Mower JP, Govindu M, Zhang J, Weng ML, Ruhlman TA (2016) Variable presence of the inverted repeat and plastome stability in Erodium. Ann Bot 117:1209–1220
Bock R (2007) Structure, function, and inheritance of plastid genomes. In: Bock R (ed) Cell and molecular biology of plastids. Springer Berlin Heidelberg, Berlin, Heidelberg, pp 29–63
Bouchenak-Khelladi Y, Maurin O, Hurter J, van der Bank M (2010) The evolutionary history and biogeography of Mimosoideae (Leguminosae): an emphasis on African acacias. Mol Phylogen Evol 57:495–508
Cai ZQ, Guisinger M, Kim HG, Ruck E, Blazier JC, McMurtry V, Kuehl JV, Boore J, Jansen RK (2008) Extensive reorganization of the plastid genome of Trifolium subterraneum (Fabaceae) is associated with numerous repeated sequences and novel DNA insertions. J Mol Evol 67:696–704
Chase MW, Cowan RS, Hollingsworth PM, van den Berg C, Madrinan S, Petersen G, Seberg O, Jorgsensen T, Cameron KM, Carine M, Pedersen N, Hedderson TAJ, Conrad F, Salazar GA, Richardson JE, Hollingsworth ML, Barraclough TG, Kelly L, Wilkinson M (2007) A proposal for a standardised protocol to barcode all land plants. Taxon 56:295–299
Choi IS, Kim JH, Choi BH (2015) Complete plastid genome of Astragalus mongholicus var. nakaianus (Fabaceae). Mitochondr DNA. doi:10.3109/19401736.2015.1053118
Chumley TW, Palmer JD, Mower JP, Fourcade HM, Calie PJ, Boore JL, Jansen RK (2006) The complete chloroplast genome sequence of Pelargonium × hortorum: organization and evolution of the largest and most highly rearranged chloroplast genome of land plants. Mol Biol Evol 23:2175–2190
Cosner ME, Jansen RK, Palmer JD, Downie SR (1997) The highly rearranged chloroplast genome of Trachelium caeruleum (Campanulaceae): multiple inversions, inverted repeat expansion and contraction, transposition, insertions/deletions, and several repeat families. Curr Genet 31:419–429
Curci PL, De Paola D, Danzi D, Vendramin GG, Sonnante G (2015) Complete chloroplast genome of the multifunctional crop globe artichoke and comparison with other Asteraceae. PLoS One 10:e0120589
Delsuc F, Brinkmann H, Philippe H (2005) Phylogenomics and the reconstruction of the tree of life. Nat Rev Genet 6:361–375
Dong WP, Liu J, Yu J, Wang L, Zhou SL (2012) Highly variable chloroplast markers for evaluating plant phylogeny at low taxonomic levels and for DNA barcoding. PLoS One 7:e35071
Downie SR, Jansen RK (2015) A comparative analysis of whole plastid genomes from the Apiales: expansion and contraction of the inverted repeat, mitochondrial to plastid transfer of DNA, and identification of highly divergent noncoding regions. Syst Bot 40:336–351
Downie SR, Palmer JD (1992) Use of chloroplast DNA rearrangements in reconstructing plant phylogeny. In: Soltis PS, Soltis DE, Doyle JJ (eds) Molecular systematics of plants. Springer, Chapman & Hall, New York, NY, pp 14–35
Doyle JJ (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem bull 19:11–15
Doyle JJ, Doyle JL, Ballenger JA, Palmer JD (1996) The distribution and phylogenetic significance of a 50-kb chloroplast DNA inversion in the flowering plant family Leguminosae. Mol Phylogen Evol 5:429–438
Drabkova L, Kirschner J, Vlcek C, Paces V (2004) TrnL-trnF intergenic spacer and trnL intron define major clades within Luzula and Juncus (Juncaceae): importance of structural mutations. J Mol Evol 59:1–10
Dugas DV, Hernandez D, Koenen EJM, Schwarz E, Straub S, Hughes CE, Jansen RK, Nageswara-Rao M, Staats M, Trujillo JT, Hajrah NH, Alharbi NS, Al-Malki AL, Sabir JSM, Bailey CD (2015) Mimosoid legume plastome evolution: IR expansion, tandem repeat expansions, and accelerated rate of evolution in clpP. Sci Rep 5:16958
Duvall MR, Fisher AE, Columbus JT, Ingram AL, Wysocki WP, Burke SV, Clark LG, Kelchner SA (2016) Phylogenomics and plastome evolution of the chloridoid grasses (Chloridoideae: Poaceae). Int J Plant Sci 177:235–246
