Abstract
Consumption of foodstuff with low contents of saturated fatty acids is considered beneficial for human health. Reducing saturated fatty acid content in oilseed rape (canola) and other oil and protein crops is a relevant breeding aim. The objective of this work was to study the genetic variation and inheritance of saturated fatty acids in two DH populations of oilseed rape, to map QTL and to identify candidate genes. In addition, the correlation to other seed quality traits was studied. To this end, two half-sib DH populations were tested in up to five field environments in north-western Europe and seeds harvested from open-pollinated seeds were analyzed. Genotyping was performed using Illumina Brassica 15 K SNP chip. In both populations, significant effects for the genotypes and for the environments were detected, and heritability ranged from 68 to 89% for the predominant palmitic acid and stearic acid content. Up to 48 QTL for different fatty acids, oil and acid detergent lignin (ADL) content were mapped in the two populations. Co-locating QTL for palmitic acid, stearic acid, the C16/18 fatty acid ratio, the FATB/A ratio, oil and ADL content were identified on different chromosomes. A large number of candidate genes were identified within the vicinity of QTL flanking markers. Identification of several co-locating QTL positions, of associated candidate genes and SNP markers should facilitate oilseed rape breeding for low saturated fatty acid content.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
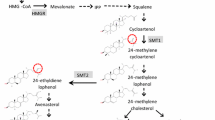
Oilseed rape (Brassica napus L., AACC, 2n = 38) is an allotetraploid species of winter and spring genotypes. It is the third largest source of vegetable oil after oil palm and soybean (FAOSTAT 2020). The production of oilseed rape is predominantly in most part of Europe, Canada, Australia, China and India, with over 36.5 million ha of cultivation worldwide with annual global production of about 72 million tons (FAOSTAT 2020). Even though the by-product after oil extraction from the seed is the protein rich meal, the major interest in oilseed rape used to be the seed oil. Recently, other seed quality traits like increasing the protein and reducing the fibre content are gaining importance (Balalić et al. 2017). The oil quality and its nutritive value is determined by the relative composition of the fatty acid content present in the oil (Vick et al. 2002). Modern cultivars of rapeseed contain about 60–65% monounsaturated fatty acids of mainly oleic acid (18:1), 30–35% polyunsaturated fatty acids of mainly linoleic acid (18:2) and linolenic acid (18:3), and 5–8% saturated fatty acids (SFA) (Chen et al. 2018). Monounsaturated 18:1 and polyunsaturated 18:2 are beneficial to human health and are less susceptible to oxidation during frying (Merrill et al. 2008; Zhao et al. 2019), so are desirable traits for improvement. However, presence of high SFA content in the oil, possess health risk, as it is reported to elevate the Low Density Lipoprotein (LDL) in blood serum causing coronary heart disease (Rebetzke et al. 2001). From the human nutritional point of view, the content of SFA should be as low as possible. Palmitic (16:0) and stearic acid (18:0) are the major components of SFA in Brassica napus (L.) and its composition in the oil content is receiving interest because dietary recommendations limit its intake to less than 7% (Beaith et al. 2005). Like other fatty acids, the variation of SFA content is quantitatively inherited in plants, which depends on the genotype, the environment and their interactions (Rebetzke et al. 2001). As a result, genetic reduction of SFA will increase the integrity of the oil content and the competitive ability with other vegetable oils utilized for nutritional purpose (Gororo et al. 2014). A low SFA content of less than 7% is critical for labelling rapeseed oil as low SFA product in the USA (Rakow and Raney 2005). In plant biosynthesis, fatty acid synthesis (FAS) takes place in the plastid while oil synthesis in form of triacylglycerol (TAG) takes place in the cytosol at the endoplasmic reticulum. Several enzymes are involved in the biosynthetic pathways producing fatty acids, which regulate the composition of different fatty acids in the oil (Gororo et al. 2014). As a result, genetic modifications of fatty acids can be achieved by studying enzyme activities in the respective biosynthetic pathways (Stoll et al. 2005). Mapping of QTL linked to genes encoding enzymes in the fatty acids and TAG synthesis will be of great interest to elucidate the complexity involved in the accumulation of fatty acids in the oil. Genetic basis of fatty acid biosynthesis and pathway have been characterized in Arabidopsis thaliana (Li-Beisson et al. 2013). For the SFAs, they are generally synthesized in the plastid and exported to cytoplasm to be incorporated into TAG. In the plastids, palmitoyl-ACP (16:0-ACP) is the primary product of series of fatty acid synthesis (FAS) catalyzed by two condensing enzymes 3-ketoacyl-ACP synthase I and III (KASI and KASIII). Palmitoyl-ACP is either desaturated to palmitoleic acid (16:1) or further elongated to stearoyl-ACP (18:0-ACP) catalyzed by the enzyme 3-keto-acyl-synthase II (KASII). The competitive efficiency of the KASII enzyme can be calculated as the ratio of 16:0 + 16:1/18:1 + 18:0 + 18:2 + 18:3 + 20:0 + 20:1 + 22:0 + 22:1 (C16/18). Stearoyl-ACP (18:0-ACP) in turn is desaturated by stearoyl-CoA desaturase (SAD) to oleoyl-ACP (18:1-ACP) before being hydrolyzed and activated to 18:1-CoA catalyzed by acyl carrier protein thioesterase- A (FATA) enzyme; 18:1-CoA in the cytoplasm will be either incorporated to TAG synthesis or desaturated to linoleic acid (18:2). On the other hand, palmitoyl-ACP (16:0-ACP) and stearoyl-ACP can also be released into free 16:0 and 18:0, respectively, by the acyl carrier protein thioesterase-B (FATB) enzyme and exported to the cytoplasm where it is activated to 16:0-CoA and 18:0-CoA for incorporation to TAG as well. Since FATB and FATA thioesterases preferentially accepts saturated fatty acids (Sun et al. 2014) and unsaturated fatty acids, respectively, the FATB/A ratio can be calculated as 16:0 + 18:0 + 20:0 + 22:0/16:1 + 18:1 + 20:1 + 22:1. Otherwise, relative accumulation of palmitic acid and stearic acid in the oil is influenced by the competition between KASII and FATB (Ohlrogge and Browse 1995; Möllers and Schierholt 2002) along with activities of SAD (Karim Zarhloul et al. 2006). Reduced ratio of C16- to C18-fatty acids (C16/18 ratio) may indicate increased KASII-over FATB-activities. As a result, more palmitic acid will be converted to stearic acid and subsequently elongated or desaturated to C18 fatty acids. TAG metabolism involved three acylation steps involving glycerol-3-phosphate acyltransferase (GPAAT), lysophosphatidic acid acyltransferase (LPAT) and diacylglycerol acyltransferase (DGAT1) which finally results in accumulation of oil in form of TAG (Ohlrogge and Jaworski 1997).
