Abstract
Seven hundred fifty-two to one thousand ninety-seven base pairs of the trnL intron and trnL–trnF intergenic spacer of the chloroplast DNA of 55 Juncaceae taxa (Juncus, Luzula, Rostkovia, and Oxychloë) was sequenced. Seventeen structural mutations (13 indels marked A to M, 3 parts of the trnF pseudogene, and insertion “o” within a pseudogene) within the chloroplast trnL–trnF region were examined as possible indicators for phylogenetic relationships in Juncaceae. Juncus trifidus (section Steirochloa) was clearly separated from the other taxa by two large (>80 bp) indels. The “Southern Hemisphere clade” was strongly supported by a unique insertion (334 bp) in the trnL intron. The monophyly of Luzula was supported by three small (<10 bp) indels in the trnL-F spacer. They were found in all 22 examined members that represent the taxonomic and geographical diversity of the genus Luzula. A tandemly duplicated tRNA pseudogene was found in the Juncus subgenus Juncus species and is supported by four small unique indels too. The acceptor stem and D-domain-encoding regions are separated by a unique 8-bp insertion. The T-domain and acceptor stem-encoding regions were not found in the pseudogene repeats. Only the Juncus sections Ozophyllum and Iridifolii contain the 5′ acceptor stem, D-domain, and anticodon domain of the tRNAF encoding DNA. The structural mutations in the trnL intron and the trnL–trnF intergenic spacer are useful for phylogenetic reconstruction in the Juncaceae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chloroplast DNA regions are widely used for phylogenetic studies at all taxonomic levels. The trnL–trnF intergenic spacer and trnL intron have been used from intra- and interspecific levels to the subfamilial and tribal levels (reviewed by Mes et al. 2000). Generally, substitutions are more often used in phylogenetic studies than structural mutations (indels). However, small indels were in some cases informative for the reconstruction of phylogenetic trees (Vijverberg et al. 1999) because the resulting gaps from the noncoding plastome regions in the aligned matrix were found to be phylogenetically relevant (Kelchner 2000). On the other hand, Vijverberg and Bachmann (1999) and Mes et al. (2000) show that some indels in noncoding regions cannot be reliably used for phylogenetic comparisons.
Most structural mutations in cpDNA are small indels, from 1 to 10 bp (reviewed by Vijverberg and Bachmann 1999). Larger indels are centred in noncoding regions only. Small indels associated with hairpins may be highly susceptible to reversal or parallelism within individual groups (Kelchner 2000). Applequist (2002) analyzed deletions in the trnT–trnL region (350 and 250 bp) and found them phylogenetically informative. He showed that they define clades at the subfamily level. Phylogenetically significant 70-bp deletions at the species level were found by Mes et al. (1994) in the trnL–trnF spacer. Several studies focused on tandemly repeated pseudogenes such as in the case of rbcL (Nickrent et al. 1998). Tandemly duplicated, triplicated, or quadruplicated tRNA pseudogenes are most frequent. The tRNA copies adjacent to the trnF (GAA) gene were found in Ozyza sativa (Hiratsuka et al. 1989), Triticum aestivum (Quigley and Weil 1985), Microseris (Vijverberg 1999), Taraxacum (Wittzell 1999), and gymnosperms (Tsudzuki et al. 1994; Hipkins et al. 1995).
Other promising objects of phylogenetic studies are secondary structures of various RNAs. Stem–loop structures have been widely discussed in ribosomal DNA (Soltis and Soltis 1998). Of special interest are ITS and 18 rDNA regions (e.g., Baldwin 1995; Soltis and Soltis 1998). Intergenic spacers as well as introns in the chloroplast DNA contain several stem–loop encoding structures (Kelchner 2000). Several potential stem–loops in the trnL–trnF region and nine such structures in the trnL intron have been described by Gielly and Taberlet (1994). Loops of the stem–loop secondary structures in the noncoding DNA regions are often associated with mutational hot spots. These mutations include nucleotide substitutions and indels (Kelchner 2000). Mes et al. (2000) distinguish phylogenetically misleading indels from true synapomorphies by folding energy calculations.
