Abstract
The plastid genome of Trifolium subterraneum is 144,763 bp, about 20 kb longer than those of closely related legumes, which also lost one copy of the large inverted repeat (IR). The genome has undergone extensive genomic reconfiguration, including the loss of six genes (accD, infA, rpl22, rps16, rps18, and ycf1) and two introns (clpP and rps12) and numerous gene order changes, attributable to 14–18 inversions. All endpoints of rearranged gene clusters are flanked by repeated sequences, tRNAs, or pseudogenes. One unusual feature of the Trifolium subterraneum genome is the large number of dispersed repeats, which comprise 19.5% (ca. 28 kb) of the genome (versus about 4% for other angiosperms) and account for part of the increase in genome size. Nine genes (psbT, rbcL, clpP, rps3, rpl23, atpB, psbN, trnI-cau, and ycf3) have also been duplicated either partially or completely. rpl23 is the most highly duplicated gene, with portions of this gene duplicated six times. Comparisons of the Trifolium plastid genome with the Plant Repeat Database and searches for flanking inverted repeats suggest that the high incidence of dispersed repeats and rearrangements is not likely the result of transposition. Trifolium has 19.5 kb of unique DNA distributed among 160 fragments ranging in size from 30 to 494 bp, greatly surpassing the other five sequenced legume plastid genomes in novel DNA content. At least some of this unique DNA may represent horizontal transfer from bacterial genomes. These unusual features provide direction for the development of more complex models of plastid genome evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During the past 5 years there has been a rapid increase in the availability of complete plastid genome sequences of angiosperms, due largely to the development of faster and cheaper methods for whole-genome sequencing (Jansen et al. 2005; Moore et al. 2006). The 90 angiosperm plastid genome sequences already available in GenBank (http://www.ncbi.nlm.nih.gov/genomes/ORGANELLES/plastids_tax.html) have provided valuable insight into plastid genome evolution (Saski et al. 2005; Chumley et al. 2006; Lee et al. 2007; Raubeson et al. 2007; Haberle et al. 2008; Wang et al. 2008) as well as phylogenetic relationships (Goremykin et al. 2003; Leebens-Mack et al. 2005; Chang et al. 2006; Hansen et al. 2007; Jansen et al. 2007; Moore et al. 2007). These genome sequences support the view that the plastid genome is highly conserved in both gene order and content, with the majority having two copies of a large (usually 25-kb) inverted repeat (IR) separating the small and large single-copy regions (SSC and LSC, respectively). The ancestral angiosperm genome (Raubeson et al. 2007) has 115 different genes, 18 of which contain introns. This ancestral organization is conserved across angiosperms (Jansen et al. 2007) ranging from the earliest-diverging lineages, Amborella (Goremykin et al. 2003) and Nymphaeales (Goremykin et al. 2004; Raubeson et al. 2007), to more derived lineages, including the asterid Nicotiana (Shinozaki et al. 1986) and the monocot Acorus (Goremykin et al. 2005).
Recent studies have revealed exceptions to the prevailing view that angiosperm plastid genome organization is highly conserved. The most highly rearranged genomes from photosynthetic angiosperms published to date are Pelargonium (Geraniaceae [Chumley et al. 2006]) and Trachelium (Campanulaceae [Haberle et al. 2008]). The extent of rearrangement in these two lineages is different, but the underlying mechanisms, inversions and expansion of the IR, are generally similar. Another common feature of these extensively rearranged plastid genomes is the prevalence of repeated sequences, many of which are associated with rearrangement endpoints. Previous studies have provided evidence that recombination between inverted repeats or tRNA genes has caused inversions (Bowman and Dyer 1986; Hiratsuka et al. 1989; Hupfer et al. 2000; Pombert et al. 2005, 2006). Furthermore, although transposition via transposable elements (TEs) has been suggested as a possible mechanism for gene order changes for a number of highly rearranged angiosperm plastid genomes (Milligan et al. 1989; Cosner et al. 1997), no direct evidence for this process has been uncovered. In fact, the inactive “Wendy” element in Chlamydomonas (Fan et al. 1995) remains the only known example of a TE in any plastid genome.
Early gene mapping studies (Milligan et al. 1989) suggested that the plastid genome of Trifolium subterraneum is unusual among land plants in several respects. Milligan et al. (1989) reported three unusual features of this genome. First, the gene order is highly rearranged, with 10 clusters of genes rearranged in both order and orientation relative to another legume, Medicago. Eight large inversions were proposed to explain the reorganization of these gene clusters. Second, a family of dispersed repeats unique to Trifolium and two closely related species was identified and proposed to originate from TEs. Third, unique repeated elements and unique single-copy sequences may represent rare instances of transfer of sequences into a plastid genome.
