Abstract
Biotechnological application of xylanolytic enzymes is normally hindered by their temperature-dependent catalytic property. To satisfy the industrial demands, xylanases that can perform catalysis under cold condition are attracting attention. In this study, the biochemical properties of a predicted xylanase (laXynA) encoded in the genome of marine bacterium Luteimonas abyssi XH031T were characterized. Structure modeling and structure-based sequence alignment indicated that laXynA belongs to the glycoside hydrolase family 10, and it is 20–26% identical to other characterized cold-active xylanases in the same family. Recombinant laXynA was successfully produced in Escherichia coli system by autoinduction and purified by Ni-affinity chromatography. The isolated enzyme showed an optimum temperature of 30 °C toward beechwood xylan and retained important percentage of optimal activity at low temperatures (64, 55, and 29% at 10, 5, and 0 °C, respectively). A remarkable characteristic of laXynA was extreme halophilicity as demonstrated by fourfold enhancement on xylanase activity at 0.5 M NaCl and by maintaining nearly 100% activity at 4 M NaCl. Thin layer chromatography analysis demonstrated that laXynA is an endo xylanase. This study is the first to report the over-expression and characterization of a cold-active xylanase from Luteimonas species. The enzymatic property revealed the cold-active nature of laXynA. The enzyme is a promising candidate in saline food processing application.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Xylan is one of the major structural polysaccharides in plant cells and the second most abundant polysaccharide after cellulose in nature. Given its structural complexity, enzymatic hydrolysis of xylan needs cooperative action of a variety of enzymes (Moreira and Filho 2016). Among these xylanolytic enzymes, endo-xylanase (EC 3.2.1.8), which is normally termed xylanase, is particularly important because it catalyzes cleavage of the internal β-1,4-glycosidic linkages of xylan backbone, thereby generating xylooligosaccharides (XOS) with low degree of polymerization (DP) (Moreira and Filho 2016). On the basis of amino acid sequence and three-dimensional structure, majority of endo-xylanases are categorized as glycosyl hydrolase family 10 and 11 (GH10 and GH11) (http://www.cazy.org/). To date, a large number of endo-xylanases have been characterized and applied in various fields of industry, such as pulp biobleaching (Walia et al. 2017), waste paper deinking (Dhiman et al. 2014), animal feed production (Harris and Ramalingam 2010), bread making (Butt et al. 2008), biofuel production (Bhalla et al. 2015), and prebiotic production (Jain et al. 2015).
Generally, an enzyme performance is highly sensitive to reaction temperature. Biotechnological application of xylanase (similar to many other enzymes) is normally hindered by its temperature-dependent catalytic property. Therefore, in recent years, the use of extremophilic xylanases to satisfy the industrial demands for xylanase that can catalyze under harsh conditions has been attracting interest. Currently, a large number of thermophilic xylanases have been exploited (Kumar et al. 2018). By contrast, less attention is given to psychrophilic or cold-active xylanases, despite their considerable potential applications in many industrial processes. Cold-active xylanase can be especially used in industrial processes where undesirable chemical reactions occur at high temperature (Santiago et al. 2016), and they are particularly suitable in food and feed industry applications (Cavicchioli et al. 2002, 2011). An example of cold-active xylanase application is its usage for improving dough stability and flexibility and for increasing bread volume and crumb structure (Collins et al. 2006; Butt et al. 2008; Dornez et al. 2011). In addition, cold-active xylanase possesses the general advantage of a cold-active enzyme for application, such as energy-saving characteristic and inherently broad substrate specificity relative to its thermophilic counterparts (Santiago et al. 2016).
To date, the number of cold-active xylanases is largely limited (Santiago et al. 2016). No more than 20 cold-active xylanolytic enzymes have been heterogeneously expressed and characterized (Santiago et al. 2016). These cold-active xylanases were generally obtained from cultured psychrophilic microorganisms or metagenomes (Vester et al. 2015). Mining of enzyme from unexplored genome databases (such as GenBank) can be an alternative method to characterize novel enzymes (Lauro et al. 2010; Gong et al. 2013). Therefore, we provided special attention to sequenced genomes of marine microorganisms that are considered reservoirs of novel and extremophilic enzymes (Littlechild 2015). Luteimonas abyssi, XH031T, a novel species of Luteimonas, was isolated from deep-sea sediment of the South Pacific Gyre (genome accession number in GenBank: NZ_KQ759763) (Fan et al. 2014). According to genome annotation, a xylanase (GenBank accession numbers: WP_082672697.1, termed as laXynA) is presumably produced by the strain. In the present study, the amino acid sequence and structural features of putative xylanase were analyzed. The biochemical properties, kinetic parameters, and cleavage patterns were characterized. Our results indicated that laXynA is a typical cold-active endo-xylanase belonging to the GH10 family. Additionally, extreme halophilicity is a prominent characteristic of laXynA.
Materials and methods
Bacterial strains, gene, and chemicals
Escherichia coli DH5α (Invitrogen) and BL21 Rosetta (DE3) (Novagen) were used as host cells for gene cloning and heterologous expression, respectively. DNA fragments encoding full open reading frame (ORF) of laXynA on the pUC57-simple vector (pUC57-laXynA, restriction sites EcoR1 and Not1 were added to the flanking 5′- and 3′-ends of the ORF, respectively) were synthesized by GENEWIZ (Suzhou, China). Xylose (X1) was purchased from Merck Life Science (Shanghai). Xylobiose (X2), xylotriose (X3), xylotetraose (X4), xylopentaose (X5), xylohexaose (X6), and beechwood xylan were purchased from Megazyme (Ireland).
