Abstract
Overfishing and unsustainable practices have caused drastic declines in shark populations worldwide; these decreases are largely attributed to the demand for shark products (e.g., fins and meat) and shark bycatch associated with the global fishing industry. In an effort to understand the species composition of the shark fin trade in Singapore – a globally significant trade hub, we collected and genetically identify a total of 6840 shark fins collected between January 2021 and February 2022. We then adopted a Bayesian modelling approach to understand how the identified species contributed to the overall trade within Singapore. The Singapore market appears distinct in terms of species composition when compared to the markets of Hong Kong and mainland China. In Singapore 81% of samples belonged to ten species, whereas, in Hong Kong and mainland China a single species dominated the trade. Of those identified in Singapore, the IUCN considers six to be threatened species and eight are listed in CITES Appendix II (in online). The differences in species composition between East and Southeast Asian markets suggest that different supply chains are sustaining these hubs. Given these differences, it is important that conservation, sustainable fisheries management and policy decisions are based upon the findings from multiple trade centres.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The oceans are a significant source of protein that billions of people and coastal communities rely upon each day (Roberson et al. 2020). Biologically and genetically diverse ecosystems, along with sustainably managed extractive activities are critical to the maintenance of ecosystem functioning and the preservation of ecosystem services (Roger et al. 2012; Ceballos et al. 2020; Ceballos and Ehrlich 2023). However, marine ecosystems are increasingly subjected to a variety of unsustainable practices across all habitats globally (Roberson et al. 2020; Pacoureau et al. 2021; Sherman et al. 2023). For example, coral reef habitats are succumbing to rapid human induced climate change and poor land use practices (Abelson 2020), and fisheries throughout the world are collapsing as government subsidized industrial scale fishing, Illegal, Unregulated and Unreported (IUU) fishing, and the associated bycatch removes biodiversity at unprecedented and unsustainable rates (Yan et al. 2021; Welch et al. 2022).
As apex predators and keystone species (Bres 1993), sharks, rays and chimeras (class Chondrichthyes, herein ‘sharks’) are key functional components in marine and coastal ecosystems (Bornatowski et al. 2014; Sherman et al. 2023). Unfortunately, they are also a prime example of a species that humans have driven to the brink of extinction through unsustainable practices (Simpfendorfer and Dulvy 2017; Sherman et al. 2023). The vast majority of shark species are now considered at serious risk of extinction, with some populations experiencing catastrophic declines of 70% or more since 1970 (Pacoureau et al. 2021). Underlining the size of these precipitous declines, sharks are currently the planet’s second most threatened vertebrate lineage (Pacoureau et al. 2021), with once common species of shark now absent on many reefs across the world (Simpfendorfer et al. 2023). Sharks are particularly vulnerable to overfishing and other unsustainable practices because of their slow maturation rates, low reproductive output and lengthy gestation period (Liu et al. 2021). The removal and ensuing absence of apex predators in marine ecosystems can disrupt the delicate balances that maintain ecosystem function and stability. For example, removal can cause the amplification of trophic cascades throughout food chains (Stevens 2000; Bornatowski et al. 2014), and even the grazing behavior of turtles can be altered through the loss of the fear-induced effects that large apex predators such as sharks create (Nowicki et al. 2021).
A significant driver contributing to the overfishing of sharks is the continuing demand for shark fins and shark products (Sherman et al. 2023). Specifically, East and Southeast Asia are regions where significant shark fin consumption occurs, here they are frequently served at celebratory events, or sold as health-promoting tonics (Wainwright et al. 2018). Compounding the unsustainable nature of shark fishing, in particular the fin trade, are the challenges associated with determining what species of shark a fin came from (Drescher et al. 2022). Without an understanding of the species involved in the trade; it becomes very difficult to sustainably manage shark fisheries. Even when species-specific trade regulations are imposed, they are easily circumvented through the use of broad, ambiguous or generic labelling of the traded product (e.g., dried seafood, or dried fish product) (Neo et al. 2022). Further complicating management, very few countries keep or maintain accurate catch data detailing the species and the number caught (Fields et al. 2018). Consequently, understanding the composition of the shark fin trade and designing sustainably managed shark fisheries is challenging, because of this it is estimated that 79% of all fins traded globally come from unsustainably managed sources (Oliver et al. 2015).
DNA barcoding is a routinely applied technique that can aid in the identification of species that are challenging to identify by visual methods alone (Van Houtan et al. 2020). This method generates accurate and reliable results, and has been applied successfully in numerous surveys throughout the world to identify the species of shark that a dried fin, meat or other shark products come from (Cardeñosa et al. 2017, 2022; French and Wainwright 2022). Findings from these barcoding studies further demonstrate the unsustainable nature of the shark fin trade, with many reporting the frequent occurrence of species that are assessed as Critically Endangered or Endangered in the IUCN Red List, or species that have a degree of control imposed upon their international trade through CITES regulations (Wainwright et al. 2018; Hobbs et al. 2019; Haque et al. 2019; Choy and Wainwright 2022; Sharrad et al. 2023).