In this study, we present the complete plastid genome sequence of Trifolium subterraneum and provide detailed comparisons of repeats and unique DNA with other sequenced plastid genomes, including those of five other legumes. These comparisons confirm size estimates and overall genome architecture proposed previously (Milligan et al. 1989) but also show that the Trifolium genome is more highly rearranged than previously suggested. The genome includes 19.5% (ca. 28 kb) of repetitive DNA, five times more than any other angiosperm plastid genome sequenced to date, and these repeats are associated with rearrangements. However, we find no evidence of TEs, suggesting that inversion and gene duplication/loss are the primary mechanisms for gene order changes. The Trifolium plastid genome also includes 19.5 kb of unique DNA, some of which may represent lateral DNA transfer from other organisms.
Materials and Methods
Plastid DNA Isolation
Trifolium subterraneum seeds were obtained from the U.S. Department of Agriculture Plant Introduction Service. Plastid DNA was extracted from freshly harvested leaves using a modification of the procedure outlined by Palmer (1986), involving the use of a high-salt extraction buffer recommended by Bookjans et al. (1984) and elimination of the sucrose-gradient step.
Genome Assembly and Annotation
Purified plastid DNA was sheared into ≈3-kb fragments using a Hydroshear device (Gene Machines, San Carlos, CA, USA). These fragments were then end-repaired, gel-purified, ligated into pUC18 plasmid vectors to create a DNA library, introduced into competent E. coli by electroporation, and plated onto nutrient media with antibiotic selection. The resulting colonies were randomly selected and processed robotically for end sequencing using Big Dye (Applied Biosystems, Foster City, CA, USA) chemistry and an ABI 3730 XL sequencer at the DOE Joint Genome Institute (Walnut Creek, CA, USA). Sequences from randomly chosen clones were processed using PHRED and assembled based on overlapping sequences into a draft genome using PHRAP (Ewing and Green 1998). Quality of the sequence and assembly was verified using CONSED (Gordon et al. 1998). Most regions of the genome had 6- to 12-fold coverage, and areas with gaps or low depth of coverage were PCR amplified and sequenced at The University of Texas at Austin. Additional sequences were added until a completely contiguous consensus was created, representing the entire plastid genome, with a minimum of 2 × coverage and a consensus quality score of ≥Q40. The genome was annotated using DOGMA (Dual Organellar GenoMe Annotator [Wyman et al. 2004]; http://dogma.ccbb.utexas.edu/).
Identification of Repeats and Unique DNA
BLAST (default parameters) comparisons of 15 representative angiosperm plastid genomes were performed against themselves to identify repeated sequences in each genome. Repeats ≥30 bp were plotted. For the plastid genomes with two copies of the inverted repeat, one copy was removed. The genomes examined included the six legumes Cicer arietinum (NC_011163), Glycine max (NC_007942), Lotus japonicus (NC_002694), Medicago truncatula (NC_003119), Phaseolus vulgaris (NC_009259), and Trifolium subterraneum (EU849487) and nine other eudicots, Arabidopsis thaliana (NC_000932), Cucumis sativus (NC_007144), Eucalyptus globules (NC_008115), Morus indica (NC_008359), Nicotiana tabacum (NC_001879), Pelargonium x hortorum (NC_008454), Populus alba (NC_008235), Trachelium caeruleum (NC_010442), and Vitis vinifera (NC_007957).
BLAST searches of Trifolium were also performed against all plastid genomes on GenBank (using release number 164) to identify unique sequences within the six legume plastid genomes. Unique DNA is defined as DNA 30 bp or longer not shared by legumes or any of the other available 131 plastid genomes. BLASTN and BLASTX analyses of the Trifolium unique DNA were performed against GenBank.
Identification of Transposable Elements
To identify putative TEs, the Plant Repeat Database was downloaded from http://www.tigr.org/tdb/e2k1/plant.repeats/ and BLAST searches of Trifolium against this database were performed. BLAST searches were performed with an e value of 10, and repetitive or low-complexity sequences were not filtered. In addition, the program LTR STRUC (McCarthy and McDonald 2003) was used to search for long terminal repeat (LTR) retrotransposons in all 131 plastid genomes available at GenBank to locate putative TEs.
Estimation of Number of Inversions
The genome comparison tool GRIMM (Tesler 2002) was used to estimate the minimum number of inversions required to derive the Trifolium gene order from that of Medicago. The GRIMM algorithm is limited in that it cannot accommodate gene duplications or other differences in gene content between the input genomes. Gene content, order, and orientation were determined for each genome using DOGMA, and an ordered matrix of each gene and its relative strand orientation was created. To equalize the content of the genomes, two genes (accD and rps18) absent from Trifolium were removed from Medicago; likewise, it was necessary to exclude the many pseudogenes (three fragments of psbN, six of rpl23) present in Trifolium from the analysis. Novel DNA, representing ~20% of Trifolium, was also excluded from the analysis. Therefore, missing and duplicated genes in Trifolium were excluded from the input file for Medicago, resulting in a comparison of 107 shared genes.