Sequence and structural analyses
The amino acid sequence of laXynA was compared with deposited sequences in the protein database (NCBI) using the BLASTp algorithm on the BLAST server (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Signal peptide prediction was conducted using SignalP 4.1 (http://www.cbs.dtu.dk/services/SignalP/). A three-dimensional model of laXynA was generated by homology modeling using SWISS-MODEL (https://swissmodel.expasy.org/interactive). The credibility of the structural model was evaluated by MolProbity (http://molprobity.biochem.duke.edu/) (Chen et al. 2010). The secondary structure of laXynA was assigned using the DSSP program (http://swift.cmbi.ru.nl/gv/dssp/) (Kabsch and Sander 1983). Structure-based sequence alignment was performed on the T-Coffee server (http://tcoffee.crg.cat/apps/tcoffee/do:regular) and ESPript3.0 (http://espript.ibcp.fr/ESPript/cgi-bin/ESPript.cgi). Structural figures were created using PyMOL (Schrödinger).
Expression and purification of laXynA
pUC57-laXynA plasmids (amplified in E. coli DH5α) were double-digested with EcoR1 and Not1. Gel purified DNA fragments encoding laXynA from the digestion were inserted into pET-28a expression vector. The recombinant plasmid, named pET-28a-laXynA, was verified by DNA sequencing and transformed into E. coli BL21 Rosetta (DE3) competent cells. A 5 ml overnight culture of transformed E. coli BL21 Rosetta (DE3) in Luria–Bertani medium was inoculated into 500 ml ZYM 5052 autoinduction medium in a 2-l flask containing antibiotic kanamycin and chloramphenicol with the final concentration of 100 and 34 µg/ml, respectively. The culture grew at 37 °C until the optical density at 600 nm approximately reached 1. Subsequently, the culture temperature was changed to 20 °C and maintained for 16 h. The cells were harvested by spinning the tubes at 4 °C. The cell pellet resuspended in lysis buffer (500 mM NaCl, 20 mM imidazole, 20 mM Na2HPO4, pH 7.4) was broken by a high-pressure homogenizer. The supernatant of the cell lysate was filtered over a 0.45-µm filter and applied to a 5-ml HisTrap column (GE Healthcare). After equilibrating the column with the lysis buffer on an Äkta prime purifier (GE Healthcare), the recombinant protein was eluted with an elution buffer (500 mM NaCl, 500 mM imidazole, 20 mM Na2HPO4, pH 7.4) in a gradient elution mode. Elution fractions were collected and pooled as indicated by sodium dodecylsulfate polyacrylamide gel electrophoresis (SDS–PAGE). A 10 kDa cut-off concentrator (Millipore) was used to exchange the buffer of the eluted protein for a phosphate buffer (10 mM Na2HPO4, pH 7.4) and concentrate the yielded enzyme properly. The final isolated enzyme was stored in 1-ml aliquots at − 80 °C. The protein concentration was determined by a Bradford protein assay kit (Beyotime, China).
Enzymatic assay and kinetic parameter determination
Xylanase activity was quantified by using the 3,5-dinitrosalicylicacid (DNS) method as described by Bailey et al. (1992). A standard reaction mixture consisted of 0.5 ml of diluted enzyme (approximate 1:20), 1 ml of buffer, and 0.5 ml of beechwood xylan (10 mg/ml). After incubation in a water bath for 15 min, the reaction was stopped with 2 ml of DNS reagent. Afterward, the mixture was heated at 100 °C for 5 min and cooled to room temperature. The absorption of the reaction mixture at 520 nm was measured on a UV/Vis spectrophotometer. Xylose standard was used to plot the calibration curve. One unit (U) of xylanase activity was defined as the amount of enzyme that can release 1 µmol of reducing sugars from beechwood xylan equivalent to xylose per minute. The specific activity was presented as units per milligram of protein. All activity assays were conducted three times.
The Michaelis–Menten constant (Km), maximum rate of reaction (Vmax), and turnover number (kcat) for the isolated laXynA were determined using beechwood xylan as the substrate at concentrations varying from 0.5 to 6 mg/ml in McIlvaine buffer (pH 7) for 15 min. The enzyme kinetic constant values were calculated by constructing double-reciprocal plots.
Effects of temperature and pH on the activity and stability of laXynA
The optimal pH of laXynA was determined at 30 °C in buffers at pH 3–11. pH stability was monitored by assessing the residual activity at pH 7 after incubating the enzyme in buffers with various pH at 4 °C for 2 h. The pH gradient buffers used for enzymatic assays included McIlvaine buffer for pH 3–8 and glycine-NaOH buffer (50 mM) for pH 9–11. The optimal temperature was estimated over the range of 0–90 °C in the McIlvaine buffer of pH 7. The temperature stability of laXynA was evaluated by incubating the enzyme for 2 h at 10–90 °C, cooling on ice, and measuring the residual activity at 30 °C.
Effects of metal ions and NaCl on activity
The effects of different metal ions on the activity of laXynA were investigated by adding 5 mM (final concentration) FeSO4, MnSO4, CaCl2, CoCl2, or MgSO4 to the standard reaction mixture. The effect of salt concentration on xylanase activity was evaluated with 0–4 M NaCl in the reaction mixture at pH 7 and 40 °C.
