Abstract
Purpose
To report genetic characteristics and associated risk of chromosomal breaks due to chromosomal rearrangements in large samples.
Methods
MicroSeq, a technique that combines chromosome microdissection and next-generation sequencing, was used to identify chromosomal breakpoints. Long-range PCR and Sanger sequencing were used to precisely characterize 100 breakpoints in 50 ABCR carriers.
Results
In addition to the recurrent regions of balanced rearrangement breaks in 8q24.13, 11q11.23, and 22q11.21 that had been documented, we have discovered a 10-Mb region of 12q24.13-q24.3 that could potentially be a sparse region of balanced rearrangement breaks. We found that 898 breakpoints caused gene disruption and a total of 188 breakpoints interrupted genes recorded in OMIM. The percentage of breakpoints that disrupted autosomal dominant genes recorded in OMIM was 25.53% (48/188). Fifty-four of the precisely characterized breakpoints had 1–8-bp microhomologous sequences.
Conclusion
Our findings provide a reference for the evaluation of the pathogenicity of mutations in related genes that cause protein truncation in clinical practice. According to the characteristics of breakpoints, non-homologous end joining and microhomology-mediated break-induced replication may be the main mechanism for ABCRs formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chromosome breakage and reconnection often result in apparently balanced chromosome rearrangements (ABCRs), such as reciprocal translocations and inversions, which have a prevalence of 0.52% in infants [1]. Chromosome rearrangements are often detected in prenatal diagnosis [2,3,4], children with developmental abnormalities [5,6,7], couples with recurrent miscarriages [8, 9], or infertile men [10]. Chromosome structure changes can occur during the processes of DNA replication, recombination, or repair [11, 12]. Generally, ABCRs with abnormal phenotypes are the result of disruption of autosomal dominant (AD) genes or haploinsufficient genes [13,14,15]. Most individuals with ABCRs present no abnormal phenotypes. During meiosis, errors in the pairing and separation of homologous chromosomes can lead to the production of chromosomally unbalanced gametes, resulting in spontaneous abortions or the birth of children with intellectual disabilities and developmental abnormalities [16,17,18]. Although prenatal diagnosis and preimplantation genetic testing have been successfully applied to prevent the birth of children with unbalanced rearrangements, the evaluation of genetic risks due to ABCRs remains of great significance. By sequencing the breakpoints of ABCRs and determining whether they disrupt genes, we can more accurately assess the risk of transmitting Mendelian genetic diseases, which is essential for providing precise personalized genetic counseling to prevent birth defects.
Given the intricate relationship between genes and diseases, reports have indicated that a few genes associated with autosomal dominant (AD) diseases can also demonstrate an autosomal recessive (AR) inheritance pattern. For example, hereditary spastic paraplegia (HSP), caused by mutations in the ERLIN2 gene, can be inherited in either a dominant or recessive manner. However, dominantly inherited ERLIN2 mutations are more likely to be associated with typical HSP symptoms, which may be due to the dominant negative effect of mutations in the SPFH structural domain of the ERLIN2 gene [19]. The p.T258M heterozygous mutation in the KIF1A gene was observed to have a dominant negative effect in a study of a familial complex HSP [20]. However, despite our limited knowledge of the genome, there is still disagreement about whether truncating mutations in certain AD genes are pathogenic. In a large cohort of triple-negative breast cancer patients, nine individuals were found to have only protein-truncating mutations of the BARD1 gene [21]. Despite the low mutation rate of BARD1 in the general population, the correlation between these gene mutations and breast cancer has yet to be fully explored [22]. Moreover, different types of mutations of the same gene can result in disparate disease phenotypes. For example, the p.I157T missense mutation in the multiorgan cancer susceptibility gene CHEK2 is associated with a lower risk of breast cancer than the c.1100delC truncating mutation [23]. Follow-up of carriers with ABCRs that interrupt AD genes may provide insight into whether the truncating mutations in these AD genes are pathogenic.
The main mechanisms that lead to chromosomal rearrangements include nonhomologous end joining (NHEJ), non-allelic homologous recombination (NAHR), replication fork stalling and template switching (FoSTeS), and microhomology-mediated break-induced replication (MMBIR) [24,25,26,27,28,29]. NHEJ can repair double-stranded DNA breaks by directly linking the broken ends, which is thought to be the primary cause of chromosome translocations and other chromosomal structural aberrations [30]. Another investigation of balanced chromosome rearrangements revealed that microhomologous sequences were present at 74% (17/23) of the translocation breakpoints [31], indicating that NHEJ is a major contributor to chromosome rearrangements. Homologous recombination between region-specific low-copy repeat sequences (LCR) typically leads to NAHR [24], which has been identified as the cause of many genomic diseases [26]. FoSTeS and MMBIR are involved in complex chromosome variations. FoSTeS is an active replication fork stalling mechanism during DNA replication. It involves switching to another DNA template with microhomologous sequences [27], thus connecting the DNA being replicated to another DNA. During DNA replication, MMBIR can repair single-strand breaks (SSBs). However, structural variations can be introduced, as single-stranded DNA with microhomology to the 3′ DNA of SSB may be mistakenly involved in the repair process [28].