There have been several studies on the expression of genes encoding some fatty acid synthase enzymes and the phenotypic expression of SFA in the seed oil content of crops like maize (Zheng et al. 2014), Arabidopsis (Fahy et al. 2013; Yao et al. 2003), soybean (Wilson et al. 2001) and oilseed rape (Moon et al. 2000; Bondaruk et al. 2007; Fahy et al. 2013; Sun et al. 2014). In oilseed rape, mutagenesis was successfully used to study the FAS genes involved in accumulation of saturated fatty acids (Beaith et al. 2005; Yao et al. 2003; Schnurbusch et al. 2000). Low SFA Brassica napus line was developed from the interspecific cross with Brassica rapa (Rakow and Raney 2005). Using a doubled haploid (DH) population, Möllers and Schierholt (2002) explained the pleiotropic effect of KASII of a high oleic acid (18:1) mutation on palmitic acid (16:0) and oil content. Significant genetic variation among winter oilseed rape cultivars was reported by Dimov and Möllers (2010). Sun et al. (2014) observed high accumulation of palmitic and stearic acid in oilseed rape by simultaneously overexpressing acyl carrier protein thioesterase-B (FATB) alleles and suppressing stearoyl-CoA desaturase (SAD) genes. However, studies on QTL linked to genes encoding enzymes involved in the accumulation of SFA in winter oilseed rape are limited. Most reported articles focus on erucic acid (Lu et al. 2020), oleic acid (Zhao et al. 2019; Yao et al. 2020), and fatty acids in general (Gazave et al. 2020; Chen et al. 2018; Teh and Möllers 2016; Yan et al. 2011; Zhao et al. 2008). Minor QTL were previously reported on A08 and C03 for both 16:0 and 18:0 using linkage mapping and association mapping studies (Bao et al, 2018; Guan et al, 2019), while Javed et al. (2016) mapped a major QTL for 16:0 content on C08 and 20:0 on A10.
In this study, two doubled haploid populations derived from crosses between canola cultivar Adriana with erucic acid DH line SGEDH13 and with the Chinese canola cultivar Zheyou 50, were investigated. The aim of this work was to (1) Study the genetic variation and inheritance of saturated fatty acids and their correlations with other fatty acids and oil content in two DH populations, to map QTL and to identify candidate genes involved in the expression of these traits.
Materials and methods
Plant material
The study material consisted of two DH populations. The first population consisted of 170 F1 derived doubled haploid (DH) lines from a cross between the canola cultivar Adriana and the DH line SGEDH13, using microspore culture. Adriana is a German winter rapeseed line cultivar and is among the canola cultivars analyzed by Dimov and Möllers (2010), characterized by high oil, reduced palmitic acid (16:0) and high oleic acid content (18:1). SGEDH13 is a DH line derived from the cross between DH line SGDH14 (Zhao et al. 2005) and inbred line 617 of the German winter rapeseed canola cultivar Express analyzed by Behnke et al. (2018). SGEDH13 is characterized by high oil content, normal palmitic acid (16:0) and medium erucic acid (22:1) contents caused by the presence of only one erucic acid gene. The DH population was termed ASG, an acronym derived from the letters from both parents Adriana and SGEDH13. The second DH population consisted of 95 F1 derived doubled haploid lines derived by microspore culture from a cross between Adriana and Zheyou 50; henceforth referred to as AZH DH population. Zheyou 50 is a semi-winter cultivar from China characterized by high oil and normal oleic acid (18:1) and palmitic acid (16:0) contents. Both DH populations were developed at the Division of Crop Plant Genetics, Georg-August University, Göttingen, Germany.
Field experiments
The ASG DH lines and the parents were tested in three growing seasons (2015/16, 2016/17, and 2017/18) in five field environments located in north-western Germany and Poland. The AZH DH population was evaluated in four consecutive seasons in one environment in north-western Germany. The field experiments were conducted in small plots as a Randomized Complete Block design without replication. Each genotype was sown with 100 seeds in a row of 5 m length; distance between the rows was 75–90 cm. At maturity, open pollinated seeds were bulk harvested from each genotype from the terminal raceme and three upper most primary branches of ten healthy plants. The harvested seeds were de-husked and cleaned and stored at room temperature for seed quality trait analysis using gas chromatography (GC) and near-infrared reflectance spectroscopy (NIRS).
Measurement of fatty acid composition using gas chromatography
The fatty acid composition was analyzed using gas chromatography as described by Rücker and Röbbelen (1996) with a few modifications. About 200 mg of seeds per genotype were weighted into a clean propylene tube and a small stainless-steel rod was added to the seeds in the tube. 1000 µl of Na-methylate in methanol (0.5 mol/l; 9 ml 5.56 mol sodium methylate in 1 l methanol -71 FLUKA 71748- + 10 ml isooctane ad 100 ml with methanol p.a., + 100 µl 1% bromocresol green in methanol) was dispensed into each tube with the seeds and tightly closed with screw cap. The sample was then ground for 1 min using a custom-built Wartburg-Engine vertical shaker at the Institute of Applied Plant Nutrition, Georg-August-University Göttingen, in order to extract the oil. The ground samples were then incubated for about 10 min at room temperature. Afterwards, 300 µl each of 5% NaHSO4 and iso-octane solutions were dispensed into each tube. The tubes were then vortexed to allow a thorough mixture and extraction of fatty acid methyl esters. The tubes were then centrifuged at 1000 rpm for 10 min. Then, about 200 µl of the clear upper phase was removed using a Hamilton syringe and dispensed into septum vials and tightly closed with crimp caps. The extracted fatty acids were then stored in the refrigerator until the gas chromatograph analyzer was ready. Using an auto-sampler 2 µl of the fatty acid extract was injected into the gas chromatograph (Trace GC ultra, Thermo Electron corporation) equipped with a 25 m × 0.25 mm I.D. FFAP column (Macherey & Nagel, 0.25 µm film thickness, 210 °C, split injection ratio (1:70), Hydrogen carrier gas: 150 kPa, injection/detector: temperature 230 °C). Total analytical time for each sample was 6 min. The fatty acids were expressed as the percentage of the total fatty acid content in the seed oil. The fatty acids that were considered to calculate SFA, total fatty acid content, C16/18 (KASII) and FATB/A ratio are listed in Table 1.
Measurement of oil and ADL content using near infrared reflectance spectroscopy (NIRS)
Seed oil content was determined by scanning 2–3 g of seed samples with a NIRS monochromator as described in Behnke et al. (2018). Seed oil content was expressed at 9% moisture content. The acid detergent lignin (ADL) content was determined using the calibration equation developed by Dimov et al. (2012).
Correction for the effect of erucic acid on oil and palmitic acid
ASG DH population segregated for erucic acid with 102 and 68 genotypes with high and low erucic acid content (< 2%), respectively. The expression of the erucic acid alleles has pleiotropic effect on contents of oil and palmitic acid. The corrected values of these traits were determined using the regression method described in Behnke et al. (2018). The corrected oil and palmitic acid content was estimated by fitting a linear regression as follows: Oil content corrected for Erucic acid (Oilc) = NIRS oil content − (slope of the linear regression between NIRS oil content × GC erucic acid content) and for the palmitic acid: Palmitic acid corrected for erucic acid (16:0c) = Palmitic acid content − (slope of linear regression between palmitic and erucic acid × GC erucic acid content (Fig. S1).
Statistical analysis
Analysis of Variance (ANOVA) was performed for the data using PLABSTAT version 3A software (Utz 2011), considering location – year as an environment using the following simple linear model:
where Yij is the trait value of ith genotype in jth environment and µ is the overall mean, gi is the effect of the ith genotype (i = 1,2…), while ej is the effect of j environment and geij is the interaction between ith genotype and jth environment and the random error. Both genotype and environment were considered as random factors. Broad sense heritability (h2) was calculated for each trait using \(h^{2} = \frac{{\sigma_{g}^{2} }}{{\left( {\sigma_{g}^{2} + \frac{{\sigma_{ge}^{2} }}{E}} \right)}}\).
Where \({\upsigma }_{g}^{2}\) is the variance component for the genotype and \({\upsigma }_{ge}^{2}\) is the variance component for the genotype × environment interaction and random error, and E is the number of environments. The mean values across the environments were used to calculate the Spearman rank correlation coefficient using R 4.0.3 Package (R Core Team 2018). The normality of the distribution was tested using Shapiro test using the R default script.