While DNA barcoding surveys provide a valuable snapshot of the species present at the time of collection, their utility is limited to a small window in time. These one-off, single point in time surveys have been regularly performed throughout Indonesia (Sembiring et al. 2015), Singapore (Wainwright et al. 2018; Choo et al. 2021; Shen et al. 2024), Malaysia (Seah et al. 2022; Loh et al. 2023), Thailand (Klangnurak et al. 2023), Papua New Guinea (Appleyard et al. 2018) and the wider Asia Pacific region (Liu et al. 2013; Bineesh et al. 2017; Thu et al. 2019). In these surveys, threatened species of shark were uncovered along with those that are listed on one of the CITES Appendices, however, it is not possible to reconstruct the full species composition of a trade hub (e.g., Singapore) from these data. As a result of this limitation, Bayesian modeling techniques have been developed to provide a more complete picture of the entire species composition within a given trade hub (Fields et al. 2018). Such understanding will aid regulatory bodies, agencies and policy makers to implement more accurate catch quotas and develop sustainable management and conservation plans. Here, we survey the shark fin market in Singapore, and implement a Bayesian modelling approach to understand the species composition and the contribution of each species within this globally significant trade hub.
Methods
Sample collection
An exhaustive list of shops selling dried shark fins in Singapore was compiled and updated as new shops opened (additions) or closed (removal). We assigned each shop a number, and a random list of 10 shops to visit and collect samples from was created bi-monthly. At each shop, a haphazardly chosen set of at least 20 fins from each available price point, size class, grade, or mixed container was collected; this resulted in totals of between 40 and 181 fins collected per shop, with an average of 97 fins. Note however, due to cultural norms, proprietor reluctance to be involved in work exploring the shark fin trade and the perceived negativity that our work may have on profitable business practices, it was not possible to truly select fins at random, hence the haphazard nature of our collection within shops. All fins were then assigned numbers and 20 fins from each shop were randomly picked and DNA barcoded. All collections were made between January 2021 and February 2022.
DNA barcoding
DNA was extracted from an approximate 25 mg piece of tissue using the BioBasic EZ-10 96 Well Plate Animal Genomic DNA Isolation Kit following the manufacturer’s protocol. Due to the processed nature of the fins and the likely degraded nature of the extracted DNA, we elected to amplify an approximate 350 base pair (bp) fragment of cytochrome c oxidase subunit I (COI) gene using the mlCOIintF (5′-GGW ACW GGW TGA ACW GTW TAY CCY CC-3′) (Leray et al. 2013) and reverse primer LoboR1 (5′-TAA ACY TCW GGR TGW CCR AAR AAY CA-3′) (Lobo et al. 2013). Each Polymerase Chain Reaction (PCR) was performed in a 25 μL volume containing: 12.5 μL GoTaq mastermix green, 7.5 μL of nuclease-free water, and 2 μL of undiluted DNA template, 1 μL Bovine Serum Albumin (BSA) and 1 μL each of the forward primer and the reverse primer, both at 10 mM. Our thermal cycling profile consisted of: 5 repeats of 94 °C for 30 s, 48 °C for 2 min, 72 °C for 1 min, then 35 repeats of 94 °C for 30 s, 54 °C for 2 min, 72 °C for 1 min, then 72 °C for 5 min. If a sample failed to amplify, the amount of input DNA was varied by dilution with water at 1:10, 1:20, 1:100, 1:200. If DNA dilution was unsuccessful, more DNA template was added incrementally until amplification was achieved, up to a total 9.5 μL, with adjustment in the amount of nuclease free water added to maintain a total volume of 25 μL. If amplification continued to fail, DNA was extracted again and the process was repeated until successful. We used PCR negatives throughout to detect possible contamination issues, and no contamination was observed. Enzymatic cleaning and bidirectional Sanger sequencing was performed by Bio-Basic Asia Inc.
Sequence identification
Geneious Prime (v2023.0.4) (Kearse et al. 2012) was used to view and select high quality sequences for downstream analysis. We define a high quality sequence as one that contains no ambiguous base calls, and has only well-defined peaks. Species identifications were considered positive if the Barcode of Life Data System (BOLD) returned a 100% match for a single species, and the same species was then identified in GenBank as the top match. When an exact species match was not possible, we used the top matching genus returned in both databases.
