Abstract
A series of novel bivalent metal complexes M(L1)2 and M(L2)2 where M = Cu(II), Ni(II), Co(II) and L1 = 2-((benzo [d] thiazol-6-ylimino)methyl)-4-bromophenol [BTEMBP], L2 = 1-((benzo [d] thiazol-6-ylimino)methyl) naphthalen-2-ol [BTEMNAPP] were synthesized. All the compounds have been characterized by elemental analysis, SEM, Mass, 1H NMR, 13C NMR, UV–Vis, IR, ESR, spectral data and magnetic susceptibility measurements. Based on the analytical and spectral data four-coordinated square planar geometry is assigned to all the complexes. DNA binding properties of these complexes have been investigated by electronic absorption spectroscopy, fluorescence and viscosity measurements. It is observed that these binary complexes strongly bind to calf thymus DNA by an intercalation mode. DNA cleavage efficacy of these complexes was tested in presence of H2O2 and UV light by gel electrophoresis and found that all the complexes showed better nuclease activity. Finally the compounds were screened for antibacterial activity against few pathogens and found that the complexes have potent biocidal activity than their free ligands.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Benzothiazoles have scaffold importance in medicine as it constitutes bioactive pharmacophore in many drugs due to their broad range of biological activity such as anticancer, anti-HIV activity, anti-inflammatory, antimicrobial, antimalarial, analgesic, anticonvulsant, antileishmanial, antitubercular, anticancer lipid peroxidation inhibitor activities [1–11]. A large number of benzothiazole derivatives possess potent anticancer activity and were considered as mechanistically distinct class of clinically proven chemotherapeutic agents [12–14]. To design effective chemotherapeutic agents and better anticancer drugs, the choice of metal ion and planarity of ligand are the most important factors [15]. A plenty of biological experiments were performed so far to suggest that DNA is the primary intracellular target of many anticancer drugs, because the interaction between complexes (small molecules) and DNA can induce DNA damages in cancer cells, blocking the division of aggressive growing cells leading cell death [16–21]. For DNA activity, the most important step is DNA binding interactions with the ligands as well as metal complexes [22, 23]. Metal complexes can bind to DNA through non-covalent modes, such as groove binding, electrostatic binding, intercalative binding [24–27]. Palaniandavar et al. pointed out that Cu(II) complexes are the best alternatives to cis platin. Nickel complexes play an important role in bioinorganic chemistry and redox enzyme systems, and may provide the basis for models of the active sites of biological systems [28]. Cobalt play an important role in biological system (constituent of coenzyme B12) and its complexes exhibit various biological properties viz., antifungal, antitumor, antiproliferative, antiviral, antimicrobial, antioxidant and anticancer [29–31]. DNA cleavage, cytotoxic and antimicrobial studies of binary and ternary Cu(II) complexes of isoxazole Schiff bases and heterocyclic compounds were reported earlier from our laboratory [32, 33].
Keeping in view the above facts, we herein report the synthesis, structural characterization, DNA binding, cleavage and biological studies of Cu(II), Ni(II) and Co(II) complexes of benzothiazole Schiff bases.
Experimental
Materials
All starting moieties, metal salts used for the preparation of ligands and metal complexes were procured from Sigma-Aldrich Bangalore, India. The solvents used for physical measurements were purified according to literature methods [34]. The CT-DNA and supercoiled pBR322 DNA were purchased from Genei, Bangalore and stored at 4 °C. Tris–HCl/NaCl buffer, ethidium bromide obtained from Merck, Hyderabad, India.
Instrumentation
The elemental analysis of the compounds was performed on a Perkin Elmer 240C (USA) elemental analyzer. Metal content of the complexes was estimated by atomic absorption spectroscopy using GBC Avanta 1.0 AAS. Melting points of the compounds were determined on a Polmon instrument (model No. MP-96). 1H and 13C NMR spectra of the ligands were recorded on a Bruker 400 MHz NMR instrument using TMS as internal standard. Morphology and particle-size of the compounds were determined on Zeiss scanning electron microscope. Surface elements of compounds were analyzed by the INCA EDX instrument. ESI mass spectra were recorded on a Vergleichbare Gerate (VG) micro mass 7070–H instrument, IR spectra of the compounds were carried out using KBr discs in the range of 4000–400 cm−1on a Perkin-Elmer Infrared model 337. Electronic spectra were recorded on a Shimadzu UV-Vis 1601 spectrophotometer using DMSO as solvent. Fluorescence spectra were recorded on a Shimadzu RF-5301PC spectrofluorometer. Magnetic susceptibilities of the complexes were determined on a Gouy balance model 7550 using Hg[Co(NCS)4] as standard. Thermograms of complexes were carried out on a Mettler Toledo Star system in the temperature range of 30–1000 °C. EPR spectra of the copper complexes were recorded using JES-FA200 ESR spectrometer (JEOL-Japan) at liquid nitrogen temperature (77 K).
Synthesis of Schiff Bases and Binary Metal Complexes
Synthesis of Schiff Bases L1 and L2
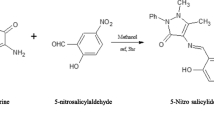
To a hot methanolic solution (50 ml) of 6-Amino benzothiazole (10 mM), hot methanolic solution (50 ml) of 5-bromo salicylaldehyde (10 mM)/2-hydroxy naphthaldehyde (10 mM) was added drop wise with constant stirring and refluxed on an oil bath for 4 h. The resulting solid product was isolated by filtration and recrystallized from hot methanol.
-
L 1 ; (C14H9BrN2OS) Anal. Calc (%): C, 50.46; H, 2.72; N, 8.41; S, 9.62. found: C, 50.68; H, 2.92; N, 8.62; S, 9.92. IR (KBr): (νO-H) 3438, (νCH=N) 1616, (νC-O) 1167. UV-Vis; λmax/nm (cm−1): 270 (37,037), 360 (27,777). 1H-NMR (CDCl3) (δ): 13.12 (s, 1H); 9.01 (s, 1H); 8.63 (s, 1H); 8.16 (s, 1H); 7.86 (d, 1H); 7.54–7.46 (m, 3H); 6.95(d, H) (Shown in Fig. S1). C13 -NMR (CDCl3) (δ): 161.8, 160.1, 154.0, 152.4, 145.7, 136.0, 135.0, 134.3, 124.3, 120.5, 120.0, 119.3, 114.4, 110.6. ESI-MS (m/z): Calc: 333. Found: 335 (M + 2). MP-170 °C.