In this study, we analyzed 2441 chromosomal structural rearrangement breakpoints in 1219 ABCR carriers using a modified MicroSeq technique and found that the 10-Mb region of 12q24.13-q24.3 may be a region sparse for balanced rearrangement breaks. Further analysis of the breakpoints revealed that 42.72% (898/2102) of breakpoints disrupted known genes, and in the number of breakpoints that interrupted the genes recorded in OMIM, 25.53% (48/188) of breakpoints interrupted the AD genes. We also identified 100 chromosomal structural breakpoints in 50 balanced rearrangement carriers and characterized sequences near the breakpoints. This result suggests that NHEJ and MMBIR may be the primary mechanisms underlying ABCR formation.
Materials and methods
Study subjects
The study was conducted after approval by the Reproductive and Genetic Hospital of CITIC Xiangya Ethics Committee (LL-SC-2016-002) and informed consent was obtained from the patients, including 1182 cases of balanced translocation, 12 cases of inversion, and 3 cases of insertional translocation. Robertsonian translocations were excluded from this study since they only involved centric fusions between acrocentric chromosomes. All patients sought PGT treatment after detection of ABCRs for primary or secondary infertility or spontaneous abortion after marriage, and no significant phenotypic abnormalities were observed in the clinical records.
Genomic DNA preparation
Genomic DNA from blood from the patients was extracted by Qiagen Blood DNA Kits (Qiagen), according to manufacturer’s instruction. Quality and quantity of the extracted DNA were measured using NanoDrop 1000 Spectrophotometer and agarose gel electrophoresis. For all subjects, DNA extracted from whole blood was used for junction breakpoint PCR experiments.
Preparation of metaphase chromosome division phase in peripheral blood lymphocytes
Karyotypes were determined from G-banding analysis using standard protocol according to the ISCN nomenclature. Metaphase chromosomes were prepared from cultured lymphocyte cells obtained from carriers by standard techniques.
Modified MicroSeq
Chromosome microdissection-based next-generation sequencing (MicroSeq) was performed as described previously [32] with certain improvements. The improvements include (1) adding 10 μL of nucleic acid-free sterilized water dropwise to the split phase region before microdissection of the target fragment to facilitate firm attachment of the cut fragment to the capillary glass needle tip during excision and (2) changing the DOP-PCR amplification to the application of the PicoPLEX or WGA4 kit, which greatly reduced the amplification time of the cut product. The Ion Xpress™ Plus Fragment Library Kit (Thermos Fisher Scientific) was used to construct the sequencing library following the manufacturer’s instructions. The library DNA was amplified using the Ion PGM™ Template OT2 200 Kit (Thermos Fisher Scientific) for 5.5 h according to the manufacturer’s instructions. About 8–10 identical derived chromosomes were used for amplification, and amplification according to the WGA4 kit instructions procedure yielded approximately 2.8 μg of DNA product. About 2 ng of DNA product was used for each sequencing to construct the sequencing library.
The universal forward and reverse primer sequences (CCGACTCGAG) contained in the read sequences were removed and sequences were aligned to the human chromosome HG19 using the Burrows-Wheeler Alignment (BWA) software, removing sequences that were repeatedly aligned to multiple positions. Based on the number of sequence matches, the location of the derivative chromosome breakpoint was detected, and the characteristics of the region at that location included: (1) both derived chromosomes had sequences at that location; (2) each derived chromosome had a significantly higher number of sequences in that region than the other regions.
Cross-breakpoint PCR
The primers were designed to amplify cross-breakpoint fragments based on the specific sequences of NGS sequencing reads compared to HG19, with reference to the range of breakpoints predicted by the chromosome location, and the effective cross-breakpoint PCR primer sequences are shown in Supplementary Table S1. Five hundred nanograms of carrier and normal human genomic DNA was used as template in a 50-μL (1 × Taq Plus PCR Green) reaction system, programmed to denaturation at 95 °C for 5 min, 35 cycles for amplification (denaturation at 94 °C for 30 s, annealing at 58 °C for 30 s, extension at 72 °C for 30 s), and finally extension at 72 °C for 5 min.
The carrier-specific single PCR product bands were purified using the GelExtraction Kit (Qiagen) and sequenced by bi-directional Sanger sequencing, and the trans-breakpoint sequences were aligned with the human chromosome reference sequences using the Blast online software.
