Abstract
Gastric cancer (GC) is a prevalent and heterogeneous malignancy in the digestive system. Increasing studies have suggested that circular RNAs are implicated in GC pathogenesis. This study aimed to explore the biological role and underlying mechanism of circRNA zinc finger protein 131 (circZNF131) in GC. The expression pattern of circZNF131, microRNA-186-5p (miR-186-5p), and 6-phosphofructo-2-kinase/fructose-2, 6-bisphosphatase 2 (PFKFB2) mRNA in GC tissues and cells was detected by quantitative real-time polymerase chain reaction. The stability of circZNF131 was verified using ribonuclease R assay. Functional experiments were performed by colony formation assay for cloning ability analysis, transwell assay and wounding healing assay for cell metastasis, and flow cytometry for cell apoptosis. Glycolysis metabolism was investigated by determining the levels of glucose uptake and lactate production. The protein detection of apoptosis- or glycolysis-associated markers, PFKFB2, and Ki-67 was implemented by western blot or immunohistochemistry. Dual-luciferase reporter assay was conducted to identify the interaction between miR-186-5p and circZNF131 or PFKFB2. The role of circZNF131 on tumor growth in nude mice was investigated via xenograft tumor assay. Expression analysis indicated that circZNF131 was upregulated in GC tissues and cells in a stable structure. Functional analyses showed that circZNF131 knockdown suppressed GC cell colony formation ability, migration, invasion and glycolysis metabolism, and induced cell apoptosis. Mechanically, miR-186-5p was a target of circZNF131, and miR-186-5p could bind to PFKFB2. Rescue experiments presented that miR-186-5p inhibition reversed the effects of circZNF131 knockdown on GC cell growth and glycolysis, and PFKFB2 overexpression abolished the impacts of miR-186-5p restoration on GC cell progression. Moreover, circZNF131 could positively modulate PFKFB2 expression via sponging miR-186-5p. In vivo, circZNF131 knockdown hindered GC tumor growth by regulating the miR-186-5p/PFKFB2 axis. circZNF131 could exert an oncogenic role in GC malignant development through the miR-186-5p/PFKFB2 axis, which might provide novel targets for GC treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer (GC) is a familiar pernicious tumor in the digestive system with poor prognosis, which ranks third in terms of cancer-related death globally (Smyth et al. 2020; Zhang and Zhang 2017). Although lots of great advancements have been acquired in the diagnostic and therapeutic techniques, the prognosis of advanced GC patients is still poor (Yoon and Kim 2015). Thus, identifying novel therapeutic targets for GC remedy and finding the underlying mechanism of GC tumorigenicity are indispensable.
Circular RNAs (circRNAs) are a novel class of RNA transcripts with covalent closed-loop structures, and they can regulate post-transcriptional gene expression, thus, to drive tumor progression via exerting sponge effects on microRNAs (miRNAs) in cancer cells (Han et al. 2018; Hsiao et al. 2017; Qu et al. 2015). Numerous studies have illuminated that plentiful circRNAs are differently expressed and associated with the cellular processes of multiple malignancies, including GC (Fang et al. 2019; Wei et al. 2020). For instance, circ_0001829 facilitated the advancement of GC via miR-155-5p/SMAD2 axis (Niu et al. 2020). And circNHSL1 promoted GC progression via the miR-149-5p/YWHAZ pathway (Hui et al. 2020). Moreover, circRNA zinc finger protein 131 (circZNF131), also called hsa_circ_0001479, has been reported to be upregulated in the tissues and blood of GC patients (Li et al. 2018; Liu et al. 2019b). However, the detail role of circZNF131 in the pathogenesis of GC has been uncovered.
MiRNAs are a type of small non-coding RNAs containing approximate 22 nucleotides and can modulate gene expression through binding to target mRNAs in cells (Bartel 2004; Yan et al. 2020). Emerging research has presented that miRNAs had abnormal expression and are implicated in carcinogenesis and development in GC (Azarbarzin et al. 2020; Ishiguro et al. 2014). For example, miR-4636 was downregulated, and its overexpression restricted GC cell proliferation and metastasis (Tang et al. 2021). Meanwhile, miR-186-5p has been attested to be concerned with cell vicious behaviors by targeting NEK2 and serve as an anti-tumor factor in GC (Eun et al. 2018; Ouyang et al. 2019). Furthermore, Liu et al. have shown that miR-186-5p exerted a key part in cell growth and glycolysis in GC through being sponged by circ-NRIP1 (Liu et al. 2020b). Nevertheless, it was still unclear about the correlation between miR-186-5p and circZNF131, and whether miR-186-5p was involved in the regulatory pathway of circZNF131 in GC advancement.
Herein, we not only inspected the expression profile of circZNF131 in GC, but also explored the functions of circZNF131 in the development of GC both in vitro and in vivo. Furthermore, we provided a potential mechanism of circZNF131 action in GC associated with miR-186-5p and 6-phosphofructo-2-kinase/fructose-2, 6-bisphosphatase 2 (PFKFB2), aiming to offer feasible strategy for GC therapy.
Materials and Methods
Patient Specimens
The tumor specimens (n = 35) and surrounding para-carcinoma specimens (n = 35) were collected from GC patients undergoing radical resection at Jinan Central Hospital Affiliated to Shandong First Medical University. All participators provided the written informed consent before the operation. This study got the authorization from the Ethics Committee of Jinan Central Hospital Affiliated to Shandong First Medical University.