Analysis of hydrolytic product
Hydrolysis products of beechwood xylan and XOS with backbone length of 2–6 (X2–X6) by laXynA were investigated by thin layer chromatography (TLC). A reaction mixture consisting of 12.5 µl of appropriately diluted enzyme, 12.5 µl of substrate, and 25 µl of McIlvaine buffer (pH 7) was incubated at 40 °C for 1 h in a thermal cycler and the reaction was stopped by heating at 95 °C for 10 min. An appropriate volume of the sample was spotted onto a Silica Gel plate (Merck, Germany). For each of the substrates, a reaction with heat-inactivated laXynA (95 °C, 10 min) was used as negative control. The TLC plate was developed with n-butanol/islpropanol/acetic acid/water (7:5:2:4, v/v). Spots were visualized by spraying the plate with a solution of 3% (w/v) urea in n-butanol/ethanol/water/phosphoric acid (80:8:5:7, v/v) and heated at 115 °C for 20 min in an oven.
Results
Recombinant laXynA production and purification
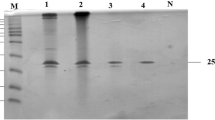
According to the genome annotation of L. abyssi, XH031T, the ORF of laXynA is 1098 bp in length, and it encodes a putative xylanase with 365 residues. The theoretical molecular weight and isoelectric point are 39590.12 Da and 4.53, respectively. Recombinant laXynA was produced in E. coli and purified using His tag affinity chromatography. SDS–PAGE analysis of the fractions eluted by imidazole indicated that the recombinant laXynA displayed a band of approximately 45 kDa, which is larger than the calculated molecular mass of 39590.12 Da because of the additional 34 amino acids in the N-terminus of laXynA derived from pET-28a vector (Fig. 1). The relatively pure fractions were pooled and the protein was applied in the enzymatic assays. The protein concentration of the final isolated recombinant laXynA was about 1 mg/ml.
Biochemical characterization of recombinant laXynA
With beechwood xylan as the substrate, laXynA showed the highest activity at 30–40 °C, peaking at 30 °C (Fig. 2a). The enzyme retained more than 64, 55, and 29% of the maximum activity at 10, 5, and 0 °C, respectively. The effect of pH on the activity of laXynA was investigated at a pH range of 3–11. As shown in Fig. 2b, laXynA is a neutral xylanase (optimal pH of 6.5). However, the enzyme was active across a broad pH range as indicated by the retention of around 40% activity between pH 6 and 9.
Effects of pH and temperature on laXynA activity and stability. a Effect of temperature on the activity of laXynA. b Effect of pH on the activity of laXynA. c Temperature stability of laXynA. The 100% activity was obtained from the enzyme under normal storage temperature (4 °C). d pH stability of laXynA. The 100% specific activity is approximately 30 U/mg. Data are means ± standard deviations from three repeats. The figure was prepared using GraphPad Prism
LaXynA displayed low thermostability as demonstrated by a loss of more than 75% activity after 2 h incubation at 40 °C and almost complete loss of activity after 2 h incubation at 90 °C (Fig. 2c). LaXynA was stable at pH ranging from 6 to 11. It retained more than 90% of the maximum activity after 2 h incubation at this pH range at 4 °C (Fig. 2d).
Among the tested metal ions, Mg2+, Ca2+, and Fe2+ show no evident effect on the activity of laXynA at a concentration of 5 mM; Mn2+ showed negative effect of 0.25-fold whereas Co2+ enhanced the activity slightly (Fig. 3a).
NaCl (0.5–3 M) exerted positive effect on the activity of laXynA (Fig. 3b). When 0.5 M NaCl was added into the reaction mixture, laXynA activity increased by about fourfold relative to the reaction condition with no NaCl added. The degree of improvement decreased with the increased NaCl concentration (Fig. 3b). Nevertheless, the positive effect was present until the NaCl concentration reached 4 M.
Kinetic study on laXynA
A linear relationship between xylanase activity and reaction time was observed from 0 to 15 min at a beechwood xylan concentration of 10 mg/ml (data not shown). Therefore, a reaction time of 15 min was adopted for kinetic assays. Kinetic values were determined at three different temperatures. The maximum velocity of the isolated laXynA at 10 °C was 9 U/mg. This value significantly increased to 18 and 49 U/mg when the reaction temperature was changed to 25 and 40 °C, respectively (Table 1). The Km values for laXynA determined at 10 and 40 °C were 3.2 mg/ml, and the value decreased to 1.6 mg/ml at 25 °C. The kcat values increased with the increase in tested temperatures. The kcat/Km values at 25 and 40 °C were 4.1 and 5.6 times that at 10 °C, respectively (Table 1).
Hydrolytic property of laXynA
Hydrolytic property of laXynA was investigated with beechwood xylan and XOS (X2–X6) as the substrates. The main hydrolytic products of beechwood xylan by laXynA were X2, X4, and a few of X1 after 1 h reaction at 40 °C (Fig. 4). No xylosidase activity was observed for laXynA. X3 was hydrolyzed to X2 and X1 (Fig. 4). The majority of X4 was hydrolyzed to X2. The hydrolytic products of X5 were dominated with X2, X1, X3, and X4. A similar product profile for X5 was observed for X6 under the same reaction condition (Fig. 4).