Results
Overall distribution of the 2441 breakpoints
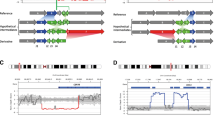
We retrospectively analyzed the 2441 chromosomal rearrangement breakpoints from MicroSeq localization data of 1219 ABCR carriers, including 2364 breakpoint regions in 1182 cases of reciprocal translocations, 68 breakpoint regions in 34 cases of inversions, and 9 breakpoint regions in 3 cases of insertion of translocations. Reciprocal translocation was the major chromosome rearrangement, accounting for 97% of the total (Table 1). Approximately 3–4 million reads with up to 40% coverage were obtained from NGS sequencing. Given that the WGA4 kit is an optimized amplification kit based on the principle of degenerate oligonucleotide-primed polymerase chain amplification (DOP-PCR), the distances between randomly bound sites in the human genome of degenerate oligonucleotide primers are on average approximately 3–5 kb. Thus, it is possible to pinpoint the location of breakpoints for chromosomal rearrangements that were located on specific chromosomes to an average of 3.8 kb. The distribution of breakpoints on chromosomes is shown in Fig. 1a. The karyotype analysis and breakpoint data of 1219 cases are summarized in Supplementary Table S2.
The ABCRs encompass known recurrent rearrangement regions, such as 22q11.21 (chr22:20326658-20327753 bp, 41/1219), 8q24.13 (chr8:125493713-125711433 bp, 7/1219), and 11q11.23 (chr11:116682059-116683162 bp, 37/1219) (Fig. 1b, c, and d). We also found that a 10-Mb region located on 12q24.13-q24.3 was likely to have a reduced amount of balanced chromosome rearrangements (Fig. 1e). Chromosome 9 had a large breakpoint gap region at 9q12 due to the presence of heterochromatin (Fig. 1f). Our analysis revealed that 89.19% (33/37) of the rearrangements that occurred in 11q11.23 were exchanged with 22q11.21, and 80.49% (33/41) of the rearrangements that occurred in the recurrent rearrangement region on 22q11.21 were exchanged with 11q11.23 (Fig. 2). This suggests that chromosomal recurrent rearrangement regions that are abundant in AT repeat palindrome sequences are likely to interchange with one another.
Distribution of breakpoints on chromosomes. a Distribution of breakpoints on chromosomes other than sex chromosomes. b Distribution of breakpoints on chromosome 22 shows a recurrent breakpoint region of 22q11.21. c Distribution of breakpoints on chromosome 8 shows a recurrent breakpoint region of 8q24.13. d Distribution of breakpoints on chromosome 11 shows a recurrent breakpoint region of 11q11.23. e Distribution of breakpoints on chromosome 12 shows a large sparse region of q24.13-q24.3. f Distribution of breakpoints on chromosome 9 shows a large breakpoint gap of 9q12 due to the presence of heterochromatin
Breakpoint-disrupted genes
Of the 2441 chromosomal breakpoints that were analyzed, 105 breakpoints had poor sequence alignment results, and 85 breakpoints were located in genomic regions with complex structures or segmental duplications based on data from T2T-CHM13 and GRCh38, which could not be accurately determined by short-read sequencing. These breakpoints were not suitable for gene disruption analysis. In addition, a total of 149 breakpoints were identified in the heterochromatin region, with the majority (66.44%, 99/149) situated in the centromere. Additionally, 15.44% (23/149) were located in the short arm of the D/G group, 7.38% (11/149) were located in the telomere, and 10.74% (16/149) were located in the secondary constrictions (Fig. 3a). The MicroSeq technique can narrow down the breakpoints to an average area of 3.8 kb in the remaining 2102 (86.11%) breakpoints that were located in the euchromatin region, thereby making it feasible to determine whether the genes are disrupted. Among these breakpoints, 54.14% (1138/2102) were situated between two adjacent genes, which indicates that these breakpoints did not disrupt genes. Moreover, 42.72% (898/2102) of the breakpoint ranges were observed within the same genes, suggesting that these chromosomal breaks had caused gene disruption. Only 3.14% (66/2102) of the breakpoint ranges had one side inside the gene and the other side outside the gene, making it unclear whether gene disruption had occurred (Fig. 3b, Supplementary Fig. S1). Given that the proportion of protein-coding genes in the genome is less than 1.5%, our data imply that chromosomal rearrangements are more likely to occur within genes than in intergenic regions.
Classification of breakpoints location. a Classifications of breakpoints located in specific euchromatin. b Proportion of 2102 breakpoints which can be determine whether the genes are disrupted. c Classifications of diseases associated with the 48 AD genes which were interrupted by breakpoints. Red font indicates that the genes belong to haploinsufficiency genes
To investigate the genetic risks of the ABCRs, we analyzed the 898 breakpoints that interrupted genes. It was discovered that 8.94% (188/2102) of breakpoints interrupted Mendelian disease-related genes, as recorded in the OMIM database (Fig. 3b). Among the 188 breakpoints, 53.19% (100/188) disrupted AR genes, 25.53% (48/188) disrupted AD genes, 11.70% (22/188) disrupted genes of two or more different inheritance patterns, and 9.57% (18/188) disrupted genes with unknown inheritance patterns (Fig. 3b, Supplementary Table S3).