DNA extraction and SNP markers
The DNA from all the DH and the parental lines was extracted at TraitGenetics GmbH (https://traitgenetics.com/) using a proprietary protocol. Genotyping of the DH populations and parental lines were performed with an Illumina Infinium Brassica 15 K SNP array chips, which comprises of 13,715 SNP markers at TraitGenetics GmbH (https://traitgenetics.com/; Clarke et al. 2016). The assay data were analyzed using Illumina’s GenomeStudio Software v2011.1 applying a proprietary cluster file.
Linkage map construction
The linkage map was constructed using ASMap package in R (Taylor and Butler 2017) based on the minimum spanning tree (MST) algorithm (Wu et al. 2008). In total 43.66% and 44.61% of the markers were polymorphic in the ASG and AZH population, respectively. Heterozygous genotype calls were manually set to missing values and markers with more than 5% missing calls were deleted. Genotypes were checked for too many double cross overs and markers which were not polymorphic between the two parents were all removed before the map construction. Co-segregating markers and those with strongly distorted segregation were also initially excluded from map construction. After the map was constructed, co-segregated and distorted markers that deviated from 1:1 ratio were pushed back to have a full map. Kosambi distance (1944) was used for the final map construction. The threshold distance between markers of 25 cM was used to cluster them into linkage groups. MST algorithm (Wu et al. 2008) implemented in R was used for marker ordering within a linkage group. Marker order and chromosome assignment was compared with the map of Clarke et al. (2016). The length covered by the linkage map in both populations was smaller compared to the length covered by the linkage map in SGDH14 × Express and Sansibar × Oase populations which are 2651 cM and 2350 cM, respectively (Behnke et al. 2018; Teh and Möllers 2016). However, maps published by Wang et al. (2015) and Zhao et al. (2012) were of similar sizes as for the present two populations.
QTL mapping
QTL was analyzed using R/qtl package in R (Broman et al. 2003; Broman and Sen 2009) using multiple interval mapping (Kao et al. 1999). At first, interval mapping was used to perform a 1-D genome scan with a single-QTL model to detect loci with marginal effects. A log of odd (LOD) scored was calculated for QTL at each test position using Haley–Knott regression model (1992), using a walking speed of 1 cM. The threshold of the significance of the LOD score (p = 0.05) was initially determined using the 50,000-permutation test to get a genome-wide LOD significance threshold. In the second step, a 2-D genome scan using a two-QTL model, testing for additive effect at a locus, while considering the second locus and epistatic effect between pairs of loci, this step also calculated the maximum LOD score for the full model, for additive model and a test of epistasis as well as the LOD scores to test for evidence of the presence of the second locus. The LOD significance threshold are also calculated using the 1000-permutation test (p = 0.05). A QTL and interaction between the QTL were considered significant when the LOD were greater than the threshold. Finally, a multiple QTL model was fitted, including all the QTL detected and the interactions (Kao et al. 1999). Having fit the multiple QTL model for a putative QTL, an additional genome-wide scan was carried out to detect other QTL, while controlling the effect of the already mapped QTL. When there were additional QTL, a new multiple QTL model was fit, incorporating the newly detected QTL. Most of the additional QTL detected had minor effects. QTL that was not significant after fitting models involving all QTL were removed. The percentage of phenotypic variance explained by individual, combined QTL and the interactions for a trait was calculated in the fitted model. The identified QTL were named according to earlier published results (Javed et al. 2016; Chen et al. 2018; Chao et al. 2022) where q relates to QTL followed by the trait and then the rapeseed chromosome number (A1–A10 and C1–C9). If more than one QTL was identified on the same chromosome for the same trait, a serial number was suffixed.
Result
Phenotypic analysis
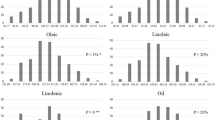
The genotypic and environmental variance components were statistically significant for all traits studied in both DH populations (Tables S5, S6, S7). The heritabilities for the saturated fatty acids in both populations ranged from 33% for behenic (22:0) to 89% for palmitic acid (16:0). Since the ASG population segregated for one erucic acid gene, the heritability was very high for oleic and erucic acid in ASG population. Despite significant differences for all traits, the means of the two populations were quite similar (Table 1). Palmitic acid was the most prominent saturated fatty acid followed by stearic acid. The distribution of all the saturated fatty acids showed transgressive segregation in both populations and were normally or near normally distributed; both populations showed bimodal distributions for ADL content (Figs. S2, S3). ASG population showed a bimodal distribution for 18:1 (Fig. S2) and for 22:1 content that were consistent with 1:1 segregation hypothesis in chi square tests (data not shown).
Close negative correlations between saturated fatty acids, the FATB/A ratio and the 20:1 and 22:1 content were found in the ASG population. In contrast a close positive correlation to oil content were found (Table 2). Since the presence of erucic alleles confounded traits like seed oil and 16:0 content, both traits were corrected for 22:1 value using regression method. Following correction, the value for palmitic acid content was higher and for oil content it was lower (Table 1). Through the correction, the negative correlation between 16:0 and oil content remained negative, although insignificantly. The correlation between 16:0 and 18:1 was negative (− 0.43**) among the 68 erucic acid free genotypes (Table S8). Stearic acid and SFA content were negatively correlated with oil content in the ASG population; this remained significant even after correction. Also, stearic acid and SFA contents were negatively correlated with oil content in AZH population (Table 3). The C16/18 ratio was negatively correlated with oleic acid content in both populations (Tables 2, 3), and which remained negative among the erucic acid free genotypes of the ASG population (Table S8). The C16/18 ratio was not correlated to oil content in both populations. Noteworthy, ADL content was negatively correlated to oil content in ASG and AZH population (Tables 2, 3, S8).
Linkage map
The final map covered 19 chromosomes with a distance of 1683 and 1703.2 cM and with 3763 and 5743 SNP markers for ASG and AZH population, respectively (Tables S1, S2). For QTL mapping, a framework map of 870 and 766 markers was used for ASG and AZH population, respectively (Tables S3, S4). In the ASG genetic map, 51.5% SNP markers showed significant deviation from the 1:1 segregation ratio, of which 42.4% favored Adriana alleles (Table S3). For the AZH population, 35.2% SNP markers deviated from the 1:1 segregation ratio of which 51.5% favored Adriana alleles (Table S4).