Hydrolytic products of beechwood xylan and XOS by TLC. Lanes 1 and 2, samples of beechwood xylan incubated with heat-inactivated and untreated laXynA, respectively; lane 3, sample of xylose incubated with untreated laXynA; lanes 4 and 5, samples of xylobiose (X2) incubated with heat-inactivated and untreated laXynA, respectively; lanes 6 and 7, samples of xylotriose (X3) incubated with heat-inactivated and untreated laXynA, respectively; lanes 8 and 9, samples of xylotetraose (X4) incubated with heat-inactivated and untreated laXynA, respectively; lanes 10 and 11, samples of xylopentaose (X5) incubated with heat-inactivated and untreated laXynA, respectively; lanes 12 and 13, samples of xylohexaose (X6) incubated with heat-inactivated and untreated laXynA, respectively. Migration positions of XOS (X1–X6) are indicated by height of XOS abbreviations
Sequence analysis and structural modeling of laXynA
Signal peptide analysis revealed the presence of a signal peptide with cleavage site between Gly24 and Asp25 in laXynA. The cleavage will produce a mature protein of 341 amino acids. BLASTp search against GenBank protein database (mature laXynA as the query sequence) demonstrated that laXynA belongs to the GH10 family and only consists of a catalytic domain. The search also showed that laXynA shared the highest identity (68%) with endo-xylanase from Xanthomonas sp. (XynB).
A structural model of laXynA containing residues 52–307 was built by the SWISS-MODEL server using crystal structure of XynB from Xanthomonas species (Protein Data Bank entry: 4PN2) (Santos et al. 2014) as the template (Fig. 5). The model showed a small MolProbity core of 1.25 (99th percentile), which indicated good quality of the overall structure. The predicted structure of laXynA exhibited a featured (β/α)8-barrel fold of GH10 xylanases (Fig. 5). The secondary structure of laXynA was assigned by DSSP on the basis of the three-dimensional model. A structure-based alignment of catalytic domains indicated that laXynA was not well-conserved among the characterized cold-active GH10 xylanases (Fig. 6). LaXynA shared 20–26% sequence identity with other cold-active GH10 xylanases. The most notable difference between laXynA and other cold-active GH10 xylanases was located in the loop regions, particularly the loops connecting β1 and β2, and β7 and α7 (Fig. 6).
Overall structural model (left) and detail of substrate-binding cleft (right) of laXynA. The carbon atoms of catalytic residues in the overall structure are shown in green. The approximate positions of putative subsites are indicated by large black numbers. The carbon atoms of the amino acids constituting each subsites are shown in different colors, with yellow, green, magenta, and brown for − 2, − 1, + 1, and + 2 subsites, respectively. The carbon atoms of 310-helix where Phe251 is located is shown in cyan. The figure was prepared using PyMOL
Structure-based alignment of the catalytic domain of laXynA with other characterized cold-active GH10 xylanases. The accession numbers of the xylanase sequences are as follows: WP_082672697.1, the xylanase in this study; ADN44261.1, XynGR40 from goat rumen contents; Q21DH6, Xyn10C from Saccharophagus degradans 2–40; AGC01501.1, XynAGN16 from Arthrobacter sp. GN16; WP_013072455.1, XynA from Zunongwangia profunda; WP_012025843.1, Xyn10A from Flavobacterium johnsoniae; AFE82288.1, XynAHJ2 from Bacillus sp. HJ2; AEB69780.1, XynA from Sorangium cellulosum So9733-1; AGA16736.1, Xyn10A from Bacillus sp. SN5; ACN76857.1, XynA from Glaciecola mesophila KMM 241; AAY98787.1, Xyn10 from Flavobacterium sp. Conserved and identical amino acids are highlighted in yellow and red, respectively. Acid/base catalyst and catalytic nucleophile are indicated with red asterisks (*); the putative residues involving substrate-binding are indicated with green asterisks (*). The secondary structural elements are displayed above the corresponding sequences. Gaps are indicated by dashes. The figure was generated by ESPript3.0
The putative catalytic acid–base Glu164 and nucleophile Glu280 were situated at the end of β-strand 4 and 7, respectively (Figs. 5, 6), consistent with the active-site topology of GH10 xylanase (Pollet et al. 2010). Amino acids, either participating in the catalytic reaction or substrate binding in the glycon region of substrate-binding cleft (subsites − 2 and − 1), were highly conserved among cold-active GH10 xylanases (Fig. 6). In the aglycon region, two aromatic amino acids, namely, Tyr216 and Phe251, can be assumed to be the binding sites for + 1 and + 2 xylose residues, respectively. Tyr216 was highly conserved among GH10 xylanases, and Phe251 situated at the bottom of active cleft was only found in laXynA (Figs. 5, 6). A 310-helix, where Tyr216 was located, was also distinct for laXynA (Figs. 5, 6).
Discussion
Xylanase, in particular endo-xylanase, plays the most important role in xylan degradation in nature. Xylanolytic enzymes have also shown considerable potential in applications, such as animal feed, food processing, and textile (Dhiman et al. 2008; Cavicchioli et al. 2011). Exploring novel xylanases and understanding their enzymatic properties are critical for their efficient and effective usage. L. abyssi XH031T is a recently isolated marine bacterium from deep-sea sediment of the South Pacific (Fan et al. 2014). The organism extensively produces cold-active enzymes, including polysaccharide hydrolyases, such as amylase, cellulose, and chitinase (Zhang et al. 2015). In the present study, laXynA, a predicted xylanase encoded in the genome of L. abyssi XH031T, was characterized. Amino acid sequence alignment and three-dimensional structure modeling revealed that laXynA belongs to the GH10 family. It showed the highest identity of 68% with endo-xylanase from Xanthomonas sp. but relatively low identity with other characterized cold-active GH10 xylanases. The optimum temperature of laXynA was approximately 30 °C, and the xylanase retained important percentage of optimal activity (more than 55%) at 5 °C. Meanwhile, laXynA was found to be thermolabile. These properties of laXynA are similar to those of cold-active xylanases characterized from various microbes (Santiago et al. 2016). These results indicated that laXynA is a new member of cold-active xylanase (Table 2).