Cell Culture
Human normal gastric epithelial cell line (GES-1) and GC cell lines (AGS and HGC-27) were obtained from Procell Co., Ltd (Wuhan, China). GES-1 cells were trained in RPMI-1640 medium (Hyclone, Logan, UT, USA) plus 10% FBS (Hyclone) and 1% streptomycin/penicillin (P/S, Solarbio, Beijing, China). AGS cells were cultivated in Ham’s F-12 medium (Procell) plus 10% FBS and 1% P/S. HGC-27 cells were cultivated in RPMI-1640 medium plus 20% FBS and 1% P/S. These cells were cultured in a humid 37 °C environment containing 5% CO2.
Cell Transfection
Small interference RNA (siRNA) against circZNF131 (si-circZNF131), miR-186-5p mimic or inhibitor (miR-186-5p or in-miR-186-5p), and their matched controls (si-NC, miR-NC, in-miR-NC) were obtained from Ribobio (Guangzhou, China). PFKFB2 sequence was inserted into empty pcDNA3.1 vector (pcDNA; Invitrogen, Carlsbad, CA, USA) to obtain PFKFB2 overexpression vector (PFKFB2), with pcDNA used as the control. For circZNF131 stable knockdown, circZNF131 lentiviral short hairpin RNA (sh-circZNF131) and corresponding contrast (sh-NC) were also obtained from Ribobio and packaged into lentiviral vectors. AGS and HGC-27 cells were transduced with these oligonucleotides or plasmids using Lipofectamine 3000 (Invitrogen).
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
Isolation of total RNA from tissues and cells was completed by TRI Reagent (Sigma-Aldrich, St. Louis, MO, USA). RNA was reversely transcribed into cDNA employing the QuantiTect Reverse Transcription Kit (Qiagen, Hilden, Germany), and then QuantiTect SYBR Green PCR Kit (Qiagen) was employed for qRT-PCR process with primers. The relative expression was estimated via the 2−∆∆Ct method after normalization by inner reference β-actin or U6. Primers were listed: circZNF131, F: 5′-GAAGCAAAGAACTGGGAAAAA-3′, and R: 5′-CAATCGGTCGAGGATCATTT-3′; miR-186-5p, F: 5′-CCGCGCGCAAAGAATTCTCCTTT-3′, and R: 5′-ATCCAGTGCAGGGTCCGAGG-3′; PFKFB2, F: 5′-AGTCCTACGACTTCTTTCGGC-3′, and R: 5′-TCTCCTCAGTGAGATACGCCT-3′; U6, F: 5′-CTCGCTTCGGCAGCACA-3′, and R: 5′-AACGCTTCACGAATTTGCGT-3′. β-actin, F: 5′-GTCACCGGAGTCCATCACGAT-3′, and R: 5′-TCACCAACTGGGACGACATG-3′.
Ribonuclease R (RNase R) Treatment
Total RNA of AGS and HGC-27 cells was disposed with RNase R (3 U/μg; Epicentre, Madison, WI, USA) or not for 0.5 h at 37 °C. Then, the expression of circZNF131 and linear β-actin was detected by qRT-PCR assay.
Colony Formation Assay
Transfected AGS and HGC-27 cells (200 cells/well) were plated into 6-well plates and cultivated for another 14 days. The medium was replaced with fresh medium every 3 days. Subsequently, the formed colonies were immobilized with methanol (Sigma-Aldrich) and then dyed with crystal violet (0.1%, Sigma-Aldrich) for 20 min following the abstersion with PBS (Solarbio). The colonies were photographed, and a microscope (Olympus, Tokyo, Japan) was employed to count the number of colonies (> 50 cells) in each group.
Cell Migration and Invasion Assays
Transwell chambers (Corning Inc., Corning, NY, USA) pre-coated with or without Matrigel (BD Biosciences, San Diego, CA, USA) were utilized to detect cell migratory and invasive abilities, respectively. AGS and HGC-27 cells in serum-free medium were plated into the membrane of upper chamber. Meanwhile, the bottom of chamber was supplemented with culture medium plus FBS. At 24 h post-incubation, the migratory and invasive cells passed through the membranes into the lower chamber were fixated with 4% paraformaldehyde (Solarbio) and dyed with crystal violet (0.1%, Sigma-Aldrich) for 30 min. Last, the migratory and invasive cells were photographed and counted under an inverted microscope (100 × ; Olympus).
Wound Healing Assay
Cell migratory capacity was also analyzed by wound healing assay. Transfected AGS and HGC-27 cells (2 × 105) were plated into 6-well plates. Then, two scratches were gently and vertically generated in the monolayer AGS and HGC-27 cells with a pipette tip. Next, the monolayer with a wound was gently washed using PBS for twice to wipe out the floating cells. The residual cells were trained in culture medium without serum for another 24 h. The representative images of migration distance at 0 h and 24 h were, respectively, obtained with a microscope (100 × ; Olympus), and the wound closure rate (%, migration rate) was assessed using the Image J software (NIH, Bethesda, MA, USA).
Cell Apoptosis Assay
Annexin V-FITC Apoptosis Detection Kit (BD Biosciences) was applied to detect the apoptotic cells. After transfection for 72 h, 1 × 105 AGS and HGC-27 cells were re-suspended in 1 × binding buffer after harvest. Subsequently, cells were dyed with Annexin V-FITC and propidium iodide (PI) for 15 min and keep out light. The apoptotic cells were monitored by a flow cytometer (BD Biosciences).
Lactate Production and Glucose Uptake
To monitor glycolysis metabolism, the level of lactate production or glucose uptake was severally detected via the Lactate Assay Kit (Abcam, Cambridge, UK) or Glucose Uptake Assay Kit (Abcam).