The Juncaceae is a cosmopolitan family, comprising seven genera and 442 species (Kirschner et al. 2002a, b, c). Only two of the genera, Juncus L. and Luzula DC., are almost truly cosmopolitan. Of the remaining five genera, three are restricted to South America (Distichia Nees & Meyen, Oxychoë Philippi, and Patosia Buchenau) and two are found in both South America and New Zealand (Marsippospermum Desv. and Rostkovia Desv.). In this study we first deliberate on the usefulness of these structural mutations within the trnL-F region. We then evaluated the length variations and secondary structures of the trnL intron and trnL–trnF intergenic spacer in the Juncaceae family. Therefore we used the noncoding regions for determining subgeneric and sectional relationships between Luzula and Juncus.
This is the first study of the noncoding trnL–trnF regions in the Juncaceae family and it is part of the Juncaceae phylogeny project (phylogenetic analysis of two cosmopolitan plant genera, Luzula and Juncus, Juncaceae; for details see http://www.ibot.cas.cz/juncaceae/welcome.htm). Other regions of cpDNA in the Juncaceae were examined by Drábková et al. (2003) and earlier by Plunkett et al. (1995) and Muasya et al. (2000).
Materials and Methods
Total DNA of 55 taxa (Table 1) was extracted either from herbarium specimens or from fresh leaves following the procedure of Doyle and Doyle (1987) as modified by Drábková et al. (2002). DNA was extracted from approximately 0.1 g of dried or 1 g of fresh samples. The trnL–trnF region and the trnL intron were amplified from total DNA by PCR using the primers c, d, e, and f described by Taberlet et al. (1991). PCR products were directly sequenced on a CEQ2000XL automated sequencer (Beckmann). DNA sequences were assembled and aligned using the GeneSkipper (EMBL, Heidelberg) and BioEdit (Hall 1999) programs. Secondary structures were determined using the Mulfold program (Zuker 1989). Default settings were used (folding temperature, 37°C; [Na+] = 1.0 M; [Mg2+] = 0) for folding structures.
Analysis of secondary structures together with the sequence variability was used to identify DNA regions undergoing high mutational rates and to distinguish them from the more evolutionarily conserved regions. Phylogenetic analyses were performed with only informative characters included under the parsimony criterion as implemented in PAUP* version 4.0b10 (Swofford 2002). Due to the size of the matrix only heuristic searches were carried out. The TBR swapping and steepest descent analyses were used. One analysis was performed only with substitutions and indels as missing data; the second analysis included indels coding as binary data. Phylogenetic trees were constructed using parsimony with Carex spp. as outgroup based on previous data that confirmed the Cyperaceae as sister group of the Juncaceae (Simpson 1995; Munro and Linder 1998).
The nucleotide sequences dealt with in this communication are deposited in GenBank (accession Nos. AY437928–AY437981). A number of samples are documented by specimens deposited in the herbarium of the Institute of Botany, Academy of Sciences, Czech Republic (PRA).
Results
In total, we sequenced regions from 332 to 654 bp in length of the trnL intron and from 147–540 bp in length of the trnL–trnF intergenic spacer of the plastome DNA of 55 taxa representing most of the subgenera and sections of Luzula and Juncus and also Rostkovia magellanica and two species of Oxychloë, namely, O. andina and O. bisexualis (see Table 1). Individual taxa were chosen to reflect the morphological and geographical diversity of both genera. For details on nomenclature and taxonomy of the taxa studied, see Kirschner et al. (2002a–c).
After alignment, we obtained 1553 characters for each taxon. In total, we worked with 987 informative positions (for details see Methods), and of these 766 substitutions were parsimony informative. Seventeen structural mutations (including pseudogene duplications) within the chloroplast trnL–trnF region were examined as possible indicators for phylogenetic relationships in Juncaceae.