An excellent feature of laXynA is its extremely halophilicity. The activity of the enzyme was considerably enhanced by NaCl at 0.5–3 M (Fig. 3b). The highest activity of laXynA was observed in the presence of about 0.5 M NaCl, which is close to the salt concentration of seawater (about 0.6 M NaCl); this result indicated that laXynA likely functions in seawater. To date, a few halophilic xylanases have been reported. Among them, Xyn10C from S. degradans 2–40 (cold-active) (Ko et al. 2016), XynA from Z. profunda (cold-active) (Liu et al. 2014), and XynFCB from Thermoanaerobacterium saccharolyticum NTOU1 (thermostable) (Hung et al. 2011a) display their highest activity at 2–3 M NaCl; several halophilic xylanases, including XynA from G. mesophila KMM 241 (cold-active) (Guo et al. 2009), Xyn10A from Bacillus sp. SN5 (cold-active) (Bai et al. 2012), and XynA from T. saccharolyticum NTOU1 (thermostable) (Hung et al. 2011b), just like observation on laXynA, display the highest activity at about 0.5 M NaCl. For all of these halophilic xylanases, the degrees of activity enhancement by NaCl were 1.2–1.9-fold (Liu et al. 2014). By contrast, the activity enhancement by NaCl for laXynA is the most significant (approximately fourfold). Notably, laXynA and all the other halophilic xylanases mentioned in this study are produced by microorganisms inhabiting marine environments or soda lakes, which indicated that the halophilic property was derived from environment-driven adaption. General molecular mechanism of protein adaptation to high salinity based on bioinformatic analysis may be attributed to the excess surface-exposed acid over basic amino acids (DasSarma and DasSarma 2015). LaXynA is a good halophilic protein model to verify the validity of the proposed mechanism. More negatively charged (52, Glu + Asp) residues than the positively charged (23, Arg + Lys) ones were present on laXynA, and all of the acidic amino acids were located on solvent-accessible surface. Our modeled structure revealed that the overall surface of laXynA was largely covered with negative electrostatic potential (Fig. 7).
Surface electrostatic of laXynA. a Surface presentation of active-site cleft side of laXynA. b 90° rotated view relative to (a) c 90° rotated view relative to (b) d 90° rotated view relative to (c) The surface electrostatic is colored from blue (positive potential) to red (negative potential). The figure was prepared using PyMOL
In principle, cold-active enzymes exhibit higher Km value than those of their thermostable counterparts (Georlette et al. 2004). The Km values of thermostable xylanases are normally in the range of 0.1–5 mg/ml (Basit et al. 2018). Nevertheless, most of characterized cold-active xylanases show relatively low Km values (< 5 mg/ml) (Table 2). LaXynA, as a cold-active xylanase, showed relatively low Km value of 1.6 mg/ml determined at 25 °C toward beechwood xylan. Under the similar experimental condition, low Km value was also found for cold-active xylanases, such as XynGR40 from goat rumen contents (1.8 mg/ml) (Wang et al. 2011), XynA from G. mesophila KMM 241 (1.22 mg/ml) (Guo et al. 2009), and Xyn10 from Flavobacterium sp. (1.8 mg/ml) (Lee et al. 2006a) (Table 2). Low Km value (< 1 mg/ml) was also observed for cold-active xylanase; Xyn10A from Bacillus sp. SN5 showed a Km value of 0.5 mg/ml (Bai et al. 2012). Unusually, laXynA showed a twofold higher Km value determined at 10 °C (3.2 mg/ml) and 40 °C (3.2 mg/ml) than that obtained at 25 °C (1.6 mg/ml). This result suggested that the affinity between the enzyme and xylan (or several XOS with particular lengths) reached the optimum at around 25 °C. Temperature-induced structural changes may account for this particular substrate-binding behavior. It has been shown that a temperature-dependent structural modification on substrate-binding cleft of xylanase 10B from Thermotoga petrophila changed the XOS binding at the aglycone subsites (Santos et al. 2010). Several similar studies on cold-active xylanases also showed the varied Km values at different measuring. However, the of Km values changed in different tendencies. XynAGN16L (1.46 and 2.21 mg/ml towards beechwood xylan at 10 and 30 °C, respectively) (Zhou et al. 2015a) and XynA from G. mesophila KMM 241 (0.78 and 1.22 mg/ml toward beechwood xylan at 4 and 30 °C, respectively) (Guo et al. 2009) showed an increased Km value at higher temperature; by contrast, the Km values of xynGR40 (2.2 and 1.8 mg/ml toward beechwood xylan at 10 and 30 °C, respectively) (Wang et al. 2011) and Xyn10 from F. johnsoniae (10.6 and 8.4 mg/ml towards birchwood xylan at 10 and 30 °C, respectively) (Chen et al. 2013) decreased as a result of rising temperature.