Most AD-disrupted ABCR carriers are phenotypically normal
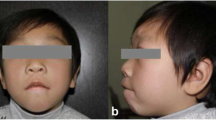
We then conducted a follow-up of 48 carriers with ABCR-caused AD gene interruptions and discovered that only one patient (MD20185) had a partial Marfan syndrome phenotype, as he had a chromosome 15 translocation with the breakpoint located in the region of chr15:48795511-48797005 bp, interrupting the FBN1 gene. Cardiac ultrasound revealed slight tricuspid regurgitation in the patient, with no other significant abnormalities observed. The patient is 1.84 m tall, has been highly myopic since childhood (left 850/right 900), has crowded and uneven mandibular teeth, stands upright with hands that do not drop below the knees, and does not have an arachnodactyly phenotype (finger length is normal). His brother, who has two children, has the same phenotype, as revealed by the family history. His father’s teeth have been lost, so the alignment is unknown. These characteristics overlap with some of the phenotypes of Marfan syndrome, albeit the phenotype is mild.
The remaining 47 carriers appeared to be consistent with normal at the stage where they sought to reproduce. All carriers reported no unusual discomfort and declined further examinations. Among the 48 AD genes disrupted by breakpoints, only 6 genes have HI scores of 3 in ClinGen (https://www.clinicalgenome.org/), which means they have sufficient evidence for haploinsufficiency. In addition, 23 of 48 AD genes had a pLI score > 0.9 according to gnomAD (http://www.gnomad-sg.org/), indicating that these genes are not resistant to protein truncation mutations (Supplementary Table S4). However, no one presented with an anomalous phenotype at the time of presentation except the carrier with interrupted FBN1. We screened 6 genes (ZFPM2, TCF12, PTPN11, CDON, KCNJ6, and KCNQ5) related to early-onset and severe teratogenic diseases to assess the pathogenicity of the gene interruption, including two 2 haploinsufficiencies. We excluded 41 genes related to late-onset or cryptic diseases for which the pathogenicity of translocations was difficult to evaluate (Fig. 3c, Supplementary Table S5). Patients with mutations in the six genes primarily display phenotypes that are associated with considerable anomalies in facial features, stature, limb development, and intellectual capacity. None of these ABCR carriers exhibits abnormal phenotypes in the aspects mentioned above. We compared our rearrangement sites to the potential pathogenic truncated mutation sites reported in the Human Gene Mutation Database (HGMD). Additionally, we investigated PubMed and OMIM to determine the penetrance of diseases. Reports of truncated mutations in the rearrangement positions of ZFPM2 and TCF12 have been documented. The breakpoints of PTPN11 and CDON are positioned after the reported truncated mutation sites. Furthermore, the breakpoints of KCNJ6 and KCNQ5 are both situated before the reported truncated mutation sites (Table 2). Our findings suggest that truncation mutations at arrangement regions in PTPN11, CDON, KCNJ6, and KCNQ5 may not cause any abnormal phenotype in carriers at reproductive age.
It was found that both of the breakpoints that interrupted TCF12 and ZFPM2 were between the positions of potential truncated mutation sites, which means that they have varying degrees of truncated mutations leading to associated diseases. The reason that the ABCR carriers do not present with abnormal phenotypes can be explained by the incomplete penetrance of related diseases (Table 2). Breakpoints that interrupted PTPN11 and CDON were behind potential truncated mutation sites. It is possible that the protein truncated by rearrangements still functions as a partial protein, which could account for the normal phenotypes of carriers; thus, this truncated mutation may have little effect on protein functions. Furthermore, both rearranged sites of KCNJ6 and KCNQ5 were in front of the potential truncated mutations, and both genes are haploinsufficient. Additionally, related diseases have not been reported with incomplete penetrance (Table 2). The breakpoint site of KCNJ6 was located in the early region of the 5′UTR, which suggests the possibility that the expression of this gene can be executed on another chromatin after translocation. The breakpoint site of KCNQ5 was located in introns 1–2. It is possible that the truncated protein will be degraded, and its allele is sufficient for normal physiological functions. The above results suggest that truncation mutations at arrangement regions in PTPN11, CDON, KCNJ6, and KCNQ5 may not cause an abnormal phenotype at reproductive age in carriers.
Mutational signatures underlying mechanisms of rearrangement formation
To investigate the mechanisms leading to balanced chromosomal rearrangements, a total of 100 chromosomal breakpoints carried by 50 of these patients were pinpointed and characterized using long fragment PCR combined with Sanger sequencing (Supplementary Table S6). Of these, 54 (54%, 54/100) rearrangement breakpoints had 1–8 bp of microhomologous sequence at the breakpoints, with a predominance of 2–3-bp microhomology, which were accompanied by base deletions (1–14 bp), insertions (1–21 bp), or duplications (1–8 bp) at both ends or individually, and 30 (30%, 30/100) break junctions were blunt end junctions (Fig. 4a–b). NHEJ is a way to repair double-stranded DNA breaks, as its template-independent direct joining of broken DNA ends is prone to cause structural chromosome variations. MMBIR can repair SSB during DNA replication through a break-induced replication mechanism based on microhomologous sequences. This result shows that there is no large segment homology at the breakpoints of these chromosomes and that there is only microhomology with minor imbalance features, suggesting that the possible mechanism of rearrangement is related to the combination of NHEJ and MMBIR.