Bayesian modelling analysis
Using an approach similar to that used in Hong Kong (Fields et al. 2018), let \({N}_{jk}\) be the number of fins for day j and shop k, j = 1,…,J, k = 1,…,K. For shops k which are not sampled on day j, we define \({N}_{jk}\) = 0. Write \({y}_{ijk}\) for the number of fins of species i (i = 1,…,I) on day j for shop k, and write \({y}_{jk}=({y}_{1jk},\dots ,{y}_{Ijk})\). In our analysis, rare species – those with fewer than 10 identifications – are aggregated into a single “Others” category. We then constructed a model where we condition on \({N}_{jk}\) and assume that the \({y}_{jk}\) are multinomially distributed,
where
And for i = 2,…,I, \({\alpha }_{i}\) are species effects, \({\beta }_{ij}\), j = 1,…,J, are species-day interactions and \({\gamma }_{ik}\), k = 1,…,K, are species-shop interactions. We identify the model by setting \({\alpha }_{1}\)=0, \({\beta }_{1j}\)= 0, j = 1,…,J, and \({\gamma }_{1k}\) = 0, k = 1,…,K. We consider Bayesian inference, with priors similar to those in Fields et al. (2018), \({\alpha }_{i}\sim \) N (0, 10, 000), i = 2,…,I, \({\beta }_{ij}\sim N(0,{\sigma }_{\beta }^{2})\), i = 2,…,I, j = 1,…,J, \({\sigma }_{\beta }\sim U[\mathrm{0.0001,100}]\), \({\gamma }_{ik}\sim N(0,{\sigma }_{\gamma }^{2})\), i = 2,…,I, k = 1,…,K, \({\sigma }_{\gamma }\sim U[\mathrm{0.0001,100}]\).
Fields et al. (2018) fit their model using a variant of the so-called “Poisson trick”, which however leads to a highly parametrized model and slow mixing in Markov chain Monte Carlo (MCMC) algorithms for drawing samples from the posterior distribution. Hence, here we fit the multinomial model directly, see Gelman et al. (2013) for an introduction to Bayesian methods and MCMC computational algorithms. The model was fitted using the rjags R package V4-13, and we ran four chains from different starting values, with 5000 warm-up iterations and a further 10,000 sampling iterations each. Convergence was assessed by using trace plots and the Gelman-Rubin diagnostic (Gelman et al. 2013). For summarizing the importance of each species, the raw observed proportions for each species can be highly variable and the Bayesian model produces smoothed proportions. To get model-based estimates, we considered the expected proportion of species in a posterior predictive replicate of the data (Gelman et al. 2013), obtaining point estimates and credible intervals for these quantities. All code for recreating the above analyses can be found in the Supplemental Material.
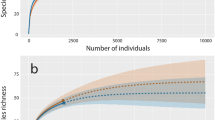
To determine completeness of our data set we used the iNEXT (Chao et al. 2014; Hsieh et al. 2016) online package to generate a rarefaction curve, with 100,000 bootstrap samples and a 95% confidence interval. Using the abundance data collected for each species, we used SpadeR Online (Chao et al. 2016) to estimate the number of species within the Singapore’s market (Table 2).
Results
DNA barcoding
Of the 6840 barcoded fins, we identified 5888 fins to species level but were unable to resolve 952 fins beyond the genus level. Of these 952 genus level identifications, 742 fins belonged to genus Carcharhinus; 196 fins were in the Mustelus genus and the remaining 14 genus level identifications belonged to the genera Glyphis (n = 6), Centrophorus (n = 3), Glaucostegus (n = 2) & Acroteriobatus (n = 3). In total, we identified 52 species of sharks, belonging to 28 genera via DNA barcoding. Using the updated CITES listings that will enter into force with effect from 25 November 2023 (Stokstad 2022), 39 of the species identified are listed on Appendix II (in online), while 36 of the identified species are assessed to be threatened (Critically Endangered = 7, Endangered = 10, Vulnerable = 19) in the IUCN Red List (Table 1). The top 10 species identified in this work accounted for 84% of all the identifications made at the species level. In particular, the silky shark (Carcharhinus falciformis) was the most frequently observed species, with 1171 occurrences accounting for 20% of all the identifications made at the species level.
Modelled species richness in Singapore
Our rarefaction curve (Fig. 1) indicates that our sampling strategy was robust and sufficient to capture the diversity of shark species traded within Singapore’s market and our data is suitable for the modelling approach applied. Further supporting this, modelled species richness (Table 2) is close to the species richness determined by DNA barcoding. In our modelling approach, the top 10 identified species accounted for 81% of the species found within Singapore’s market (Fig. 2). These 10 species come from oceanic and pelagic habitats, and include a mix of species classified as large or small-bodied species at maturity, implying that our work and sampling strategy is not biased towards lower value small-bodied sharks or sharks from one habitat over the other. Carcharhinus spp. accounted for 46% of all the modelled observations. Of the modelled top 10 species, 60% are considered threatened in the IUCN Red List, with 8 species listed on CITES Appendix II (in online), although this top 10 list contains one group (Carcharhinus spp) which could not be fully resolved to the species level. However, all members of this genus are listed on CITES Appendix II.