The kcat of laXynA increased with the increase in tested temperatures (10, 25, and 40 °C) (Table 1). This phenomenon was also observed for XynA from G. mesophila KMM 241 (Guo et al. 2009), XynGR40 (Wang et al. 2011), and XynAGN16L (Zhou et al. 2015a). LaXynA displayed an intermediate kcat value of 35.6 s−1 at 40 °C. Actually, the kcat values of cold-active xylanases toward xylan determined at similar temperature covered a considerably broad range (6.84–1247 s−1). The kcat value of laXynA is much higher than that of the cold-active xylanases, such as Xyn10 from S. cellulosum So9733-1 (6.84 s−1 toward beechwood xylan at 30 °C) (Wang et al. 2012) and F. johnsoniae. (10.7 s−1 toward beechwood xylan at 35 °C) (Chen et al. 2013). The kcat values are higher but similar to that of laXynA for XynAGN16 (49.2 s−1 toward beechwood xylan at 45 °C) (Zhou et al. 2015b) and XynA from Z. profunda (47.3 s−1 toward beechwood xylan at 30 °C) (Liu et al. 2014). Several cold-active xylanases, mainly belonging to the GH8 family, such as XynA from G. mesophila KMM 241 (609 s−1 toward beechwood xylan at 35 °C) (Guo et al. 2009) and xylanases from Pseudoalteromonas haloplanktis (1247 s−1 toward birchwood xylan at 25 °C) (Collins et al. 2002), show much larger kcat values than that of laXynA.
In principle, to maintain activity at low temperature, an increase in kcat is necessary for cold-active enzymes (Georlette et al. 2004; Santiago et al. 2016). Therefore, among the characterized cold-active xylanases, the xylanase from P. haloplanktis most satisfactorily satisfies the evolutionary principle of a cold-adapted enzyme, as demonstrated by its considerably large Km and kcat values. LaXynA and most of other cold-active xylanases exhibit a relatively small Km and kcat values, which implied that their adaption to cold environment is not sufficient or in a unusual evolutionary road.
The hydrolytic product of xylan produced by most of GH10 endo-xylanases is a mixture of XOS (low DP less than five) with a small percentage of xylose (Linares-Pasten et al. 2018). LaXynA degraded beechwood xylan to X1, X2, and X4 after 1 h incubation at 40 °C, which suggested its endo-xylanase property (Fig. 4). Structural data combined with kinetic activity on XOS have demonstrated that generally, 4–7 subsites are present in the active-site cleft of GH10 endo-xylanases (Pollet et al. 2010), and the highly conserved − 2, − 1, and + 1 subsites play a crucial role in glycosidic bond cleavage (Ducros et al. 2000). In accordance with this canonical property of GH10 endo-xylanase, subsites − 2 to + 1 in the active cleft of laXynA can execute the catalytic cleavage of X3 (Fig. 4). The dominant hydrolytic product of X4 by laXynA was X2, which demonstrated that + 2 to − 2 binding and cleavage was preferred by laXynA (Fig. 4). This preferred cleavage mode can be explained by the presence of Phe251 in the + 2 subsite of laXynA. This aromatic residue likely formed hydrophobic stacking interaction with + 2 xylose residue, which is consistent with the dominant stacking interactions in the aglycon region of GH10 endo-xylanases (Zolotnitsky et al. 2004). In contrast, cold-active endo-xylanases such as XynA (GH10) and XynB (GH8) from G. mesophila KMM 241 were unable to hydrolyze X3 and X4, thereby suggesting the requirement of at least four and five subsites for effective cleavage (Guo et al. 2009, 2013). At least four subsites are also required for cleavage for cold-active Xyn8 (GH8) from an environmental genomic DNA library (Lee et al. 2006b).
Conclusions
We expressed and characterized a new member of cold-active endo-xylanases (laXynA) from marine microorganism. Several characteristics of laXynA are attractive for industry applications. LaXynA degraded xylan to small XOS with a small proportion of xylose, which suggested that it can be used for prebiotic XOS production. LaXynA exhibited excellent activity in high-salinity environment, which implied a potential usage in biotechnological processes of sea food and saline food. However, similar to most of other cold-active xylanases, the catalytic efficiency of laXynA is not as high as that of their mesophilic counterparts (Georlette et al. 2004). Therefore, to search for highly active cold-active xylanases or modify the current cold-active xylanases by protein engineering may be significant. This study reported the over-expression and biochemical characterization of a cold-active xylanase from a Luteimonas species for the first time. This study can also help in elucidating hemicellulose utilization in deep-sea sediment of the South Pacific.
