Abstract
Chromosomal translocations, rearrangements involving the exchange of segments between chromosomes, were documented in humans in 1959. The first accurately reported clinical phenotype resulting from a translocation was that of Down syndrome. In a small percentage of Down syndrome cases, an extra 21q is provided by a Robertsonian translocation chromosome, either occurring de novo or inherited from a phenotypically normal parent with the translocation chromosome and a balanced genome of 45 chromosomes. Balanced translocations, including both Robertsonian and reciprocal translocations, are typically benign, but meiosis in germ cells with balanced translocations may result in meiotic arrest and subsequent infertility, or in unbalanced gametes, with attendant risks of miscarriage and unbalanced progeny. Most reciprocal translocations are unique. A few to several percent of translocations disrupt haploinsufficient genes or their regulatory regions and result in clinical phenotypes. Balanced translocations from patients with clinical phenotypes have been valuable in mapping disease genes and in illuminating cis-regulatory regions. Mapping of discordant mate pairs from long-insert, low-pass genome sequencing now permits efficient and cost-effective discovery and nucleotide-level resolution of rearrangement breakpoints, information that is absolutely necessary for interpreting the etiology of clinical phenotypes in patients with rearrangements. Pathogenic translocations and other balanced chromosomal rearrangements constitute a class of typically highly penetrant mutation that is cryptic to both clinical microarray and exome sequencing. A significant proportion of rearrangements include additional complexity that is not visible by conventional karyotype analysis. Some proportion of patients with negative findings on exome/genome sequencing and clinical microarray will be found to have etiologic balanced rearrangements only discoverable by genome sequencing with analysis pipelines optimized to recover rearrangement breakpoints.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Cytogenetics
- Karyotype
- Balanced translocation
- Breakpoint
- Congenital anomaly
- Gene mapping
- Mate-pair sequencing
1.1 Introduction and Background
Chromosomal translocations encompass a diverse set of rearrangements involving the exchange of segments between chromosomes, and are common in humans. Balanced translocations, those without accompanying copy number variation, usually have no phenotypic consequence. Estimates vary, but about one in every 300–500 individuals has a balanced reciprocal translocation, and about one per 1000 has a balanced Robertsonian translocation (the joining of complete long arms of two acrocentric chromosomes in β-satellite sequences in the short arms) ([16, 24, 27, 28, 36, 51, 57, 60, 89]). From a multicenter study of 377,357 amniocenteses, the incidence of a de novo reciprocal translocation has been estimated at about 1 per 2000, and 1 per 9000 for a Robertsonian translocation [92]. Warburton also estimated the risk of congenital abnormality associated with a balanced reciprocal translocation to be about 6% (an approximate 2- to 3-fold increase over the general population risk), and that from balanced Robertsonian translocation to be negligible. Congenital anomaly or other clinical phenotype in an individual with a balanced translocation may result from any of a number of possible effects of a translocation, including direct gene disruption, creation of a fusion gene, dysregulation of a gene separated from its normal extragenic cis-regulatory elements, or dysregulation of a gene placed in an environnment of altered chromatin modification. Balanced translocations constitute an important class of mutation that are etiologic in hundreds of Mendelian diseases, and many cancers. When ascertained from individuals with clinical phenotypes, translocations have been invaluable biological tools in mapping disease loci and cis-regulatory regions of disease genes. Because missegregation of balanced translocations in meiosis may result in meiotic arrest or unbalanced gametes, individuals with balanced translocations have higher risks of subfertility and infertility, miscarriage, and genomic imbalance in their offspring.
1.2 Early Observations of Translocations
The field of human cytogenetics was very young in 1959 and 1960, when the first human translocations were reported. The correct diploid human chromosome number had only been established by Tjio and Levan in 1956 [86], shortly afterwards confirmed by Ford and Hamerton [17]. This foundational achievement depended upon a number of technical advances, including improved methods of tissue culture, use of colchicine for inducing mitotic arrest [18], and most critically, incubation of cells in hypotonic solution for better chromosome spreading [29, 30]. Still, only slow progress was made in the late 1950s in identifying and characterizing human chromosomal abnormalities. Until the advent of quinacrine mustard banding [7] and Giemsa banding [76, 81] in the 1970s, chromosomes could be reliably ordered by size and centromere position only into seven groups (when this system was used, the groups were denoted A-G). Distinguishing chromosomes within groups was difficult; many specific chromosome assignments published in those years were speculative, and some were almost certainly wrong. Before the development and widespread adoption of techniques for generating sufficient numbers of mitotic cells in cultures of peripheral leukocytes [32, 53, 59], bone marrow or testis, neither trivial to ask of patients or their family members, were the tissues generally sought for karyotyping. Thus, early karyotyping was usually limited to patients, so it was not possible to assess the segregation of a chromosomal rearrangement with a phenotype in a pedigree. In spite of these difficulties, trisomy 21 was convincingly established as the chromosomal basis of Down syndrome [OMIM #190685] in 1959, on the collective evidence of 30 cases (in order of publication: [43, 35, 19, 4]), and identification of other aneuploidy syndromes quickly followed.
Lejeune’s group was also the first to report a human translocation. They found, in a patient with intellectual and speech disabilities and “polydysspondylie” (spondylocostal dysostosis, a skeletal dysplasia with six known loci, most demonstrating recessive inheritance [e.g. OMIM #277300]), a karyotype of 45 chromosomes with what is now known to be a Robertsonian translocation chromosome, which they interpreted as composed of 22q and either 14q or 15q ([88], described in [87]). Common and easily identifiable, particularly in a balanced karyotype of 45 chromosomes, it is not surprising that the first reported human translocation was a Robertsonian. However, the phenotype in this case was almost certainly not related to the translocation; we now know that most individuals with a Robertsonian translocation and a balanced karyotype are phenotypically normal, and no clinical phenotypes have yet to be associated convincingly with any balanced Robertsonian translocation, with the important exceptions of increased risk of miscarriage and reduced fertility.
The second reported human translocation was also a Robertsonian, interpreted as comprising chromosomes 21 and 14, identified in an unbalanced karyotype of a girl with Down syndrome and 46 chromosomes, with the translocation chromosome providing the extra copy of 21q etiologic for Down syndrome [67]. Because the maternal age effect in Down syndrome births [64] was by then well known, Polani et al. chose the children of young mothers to investigate, hoping to increase the chances of uncovering additional karyotypically-visible etiologies for Down syndrome. From the two cases they successfully karyotyped, they were lucky to have ascertained one case of translocation Down syndrome; a translocation chromosome is present in only about 3–4% of Down syndrome karyotypes, even among young mothers. Because the phenotypes of trisomy 21 Down syndrome and translocation Down syndrome are indistinguishable, Polani et al. were able to make a strong and correct case for the participation of chromosome 21 in this rearrangement.