QTL analysis of the ASG population
There were total of 48 QTL mapped for different fatty acids, oil and ADL content in this population (Table 4). 7 QTL were identified for 16:0 on 7 chromosomes which cumulatively explained 74.6% of the phenotypic variance (Table 4, Fig. S4). Except for QTL q16:0-C3 and q16:0-C6, all QTL showed negative additive effects with Adriana alleles reducing 16:0 content. The position of major QTL q16:0-C3 co-located with positions of QTL q18:0-C3, q18:1-C3, and q16/18-C3 ratio with positive additive effects. QTL q16:0-C3 also co-located with QTL q22:1-C3, q20:1-C03, q18:2-C3, q18:3-C3-1, and qOil-C3 with opposite additive effects, indicating that q20:1-C03 and q22:1-C3 alleles led to a decrease in palmitic, stearic, oleic acid and derived polyunsaturated fatty acid contents, to a decrease in C16/18 ratio and to an increase in oil content. The erucic acid QTL allele q22:1-C3 of SGEDH13 also led to a decrease the FATB/A ratio. Following correction of the 22:1-effect, QTL q16:0-C3, q18:0-C3 and qOil-C3 became insignificant and the additive effect and the percentage of explained phenotypic variance of the remaining four QTL for 16:0c increased. All four remaining QTL following correction had negative additive effects with Adriana alleles leading to a reduction of 16:0 content (Fig. S4). QTL q16:0-A1 and q16:0c-A1 co-located with QTL q18:2-A1 with the same direction of the additive effect, which agrees with the positive correlation between 18:2 and 16:0. QTL q16:0c-A9 and q16:0c-C9 co-located with QTL q16/18-A9 and q16/18-C9 with the same direction of the additive effect. Following correction of the erucic acid alleles, only two QTL for oil content remained significant. Interestingly, QTL qOil-C5 and qOilc-C5 mapped at the same position as QTL qADL-C5 with opposite additive effects, indicating that a reduction in ADL content by SGEDH13 alleles led to an increase in oil content. QTL qADL-C5 mapped also at the same position as QTL q18:0-C5 with the same sign of the additive effect, which is in line with the positive correlation between these two traits (Table 2). Noteworthy, QTL qADL-C5 also mapped at the same position as QTL q18:2-C5 and q18:3-C5, but with opposite additive effects. The increase in oil, 18:2 and 18:3 content caused by QTL qADL-C5 is congruent with the negative correlation between these traits. For arachidic acid (20:0) and behenic acid (22:0) content no significant QTL were detected. The QTL qFATB/A-A9 with the Adriana allele causing an increase in the ratio co-located with QTL q16:0-A9 and q16/18-A9 with the same direction of the additive effect.
QTL analysis of the AZH population
There were total of 42 QTL mapped for different fatty acids, oil and ADL content in this population (Table 5). Two QTL for 16:0 mapped on chromosome C08 and C09 which cumulatively explained 26.5% of the phenotypic variance (Table 5, Fig. S5). QTL q16:0-C8 on C08 had negative additive effect indicating that the alleles reducing 16:0 were derived from parent Adriana while QTL q16:0-C9 showed positive additive effect. q16:0-C8 mapped at the same position as QTL q16/18-C8 with the same direction of the additive effect. The two QTL q18:0-C3 and q18:0-C9 with combined negative additive effects explained 30.76% of the phenotypic variance and did not show overlapping confidence intervals with QTL of any other trait. For Arachidic acid (20:0) content, five QTL with positive and negative additive effects together explained 60.9% of the phenotypic variation (Table 5, Fig. S5). This is in line with the transgressive segregation found in the population (c.f. Table 1). QTL q20:0-C9 co-located with QTL q22:0-C9 on C09. QTL q20:0-A2 and q20:0-C6 mapped at same positions as qFATB/A-A2 and qFATB/A-C6, respectively, with opposite direction of the additive effect. There was epistatic interaction between QTL q20:0-A2 and q20:0-A3 reducing 20:0 content by 0.04%. Three QTL for oleic acid content with positive additive effect collectively accounted for 49.6% of the phenotypic variance. The large QTL q18:1-A1 mapped very close to QTL qOil-A1 for oil content with same direction of the additive effect and with QTL q18:2-A1 and q18:3-A1 with opposite direction of additive effects. This is in line with the positive correlation between 18:1 and oil content and their negative correlation to 18:2 and 18:3 (Table 3). Altogether five QTL with additive and two QTL with epistatic effects for oil content were detected. Among these five QTL, confidence intervals of three QTL on A04, A05 and C05 overlapped with QTL for acid detergent lignin content (ADL), partly with opposite signs of the additive effect. This is in support with the transgressive segregation of ADL content in the DH population and with the negative correlation between ADL and oil content. Remarkably is the co-location of QTL qADL-C5 with QTL qOil-C5, q18:1-C5, and with q18:3-C5 with Zheyou 50 alleles leading to an increase in oil and polyunsaturated fatty acid content and a decrease in 18:1 content.
Identification of possible candidate genes
In ASG population on A09, confidence intervals of QTL q16:0-A9, q16:0c-A9 and q16/18-A9 corresponded to physical positions from 62.48 to 63.02 Mbp on ZS11 reference genome (Song et al. 2020, Fig. 1). Inspecting this physical region, the FATB homolog BnaA09G0665700ZS was identified at a 5 kbp distance from the SNP marker Bn-A09-p35485458 (Table S9, Fig. 1). The physical position of the additional QTL q16:0-C9, q16:0c-C9, q16/18-C9 on C09 ranged from 9.56 to 22.71 Mbp. Also, KASIII gene homolog BnaC09G0135500ZS and DGAT1 homolog BnaC09G0126800ZS were identified within this region. The FAD2 homolog BnaA01G0369500ZS and the GPAT5 homolog BnaA01G0373800ZS was located within the confidence interval of the minor QTL q16:0-A1 on A01 in the ASG population. On A06 in the ASG population FATB (BnaA06G0050800ZS) was found near the QTL confidence interval of QTL q16:0-A6. Furthermore, in the ASG population, the two KASII homolog copies BnaA02G0213600ZS and BnaA02G0213500ZS were located nearby QTL q16:0-A2, q16:0c-A2 and q16/18-A2. Candidate genes for QTL q18:0-C5, qOil-C5, qOilc-C5, and qADL-C5 for oil and fiber content on C05 are the KASII homolog BnaC05G0503400ZS, the GPAT5 homolog BnaC05G0482300ZS, the CCR1 homolog BnaC05G0501000ZS, the CESA3 homolog BnaC05G0493500ZS, and the PAL4 homolog BnaC05G0498400ZS.
Physical map positions of SNP markers, of QTL confidence intervals (CI, red bar) for SFA and other seed quality traits, and positions of candidate genes (Loci names in red) for ASG Population. The SNP markers in green were also identified within the same region for erucic acid (on C03) and ADL (on C05) contents by Behnke et al. (2018). (Color figure online)
In AZH population, for q16:0-C8 and QTL q16/18-C8 on C08, the FATB homolog BnaC08G0530500ZS was located at 15 kb distance to marker Bn-scaff_21269_1-p122418 (Fig. 2). Similarly, SAD homolog BnaC03G0176900 was found within the QTL q18:0-C3, located 425 kb away from marker Bn-scaff_21312_1-p767376 (Fig. 2, Table S9). Candidate genes for QTL q18:1-A1 and qOil-A1 on A01 are the FAD2 homolog BnaA01G0369500ZS, the LPAAT5 homolog BnaA01G0323800ZS and the GPAT5 homolog BnaA01G0373800ZS (Fig. 2). No candidate genes were identified for saturated fatty acids QTL on C09. On A04, the PAL2 homolog BnaA04G0070500ZS mapped within the confidence interval of QTL qADL-A4. On A05, both FAD2 (BnaA05G0427800ZS) and LPAAT5 (BnaA05G0429000ZS) homologs were identified near SNP Bn-A05-p21370435 on the peak of the QTL region for QTL qOil-A5. No oil candidate gene for the major QTL qOil-C2 on C02 in the AZH population was identified.