-
L 2 ; (C18H12N2OS) Anal. Calc (%): C, 71.03; H, 3.97; N, 9.20; S, 10.54. found: C, 71.32; H, 4.23; N, 9.42; S, 10.84. IR (KBr): (νO-H) 3432, (νCH=N) 1619, (νC-O) 1154. UV-Vis; λmax/nm (cm−1):, 243 (41,152),320 (31,250), 360 (25,974). 1H-NMR (CDCl3) (δ): 15.33 (s, 1H); 9.40 (s, 1H); 8.973 (s, 1H); 8.16–7.09 (m, 9H). C13-NMR (CDCl3) (δ): 167.9, 156.4, 154.0, 151.9, 144.3, 136.4, 135.2, 133.0, 129.4, 128.16, 127.5, 124.4, 123.7, 121.3, 119.8, 119.0, 113.3, 109.2. ESI-MS (m/z): Calc: 304. Found: 305(M + 1). MP-190 °C.
Synthesis of Binary Metal Complexes
A hot methanolic solution of Copper acetate mono hydrate (10 mM) was added to a hot methanolic solution of Schiff base ligands (L1/L2) (20 mM) and the resulting mixture was refluxed for 2–4 h. The solid product obtained was separated, washed thoroughly with methanol and dried in vacuum. Similar experimental protocol was used for Ni(II) and Co(II) complexes using Nickel acetate tetra hydrate and Cobalt acetate tetra hydrate. The synthetic procedure of ligands and respective complexes was shown in Scheme 1.
-
1)
[Cu(BTEMBP)2] (1a); (C28H16Br2N4O2S2Cu) Anal. Calc(%): C, 46.20; H, 2.22; N, 7.70; S, 8.81; Cu, 8.73. Found: C, 46.40; H, 2.44; N, 7.50; S, 8.6; Cu, 8.75. IR (KBr): ν(CH=N) 1596, ν(C-O) 1175, ν(M-O) 504, ν(M-N) 435. UV-Vis; λmax/nm (cm−1)(DMSO): 258 (38,759), 357 (28,011), 573 (17,452). μeff (BM): 1.72. ESI-MS (m/z): Clac: 727. Found: 727 [M]+. MP-255 °C.
-
2)
[Ni(BTEMBP)2] (1b); (C28H16Br2N4O2S2Ni) Anal. Calc(%): C, 46.51; H, 2.23; N, 7.75; S, 8.87; Ni, 8.12. Found: C, 46.42; H, 2.52; N, 7.80; S, 8.66; Ni, 8.16. IR (KBr): ν(CH=N) 1592, ν(C-O) 1180, ν(M-O) 537, ν(M-N) 423. UV-Vis; λmax/nm (cm−1)(DMSO): 258 (38,759), 300 (33,333), 525 (19,047), 542 (18,450). μeff (BM): dia. ESI-MS (m/z): Calc: 719. Found: 737 [M + NH4] +. MP-300 °C.
-
3)
[Co(BTEMBP)2] (1c); (C28H16Br2N4O2S2Co) Anal. Calc(%): C, 46.49; H, 2.23; N, 7.75; S, 8.87; Co, 8.15. Found: C, 46.45; H, 2.45; N, 7.85; S, 8.57; Co, 8.17. IR (KBr): ν(CH=N) 1593, ν(C-O) 1176, ν(M-O) 500, ν(M-N) 432. UV-Vis; λmax/nm (cm−1)(DMSO): 260 (38,461), 359 (27,855), 543 (18,416). μeff (BM): 2.11. ESI-MS (m/z): Calc: 720. Found: 721 [M + 1]. MP-270 °C.
-
4)
[Cu(BTEMNAPP)2] (2a); (C36H22N4O2S2Cu) Anal. Calc.(%): C, 64.51; H, 3.31; N, 8.36; S, 9.57. Cu, 9.48. Found: C, 64.62; H, 3.12; N, 8.15; S, 9.87; Cu, 9.46. IR (KBr): ν(CH=N) 1600, ν(C-O) 1190, ν(M-O) 564, ν(M-N) 451. UV- UV-Vis; λmax/nm (cm−1)(DMSO): Vis; λmax/nm (cm−1): 264 (37,878), 326 (30,674), 400 (25,510), 563 (17,761). μeff (BM): 1.8. ESI-MS (m/z): Calc: 669. Found: 692 [M + Na]+. MP-280 °C.
-
5)
[Ni(BTEMNAPP)2] (2b); (C36H22N4O2S2Ni) Anal. Calc.(%): C, 64.98; H, 3.33; N, 8.42; S, 9.64; Ni, 8.82. Found: C, 64.76; H, 3.68; N, 8.24; S, 9.84; Ni, 8.79. IR (KBr): ν(CH=N) 1602, ν(C-O) 1180, ν(M-O) 537, ν(M-N) 423. UV-Vis; λmax/nm (cm−1)(DMSO): 263 (38,022), 325 (30,769), 407 (25,000), 555 (18,018), 572 (17,482). μeff (BM): dia. ESI-MS (m/z): Calc: 664. Found: 664 [M]+. MP-320 °C.
-
6)
[Co(BTEMNAPP)2] (2c); (C36H22N4O2S2Co) Anal. Calc.(%): C, 64.98; H, 3.33; N, 8.42; S, 9.64; Co, 8.85. Found: C, 64.75; H, 3.12; N, 8.57; S, 9.92; Co, 8.87. IR (KBr) ν(CH=N) 1606, ν(C-O) 1188, ν(M-O) 483, ν(M-N) 425. UV-Vis; λmax/nm (cm−1)(DMSO): 277 (36,101), 326 (35,971), 435 (22,988), 567 (17,636). μeff (BM): 2.18. ESI-MS (m/z) Calc: 665. Found: 665 [M]+. MP-290 °C.