Structural Mutations in the trnL Intron
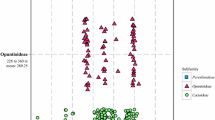
The trnL intron length range was detected in both Luzula and Juncus (Fig. 1B, Table l). Most of these structural mutations are Juncus specific (indels B–G; Fig. 3) and none of them is shared by two genera. One insertion type (A; Fig. 3) is identical in several Luzula species and Rostkovia magellanica. All Juncus species are characterized by a 7-bp deletion. Carex species differ by only one substitution (AAAGATA). Insertion type A is typical for Luzula section Luzula, Alpinae, Thyrsanochlamydeae, Nodulosae, Anthelaea (Fig. 3). J. trifidus is unique among the other species by a 322-bp deletion (C; Fig. 3). Another long deletion (D; 334 bp) was found in the “Southern Hemisphere clade,” which consists of Juncus species of the section Graminifolii (J. lomatophyllus, J. capensis), Rostkovia magellanica, and two Oxychloë (O. andina and O. bisexualis). Insertion E (22 bp) is typical of Juncus sections Ozophyllum, Iridifolii, and one representative of Graminifolii. For both subgenus Juncus and subgenus Agathryon the 4-bp insertion is typical. On the other hand Juncus subgenus Agathryon has an additional 5-bp deletion close to the 3′ end of the intron.
Structural Mutations in the trnL–trnF Intergenic Spacer
The varying length in the trnL–trnF intergenic spacer of Luzula and Juncus species is described in Fig. 1A and Table 1. Six main structural mutations (H–M) were investigated in the trnL–trnF region, and in addition, three trnF pseudogenes were found (Figs. 2 and 4). Compared to the trnL intron (Fig. 3) the trnL–trnF spacer contains shorter indels (from 3 to 85 bp). All of them are genus specific: deletions H, K, and M are typical of the genus Luzula; deletions I and L and insertion J are Juncus specific (Fig. 4). The first 5-bp deletion (H) was found in all Luzula species studied (Fig. 4). The 9-bp deletion (I) is typical of subgenus Juncus only. Insertion J is typical of J. trifidus of section Steirochloa. This insertion consists of an 84-bp repetition of the (TATATAAT)6 motif in combination with an “A” duplication. The 5-bp deletion K was found in several Luzula species from the sections Anthelea, Luzula, Alpinae, Thyrsanochlamydeae, and Nodulosae. It was not found in subgenus Marlenia, Pterodes and section Diprophyllateae. The 8-bp deletion (L) was typical of the entire Iridifolii and Ozophyllum sections and one taxon of the Graminifolii. The 10-bp indel M was found in all Luzula representatives (22 species). Indels occurred with prevalence in A/T-rich stem regions of hairpins (Figs. 4 and 6). It was used for understanding position homology in an aligment (especially in difficult taxa, e.g., J. trifidus).
Structural mutations in the trnL–trnF intergenic spacer (n, o, p, q). Arrows show main mutational hot spots in the trnL–trnF region. n: 1427–1436 bp Duplication of acceptor stem. o: 1437–1444 bp Insertion. p: 1445–1453 bp Duplication of D-domain. q: 1454–1479 bp Duplication of anticodon domain. Base pair positions are counted from primer “c” site 1–1553 in alignment (Taberlet 1994).
Characteristics of structural mutation in the trnL intron. Numbers above boxes show the length of regions (bp). Structural mutations: A, 216–222 bp Insertion;B, 277–362 bp Deletion; C, 345–667 bp Deletion; D, 404–738 bp Insertion;E, 429–451 bp Insertion; F, 574–577 bp Insertion; G, 734–738 bp Deletion (for taxon details see Table 1).
Characteristics of structural mutation in the trnL–trnF intergenic spacer. Numbers above boxes indicate the length of regions (bp). TrnF pseudogene details are given in Fig. 2. Structural mutations: H, 937–940 bp Deletion; I, 1063–1070 bp Deletion; L, 1241–1225 bp Insertion; K, 1146–1150 bp Deletion; L, 1189–1296 bp Deletion; M, 1207–1216 bp Deletion (for taxon details see Table 1).