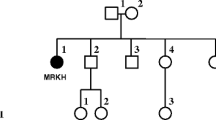
Penrose et al. [65] were the first to demonstrate segregation of a balanced translocation in a family. A grandmother, mother, and daughter, all phenotypically normal, each had 45 chromosomes with a translocation chromosome interpreted as rob(15;21) [although it may have been the much more common rob(14;21)] (Fig. 1.1). The mother also had two children with translocation Down syndrome, and had reported two miscarriages. Carter et al. [6] also reported transmission of rob(15;21), in a three-generation pedigree where the mothers of two first cousins with Down syndrome had each inherited the translocation chromosome from their mother. Other Robertsonian translocations besides those associated with chromosome 21 were also identified around the same time. For example, Lejeune et al. [44] found, in a man with a 46, XXY karyotype and Klinefelter syndrome, a translocation between a D group chromosome (chr13–15) and chr22. Transmission of a rob(13;15) was described in a pedigree including 10 individuals with balanced karyotypes and the translocated chromosome; among those with the translocation were eight phenotypically normal individuals. Although the primary amenorrhea in the proband was unlikely to have been related to the translocation, azoospermia in Robertsonian translocation carriers is very common and likely accounted for that reported phenotype in one of the males in the pedigree [91].
The first chromosomal rearrangements to be identified in the early days of cytogenetics were Robertsonian translocations. These photomicrographs published in 1960 show a balanced karyotype of 45 chromosomes with a Robertsonian translocation involving chromosome 21, derived from skin cells of a phenotypically normal woman who had two children with translocation Down syndrome. This was the first publication to document inheritance of a translocation, in a three-generation family. (Reproduced from Ref. [65])
It was recognized early on that translocations between chromosomes would only be detectable if the lengths of the exchanged chromosomal segments were sufficiently different, and large enough, to change the length of the translocated chromosome arms substantially, or to result in an appreciable change in the location of the centromere. In 1962, Edwards et al. [14] reported the first two such cases; the first, a translocation between chromosomes 4 and 9, and the second, between chromosomes 1 and 6. In each case, the balanced reciprocal translocation in a phenotypically normal parent was ascertained through investigation of genomically unbalanced offspring with mental retardation and congenital anomalies. Schmid [74] ascertained a balanced translocation carrier solely on the basis of a history of miscarriage. Among 10 couples who had experienced miscarriage and had one or more phenotypically normal children, one translocation carrier (a male) with an apparently balanced karyotype of 46 chromosomes was identified. The translocation involved one chromosome 21 or 22, and was non-Robertsonian, but the sizes of the exchanged segments were too small to reveal the identity of the larger translocation partner.
1.3 Clinical Relevance of Translocations
1.3.1 Balanced Translocation Carriers Have Higher Risks of Infertility, Miscarriage, and Unbalanced Progeny
Most reciprocal translocations arise on paternal chromosomes in spermatogenesis, and most Robertsonian translocations arise on maternal chromosomes in oogenesis [1, 63, 84]. Thomas et al. [84] detected a significant paternal age effect in the de novo occurrence of reciprocal translocations. Most individuals with balanced translocations are phenotypically normal, and balanced translocations may be transmitted in families through many generations. This was observed in the earliest pedigrees of translocation Down syndrome, and it is still the case that many balanced chromosomal rearrangements are ascertained in couples with an unbalanced fetus or child. In meiosis when a balanced translocation is present, the translocated chromosomes associate with their homologs as a trivalent (for Robertsonians) or tetravalent (for reciprocal translocations), and segregation results in normal and balanced, or unbalanced, gametes. Men with azoospermia or severe oligospermia (no viable sperm or low sperm count) have an incidence of balanced translocations that is many times higher than that in the general male population [8, 90, 96], evidence that checkpoints in male gametogenesis lead to apoptosis of unbalanced gametes. The proportion of normal, balanced, and unbalanced gametes produced by men with balanced translocations varies from practically none to nearly all unbalanced gametes [54, 66], with Robertsonian translocations typically yielding the lowest proportion of unbalanced gametes, and reciprocal translocations the highest. As most reciprocal translocations are unique, it is difficult to assess the reproductive risk of unbalanced progeny for couples in whom one partner is known to have a translocation [96], though Boué and Gallano [5], from assessing pregnancy outcomes in more than 1200 couples where one partner had a translocation, estimated the overall risk of unbalanced progeny to be 10% for a term pregnancy, and 11.5% for a fetus. A balanced translocation in one parent is significantly associated with miscarriage; parental balanced translocation was identified in ~3–4% of couples with recurrent miscarriage [62, 80]. In translocation Down syndrome, about two thirds of cases arise de novo, with about half due to rob(14;21) and half due to rea(21;21) [55], although most or all rea(21;21) are isochromosomes and not true Robertsonian translocations [77]. In inherited translocation Down syndrome, the translocation chromosome is more frequently inherited from the mother, indicating that checkpoints in oogenesis are not as stringent as those in spermatogenesis. Recurrence risk for Down syndrome from inherited Robertsonian translocations is believed to be 10–15% if the mother has the translocation, and less than 1% if the father has the translocation.
Only a few reciprocal translocations have been found to be recurrent. Emanuel syndrome [OMIM #609029] is characterized by multiple congenital anomalies and an unbalanced karyotype with a supernumerary der(22) inherited from an unaffected parent with a t(11;22)(q23;q11.2) [20, 94]. Breakpoints in palindromic AT-rich repeat (PATRR) sequences on chromosomes 11 and 22 suggest that palindrome-mediated formation of secondary structure promotes double-strand breakage and resulting translocation [37]. PATRRs have been recognized on chromosomes 1, 4, 8, 11, 17 and 22, and rare recurrent translocations between PATRRs on chromosomes 8 and 22 [that may result in progeny with supernumerary der(22)t(8;22) syndrome, OMIM #613700] have been reported [78], as well as rare translocations between PATRRs on chromosomes 17 and 22, in individuals with neurofibromatosis type I [NF1, OMIM #162200] [37].
1.3.2 Some Somatic Translocations Initiate Transformation of Cancer Cells
Although interest in cancer had driven much early progress in cytogenetic techniques, the complex, abundant, and variable chromosomal aberrations observed in many cancers were difficult to interpret, and also led investigators to believe that chromosomal rearrangements might all be secondary to the events that initiate tumorigenesis. Two early successes in deciphering the molecular biology of oncogenic translocations were groundbreaking in understanding the importance of chromosomal rearrangements in driving tumorigenesis.
Although not initially recognized as such, the first translocation associated to a specific cancer was the Philadelphia chromosome (Ph), uniquely associated with chronic myelogenous leukemia [CML, OMIM #608232] [59]. Its identity as a der(22) involved in a translocation with chr9 was only uncovered 13 years later, once banding techniques had been developed [73]. In 1982, the ABL oncogene [OMIM *189980], known to map to chr9, was shown to be translocated to the Ph chromosome 22 [10]. Fine-mapping of ABL [25] and the chr22 breakpoint cluster region [23] led to identification of an ABL-BCR fusion gene that results from translocation [79], and the fusion gene’s oncogenic activity in CML was demonstrated [40]. More than 90% of CML cases have a t(9;22), and presence of the Ph chromosome aids in diagnosis; cryptic rearrangements are likely responsible for the remaining 10%.