DNA Binding Studies
Electronic Absorption Study
The UV–Visible absorption experiments were carried out by maintaining the constant concentration of metal complexes and varying the concentration of the CT-DNA. A Tris-buffer solution of CT-DNA gave a ratio of 1.8–1.9 of UV absorbance at 260 and 280 nm, indicating that the DNA was suitably free of protein [35]. The DNA concentration per nucleotide was measured spectrophotometrically by using known molar extinction coefficient value 6600 M−1 cm−1 at 260 nm [36]. The stock solution of CT-DNA was prepared by diluting DNA in Tris–HCl/NaCl buffer (pH = 7.2, 50 mM NaCl 5 Mm Tris–HCl,). To measure the absorbance of complex and to eliminate the absorbance of CT-DNA itself, equal quantity of CT-DNA was added to both the complex solution and the reference solution. From the absorption data, the intrinsic binding constant (Kb) was calculated by a plot made between [DNA]/(ε a-ε f) and [DNA] [37].
Fluorescence Quenching Study
Competitive CT-DNA in ethidium bromide (EB) displacement study was carried out by pretreating the DNA (15 μM) containing EB (3 μM) in Tris–HCl buffer. To this metal complexes were added in succssive additions. Then samples were excited at 320 nm. The relative binding of the complexes to CT-DNA was determined by Stern–Volmer equation, I0/I = 1 + KSV r, where I0 and I are the fluorescence intensities in the absence and presence of complexes respectively, KSV is a linear Stern–Volmer constant, and r is the concentration of complex to that of DNA.
Viscosity Study
Viscosity measurements were performed on Ostwald capillary viscometer immersed in a thermostatic waterbath at constant temperature (30 ± 1 °C). Metal complexes concentration was varied (0-100 μM) and DNA (100 μM) concentration was kept constant. The relative viscosities were calculated using formula (η/η0)1/3 where η = (t-t0) where t0 and t represent the flow time of DNA solution in the absence and presence of complex and η0 is the viscosity of CT-DNA alone. Average flow time was recorded after triplicates of sample with a digital stopwatch.
DNA Cleavage
Agarose gel electrophoresis was universally accepted method to delineate the interaction between synthesized metal complexes and the supercoiled (SC) pBR322 plasmid DNA. The experiment involves incubating the samples (20 μM) with pBR 322 plasmid DNA (0.2 μg/μL) in Tris-HCl/NaCl buffer (pH 7.2) at 37 °C for 2 h. After incubation bromophenol blue dye was added and then electrophoresed at 50 V for 1 h in Tris buffer using 1 % agarose gel. The resulting bands were blemished with ethidium bromide and images were taken under UV light.
Antibacterial Assay
All synthesized compounds were screened against different bacterial strains by the disc diffusion method using nutrient agar as the medium [38]. In typical procedure, standard and stock solutions of compounds were prepared by dissolving the compounds in DMSO. The plates were inoculated with microorganisms and filled with test compounds for 24 h at 30 °C. During the incubation period, the test solution diffused and the growth of the inoculated microorganisms was affected. The activity was measured in terms of inhibition zone.
Results and Discussion
FT-IR Spectroscopy
IR spectra of compounds provide information about nature of binding mode and functional group attached to metal ion. The binding modes were evaluated by comparing some important IR spectral bands of free ligands with their corresponding metal complexes (Table 1) and (Shown in Fig. S2). The azomethine stretching frequencies of L 1 and L 2 at 1616 and 1619 cm−1 are shifted to higher frequency region to the extent of 10–15 cm−1 in the complexes indicating the nitrogen of azomethine is coordinated to the metal ion [39–42]. The broad bands at 3438 and 3432 cm−1 due to free ν(O-H) stretching frequencies of L 1 and L 2 are disappeared upon complexation indicating coordination through phenolic hydroxyl groups. A medium intensity band around 1167 cm−1due to phenolic ν(C–O) group of the ligand is shifted towards positive side by 15–20 cm−1 [43]. A lower frequency region new bands are observed at 430 and 530 cm−1 due to ν(M-N) and ν(M-O) [44, 45] respectively.
Electronic Spectra and Magnetic Susceptibility
The geometry of metal complexes can be deduced from absorption spectra and magnetic data. Electronic spectra of the ligands and their metal complexes were measured in DMSO solution. The electronic spectra of ligands L 1 and L 2 shows two bands (270 and 360 nm) and three bands (243, 320 and 360 nm) respectively assigned to intraligand π-π* and π-π* transitions. The metal complexes Cu(II), Ni(II) and Co(II) show d-d bands at the range of 400–600 nm (Shown in Fig. S3). Due to 2B1g-2Eg for Cu(II) complexes [46], 1A1g-1A2g and 1A1g-1B1g for Ni(II) complexes [47] and 1A1g-1B1g transition for Co(II) complexes [48, 49]. The magnetic moments of Cu(II) and Co(II) are found to be 1.72 (1a), 1.8 (2a), 2.11 (1c) and 2.18 (2c) respectively whereas Ni (II) complexes are diamagnetic [50]. The electronic transitions and magnetic moments of these complexes are the characteristic features of square planar geometry.
SEM
The SEM analysis was carried out to check the surface morphology of Schiff bases and their metal complexes. All the compounds were analyzed by EDX analysis to find the elements present on surface. Figure 1 depicts the SEM photographs of the synthesized ligand L 1 , and its metal complexes. The broken ice block like particles are observed in the ligand L 1 , the grass like surface is observed in 1a complex, irregular small rock circles like surface is observed in 1b complex and the complex 1c faceted with agglomeration of smaller and larger spherical particles. Figure S4 depicts the SEM photographs of the synthesized ligand L 2 , and its metal complexes. An elongated flake are observed in ligand L 2, needle like particles are observed in 2a complex, the bundle of irregularly broken ice piece shape particles are observed in 2b complex and the spherical rock like particles are observed in 2c. The SEM micrographs revealed that the surface morphology of metal complexes differ from ligand and each other due to the complexation and change of metal ion.