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5:1–19
Eisen JA (1998) Phylogenomics: improving functional predictions for uncharacterized genes by evolutionary analysis. Genome Res 8:163–167
Eisen JA, Fraser CM (2003) Phylogenomics: intersection of evolution and genomics. Science 300:1706–1707
Erixon P, Oxelman B (2008) Whole-gene positive selection, elevated synonymous substitution rates, duplication, and indel evolution of the chloroplast clpP1 gene. PLoS One 3:e1386
Frazer KA, Pachter L, Poliakov A, Rubin EM, Dubchak I (2004) VISTA: computational tools for comparative genomics. Nucleic Acids Res 32:W273–W279
Gao X, Zhu YP, Wu BC, Zhao YM, Chen JQ, Hang YY (2008) Phylogeny of Dioscorea sect. Stenophora based on chloroplast matK, rbcL and trnL-F sequences. J Syst Evol 46:315–321
Givnish TJ, Spalink D, Ames M, Lyon SP, Hunter SJ, Zuluaga A, Iles WJD, Clements MA, Arroyo MTK, Leebens-Mack J, Endara L, Kriebel R, Neubig KM, Whitten WM, Williams NH, Cameron KM (2015) Orchid phylogenomics and multiple drivers of their extraordinary diversification. Proc R Soc B Biol Sci 282:20151553
Goulding SE, Olmstead RG, Morden CW, Wolfe KH (1996) Ebb and flow of the chloroplast inverted repeat. Mol Gen Genet 252:195–206
Greiner S, Wang X, Rauwolf U, Silber MV, Mayer K, Meurer J, Haberer G, Herrmann RG (2008) The complete nucleotide sequences of the five genetically distinct plastid genomes of Oenothera, subsection Oenothera: I. Sequence evaluation and plastome evolution. Nucleic Acids Res 36:2366–2378
Grewe F, Guo WH, Gubbels EA, Hansen AK, Mower JP (2013) Complete plastid genomes from Ophioglossum californicum, Psilotum nudum, and Equisetum hyemale reveal an ancestral land plant genome structure and resolve the position of Equisetales among monilophytes. BMC Evol Biol 13. doi:10.1186/1471-2148-13-8
Guisinger MM, Kuehl JV, Boore JL, Jansen RK (2011) Extreme reconfiguration of plastid genomes in the angiosperm family Geraniaceae: rearrangements, repeats, and codon usage (vol 28, pg 583, 2011). Mol Biol Evol 28:1543
Guo W, Grewe F, Cobo-Clark A, Fan W, Duan Z, Adams RP, Schwarzbach AE, Mower JP (2014) Predominant and substoichiometric isomers of the plastid genome coexist within Juniperus plants and have shifted multiple times during cupressophyte evolution. Genome Biol Evol 6:580–590
Guo XW, Castillo-Ramirez S, Gonzalez V, Bustos P, Fernandez-Vazquez JL, Santamaria RI, Arellano J, Cevallos MA, Davila G (2007) Rapid evolutionary change of common bean (Phaseolus vulgaris L.) plastome, and the genomic diversification of legume chloroplasts. BMC Genomics 8. doi:10.1186/1471-2164-8-228
Haberle RC, Fourcade HM, Boore JL, Jansen RK (2008) Extensive rearrangements in the chloroplast genome of Trachelium caeruleum are associated with repeats and tRNA genes. J Mol Evol 66:350–361
Hilu KW, Black C, Diouf D, Burleigh JG (2008) Phylogenetic signal in matK vs. trnK: a case study in early diverging eudicots (angiosperms). Mol Phylogen Evol 48:1120–1130
Hipkins VD, Marshall KA, Neale DB, Rottmann WH, Strauss SH (1995) A mutation hotspot in the chloroplast genome of a conifer (Douglas-fir, Pseudotsuga) is caused by variability in the number of direct repeats derived from a partially duplicated tRNA gene. Curr Genet 27:572–579
Hollingsworth PM, Graham SW, Little DP (2011) Choosing and using a plant DNA barcode. PLoS One 6:e19254
Hoot SB, Palmer JD (1994) Structural rearrangements, including parallel inversions, within the chloroplast genome of Anemone and related genera. J Mol Evol 38:274–281
Jansen RK, Cai Z, Raubeson LA, Daniell H, Depamphilis CW, Leebens-Mack J, Muller KF, Guisinger-Bellian M, Haberle RC, Hansen AK, Chumley TW, Lee SB, Peery R, McNeal JR, Kuehl JV, Boore JL (2007) Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc Natl Acad Sci U S A 104:19369–19374