Western Blot Assay
RIPA Lysis Buffer (Sigma-Aldrich) was utilized for total protein extraction. After isolated by SDS-PAGE (Solarbio), the protein was transferred on PVDF membrane. Next, the membrane was reacted with the primary antibodies after blocked by 5% skim milk overnight at 4 °C. Then, the membrane was added with HRP-conjugated secondary antibody (1:10,000; ab205718; Abcam) for 1.5 h. The protein signals were detected by enhanced chemoiluminescence (ECL; Sangon, Shanghai, China) reagents, and the band intensity was analyzed on Image J software (NIH). The primary antibodies included anti-β-actin antibody (1:200; ab115777; Abcam), anti-B-cell lymphoma-2 antibody (anti-Bcl-2; 1:1000; ab32124; Abcam), anti-Bcl-2-associated X protein antibody (anti-Bax; 1:2000; ab263897; Abcam), anti-glucose transporter-1 antibody (anti-GLUT1; 1:100,000; ab115730; Abcam), anti-hexokinase 2 antibody (anti-HK-2; 1:10,000; ab227198; Abcam), and anti-PFKFB2 antibody (1:2000; ab234865; Abcam). β-actin was deemed as the internal control in this assay.
Dual-Luciferase Reporter Assay
The luciferase report plasmids of wild-type (WT) or mutant (MUT) circZNF131 and PFKFB2 3′-untranslated region (3′UTR) were constructed based on PGL4 vector (Promega, Madison, WI, USA), naming as circZNF131 WT, circZNF131 MUT, PFKFB2 3′UTR WT, or PFKFB2 3′UTR MUT. Then, the above plasmids were co-transduced with miR-186-5p or miR-NC into AGS and HGC-27 cells. After co-transfection for 48 h, the luciferase detection was performed using Dual-Luciferase Reporter Assay System (Promega).
Xenograft Tumor Model
The BALB/c nude mice (male, 5–6 weeks old) were acquired from Vital River Laboratory Animal Technology (Beijing, China) and divided into two groups (sh-circZNF131 group and sh-NC group; n = 5 per group). 4 × 106 HGC-27 cells with transfection of sh-circZNF131 vector or sh-NC vector were subcutaneously inoculated into the flank of the mice. The tumor volume (0.5 × length × width2) was measured once a week for 4 times after cell injection. After 4 weeks, all mice were euthanatized, and xenografts were removed from mice for weight and expression detection. This animal experiment was empowered by the Institutional Animal Care and Use Committee (IACUC) of Jinan Central Hospital Affiliated to Shandong First Medical University.
Immunohistochemistry (IHC) Assay
The formalin-fixed, paraffin-embedded tissue sections from nude mice were cut into 5 µm slides. The slides were incubated with primary antibody against PFKFB2 (1:200; ab234865; Abcam) and Ki-67 (1:200, ab16667, Abcam) overnight at 4 °C after antigen retrieval, following by HRP-conjugated secondary antibody (1:20,000, ab205718, Abcam) at room temperature for 1 h. Then the slides were stained with 3, 3′-diaminobenzidine solution (DAB; Beyotime, Shanghai, China) for 10 min, and counterstained with hematoxylin (Beyotime). The representative images were obtained under a microscope (100 × ; Olympus).
Statistical Analysis
All data were presented as mean ± standard deviation (SD) with at least three duplicates. The data comparison was implemented by Student’s t test or one-way analysis of variance followed by Tukey’s post hoc test. All statistical analyses in this study were conducted using GraphPad Prism version 6.0 software (GraphPad Software, La Jolla, CA, USA). P < 0.05 was defined as statistically significant.
Results
circZNF131 was Highly Expressed in GC Tissues and Cells
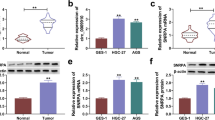
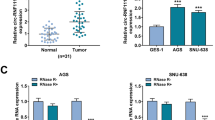
To confirm the dysregulation of circZNF131 in GC, the expression pattern of circZNF131 in collected tissues samples and cultured cells was tested by qRT-PCR analysis. As depicted in Fig. 1A, the expression of circZNF131 was remarkably increased in GC tumor tissues relative to adjacent normal tissues. Meanwhile, its level was elevated in human GC cells (AGS and HGC-27) versus that in normal GES-1 cells (Fig. 1B). Besides, the stability of circZNF131 was further investigated in AGS and HGC-27 cells, and the result indicated that circZNF131 was much more stable than linear β-actin mRNA after RNase R digestion treatment (Fig. 1C, D). The above data showed that circZNF131 was an upregulated circRNA in GC tissues and cells in a stable structure.
CircZNF131 was upregulated in GC tissues and cells. A, B The qRT-PCR was adopted to measure the expression of circZNF131 in GC tissues (n = 35) and paired normal tissues (n = 35), as well as GC cells (AGS and HGC-27) and normal GES-1 cells. C, D The expression of circZNF131 and linear β-actin mRNA was tested by qRT-PCR in AGS and HGC-27 cells after RNase R treatment. *P < 0.05
circZNF131 Knockdown Suppressed Colony Formation, Migration, Invasion, and Glycolysis Metabolism While Induced Cell Apoptosis in GC Cells
Next, the siRNA-mediated circZNF131 knockdown was implemented to explore the biological function of circZNF131 in GC cells. The qRT-PCR result exhibited that circZNF131 level was significantly decreased after si-circZNF131 transfection in comparison to si-NC transfection in AGS and HGC-27 cells (Fig. 2A), implying the successful knockdown of circZNF131. As the outcomes of circZNF131 downregulation, colony formation assay displayed that the colony forming ability of AGS and HGC-27 cells was notably restrained (Fig. 2B). Meanwhile, transwell assay manifested that cell migration and invasion were impeded by circZNF131 knockdown (Fig. 2C, D). The migratory ability of AGS and HGC-27 cells was also inspected by wound healing assay through measuring the percentage of wound closure, which was deceased after circZNF131 silence (Fig. 2E). The above-mentioned results about migration and invasion proved that cell metastasis was inhibited by circZNF131 knockdown. By performing flow cytometry analysis, cell apoptosis was promoted in si-circZNF131 group compared with si-NC group (Fig. 2F). To further verify the acceleration of apoptosis in AGS and HGC-27 cells, western blot assay was proceeded to examine the expression of apoptosis-associated markers Bcl-2 and Bax. The result showed that the level of Bcl-2 was reduced while the expression of Bax was increased after circZNF131 interference (Fig. 2G, H). Moreover, the effect of circZNF131 on glycolysis was analyzed, the data depicted that the levels of lactate production and glucose uptake were notably suppressed (Fig. 2I, J), and the expression of glycolysis-associated markers HK-2 and GLUT1 was dramatically reduced in AGS and HGC-27 cells in the presence of si-circZNF131 (Fig. 2K, L), evidencing that circZNF131 deletion impeded the glycolysis metabolism. Taken together, circZNF131 knockdown exerted inhibiting effects on cell malignant behaviors and glycolysis.