Mass Spectra
Mass spectra provide a preliminary clue for structure elucidation of compounds. The mass spectra of ligands L1 and L2 gave the molecular ion peak at m/z = 335, m/z = 305 respectively. The molecular ion peaks at m/z 727(M+) and 692 (M + Na) of copper complexes are confirmed its stoichiometry as [Cu(L1)2 and Cu(L2)2]. For the Ni(II), Co(II) complexes peaks are observed at m/z 737 [(L1)2Ni + NH4], 721 [(L1)2Co + H], 664 [(L2)2Ni] and 665 [(L2)2Co] (Shown in Fig. S5). The mass spectral results and elemental analysis are in good agreement with 1:2 stoichiometry for the formation of complexes.
Thermogravimetric Analysis
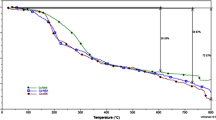
Thermal behaviors of all the complexes are studied by thermogravimetric analysis (TG) under nitrogen atmosphere in the temperature range 30–1000 °C. The TG curves indicated that Cu(II), Ni(II) and Co(II) metal complexes undergo pyrolysis in two stages. The thermograms of complexes (1a, 1b, 1c, 2a, 2b and 2c) show similar decomposition pattern in two steps. The first step corresponds to the removal of starting material. The second step corresponds to removal of total ligand moiety and above this temperature left out was metal oxide (MO) residue. Representative thermograms of 1a and 2a complexes are shown in Fig. 2. In complexes 1a, 2a the first step corresponds to departure of partial ligand moiety in the range of 266–390 °C. Second step degradation within the temperature range of 321–774 °C corresponds to removal of total ligand moiety and remaining left out was due to metal oxide (MO) residue.
ESR Spectra
The ESR spectra of Cu(II) complexes provide very useful information in studying the environment of Cu (II) ion. ESR spectra of 1a and 2a complexes are given in Fig. 3. The g values of the Cu(II) complexes could be used to obtain the ground state [51]. g║, g⊥, Δg, gav and G of the copper complexes have been calculated and presented in Table 2. The trend in the observed g values of copper(II) complexes at liquid nitrogen temperature (77 K) is g║ > g⊥ > ge(2.0023). This trend provides an evidence of localization of the unpaired electron in dx 2 – y 2 orbital indicating square planar geometry. For Cu(II) complexes the G [G = (g║-2.0023)/(g⊥-2.0023)] value found to be 2.442 (1a) and 2.296 (2a) suggesting considerable exchange interactions in Cu(II) complexes [52]. As g║ < 2.3 these complexes are covalent in nature [53].
DNA Binding Studies
Electronic Absorption Study
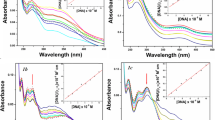
UV-Vis spectroscopy has proven to be one of the most useful methods to determine the binding characteristics of metal complexes with CT-DNA [54, 55]. Absorption spectra of 1a, 1b and 2c are given in Fig. 4. The changes in absorption spectra of metal complexes (10 μM) were recorded in the absence and presence of CT-DNA. Upon increasing the concentration of CT-DNA, (0–10 μM) change in absorbance is effected, resulting in hypochromism (17–36 %) with slight red-shift (2-4 nm) (bathochromism) in all metal complexes indicating intercalative way of binding between DNA duplex and complexes [56, 57]. It was marked that the complexes bind to the base pair of DNA (intercalative), where the π* orbital of the intercalated ligand on the complexes can couple with π orbital of the base pairs, thus decreasing the π–π* transition energies. On the other hand, the coupled π* orbital is partially filled by electrons, thus decreasing the transition probabilities [58]. The extent of the hypochromism depends upon strength of intercalative binding [59]. The electronic absorption spectra of 1a (401 nm), 1b (406 nm), 1c (408 nm), 2a (444 nm), 2b (440 nm) and 2c (435 nm) shows intensive absorption bands. The intrinsic binding constant Kb of the complexes with CT-DNA is determined according to the following equation,
Where [DNA] is the concentration of DNA in the base pairs, Kb is the intrinsic binding constant, εa is apparent coefficient of Aobsd/[complex], εf and εb correspond to the extinction coefficients of the free and fully bound forms of the complex, respectively. Using this formula, the binding constant are found to be 9.6 ± 0.14 × 104 M−1 (1a), 9.39 ± 0.17 × 104 M−1 (2a), 2.3 ± 0.13 × 104 M−1 (1b), 2.12 ± 0.18 × 104 M−1 (2b), 5.73 ± 0.2 × 104 M−1 (1c) and 4.88 ± 0.18 × 104 M−1 (2c). From the above values it is clear that, the copper complexes are strongly bind to DNA than the nickel and cobalt complexes.
Fluorescence Quenching Study
The evidences in favour of intercalative binding mode were also obtained by fluorescence titration studies. EB is a sensitive fluorescence probe, its fluorescence intensity is enhanced in the presence of DNA due to strong intercalation between the DNA base pairs [60, 61]. The experiment involves pretreating EB with DNA then followed by successive addition of complexes to the EB-DNA system. The fluorescence intensity is decreased due to complex competitive binding to EB-bound DNA system due to displacement of bound EB from DNA [62, 63]. As shown in Fig. 5, the fluorescence emission intensities at 619 nm (330 nm excitation) decreased with the increasing complex concentrations, which suggested that complexes could displace DNA-bound EB and bind to CT-DNA at the intercalation sites with almost the same affinity [64]. The Ksv is calculated from the slope of plot I0/I versus r [65]. The apparent binding constant is found to be 8.4–3.32 × 103 M−1. It may be due to the complexes interacting with DNA through intercalation binding, so releasing some free EB from the EB-DNA complex, which is consistent with the above absorption spectral results.
Viscosity Studies
In order to get further confirmation of the intercalative binding nature of the complexes with CT-DNA, viscosity measurements have been performed. Viscosity studies give valuable information regarding binding mode of metal complexes with DNA in the absence of crystallographic structural data [66]. However, a partial and non-classical intercalation of ligand may bend the DNA helix, resulting in the decrease of its effective length and concomitantly its viscosity decrease [66]. The effects of investigated metal complexes on the viscosity of DNA at 30 ± 1 °C are shown in Fig. 6. The experimental results showed that the relative viscosity of CT-DNA increased steadily on successive addition of increasing concentration of complexes. The Cu(II) complexes intercalate stronger than the Ni(II) and Co(II) complexes leading to the greater increase in viscosity of the CT-DNA. All the complexes intercalated between DNA duplex causing an increase in the viscosity of DNA solution. In conclusion, it is observed from experiment that, the complexes (1a-2c) may bind with DNA through intercalative mode.