Molecular characterization of 100 precise breakpoints. a Schematic representation of types of chromosomal equilibrium rearrangements. i Flat end joining; ii schematic representation of end joining containing only microhomology; iii schematic representation of end joining containing microhomology accompanied by insertion or deletion. b Bar chart of classification of 100 precise breakpoint characters
Discussion
Carrier screening for healthy populations, in combination with prenatal and preimplantation diagnosis to avoid birth defects, is an effective approach to reduce the incidence of Mendelian genetic disorders [33,34,35,36,37]. Our breakpoint data analysis revealed that 42.72% (898/2102) of breakpoints explicitly interrupted genes, and this percentage is significantly higher than the 1.5% of gene sequences in the human genome, suggesting that chromosomal rearrangements are more likely to occur in gene sequences. In this study, 8.22% (93/1132) of phenotype-free ABCR carriers were found to have an interruption in the AR gene (Table 3). All individuals had a normal phenotype, likely due to a fully functional allele on the normal homologous chromosome. If the carrier’s normal spouse carries a heterozygous pathogenic mutation in the broken gene, the offspring will be at high risk of suffering from the related genetic diseases since they may inherit both the broken genes from the carrier and the pathogenic mutations from his or her spouse [38]. Therefore, it is recommended that the spouse of the carrier be screened for the corresponding gene, allowing for an accurate assessment of the genetic risk of the broken gene in the event of the carrier couple having offspring and the selection of an appropriate approach for childbirth.
In this study, with the exception of one patient who carried an interrupted FBN1 gene, the rest of the patients with chromosomal rearrangements were tentatively not found to have abnormal phenotypes at the stage where they were seeking reproduction. These findings are distinct from the results of other research showing that at least 33.9% of carriers of balanced chromosomal abnormalities (BCAs) presented with congenital anomalies that could be caused by genes interrupted by breakpoints overlapping with known microdeletion/microduplication syndromes or associated with dominant developmental disorders [39]. We hypothesize that there may be a few reasons that a patient may not have a phenotype. For example, the truncated protein is degraded and the protein is not dose-sensitive or the disease caused by the truncated mutation has a late onset or incomplete penetrance. [40, 41]. Additionally, two ABCR carriers had rearrangements disrupting the ZFPM2 and TCF12 genes in truncated regions reported by others, yet they did not demonstrate any symptoms, suggesting that the related diseases may not be fully penetrant. Furthermore, two asymptomatic ABCR carriers had chromosome rearrangements with in breaks in PTPN11 and CDON subsequent to the truncation sites repoted by others. It is possible that the lack of phenotype in these ABCR carriers is due to the rearranged truncated proteins retaining some of the capabilities of the normal proteins. Additionally, the breakpoints of KCNJ6 and KCNQ5 in two ABCR carriers are located prior to the reported truncation sites, and neither gene is indicated to be haploinsufficient. No incomplete penetrance has been reported for the related diseases. The rearrangement of KCNJ6 occurs in the 5′UTR, thus KCNJ6 gene expression is still possible despite the rearrangement. The rearrangement of KCNQ5 is located in the intron 1–2 region, and the truncated KCNQ5 protein may be degraded. However, the other normal KCNQ5 allele may be sufficient to fulfill its normal functions. Our findings suggest that most truncation mutations in rearrangement regions in ABCR carriers may be benign.
Topologically associated domains (TADs) on chromosomes can facilitate the interaction between enhancers and promoters to regulate gene expression, and chromosomal rearrangements that disrupt the TAD structure can lead to abnormal gene expression [39, 42, 43]. For example, the expression of some developmental genes was dysregulated by TAD structure, which was interrupted by breakpoints in 7.3% of subjects with congenital malformations in the study of Redin et al. [39]. It has been shown that some BCA carriers present a fertility disorder phenotype without disrupting reproduction-related genes, possibly because the breakpoints interrupt the TAD structure and thus aberrantly regulate the expression of genes related to reproduction [10]. For patients with ABCRs with infertility, no reproduction-related genes were interrupted in our research, and we did not exclude the possibility that breakpoints may disrupt the TAD structure of genes associated with reproduction. However, there are some cases with altered gene expression that cannot be explained by position effects [44]. Similarly, ABCR carriers with chromosomal rearrangement-disrupted AD genes in other studies did not present any abnormal phenotypes [3]. Further investigation is necessary to determine whether the genes interrupted by chromosome rearrangements have a different physiological significance or expression patterns than other types of mutations. Our data may provide an informative value for clinical genetic counseling regarding the risk of diseases associated with genes affected by chromosome rearrangement.
For ABCR carriers with an interrupted AD gene, it is necessary to determine whether the gene-related diseases are late-onset. If the carrier does not present any phenotype and the corresponding diseases are not late-onset, the mutation at the region may be considered a nonpathogenic mutation. In cases where the carrier has a breakpoint-associated genetic disorder phenotype, prenatal diagnosis or preimplantation genetic testing (PGT) should be suggested to ensure the birth of a child free from chromosome rearrangements. Upon learning that his chromosome rearrangement interrupted the FBN1 gene, resulting in his mild Marfan syndrome phenotypes, case MD20185 chose PGT to have an ABCR-free baby. After the first cycle of PGT treatment, the patient obtained three paternally balanced translocation carrier embryos but no normal embryos. Subsequently, after receiving genetic counseling, the couple decided to initiate a new PGT cycle and chose to transfer only normal embryos.