Table 2A shows the trnF gene of one representative of the Juncaceae (J. articulatus) demonstrating that there is a duplication of “G” and two transitions (A→G, T→C) in Juncaceae compared to the homologous region of Nicotiana tabacum. Our study shows (see Table 1B) that trnF pseudogenes of differing lengths in the Juncus species occur only in subgenus Juncus. This diversity was not observed in subgenera Agathryon and Luzula. No variation was found in the 5′ sequence of the acceptor stem of the trnF pseudogene in Juncus and Luzula species.
There is an insertion of 8 bp between the acceptor stem and the D-domain in the trnF pseudogene and in all examined species of subgenus Juncus. The T-domain and the acceptor stem at the 3′end of the gene were not found in the pseudogenes. The trnF pseudogene varies in length in different species (Table 2). Sections Ozophyllum and Iridifolii contain at least part of the D-domain and anticodon domain of trnF pseudogene structures (ψ1). Juncus ensifolius (section Iridifolii belongs to a separate group (ψ2) which lacks most of the anticodon domain. Section Stygiopsis (J. biglumis, J. triglumis, J. castaneus, and J. stygius) contains the shortest pseudogene (ψ3), which lacks the anticodon and most of the D-domain. Two mutations were found among the Juncus trnF pseudogenes: A→C, C→T substitutions are found in the D-domain and anticodon domain, respectively. The acceptor stem, D-domain, and anticodon domain contain a few substitutions in comparison with the trnF gene (see Table 2).
Phylogenetic Analysis
We constructed a phylogenetic tree (simplified in Fig. 5) based on substitutions and indels. We included in our study geographically diverse species and concluded that the large indels are autapomorphies for some species (e.g., 322-bp deletion for J. trifidus) and synapomorphies for others (e.g., 334-bp insertion for the Southern Hemisphere clade) and that small structural rearrangements seem to be more often homoplasious (e.g., 7-bp insertion for Luzula and Rostkovia).
Simplified phylogenetic tree of Juncaceae based on substitutions. Structural mutations are marked on the tree. Light gray boxes designate positions in the trnL–trnF region; dark gray in the trnL region; black circles, homoplasious deletion; ψ1–3 parts of the pseudogene (see Table 2). Tree characteristics: CI = 0.61, RI = 0.87, RC = 0.54, HI = 0.39. Numbers above branches indicate bootstrap values.
The monophyly of Luzula suggests that deletions in the trnL–trnF spacer occurred in an ancestor of all Luzula species (three <10-bp indels). This is satisfactorily confirmed by morphology (Fig. 7). On the other hand, Juncus is nonmonophyletic as shown by analysis of both the trnL–trnF and the rbcL regions (Drábková et al. 2003). The subgenus Juncus (and Agathryon) clades are amply supported by specific indels. Additionally, it seems that the tRNA pseudogenes in Juncaceae evolved more recently since they occur in the youngest part of the tree (Fig. 5) in subgenus Juncus. Generally genus Juncus (excluding Juncus trifidus) forms a recently evolved clade. When the tree is based on both substitutions and indels in the whole trnL–trnF region, the results are better (CI = 0.61, RI = 0.87, RC = 0.54, HI = 0.39) than when only substitutions are used (CI = 0.55, RI = 0.82, RC = 0.45, HI = 0,45). The inclusion of indels clearly improved statistical support of the phylogenetic tree (nevertheless the tree topology of the main clades remain unchanged).
J. trifidus forms a sister group of the whole genus Luzula and, according to the trnL–trnF spacer analysis, also a sister group of the main Juncus clade (for more details see Drábková et al. 2003). J. trifidus is characterized by two main autapomorphic indels, one in the trnL intron (322-bp deletion) and the other in the trnL–trnF spacer (84-bp insertion that forms a hairpin; see Fig. 6). These two structural events are unique among the species of Juncus and Luzula. The Southern Hemisphere clade that was inferred from the rbcL data (Drábková et al. 2003) is supported by autapomorphic substitutions in the trnL–F region. It is also characterized by a unique 334-bp deletion. In the small South American genus Oxychloë (represented by O. andina and O. bisexualis), the deletion is even 144 bp longer, the longest indel within this region in the whole Juncaceae family. This separates it from other species of this clade (not shown).