Jansen RK, Ruhlman TA (2012) Plastid genomes of seed plants. In: Bock R, Knoop V (eds) Genomics of chloroplasts and mitochondria. Springer Netherlands, Dordrecht, pp 103–126
Jansen RK, Wojciechowski MF, Sanniyasi E, Lee SB, Daniell H (2008) Complete plastid genome sequence of the chickpea (Cicer arietinum) and the phylogenetic distribution of rps12 and clpP intron losses among legumes (Leguminosae). Mol Phylogen Evol 48:1204–1217
Jian SG, Soltis PS, Gitzendanner MA, Moore MJ, Li R, Hendry TA, Qiu YL, Dhingra A, Bell CD, Soltis DE (2008) Resolving an ancient, rapid radiation in Saxifragales. Syst Biol 57:38–57
Jo YD, Park J, Kim J, Song W, Hur CG, Lee YH, Kang BC (2011) Complete sequencing and comparative analyses of the pepper (Capsicum annuum L.) plastome revealed high frequency of tandem repeats and large insertion/deletions on pepper plastome. Plant Cell Rep 30:217–229
Käss E, Wink M (1996) Molecular evolution of the Leguminosae: phylogeny of the three subfamilies based on rbcL-sequences. Biochem Syst Ecol 24:365–378
Kato T, Kaneko T, Sato S, Nakamura Y, Tabata S (2000) Complete structure of the chloroplast genome of a legume, Lotus japonicus. DNA Res 7:323–330
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kazakoff SH, Imelfort M, Edwards D, Koehorst J, Biswas B, Batley J, Scott PT, Gresshoff PM (2012) Capturing the biofuel wellhead and powerhouse: the chloroplast and mitochondrial genomes of the leguminous feedstock tree Pongamia pinnata. PLoS One 7:e51687
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Kim KJ, Jansen RK (1995) NdhF sequence evolution and the major clades in the sunflower family. Proc Natl Acad Sci U S A 92:10379–10383
Kim YD, Jansen RK (1994) Characterization and phylogenetic distribution of a chloroplast DNA rearrangement in the Berberidaceae. Plant Syst Evol 193:107–114
Koch MA, Dobes C, Matschinger M, Bleeker W, Vogel J, Kiefer M, Mitchell-Olds T (2005) Evolution of the trnF(GAA) gene in Arabidopsis relatives and the Brassicaceae family: monophyletic origin and subsequent diversification of a plastidic pseudogene. Mol Biol Evol 22:1032–1043
Kress WJ, Wurdack KJ, Zimmer EA, Weigt LA, Janzen DH (2005) Use of DNA barcodes to identify flowering plants. Proc Natl Acad Sci U S A 102:8369–8374
Kurtz S, Choudhuri JV, Ohlebusch E, Schleiermacher C, Stoye J, Giegerich R (2001) REPuter: the manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res 29:4633–4642
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359
Lavin M, Doyle JJ, Palmer JD (1990) Evolutionary significance of the loss of the chloroplast-DNA inverted repeat in the Leguminosae subfamily Papilionoideae. Evolution 44:390–402
Lee HL, Jansen RK, Chumley TW, Kim KJ (2007) Gene relocations within chloroplast genomes of Jasminum and Menodora (Oleaceae) are due to multiple, overlapping inversions. Mol Biol Evol 24:1161–1180
Lei WJ, Ni DP, Wang YJ, Shao JJ, Wang XC, Yang D, Wang JS, Chen HM, Liu C (2016) Intraspecific and heteroplasmic variations, gene losses and inversions in the chloroplast genome of Astragalus membranaceus. Sci Rep 6:21669
Lewis GP, Schrire BD, Mackinder BA, Lock M (2005) Legumes of the world. Royal Botanic Gardens, Kew, Richmond, U.K
Li FW, Kuo LY, Pryer KM, Rothfels CJ (2016) Genes translocated into the plastid inverted repeat show decelerated substitution rates and elevated GC content. Genome Biol Evol 8:2452–2458
Li JH (2008) Phylogeny of Catalpa (Bignoniaceae) inferred from sequences of chloroplast ndhF and nuclear ribosomal DNA. J Syst Evol 46:341–348
Lin CP, Wu CS, Huang YY, Chaw SM (2012) The complete chloroplast genome of Ginkgo biloba reveals the mechanism of inverted repeat contraction. Genome Biol Evol 4:374–381
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW—a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res 41:W575–W581