Three translocations associated with Burkitt’s lymphoma [OMIM #113970] and involving 8q24 were characterized in the late 1970s, with the causative gene rearrangement [83] identified prior to that of the Ph chromosome in CML. The common translocation, t(8;14)(q24;q32), places the intact coding exons of MYC [OMIM *190080], a gene encoding a cell growth and cell cycle transcription factor, close to the enhancer of the immunoglobulin heavy chain locus [IgH locus, see OMIM *147100], driving constitutive expression in B-lymphocytes and conferring oncogenicity. The t(2;8) and t(8;22) translocations place MYC in proximity to enhancers at the IgK or IgL loci, respectively, with similar effect. The pathogenicity of the CML and Burkitt’s translocations is thus explained by two distinct, important, and generalizable models: in CML, the formation of a fusion gene results in pathogenic gain of function of a novel chimeric protein; in Burkitt’s lymphoma, alteration of the regulatory environment of a gene drives its misexpression, resulting in pathogenic gain of function of the normal protein.
1.3.3 Balanced Reciprocal Translocations Are Etiologic for Many Clinical Phenotypes
The earliest pedigrees with translocations were those of translocation Down syndrome, where trisomy for 21q was clearly implicated in the clinical phenotype, and parents and other family members with the translocated chromosome and balanced karyotypes were phenotypically normal. Early pedigrees of reciprocal translocations likewise showed phenotypically normal parents with apparently balanced translocations whose children with clinical phenotypes had unbalanced karyotypes. These pedigrees, though perhaps surprising at first, established the paradigm that rearrangement without apparent loss or gain of chromosomal material had no consequence other than that of the contribution of unbalanced gametes to risks of infertility, miscarriage, and unbalanced offspring. Challenging that paradigm, Jacobs [34] presented epidemiological evidence suggesting that de novo balanced rearrangements (translocations and inversions) were overrepresented among mentally retarded individuals compared to consecutive or random newborns and individuals ascertained for unspecified reasons. Funderburk et al. [21] found balanced rearrangements to be significantly overrepresented among mentally retarded individuals when compared to children of normal intelligence with psychiatric indications. Other studies of outcomes where balanced rearrangements were ascertained from surveys of consecutive newborns [50, 56, 85] were underpowered, failing to ascertain large enough numbers of subjects with balanced rearrangements to interpret correctly an effect that we now know to apply to only a few to several percent of individuals with balanced, non-Robertsonian, rearrangements [92]; Warburton’s robust estimate of the risk of congenital anomaly associated with such rearrangement required the ascertainment of outcomes of 377,357 amniocenteses from multiple clinical centers.
A high burden of proof is required for assigning etiology of phenotype to a balanced reciprocal translocation, given that most are both unique and without phenotypic consequence. Nonetheless, a diagnosis can be made when direct disruption of a known disease gene can be documented and correlated to a specific phenotype, as abundant case reports attest. Individuals with balanced translocations and clinical phenotypes have been useful in mapping a number of Mendelian disease loci, particularly those with severe phenotypes that usually occur de novo, precluding linkage analysis. For instance, the locus for neurofibromatosis type 1 (NF1, OMIM #162200) was identified on the basis of two balanced reciprocal translocations, both with breakpoints in 17q11.2 [42, 75]. Similarly, various rearrangements were reported in patients with Sotos syndrome [OMIM #117550], but two translocations with breakpoints in 5q35 [33, 52] directed investigation to that region; NSD1 [OMIM *606681] was cloned, shown to be disrupted in the translocation of the tested patient, and point mutations and genomic deletions of NSD1, now known to explain about 90% of Sotos syndrome cases, were found in the majority of a cohort of patients [41].
Translocations have also been productive in identifying new candidate genes underlying common clinical phenotypes that may arise from dysfunction of any number of genes, as in autism spectrum disorder, cardiac defects, and orofacial clefting [26, 71]. Two cases described by Kim et al. [38] contributed to identifying a role for neurexin 1 [NRXN1, OMIM *600565] in autism spectrum disorder; NRXN1 has since also been strongly implicated in other neurodevelopmental disorders, including schizophrenia and intellectual disability. Quintero-Rivera et al. [69] marshalled substantial evidence that matrin 3 [MATR3, OMIM *164015], previously associated with amyotrophic lateral sclerosis [ALS21; OMIM #606070], is etiologic for cardiac left ventricle outflow tract (LVOT) defects in a child with a t(1;5) disrupting MATR3 on 5q. Interestingly, AHDC1 [AT-hook DNA-binding motif-containing protein 1, OMIM *615790] on 1q was also disrupted by this translocation, and several aspects of the child’s phenotype, including intellectual disability, facial dysmorphisms, and respiratory and sleep disturbances, were concordant with those reported for a newly-described syndrome [Xia-Gibbs syndrome, OMIM #615829] attributed to heterozygous mutation in AHDC1 in only four cases [93].
1.3.4 Resolution of Translocation Breakpoints by Sequencing Provides New Information
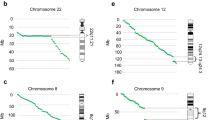
Whole-genome sequencing now permits discovery and precise localization of rearrangement breakpoints [58, 71, 82] (Table 1.1). Particularly, analysis of discordant mate-pair mappings from low-coverage, long-insert whole-genome sequencing is a cost-effective means of doing so [82], and is likely to one day displace standard karyotyping. Sequence-level breakpoint mapping provides identities of genes directly disrupted in a rearrangement, and of nearby genes that may be dysregulated by altered positioning of cis-regulatory enhancers, other regulatory elements, or regions of chromatin modification. Sequencing discovers complexity that is cryptic to karyotyping and imbalances that are below the resolution of clinical microarrays. De novo balanced translocations detected on prenatal karyotype can be assessed in a timely manner by this approach; for example, in prenatal cases reported by Ordulu et al. [61], sequencing supported or confirmed a suspected genetic diagnosis in most of the cases referred for abnormal prenatal findings. Redin et al. [71] reported sequenced breakpoints in 248 of 273 subjects, the majority ascertained via the Developmental Genome Anatomy Project (DGAP), a long-running effort to identify genes important in development by investigating apparently balanced rearrangements in patients with a wide variety of phenotypes, including neurodevelopmental disorders and structural congenital anomalies [26]. Redin et al. were able to make high-confidence correlations of genes to phenotypes in about a quarter of the subjects, and identify, in another 20%, likely candidate genes based on gene disruption or predicted position effects. They also documented additional complexity, cryptic to karyotype, in more than 20% of the rearrangements that they analyzed. This complexity included genomic gains or losses, some proportion of which would be invisible to clinical microarray. Among 65 subjects (26% of the total) with complex rearrangements (three or more breakpoints), 13 had multiple breakpoints characteristic of the “shattering” phenomena of chromothripsis or chromoplexy; in one case, 57 breakpoints were mapped. Overall, about 80% of the 248 analyzed rearrangements were balanced or nearly so, with less than 10 kb of genomic imbalance, indicating that the majority of these pathogenic mutations fail to leave even a footprint on clinical microarray.