Discussion
In this study we wanted to show the usefulness of the structural mutations and length variations within the trnL–F region. We evaluated variation of the trnL intron and trnL–trnF intergenic spacer in the Juncaceae family with respect to determination of subgeneric and sectional relationships within Luzula and Juncus. Results were compared with those obtained from the analysis of morphological and other taxonomic data and the data from rbcL (Drábková et al. 2003).
Both cosmopolitan genera, Juncus and Luzula, are morphologically well defined (shared characters for both, but not for other members [see below] of the Juncaceae: flowers in multiflowered inflorescence, anthers not mucronate or if anthers minutely mucronat, then auricules lacerate; Juncus— capsule many-seeded, leaves not hairy, flowers with or without bracteoles, leaf sheath open; Luzula—capsule three-seeded, leaves ciliate, flowers with basal bracteoles, leaf sheath closed). The genus Juncus is traditionally subdivided in to two subgenera, Juncus and Agathryon (Fig. 7), based on shared inflorescence characters, which were taken to be homologous (racemose or cymose inflorescence and presence or absence of a pair of floral bracteoles). Our study of the trnF in Juncaceae has been especially informative in that it shows a range of structural mutations in a single lineage of Juncus, namely, in subgroups Juncus sections Ozophyllum and Iridifolii. This is interesting because these two sections contain many species that share morphological features that are considered to be homoplasious (leaves pluritubulose, septate, vascular bundles usually in subepidermal position, lower bract not appearing as a stem prolongation). Molecular data confirmed monophyly of Luzula, but the traditionally recognized subgenus Pterodes was found dispersed throughout the Luzula clade. On the other hand, subgenus Luzula formed one well-defined clade, and the monotypic subgenus Marlenia occupied (in some cases) a basal position in the Luzula branch. The monophyly of Juncus is violated by the basal position of Juncus trifidus (Fig. 5). J. trifidus L. and the closely related J. monanthos Jacq. are generally considered to form a taxonomically distinct group. Rouy (1912) and Novikov (1990) placed them in a separate section or subsection Trifidi, respectively (Fig. 7). Placement of the Southern Hemisphere clade inside the Juncaceae as a sister group to Juncus with an ancestral position of Luzula was the second main difference from expectations based on morphological data. The accepted taxonomic circumscription of these five small, mainly South American genera is based on solitary flowers, mucronate anthers, and never lacerate auricles (for differences from others see above). The unique position within the Southern Hemisphere clade occupied two species from the section Graminifolii, J. capensis and J. lomatophyllus, distributed in South Africa. Our finding of differences between our data and widely accepted taxonomic classification in general topology of the Juncaceae (compare Figs. 6 and 7) was confirmed from both data sets (trnL–F, rbcL [Drábková et al. 2003]).
Circle graph of secondary structures of the trnL–trnF intergenic spacer of Juncus ensifolius (A), J. trifidus (B), and Luzula elegans (C), obtained from Mfold (Zuker; http://www.bioinfo.rpi.edu/applications/mfold/old/dna). Domains commonly found in these structures in all examined species (55) are in boxes in structure A. Arrows show main indel regions. Circle shows 84-bp insertion in J. trifidus.
Relationships of (A) Juncaceae (Thurniaceae, including Prioniaceae) and (B) Juncus and Luzula clades based on morphological data (according to Kirschner et al. 1999; Kirschner 20002a; Novikov 1999; separate position of section Trifidii).
Some authors have shown the potential usefulness of structural mutations for reconstruction of phylogenetic relationships in plants (e.g., Soltis and Soltis 1998). Others maintain that structural mutations cannot be used as reliable indicators of phylogenetic distance (Vijverberg and Bachmann 1999). The homoplasy of indels was investigated in some plant species. It was shown by Mes et al. (2000) that only homoplasious indels are found in Taraxacum if presented in stabilized secondary structures. However, only a small number of Taraxacum individuals were used in this study, while for a meaningful evaluation, more extensive sampling of the study group is required (Vijverberg and Bachmann 2000). Our study of Juncaceae shows that some indels in noncoding chloroplast DNA are phylogenetic indicators, because the revealed clades represent morphological groups (for summary see Fig. 5).