Physical map positions of SNP markers, QTL confidence intervals (CI, red bar) for SFA and other seed quality traits, and positions of candidate genes (Loci names in red) for AZH Population. The SNP markers in green were also identified within the same region for ADL content by Behnke et al. (2018). (Color figure online)
Discussion
To avoid pollen genotype effects (xenia) on seed quality traits in oilseed rape, one usually avoids cross-pollination by bagging inflorescences at the beginning of flowering. Bagging of inflorescences, however, is changing light quality and intensity as well as the microclimate inside the bags. This directly influences different seed quality traits. In addition, bagged plants are usually more prone to Sclerotinia stem rot disease that leads to premature senescence and seed quality changes. To avoid these kinds of negative effects in this study, open pollinated seeds were harvested from main inflorescences of healthy plants at full maturity. With the exception of the xenia effect of the erucic acid alleles on oil content and fatty acid composition, xenia effects appear to be only of minor relevance. Since the ASG population segregated for erucic acid content, a regression method was applied to correct for strong effects of the erucic acid allele on oil and palmitic acid content. As the conditional mapping approach developed by Zhu (1995) and applied by others (Zhao et al. 2005; Behnke et al. 2018; Chen et al. 2018), the regression allows for the correction of individual correlated traits.
Genetically reducing saturated fatty acid content and increasing oil content are continuous aims in oilseed rape breeding. In this study, significant and transgressive genetic variation has been found for most of the traits. Heritability values for saturated fatty acids ranged from 33% for 22:0 to 89% for 16:0 in both populations (Table S5), while for seed oil content it ranged from 74% in AZH to 93% in the ASG population. Low heritability values for 20:0 and 22:0 were caused by values of 1% and less at the detection level (Kaur et al. 2020). High heritability values for main fatty acids, oil and ADL content show that contents are generally more influenced by the genotype than by the GxE interaction (Gazave et al. 2020; Behnke et al. 2018; Teh and Möllers 2016).
The only major QTL for the prevailing saturated palmitic acid content (16:0) in the ASG population was caused by the pleiotropic effect of the erucic acid allele on chromosome C03. Results from the QTL mapping revealed that the erucic acid allele led to an increase in oil content and to a decrease in 16:0, 20:0, 22:0 content and the C16/18 ratio (Table 4, Fig. S4). Following correction of the effect of the erucic acid allele on 16:0, the percentage of explained phenotypic variance dropped from originally 75 to 45%. Obviously, there are a larger number of small, insignificant genetic loci that influence 16:0 content. This explains the transgressive segregation for 16:0 found in both populations. Similar results were described by Behnke et al. (2018) and Miao et al. (2019). In a diversity set of spring Brassica napus L. Gazave et al. (2020) reported lower values for 16:0, and for winter oilseed rape cultivars Dimov and Möllers (2010) described similar results for SFA content as in the present study.
Reducing SFA in oilseed rape would involve lowering levels of palmitic acid (Pandian et al. 2003). The negative correlation between the palmitic acid and oleic acid content for the erucic acid free genotypes of the ASG population (− 0.43**, Table S8) and the AZH population ( − 0.38**, Table 3) shows that the palmitic acid content can be reduced by breeding for higher oleic acid content in the seed oil. QTL underlying this negative correlation in ASG population are located on A09 with candidate gene FATB. The negative sign of the additive effects shows that the Adriana allele led to reduced 16:0 contents and C16/18 ratio (Table 4, Fig. S4). Earlier reports (Zhao et al. 2008; Teh and Möllers 2016) also identified QTL for 16:0 on A09 and suggested FATB gene as the underlying candidate gene. Zhao et al. (2019) mapped a QTL for increased 18:1 content on A09 at a different physical region in ZS11, that also did not affect 16:0 or 18:0 content. In the AZH population there were no such QTL on A09. However, in AZH population on C08 there were QTL for 16:0 and C16/18 ratio with the Adriana allele causing reduced contents and ratio. Again, FATB was identified as possible candidate gene on C08 (Fig. 2). Obviously, there are allelic differences in FATB thioesterases preferentially accepting 16:0-ACP or 18:0-ACP as substrates (Bonaventure et al. 2003). Overexpression of Cupea paucipetala FATB (cpFATB) in oilseed rape was found to increase 16:0 and other saturated fatty acids by four times (Nam et al. 2019), while knocking out FATB alleles by genome editing led to a significant reduction in 16:0 and 18:0 contents in soybean seeds (Ma et al. 2021). In a GWAS study Gacek et al. (2017) identified the FATB gene BnaA05g23790D as one promising candidate gene for oleic acid and linoleic acid content. However, a search for FATB genes in the Darmor-bzh reference genome revealed some confusion regarding the annotation of FATB and FATA genes (Gacek et al. 2017). Studies on expression of Brassica napus FATB and FATA in different tissues and stages of growth will help to understand the relationship between their mutation and phenotypic expression of palmitic and stearic acid contents under natural condition. Negative correlation between palmitic acid and oleic acid content was also reported in other studies in oilseed rape (Chen et al. 2018; Zhao et al. 2019). As prevailing saturated fatty acid, palmitic acid is closely positive correlated with SFA content in the complete ASG population and in the erucic acid free sub-population. There is no such correlation in the AZH population which may be explained by closer correlations between SFA and 18:0, 20:0 and 22:0. Other candidate genes that could be responsible for changes in 16:0, 18:0 and the C16/18 ratio are FATB, KASI, KASII, and KASIII. KASI is responsible for condensation cycles of 4:0-ACP to 16:0-ACP, KASII is involved in the elongation of 16:0-ACP to 18:0-ACP, and KASIII is catalyzing in the initial condensation of acetyl-CoA with malonyl-ACP (Hölzl and Dörmann 2019). No FATA candidate gene was identified in any of the QTL confidence intervals in both populations. FATA copies of Arabidopsis thaliana in rapeseed genome ZS11 are located on A07, A09, C03, C04, C06 and C07 (Song et al. 2020), but none was located within the QTL CI identified on A07 and C06 for FATB/A (Tables 4, 5). In contrast, KASII candidate locus was identified in the QTL region on A02 and KASI and KASIII candidate genes were identified in the QTL region on C09 in ASG population. Since KASIII is involved in the initial condensation reaction (Hölzl and Dörmann 2019), it appears unlikely as candidate gene for 16:0 and C16/18 ratio. In the AZH population only KASI was identified as candidate gene for QTL Arachidic-1 on A02 (Fig. 2).
Furthermore, SFA content can be reduced by selecting for higher oil content, since oil content is positively correlated with oleic acid content in the erucic acid free ASG sub-population (Table S8) and in the AZH population (Table 3). In both populations on A01 a QTL region for oleic acid conspicuously overlapped with a QTL for oil content. In this study, FAD2, LPAAT, DGAT2 and GPAAT5 were identified as candidate genes for the QTL on A01. FAD2 gene encodes for oleoyl-CoA desaturase, GPAAT5, and DGAT2 are involved in TAG synthesis taking place in the endoplasmic reticulum. Teh and Möllers (2016) mapped LPAAT and FAD2 on A01 when they analyzed Sansibar × Oase DH population. If TAG synthesis is efficient, oleic acid will be assembled more in TAG and will be less desaturated to 18:2 and 18:3, which finally leads to increased oleic acid content in the seed oil content (Möllers and Schierholt 2002; Zhao et al. 2012). No correlation between 16:0 and oil content have also been reported by Teh and Möllers (2016), whereas Möllers and Schierholt (2002) found a negative correlation in a DH population segregating for a major QTL affecting 18:1 content. Negative correlations between 16:0 and oil content were also found in two different sets of European winter oilseed rape cultivars (Dimov and Möllers 2010). Loose negative correlations between oil content and 16:0c in ASG, 16:0 in AZH population and 18:0 was not reflected by co-locating QTL in both populations. The close correlation between C16/18 ratio and 16:0c in ASG population and 16:0 in AZH population suggests little variation in the KASII activity, and hence flux from palmitic acid to stearic acid, which is evident by the lack of correlation between C16/18 ratio and oil content.