Some balanced translocations are also cryptic to karyotyping. In an unusual case [68], a child with multiple congenital anomalies and some features overlapping those of cri-du-chat syndrome [OMIM #123450] was found by FISH to have a 4.6 Mb deletion at the terminus of 5p, much smaller than the canonical cri-du-chat region. Two previous pregnancies of the mother had been terminated, one for complete lissencephaly [OMIM #607432], and one for intrauterine growth restriction. The karyotypes of both of the parents and the affected child were unremarkable. However, FISH of the mother’s chromosomes, including a probe to 17p (a known lissencephaly locus), indicated a balanced translocation between 5p and 17p. The affected daughter had inherited the der(5) chromosome, resulting in gain of 17p as well as loss of 5p, with phenotypic features attributable to each. The authors hypothesized that the fetus with lissencephaly had inherited the der(17) chromosome, with presumed heterozygous loss of PAFAH1B1 [OMIM *601545] responsible for the lissencephaly phenotype. A cautionary tale, this cryptic translocation was hypothesized and uncovered only through thoughtful assessment of a “peculiar” pedigree and distinctive phenotypes already strongly linked to known loci. Without exceptional circumstances such as these, cryptic balanced translocations will remain undetected until whole-genome sequencing designed to ascertain rearrangements becomes routine.
1.3.5 Mapping the Regulatory Genome with Translocations
Whole exome sequencing of patients with conditions of suspected single-gene etiology currently yields known or likely candidate pathogenic mutations in about half. Extragenic regulatory mutation is likely to constitute a significant proportion of this “missing” mutation. Mammalian genomes are now well-known to be looped, folded, and scaffolded into three-dimensional architectures that influence gene expression through the spatial control of interactions of gene promoters with extragenic enhancers and other regulatory elements [12, 47, 70]. Chromosomes are physically organized into topologically-associated domains (TADs), “neighborhoods” of typically a megabase or smaller, that demonstrate higher frequencies of chromatin-chromatin contacts within domains than across domains [48, 49]. Enhancer-promoter contact frequencies define TADs, delimited by boundary elements that discourage promiscuous interactions of enhancers with non-target neighboring promoters; critical gene-regulatory enhancers may be arrayed along a chromosome at distances of several 100 kilobases to a megabase or more from the genes they regulate, particularly in the case of genes important in developmental processes [11].
Translocations can displace important enhancers or other regulatory elements from the genes they regulate. This has been demonstrated for several genes where translocations were central in flagging locations of distant cis-regulatory elements. An enhancer controlling limb expression of sonic hedgehog (Shh/SHH [OMIM *600725]) was localized to an intron of a distant gene and implicated in Shh/SHH expression in both mouse and human by a combination of evidence, including a translocation in a patient with preaxial polydactyly type II [PPD2, OMIM #174500], where one breakpoint mapped very close to the enhancer, nearly 1 Mb away from SHH itself [45, 46]. A downstream regulatory region was identified by mapping translocation and inversion breakpoints in patients with aniridia [OMIM #106210] where the causative gene, PAX6 [OMIM *607108], was found to be left intact by the rearrangements [15, 39]. Structural variants, including translocations, in patients with campomelic dysplasia or acampomelic campomelic dysplasia [CD and ACD, OMIM #114290] indicate that a complex SOX9 [OMIM *608160] regulatory landscape exists as far as 2 Mb upstream and 500 kb downstream of the gene itself; severity of the phenotype in translocation patients is broadly correlated with distance of the translocation breakpoint from SOX9 itself [22]. Other phenotypes are associated with copy number variation or rearrangements around SOX9, including Pierre-Robin sequence [PRS, OMIM %261800] and 46,XX and 46,XY disorders of sex development (DSDs) [OMIM #278850 and #613080], all of which can occur with or without accompanying CD or ACD. Translocations of patients with isolated PRS have been localized to two separate regions, about 1 Mb and 400 kb upstream of SOX9 [2, 61]; likewise, deletions and duplications in DSD patients define critical regulatory regions for sex development between 500 and 640 kb upstream of SOX9.
Brief descriptions belie the complexity of cis-regulatory regions, the variations that may occur among phenotypes of patients with rearrangements variously disrupting a regulatory locus, and the many mechanisms proposed to explain the pathogenicity of those rearrangements [3]. Beyond simply removing an enhancer from its cognate promoter, translocations and other rearrangements change the physical conformation of the regulatory locus, altering or disrupting chromatin loops that may affect the function of enhancers and other elements that remain between a cognate promoter and a rearrangement breakpoint. Enhancers brought into a locus by a rearrangement may make spurious contacts with an existing promoter, altering its expression. New chromatin conformations may alter the maintenance of the epigenetic landscape, resulting in gene expression changes via classical position effect mechanisms. Thus, predicting the effects of translocations that disrupt extragenic sequences is not trivial. Zepeda-Mendoza et al. [95] collated genome-wide datasets of enhancer marks, DNAse-hypersensitivity sites, and TAD boundaries predicted by chromatin contacts [13], along with haploinsufficiency and triplosensitivity scores [31] and phenotype information from DECIPHER [9] and ClinGen [72] to identify and prioritize candidate genes that may be etiologic in patients harboring balanced rearrangements with intergenic breakpoints. The success of this approach relies on the existence of multiple, diverse datasets that inform our understanding of the complexities of gene regulation and function, recognizes that rearrangements may result in dysfunction of genes located at distances up to several Mb, and depends most critically on the availability of large collections of patient phenotype-genotype information.
1.4 Summary
Much Mendelian disease is rare. Rarer still are patients whose congenital anomalies or neurodevelopmental disorders are caused by balanced translocations that disrupt genes or their cis-regulatory regions. Approaches to diagnosing genetic etiologies that discount the contribution of balanced chromosomal rearrangements disenfranchise those patients, lengthening diagnostic odysseys and adding medical costs, and sometimes compromising patient care. Reciprocal translocations and other balanced rearrangements represent an important class of pathogenic variation that is cryptic to chromosomal microarray, cannot be ascertained from exome sequencing data, and may be incompletely described or even undetected by karyotyping. Nucleotide-level resolution of rearrangement breakpoints is essential for interpreting the etiology of phenotypes in patients with balanced rearrangements.