In this report we have dealt with indels, ranging from 3 to 334 bp. The short indels in the trnL–trnF region may have evolved by replication slippage or unequal crossing-over (e.g., Kelchner and Wendel 1996). The longer structural mutations may result from duplications through slipped-strand mispairing. Substitutions in the trnL–trnF spacer are the results of point mutations.
Forty-two stem–loop structures were found throughout the trnL-F region. In the trnL–trnF spacer, 11 or 14 hairpins were found, of which 3 are associated with the main indel (Fig. 6). As reviewed by Kelchner (2000) some of the large loops are associated with the length-variable regions as in the case of the Juncus trifidus 5′-...AAAATATATAAT...-3′region (insertion of 84 bp in the trnL–trnF spacer—J). These stem–loop secondary structures not only are specific for the GC-rich DNA regions (Kelchner 2000) but also are found in the AT-rich regions in the Juncaceae (Fig. 6). The stem length is often constant and the length variability appears in the loop. In Luzula section Luzula the stem–loop structures are often conserved in the trnF region. More variability was found in Juncus. These facts highly correspond with more conservative regions found within the genus Luzula (trnL–F and rbcL) and the more variable genus Juncus, which is nonmonophyletic.
In the Juncaceae, the trnL–trnF spacer region was found to be more variable than the trnL intron. This is in agreement with the hypothesis of Bakker et al. (2000) that nucleotide substitutions in introns accumulate more uniformly than in spacer regions (see also Applequist 2002; Gielly and Taberlet 1994). This has been verified for six distant angiosperm groups (Actea, Digitalis, Drosera, Panicoideae, Pelargonium, and Huperzia).
In conclusion, structural mutations in the trnL intron and the trnL–trnF intergenic spacer are good phylogenetic indicators in the Juncaceae. The tRNA pseudogene is useful for phylogenetic reconstructions at the level of Juncus subgenus Juncus.
References
WL Applequist RS Wallace (2002) ArticleTitleDeletions in the plastid trnL–trnF intergenic spacer define clades within Cactaceae subfamily Cactoideae Pl Syst Evol 231 153–162 Occurrence Handle10.1007/s006060200017 Occurrence Handle1:CAS:528:DC%2BD38Xks1yis7Y%3D
FT, Bakker et al. (2000) ArticleTitlePatterns of nucleotide substitution in angiosperm cpDNA trnL(UAA)–trnF(GAA) regions Mol Biol Evol 17 IssueID8 1146–1155 Occurrence Handle1:CAS:528:DC%2BD3cXlvFChu7c%3D Occurrence Handle10991703
BG Baldwin et al. (1995) ArticleTitleThe ITS region of nuclear ribosomal DNA: A valuable source of evidence on angiosperm phylogeny Ann Mo Bot Gard 82 247–277
JJ Doyle JL Doyle (1987) ArticleTitleA rapid DNA isolation procedure for small quantities of fresh leaf tissue Phytochem Bull 19 11–15
L Dráková J Kirschner Č Vlček (2002) ArticleTitleHistorical herbarium specimens in molecular taxonomy of the Juncaceae: A comparison of DNA extraction and amplification protocols Plant Mol Biol Rep 20 IssueID2 161–175
L Drábková J Kirschner O Seberg G Petersen Cø Vlček (2003) ArticleTitlePhylogeny of the Juncaceae based on rbc L sequences, with special emphasis on Luzula DC. and Juncus L Pl Syst Evol 240 133–147 Occurrence Handle10.1007/s00606-003-0001-6