CircZNF131 knockdown suppressed cancer progression and glycolysis in GC cells. AGS and HGC-27 cells were transfected with si-circZNF131 and si-NC. A The efficiency of circZNF131 knockdown was determined by qRT-PCR. B The number of colonies in transfected cells was tested using colony formation assay. C–E Cell metastatic capacity was examined by transwell assay for migration and invasion (C and D) and wound healing assay for migration (E). F Flow cytometry was conducted for the detection of cell apoptosis rate. G, H Western blot assay was performed to assess the expression of apoptosis-associated proteins (Bcl-2 and Bax). I, J Lactate production and glucose uptake were examined using corresponding detection kits. K, L The expression of glycolysis-related protein (HK-2 and GLUT1) was measured by western blot. *P < 0.05
circZNF131 Served as a Sponge for miR-186-5p
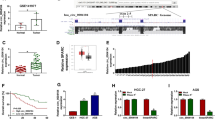
To further elucidate the mechanism of circZNF131 in GC advancement, the target miRNAs of circZNF131 were identified through starBase online database (http://starbase.sysu.edu.cn). According to the prediction results and related published literatures, 5 miRNAs (miR-186-5p, miR-448, miR-605-3p, miR-545-5p, and miR-4766-5p) that have complementary sequences with circZNF131 and have been studied in GC were selected. Through the expression analysis of above alternative 5 miRNAs in 3 paired GC tumor tissues and adjacent normal tissues, the result demonstrated that miR-186-5p, miR-448, miR-605-3p, and miR-4766-5p were all downregulated in GC tissue samples (Supplementary Fig. 1A). Therein, miR-186-5p with the most downregulation was chosen as the molecular target in our study. As depicted in Fig. 3A, the putative potential complementary sites between circZNF131 and miR-186-5p were presented. To affirm the interaction between them in GC cells, dual-luciferase reporter assay was carried out via constructing circZNF131 WT and circZNF131 MUT reporter plasmids. The overexpression efficiency of miR-186-5p in AGS and HGC-27 cells was confirmed by qRT-PCR assay (Fig. 3B). As presented in Fig. 3C, D, the relative luciferase activity was markedly repressed in AGS and HGC-27 cells after the co-transfection of miR-186-5p and circZNF131 WT but not circZNF131 MUT, verifying the interaction between circZNF131 and miR-186-5p. Contrary to circZNF131, the expression of miR-186-5p was downregulated in GC tissues and cells compared with that in normal tissues and cells, respectively (Fig. 3E, F). In addition, miR-186-5p expression was remarkably raised in both AGS and HGC-27 cells with circZNF131 knockdown (Fig. 3G). Thus, these outcomes indicated that circZNF131 was a molecular sponge for miR-186-5p in GC cells.
CircZNF131 acted as a sponge for miR-186-5p. A The binding sites between circZNF131 and miR-186-5p were predicted by starBase. B The efficiency of miR-186-5p overexpression was examined using qRT-PCR. C, D Dual-luciferase reporter assay was implemented to determine the luciferase activity in AGS and HGC-27 cells co-transfected with circZNF131 WT or circZNF131 MUT and miR-186-5p or miR-NC. E, F The expression of miR-186-5p in normal tissues, GC tissues, and cell lines (GES-1, AGS and HGC-27) was detected by qRT-PCR. G The effect of circZNF131 knockdown on the expression of miR-186-5p was analyzed by qRT-PCR. *P < 0.05
Blocking miR-186-5p Partially Reversed the Effects of circZNF131 Knockdown in GC Cells
Based on the interaction between miR-186-5p and circZNF131, the rescue experiments were performed to verify whether the function of circZNF131 in GC cells was related with miR-186-5p. The expression of miR-186-5p was enhancive in AGS and HGC-27 cells transfected with si-circZNF131, while the impact was abrogated by the co-transfection of in-miR-186-5p (Fig. 4A), hinting the high transfection efficiency of in-miR-186-5p. Except for that, cellular behaviors suggested that the inhibiting actions of circZNF131 knockdown on colony formation (Fig. 4B), migration (Fig. 4C, E), invasion (Fig. 4D), lactate production (Fig. 4I), glucose uptake (Fig. 4J), and protein expression of Bcl-2 (Fig. 4G, H), HK-2, and GLUT1 (Fig. 4K, L) as well as the enhancing effect on Bax expression (Fig. 4G, H) and cell apoptosis (Fig. 4F) of AGS and HGC-27 cells were all overturned by miR-186-5p deficiency. All these findings collectively clarified that circZNF131 played its regulatory role in GC cells via targeting miR-186-5p.