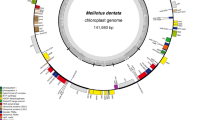
Both pairwise BLAST and the Mauve genome alignment algorithm (Darling et al. 2004) revealed many clusters of genes, or local collinear blocks, that are in the same order in both Medicago and Trifolium. Each cluster included two or more genes, for a total of 16 clusters or collinear, unrearranged blocks of genes (Fig. 1). In addition, three genes (trnL-caa, trnI-cau, and clpP) occur singly (i.e., alone and not part of a cluster), separated from other genes by novel DNA. These 16 gene clusters were also coded for analysis with GRIMM.
Gene map of the Trifolium subterraneum plastid genome. Genes on the outside of the map are transcribed in the clockwise direction and genes on the inside of the map are transcribed in the counterclockwise direction. Boxed areas indicate the 16 clusters of genes discussed in the text and also presented in Figs. 2 and 3
Results
Organization of the Trifolium Plastid Genome
The complete plastid genome of Trifolium (GenBank accession number EU849847) has lost one copy of the IR, but this loss has not greatly reduced its overall size relative to those genomes containing two copies of the IR because the novel DNA in Trifolium is cumulatively similar in length to the typical angiosperm IR. Trifolium, like two relatives whose plastid genomes are being compared here, Cicer and Medicago, has only one copy of the IR. These three genomes are members of a large clade (IRLC [Wojciechowski et al. 2004]) of papilionoid legumes that is marked by the loss of one copy of the IR. The Trifolium subterraneum genome is 144,763 bp (Fig. 1), ~20 kb longer than other legumes with which it shares only a single copy of the IR (Table 1). The overall GC content of the six sequenced legume plastid genomes is similar, ranging from 34% to 36% (Table 1). The Trifolium plastid genome contains 111 different genes. Six genes (accD, infA, rpl22, rps16, rps18, and ycf1) are missing, and two genes (clpP, rps12) have lost an intron relative to the ancestral angiosperm plastid genome (Raubeson et al. 2007). Two of these genes, infA and rpl22, are missing from all legumes (Doyle et al. 1995; Millen et al. 2001), and a third, rps16, has been lost from many papilionoid legumes (Doyle et al. 1995).
The Trifolium genome is highly rearranged in both gene order and orientation. Comparison of the Trifolium plastid genome with Medicago reveals that it is composed of 16 clusters of genes (Figs. 1 and 2); genes within each cluster are in the same order as in Medicago, whereas the clusters themselves have been extensively shuffled across the genome (Fig. 2). Considering only these 16 clusters (Fig. 1), GRIMM proposes 14 inversions to derive the Trifolium gene order from that of its close relative Medicago, some of which break up clusters of genes that are adjacent to each other in the remaining copy of the IR (Fig. 2). Four additional inversions (18 total) are required when including the three genes appearing alone, apart from the blocks of genes. The latter scenario, with 18 inversions separating Trifolium from the gene order of Medicago, is supported by GRIMM analysis of the position and orientation of 107 genes common to both genomes.
Comparison of the gene order and orientation of plastid genes between Medicago truncatula (top) and Trifolium subterraneum (bottom). The arrows refer to the clusters of genes in Fig. 1, with the reversed arrows indicating an inverted orientation. The scale bar at the bottom shows the coordinates of the clusters of genes on the genome. Bars on the Medicago and Trifolium maps indicate clusters of genes that are normally located in the inverted repeat (IR) of legumes (Saski et al. 2005)
It is not surprising that using both collinear blocks and individual genes in the GRIMM analyses converged on the same number of inversions, because they both exclude duplicated genes. There is still no comprehensive method for comparing genomes with unequal gene contents. Modeling the evolution of these rearranged genomes with confidence will require an improved algorithm capable of incorporating gene/intron losses, gene duplications, and, for the majority of plastid genomes, variation in the IR boundaries. Current models of plastid genome evolution based solely on inversions between a reduced, common set of genes cannot explain the proliferation of repeats such as pseudogenes in this or other rearranged genomes (e.g., Pelargonium [Chumley et al. 2006] and Trachelium [Haberle et al. 2008]). In Trifolium, the endpoints of most rearranged gene clusters are flanked by repeated sequences, tRNAs or pseudogenes (Fig. 3). More sophisticated models of plastid genome evolution may elucidate the roles of multiple pseudogenes and other types of repeats in the evolution of this genome and reveal the purely inversion-based model to be unsuitable for Trifolium.