LPWG (2013) Legume phylogeny and classification in the 21st century: progress, prospects and lessons for other species-rich clades. Taxon 62:217–248
Luckow M, Miller JT, Murphy DJ, Livshultz T (2003) A phylogenetic analysis of the Mimosoideae (Leguminosae) based on chloroplast DNA sequence data. In: Klitgaard BB, Bruneau A (eds) Advances in legume systematics, part 10. Higher level systematics. Royal Botanic Gardens, Kew, Richmond, U.K., pp 197–220
Luckow M, White PJ, Bruneau A (2000) Relationships among the basal genera of mimosoid legumes. In: Herendeen PS, Bruneau A (eds) Advances in legume systematics, part 9. Royal Botanic Gardens, Kew, Richmond, U.K., pp 165–180
Luo Y, Ma PF, Li HT, Yang JB, Wang H, Li DZ (2016) Plastid phylogenomic analyses resolve Tofieldiaceae as the root of the early diverging monocot order Alismatales. Genome Biol Evol 8:932–945
Maddison WP, Maddison DR (2016) Mesquite: a modular system for evolutionary analysis. Version 3.10. http://mesquiteproject.org
Magee AM, Aspinall S, Rice DW, Cusack BP, Semon M, Perry AS, Stefanovic S, Milbourne D, Barth S, Palmer JD, Gray JC, Kavanagh TA, Wolfe KH (2010) Localized hypermutation and associated gene losses in legume chloroplast genomes. Genome Res 20:1700–1710
Martin GE, Rousseau-Gueutin M, Cordonnier S, Lima O, Michon-Coudouel S, Naquin D, de Carvalho JF, Ainouche M, Salmon A, Ainouche A (2014) The first complete chloroplast genome of the Genistoid legume Lupinus luteus: evidence for a novel major lineage-specific rearrangement and new insights regarding plastome evolution in the legume family. Ann Bot 113:1197–1210
Miller JT, Grimes JW, Murphy DJ, Bayer RJ, Ladiges PY (2003) A phylogenetic analysis of the Acacieae and Ingeae (Mimosoideae: Fabaceae) based on trnK, matK, psbA-trnH, and trnL/trnF sequence data. Syst Bot 28:558–566
Miller JT, Seigler D (2012) Evolutionary and taxonomic relationships of Acacia s.l. (Leguminosae: Mimosoideae). Aust Syst Bot 25:217–224
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. Gateway Computing Environments Workshop (GCE), 2010. IEEE, pp 1–8. doi: 10.1109/GCE.2010.5676129
Moore MJ, Bell CD, Soltis PS, Soltis DE (2007) Using plastid genome-scale data to resolve enigmatic relationships among basal angiosperms. Proc Natl Acad Sci U S A 104:19363–19368
Moore MJ, Soltis PS, Bell CD, Burleigh JG, Soltis DE (2010) Phylogenetic analysis of 83 plastid genes further resolves the early diversification of eudicots. Proc Natl Acad Sci U S A 107:4623–4628
Newmaster SG, Fazekas AJ, Ragupathy S (2006) DNA barcoding in land plants: evaluation of rbcL in a multigene tiered approach. Can J Bot 84:335–341
Newmaster SG, Ragupathy S (2009) Testing plant barcoding in a sister species complex of pantropical Acacia (Mimosoideae, Fabaceae). Mol Ecol Resour 9:172–180
Ogihara Y, Terachi T, Sasakuma T (1988) Intramolecular recombination of chloroplast genome mediated by short direct-repeat sequences in wheat species. Proc Natl Acad Sci U S A 85:8573–8577
Palmer JD (1991) Plastid chromosomes: structure and evolution. In: Biogorad L, Vasil IK (eds) Cell culture and somatic cell genetics of plants, The molecular biology of plastids, vol 7. Academic Press, San Diego, California, pp 5–53
Palmer JD, Nugent JM, Herbon LA (1987a) Unusual structure of geranium chloroplast DNA: a triple-sized inverted repeat, extensive gene duplications, multiple inversions, and two repeat families. Proc Natl Acad Sci U S A 84:769–773
Palmer JD, Osorio B, Aldrich J, Thompson WF (1987b) Chloroplast DNA evolution among legumes—loss of a large inverted repeat occurred prior to other sequence rearrangements. Curr Genet 11:275–286
Palmer JD, Osorio B, Thompson WF (1988) Evolutionary significance of inversions in legume chloroplast DNAs. Curr Genet 14:65–74
Patel RK, Jain M (2012) NGS QC toolkit: a toolkit for quality control of next generation sequencing data. PLoS One 7:e30619