References
Bai W, Xue Y, Zhou C, Ma Y (2012) Cloning, expression and characterization of a novel salt-tolerant xylanase from Bacillus sp. SN5. Biotechnol Lett 34(11):2093–2099
Bailey MJ, Biely P, Poutanen K (1992) Interlaboratory testing of methods for assay of xylanase activity. J Biotechnol 23:257–270
Basit A, Liu J, Rahim K, Jiang W, Lou H (2018) Thermophilic xylanases: from bench to bottle. Crit Rev Biotechnol 17:1–14. https://doi.org/10.1080/07388551.2018.1425662
Bhalla A, Bischoff KM, Sani RK (2015) Highly thermostable xylanase production from a thermophilic Geobacillus sp. strain WsUcF1 utilizing lignocellulosic biomass. Front Bioeng Biotechnol. https://doi.org/10.3389/fbioe.2015.00084
Butt MS, Tahir-Nadeem M, Ahmad Z, Sultan MT (2008) Xylanases and their applications in baking industry. Food Technol Biotechnol 46:22–31
Cavicchioli R, Siddiqui KS, Andrews D, Sowers KR (2002) Low-temperature extremophiles and their applications. Curr Opin Biotechnol 13:253–261
Cavicchioli R, Charlton T, Ertan H, Mohd Omar S, Siddiqui KS, Williams TJ (2011) Biotechnological uses of enzymes from psychrophiles. Microb Biotechnol 4:449–460
Chen VB, Arendall WB 3rd, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, Murray LW, Richardson JS, Richardson DC (2010) MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D 66:12–21
Chen S, Kaufman MG, Miazgowicz KL, Bagdasarian M, Walker ED (2013) Molecular characterization of a cold-active recombinant xylanase from Flavobacterium johnsoniae and its applicability in xylan hydrolysis. Bioresour Technol 128:145–155
Collins T, Meuwis MA, Stals I, Claeyssens M, Feller G, Gerday C (2002) A novel family 8 xylanase, functional and physicochemical characterization. J Biol Chem 277:35133–35139
Collins T, Hoyoux A, Dutron A, Georis J, Genot B, Dauvrin T, Arnaut F, Gerday C, Feller G (2006) Use of glycoside hydrolase family 8 xylanases in baking. J Cereal Sci 43:79–84
DasSarma S, DasSarma P (2015) Halophiles and their enzymes: negativity put to good use. Curr Opin Microbiol 25:120–126
Del-Cid A, Ubilla P, Ravanal MC, Medina E, Vaca I, Levicán G, Eyzaguirre J, Chávez R (2014) Cold-active xylanase produced by fungi associated with Antarctic marine sponges. Appl Biochem Biotechnol 172:524–532
Dhiman SS, Sharma J, Bindu B (2008) Industrial applications and future prospects of microbial xylanases: a review. BioResources 3:1377–1402
Dhiman SS, Garg G, Sharma J, Kalia VC, Kang YC, Lee JK (2014) Reduction in acute ecotoxicity of paper mill effluent by sequential application of xylanase and laccase. PLoS ONE 9:1–13
Dornez E, Verjans P, Arnaut F, Delcour JA, Courtin CM (2011) Use of psychrophilic xylanases provides insight into the xylanase functionality in bread making. J Agric Food Chem 59:9553–9562
Ducros V, Charnock SJ, Derewenda U, Derewenda ZS, Dauter Z, Dupont C, Shareck F, Morosoli R, Kluepfel D, Davies GJ (2000) Substrate specificity in glycoside hydrolase family 10. Structural and kinetic analysis of the Streptomyces lividans xylanase 10A. J Biol Chem 275:23020–23026
Fan X, Yu T, Li Z, Zhang XH (2014) Luteimonas abyssi sp. nov., isolated from deep-sea sediment. Int J Syst Evol Microbiol 64:668–674
Georlette D, Blaise V, Collins T, D’Amico S, Gratia E, Hoyoux A, Marx JC, Sonan G, Feller G, Gerday C (2004) Some like it cold: biocatalysis at low temperatures. FEMS Microbiol Rev 28:25–42
Gong J, Lu Z, Li H, Zhou Z, Shi J, Xu Z (2013) Metagenomic technology and genome mining: emerging are as for exploring novel nitrilases. Appl Microbiol Biotechnol 97:6603–6611
Guo B, Chen XL, Sun CY, Zhou BC, Zhang YZ (2009) Gene cloning, expression and characterization of a new cold-active and salt-tolerant endo-β-1,4-xylanase from marine Glaciecola mesophila KMM 241. Appl Microbiol Biotechnol 84:1107–1115
Guo B, Li PY, Yue YS, Zhao HL, Dong S, Song XY, Sun CY, Zhang WX, Chen XL, Zhang XY, Zhou BC, Zhang YZ (2013) Gene cloning, expression and characterization of a novel xylanase from the marine bacterium, Glaciecola mesophila KMM241. Mar Drugs 11:1173–1187
Harris AD, Ramalingam C (2010) Xylanases and its application in food industry: a review. J Exp Sci 1:1–11
Hung KS, Liu SM, Tzou WS, Lin FP, Pan CL, Fang TY, Sun KH, Tang SJ (2011a) Characterization of a novel GH10 thermostable, halophilic xylanase from the marine bacterium Thermoanaerobacterium saccharolyticum NTOU1. Process Biochem 46:1257–1263
Hung KS, Liu SM, Fang TY, Tzou WS, Lin FP, Sun KH, Tang SJ (2011b) Characterization of a salt-tolerant xylanase from Thermoanaerobacterium saccharolyticum NTOU1. Biotechnol Lett 33:1441–1447
Jain I, Kumar V, Satyanarayana T (2015) Xylooligosaccharides: an economical prebiotic from agroresidues and their health benefits. Indian J Exp Biol 53:131–142
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637
Ko JK, Ko H, Kim KH, Choi IG (2016) Characterization of the biochemical properties of recombinant Xyn10C from a marine bacterium Saccharophagus degradans. Bioprocess Biosyst Eng 39:2–40, 677–684
Kumar V, Dangi AK, Shukla P (2018) Engineering thermostable microbial xylanases toward its industrial applications. Mol Biotechnol 60:226–235
Lauro FM, Allen M, Wilkins D, Williams TJ, Cavicchioli R (2010) Genetics, genomics and evolution of psychrophiles. In: Horikoshi K (ed) Extremophiles handbook. Springer, Tokyo, pp 865–890