References
Bandyopadhyay R, Heller A, Knox-DuBois C, McCaskill C, Berend SA, Page SL, Shaffer LG (2002) Parental origin and timing of de novo Robertsonian translocation formation. Am J Hum Genet 71(6):1456–1462. https://doi.org/10.1086/344662
Benko S, Fantes JA, Amiel J, Kleinjan DJ, Thomas S, Ramsay J, Jamshidi N, Essafi A, Heaney S, Gordon CT, McBride D, Golzio C, Fisher M, Perry P, Abadie V, Ayuso C, Holder-Espinasse M, Kilpatrick N, Lees MM, Picard A, Temple IK, Thomas P, Vazquez MP, Vekemans M, Roest Crollius H, Hastie ND, Munnich A, Etchevers HC, Pelet A, Farlie PG, Fitzpatrick DR, Lyonnet S (2009) Highly conserved non-coding elements on either side of SOX9 associated with Pierre Robin sequence. Nat Genet 41(3):359–364. https://doi.org/10.1038/ng.329
Bhatia S, Kleinjan DA (2014) Disruption of long-range gene regulation in human genetic disease: a kaleidoscope of general principles, diverse mechanisms and unique phenotypic consequences. Hum Genet 133(7):815–845. https://doi.org/10.1007/s00439-014-1424-6
Book JA, Fraccaro M, Lindsten J (1959) Cytogenetical observations in mongolism. Acta Paediatr 48:453–468
Boué A, Gallano P (1984) A collaborative study of the segregation of inherited chromosome structural rearrangements in 1356 prenatal diagnoses. Prenat Diagn 4 Spec No:45–67
Carter CO, Hamerton JL, Polani PE, Gunalp A, Weller SD (1960) Chromosome translocation as a cause of familial mongolism. Lancet 2(7152):678–680
Caspersson T, Zech L, Johansson C, Modest EJ (1970) Identification of human chromosomes by DNA-binding fluorescent agents. Chromosoma 30(2):215–227
De Braekeleer M, Dao T-N (1990) Cytogenetic studies in couples experiencing repeated pregnancy losses. Hum Reprod 5(5):519–528
DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans using Ensembl Resources. Firth HV et al (2009) Am J Hum Genet 84 524–533. https://doi.org/10.1016/j.ajhg.2009.03.010
de Klein A, van Kessel AG, Grosveld G, Bartram CR, Hagemeijer A, Bootsma D, Spurr NK, Heisterkamp N, Groffen J, Stephenson JR (1982) A cellular oncogene is translocated to the Philadelphia chromosome in chronic myelocytic leukaemia. Nature 300(5894):765–767
de Laat W, Duboule D (2013) Topology of mammalian developmental enhancers and their regulatory landscapes. Nature 502(7472):499–506. https://doi.org/10.1038/nature12753
Dekker J, Rippe K, Dekker M, Kleckner N (2002) Capturing chromosome conformation. Science 295(5558):1306–1311. https://doi.org/10.1126/science.1067799
Dixon JR, Selvaraj S, Yue F, Kim A, Li Y, Shen Y, Hu M, Liu JS, Ren B (2012) Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485(7398):376–380. https://doi.org/10.1038/nature11082
Edwards J, Fraccaro M, Davies P, Young R (1962) Structural heterozygosis in man: analysis of two families. Annals Hum Genet Lond 26:163–178
Fantes J, Redeker B, Breen M, Boyle S, Brown J, Fletcher J, Jones S, Bickmore W, Fukushima Y, Mannens M (1995) Aniridia-associated cytogenetic rearrangements suggest that a position effect may cause the mutant phenotype. Hum Mol Genet 4(3):415–422
Forabosco A, Percesepe A, Santucci S (2009) Incidence of non-age-dependent chromosomal abnormalities: a population-based study on 88965 amniocenteses. Eur J Hum Genet 17(7):897–903. https://doi.org/10.1038/ejhg.2008.265
Ford CE, Hamerton JL (1956a) The chromosomes of man. Nature 178(4541):1020–1023
Ford CE, Hamerton JL (1956b) A colchicine, hypotonic citrate, squash sequence for mammalian chromosomes. Stain Technol 31(6):247–251
Ford CE, Jones KW, Miller OJ, Mittwoch U, Penrose LS, Ridler M, Shapiro A (1959) The chromosomes in a patient showing both mongolism and the Klinefelter syndrome. Lancet 1(7075):709–710
Fraccaro M, Lindsten J, Ford CE, Iselius L (1980) The 11q;22q translocation: a European collaborative analysis of 43 cases. Hum Genet 56(1):21–51
Funderburk SJ, Spence MA, Sparkes RS (1977) Mental retardation associated with “balanced” chromosome rearrangements. Am J Hum Genet 29(2):136–141
Gordon CT, Attanasio C, Bhatia S, Benko S, Ansari M, Tan TY, Munnich A, Pennacchio LA, Abadie V, Temple IK, Goldenberg A, van Heyningen V, Amiel J, FitzPatrick D, Kleinjan DA, Visel A, Lyonnet S (2014) Identification of novel craniofacial regulatory domains located far upstream of SOX9 and disrupted in Pierre Robin sequence. Hum Mutat 35(8):1011–1020. https://doi.org/10.1002/humu.22606
Groffen J, Stephenson JR, Heisterkamp N, de Klein A, Bartram CR, Grosveld G (1984) Philadelphia chromosomal breakpoints are clustered within a limited region, bcr, on chromosome 22. Cell 36(1):93–99
Hamerton JL, Canning N, Ray M, Smith S (1975) A cytogenetic survey of 14,069 newborn infants I. Incidence of chromosome abnormalities. Clin Genet 8(4):223–243
Heisterkamp N, Stephenson JR, Groffen J, Hansen PF, de Klein A, Bartram CR, Grosveld G (1983) Localization of the c-ab1 oncogene adjacent to a translocation break point in chronic myelocytic leukaemia. Nature 306(5940):239–242
Higgins AW, Alkuraya FS, Bosco AF, Brown KK, Bruns GA, Donovan DJ, Eisenman R, Fan Y, Farra CG, Ferguson HL, Gusella JF, Harris DJ, Herrick SR, Kelly C, Kim HG, Kishikawa S, Korf BR, Kulkarni S, Lally E, Leach NT, Lemyre E, Lewis J, Ligon AH, Lu W, Maas RL, MacDonald ME, Moore SD, Peters RE, Quade BJ, Quintero-Rivera F, Saadi I, Shen Y, Shendure J, Williamson RE, Morton CC (2008) Characterization of apparently balanced chromosomal rearrangements from the developmental genome anatomy project. Am J Hum Genet 82(3):712–722. https://doi.org/10.1016/j.ajhg.2008.01.011
Hook EB, Hamerton JL (1978) Analyses of data on rates of cytogenetic disorders in live births. Am J Hum Genet 30(3):330–331
Hook EB, Schreinemachers DM, Willey AM, Cross PK (1983) Rates of mutant structural chromosome rearrangements in human fetuses: data from prenatal cytogenetic studies and associations with maternal age and parental mutagen exposure. Am J Hum Genet 35(1):96–109