Perry AS, Wolfe KH (2002) Nucleotide substitution rates in legume chloroplast DNA depend on the presence of the inverted repeat. J Mol Evol 55:501–508
Peterson PM, Romaschenko K, Johnson G (2010) A classification of the Chloridoideae (Poaceae) based on multi-gene phylogenetic trees. Mol Phylogen Evol 55:580–598
Plunkett GM, Downie SR (2000) Expansion and contraction of the chloroplast inverted repeat in Apiaceae subfamily Apioideae. Syst Bot 25:648–667
Raubeson LA, Jansen RK (2004) Chloroplast genomes of plants. In: Henry RJ (ed) Plant diversity and evolution: genotypic and phenotypic variation in higher plants. CABI Publishing, Wallingford, Oxfordshire, UK, pp 45–68
R Development Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical computing, Vienna. http://www.r-project.org/
Ruhfel BR, Gitzendanner MA, Soltis PS, Soltis DE, Burleigh JG (2014) From algae to angiosperms-inferring the phylogeny of green plants (Viridiplantae) from 360 plastid genomes. BMC Evol Biol 14. doi:10.1186/1471-2148-14-23
Sabir J, Schwarz E, Ellison N, Zhang J, Baeshen NA, Mutwakil M, Jansen R, Ruhlman T (2014) Evolutionary and biotechnology implications of plastid genome variation in the inverted-repeat-lacking clade of legumes. Plant Biotechnol J 12:743–754
Saski C, Lee SB, Daniell H, Wood TC, Tomkins J, Kim HG, Jansen RK (2005) Complete chloroplast genome sequence of Glycine max and comparative analyses with other legume genomes. Plant Mol Biol 59:309–322
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:W686–W689
Shen GF, Chen K, Wu M, Kung SD (1982) Nicotiana chloroplast genome. 4. N. accuminata has larger inverted repeats and genome size. Mol Gen Genet 187:12–18
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Stein DB, Palmer JD, Thompson WF (1986) Structural evolution and flip-flop recombination of chloroplast DNA in the fern genus Osmunda. Curr Genet 10:835–841
Sveinsson S, Cronk Q (2014) Evolutionary origin of highly repetitive plastid genomes within the clover genus (Trifolium). BMC Evol Biol 14. doi:10.1186/s12862-014-0228-6
Swofford D (2002) PAUP*: phylogenetic analysis using parsimony (* and other methods), version 4. Sinauer Associates, Sunderland, MA. doi:10.1111/j.0014-3820.2002.tb00191.x
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Vijverberg K, Bachmann K (1999) Molecular evolution of a tandemly repeated trnF(GAA) gene in the chloroplast genomes of Microseris (Asteraceae) and the use of structural mutations in phylogenetic analyses. Mol Biol Evol 16:1329–1340
Wakasugi T, Tsudzuki J, Ito S, Nakashima K, Tsudzuki T, Sugiura M (1994) Loss of all ndh genes as determined by sequencing the entire chloroplast genome of the black pine Pinus thunbergii. Proc Natl Acad Sci U S A 91:9794–9798
Wang RJ, Cheng CL, Chang CC, Wu CL, Su TM, Chaw SM (2008) Dynamics and evolution of the inverted repeat-large single copy junctions in the chloroplast genomes of monocots. BMC Evol Biol 8. doi:10.1186/1471-2148-8-36
Weng ML, Blazier JC, Govindu M, Jansen RK (2014) Reconstruction of the ancestral plastid genome in Geraniaceae reveals a correlation between genome rearrangements, repeats, and nucleotide substitution rates. Mol Biol Evol 31:645–659
Wicke S, Muller KF, de Pamphilis CW, Quandt D, Wickett NJ, Zhang Y, Renner SS, Schneeweiss GM (2013) Mechanisms of functional and physical genome reduction in photosynthetic and nonphotosynthetic parasitic plants of the broomrape family. Plant Cell 25:3711–3725
Wickett NJ et al (2014) Phylotranscriptomic analysis of the origin and early diversification of land plants. Proc Natl Acad Sci U S A 111:E4859–E4868
Williams AV, Boykin LM, Howell KA, Nevill PG, Small I (2015) The complete sequence of the Acacia ligulata chloroplast genome reveals a highly divergent clpP1 gene (vol 10, e0125768, 2015). PLoS One 10:e0138367
Wilson CA (2009) Phylogenetic relationships among the recognized series in Iris section Limniris. Syst Bot 34:277–284