Distribution of genes and repeats on the Trifolium plastid genome plotted against GC content. The thin line/dots represents genes and the thick line/dots represents repeats. The vertical dashed lines indicate the boundaries of clusters of genes as numbered in Figs. 1 and 2, with the genes at the boundaries labeled. Coordinates indicate positions in the genome (kb). The horizontal dashed line shows the overall GC content in the Trifolium plastid genome
Repeated Sequences
The great proliferation of repeated sequences is one of the most remarkable features of the Trifolium subterraneum plastid genome and these repeats contribute to the unusually large size of the genome in comparison to other legumes lacking one copy of the IR. Among 15 representative eudicot plastid genomes examined, Trifolium exhibits the highest proportion of repeats (19.5%; ca. 28 kb) (Fig. 4). Our genome sampling included six legumes, eight nonlegume rosids, and an asterid. Aside from the highly rearranged plastid genome of the rosid Pelargonium (Geraniaceae), nonlegume plastid genomes contain <5% repeated DNA. However, legumes generally contain more repeated DNA; >5% of the genomes comprise repeated sequence. We divided repeats into four groups based on length: 30–50, 51–100, 101–150, and ≥151 bp (Fig. 5). Most repeats are 30–50 bp in size and occur across all 15 genomes, whereas long repeats (≥151 bp) are more prevalent in the highly rearranged genomes of Trifolium and Pelargonium. Repeats in Trifolium are not evenly distributed across the genome, and most repeats are located within A + T-rich intergenic regions, as well as at rearrangement endpoints (Fig. 3).
Nine genes (psbT, rbcL, clpP, rps3, rpl23, atpB, psbN, ycf3, and trnI-cau) have also been duplicated partially or completely in Trifolium, two of which are not merely duplicated but are present with multiple nonfunctional copies. First, three degenerate copies of the photosynthetic gene psbN are found in Trifolium. Two of these pseudogenes are complete, but with single-base-pair insertions resulting in a frame shift. The two nonfunctional copies are located within repeats differing by a 7-bp deletion and are flanked by two unique DNA fragments. One of the nonfunctional copies of psbN is inserted into part of the highly conserved S10 operon (cluster 6 in Fig. 1).
Repeats of the ribosomal protein gene rpl23 result in some of the most unusual structural features of the Trifolium plastid genome. This family of dispersed repeats was previously identified by Milligan et al. (1989). Analyses of the complete sequence of Trifolium show that portions of rpl23 are repeated six times across the genome, another genomic anomaly not readily explicable through inversions. There is one intact, full-length copy of rpl23 and six nonfunctional, partial copies, with these six partial copies located within dispersed repeats ranging in size from 88 to 2,399 bp (Table 2). Repeats 2–5 are flanked by DNA that is not found in any other sequenced plastid genome.
Transposable Elements
BLAST results showed no significant sequence similarity between Trifolium subterraneum and TEs in the Plant Repeat Database. Furthermore, repeated sequences of Trifolium do not have characteristic inverted repeats that are known to flank some TEs, suggesting that TEs are not found in Trifolium. Thus, the high incidence of dispersed repeats and the extensive rearrangements of the genome are not likely the result of transposition. In addition, we found no evidence of TEs in the other 131 publicly available plastid genomes based on either BLAST similarity or the presence of flanking inverted repeats.
Unique DNA
BLAST comparisons of each of the six legume plastid genomes against 131 available plastid genomes identified unique DNA (30 bp or longer) in each of the legume genomes (Fig. 6). These comparisons identified 19,551 bp of unique DNA in 160 fragments of the Trifolium plastid genome, ranging in size from 30 to 494 bp. The proportion of unique DNA in Trifolium is 2.5–10 times higher than in any other legume.
BLASTN analyses of the 160 unique fragments of Trifolium resulted in no matches to fragments >50 bp, however, there were numerous hits to small fragments, ranging from 14 to 38 bp (results not shown). In view of the very short length of the fragments in the BLASTN results, we performed BLASTX analyses of these same 160 fragments, which resulted in matches for 24 of the sequences (Supplemental Table 1). In most cases, the BLASTX hits had a very low amino acid sequence identity (25–40% amino acid identity for < 50 amino acids) but 18 hits showed either higher sequence identity (>50%) for a short polypeptide (25–50 amino acids) or lower sequence identity (30–50%) for a longer polypeptide (50–137 amino acids). Several of the most significant BLAST hits are notable in terms of their putative identification. Fragment 98, a 427-bp DNA sequence located between Trifolium gene clusters 3 and 11 (Fig. 1), had 147 hits, many of which matched unidentified hypothetical proteins. However, several of these hits had sequence identities ranging from 50% to 65%, and six of the top matches were to batB proteins, which are involved in the amino acid autotransporter system in bacteria (Henderson et al. 2004). Eight other significant hits (62% sequence identity for 27 amino acids) for the same fragment matched a bacterial chitinase gene. Fragment 104, which is 164 bp long and again occurs between gene clusters 3 and 11, has a 46% to 48% sequence identity to a 45-amino acid segment of the plastid gene ycf1 from Medicago. No intact copy of this gene is present in the Trifolium plastid genome.