Chromosomal rearrangement breakpoints contain essential details regarding structural variation. If the breakpoints of chromosomal rearrangements were usually located in regions of nonallelic homologous sequences, this usually suggested that the rearrangements were likely caused by NAHR between two low-copy repetitive sequences [26, 45]. Given the limitation of MicroSeq that could not confidently map the breakpoints which were located in segmental duplications, the NAHR was not been excluded in this study. Our research revealed that 54% of the breakpoints featured microhomology with minor imbalances. These findings indicate that NHEJ or combination with MMBIR are possible mechanisms of rearrangement, as previously suggested [3, 5, 31, 46, 47]. One of the limitations of MicroSeq is that it can only detect breakpoints of known obvious chromosome rearrangements and construct single nucleotide polymorphism (SNP) haplotypes linked to breakpoint regions. However, it cannot identify unknown chromosomal structure variants on a genome-wide scale in a blind way. Moreover, MicroSeq was unable to directly identify the orientation of complex or cryptic imbalances that may exist in breakpoints. Combination with controlled polymerizations by adapter-ligation (CP-AL) techniques is required to assemble a complete spectrum of chromosomal structural variation [48, 49]. Future technologies that directly identify all precise breakpoints at the base level on a genome-wide scale need to be developed, and these technologies that can identify breakpoint-linked SNPs will have more general clinical applications.
Despite our attempts to contact asymptomatic ABCR carriers after the gene interruption was identified, the majority of patients declined further examination, which prevented us from gathering any potential concealed phenotypes. Consequently, the clinical significance of most late-onset or mild AD gene interruptions has not been accurately assessed; we will continue to monitor the subsequent health status of patients with chromosomal rearrangements in an attempt to elucidate the pathogenicity of chromosomal rearrangements. In addition, this study did not investigate the positional effects of chromosomal rearrangements. Moreover, for those ABCR carriers whose haploinsufficient genes or AD genes have been disrupted by chromosomal rearrangements, we were unable to obtain samples to determine if the expression levels of related genes and downstream gene functions had been altered. Whether the AD OMIM genes have a new pathogenic mechanism or whether the genetic calling of chromosomal rearrangement is correct is one of the limitations of this study as we cannot give an accurate answer without the study of the exact mechanism. Nevertheless, this study offers a gene-level breakpoint map of chromosomal rearrangements in a cohort of ABCR carriers, thereby providing reference data to assess the genetic risks associated with the chromosomal rearrangement breakpoints in this population. Moreover, it suggests that NEHR and MMBIR may be the primary mechanisms underlying the formation of apparently balanced chromosomal rearrangements.
References
Jacobs PA, et al. Estimates of the frequency of chromosome abnormalities detectable in unselected newborns using moderate levels of banding. J Med Genet. 1992;29:103–8. https://doi.org/10.1136/jmg.29.2.103.
Giardino D, et al. De novo balanced chromosome rearrangements in prenatal diagnosis. Prenat Diagn. 2009;29:257–65. https://doi.org/10.1002/pd.2215.
Halgren C, et al. Risks and recommendations in prenatally detected de novo balanced chromosomal rearrangements from assessment of long-term outcomes. Am J Hum Genet. 2018;102:1090–103. https://doi.org/10.1016/j.ajhg.2018.04.005.
David D, et al. Comprehensive clinically oriented workflow for nucleotide level resolution and interpretation in prenatal diagnosis of de novo apparently balanced chromosomal translocations in their genomic landscape. Hum Genet. 2020;139:531–43. https://doi.org/10.1007/s00439-020-02121-x.
Gijsbers AC, et al. Whole genome paired-end sequencing elucidates functional and phenotypic consequences of balanced chromosomal rearrangement in patients with developmental disorders. Eur J Med Genet. 2019;56:526–35. https://doi.org/10.1136/jmedgenet-2018-105778.
Schluth-Bolard C, et al. Cryptic genomic imbalances in de novo and inherited apparently balanced chromosomal rearrangements: array CGH study of 47 unrelated cases. Eur J Med Genet. 2009;52(5):291–6. https://doi.org/10.1016/j.ejmg.2009.05.011.
Feenstra I, et al. Balanced into array: genome-wide array analysis in 54 patients with an apparently balanced de novo chromosome rearrangement and a meta-analysis. Eur J Hum Genet. 2011;19:1152–60. https://doi.org/10.1038/ejhg.2011.120.
Dong Z, et al. Genome sequencing explores complexity of chromosomal abnormalities in recurrent miscarriage. Am J Hum Genet. 2019;105:1102–11. https://doi.org/10.1016/j.ajhg.2019.10.003.