MiR-186-5p inhibition relieved the effects of circZNF131 knockdown in GC cells. All experiments were conducted in AGS and HGC-27 cells with respective transfection of si-NC, si-circZNF131, si-circZNF131+in-miR-NC, or si-circZNF131+in-miR-186-5p. A The expression of miR-186-5p in AGS and HGC-27 cells after transfection was assessed via qRT-PCR. B Examination of cell cloning ability was carried out by colony formation assay. C–E Transwell migration and invasion assays (C and D) and wound healing assay (E) were implemented to evaluate cell metastasis. F Flow cytometry was used to measure cell apoptosis. G, H The protein expression levels of Bcl-2 and Bax were detected by western blot. I–L The glycolysis metabolism was analyzed through detecting the levels of lactate production and glucose uptake using corresponding detection kits (I and J), and the expression of HK-2 and GLUT1 using western blot (K and L). *P < 0.05
PFKFB2 was a Target Gene of miR-186-5p in GC Cells
Additionally, the potential downstream target mRNAs of miR-186-5p were sought using tool Diana (http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=MicroT_CDS/index). Similar to the screening of candidate miRNAs, 5 five genes (PFKFB2, CASK, ERBB4, HMGA2, and FOXK1) that have binding sites with miR-186-5p and have been reported in GC were investigated. As expected, PFKFB2, CASK, ERBB4, and HMGA2 were all upregulated in GC tissue samples compared to that in adjacent normal tissues (Supplementary Fig. 1A). Among them, PFKFB2 with the most upregulation was chosen for further research. The special binding sites between miR-186-5p and PFKFB2 3’UTR are displayed in Fig. 5A. The dual-luciferase reporter assay further verified the putative target binding between them, as described by the decline of luciferase activity in AGS and HGC-27 cells co-transfected with PFKFB2 3′ UTR WT and miR-186-5p (Fig. 5B, C). Meanwhile, the mRNA and protein expression of PFKFB2 were increased in GC tissues compared to normal tissues (Fig. 5D, E). Conformably, PFKFB2 was upregulated in GC cells contraposed to that in normal cells (Fig. 5F). Furthermore, the expression of PFKFB2 was inhibited by miR-186-5p overexpression in AGS and HGC-27 cells (Fig. 5G), indicating the negative regulation of miR-186-5p on PFKFB2 in GC cells. Besides, PFKFB2 expression was downregulated by circZNF131 knockdown in AGS and HGC-27 cells, which was subsequently rescued by miR-186-5p inhibition (Fig. 5H). In short, these results manifested that circZNF131 positively modulated the level of PFKFB2 via targeting miR-186-5p.
PFKFB2 was a target of miR-186-5p in GC cells. A Schematic diagram showed the binding sites between miR-186-5p and PFKFB2 3’UTR. B, C The interaction between miR-186-5p and PFKFB2 was verified using dual-luciferase reporter assay. D, E The expression of PFKFB2 in GC tissues and normal tissues was detected by qRT-PCR and western blot. F The expression of PFKFB2 in GES-1, AGS, and HGC-27 cells was detected by western blot. G PFKFB2 protein expression was examined via western blot in AGS and HGC-27 cells after transfection of miR-NC or miR-186-5p. H Western blot was conducted to examine PFKFB2 protein expression after transfection of si-circZNF131, si-circZNF131+in-miR-186-5p or their matched controls. *P < 0.05
Restoring PFKFB2 Partially Abrogated the Suppressive Influences of miR-186-5p Overexpression on Cell Development and Glycolysis
For determining whether miR-186-5p exerted its anti-cancerous role in GC cells by modulating PFKFB2, AGS and HGC-27 cells were transfected with miR-186-5p alone or together with PFKFB2 overexpression vector to perform rescue experiments. As exhibited in Fig. 6A, the miR-186-5p-mediated inhibition of PFKFB2 protein expression was found to be alleviated by the addition of PFKFB2 overexpression vector, which revealed that the overexpression of PFKFB2 in GC cells was successful. Through the functional investigation, miR-186-5p overexpression was shown to evoke the repressive effects on cell clonal ability (Fig. 6B), migration (Fig. 6C, E), invasion (Fig. 6D), and Bcl-2 expression (Fig. 6G, H), together with accelerating effects on apoptosis (Fig. 6F) and Bax abundance (Fig. 6G, H), whereas all these effects were mitigated by PFKFB2 introduction. In addition, the miR-186-5p overexpression-induced inhibitory effects on lactate production (Fig. 6I), glucose uptake (Fig. 6J), and expression of HK-2 and GLUT1 (Fig. 6K, L) were counteracted by restoring PFKFB2 in AGS and HGC-27 cells. Collectively, miR-186-5p could act as a tumor inhibitor in GC by decreasing PFKFB2 expression.