Discussion
The Trifolium subterraneum plastid genome is unusual in several respects compared to other legumes and angiosperms. It is among the most highly rearranged angiosperm plastid genome sequenced to date. Trifolium, like many other papilionoid legume plastid genomes, such as Cicer and Medicago, has lost one copy of the IR. The Trifolium genome is larger than those of other papilionoid legumes having one copy of the IR. The gene order of Trifolium is also rearranged relative to that of the closely related genus Medicago, which is similar to the ancestral angiosperm genome organization (Raubeson et al. 2007), except for the presence of a large, 50-kb inversion that occurred early in the divergence of papilionoid legumes (Palmer and Thompson 1981; Doyle et al. 1996; Jansen et al. 2008). The Medicago plastid genome is 124 kb, whereas the Trifolium plastid genome is significantly longer, at 144 kb. However, despite this increase in genome size, the gene content of Trifolium has decreased. Six genes and two introns are missing relative to the complement of genes in the ancestral angiosperm plastid genome (Raubeson et al. 2007). In terms of gene loss, Passiflora (Jansen et al. 2007) and Trifolium have the highest number of losses among photosynthetic angiosperms. Passiflora is missing a total of eight genes and four of the six gene losses in Trifolium are shared with Passiflora (accD, infA, rps18, and ycf1). Notably, accD is also partially or completely missing in several lineages, including grasses (Katayama and Ogihara 1996), Acorus (Goremykin et al. 2005), Lobeliaceae (Knox and Palmer 1999), Campanulaceae (Cosner et al. 1997, 2004; Haberle et al. 2008), Oleaceae (Lee et al. 2007), and Pelargonium (Chumley et al. 2006).
Repetitive DNA composed of active or inactive TEs is a major component of plant nuclear genomes, comprising up to 50–60% of maize (San Miguel and Bennetzen 1998; Meyers et al. 2001) and 70% of barley (Vicient et al. 1999), and TEs contribute significantly to nuclear genome size variation (Zhang and Wessler 2004). Additionally, TEs can facilitate genome rearrangements, including inversions, duplications, or deletions of DNA. Aside from an inactive TE, the “Wendy” element in the plastid genome of the alga Chlamydomonas (Fan et al. 1995), TEs have not been identified in plastid genomes. Milligan et al. (1989) proposed that TEs may contribute to the extent of repetitive DNA and to rearrangement in Trifolium. Our results suggest that repeats in Trifolium are not the product of TEs; neither BLAST searches of the Plant Repeat Database nor searches for flanking inverted repeats, characteristic of some elements, indicated the presence of TEs in the plastid genome of Trifolium.
Milligan et al. (1989) also noted the unprecedented extent of unique DNA in the Trifolium plastid genome. All of the sequenced legume plastid genomes contain some unique DNA (Fig. 6) but the origin of this DNA is unclear. There are four possible explanations for the origin of this unique DNA in Trifolium. First, much of this DNA could simply represent noncoding plastid DNA that does not have any matches among publicly available plastid genomes sequences, including five other completely sequenced legume plastid genomes. Although it is difficult to disprove that the unique DNA is merely noncoding plastid DNA, one would expect noncoding plastid DNA from Trifolium to yield some BLAST matches with the closely related legumes Cicer and Medicago (Wojciechowski et al. 2004). Furthermore, this explanation immediately raises further questions as to how Trifolium came to contain such a great abundance of noncoding plastid DNA. Second, some of this unique DNA could represent pseudogenes from the six genes that have been lost from Trifolium. The BLASTX results for unique fragment 104 provides some support for this explanation because this fragment matches a 45-amino acid portion of ycf1 from the Medicago plastid genome (Supplemental Table 1). In Trifolium. this sequence is located adjacent to trnN-GUU, the location of ycf1 in unrearranged angiosperm plastid genomes. Third, the unique DNA could represent intracellular transfer of DNA into the plastid from the Trifolium mitochondrial and nuclear genomes. BLASTX comparisons (Supplemental Table 1) did not identify any matches to sequences from either of these genomes. However, the absence of BLAST hits could be due to the paucity of mitochondrial and nuclear genome sequences for legumes, although there are considerable nuclear data available for Medicago truncatula. Fourth, the unique DNA could have originated via horizontal transfer from other organisms. Horizontal gene transfers into the plastid genome are extremely rare, with only two instances documented, both of which are ancient (Rice and Palmer 2006). Our BLASTX comparisons did identify several strong matches to bacterial genes, including the batB amino acid transport genes from proteobacteria. These genes are involved in the autotransporter secretion pathway in gram-negative bacteria including animal and plant pathogens. Although it is highly speculative at this time, it is possible that some portion of the unique DNA in the Trifolium plastid genome is derived from horizontal transfer from bacterial plant pathogens. Additional investigations into the origin of the unique DNA in the plastid genome of Trifolium and other legumes are needed to clarify the origin of these sequences.