L Gielly P Taberlet (1994) ArticleTitleThe use of chloroplast DNA to resolve plant phylogenies: Noncoding versus rbcL sequences Mol Biol Evol 11 IssueID5 769–777 Occurrence Handle1:CAS:528:DyaK2cXmtVWjtLk%3D Occurrence Handle7968490
TA Hall (1999) ArticleTitleBioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT Nucl Acids Symp Ser 41 95–98 Occurrence Handle1:CAS:528:DC%2BD3cXhtVyjs7Y%3D
VD, Hipkins et al. (1995) ArticleTitleA mutation hotspot in the chloroplast genome of a conifer (Douglas-fir: Pseudotsuga) is caused by variability in the number of direct repeats from a partially duplicated tRNA gene Curr Genet 27 527–579
J, Hiratsuka et al. (1989) ArticleTitleThe complete sequence of the rice (Oryza sativa) chloroplast genome: Intermolecular recombination between distinct tRNA genes accounts for a major plastid DNA inversion during the evolution of cereals Mol General Genet 217 185–194 Occurrence Handle1:CAS:528:DyaL1MXlt1yjs7k%3D
SA Kelchner (2000) ArticleTitleThe evolution of non-coding chloroplast DNA and its application in plant systematics Ann Miss Bot Gard 87 482–498
SA Kelchner JF Wendel (1996) ArticleTitleHairpins create minute inversions in non-coding regions of chloroplast DNA Curr Genet 30 259–262 Occurrence Handle10.1007/s002940050130 Occurrence Handle1:CAS:528:DyaK28XlvVCgtL8%3D Occurrence Handle8753656
J, Kirschner et al. (1999) ArticleTitleSupraspecific division of the genus Juncus (Juncaceae) Folia Geobot 34 377–390
J, Kirschner et al. (2002a) Juncaceae 1: Rostkovia to Luzula, species plantarum: Flora of the world, part 6 ABRS Canberra 1–237
J, Kirschner et al. (2002b) Juncaceae 2: Juncus subg. Juncus, species plantarum: Flora of the world, part 7 ABRS Canberra 1–237
J, Kirschner et al. (2002c) Juncaceae 3: Juncus subg. Agathryon, species plantarum: Flora of the world, part 8 ABRS Canberra 1–192
THM, Mes et al. (2000) ArticleTitleHairpins involving both inverted and direct repeats are associated with homoplasious indels in non-coding chloroplast DNA of Taraxacum (Lactuceae: Asteraceae) Genome 43 634–641 Occurrence Handle10.1139/gen-43-4-634 Occurrence Handle1:CAS:528:DC%2BD3cXmsVWitbc%3D Occurrence Handle10984175
THM Mes H t′Hart (1994) ArticleTitle Sedum surculosum and S. jaccardianum (Crassulaceae) share a unique 70 bp deletion in the chloroplast DNA trnL (UAA)– trnF(GAA) intergenic spacer Pl Syst Evol 193 213–221 Occurrence Handle1:CAS:528:DyaK2MXkvVSktL8%3D
AM Muasya JJ Bruhl Simpson DA A Culham MW Chase (2000)) Suprageneric phylogeny of Cyperaceae: A combined analysis PJ Rudall PJ Cribb DF Cutler CJ Humpries (Eds) Monocotyledons: Systematics and evolution Royal Botanic Gardens Kew 593–601
SL Munro HP Linder (1998) ArticleTitleThe phylogenetic position of Prionium (Juncaceae) within the order Juncales based on morphological and sequence data Syst Bot 23 IssueID1 43–55
DL, Nickrent et al. (1998) Molecular phylogenetic and evolutionary studies of parasitic plants DE Soltis PS Soltis JJ Doyle (Eds) Molecular systematics of plants II: DNA sequencing Kluwer Academic New York 211–242
VS Novikov (1990) ArticleTitleKonspect sistemy roda Juncus (Juncaceae), [Synopsis of the genus Juncus (Juncaceae)] Bjul Mosk Obšč Ispyt Prir Otd Biol 95 IssueID5 111–125
F Quigley JH Weil (1985) ArticleTitleOrganization and sequence of five tRNA genes and of an unidentified reading frame in the wheat chloroplast genome: Evidence for gene rearrangements during evolution of genomes Curr Genet 9 495–503 Occurrence Handle1:CAS:528:DyaL2MXltlKit78%3D Occurrence Handle3870931