Noteworthy, 16:0, the C16/18 ratio and oil content in both populations were negatively correlated with ADL content. ADL content was positively correlated with 18:0 content; and 18:0 content was negatively correlated with oil content, because it was mainly desaturated to 18:1, which was positively correlated with oil content. Furthermore, reduced ADL content was accompanied by reduced oleic acid content and increased contents of polyunsaturated fatty acids in the AZH (Table 3) and in the erucic acid free ASG subpopulation (Table S8). This finding is opposite to the observed positive correlation between 18:1 and oil content (Möllers and Schierholt 2002). There are a number of candidate genes located within the various overlapping QTL confidence intervals on C05 in both populations. The QTL qADL-C5 for lignin content in SGEDH13 is derived from the ancestor population SGDH14 × Express 617, in which the QTL was mapped exactly at the same position on C05 (Behnke et al. 2018). Surprisingly, Zheyou 50 carries the same QTL at C05. This may be explained by the origin of both SGDH14 and Zheyou 50 from the Zhejiang Academy of Agricultural Sciences (Hangzhou, China). Since Zheyou 50 is of canola quality, it is possible that Zheyou 50 may be derived from one of the DH lines of SGDH14 × Express 617 (Behnke et al. 2018). The low ADL content of DH lines with the SGEDH13 or Zheyou 50 QTL allele at C05 can be caused by allelic differences in the PAL4, CESA3 and CRR1 candidate genes. Synthesis of cellulose is mediated by the cellulose synthase (CESA) gene family, of which CESA3 is one. Both, the Phenylalanine-Ammonium Lyase (PAL) and Cinnamoyl-CoA-Reductase (CRR) are involved in the Phenylpropanoid biosynthesis pathway (Liu et al. 2012). Hence, all three genes are candidates for reduced lignin content. Furthermore, a QTL for reduced seed hull proportion was mapped at C05 at the same position in the DH population SGDH14 × Express617 (Behnke et al. 2018). It may be that a reduced lignin content or reduced seed hull proportion has led to an increased oil content. On the other hand, it is possible that the KASII gene expression caused increased 18:0, decreased 16:0 and C16/18 ratio and higher oil content. Own, unpublished results (manuscript in preparation), has shown that in SGDH14 and SGEDH13 a homoeologous non-reciprocal translocation has occurred by which a 200 kb large fragment carrying the PAL4 gene from A05 has been transferred to C05, replacing the original C05 fragment. Gene conversion or homoeologous nonreciprocal translocations (HNRT) in rapeseed have been described by Stein et al. (2017). Again, the GPAT5 homolog (BnaC05G0482300) may be underlying the QTL for oil content. GPAT is the first transferase involved in the acylation of glycerol-3-phosphate to lysophosphatidic acid in the oil synthesis at the endoplasmic reticulum. In the AZH population there are in addition co-locating QTL for reduced ADL and increased oil content on A04 with a PAL2 candidate gene (Figs. 2, S5).
The genetic combination of QTL alleles for reduced SFA content could result in further reduced saturated fatty acid contents in canola oil. Additional genetic variation for reduced SFA content may be identified in canola genetic resources which includes oilseed rape resynthesized from its diploid progenitors Brassica rapa and Brassica oleracea (Jesske et al. 2013). In this context, the correction of the erucic acid allele xenia effects on oil and saturated fatty acid content by regression is valuable in determining the corrected saturated fatty acid content. Homoeologous nonreciprocal translocations in resynthesized rapeseed (Stein et al. 2017) may lead to a replacement of a FATB by a FATA and as a consequence to a decreased 16:0 and increased 18:1 content and probably oil content (Rodríguez-Rodríguez et al. 2014). Recently, Cargill has released specialty low saturate canola oil with 3.5–5% total saturated fatty acid content (Patent US20190174788). In conclusion, the present work has elaborated the interactions between different fatty acids and oil content in rapeseed and has identified QTL alleles at different loci that can be applied to develop new canola breeding material with reduced saturated fatty acid content.
Data availability
The datasets of the current study are available from the corresponding author on reasonable request.
References
Balalić I, Marjanović-Jeromela A, Crnobarac J, Terzić S, Radić V, Miklič V, Jovičić D (2017) Variabilty of oil and protein content in rapeseed cultivars affected by seeding date. Emir J Food Agric 29(6):404–410. https://doi.org/10.9755/ejfa.2017-01-229
Bao B, Chao H, Wang H, Zhao W, Zhang L, Raboanatahiry N, Wang X, Wang B, Jia H, Li M (2018) Stable, environmental specific and novel QTL identification as well as genetic dissection of fatty acid metabolism in Brassica napus. Front Plant Sci 9:1018. https://doi.org/10.3389/fpls.2018.01018
Beaith ME, Fletcher RS, Kott LS (2005) Reduction of saturated fats by mutagenesis and heat selection in Brassica napus L. Euphytica 144(1–2):1–9. https://doi.org/10.1007/s10681-005-4559-7
Behnke N, Suprianto E, Möllers C (2018) A major QTL on chromosome C05 significantly reduces acid detergent lignin (ADL) content and increases seed oil and protein content in oilseed rape (Brassica napus L.). Theor Appl Genet 131(11):2477–2492. https://doi.org/10.1007/s00122-018-3167-6
Bonaventure G, Salas JJ, Pollard MR, Ohlrogge JB (2003) Disruption of the FATB gene in Arabidopsis demonstrates an essential role of saturated fatty acids in plant growth. Plant Cell 15:1020–1033. https://doi.org/10.1105/tpc.008946
Bondaruk M, Johnson S, Degafu A, Boora P, Bilodeau P, Morris J, Wiehler W, Foroud N, Weselake R, Shah S (2007) Expression of a cDNA encoding palmitoyl-acyl carrier protein desaturase from cat’s claw (Doxantha unguis-cati L.) in Arabidopsis thaliana and Brassica napus leads to accumulation of unusual unsaturated fatty acids and increased stearic acid content in the seed oil. Plant Breed 126(2):186–194. https://doi.org/10.1111/j.1439-0523.2007.01316.x
Broman KW, Sen S (2009) A guide to qtl mapping with R. Springer, New York
Broman KW, Wu H, Sen Ś, Churchil GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinformatics 19(7):889–890. https://doi.org/10.1093/bioinformatics/btg112
Chao H, Guo L, Zhao W et al (2022) A major yellow-seed QTL on chromosome A09 significantly increases the oil content and reduces the fiber content of seed in Brassica napus. Theor Appl Genet 135:1293–1305. https://doi.org/10.1007/s00122-022-04031-0
Chen F, Zhang W, Yu K, Sun L, Gao J, Zhou X, Peng Q, Fu S, Hu M, Long W, Pu H, Chen S, Wang X, Zhang J (2018) Unconditional and conditional QTL analyses of seed fatty acid composition in Brassica napus L. BMC Plant Biol. https://doi.org/10.1186/s12870-018-1268-7