Hsu TC (1952) Mammalian chromosomes in vitro I. The karyotype of man. J Hered 44:167–172
Hsu TC, Pomerat CM (1953) Mammalian chromosomes in vitro II. A method for spreading the chromosomes of cells in tissue culture. J Heredity 44:23–29
Huang N, Lee I, Marcotte EM, Hurles ME (2010) Characterising and predicting haploinsufficiency in the human genome. PLoS Genet 6(10):e1001154. https://doi.org/10.1371/journal.pgen.1001154
Hungerford DA, Donnelly AJ, Nowell PC, Beck S (1959) The chromosome constitution of a human phenotypic intersex. Am J Hum Genet 11:215–236
Imaizumi K, Kimura J, Matsuo M, Kurosawa K, Masuno M, Niikawa N, Kuroki Y (2002) Sotos syndrome associated with a de novo balanced reciprocal translocation t(5;8)(q35;q24.1). Am J Med Genet 107(1):58–60
Jacobs PA (1974) Correlation between euploid structural chromosome rearrangements and mental subnormality in humans. Nature 249(453):164–165
Jacobs PA, Baikie AG, Court Brown WM, Strong JA (1959) The somatic chromosomes in mongolism. Lancet 1(7075):710
Jacobs PA, Browne C, Gregson N, Joyce C, White H (1992) Estimates of the frequency of chromosome abnormalities detectable in unselected newborns using moderate levels of banding. J Med Genet 29(2):103–108
Kato T, Kurahashi H, Emanuel BS (2012) Chromosomal translocations and palindromic AT-rich repeats. Curr Opin Genet Dev 22(3):221–228. https://doi.org/10.1016/j.gde.2012.02.004
Kim HG, Kishikawa S, Higgins AW, Seong IS, Donovan DJ, Shen Y, Lally E, Weiss LA, Najm J, Kutsche K, Descartes M, Holt L, Braddock S, Troxell R, Kaplan L, Volkmar F, Klin A, Tsatsanis K, Harris DJ, Noens I, Pauls DL, Daly MJ, MacDonald ME, Morton CC, Quade BJ, Gusella JF (2008) Disruption of neurexin 1 associated with autism spectrum disorder. Am J Hum Genet 82(1):199–207. https://doi.org/10.1016/j.ajhg.2007.09.011
Kleinjan DA, van Heyningen V (2005) Long-range control of gene expression: emerging mechanisms and disruption in disease. Am J Hum Genet 76(1):8–32. https://doi.org/10.1086/426833
Konopka JB, Watanabe SM, Witte ON (1984) An alteration of the human c-abl protein in K562 leukemia cells unmasks associated tyrosine kinase activity. Cell 37(3):1035–1042
Kurotaki N, Imaizumi K, Harada N, Masuno M, Kondoh T, Nagai T, Ohashi H, Naritomi K, Tsukahara M, Makita Y, Sugimoto T, Sonoda T, Hasegawa T, Chinen Y, Tomita Ha HA, Kinoshita A, Mizuguchi T, Yoshiura Ki K, Ohta T, Kishino T, Fukushima Y, Niikawa N, Matsumoto N (2002) Haploinsufficiency of NSD1 causes Sotos syndrome. Nat Genet 30(4):365–366. https://doi.org/10.1038/ng863
Ledbetter DH, Rich DC, O’Connell P, Leppert M, Carey JC (1989) Precise localization of NF1 to 17q11.2 by balanced translocation. Am J Hum Genet 44(1):20–24
Lejeune J, Gautier M, Turpin R (1959) Study of somatic chromosomes from 9 mongoloid children. C R Hebd Seances Acad Sci 248(11):1721–1722
Lejeune J, Turpin R, Decourt J (1960) Chromosome aberrations and human diseases XXY Klinefelter’s syndrome from 46 chromosomes caused by T-T centromeric fusion. C R Hebd Seances Acad Sci 250:2468–2470
Lettice LA, Heaney SJ, Purdie LA, Li L, de Beer P, Oostra BA, Goode D, Elgar G, Hill RE, de Graaff E (2003) A long-range Shh enhancer regulates expression in the developing limb and fin and is associated with preaxial polydactyly. Hum Mol Genet 12(14):1725–1735
Lettice LA, Horikoshi T, Heaney SJ, van Baren MJ, van der Linde HC, Breedveld GJ, Joosse M, Akarsu N, Oostra BA, Endo N, Shibata M, Suzuki M, Takahashi E, Shinka T, Nakahori Y, Ayusawa D, Nakabayashi K, Scherer SW, Heutink P, Hill RE, Noji S (2002) Disruption of a long-range cis-acting regulator for Shh causes preaxial polydactyly. Proc Natl Acad Sci U S A 99(11):7548–7553. https://doi.org/10.1073/pnas.112212199
Liang S, Tippens ND, Zhou Y, Mort M, Stenson PD, Cooper DN, Yu H (2017) iRegNet3D: three-dimensional integrated regulatory network for the genomic analysis of coding and non-coding disease mutations. Genome Biol 18(1):10. https://doi.org/10.1186/s13059-016-1138-2
Lupiáñez DG, Kraft K, Heinrich V, Krawitz P, Brancati F, Klopocki E, Horn D, Kayserili H, Opitz JM, Laxova R, Santos-Simarro F, Gilbert-Dussardier B, Wittler L, Borschiwer M, Haas SA, Osterwalder M, Franke M, Timmermann B, Hecht J, Spielmann M, Visel A, Mundlos S (2015) Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161(5):1012–1025. https://doi.org/10.1016/j.cell.2015.04.004
Lupiáñez DG, Spielmann M, Mundlos S (2016) Breaking TADs: how alterations of chromatin domains result in disease. Trends Genet 32(4):225–237. https://doi.org/10.1016/j.tig.2016.01.003
MacGregor DJ, Imrie S, Tolmie JL (1989) Outcome of de novo balanced translocations ascertained prenatally. J Med Genet 26(9):590–591
Mackie Ogilvie C, Scriven PN (2002) Meiotic outcomes in reciprocal translocation carriers ascertained in 3-day human embryos. Eur J Hum Genet 10(12):801–806. https://doi.org/10.1038/sj.ejhg.5200895
Maroun C, Schmerler S, Hutcheon RG (1994) Child with Sotos phenotype and a 5:15 translocation. Am J Med Genet 50(3):291–293. https://doi.org/10.1002/ajmg.1320500313
Moorhead PS, Nowell PC, Mellman WJ, Battips DM, Hungerford DA (1960) Chromosome preparations of leukocytes cultured from human peripheral blood. Exp Cell Res 20:613–616
Morel F, Douet-Guilbert N, Le Bris MJ, Herry A, Amice V, Amice J, De Braekeleer M (2004) Meiotic segregation of translocations during male gametogenesis. Int J Androl 27(4):200–212. https://doi.org/10.1111/j.1365-2605.2004.00490.x
Mutton D, Alberman E, Hook EB (1996) Cytogenetic and epidemiological findings in down syndrome, England and Wales 1989 to 1993. National down Syndrome Cytogenetic Register and the Association of Clinical Cytogeneticists. J Med Genet 33(5):387–394
Nielsen J, Krag-Olsen B (1981) Follow-up of 32 children with autosomal translocations found among 11,148 consecutively newborn children from 1969 to 1974. Clin Genet 20(1):48–54