In summary, the organization of the plastid genome of Trifolium is unusual relative to that of other angiosperms. Compared with closely related Medicago, Trifolium shows a highly accelerated rate of genomic rearrangements, with 14–18 inversions, six gene losses, and two intron losses. In addition, the genome contains a large number of repetitive sequences and unique DNA of uncertain origin. Some repeats are associated with the endpoints of rearrangements, and the unique DNA likely represents recently derived segments of the plastid genome, highly divergent remnants of former genes, intracellular transfers from the mitochondrion or nucleus, or horizontal transfers from other genomes, possibly pathogenic bacteria. The Trifolium plastid genome is an excellent model system for examining mechanisms of rearrangements and the evolution of repeats and unique DNA.
References
Bookjans G, Stummann BM, Henningsen KW (1984) Preparation of chloroplast DNA from pea plastids isolated in a medium of high ionic strength. Anal Biochem 141:244–247
Bowman CM, Dyer TA (1986) The location and possible evolutionary significance of small dispersed repeats in wheat ctDNA. Curr Genet 10:931–941
Chang C-C, Lin H-C, Lin I-P, Chow T-Y, Chen H-H, Chen W-H, Cheng C-H, Lin C-Y, Liu S-M, Chang C-C, Chaw S-M (2006) The chloroplast genome of Phalaenopsis aphrodite (Orchidaceae): comparative analysis of evolutionary rate with that of grasses and its phylogenetic implications. Mol Biol Evol 23:279–291
Chumley TW, Palmer JD, Mower JP, Fourcade HM, Calie PJ, Boore JL, Jansen RK (2006) The complete chloroplast genome sequence of Pelargonium × hortorum: organization and evolution of the largest and most highly rearranged chloroplast genome of land plants. Mol Biol Evol 23:2175–2190
Cosner ME, Jansen RK, Palmer JD, Downie SR (1997) The highly rearranged chloroplast genome of Trachelium caeruleum (Campanulaceae): multiple inversions, inverted repeat expansion and contraction, transposition, insertions/deletions, and several repeat families. Curr Genet 31:419–429
Cosner ME, Raubeson LA, Jansen RK (2004) Chloroplast DNA rearrangements in Campanulaceae: phylogenetic utility of highly rearranged genomes. BMC Evol Biol 4:1–17
Darling ACE, Mau B, Blattner FR, Perna NT (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genet Res 14:1394–1403
Doyle JJ, Doyle JL, Palmer JD (1995) Multiple independent losses of two genes and one intron from legume chloroplast genomes. Syst Bot 20:272–294
Doyle JJ, Doyle JL, Palmer JD (1996) The distribution and phylogenetic significance of a 50-kb chloroplast DNA inversion in the flowering plant family Leguminosae. Mol Phylogenet Evol 5:429–438
Ewing B, Green P (1998) Base-calling of automated sequencer traces using phred II. error probabilities. Genome Res 8:186–194
Fan WH, Woelfle MA, Mosig G (1995) Two copies of a DNA element, Wendy, in the chloroplast chromosome of Chlamydomonas reinhardtii between rearranged gene clusters. Plant Mol Biol 29:63–80
Gordon D, Abajian C, Green P (1998) Consed: a graphical tool for sequence finishing. Genome Res 8:195–202
Goremykin VV, Hirsch-Ernst KI, Wolfl S, Hellwig FH (2003) Analysis of the Amborella trichopoda chloroplast genome sequence suggests that Amborella is not a basal angiosperm. Mol Biol Evol 20:1499–1505
Goremykin VV, Hirsch-Ernst KI, Wolfl S, Hellwig FH (2004) The chloroplast genome of Nymphaea alba: whole-genome analyses and the problem of identifying the most basal angiosperm. Mol Biol Evol 21:1445–1454
Goremykin VV, Holland B, Hirsch-Ernst KI, Hellwig FH (2005) Analysis of Acorus calamus chloroplast genome and its phylogenetic implications. Mol Biol Evol 22:1813–1822
Haberle RC, Fourcade HM, Boore JL, Jansen RK (2008) Extensive rearrangements in the chloroplast genome of Trachelium caeruleum are associated with repeats and tRNA genes. J Mol Evol 66:350–361