Bar plot showing the relative frequency of each species as predicted by Bayesian modelling with 95% credible intervals indicated. Asterisks above bars indicates species that listed under CITES Appendix II (in online). Species with fewer than 10 identifications are aggregated into the single category “other”. Those samples that could not be resolved to the species level are recorded at the genus level only (e.g., Carcharhinus spp.)
Discussion
Global shark populations have undergone precipitous declines over recent decades. As apex predators, such population declines can create trophic cascades which disrupt the delicate balances maintaining ecosystem functioning and stability, ultimately this will negatively impact the human communities that rely upon coastal habitats and the services they provide (Sherman et al. 2023; Simpfendorfer et al. 2023). These declines are largely attributed to practices such as overfishing, and are indicative of the unsustainable practices that humans continue to inflict upon the ocean and our planet.
Despite a number of shark species having a degree of control exerted upon their international trade through treaties that are designed to prevent their overexploitation (e.g., CITES), shark populations continue to decline. A significant barrier that hinders the sustainable management of sharks is the difficulty associated with making accurate species identifications from dried and processed fins from which the distinguishing morphological features are generally absent. Frequently, even for experts and enforcement officials, identifying the correct species of shark that a fin came from via visual methods is difficult and often impossible (Klangnurak et al. 2023). Due to these difficulties, many threatened or trade-regulated species of sharks continue to be sold in the shark fin markets throughout the world. DNA barcoding is now a frequently used technique that can overcome these identification difficulties; and similar to other studies that have used this technique to examine the species sold in shark fin markets, this study also found a number of threatened (Critically Endangered, Endangered or Vulnerable IUCN Red Listed species) and CITES-listed species available for sale in Singapore. However, unlike other studies performed in Southeast Asia, we used a Bayesian modelling framework to understand the species composition and the respective contributions of each species to the shark fin market in Singapore between January 2021 and February 2022. This approach is necessary because it is impossible to exhaustively sample the entire market through physical collections. This yearlong sample collection scheme and modelling approach facilitated a better, and more complete understanding of the species composition of the shark fin market in Singapore, thereby providing increasingly comprehensive data that can be used to develop more accurate management and conservation strategies through the implementation of policies that promote sustainable fishing and resource use (Fields et al. 2018; Cardeñosa et al. 2020).
Between January 2021 and February 2022, we estimated that approximately 50% of all the shark fins sold in Singapore that could be identified to the species level came from five species (17% Carcharhinus falciformis, 13% Sphyrna lewini, 9% Galeorhinus galeus, 7% Rhizoprionodon acutus, 7% Carcharhinus sorrah). This list excludes the Carcharhinus identifications that could only be resolved to the genus level, but it is worth noting that this category accounted for 11% of all the modelled identifications and was the third most commonly encountered. Of the five species level identifications listed above, four are listed under CITES Appendix II (in online) and four are currently assessed as Endangered on the IUCN Red List. These five species were also frequently found in one-off surveys performed in Singapore between 2018 and 2022 (Wainwright et al. 2018; Liu et al. 2021; Drescher).
A Bayesian approach similar to the one used in this study was also implemented in the shark fin markets of Hong Kong (Fields et al. 2018) and mainland China (Guangzhou) (Cardeñosa et al. 2020) to assess the species composition in these regions respectively between 2014–2015, and 2015–2017. The rarefaction curves in these studies suggested that the majority of the shark diversity was recovered within the respective markets. In Hong Kong, DNA barcoding identified 67 species, with models predicting at least 76 species are sold domestically. Whereas barcoding revealed 43 species in Guangzhou, with models estimating at least 48 species in the domestic market. In comparison, within Singapore, our barcoding work identified 52 species, with 56 species predicted by the model. Notably, the markets in Hong Kong and Guangzhou were similar to each other in terms of species composition and the proportion that each species contributes, but both differed markedly from Singapore.
The markets of Hong Kong and Guangzhou are dominated by the same single species, the blue shark (Prionace glauca); in Hong Kong, close to 50% of the fins comprised of this single species, and in Guangzhou, approximately 32% of the fins came from the blue shark (Fields et al. 2018; Cardeñosa et al. 2020). Contrastingly, in Singapore, the blue shark accounted for only three percent of the species identified. This low incidence corresponds with findings from recent one-off barcoding research performed in Singapore, which found no P. glauca in 294 barcoded fins collected in Singapore (Drescher et al. 2022). Conservation concerns resulting from the low occurrence of P. glauca in surveys performed in Singapore and the potential negative impacts of unsustainable fishing on this species were previously raised (Drescher et al. 2022). These concerns are further supported by the acknowledgement that P. glauca is one of the most heavily exploited shark species in the world, making up approximately 90% of all oceanic shark catches with scientific evidence indicating that P. glauca is exploited at a level that already exceeds its maximum sustainable yield (Silva et al. 2021). Considering the close proximity of the Hong Kong and Guangzhou markets, it is highly plausible that both markets are supplied by the same, or similar supply chains, consequently P. glauca was the dominant species in both markets, with both showing similar species composition too.