Nielsen J, Wohlert M (1991) Chromosome abnormalities found among 34,910 newborn children: results from a 13-year incidence study in Arhus, Denmark. Hum Genet 87(1):81–83
Nilsson D, Pettersson M, Gustavsson P, Förster A, Hofmeister W, Wincent J, Zachariadis V, Anderlid BM, Nordgren A, Mäkitie O, Wirta V, Käller M, Vezzi F, Lupski JR, Nordenskjöld M, Lundberg ES, Carvalho CMB, Lindstrand A (2017) Whole-genome sequencing of cytogenetically balanced chromosome translocations identifies potentially pathological gene disruptions and highlights the importance of microhomology in the mechanism of formation. Hum Mutat 38(2):180–192. https://doi.org/10.1002/humu.23146
Nowell PC, Hungerford DA (1960) Chromosome studies on normal and leukemic human leukocytes. J Natl Cancer Inst 25:85–109
Oliver-Bonet M, Navarro J, Carrera M, Egozcue J, Benet J (2002) Aneuploid and unbalanced sperm in two translocation carriers: evaluation of the genetic risk. Mol Hum Reprod 8(10):958–963
Ordulu Z, Kammin T, Brand H, Pillalamarri V, Redin CE, Collins RL, Blumenthal I, Hanscom C, Pereira S, Bradley I, Crandall BF, Gerrol P, Hayden MA, Hussain N, Kanengisser-Pines B, Kantarci S, Levy B, Macera MJ, Quintero-Rivera F, Spiegel E, Stevens B, Ulm JE, Warburton D, Wilkins-Haug LE, Yachelevich N, Gusella JF, Talkowski ME, Morton CC (2016) Structural chromosomal rearrangements require nucleotide-level resolution: lessons from next-generation sequencing in prenatal diagnosis. Am J Hum Genet 99(5):1015–1033. https://doi.org/10.1016/j.ajhg.2016.08.022
Ozawa N, Maruyama T, Nagashima T, Ono M, Arase T, Ishimoto H, Yoshimura Y (2008) Pregnancy outcomes of reciprocal translocation carriers who have a history of repeated pregnancy loss. Fertil Steril 90(4):1301–1304. https://doi.org/10.1016/j.fertnstert.2007.09.051
Page SL, Shaffer LG (1997) Nonhomologous Robertsonian translocations form predominantly during female meiosis. Nat Genet 15(3):231–232. https://doi.org/10.1038/ng0397-231
Penrose LS (1933) The relative effects of paternal and maternal age in mongolism. Reproduced in 2009. J Genet 88(1):9–14
Penrose LS, Ellis JR, Delhanty JD (1960) Chromosomal translocations in mongolism and in normal relatives. Lancet 2(7147):409–410
Perrin A, Morel F, Douet-Guilbert N, Le Bris MJ, Amice J, Amice V, De Braekeleer M (2010) A study of meiotic segregation of chromosomes in spermatozoa of translocation carriers using fluorescent in situ hybridisation. Andrologia 42(1):27–34. https://doi.org/10.1111/j.1439-0272.2009.00951.x
Polani PE, Briggs JH, Ford CE, Clarke CM, Berg JM (1960) A Mongol girl with 46 chromosomes. Lancet 1(7127):721–724
Primerano A, Colao E, Villella C, Nocera MD, Ciambrone A, Luciano E, D’Antona L, Vismara MFM, Loddo S, Novelli A, Perrotti N, Malatesta P (2015) A cryptic balanced translocation (5;17), a puzzle revealed through a critical evaluation of the pedigree and a FISH focused on candidate loci suggested by the phenotype. Mol Cytogenet 8:70. https://doi.org/10.1186/s13039-015-0172-1
Quintero-Rivera F, Xi QJ, Keppler-Noreuil KM, Lee JH, Higgins AW, Anchan RM, Roberts AE, Seong IS, Fan X, Lage K, Lu LY, Tao J, Hu X, Berezney R, Gelb BD, Kamp A, Moskowitz IP, Lacro RV, Lu W, Morton CC, Gusella JF, Maas RL (2015) MATR3 disruption in human and mouse associated with bicuspid aortic valve, aortic coarctation and patent ductus arteriosus. Hum Mol Genet 24(8):2375–2389. https://doi.org/10.1093/hmg/ddv004
Rao SS, Huntley MH, Durand NC, Stamenova EK, Bochkov ID, Robinson JT, Sanborn AL, Machol I, Omer AD, Lander ES, Aiden EL (2014) A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159(7):1665–1680. https://doi.org/10.1016/j.cell.2014.11.021
Redin C, Brand H, Collins RL, Kammin T, Mitchell E, Hodge JC, Hanscom C, Pillalamarri V, Seabra CM, Abbott MA, Abdul-Rahman OA, Aberg E, Adley R, Alcaraz-Estrada SL, Alkuraya FS, An Y, Anderson MA, Antolik C, Anyane-Yeboa K, Atkin JF, Bartell T, Bernstein JA, Beyer E, Blumenthal I, Bongers EM, Brilstra EH, Brown CW, Brüggenwirth HT, Callewaert B, Chiang C, Corning K, Cox H, Cuppen E, Currall BB, Cushing T, David D, Deardorff MA, Dheedene A, D’Hooghe M, de Vries BB, Earl DL, Ferguson HL, Fisher H, FitzPatrick DR, Gerrol P, Giachino D, Glessner JT, Gliem T, Grady M, Graham BH, Griffis C, Gripp KW, Gropman AL, Hanson-Kahn A, Harris DJ, Hayden MA, Hill R, Hochstenbach R, Hoffman JD, Hopkin RJ, Hubshman MW, Innes AM, Irons M, Irving M, Jacobsen JC, Janssens S, Jewett T, Johnson JP, Jongmans MC, Kahler SG, Koolen DA, Korzelius J, Kroisel PM, Lacassie Y, Lawless W, Lemyre E, Leppig K, Levin AV, Li H, Liao EC, Lim C, Lose EJ, Lucente D, Macera MJ, Manavalan P, Mandrile G, Marcelis CL, Margolin L, Mason T, Masser-Frye D, McClellan MW, Mendoza CJ, Menten B, Middelkamp S, Mikami LR, Moe E, Mohammed S, Mononen T, Mortenson ME, Moya G, Nieuwint AW, Ordulu Z, Parkash S, Pauker SP, Pereira S, Perrin D, Phelan K, Aguilar RE, Poddighe PJ, Pregno G, Raskin S, Reis L, Rhead W, Rita D, Renkens I, Roelens F, Ruliera J, Rump P, Schilit SL, Shaheen R, Sparkes R, Spiegel E, Stevens B, Stone MR, Tagoe J, Thakuria JV, van Bon BW, van de Kamp J, van Der Burgt I, van Essen T, van Ravenswaaij-Arts CM, van Roosmalen MJ, Vergult S, Volker-Touw CM, Warburton DP, Waterman MJ, Wiley S, Wilson A, Yerena-de Vega MC, Zori RT, Levy B, Brunner HG, de Leeuw N, Kloosterman WP, Thorland EC, Morton CC, Gusella JF, Talkowski ME (2017) The genomic landscape of balanced cytogenetic abnormalities associated with human congenital anomalies. Nat Genet 49(1):36–45. https://doi.org/10.1038/ng.3720
Rehm HL, Berg JS, Brooks LD, Bustamante CD, Evans JP, Landrum MJ, Ledbetter DH, Maglott DR, Martin CL, Nussbaum RL, Plon SE, Ramos EM, Sherry ST, Watson MS, ClinGen (2015) ClinGen--the clinical genome resource. N Engl J Med 372(23):2235–2242. https://doi.org/10.1056/NEJMsr1406261
Rowley JD (1973) Letter: a new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and Giemsa staining. Nature 243(5405):290–293