Miao ZY, et al. Cytogenetic analysis of 2959 couples with spontaneous abortion and detailed analysis of rare karyotypes. J Genet. 2022;101:10.
Chau MHK, et al. Investigation of the genetic etiology in male infertility with apparently balanced chromosomal structural rearrangements by genome sequencing. Asian J Androl. 2022;24(3):248–54. https://doi.org/10.4103/aja2021106.
Carvalho CM, Lupski JR. Mechanisms underlying structural variant formation in genomic disorders. Nat Rev Genet. 2016;17:224–38. https://doi.org/10.1038/nrg.2015.25.
Lieber MR. The mechanism of human nonhomologous DNA end joining. J Biol Chem. 2008;283:1–5. https://doi.org/10.1074/jbc.R700039200.
Mitelman F, et al. The impact of translocations and gene fusions on cancer causation. Nat Rev Cancer. 2007;7:233–45. https://doi.org/10.1038/nrc2091.
Mitelman F, et al. Prevalence estimates of recurrent balanced cytogenetic aberrations and gene fusions in unselected patients with neoplastic disorders. Genes Chromosomes Cancer. 2005;43:350–66. https://doi.org/10.1002/gcc.20212.
Schnause AC, et al. Marfan syndrome caused by disruption of the FBN1 gene due to a reciprocal chromosome translocation. Genes (Basel). 2021;12:1836. https://doi.org/10.3390/genes12111836.
Siffroi JP, et al. Assisted reproductive technology and complex chromosomal rearrangements: the limits of ICSI. Mol Hum Reprod. 1997;3:847–51. https://doi.org/10.1093/molehr/3.10.847.
Miny P, Schloo R. Ist Sterilität eine Erblast? [Is sterility a genetic burden?]. Ther Umsch. 1999;56:265–70. https://doi.org/10.1024/0040-5930.56.5.265.
Neri G, et al. Reproductive risks for translocation carriers: cytogenetic study and analysis of pregnancy outcome in 58 families. Am J Med Genet. 1983;16:535–61. https://doi.org/10.1002/ajmg.1320160412.
Park JM, et al. An autosomal dominant ERLIN2 mutation leads to a pure HSP phenotype distinct from the autosomal recessive ERLIN2 mutations (SPG18). Sci Rep. 2020;10:3295. https://doi.org/10.1038/s41598-020-60374-y.
Cheon CK, et al. Autosomal dominant transmission of complicated hereditary spastic paraplegia due to a dominant negative mutation of KIF1A, SPG30 gene. Sci Rep. 2017;7:12527. https://doi.org/10.1038/s41598-017-12999-9.
Couch FJ, et al. Inherited mutations in 17 breast cancer susceptibility genes among a large triple-negative breast cancer cohort unselected for family history of breast cancer. J Clin Oncol. 2015;33:304–11. https://doi.org/10.1200/JCO.2014.57.1414.
Couch FJ, et al. Associations between cancer predisposition testing panel genes and breast cancer. JAMA Oncol Sep. 2017;3(9):1190–6. https://doi.org/10.1001/jamaoncol.2017.0424.
Kilpivaara O, et al. CHEK2 variant I157T may be associated with increased breast cancer risk. Int J Cancer. 2004;111:43–547. https://doi.org/10.1002/ijc.20299.
Lieber MR, et al. Flexibility in the order of action and in the enzymology of the nuclease, polymerases, and ligase of vertebrate non-homologous DNA end joining: relevance to cancer, aging, and the immune system. Cell Res. 2008;18:125–33. https://doi.org/10.1038/cr.2007.108.
Lupski JR. Genomic disorders: structural features of the genome can lead to DNA rearrangements and human disease traits. Trends Genet. 1998;14:417–22. https://doi.org/10.1016/s0168-9525(98)01555-8.
Stankiewicz P, Lupski JR. Genome architecture, rearrangements and genomic disorders. Trends Genet. 2002;18:74–82. https://doi.org/10.1016/s0168-9525(02)02592-1.
Lee JA, et al. A DNA replication mechanism for generating nonrecurrent rearrangements associated with genomic disorders. Cell. 2007;131(7):1235–47. https://doi.org/10.1016/j.cell.2007.11.037.
Hastings PJ, et al. A microhomology-mediated break-induced replication model for the origin of human copy number variation. PLoS Genet. 2009;5(1):e1000327. https://doi.org/10.1371/journal.pgen.1000327.
Burssed B, et al. Mechanisms of structural chromosomal rearrangement formation. Mol Cytogenet. 2009;15:23. https://doi.org/10.1186/s13039-022-00600-6.
Symington LS, Gautier J. Double-strand break end resection and repair pathway choice. Annu Rev Genet. 2011;45:247–71. https://doi.org/10.1146/annurev-genet-110410-132435.
Nilsson D, et al. Whole-genome sequencing of cytogenetically balanced chromosome translocations identifies potentially pathological gene disruptions and highlights the importance of microhomology in the mechanism of formation. Hum Mutat. 2017;38:180–92. https://doi.org/10.1002/humu.23146.