Hansen DR, Dastidar SG, Cai Z, Penaflor C, Kuehl JV, Boore JL, Jansen RK (2007) Phylogenetic and evolutionary implications of complete chloroplast genome sequences of four early diverging angiosperms: Buxus (Buxaceae), Chloranthus (Chloranthaceae), Dioscorea (Dioscoreaceae), and Illicium (Schisandraceae). Mol Phylogenet Evol 45:547–563
Henderson IR, Navarro-Garcia F, Desvaux M, Fernandez RC, Ala’Aldeen D (2004) Type V protein secretion pathway: the autotransporter story. Micrbiol Mol Biol Rev 68:692–744
Hiratsuka J, Shimada H, Whittier R, Ishibashi T, Sakamoto M, Mori M, Kondo C, Honji Y, Sun CR, Meng BY, Li YQ, Kanno A, Nishizawa Y, Hirai A, Shinozaki K, Sugiura M (1989) The complete sequence of the rice Oryza sativa chloroplast genome - intermolecular recombination between distinct transfer-RNA genes accounts for a major plastid DNA inversion during the evolution of the cereals. Mol Gen Genet 217:185–194
Hupfer H, Swiatek M, Hornung S, Herrmann RG, Maier RM, Chiu WL, Sears B (2000) Complete nucleotide sequence of the Oenothera elata plastid chromosome, representing plastome I of the five distinguishable Euoenothera plastomes. Mol Gen Genet 263:581–585
Jansen RK, Raubeson LA, Boore JL, dePamphilis CW, Chumley TW, Haberle RC, Wyman SK, Alverson A, Peery R, Herman SJ, Fourcade HM, Kuehl JV, McNeal JR, Leebens-Mack J, Cui L (2005) Methods for obtaining and analyzing whole chloroplast genome sequences Molecular evolution: producing the biochemical data part B. Methods Enzymol 395:348–384
Jansen RK, Cai Z, Raubeson LA, Daniell H, dePamphilis CW, Leebens-Mack J, Müller KF, Guisinger-Bellian M, Haberle RC, Hansen AK, Chumley TW, Lee S-B, Peery R, McNeal J, Kuehl JV, Boore JL (2007) Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc Natl Acad Sci USA 104:19369–19374
Jansen RK, Wojciechowski MF, Sanniyasi E, Lee S-B, Daniell H (2008) Complete plastid genome sequence of the chickpea (Cicer arietinum) and the phylogenetic distribution of rps12 and clpP intron losses among legumes (Fabaceae). Mol Phylogenet Evol 48:1204–1217
Katayama H, Ogihara Y (1996) Phylogenetic affinities of the grasses to other monocots as revealed by molecular analysis of chloroplast DNA. Curr Genet 29:572–581
Knox EB, Palmer JD (1999) The chloroplast genome arrangement Lobelia thuliniana Lobeliaceae: expansion of the inverted repeat in an ancestor of the Campanulales. Plant Sys Evol 214:49–64
Lee H-L, Jansen RK, Chumley TW, Kim K-J (2007) Gene relocations within chloroplast genomes of Jasminum and Menodora (Oleaceae) are due to multiple, overlapping inversions. Mol Biol Evol 24:1161–1180
Leebens-Mack J, Raubeson LA, Cui LY, Kuehl JV, Fourcade MH, Chumley TW, Boore JL, Jansen RK, dePamphilis CW (2005) Identifying the basal angiosperm node in chloroplast genome phylogenies: sampling one’s way out of the Felsenstein zone. Mol Biol Evol 22:1948–1963
McCarthy EM, McDonald JF (2003) LTR STRUC: a novel search and identification program for LTR retrotransposons. Bioinformatics 19:362–367
Meyers BC, Tingey SV, Morgante M (2001) Abundance, distribution, and transcriptional activity of repetitive elements in the maize genome. Genome Res 11:1660–1676
Millen RS, Olmstead RG, Adams KL, Palmer JD, Lao NT, Heggie L, Kavanagh TA, Hibberd JM, Gray JC, Morden CW, Caile PJ, Jermiin LS, Wolfe KH (2001) Many parallel losses of infA from chloroplast DNA during angiosperm evolution with multiple independent transfers to the nucleus. Plt Cell 13:645–658
Milligan BG, Hampton JN, Palmer JD (1989) Dispersed repeats and structural reorganization in subclover chloroplast DNA. Mol Biol Evol 6:355–368
Moore MJ, Dhingra A, Soltis P, Shaw R, Farmerie WG, Folta KM, Soltis DE (2006) Rapid and accurate pyrosequencing of angiosperm plastid genomes. BMC Plt Biol 6:17
Moore MJ, Bell CD, Soltis PS, Soltis DE (2007) Using plastid genome-scale data to resolve enigmatic relationships among basal angiosperms. Proc Natl Acad Sci USA 104:19363–19368