Wojciechowski MF, Sanderson MJ, Steele KP, Liston A (2000) Molecular phylogeny of the “temperate herbaceous tribes” of papilionoid legumes: a supertree approach. In: Herendeen PS, Bruneau A (eds) Advances in legume systematics, part 9. Royal Botanic Gardens, Kew, Richmond, U.K., pp 277–298
Wu CS, Chaw SM (2014) Highly rearranged and size-variable chloroplast genomes in conifers II clade (cupressophytes): evolution towards shorter intergenic spacers. Plant Biotechnol J 12:344–353
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255
Wysocki WP, Clark LG, Attigala L, Ruiz-Sanchez E, Duvall MR (2015) Evolution of the bamboos (Bambusoideae; Poaceae): a full plastome phylogenomic analysis. BMC Evol Biol 15. doi:10.1186/s12862-015-0321-5
Xi ZX, Ruhfel BR, Schaefer H, Amorim AM, Sugumaran M, Wurdack KJ, Endress PK, Matthews ML, Stevens PF, Mathews S, Davis CC (2012) Phylogenomics and a posteriori data partitioning resolve the cretaceous angiosperm radiation Malpighiales. Proc Natl Acad Sci U S A 109:17519–17524
Xiong AS, Peng RH, Zhuang J, Gao F, Zhu B, Fu XY, Xue Y, Jin XF, Tian YS, Zhao W, Yao QH (2009) Gene duplication, transfer, and evolution in the chloroplast genome. Biotechnol Adv 27:340–347
Yang JB, Li DZ, Li HT (2014) Highly effective sequencing whole chloroplast genomes of angiosperms by nine novel universal primer pairs. Mol Ecol Resour 14:1024–1031
Yang ZH (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Zhang T, Zeng CX, Yang JB, Li HT, Li DZ (2016) Fifteen novel universal primer pairs for sequencing whole chloroplast genomes and a primer pair for nuclear ribosomal DNAs. J Syst Evol 54:219–227
Zhang YJ, Ma PF, Li DZ (2011) High-throughput sequencing of six bamboo chloroplast genomes: phylogenetic implications for temperate woody bamboos (Poaceae: Bambusoideae). PLoS One 6:e20596
Zhong BJ, Yonezawa T, Zhong Y, Hasegawa M (2010) The position of Gnetales among seed plants: overcoming pitfalls of chloroplast phylogenomics. Mol Biol Evol 27:2855–2863
Zhou XM, Xu SX, Xu JX, Chen BY, Zhou KY, Yang G (2012) Phylogenomic analysis resolves the interordinal relationships and rapid diversification of the Laurasiatherian mammals. Syst Biol 61:150–164
Zhu A, Guo W, Gupta S, Fan W, Mower JP (2016) Evolutionary dynamics of the plastid inverted repeat: the effects of expansion, contraction, and loss on substitution rates. New Phytol 209:1747–1756
Acknowledgements
We would like to thank the Brisbane Botanic Garden and Kunming Botanic Garden for permission to sample fresh leaves, and Prof. Michelle van der Bank from University of Johannesburg for providing silica-gel-dried materials. This study was conducted in the Key Laboratory of the Southwest China Germplasm Bank of Wild Species, Kunming Institute of Botany, Chinese Academy of Sciences.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
This study was funded by grants from the Ministry of Science and Technology of China, Basic Research Project (No. 2013FY112600), and the Talent Project of Yunnan Province (No. 2011CI042).
Conflict of interest
The authors declare that they have no conflict of interest.
Data archiving statement
The newly sequenced plastomes have been submitted to GenBank with accession numbers KX852435–KX852445 (see Online Resource 1). The data matrix for phylogenetic reconstruction was deposited at the TreeBASE repository (http://purl.org/phylo/treebase/phylows/study/TB2:S20385).
Additional information
Communicated by Y. Tsumura
Rights and permissions
About this article
Cite this article
Wang, YH., Qu, XJ., Chen, SY. et al. Plastomes of Mimosoideae: structural and size variation, sequence divergence, and phylogenetic implication. Tree Genetics & Genomes 13, 41 (2017). https://doi.org/10.1007/s11295-017-1124-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-017-1124-1