PFKFB2 downregulation was accountable for miR-186-5p as a tumor inhibitor in GC cells. AGS and HGC-27 cells were transfected with miR-NC, miR-186-5p, miR-186-5p+pcDNA, or miR-186-5p+PFKFB2. A The expression of PFKFB2 protein after transfection was examined using western blot. B Colony formation assay was exploited to determine cell colony forming ability. C–E The analysis of cell metastatic ability was completed using migration or invasion detection by transwell assay (C and D) and migration detection by wound healing assay (E). F Cell apoptosis was examined via flow cytometry. G, H The protein expression of Bcl-2 and Bax was evaluated by western blot. I, J Lactate production and glucose uptake were tested using corresponding detection kits. K, L The expression of HK-2 and GLUT1 was measured using western blot. *P < 0.05
circZNF131 Downregulation Hindered GC Tumor Growth by Affecting the miR-186-5p/PFKFB2 Axis
To further analyze the role of circZNF131 in tumorigenicity of GC cells in vivo, xenograft tumor models were established in nude mice by subcutaneous injection of HGC-27 cells stably transfected with sh-circZNF131 or sh-NC. After cell injection for 28 days, it was observed that tumor volume and weight were significantly restrained in sh-circZNF131 group compared with sh-NC group (Fig. 7A, B), suggesting that circZNF131 downregulation inhibited tumor growth. Moreover, the expression levels of circZNF131 and PFKFB2 were declined, while the level of miR-186-5p was elevated in xenograft tumor tissues from the sh-circZNF131 group relative to that from sh-NC group (Fig. 7C–F). Besides, the IHC results also confirmed the downward trends of PFKFB2 protein and cell proliferation marker protein (Ki-67) (Fig. 7G). These data indicated that circZNF131 could promote GC tumor growth in vivo by mediating the miR-186-5p/PFKFB2 axis.
The role of circZNF131 in the tumorigenicity of human GC cells in vivo. A Tumor volume was measured once a week in mice after subcutaneous injection of HGC-27 cells stably transfected with sh-circZNF131 or sh-NC. B Tumors were weighed after tumor excision on the 28 days, and the representative images of tumors were presented. C, D The expression of circZNF131 and miR-186-5p in these xenograft tumor tissues was detected by qRT-PCR. E, F PFKFB2 expression in these tumor tissues was detected through performing western blot. G IHC assay was conducted to examine the protein expression of PFKFB2 and Ki-67 in tumor tissues. *P < 0.05
Discussion
Many reports have identified that circRNAs serve vital regulatory roles in cancer development and show a potential property as novel biomarkers in cancer (Bolha et al. 2017; Zhao and Shen 2017). Nevertheless, the action mechanism of circZNF131 in GC advancement remained unstated. This study mainly attested the oncogenic role of circZNF131 in GC progression, as well as the competing endogenous RNA (ceRNA) network of circZNF131/miR-186-5p/PFKFB2 axis in GC.
In recent years, circRNAs have been attested to generate their oncogenic or suppressive actions in the evolvement of human cancers through targeting different miRNAs (Panda 2018; Zhong et al. 2018). Also, many abnormally expressed circRNAs serve as pivotal regulators in the malignant behaviors of GC (Ouyang et al. 2020). For instance, circTMEM87A could facilitate cell propagation and metastasis by the miR-142-5p/ULK1 axis in GC (Wang et al. 2020). And circ-MAT2B promoted glycolysis and cell growth in GC by targeting miR-515-5p and upregulating HIF-1α expression (Liu et al. 2020a). Herein, we affirmed that circZNF131 was highly expressed in GC tissues and cells, and this was in line with anterior reports (Li et al. 2018; Liu et al. 2019b). Simultaneously, cellular assays here also revealed that circZNF131 knockdown-repressed GC cell colony formation and metastasis while enhanced cell apoptosis. Research has shown that glycolytic reaction could provide energy source for cancer cells, and inhibiting glycolysis might be a treatment countering cancer (Akram 2013; Ganapathy-Kanniappan and Geschwind 2013). Furthermore, this report disclosed that circZNF131 downregulation impeded the glycolysis process in GC cells. Moreover, animal experiments indicated that circZNF131 silence also inhibited GC tumor growth inhibited in vivo. All these findings provided sufficient evidence that circZNF131 exerted a carcinogenic role in GC. Therefore, circZNF131 inhibition was expected to be an effective anti-tumor therapy for GC.
Mounting studies have illuminated the abnormal expression of miRNAs in diversiform cancers and the implication of them in tumor carcinogenesis and advancement through serving as tumor accelerators or suppressors (Farazi et al. 2013; Hayes et al. 2014), including GC (Jiang et al. 2015). Besides, circRNAs could sponge miRNAs to interfere the targeted gene expression (Hansen et al. 2013). In this paper, it was validated that miR-186-5p was sponged by circZNF131. MiR-186-5p is testified to be a tumor-inhibiting factor in diverse cancers, such as ovarian cancer, esophageal cancer (Huang et al. 2020; Ma et al. 2020), and osteosarcoma (Ning and Bai 2020). Likewise, miR-186-5p was also certified to impede GC progression via targeting diverse genes (Cui et al. 2019; Liu et al. 2020b; Ouyang et al. 2019; Yuan et al. 2020). In keeping with above results, this report verified that miR-186-5p was downregulated in GC tissues and cells, and its overexpression exerted inhibiting effects on cell proliferation and metastasis. Meanwhile, this study reported that miR-186-5p introduction blocked glycolysis metabolism in GC cells. Additionally, we also uncovered that miR-186-5p inhibition partially overturned the repressive impacts of circZNF131 silence on GC cell malignant behaviors and glycolysis process. Therefore, above-mentioned results proposed that miR-186-5p might participate in circZNF131-mediated regulation of tumorigenesis in GC cells.
The circRNA/miRNA/mRNA molecular regulatory network has been reported to be associated with GC development (Yang et al. 2019). Here, PFKFB2 was identified to be targeted by miR-186-5p in GC cells. PFKFB2 is a key regulatory molecule of glycolysis, and has been evidenced to be concerned with glycolysis and cell proliferation in multifarious cancers, such as osteosarcoma (Pan et al. 2020) and pancreatic cancer (Ozcan et al. 2020). As well, Qu et al. found that PFKFB2 had high expression in GC, and miR-646 could restrain cell development and glycolysis via downregulating PFKFB2 expression (Qu et al. 2020). Another study illuminated that miR-613 could restrict cell growth and glycolysis metabolism through hindering PFKFB2 expression (Liu et al. 2019a). Consistently, we manifested the high level of PFKFB2 in GC tissues and cells. Furthermore, we first certified that circZNF131 positively modulated PFKFB2 expression via sponging miR-186-5p. Moreover, PFKFB2 overexpression relieved the inhibiting actions of miR-186-5p addition on cell progression and glycolysis. Besides, circZNF131 could expedite GC tumor growth in vivo via inhibiting miR-186-5p and upregulating PFKFB2 expression. Hence, the carcinogenic function of circZNF131 was accomplished by the regulation of miR-186-5p/PFKFB2 axis in GC.