Clarke WE, Higgins EE, Plieske J, Wieseke R, Sidebottom C et al (2016) A high-density SNP genotyping array for Brassica napus and its ancestral diploid species based on optimised selection of single-locus markers in the allotetraploid genome. Theor Appl Genet 129(10):1887–1899. https://doi.org/10.1007/s00122-016-2746-7
Dimov Z, Möllers C (2010) Genetic variation for saturated fatty acid content in a collection of European winter oilseed rape material (Brassica napus L.). Plant Breed 129(1):82–86. https://doi.org/10.1111/j.1439-0523.2009.01652.x
Dimov Z, Suprianto E, Hermann F, Möllers C (2012) Genetic variation for seed hull and fibre content in a collection of European winter oilseed rape material (Brassica napus L.) and development of NIRS calibrations. Plant Breed 131(3):361–368. https://doi.org/10.1111/j.1439-0523.2012.01951.x
Fahy D, Scheer B, Wallis JG, Browse J (2013) Reducing saturated fatty acids in Arabidopsis seeds by expression of a Caenorhabditis elegans 16:0-specific desaturase. Plant Biotechnol J 11(4):480–489. https://doi.org/10.1111/pbi.12034
Food and Agriculture Organization of the United Nations - FAOSTAT (2020). Database—crops production. Available at https://www.fao.org/faostat/en/#data/QCL. Accessed 26 Apr 2022
Gacek K, Bayer PE, Bartkowiak-Broda I, Szala L, Bocianowski J, Edwards D, Batley J (2017) Genome-wide association study of genetic control of seed fatty acid biosynthesis in Brassica napus. Front Plant Sci 7:2062
Gazave E, Tassone EE, Baseggio M et al (2020) Genome-wide association study identifies acyl-lipid metabolism candidate genes involved in the genetic control of natural variation for seed fatty acid traits in Brassica napus L. Ind Crops Prod. https://doi.org/10.1016/j.indcrop.2019.112080
Gororo N, Salisbury P, Rebetzke G, Burton W, Bell C (2014) Genotypic variation for saturated fatty acid content of Victorian canola. In: 13th Australian Research Assembly on Brassicas—conference proceedings
Guan M, Huang X, Xiao Z, Jia L, Wang S, Zhu M, Qiao C, Wei L, Xu X, Liang Y, Wang R, Lu K, Li J, Qu C (2019) Association mapping analysis of fatty acid content in different ecotypic rapeseed using mrMLM. Front Plant Sci 9:1872. https://doi.org/10.3389/fpls.2018.01872
Hölzl G, Dörmann P (2019) Chloroplast lipids and their biosynthesis. Annu Rev Plant Biol 70:51–81. https://doi.org/10.1146/annurev-arplant-050718-100202
Javed N, Geng J, Tahir M, McVetty PBE, Li G, Duncan RW (2016) Identification of QTL influencing seed oil content, fatty acid profile and days to flowering in Brassica napus L. Euphytica 207:191–211. https://doi.org/10.1007/s10681-015-1565-2
Jesske T, Olberg B, Schierholt A, Becker HC (2013) Resynthesized lines from domesticated and wild Brassica taxa and their hybrids with B. napus L.: genetic diversity and hybrid yield. Theor Appl Genet 126(4):1053–1065. https://doi.org/10.1007/s00122-012-2036-y
Kao CH, Zeng ZB, Teasdale RD (1999) Multiple interval mapping for quantitative trait loci. Genetics 152(3):1203–1216. https://doi.org/10.1093/genetics/152.3.1203
Karim Zarhloul M, Stoll C, Lühs W et al (2006) Breeding high-stearic oilseed rape (Brassica napus) with high- and low-erucic background using optimised promoter-gene constructs. Mol Breed 18:241–251. https://doi.org/10.1007/s11032-006-9032-3
Kaur H, Wang L, Stawniak N, Sloan R, Erp H, Eastmond P, Bancroft I (2020) The Impact of reducing fatty acid desaturation on the composition and thermal stability of rapeseed oil. Plant Biotechnol J 18:983–991. https://doi.org/10.1111/pbi.13263
Knott SA, Haley CS (1992) A simple regression method for mapping quantitative trait loci in line crosses using flanking markers. Heredity 69(4):315–324
Kosambi DD (1944) The estimation of map distances from recombinations values. Ann Eugen 12:172–175. https://doi.org/10.1111/j.1469-1809.1943.tb02321.x
Li-Beisson Y, Shorrosh B, Beisson F et al (2013) Acyl-lipid metabolism. Arabidopsis Book. https://doi.org/10.1199/tab.0161
Liu L, Stein A, Wittkop B, Sarvari P, Li J et al (2012) A knockout mutation in the lignin biosynthesis gene CCR1 explains a major QTL for acid detergent lignin content in Brassica napus seeds. Theor Appl Genet 124(8):1573–1586. https://doi.org/10.1007/s00122-012-1811-0
Lu S, Aziz M, Sturtevant D, Chapman KD, Guo L (2020) Heterogeneous distribution of erucic acid in Brassica napus seeds. Front Plant Sci. https://doi.org/10.3389/fpls.2019.01744
Ma J, Sun S, Whelan J, Shou H (2021) CRISPR/Cas9-mediated knockout of GmFATB1 significantly reduced the amount of saturated fatty acids in soybean seeds. Int J Mol Sci 22(8):3877. https://doi.org/10.3390/ijms22083877
Merrill LI, Pike OA, Ogden LV, Dunn ML (2008) Oxidative stability of conventional and high-oleic vegetable oils with added antioxidants. JAOCS 85(8):771–776. https://doi.org/10.1007/s11746-008-1256-4
Miao L, Chao H, Chen L, Wang H, Zhao W, Li B, Zhang L, Li H, Wang B, Li M (2019) Stable and novel QTL identification and new insights into the genetic networks affecting seed fiber traits in Brassica napus. Theor Appl Genet 132(6):1761–1775. https://doi.org/10.1007/s00122-019-03313-4
Möllers C, Schierholt A (2002) Genetic variation of palmitate and oil content in a winter oilseed rape doubled haploid population segregating for oleate content. Crop Sci 42(2):379–384. https://doi.org/10.2135/cropsci2002.0379
Moon H, Hazebroek J, Hildebrand DF (2000) Changes in fatty acid composition in plant tissues expressing a mammalian Δ9 desaturase. Lipids 35(5):471–479. https://doi.org/10.1007/s11745-000-546-6
Nam J-W, Yeon J, Jeong J, Cho E, Kim HB, Hur Y, Lee K-R, Yi H (2019) Overexpression of Acyl-ACP thioesterases, CpFatB4 and CpFatB5, induce distinct gene expression reprogramming in developing seeds of Brassica napus. Int J Mol Sci 20(13):3334. https://doi.org/10.3390/ijms20133334
Ohlrogge J, Browse G (1995) Lipid biosynthesis. Plant Cell 7:957–970. https://doi.org/10.1105/tpc.7.7.957
Ohlrogge JB, Jaworski J (1997) Regulation of fatty acid synthesis. Annu Rev Plant Physiol Plant Mol Biol 48:109–136. https://doi.org/10.1146/annurev.arplant.48.1.109
Pandian A, Liu Q, Singh S, Salisbury P, Green A (2003) Reduction of nutritionally undesirable saturates in Brassica napus and Brassica juncea oils using post-transcriptional gene silencing. In: 13th Australian Research Assembly on Brassicas conference proceedings 89–94pp. http://www.australianoilseeds.com/__data/assets/pdf_file/0004/4558/Reduction_of_nutritionally_undesirable_saturates_in_Brassica_napus.pdf
R Core Team (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. http://www.R-project.org/. Accessed 24 Nov 2021
Rakow G, Raney JP (2005) Present status and future perspectives of breeding for seed quality in Brassica oilseed crops. pp 1–5. https://www.gcirc.org/fileadmin/documents/Proceedings/IRC2003Copen/Plant%20Breeding/Seed%20Oil%20Meal%20Quality/BO5.1.pdf