Hu L, et al. Reciprocal translocation carrier diagnosis in preimplantation human embryos. EBioMedicine. 2016;14:139–47. https://doi.org/10.1016/j.ebiom.2016.11.007.
Levy B, Stosic M. Traditional prenatal diagnosis: past to present. Methods Mol Biol. 2019;1885:3–22. https://doi.org/10.1007/978-1-4939-8889-1_1.
D'Alton ME, DeCherney AH. Prenatal diagnosis. N Engl J Med. 1993;328:114–20. https://doi.org/10.1056/NEJM199301143280208.
Sullivan-Pyke C, Dokras A. Preimplantation genetic screening and preimplantation genetic diagnosis. Obstet Gynecol Clin North Am. 2018;45:113–25. https://doi.org/10.1016/j.ogc.2017.10.009.
Simpson JL, et al. Overview of preimplantation genetic diagnosis (PGD): historical perspective and future direction. Methods Mol Biol. 2019;1885:23–43. https://doi.org/10.1007/978-1-4939-8889-1_2.
Tan YQ, et al. Single-nucleotide polymorphism microarray-based preimplantation genetic diagnosis is likely to improve the clinical outcome for translocation carriers. Hum Reprod. 2013;28:2581–92. https://doi.org/10.1093/humrep/det271.
Trunca C, et al. Reproductive risk estimation calculator for balanced translocation carriers. Curr Protoc. 2022;2:e633. https://doi.org/10.1002/cpz1.633.
Redin C, et al. The genomic landscape of balanced cytogenetic abnormalities associated with human congenital anomalies. Nat Genet. 2017;49(1):36–45. https://doi.org/10.1038/ng.3720.
Wolff M, et al. Phenotypic spectrum and genetics of SCN2A-related disorders, treatment options, and outcomes in epilepsy and beyond. Epilepsia. 2019;60(Suppl 3):S59–67. https://doi.org/10.1111/epi.14935.
Kayvanpour E, et al. Genotype-phenotype associations in dilated cardiomyopathy: meta-analysis on more than 8000 individuals. Clin Res Cardiol. 2017;106:127–39. https://doi.org/10.1007/s00392-016-1033-6.
Ghavi-Helm Y. Functional consequences of chromosomal rearrangements on gene expression: not so deleterious after all? J Mol Biol. 2022;432:665–75. https://doi.org/10.1016/j.jmb.2019.09.010.
Lupiáñez DG, et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell. 2015;161:1012–25. https://doi.org/10.1016/j.cell.2015.04.004.
Dong Z, et al. Identification of balanced chromosomal rearrangements previously unknown among participants in the 1000 Genomes Project: implications for interpretation of structural variation in genomes and the future of clinical cytogenetics. Genet Med. 2018;20:697–707. https://doi.org/10.1038/gim.2017.170.
Shaw CJ, Lupski JR. Implications of human genome architecture for rearrangement-based disorders: the genomic basis of disease. Hum Mol Genet. 2004;13:R57–64. https://doi.org/10.1093/hmg/ddh073.
Chiang C, et al. Complex reorganization and predominant non-homologous repair following chromosomal breakage in karyotypically balanced germline rearrangements and transgenic integration. Nat Genet. 2012;44:390–S1. https://doi.org/10.1038/ng.2202.
Conrad DF, et al. Mutation spectrum revealed by breakpoint sequencing of human germline CNVs. Nat Genet. 2010;42:385–91. https://doi.org/10.1038/ng.564.
Dong Z, et al. Development of coupling controlled polymerizations by adapter-ligation in mate-pair sequencing for detection of various genomic variants in one single assay. DNA research: an international journal for rapid publication of reports on genes and genomes. DNA Res. 2019;26(4):313–25. https://doi.org/10.1093/dnares/dsz011.
Dong Z, et al. Deciphering the complexity of simple chromosomal insertions by genome sequencing. Hum Genet. 2021;140(2):361–80. https://doi.org/10.1007/s00439-020-02210-x.
Acknowledgements
We thank the patients and family members for their participation. Thanks to the National Natural Science Foundation of China for the strong support to this project.
Funding
This work was supported by grants from the National Natural Science Foundation of China (81873478).
Author information
Authors and Affiliations
Contributions
Conceptualization: DC, YT, LH, GL; methodology: DC, GL, YT, LH; formal analysis and investigation: DC, YX, KL, ML; writing—original draft preparation: DC, YX; writing—review and editing: YX, LH; visualization: YX; funding acquisition: LH; resources: YT, GL, LH; supervision: LH.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was approved by Reproductive and Genetic Hospital of CITIC Xiangya Ethics Committee and informed consent was obtained from the patients. The ethical approval number is LL-SC-2016-002.
Consent for publication
All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiao, Y., Cheng, D., Luo, K. et al. Evaluation of genetic risk of apparently balanced chromosomal rearrangement carriers by breakpoint characterization. J Assist Reprod Genet 41, 147–159 (2024). https://doi.org/10.1007/s10815-023-02986-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-023-02986-7