Hong Kong only had one species that occurred at a frequency greater than 5% (P. glauca at a frequency of 50%), while Guangzhou had two that occurred at a frequency greater than 5% (P. glauca & C. falciformis at frequencies of approximately 32% and 10% respectively). Contrastingly, Singapore is not dominated by a single species, and has seven species that occurred at a frequency of greater than 5% (Fig. 2). The more even distribution of species diversity in Singapore suggests a different supply chain to that of the Hong Kong and Guangzhou markets. In addition, notable species identified within the Singapore market includes S. lewini & G. galeus – both assessed as Critically Endangered by the IUCN Red List, and S. lewini is a CITES Appendix II (in online)-listed species. Together, they accounted for 22% of the fins sold in Singapore, while both species were observed in Hong Kong and Guangzhou at much lower frequencies, accounting for less than five percent in both markets (Fields et al. 2018; Cardeñosa et al. 2020).
Although the findings between Hong Kong, Guangzhou and Singapore are broadly comparable, caution must be used when looking for trends across the three markets. Our work performed in Singapore exclusively sampled dried whole fins, whereas the work in Hong Kong and Guangzhou used fin trimmings which are the excess skin, cartilage and muscle tissues that are removed from fins to make them more desirable. It is conceivable that multiple trimmings are required to remove these tissues from a single fin, hence a single individual may be represented multiple times leading to over-representation. Sampling and identifying only dried fins circumnavigates this potential confounding factor.
Moreover, the choice of PCR primers between Singapore, Hong Kong and Guangzhou was not the same – in this work, we selected a primer set that consistently amplified an approximate 350 bp region, while the work in Hong Kong and Guangzhou used primers that amplified a sequence of 120 bp. The decreased resolving power of a shorter fragment likely accounts for the higher proportion of unidentified species seen in Hong Kong and Guangzhou. Nevertheless, where species identifications are made, both primer sets are reliable and give accurate identifications, facilitating comparisons across markets.
Discrepancies across markets could also be a result of sample collections made over a variety of time periods and years. For instance, fin trimmings in Hong Kong were collected between 2014–2015 (Fields et al. 2018), and 2015–2017 in Guangzhou (Cardeñosa et al. 2020), whereas fins in Singapore were collected between 2021 and 2022. It is therefore reasonable to suggest that changing market pressures, implementation of new legislations pertaining to the documentation of landings, or new policies designed to protect sharks (e.g. increased protection through CITES listings and the associated regulations) could have influenced the species composition between markets and sampling years. However, given the surreptitious nature of the fin trade (Sadovy de Mitcheson et al. 2018) and the ease of avoiding rules and regulation onboard fishing vessels when at sea (e.g., finning bans, species-specific catch quotas) (Clarke et al. 2013; Busilacchi et al. 2022), it is questionable whether market and policy forces could have come into play. A more parsimonious explanation would be that the markets in China (mainland and Hong Kong) and Singapore are maintained by different supply chains. In fact, mainland China operates a distant-water fishing fleet that catches and lands sharks that subsequently supply a portion of the Chinese shark fin markets (Pauly et al. 2014). While Singapore does not have a similar fishing fleet, occasional landings of sharks do occur in Singapore ports (Clark‐Shen et al. 2023) albeit on a much smaller scale in comparison to China. Therefore, the vast majority of fins sold in Singapore are imported from other countries. Indeed, as the second-largest importer of shark fin by value in the world, Singapore receives shark fins from multiple and distant regions (e.g., Spain, Namibia, Uruguay) (Boon 2017), and its retail fin market is characterised by more even distribution of species in comparison to Hong Kong and Guangzhou.
This work confirms that a number of endangered or CITES-listed shark species are sold within Singapore, with our model suggesting that over 80% of the fins sold in the retail markets of Singapore could come from species that are threatened or require their international trade to be regulated in order not to further endanger their survival in the wild under the auspices of CITES. However, due to the limitations imposed upon our sample collection by the real world (e.g., the haphazard, rather than truly random nature of collections) care should be taken not to over interpret this. Nevertheless, suggesting that over 80% of the fins could come from threatened or trade regulated species is consistent with what would be expected given the ongoing exploitation of sharks, especially as over 75% of tropical and subtropical coastal species are already considered threatened (Dulvy et al. 2021). The current level of fishing pressure that threatened species of sharks are subject to is unsustainable, estimates suggest that less than 3% of global shark diversity meets the criteria determined for biological sustainability (Simpfendorfer and Dulvy 2017). If shark fishing remains largely unchecked and sharks continue to be harvested at a rate that exceeds their natural reproduction and growth rates, their already dwindling populations mean eventual collapse and extinction is inevitable. However, there is cause for hope, with the effective implementation of strong science-based management practices, sustainable fishing of sharks is possible (Simpfendorfer and Dulvy 2017; Shiffman 2023; Simpfendorfer et al. 2023). Fundamental to this, a comprehensive understanding and documentation of the species involved in the global fin trade is necessary. Understanding this can help determine appropriate catch quotas, develop species-specific conservation and recovery plans, and the implementation of appropriate trade controls under the CITES framework. It also allows assessment of the impacts and success of new regulations and policies that strive towards the sustainable management of shark populations. Modelling approaches such as those used here can contribute important information to these efforts.