Conclusion
In conclusion, this study demonstrated that circZNF131 was upregulated in GC tissues and cells. CircZNF131 enhanced tumor malignant behaviors and glycolysis through promoting the miR-186-5p-mediated PFKFB2 expression in GC. Thus, this research may elucidate a new ceRNA mechanism of circZNF131/miR-186-5p/PFKFB2 axis for understanding the malignancy of GC, and circZNF131 was identified to be a latent therapeutic target for GC treatment.
Data Availability
Please contact the correspondence author for the data request.
References
Akram M (2013) Mini-review on glycolysis and cancer. J Cancer Educ 28:454–457
Azarbarzin S, Safaralizadeh R, Khojasteh MB, Baghbanzadeh A, Baradaran B (2020) Current perspectives on the dysregulated microRNAs in gastric cancer. Mol Biol Rep 47:7253–7264
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bolha L, Ravnik-Glavac M, Glavac D (2017) Circular RNAs: biogenesis, function, and a role as possible cancer biomarkers. Int J Genom 2017:6218353
Cui N, Liu J, Xia H, Xu D (2019) LncRNA SNHG20 contributes to cell proliferation and invasion by upregulating ZFX expression sponging miR-495-3p in gastric cancer. J Cell Biochem 120:3114–3123
Eun JW, Kim HS, Shen Q, Yang HD, Kim SY, Yoon JH, Park WS, Lee JY, Nam SW (2018) MicroRNA-495-3p functions as a tumor suppressor by regulating multiple epigenetic modifiers in gastric carcinogenesis. J Pathol 244:107–119
Fang X, Wen J, Sun M, Yuan Y, Xu Q (2019) CircRNAs and its relationship with gastric cancer. J Cancer 10:6105–6113
Farazi TA, Hoell JI, Morozov P, Tuschl T (2013) MicroRNAs in human cancer. Adv Exp Med Biol 774:1–20
Ganapathy-Kanniappan S, Geschwind JF (2013) Tumor glycolysis as a target for cancer therapy: progress and prospects. Mol Cancer 12:152
Han B, Chao J, Yao H (2018) Circular RNA and its mechanisms in disease: from the bench to the clinic. Pharmacol Ther 187:31–44
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495:384–388
Hayes J, Peruzzi PP, Lawler S (2014) MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med 20:460–469
Hsiao KY, Sun HS, Tsai SJ (2017) Circular RNA—new member of noncoding RNA with novel functions. Exp Biol Med 242:1136–1141
Huang GM, Zang HL, Geng YX, Li YH (2020) LncRNA FAM83A-AS1 aggravates the malignant development of esophageal cancer by binding to miR-495-3p. Eur Rev Med Pharmacol, Sci 24:9408–9415
Hui C, Tian L, He X (2020) Circular RNA circNHSL1 contributes to gastric cancer progression through the miR-149-5p/YWHAZ axis. Cancer Manag Res 12:7117–7130
Ishiguro H, Kimura M, Takeyama H (2014) Role of microRNAs in gastric cancer. World J Gastroenterol 20:5694–5699
Jiang C, Chen X, Alattar M, Wei J, Liu H (2015) MicroRNAs in tumorigenesis, metastasis, diagnosis and prognosis of gastric cancer. Cancer Gene Ther 22:291–301
Li T, Shao Y, Fu L, Xie Y, Zhu L, Sun W, Yu R, Xiao B, Guo J (2018) Plasma circular RNA profiling of patients with gastric cancer and their droplet digital RT-PCR detection. J Mol Med 96:85–96
Liu H, Chen K, Wang L, Zeng X, Huang Z, Li M, Dong P, Chen X (2019a) miR-613 inhibits Warburg effect in gastric cancer by targeting PFKFB2. Biochem Biophys Res Commun 515:37–43
Liu J, Song S, Lin S, Zhang M, Du Y, Zhang D, Xu W, Wang H (2019) Circ-SERPINE2 promotes the development of gastric carcinoma by sponging miR-375 and modulating YWHAZ. Cell Prolif 52:e12648
Liu J, Liu H, Zeng Q, Xu P, Liu M, Yang N (2020a) Circular RNA circ-MAT2B facilitates glycolysis and growth of gastric cancer through regulating the miR-515-5p/HIF-1alpha axis. Cancer Cell Int 20:171
Liu Y, Jiang Y, Xu L, Qu C, Zhang L, Xiao X, Chen W, Li K, Liang Q, Wu H (2020b) circ-NRIP1 promotes glycolysis and tumor progression by regulating miR-186-5p/MYH9 axis in gastric cancer. Cancer Manag Res 12:5945–5956
Ma Y, Zhang D, Wu H, Li P, Zhao W, Yang X, Xing X, Li S, Li J (2020) Circular RNA PRKCI silencing represses esophageal cancer progression and elevates cell radiosensitivity through regulating the miR-186-5p/PARP9 axis. Life Sci 259:118168
Ning Y, Bai Z (2020) DSCAM-AS1 accelerates cell proliferation and migration in osteosarcoma through miR-186-5p/GPRC5A signaling. Cancer Biomark 30:29–39
Niu Q, Dong Z, Liang M, Luo Y, Lin H, Lin M, Zhong X, Yao W, Weng J, Zhou X (2020) Circular RNA hsa_circ_0001829 promotes gastric cancer progression through miR-155-5p/SMAD2 axis. J Exp Clin Cancer Res 39:280