Lee CC, Smith M, Kibblewhite-Accinelli RE, Williams TG, Wagschal K, Robertson GH, Wong DWS (2006a) Isolation and characterization of a cold-active xylanase enzyme from Flavobacterium sp. Curr Microbiol 52:112–116
Lee CC, Kibblewhite-Accinelli RE, Wagschal K, Robertson GH, Wong DW (2006b) Cloning and characterization of a cold-active xylanase enzyme from an environmental DNA library. Extremophiles 10:295–300
Linares-Pasten JA, Aronsson A, Karlsson EN (2018) Structural considerations on the use of endo-xylanases for the production of prebiotic xylooligosaccharides from biomass. Curr Protein Pept Sci 19:48–67
Littlechild JA (2015) Enzymes from extreme environments and their industrial applications. Front Bioeng Biotechnol 3:161. https://doi.org/10.3389/fbioe.2015.00161
Liu X, Huang Z, Zhang X, Shao Z, Liu Z (2014) Cloning, expression and characterization of a novel cold-active and halophilic xylanase from Zunongwangia profunda. Extremophiles 18:441–450
Liu Q, Wang Y, Luo H, Wang L, Shi P, Huang H, Yang P, Yao B (2015) Isolation of a novel cold-active family 11 xylanase from the filamentous fungus Bispora antennata and deletion of its N-terminal amino acids on thermostability. Appl Biochem Biotechnol 175:925–936
Moreira LR, Filho EX (2016) Insights into the mechanism of enzymatic hydrolysis of xylan. Appl Microbiol Biotechnol 100:5205–5214
Petrescu I, Lamotte-Brasseur J, Chessa JP, Ntarima P, Claeyssens M, Devreese B, Marino G, Gerday C (2000) Xylanase from the psychrophilic yeast Cryptococcus adeliae. Extremophiles 4:137–144
Pollet A, Delcour JA, Courtin CM (2010) Structural determinants of the substrate specificities of xylanases from different glycoside hydrolase families. Crit Rev Biotechnol 30:176–191
Santiago M, Ramírez-Sarmiento CA, Zamora RA, Parra LP (2016) Discovery, molecular Mechanisms, and industrial applications of cold-active enzymes. Front Microbiol 7:1408. https://doi.org/10.3389/fmicb.2016.01408
Santos CR, Meza AN, Hoffmam ZB, Silva JC, Alvarez TM, Ruller R, Giesel GM, Verli H, Squina FM, Prade RA, Murakami MT (2010) Thermal-induced conformational changes in the product release area drive the enzymatic activity of xylanases 10B: Crystal structure, conformational stability and functional characterization of the xylanase 10B from Thermotoga petrophila RKU-1. Biochem Biophys Res Commun 403:214–219
Santos CR, Hoffmam ZB, de Matos Martins VP, Zanphorlin LM, de Paula Assis LH, Honorato RV, Lopes de Oliveira PS, Ruller R, Murakami MT (2014) Molecular mechanisms associated with xylan degradation by Xanthomonas plant pathogens. J Biol Chem 289:32186–32200
Vester JK, Glaring MA, Stougaard P (2015) Improved cultivation and metagenomics as new tools for bioprospecting in cold environments. Extremophiles 19:17–29
Walia A, Guleria S, Mehta P, Chauhan A, Parkash J (2017) Microbial xylanases and their industrial application in pulp and paper biobleaching: a review. 3 Biotech 7:11. https://doi.org/10.1007/s13205-016-0584-6
Wang G, Luo H, Wang Y, Huang H, Shi P, Yang P, Meng K, Bai Y, Yao B (2011) A novel cold-active xylanase gene from the environmental DNA of goat rumen contents: direct cloning, expression and enzyme characterization. Bioresour Technol 102:3330–3336
Wang SY, Hu W, Lin XY, Wu ZH, Li YZ (2012) A novel cold-active xylanase from the cellulolytic myxobacterium Sorangium cellulosum So9733-1: gene cloning, expression, and enzymatic characterization. Appl Microbiol Biotechnol 93:1503–1512
Zhang L, Wang X, Yu M, Qiao Y, Zhang XH (2015) Genomic analysis of Luteimonas abyssi XH031(T): insights into its adaption to the subseafloor environment of South Pacific Gyre and ecological role in biogeochemical cycle. BMC Genom 16:1092. https://doi.org/10.1186/s12864-015-2326-2
Zhou J, Dong Y, Tang X, Li J, Xu B, Wu Q, Gao Y, Pan L, Huang Z (2012) Molecular and biochemical characterization of a novel intracellular low-temperature-active xylanase. J Microbiol Biotechnol 22:501–509
Zhou J, Liu Y, Shen J, Zhang R, Tang X, Li J, Wang Y, Huang Z (2015a) Kinetic and thermodynamic characterization of a novel low-temperature-active xylanase from Arthrobacter sp. GN16 isolated from the feces of Grus nigricollis. Bioengineered 6:111–114
Zhou JP, Shen JD, Zhang R, Tang XH, Li JJ, Xu B, Ding JM, Gao YJ, Xu DY, Huang ZX (2015b) Molecular and biochemical characterization of a novel multidomain xylanase from Arthrobacter sp. GN16 isolated from the feces of Grus nigricollis. Appl Biochem Biotech 175:573–588
Zolotnitsky G, Cogan U, Adir N, Solomon V, Shoham G, Shoham Y (2004) Mapping glycoside hydrolase substrate subsites by isothermal titration calorimetry. Proc Natl Acad Sci USA 101:11275–11280
Acknowledgements
This work was supported by the Science and Technology Supporting Programme of Wuhan Science and Technology Bureau (Grant Number 2016020101010084).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Han, Z., Shang-guan, F. & Yang, J. Characterization of a novel cold-active xylanase from Luteimonas species. World J Microbiol Biotechnol 34, 123 (2018). https://doi.org/10.1007/s11274-018-2505-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-018-2505-9