Data availability
The data used in this research is still part of an ongoing project, all DNA sequences used will be deposited in the appropriate repository upon completion. Data will be made available upon reasonable request.
References
Abelson A (2020) Are we sacrificing the future of coral reefs on the altar of the “climate change” narrative? ICES J Mar Sci 77:40–45. https://doi.org/10.1093/icesjms/fsz226
Appleyard SA, White WT, Vieira S, Sabub B (2018) Artisanal shark fishing in Milne Bay Province Papua New Guinea: biomass estimation from genetically identified shark and ray fins. Sci Rep. https://doi.org/10.1038/s41598-018-25101-8
Bineesh KK, Gopalakrishnan A, Akhilesh KV et al (2017) DNA barcoding reveals species composition of sharks and rays in the Indian commercial fishery. Mitochondrial DNA Part A 28:458–472. https://doi.org/10.3109/19401736.2015.1137900
Boon PY (2017) The shark and ray trade in Singapore. https://www.traffic.org/site/assets/files/1211/traffic-sg-sharkray-trade.pdf
Bornatowski H, Navia AF, Braga RR et al (2014) Ecological importance of sharks and rays in a structural foodweb analysis in southern Brazil. ICES J Mar Sci 71:1586–1592. https://doi.org/10.1093/icesjms/fsu025
Bres M (1993) The behaviour of sharks. Rev Fish Biol Fisheries 3:133–159. https://doi.org/10.1007/BF00045229
Busilacchi S, Butler JRA, van Putten I et al (2022) Why does illegal wildlife trade persist in spite of legal alternatives in transboundary regions? Hum Dimens Wildl 27:51–68. https://doi.org/10.1080/10871209.2021.1876963
Cardeñosa D, Fields A, Abercrombie D et al (2017) A multiplex PCR mini-barcode assay to identify processed shark products in the global trade. PLoS ONE 12:e0185368. https://doi.org/10.1371/journal.pone.0185368
Cardeñosa D, Fields AT, Babcock EA et al (2020) Species composition of the largest shark fin retail-market in mainland China. Sci Rep 10:12914. https://doi.org/10.1038/s41598-020-69555-1
Cardeñosa D, Shea SK, Zhang H et al (2022) Two thirds of species in a global shark fin trade hub are threatened with extinction: conservation potential of international trade regulations for coastal sharks. Conserv Lett 15:e12910. https://doi.org/10.1111/conl.12910
Ceballos G, Ehrlich PR (2023) Mutilation of the tree of life via mass extinction of animal genera. Proc Natl Acad Sci 120:e2306987120. https://doi.org/10.1073/pnas.2306987120
Ceballos G, Ehrlich PR, Raven PH (2020) Vertebrates on the brink as indicators of biological annihilation and the sixth mass extinction. PNAS. https://doi.org/10.1073/pnas.1922686117
Chao A, Gotelli NJ, Hsieh TC et al (2014) Rarefaction and extrapolation with Hill numbers: a framework for sampling and estimation in species diversity studies. Ecol Monogr 84:45–67. https://doi.org/10.1890/13-0133.1
Chao A, Ma K, Hsieh TC, Chiu C-H (2016) User’s Guide for Online Program SpadeR (Species-richness Prediction And Diversity Estimation in R)
Choo MY, Choy CPP, Ip YCA et al (2021) Diversity and origins of giant guitarfish and wedgefish products in Singapore. Aquat Conserv: Marine Freshw Ecosyst. https://doi.org/10.1002/aqc.3553
Choy CPP, Wainwright BJ (2022) What Is in your shark fin soup? probably an endangered shark species and a bit of mercury. Animals 12:802. https://doi.org/10.3390/ani12070802
Clarke SC, Harley SJ, Hoyle SD, Rice JS (2013) Population trends in pacific oceanic sharks and the utility of regulations on shark finning. Conserv Biol 27:197–209. https://doi.org/10.1111/j.1523-1739.2012.01943.x
Clark-Shen N, Chin A, Arunrugstichai S et al (2023) Status of Southeast Asia’s marine sharks and rays. Conserv Biol. https://doi.org/10.1111/cobi.13962
da Silva TEF, Lessa R, Santana FM (2021) Current knowledge on biology, fishing and conservation of the blue shark (Prionace glauca). Neotrop Biol Conserv 16:71–88. https://doi.org/10.3897/neotropical.16.e58691
Drescher L, Heng NJK, Chin MY et al (2022) Blood in the water: DNA barcoding of traded shark fins in Singapore. Front Mar Sci. https://doi.org/10.3389/fmars.2022.907714