DNA Cleavage Activities
The DNA cleavage activities of metal complexes with super coiled pBR322 plasmid DNA are carried out by oxidative and photolytic method [67–72]. The cleavage affinity of complexes are confirmed by comparing the band patterns obtained from untreated and treated (without and with complexes) plasmid DNA by oxidative and photolytic methods. When circular plasmid DNA is subjected to electrophoresis, the fast migration will be obtained for the covalently closed circular form. If one strand is nicked, the supercoil will relax to generate a slower moving open circular form (circular, open circular forms are known as Form I and Form II respectively). If both strands are nicked, a linear form that migrates between Form I and Form II will be generated [73].
The cleavage patterns of synthesised Cu(II), Ni(II) and Co(II) metal complexes are shown in Fig. 7. In oxidative method, no DNA cleavage was observed in lane 1 (control), lane 2 (DNA + H 2 O 2 ) and lane 3 (L 1 ) but lane 4 (1a), lane 5 (1b) and lane 6 (1c) are efficiently cleaved supercoiled DNA into nicked form. In photolytic method, no DNA cleavage is observed in lane 1, lane 2 (L 1 ) but lane 3 (1a), lane 4 (1b) and lane 5 (1c) efficiently cleaved into nicked form. In oxidative method DNA cleavage was more compared to photolytic method. Further it is observed that copper complexes promote the cleavage of supercoiled pBR322 DNA more efficiently than Nickel and Cobalt complexes. In all complexes 1a, 1b and 1c are shown greater cleavage property than 2a, 2b and 2c complexes.
a Oxidative cleavage of supercoiled pBR322 DNA(0.2 μg, 33.3 μM) at 37 °C in 5 mM Tris HCl/5 mM NaCl buffer by the metal complexes. Lane 1, DNA control; Lane 2, DNA + H2O2 (1 mM); Lane 3, DNA + H2O2 (1 mM) + L1; Lane 4, DNA + H2O2 (1 mM) + 1a (20 μM); Lane 5, DNA + H2O2 (1 mM) + 1b (20 μM); Lane 6, DNA + H2O2 (1 mM) +1c (20 μM). b Photoactivated cleavage of supercoiled pBR322 DNA (0.2 μg, 33.3 μM) at 37 °C in 5 mM Tris HCl/5 mM NaCl buffer by the complexes UV irradiation of wavelength 345 nm. Lane 1, DNA control; Lane 2 DNA + L1 (20 μM); Lane 3, DNA + 1a (20 μM); Lane 4, DNA + 1b (20 μM); Lane 5, DNA + 1c (20 μM)
Antibacterial Activity
In vitro antibacterial activities of all the ligands and their complexes were tested against Gram-positive (Bacillus subtilis) and Gram-negative bacteria (Escherichia coli, Pseudomonas putida, Klebsiella pneumonia.), shown in Fig. 8 and Table 3. All the metal complexes (20 mg/mL in DMSO) showed significant activity compared to free ligand. Here the standard drug employed was Ampicillin. The higher activity of the complexes is explained through Overtone’s concept [74] and chelation theory. According to Overtone’s concept of cell permeability, the lipid membrane that surrounds the cell favors the passage of only lipid soluble materials. Antimicrobial activity depends on liposolubility. On chelation, the polarity of the metal ion is reduced to a greater extent owing to the overlap of the ligand orbital and partial sharing of positive charge of the metal ion with donor groups. Further, it increases the delocalization of π -electrons over the whole chelating ring and augments the penetration of the complexes into lipid membranes and enzymes of microorganisms will be blocked by the metal binding sites. According to the microbial studies of the synthesized compounds the copper complexes are more active than nickel and cobalt complexes. The higher activity of copper complexes is deduced from the fact that increases in the size of the metal ion, decreases the polarization and further explained on the basis of chelation theory [75].
Conclusion
In the present investigation, two novel Schiff bases and their (Cu(II), Ni(II) and Co(II)) complexes have been synthesized and thoroughly characterized by analytical and spectral techniques. From the spectral data it is observed that all complexes tentatively adopted a square planar geometry. The binding mode of complexes with CT-DNA was investigated by UV-Vis spectroscopy, fluorescence studies and viscosity measurements. Binding studies reveal that all complexes are binding with CT-DNA through an intercalative mode. The DNA cleavage studies of Cu(II), Ni(II) and Co(II) complexes revealed that these complexes effectively cleaved supercoiled pBR322 DNA both in the presence of H2O2 and UV light. Further the antibacterial activity of all the synthesized ligands and their metal complexes revealed that all complexes showed high antibacterial activity compared to ligands.