Rebetzke GJ, Pantalone VR, Burton JW, Carter TE, Wilson RF (2001) Genetic background and environmental influence palmitate content of Soybean Seed Oil. Crop Sci 41:1731–1736. https://doi.org/10.2135/cropsci2001.1731
Rodríguez-Rodríguez MF, Salas JJ, Garcés R, Martínez-Force E (2014) Acyl-ACP thioesterases from Camelina sativa: cloning, enzymatic characterization and implication in seed oil fatty acid composition. Phytochemistry 107:7–15. https://doi.org/10.1016/j.phytochem.2014.08.014
Rücker B, Röbbelen G (1996) Impact of low linolenic acid content on seed yield of winter oilseed rape (Brassica napus L.). Plant Breed 115(4):226–230. https://doi.org/10.1111/j.1439-0523.1996.tb00908.x
Schnurbusch T, Möllers C, Becker HC (2000) A mutant of Brassica napus with increased palmitic acid content. Plant Breed 119(2):141–144. https://doi.org/10.1046/j.1439-0523.2000.00481.x
Song JM, Guan Z, Hu J et al (2020) Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat Plants 6(1):34–45. https://doi.org/10.1038/s41477-019-0577-7
Stein A, Coriton O, Rousseau-Gueutin M et al (2017) Mapping of homoeologous chromosome exchanges influencing quantitative trait variation in Brassica napus. Plant Biotechnol J 15(11):1478–1489. https://doi.org/10.1111/pbi.12732
Stoll C, Lühs W, Zarhloul MK, Friedt W (2005) Genetic modification of saturated fatty acids in oilseed rape (Brassica napus L.). Eur J Lipid Sci Technol 107(4):244–248. https://doi.org/10.1002/ejlt.200590021
Sun JY, Hammerlindl J, Forseille L, Zhang H, Smith MA (2014) Simultaneous over-expressing of an acyl-ACP thioesterase (FatB) and silencing of acyl-acyl carrier protein desaturase by artificial microRNAs increases saturated fatty acid levels in Brassica napus seeds. Plant Biotechnol J 12(5):624–637. https://doi.org/10.1111/pbi.12168
Taylor JD, Butler D (2017) R package ASMap: efficient genetic linkage map construction and diagnosis. J Stat Softw 79(6):1–29. https://doi.org/10.18637/jss.v079.i06
Teh L, Möllers C (2016) Genetic variation and inheritance of phytosterol and oil content in a doubled haploid population derived from the winter oilseed rape Sansibar × Oase cross. Theor Appl Genet 129(1):181–199. https://doi.org/10.1007/s00122-015-2621-y
Utz HF (2011) PLABSTAT: a computer program for the statistical analysis of plant breeding experiments, version 3Awin of 14 June 2011. Institute of Plant Breeding, Seed Science, and Population Genetics, University of Hohenheim, Stuttgart
Vick BA, Jan CC, Miller JF (2002) Inheritance of reduced saturated fatty acid content in sunflower oil. Helia 25(36):113–122. https://doi.org/10.2298/hel0236113v
Wang X, Long Y, Yin Y, Zhang C, Gan L, Liu L, Yu L, Meng J, Li M (2015) New insights in to the genetic networks affecting seed fatty acid concentratioms in Brassica napus. BMC Plant Biol. https://doi.org/10.1186/s12870-015-0475-8
Wilson RF, Marquardt TC, Novitzky WP, Burton JW, Wilcox JR, Kinney AJ, Dewey RE (2001) Metabolic mechanisms associated with alleles governing the 16:0 concentration of soybean oil. JAOCS 78(4):335–340. https://doi.org/10.1007/s11746-001-0265-4
Wu Y, Bhat PR, Close TJ, Lonardi S (2008) Efficient and accurate construction of genetic linkage maps from the minimum spanning tree of a graph. PLoS Genet 4(10):e1000212. https://doi.org/10.1371/journal.pgen.1000212
Yan XY, Li JN, Wang R, Jin MY, Chen L, Qian W, Wang XN, Liu LZ (2011) Mapping of QTLs controlling content of fatty acid composition in rapeseed (Brassica napus). Genes Genomics 33(4):365–371. https://doi.org/10.1007/s13258-010-0149-8
Yao K, Bacchetto RG, Lockhart KM, Friesen LJ, Potts DA, Covello PS, Taylor DC (2003) Expression of the Arabidopsis ADS1 gene in Brassica juncea results in a decreased level of total saturated fatty acids. Plant Biotechnol J 1(3):221–229. https://doi.org/10.1046/j.1467-7652.2003.00021.x
Yao M, Guan M, Zhang Z, Zhang Q, Cui Y, Chen H, Liu W, Jan HU, Voss-Fels KP, Werner CR, He X, Liu Z, Guan C, Snowdon RJ, Hua W, Qian L (2020) GWAS and co-expression network combination uncovers multigenes with close linkage effects on the oleic acid content accumulation in Brassica napus. BMC Genomics. https://doi.org/10.1186/s12864-020-6711-0
Zhao J, Becker HC, Zhang D, Zhang Y, Ecke W (2005) Oil content in a European X Chinese rapeseed population: QTL with additive and epistatic effects and their genotype-environment interactions. Crop Sci 45(1):51–59. https://doi.org/10.2135/cropsci2005.0051a
Zhao J, Dimov Z, Becker HC, Ecke W, Möllers C (2008) Mapping QTL controlling fatty acid composition in a doubled haploid rapeseed population segregating for oil content. Mol Breed 21(1):115–125. https://doi.org/10.1007/s11032-007-9113-y
Zhao J, Huang J, Chen F, Xu F, Ni X, Xu H, Wang Y, Jiang C, Wang H, Xu A, Huang R, Li D, Meng J (2012) Molecular mapping of Arabidopsis thaliana lipid related orthologues genes in Brassica napus. Thor Appl Gene 124:407–421. https://doi.org/10.1007/s00122-011-1716-3
Zhao Q, Wu J, Cai G, Yang Q, Shahid M, Fan C, Zhang C, Zhou Y (2019) A novel quantitative trait locus on chromosome A9 controlling oleic acid content in Brassica napus. Plant Biotechnol J 17(12):2313–2324. https://doi.org/10.1111/pbi.13142
Zheng P, Ali Babar MD, Parthasarathy S et al (2014) A truncated FatB resulting from a single nucleotide insertion is responsible for reducing saturated fatty acids in maize seed oil. Theor Appl Genet 127(7):1537–1547. https://doi.org/10.1007/s00122-014-2317-8
Zhu J (1995) Analysis of conditional genetic effects and variance components in developmental genetics. Genetics 141(4):1633–1639
Acknowledgements
The authors gratefully acknowledge KWS SAAT SE & Co. KGaA, Limagrain GmbH and Norddeutsche Pflanzenzucht Hans-Georg Lembke KG for performing field experiments. Special thanks to Uwe Ammermann, Dietrich Kaufmann and Rosemarie Clemens for their excellent technical support. AOY PhD scholarship was funded by the Islamic development Bank (IDB) No 600031095.
Funding
Open Access funding enabled and organized by Projekt DEAL. AOY PhD scholarship was funded by the Islamic development Bank (IDB) No 600031095.
Author information
Authors and Affiliations
Contributions
CM designed the experiment and developed the DH mapping populations. AOY, JCR and CM performed the field experiments. AOY did the NIRS and gas chromatographic analysis and analyzed the data. AOY and CM wrote the manuscript and all authors agreed on the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yusuf, A.O., Richter, JC. & Möllers, C. Genetic variation and QTL analysis of saturated fatty acids in two doubled haploid populations of oilseed rape (Brassica napus L.). Euphytica 218, 88 (2022). https://doi.org/10.1007/s10681-022-03043-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-022-03043-7