Schmid W (1962) A familial chromosome abnormality associated with repeated abortions. Cytogenetics 1:199–209
Schmidt MA, Michels VV, Dewald GW (1987) Cases of neurofibromatosis with rearrangements of chromosome 17 involving band 17q11.2. Am J Med Genet 28:771–777
Seabright M (1971) A rapid banding technique for human chromosomes. Lancet 2(7731):971–972
Shaffer LG, Jackson-Cook CK, Meyer JM, Brown JA, Spence JE (1991) A molecular genetic approach to the identification of isochromosomes of chromosome 21. Hum Genet 86(4):375–382
Sheridan MB, Kato T, Haldeman-Englert C, Jalali GR, Milunsky JM, Zou Y, Klaes R, Gimelli G, Gimelli S, Gemmill RM, Drabkin HA, Hacker AM, Brown J, Tomkins D, Shaikh TH, Kurahashi H, Zackai EH, Emanuel BS (2010) A palindrome-mediated recurrent translocation with 3:1 meiotic nondisjunction: the t(8;22)(q24.13;q11.21). Am J Hum Genet 87(2):209–218. https://doi.org/10.1016/j.ajhg.2010.07.002
Shtivelman E, Lifshitz B, Gale RP, Canaani E (1985) Fused transcript of abl and bcr genes in chronic myelogenous leukaemia. Nature 315(6020):550–554
Sugiura-Ogasawara M, Aoki K, Fujii T, Fujita T, Kawaguchi R, Maruyama T, Ozawa N, Sugi T, Takeshita T, Saito S (2008) Subsequent pregnancy outcomes in recurrent miscarriage patients with a paternal or maternal carrier of a structural chromosome rearrangement. J Hum Genet 53(7):622–628. https://doi.org/10.1007/s10038-008-0290-2
Sumner AT, Evans HJ, Buckland RA (1971) New technique for distinguishing between human chromosomes. Nat New Biol 232(27):31–32
Talkowski ME, Ernst C, Heilbut A, Chiang C, Hanscom C, Lindgren A, Kirby A, Liu S, Muddukrishna B, Ohsumi TK, Shen Y, Borowsky M, Daly MJ, Morton CC, Gusella JF (2011) Next-generation sequencing strategies enable routine detection of balanced chromosome rearrangements for clinical diagnostics and genetic research. Am J Hum Genet 88(4):469–481. https://doi.org/10.1016/j.ajhg.2011.03.013
Taub R, Kirsch I, Morton C, Lenoir G, Swan D, Tronick S, Aaronson S, Leder P (1982) Translocation of the c-myc gene into the immunoglobulin heavy chain locus in human Burkitt lymphoma and murine plasmacytoma cells. Proc Natl Acad Sci U S A 79(24):7837–7841
Thomas NS, Morris JK, Baptista J, Ng BL, Crolla JA, Jacobs PA (2010) De novo apparently balanced translocations in man are predominantly paternal in origin and associated with a significant increase in paternal age. J Med Genet 47(2):112–115. https://doi.org/10.1136/jmg.2009.069716
Tierney I, Axworthy D, Smith L, Ratcliffe SG (1984) Balanced rearrangements of the autosomes: results of a longitudinal study of a newborn survey population. J Med Genet 21(1):45–51
Tjio JH, Levan A (1956) The chromosome number of man. Hereditas 42:1–6
Turpin R, Lejeune J (1969) Human Afflications and Chromosomal Aberrations. Translation based on Les Chromosomes Humains (caryotype normal et variations pathologiques), Gauthier-Villars, 1965. International series of monographs in pure and applied biology. Modern trends in physiological sciences, vol 32. Pergamon Press, Oxford
Turpin R, Lejeune J, Lafourcade J, Gautier M (1959) Chromosome aberrations & human diseases; multiple spinal abnormalities with 45 chromosomes. C R Hebd Seances Acad Sci 248(25):3636–3638
Van Dyke DL, Weiss L, Roberson JR, Babu VR (1983) The frequency and mutation rate of balanced autosomal rearrangements in man estimated from prenatal genetic studies for advanced maternal age. Am J Hum Genet 35 (2):301–308.
Vincent MC, Daudin M, De MP, Massat G, Mieusset R, Pontonnier F, Calvas P, Bujan L, Bourrouillout G (2002) Cytogenetic investigations of infertile men with low sperm counts: a 25-year experience. J Androl 23(1):18–22. discussion 44–15
Walker S, Harris R (1962) Investigation of family showing transmission of a 13-15 chromosomal translocation (Denver classification). Br Med J 2(5296):25–26
Warburton D (1991) De novo balanced chromosome rearrangements and extra marker chromosomes identified at prenatal diagnosis: clinical significance and distribution of breakpoints. Am J Hum Genet 49(5):995–1013
Xia F, Bainbridge MN, Tan TY, Wangler MF, Scheuerle AE, Zackai EH, Harr MH, Sutton VR, Nalam RL, Zhu W, Nash M, Ryan MM, Yaplito-Lee J, Hunter JV, Deardorff MA, Penney SJ, Beaudet AL, Plon SE, Boerwinkle EA, Lupski JR, Eng CM, Muzny DM, Yang Y, Gibbs RA (2014) De novo truncating mutations in AHDC1 in individuals with syndromic expressive language delay, hypotonia, and sleep apnea. Am J Hum Genet 94(5):784–789. https://doi.org/10.1016/j.ajhg.2014.04.006
Zackai EH, Emanuel BS (1980) Site-specific reciprocal translocation, t(11;22) (q23;q11), in several unrelated families with 3:1 meiotic disjunction. Am J Med Genet 7(4):507–521. https://doi.org/10.1002/ajmg.1320070412
Zepeda-Mendoza CJ, Ibn-Salem J, Kammin T, Harris DJ, Rita D, Gripp KW, MacKenzie JJ, Gropman A, Graham B, Shaheen R, Alkuraya FS, Brasington CK, Spence EJ, Masser-Frye D, Bird LM, Spiegel E, Sparkes RL, Ordulu Z, Talkowski ME, Andrade-Navarro MA, Robinson PN, Morton CC (2017) Computational prediction of position effects of apparently balanced human chromosomal rearrangements. Am J Hum Genet. https://doi.org/10.1016/j.ajhg.2017.06.011
Zhang HG, Wang RX, Li LL, Sun WT, Zhang HY, Liu RZ (2015) Male carriers of balanced reciprocal translocations in Northeast China: sperm count, reproductive performance, and genetic counseling. Genet Mol Res 14(4):18792–18798. https://doi.org/10.4238/2015.December.28.28
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Wilch, E.S., Morton, C.C. (2018). Historical and Clinical Perspectives on Chromosomal Translocations. In: Zhang, Y. (eds) Chromosome Translocation. Advances in Experimental Medicine and Biology, vol 1044. Springer, Singapore. https://doi.org/10.1007/978-981-13-0593-1_1
Download citation
DOI: https://doi.org/10.1007/978-981-13-0593-1_1
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-0592-4
Online ISBN: 978-981-13-0593-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)