Ouyang Y, Li Y, Huang Y, Li X, Zhu Y, Long Y, Wang Y, Guo X, Gong K (2019) CircRNA circPDSS1 promotes the gastric cancer progression by sponging miR-186-5p and modulating NEK2. J Cell Physiol 234:10458–10469
Ouyang J, Long Z, Li G (2020) Circular RNAs in gastric cancer: potential biomarkers and therapeutic targets. Biomed Res Int 2020:2790679
Ozcan SC, Sarioglu A, Altunok TH, Akkoc A, Guzel S, Guler S, Imbert-Fernandez Y, Muchut RJ, Iglesias AA, Gurpinar Y, Clem AL, Chesney JA, Yalcin A (2020) PFKFB2 regulates glycolysis and proliferation in pancreatic cancer cells. Mol Cell Biochem 470:115–129
Pan X, Li H, Tan J, Weng X, Zhou L, Weng Y, Cao X (2020) miR-1297 suppresses osteosarcoma proliferation and aerobic glycolysis by regulating PFKFB2. Onco Targets Ther 13:11265–11275
Panda AC (2018) Circular RNAs act as miRNA sponges. Adv Exp Med Biol 1087:67–79
Qu S, Yang X, Li X, Wang J, Gao Y, Shang R, Sun W, Dou K, Li H (2015) Circular RNA: a new star of noncoding RNAs. Cancer Lett 365:141–148
Qu J, Yang J, Chen M, Wei R, Tian J (2020) CircFLNA acts as a sponge of miR-646 to facilitate the proliferation, metastasis, glycolysis, and apoptosis inhibition of gastric cancer by targeting PFKFB2. Cancer Manag Res 12:8093–8103
Smyth EC, Nilsson M, Grabsch HI, van Grieken NC, Lordick F (2020) Gastric cancer. Lancet 396:635–648
Tang J, Hu Y, Zhang C, Gong C (2021) miR-4636 inhibits tumor cell proliferation, migration and invasion, and serves as a candidate clinical biomarker for gastric cancer. Oncol Lett 21:33
Wang H, Sun G, Xu P, Lv J, Zhang X, Zhang L, Wang S, Cao J, Xia Y, Xuan Z, Li B, Huang X, Jiang T, Fang L, Xu Z (2020) Circular RNA TMEM87A promotes cell proliferation and metastasis of gastric cancer by elevating ULK1 via sponging miR-142-5p. J Gastroenterol 56:125–138
Wei G, Zhu J, Hu HB, Liu JQ (2020) Circular RNAs: promising biomarkers for cancer diagnosis and prognosis. Gene 771:145365
Yan C, Kong X, Gong S, Liu F, Zhao Y (2020) Recent advances of the regulation roles of MicroRNA in glioblastoma. Int J Clin Oncol 25:1215
Yang G, Zhang Y, Yang J (2019) Identification of potentially functional CircRNA-miRNA-mRNA regulatory network in gastric carcinoma using bioinformatics analysis. Med Sci Monit 25:8777–8796
Yoon H, Kim N (2015) Diagnosis and management of high risk group for gastric cancer. Gut Liver 9:5–17
Yuan X, Yang T, Xu Y, Ou S, Shi P, Cao M, Zuo X, Liu Q, Yao J (2020) SNHG10 promotes cell proliferation and migration in gastric cancer by targeting miR-495-3p/CTNNB1 axis. Dig Dis Sci 66:2627–2636
Zhang XY, Zhang PY (2017) Gastric cancer: somatic genetics as a guide to therapy. J Med Genet 54:305–312
Zhao ZJ, Shen J (2017) Circular RNA participates in the carcinogenesis and the malignant behavior of cancer. RNA Biol 14:514–521
Zhong Y, Du Y, Yang X, Mo Y, Fan C, Xiong F, Ren D, Ye X, Li C, Wang Y, Wei F, Guo C, Wu X, Li X, Li Y, Li G, Zeng Z, Xiong W (2018) Circular RNAs function as ceRNAs to regulate and control human cancer progression. Mol Cancer 17:79
Acknowledgements
None.
Funding
None.
Author information
Authors and Affiliations
Contributions
XS was responsible for drafting the manuscript. XS, XZ, and PH contributed to the analysis and interpretation of data. XS, TJ, YQ, and JZ contributed in the data collection. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no financial conflicts of interest.
Ethical Approval
Written informed consent was obtained from patients with approval by the Institutional Review Board in Jinan Central Hospital Affiliated to Shandong First Medical University.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10528_2021_10165_MOESM1_ESM.tif
Supplementary file1 Supplementary Figure 1. The expression of candidate miRNAs and genes in GC tissues. (A and B) The expression of candidate miRNAs (miR-186-5p, miR-448, miR-605-3p, miR-545-5p, and miR-4766-5p) and genes (PFKFB2, CASK, ERBB4, HMGA2, and FOXK1) in GC tissues and adjacent normal tissues was examined by qRT-PCR assay (TIF 184 kb)
Rights and permissions
About this article
Cite this article
Shen, X., Zhu, X., Hu, P. et al. Knockdown circZNF131 Inhibits Cell Progression and Glycolysis in Gastric Cancer Through miR-186-5p/PFKFB2 Axis. Biochem Genet 60, 1567–1584 (2022). https://doi.org/10.1007/s10528-021-10165-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-021-10165-5