References
Dua R, Shrivastava S, Sonwane SK, Srivastava SK (2011) Pharmacological significance of synthetic Heterocycles scaffold: a review. Adv Biol Res 5:120–144
Soni B, Ranawat MS, Sharma R, Bhandari A, Sharma S (2010) Synthesis and evaluation of some new benzothiazole derivatives as potential antimicrobial agents. Eur J Med Chem:2938–2942
Shafi S, Alam MM, Naveen M, Chaitanya M, Vanaja G, Arunasree M, Kalle RP, Alam MS (2012) Synthesis and in vitro anti-HIV activity of N-1,3-benzo[d]thiazol-2-yl-2-(2-oxo-2H-chromen-4-yl)acetamide derivatives using MTT method. Eur J Med Chem 49:324–333
Bhavsar D, Trivedi J, Parekh S, Savant M, Thakrar S, Bavishi A, Radadiya A, Vala H, Lunagariya J, Parmar M, Paresh L, Loddo R, Shah A (2011) Synthesis and in vitro anti-HIV activity of N-1,3-benzo[d]thiazol-2-yl-2-(2-oxo-2H-chromen-4-yl)acetamide derivatives using MTT method. Bioorg Med Chem Lett 21:3443–3446
Delmas F, Avellaneda A, Giorgio CD, et al. (2004) Synthesis and antileishmanial activity of (1,3-benzothiazol-2-yl) amino-9-(10H)-acridinone derivatives. Eur J Med Chem 39:685–690
Pereira GA, Massabni AC, Castellano EE (2012) A broad study of two new promising antimycobacterial drugs: Ag(I) and Au(I) complexes with 2-(2-thienyl)benzothiazole. Polyhedron 38:291–296
Burger A, Sawhey SN (1968) Antimalarials III benzothiazole amino alcohols. J Med Chem 11:270–273
Sharpe TR, Cherkofsky SC, Hewes WE, DH s, WA G, SB H, MR L, JG W (1985) Preparation and antiarthritic and analgesic activity of 4,5-diaryl-2-(substituted thio)-1H-imidazoles and their sulfoxides and sulfones. J Med Chem 28:118–1194
Amir M, Asif S, Ali I, Hassan MZ (2012) Synthesis of benzothiazole derivatives having acetamido and carbothioamido pharmacophore as anticonvulsant agents. Med Chem Res 21:2661–2670
Kashiyama E, Hutchinson I, Chua MS, Stinson S, Phillips LR, Kaur G, Sausville EA, Bradshaw TD, Westwell AD, Stevens MFG (1999) Antitumor benzothiazoles. 8.1 synthesis, metabolic formation, and biological properties of the C- and N-oxidation products of antitumor 2-(4-aminophenyl) benzothiazoles. J Med Chem 42:4172–4184
Karali N, Guzel O, Ozsoy N, Ozbey S, Salman A (2010) Synthesis of new spiroindolinones incorporating a benzothiazole moiety as antioxidant agents. Eur J Med Chem 45:1068–1077
Bradshaw TD, Chua MS, Orr S, Matthews CS, Stevens MFG (2000) Mechanisms of acquired resistance to 2-(4-aminophenyl)benzothiazole (CJM 126, NSC 34445). Br J Cancer:270–277. doi:10.1054/bjoc.2000.1231
Chua MS, Kashiyama E, Bradshaw TD, Stinson S, Brantley E, Sausville EA, Stevens MFG (2000) Role of CYP1A1 in modulation of antitumor properties of the novel agent 2-(4-amino-3-methylphenyl)benzothiazole (DF 203, NSC 674495 in human breast cancer Cells1. Cancer Res 60:5196–5203
Shi DF, Bradshaw TD, Wrigley S, McCall CJ, Lelieveld P, Stevens MFG (1996) Antitumor benzothiazoles. 3.1 synthesis of 2-(4-aminophenyl) benzothiazoles and evaluation of their activities against breast cancer cell lines in vitro and in vivo. J Med Chem 39:3375–3384
Arjmand F, Muddassir M, Khan RH (2010) Chiral preference of L-tryptophan derived metal-based antitumor agent of late 3d-metal ions (Co(II), Cu(II) and Zn(II)) in comparison to D- and DL-tryptophan analogues: their in vitro reactivity towards CT DNA. Eur J Med Chem 45:3549–3557
Metcalfe C, Thomas JA (2003) Kinetically inert transition metal complexes that reversibly bind to DNA. Chem Soc Rev 32:215–224
Rad FV, Housaindokht MR, Jalal R, Hosseini HE, Doghaei AV, Goghari SS (2014) Spectroscopic and molecular modeling based approaches to study on the binding behavior of DNA with a copper (II) complex. J Fluoresc 24:1225–1234
Navarro M, Cisneros Fajardo EJ, Sierralta A, Mestre MF, Silva P, Arrieche D, Marchan E (2003) Design of copper DNA intercalators with leishmanicidal activity. J Biol Inorg Chem 8:401–408
Corral E, Hotze ACG, Tooke DM, Spek AL, Reedijk J (2006) Ruthenium polypyridyl complexes containing the bischelating ligand 2, 2′-azobispyridine. Synthesis characterization and crystal structures. Inorg Chim Acta 359:830–838
Hotze ACG, Faiz JA, Mourtzis N, Pascu GI, Webber PRA, Clarkson GJ, Kopoulou KY, Pikramenou HMJ (2006) Far-red luminescent ruthenium pyridylimine complexes; building blocks for multinuclear arrays. Dalton Trans 24:3025–3034
Ljubijankic N, Zahirovic A, Turkusic E, Kahrovic E (2013) DNA binding properties of two ruthenium(III) complexes containing Schiff bases derived from salicylaldehyde: spectroscopic and electrochemical evidence of CT DNA intercalation. Croat Chem Acta 86(2):215–222
Uma V, Vaidyanathan VG, Nair Bull BU (2005) Synthesis, structure, and DNA binding studies of copper(II) complexes of Terpyridine derivatives. Chem Soc Jpn 78:845. doi:10.1246/bcsj.78.845
Shahabadi N, Kashanian S, Darabi F (2010) DNA binding and DNA cleavage studies of a water soluble cobalt(II) complex containing dinitrogen Schiff base ligand: the effect of metal on the mode of binding. Eur J Med Chem 45:4239
Raman N, Joseph J, Velan ASK (2006) Antifungal activities of biorelevant complexes of copper (II) with biosensitive macrocyclic ligands. Mycobiology 34(4):214–218
Ma XF, Li DD, Tian JL, Kou YY, Yan SP (2009) DNA binding and cleavage activity of reduced amino-acid Schiff base complexes of cobalt(II), copper(II), and cadmium(II). Transit Met Chem 34:475–481