GM Plunkett DE Soltis PS Soltis RE Brooks (1995) ArticleTitlePhylogenetic relationships between Juncaceae and Cyperaceae: Insights from rbcL sequence data Am J Bot 82 IssueID4 520–525
G Rouy (1912) ArticleTitleJoncacées Flore de France 13 222–223
D Simpson (1995) Relationships within Cyperales PJ Rudall PJ Cribb DF Cutler CJ Humpries (Eds) Monocotyledons: Systematics and Evolution Royal Botanic Gardens Kew 497–509
PS Soltis DE Soltis (1998) Molecular evolution of 18S rDNA in angiosperms: Implications for character weighing in phylogenetic analysis DE Soltis PS Soltis JJ Doyle (Eds) Molecular systematics of plants II: DNA sequencing Chapman and Hall New York 188–210
DL Swofford (2002) PAUP*. Phylogenetic analysis using parsimony (* and other methods) version 4.0b10
P Taberlet L Gielly J Bouvet (1991) ArticleTitleUniversal primers for amplification of three non-coding regions of chloroplast DNA Plant Mol Biol 17 1105–1109 Occurrence Handle1:CAS:528:DyaK38Xhslel Occurrence Handle1932684
J, Tsudzuki et al. (1994) ArticleTitleA new gene encoding tRNA-Pro(GGG) is present in chloroplast genome of black pine: A compilation of 32 tRNA genes from black pine chloroplast Curr Genet 26 153–158 Occurrence Handle1:CAS:528:DyaK2cXmsVOksrY%3D Occurrence Handle8001170
K Vijverberg K Bachmann (1999) ArticleTitleMolecular evolution of a tandemly repeated trnF(GAA) gene in the chloroplast genome of Microseris (Asteraceae) and the use of structural mutations in phylogenetic analyses Mol Biol Evol 16 IssueID10 1329–1340 Occurrence Handle1:CAS:528:DyaK1MXms1Ojt7o%3D Occurrence Handle10563014
K Vijverberg THM Mes K Bachmann (1999) ArticleTitleChloroplast DNA evidence for the evolution of Microseris (Asteraceae) in Australia and New Zealand after long-distance dispersal from western North America Am J Bot 86 IssueID10 1448–1463 Occurrence Handle1:CAS:528:DyaK1MXntlaku7Y%3D Occurrence Handle10523285
H Wittzell (1999) ArticleTitleChloroplast DNA variation and reticulate evolution in sexual and apomictic sections of dandelions Mol Ecol 8 2023–2035 Occurrence Handle10.1046/j.1365-294x.1999.00807.x Occurrence Handle1:CAS:528:DC%2BD3cXpvF2lsw%3D%3D Occurrence Handle10632854
M Zuker (1989) ArticleTitleOn finding all suboptimal foldings of an RNA molecule Science 244 48–52 Occurrence Handle1:CAS:528:DyaL1MXkt1SnsbY%3D Occurrence Handle2468181
Acknowledgments
We would like to thank the curators of herbariums AAU, BM, C, and K, where many Juncaceae DNA samples were collected, for their great hospitality. J.K. and L.D. thank T.H.M. Mes and J. Fehrer for fruitful comments on the manuscript. We are grateful to John Novotney for improving the English. The study was done at the DNA Laboratory, Institute of Botany, ASCR, and Laboratory of the Centre for Integrated Genomics and Institute of Molecular Genetics, ASCR, and was supported in part by Grants GACR 206/02/035, GAUK 109/2002/B/BIO, and AVOZ 6005908.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Drábková, L., Kirschner, J., Vlček, Č. et al. TrnL–trnF Intergenic Spacer and trnL Intron Define Major Clades Within Luzula and Juncus (Juncaceae): Importance of Structural Mutations. J Mol Evol 59, 1–10 (2004). https://doi.org/10.1007/s00239-004-2598-7
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00239-004-2598-7