Dulvy NK, Pacoureau N, Rigby CL et al (2021) Overfishing drives over one-third of all sharks and rays toward a global extinction crisis. Curr Biol 31:4773-4787.e8. https://doi.org/10.1016/j.cub.2021.08.062
Fields AT, Fischer GA, Shea SKH et al (2018) Species composition of the international shark fin trade assessed through a retail-market survey in Hong Kong: Shark Fin Trade. Conserv Biol 32:376–389. https://doi.org/10.1111/cobi.13043
French I, Wainwright BJ (2022) DNA Barcoding Identifies Endangered Sharks in Pet Food Sold in Singapore. Front Mar Sci 9:9
Gelman A, Carlin JB, Stern HS et al (2013) Bayesian data analysis, 3rd edn. CRC Press, Cambridge
Haque AB, Das SA, Biswas AR (2019) DNA analysis of elasmobranch products originating from Bangladesh reveals unregulated elasmobranch fishery and trade on species of global conservation concern. PLoS ONE 14:e0222273. https://doi.org/10.1371/journal.pone.0222273
Hobbs CAD, Potts RWA, Walsh MB et al (2019) Using DNA barcoding to investigate patterns of species utilisation in UK shark products reveals threatened species on sale. Sci Rep 9:1–10. https://doi.org/10.1038/s41598-018-38270-3
Hsieh TC, Ma KH, Chao A (2016) iNEXT: an R package for rarefaction and extrapolation of species diversity (Hill numbers). Methods Ecol Evol 7:1451–1456. https://doi.org/10.1111/2041-210X.12613
Kearse M, Moir R, Wilson A et al (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Klangnurak W, Arunrugstichai S, Manopawitr P, Krajangdara T (2023) DNA-based species identification of shark fins traded in thai markets. Conserv Genet 24:537–546. https://doi.org/10.1007/s10592-023-01519-0
Leray M, Yang JY, Meyer CP et al (2013) A new versatile primer set targeting a short fragment of the mitochondrial COI region for metabarcoding metazoan diversity: application for characterizing coral reef fish gut contents. Front Zool 10:34. https://doi.org/10.1186/1742-9994-10-34
Liu S-YV, Chan C-LC, Lin O et al (2013) DNA barcoding of shark meats identify species composition and CITES-listed species from the markets in Taiwan. PLoS ONE 8:e79373. https://doi.org/10.1371/journal.pone.0079373
Liu CJN, Neo S, Rengifo NM et al (2021) Sharks in hot soup: DNA barcoding of shark species traded in Singapore. Fish Res 241:105994. https://doi.org/10.1016/j.fishres.2021.105994
Lobo J, Costa PM, Teixeira MA et al (2013) Enhanced primers for amplification of DNA barcodes from a broad range of marine metazoans. BMC Ecol 13:34. https://doi.org/10.1186/1472-6785-13-34
Loh K-H, Lim K-C, Then AY-H et al (2023) Advancing DNA barcoding to elucidate elasmobranch biodiversity in Malaysian waters. Animals 13:1002. https://doi.org/10.3390/ani13061002
Neo S, Kibat C, Wainwright BJ (2022) Seafood mislabelling in Singapore. Food Control 135:108821. https://doi.org/10.1016/j.foodcont.2022.108821
Nowicki RJ, Thomson JA, Fourqurean JW et al (2021) Loss of predation risk from apex predators can exacerbate marine tropicalization caused by extreme climatic events. J Anim Ecol 90:2041–2052. https://doi.org/10.1111/1365-2656.13424
Oliver S, Braccini M, Newman SJ, Harvey ES (2015) Global patterns in the bycatch of sharks and rays. Mar Policy 54:86–97. https://doi.org/10.1016/j.marpol.2014.12.017
Pacoureau N, Rigby CL, Kyne PM et al (2021) Half a century of global decline in oceanic sharks and rays. Nature 589:567–571. https://doi.org/10.1038/s41586-020-03173-9
Pauly D, Belhabib D, Blomeyer R et al (2014) China’s distant-water fisheries in the 21st century. Fish Fish 15:474–488. https://doi.org/10.1111/faf.12032
Roberson LA, Watson RA, Klein CJ (2020) Over 90 endangered fish and invertebrates are caught in industrial fisheries. Nat Commun 11:4764. https://doi.org/10.1038/s41467-020-18505-6
Roger F, Godhe A, Gamfeldt L (2012) Genetic diversity and ecosystem functioning in the face of multiple stressors. PLoS ONE 7:e45007. https://doi.org/10.1371/journal.pone.0045007