Kozurkova M, Sabolova D, Janovec L, Mikes J, Koval J, Ungvarsky V, Stefanisinova M, Fedorocko P, Kristian P, Imrich J (2008) Cytotoxic activity of proflavinediureas: synthesis, antitumor, evaluation and DNA binding properties of 1′,1″-(acridin-3,6-diyl)-3′, 3″ dialkyldiureas. Bioorg Med Chem 16:3976–3984
Li Y, Yang Z (2010) Rare earth complexes with 3-Carbaldehyde Chromone-(benzoyl) Hydrazone: synthesis, characterization, DNA binding studies and antioxidant activity. J Fluoresc 20:329–342
Shafaatian B, Soleymanpour A, Oskouei NK, Notash B, Rezvani SA (2014) Synthesis, crystal structure, fluorescence and electrochemical studies of a new tridentate Schiff base ligand and its nickel(II) and palladium(II) complexes. Spectro chimica Acta Part A: Molecular and Biomolecular Spectroscopy 128:363–369
Bottcher A, Takeuchi T, Hardcastle KI, Meade TJ, Gray HB (1997) Spectroscopicand electrochemical study on axial and in-plane ligand effects in cobalt(III) Schiff-base complexes. Inorg Chem 36:2498–2504
Takeuchi T, Bottcher A, Quezada CM, Meade TJ, Gray HB (1999) Inhibition ofthermolysinand human alpha-thrombin by cobalt(III) Schiff base complexes. Bioorg Med Chem 7:815–819
Manikandan R, Viswanathamurthi P, Velmurugan K, Nandhakumar R, Hashimoto T, Endo A (2014) Synthesis, characterization and crystal structure of cobalt(III) complexes containing 2-acetylpyridine thiosemicarbazones: DNA protein interaction, radical scavenging and cytotoxic activities. J Photochem Photobiol B 130:205–216. doi:10.1016/j.jphotobiol.2013.11.008
Vijaykumar C, Sathishkumar K, Ramesh M, Parthasarathy T, Shivaraj (2014) DNA cleavage cytotoxic activitie and antimicrobial studies of ternary copper (II) complexes of isoxazole Schiff base and heterocyclic compounds. Hindawi Pub bioinochem and app. doi:10.1155/2014/691260
PradeepKumar M, Tejaswi S, Rambabu A, VeerendraKumar AK, Shivaraj (2015) Synthesis, crystal structure, DNA binding and cleavage studiesof copper(II) complexes with isoxazole Schiff base. Polyhedron 102:111–120
Vogel AI, Mendham J (2000) Vogel's textbook of quantitative chemical analysis. Prentice Hall, New York
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from micro-organisms. J Mol Biol 3:208–218
Reichmann ME, Rice SA, Thomas CA, Doty P (1954) A further examination of the molecular weight and size of Desoxypentose nucleic acid. J Am Chem Soc 76:3047–3053. doi:10.1021/ja01640a067
Wolfe A, Shimer GH, Meehan T (1987) Polycyclic aromatic hydrocarbons physically intercalate into duplex regions of denatured DNA. Biochemistry 26:6392–6396. doi:10.1021/bi00394a013
Satyanaryana S, Dabrowial JC, Chaires JB (1993) Tris(phenanthroline)ruthenium(II) enantiomer interactions with DNA: mode and specificity of binding. Biochemistry 32:2573
Maity D, Drew MGB, Godsell JF, Roy S, Mukhopadhyay G (2010) Synthesis and characterization of Cu(II) complexes of tetradentate and tridentate symmetrical Schiff base ligands involving o-phenelenediamine, salicylaldehyde and diacetylmonoxime. Transit Met Chem 35:197–204
Bhunora S, Mugo J, Bhaw-Luximon A, Mapolie S, Van Wyk J, Darkwa J, Nordlander E (2011) The use of Cu and Zn salicylaldimine complexes as catalyst precursors in ring opening polymerization of lactides: ligand effects on polymer characteristics. Appl Organo Metal Chem 25:133–145
Youssef NS, El-Zahany E, El-Seidy AMA, Caselli A, Fantauzzi S, Cenini S (2009) Synthesis and characterisation of new Schiff base metal complexes and their use as catalysts for olefin cyclopropanation. Inorg Chim Acta 362:2006–2014
Percy GC, Thornton DA (1973) Infrared spectra of N-aryl salicylaldimine complexes substituted in both aryl rings. J Inorg Nucl Chem 35:2319–2327
Saydam S, Yilmaz E (2006) Synthesis, characterization and thermal behavior of 4-chloromethyl-2-(2-hydroxybenzilidenehydrazino) thiazole and its complexes with Cr(III), Co(II), Ni(II) and Cu(II). Spectro chim Acta A 63:506–510
Nakamoto K (1997) Infrared and Raman spectra of inorganic and coordination compounds, fifth edn. Wiley-Interscience, New York
Bellamy LJ (1980) The infrared spectra of complex molecules, second edn. Chapmann and Hall, London
Rahaman A, Imatiyaz Y, Afazal M, Srivastva S, Saikrishna S, Farukh (2015) Enantiomeric fluoro-substituted benzothiazole Schiff base-valine Cu(II)/Zn(II) complexes as chemotherapeutic agents: DNA binding profile, cleavage activity, MTT assay and cell imaging studies. A Journal of Photochemistry and Photobiology B: Biology 143:61–73
Takjoo R, Centore R (2013) Synthesis, X-ray structure, spectroscopic properties and DFT studies of some dithiocarbazate complexes of nickel(II). J Mol Struct 1031:180–185
Lever ABP (1984) Inorganic Electronic Spectroscopy (2nd edn) Elsevier, Amsterdam
Carabineiro SA, Silva LC, Gomes PT, Pereira CJ, Veiros LF, SI p, MD T, Namorado S, RT h (2007) Synthesis and characterization of tetrahedral and square planar bis(iminopyrrolyl) complexes of cobalt(II). Inorg Chem 46:6880–6890
Chandrasekar T, Pravin N, Raman N (2015) DNA incision evaluation, binding investigation and biocidal screening of novel metallonucleases of 1,7-bis(4-hydroxy-3-methoxyphenyl)hepta-1,6-diene-3,5-dione based Knoevenagel condensate having methionine: synthesis and structural validation. J Mol Struct 1081:477–485