Palmer JD (1986) Isolation and structural analysis of chloroplast DNA. Methods Enzymol 118:167–186
Palmer JD, Thompson WF (1981) Rearrangements in the chloroplast genomes of mung bean and pea. Proc Natl Acad Sci USA 78:5533–5537
Pombert J-F, Otis C, Lemieux C, Turmel M (2005) The chloroplast genome sequence of the green alga Pseudendoclonium `akinetum (Ulvophyceae) reveals unusual structural features and new insights into the branching order of chlorophyte lineages. Mol Biol Evol 22:1903–1918
Pombert J-F, Lemieux C, Turmel M (2006) The complete chloroplast DNA sequence of the green alga Oltmannsiellopsis viridis reveals a distinctive quadripartite architecture in the chloroplast genome of early diverging ulvophytes. BMC Biol 4:3
Raubeson LA, Peery R, Chumley T, Dziubek C, Fourcade HM, Boore JL, Jansen RK (2007) Comparative chloroplast genomics: Analyses including new sequences from the angiosperms Nuphar advena and Ranunculus macranthus. BMC Genomics 8:174
Rice DW, Palmer JD (2006) An exceptional horizontal gene transfer in plastids: gene replacement by a distant bacterial paralog and evidence that haptophyte and cryptophyte plastids are sisters. BMC Evol Biol 4:31
San Miguel P, Bennetzen JL (1998) Evidence that a recent increase in maize genome size was caused by the massive amplification of intergene retrotransposons. Ann Bot 82:37–44
Saski C, Lee S-B, Daniell H, Wood TC, Tomkins J, Kim H-G, Jansen RK (2005) Complete chloroplast genome sequence of Glycine max and comparative analyses with other legume genomes. Plant Mol Biol 59:309–322
Shinozaki K, Ohme M, Tanaka M, Wakasugi T, Hayashida N, Matsubayashi T, Zaita N, Chunwongse J, Obokata J, Yamaguchishinozaki K, Ohto C, Torazawa K, Y. Meng B, Sugita M, Deno H, Kamogashira T, Yamada K, Kusuda J, Takaiwa F, Kato A, Tohdoh N, Shimada H, Sugiura M (1986) The complete nucleotide sequence of the tobacco chloroplast genome—its gene organization and expression. EMBO J 5:2043–2049
Tesler G (2002) GRIMM: genome rearrangements web server. Bioinformatics 18:492–493
Vicient CM, Suoniemi A, Anamthawat-Jonsson K, Tanskanen J, Beharav A, Nevo E, Schulman AH (1999) Retrotransposon BARE–1 and its role in genome evolution in the genus Hordeum. Plt Cell 11:1769–1784
Wang R-J, Cheng C-L, Chang C-C, Wu C-L, Su T-M, Chaw SM (2008) Dynamics and evolution of the inverted repeat-large single copy junctions in the chloroplast genomes of monocots. BMC Evol Biol 8:36
Wojciechowski MF, Lavin M, Sanderson MJ (2004) A phylogeny of legumes (Leguminosae) based on analysis of the plastid matK gene resolves many well-supported subclades within the family. Am J Bot 91:1846–1862
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinform 20:3252–3255
Zhang X, Wessler SR (2004) Genome-wide comparative analysis of the transposable elements in the related species Arabidopsis thaliana and Brassica oleracea. Proc Natl Acad Sci USA 101:5589–5594
Acknowledgments
This work was supported by Grant DEB–0120709 from the National Science Foundation to R.K.J. and J.L.B. Part of this work was performed under the auspices of the U.S. Department of Energy, Office of Biological and Environmental Research, by the University of California, Lawrence Berkeley National Laboratory, under contract DE-AC02-05CH11231. We thank Jeff Palmer for providing the purified plastid DNA used in this study and Stephen Downie and two anonymous reviewers for valuable comments on an early draft of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cai, Z., Guisinger, M., Kim, HG. et al. Extensive Reorganization of the Plastid Genome of Trifolium subterraneum (Fabaceae) Is Associated with Numerous Repeated Sequences and Novel DNA Insertions. J Mol Evol 67, 696–704 (2008). https://doi.org/10.1007/s00239-008-9180-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-008-9180-7