Sadovy de Mitcheson Y, Andersson AA, Hofford A et al (2018) Out of control means off the menu: the case for ceasing consumption of luxury products from highly vulnerable species when international trade cannot be adequately controlled; shark fin as a case study. Mar Policy 98:115–120. https://doi.org/10.1016/j.marpol.2018.08.012
Seah YG, Kibat C, Hew S, Wainwright BJ (2022) DNA barcoding of traded shark fins in Peninsular Malaysia. Rev Fish Biol Fisheries. https://doi.org/10.1007/s11160-022-09713-y
Sembiring A, Pertiwi NPD, Mahardini A et al (2015) DNA barcoding reveals targeted fisheries for endangered sharks in Indonesia. Fish Res 164:130–134. https://doi.org/10.1016/j.fishres.2014.11.003
Sharrad AE, Reis-Santos P, Austin J, Gillanders BM (2023) Umbrella terms conceal the sale of threatened shark species: A DNA barcoding approach. Food Control. https://doi.org/10.1016/j.foodcont.2023.109606
Shen K-LS, Cheow JJ, Cheung AB et al (2024) DNA barcoding continues to identify endangered species of shark sold as food in a globally significant shark fin trade hub. PeerJ 12:e16647. https://doi.org/10.7717/peerj.16647
Sherman CS, Simpfendorfer CA, Pacoureau N et al (2023) Half a century of rising extinction risk of coral reef sharks and rays. Nat Commun 14:15. https://doi.org/10.1038/s41467-022-35091-x
Shiffman DS (2023) Potential for recovery of declining reef sharks. Science 380:1104–1105. https://doi.org/10.1126/science.adi5759
Simpfendorfer CA, Dulvy NK (2017) Bright spots of sustainable shark fishing. Curr Biol 27:R97–R98. https://doi.org/10.1016/j.cub.2016.12.017
Simpfendorfer CA, Heithaus MR, Heupel MR et al (2023) Widespread diversity deficits of coral reef sharks and rays. Science 380:1155–1160. https://doi.org/10.1126/science.ade4884
Stevens J (2000) The effects of fishing on sharks, rays, and chimaeras (chondrichthyans), and the implications for marine ecosystems. ICES J Mar Sci 57:476–494. https://doi.org/10.1006/jmsc.2000.0724
Stokstad (2022) International body likely to protect many shark and ray species. https://www.science.org/content/article/international-body-likely-protect-many-shark-and-ray-species. Accessed 23 Oct 2023
Thu PT, Huang W-C, Chou T-K et al (2019) DNA barcoding of coastal ray-finned fishes in Vietnam. PLoS ONE 14:e0222631. https://doi.org/10.1371/journal.pone.0222631
Van Houtan KS, Gagné TO, Reygondeau G et al (2020) Coastal sharks supply the global shark fin trade. Biol Let 16:20200609. https://doi.org/10.1098/rsbl.2020.0609
Wainwright BJ, Ip YCA, Neo ML et al (2018) DNA barcoding of traded shark fins, meat and mobulid gill plates in Singapore uncovers numerous threatened species. Conserv Genet 19:1393–1399. https://doi.org/10.1007/s10592-018-1108-1
Welch H, Clavelle T, White TD et al (2022) Hot spots of unseen fishing vessels. Sci Adv 8:eabq2109. https://doi.org/10.1126/sciadv.abq2109
Yan HF, Kyne PM, Jabado RW et al (2021) Overfishing and habitat loss drive range contraction of iconic marine fishes to near extinction. Sci Adv 7:16026. https://doi.org/10.1126/sciadv.abb6026
Acknowledgements
CPPC gratefully acknowledges partial support for her time by a grant from the Shark Conservation Fund awarded to the Wildlife Conservation Society Singapore.
Funding
Financial support for this project came from the Minderoo Foundation, flourishing oceans project.
Author information
Authors and Affiliations
Contributions
Conceptualization: BW. Methodology: BW, CPPC. Sample collection: CPPC, NM, BW, CK. Lab work: CK, NM, JAPG, MQU, BW. Model development and testing; DJN. Writing-original draft; BW, CPPC. All authors commented upon, read and approved the final draft.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Murillo Rengifo, N., Choy, C.P.P., Gowidjaja, J.A.P. et al. Determining the species composition of the shark fin trade in Singapore: a globally significant Southeast Asian market. Rev Fish Biol Fisheries 34, 1101–1112 (2024). https://doi.org/10.1007/s11160-024-09861-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11160-024-09861-3