Patel RN, Singh N, Shukla KK, Chauhan KK, Gutierrez NJ, Castineiras A (2004) Magnetic, spectroscopic, structural and biological properties of mixed-ligand complexes of copper(II) with N,N,N ′,N″,N″-pentamethyldiethylenetriamine and polypyridine ligands. Inorg Chim Acta 357:2469–2476
Hathaway BJ, Billing DE (1970) The electronic properties and stereochemistry of mono-nuclear complexes of the copper(II) ion. Coord Chem Rev 5:143–207
Kivelson D, Neiman R (1961) ESR studies on the bonding in copper complexes. J Chem Phys 35:149–155
Barton JK, Danishefsky AT, Goldberg J (1984) Tris(phenanthroline)ruthenium(II): stereoselectivity in binding to DNA. J Am Chem Soc 106:2172–2176
Tysoe SV, Morgan RJ, Baker AD, Strekas TC (1993) Spectroscopic investigation of differential binding modes of. DELTA.- and. LAMBDA.-Ru(bpy)2(ppz)2+ with calf thymus DNA. J Phys Chem 97:1707–1711
Liu HK, Sadler PJ (2011) Metal complexes as DNA intercalators. Acc Chem Res 44:349–359
Pyle AM, Rehmann JP, Meshoyrer R, Kumar CV, Turro NJ, Barton JK (1989) Mixed-ligand complexes of ruthenium(II): factors governing binding to DNA. J Am Chem Soc 111:3053–3063
Li TR, Yang ZY, Wang BD, Qin DD (2008) Synthesis, characterization, antioxidant activity and DNA-binding studies of two rare earth(III) complexes with naringenin-2-hydroxy benzoyl hydrazoneligand. Eur J Med Chem 43:1688–1695
Sudhamani CN, BhojyaNaik HS, RavikumarNaik TR, Prabhakara MC (2009) Synthesis, DNA binding and cleavage studies of Ni(II) complexes with fused aromatic N-containing ligands. Spectro Chim Acta A 72:643–647
Meyer-Almes FJ, Porschke D (1993) Mechanism of intercalation into the DNA double helix by ethidium. Biochemistry 32:4246–4253
Lepecq JB, Paoletti C (1967) A fluorescent complex between ethidium bromide and nucleic acids: physical—chemical characterization. J Mol Biol 27:87–106
Baguley BC, Le Bret M (1984) Quenching of DNA-ethidium fluorescence by amsacrine and other antitumor agents: a possible electron-transfer effect. Biochemistry 23:937–943
Pasternack RF, Caccam M, Keogh B, Stephenson TA, Williams AP, Gibbs EJ (1991) Long-range fluorescence quenching of ethidium ion by cationic porphyrins in the presence of DNA. J Am Chem Soc 113:6835–6840
Chen J, Wang X, Chao Y, Zhu JH, Zhu YG, Li YZ, Xu Q, Guo ZJ (2007) A Trinuclear copper(II) complex of 2,4,6-tris(di-2-pyridylamine)-1,3,5-triazine shows prominent DNA cleavage activity. Inorg Chem 46:3306–3312. doi:10.1021/ic0614162
Lakowicz JR, Weber G (1973) Quenching of fluorescence by oxygen probe for structural fluctuations in macromolecules. Biochemistry 12:4161–4170
Satyanarayana S, Dabrowiak JC, Chaires JB (1992) Neither DELTA- nor LAMBDA-tris(phenanthroline)ruthenium(II) binds to DNA by classical intercalation. Biochemistry 31:9319
Li X, Bi CF, Fan YH, Zhang X, Wei XD, Meng XM (2014) Synthesis, characterization, DNA binding and cleavage properties of a ternary copper(II) Schiff base complex transition. Met Chem 39:577–584
Santra BK, Reddy PAN, Neelakanta G, Mahadevan S, Nethaji M, Chakravarty AR (2002) Oxidative cleavage of DNA by a dipyridoquinoxaline copper(II) complex in the presence of ascorbic acid. J InorgBiochem 899(3–4):191–196
Tsang SY, Tam SC, Bremner I, Burkitt MJ (1996) Copper-1,10-phenanthroline induces internucleosomal DNA fragmentation in HepG2 cells, resulting from direct oxidation by the hydroxyl radical. Biochem J 317:13–16
Bernadou J, Pratviel G, Bennis F, Girardet M, Meunier B (1989) Potassium monopersulfate and a water-soluble manganese porphyrin complex, [Mn(TMPyP)](OAc)5, as an efficient reagent for the oxidative cleavage of DNA. Biochemistry 28:7268–7275
Tamboura FB, Gaye M, Sall AS, Barry AH, Jouini T (2002) Synthesis, properties and X-ray structure for the mononuclear complex of [{(1-methyl imidazol-2-yl)methylene}-2-aminoethylpyridine]-dichloro copper (II) monohydrate. Inorg Chem Commun 5:235–238
Khoramdareh ZK, Hosseini-Yazdi SA, Spingler B, Khandar AA (2014) Copper(II) and zinc(II) complexes of mono- and tri-linked aza crown macrocycles: synthesis, characterization, X-ray structure, phosphodiester hydrolysis and DNA cleavage. Inorg Chim Acta 415:7–13
Sigman DS (1986) Nuclease activity of 1,10-phenanthroline-copper ion. Acc Chem Res 19:180–186
Anjaneyulu Y, Rao RP (1986) Preparation, characterization and antimicrobial activity studies on some ternary complexes of Cu(II) with Acetylacetone and various salicylic acids. Synth React Inorg Met Org Chem 16:257–272
Tweedy BG (1964) Plant extracts with metal ions as potential antimicrobial agents. Phytopathology 55:910–914
Acknowledgments
Authors express sincere thanks to the Head, Department of Chemistry for providing the necessary facilities, the Director, CFRD, Osmania University, Hyderabad, the Director, IICT, Hyderabad, and the SAIF, IIT Bombay for providing spectral and analytical data. We are also thankful to CSIR, New Delhi, DST-SERB and UGC-UPE (FAR) for providing financial assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vamsikrishna, N., Kumar, M.P., Tejaswi, S. et al. DNA Binding, Cleavage and Antibacterial Activity of Mononuclear Cu(II), Ni(II) and Co(II) Complexes Derived from Novel Benzothiazole Schiff Bases. J Fluoresc 26, 1317–1329 (2016). https://doi.org/10.1007/s10895-016-1818-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10895-016-1818-z