Abstract
Molecular monitoring of bacterial communities can explain and predict the stability of bioprocesses in varying physicochemical conditions. To study methanol-fed denitrification biofilters of municipal wastewater treatment plants, bacterial communities of two full-scale biofilters were compared through fingerprinting and sequencing of the 16S rRNA genes. Additionally, 16S rRNA gene fingerprinting was used for 10-week temporal monitoring of the bacterial community in one of the biofilters. Combining the data with previous study results, the family Methylophilaceae and genus Hyphomicrobium were determined as suitable target groups for monitoring. An increase in the relative abundance of Hyphomicrobium-related biomarkers occurred simultaneously with increases in water flow, NO −x load, and methanol addition, as well as a higher denitrification rate, although the dominating biomarkers linked to Methylophilaceae showed an opposite pattern. The results indicate that during increased loading, stability of the bioprocess is maintained by selection of more efficient denitrifier populations, and this progress can be analyzed using simple molecular fingerprinting.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Denitrification is an essential biotechnological process in municipal wastewater treatment plants (WWTPs) for reducing the nitrogen (N) load to recipient waters. This step-wise reduction of water-soluble nitrate (NO3 −) via nitrite (NO2 −) to gaseous nitric oxide (NO), nitrous oxide (N2O), and di-nitrogen (N2) is catalyzed by facultative anaerobic heterotrophic bacteria. Denitrification is a community process, as many denitrifiers perform only a portion of the reduction steps, reducing NO3 − to NO2 − or to N2O, and only some bacterial species are capable of the whole denitrification chain from NO3 − to N2 gas [8]. Due to the unfavorably low carbon-to-nitrogen (C:N) ratio of the water in many N removal systems, an additional organic C and energy source, usually methanol, is used in the process. In WWTPs, methanol-fed denitrification is often accomplished by filtration of the wastewater through a support material in biofilters [17].
The physicochemical and technical aspects of the methanol-utilizing denitrification processes have been comprehensively characterized [17, 20]. However, the optimal control and operation of the processes would also benefit greatly from microbiological data [22, 39], such as the identity and potential controlling factors of the taxonomic groups crucial for the system function, which could be used in process monitoring [22]. Methylotrophs play a key role in methanol-fed denitrification systems, both by directly utilizing methanol as an electron donor in denitrification as well as by transforming methanol into various organic extracellular compounds, which are utilized by co-occurring non-methylotrophic denitrifiers [22]. Of the known methylotrophic denitrifiers, the genus Hyphomicrobium (Alphaproteobateria) is frequently detected in methanol-fed denitrification systems [2, 6, 21, 27–29, 35, 38] and is thus considered a suitable target for monitoring methanol-fed denitrification [22]. In addition, bacteria within family Methylophilaceae (Betaproteobacteria) [10, 29, 33, 36] as well as within genera Methyloversatilis (Betaproteobacteria) [2] and Paracoccus (Alphaproteobacteria) [6, 21, 27] can also play a significant role in the process. However, most studies have been done at laboratory scale. Other than the studies of Neef et al. [27] and Lemmer et al. [21], which found Paracoccus and Hyphomicrobium to be important methylotrophs in a methanol-fed denitrifying sand filter of a WWTP, very little is known about the overall bacterial dynamics or about the identity and community dynamics of methylotrophic denitrifiers in full-scale biofilters. There are ecological differences between methylotrophs and non-methylotrophs [21]. In addition, the ecology of Hyphomicrobium differs from that of Methyloversatilis [2], Paracoccus [21], and Methylophilaceae [10]. This indicates that methylotrophs and non-methylotrophs as well as different taxonomic groups of methylotrophs respond differently to the temporal and inter-system variations in the physicochemical conditions confronted by the full-scale biofilters.
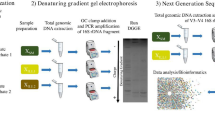
This study investigated the bacterial communities of two full-scale methanol-fed denitrifying WWTP biofilters by length heterogeneity PCR (LH-PCR) [37] and clone library and 454-pyrosequencing analysis of the 16S rRNA gene sequences. We specifically focused on the taxonomic groups of the methylotrophic bacteria that inhabited both of the biofilters as well as previously studied systems. In addition to comparing the bacterial communities of the two biofilters, we analyzed the temporal variation in the structure of the bacterial communities and linked it with the physicochemical and functional data during a 10-week follow-up period in one of the biofilters. We aimed to determine the following: (1) which methylotrophic taxonomic groups are typical for methanol-fed denitrification systems and could thus be used as target taxonomic groups for monitoring the process function in full-scale WWTP biofilters; (2) whether variations in physicochemical conditions affect the bacterial community structure; and (3) whether methylotrophs and non-methylotrophs as well as (4) different taxonomic groups of methylotrophs respond differently to these variations.
Materials and methods
Microbiological sampling
Samples were collected from the methanol-fed denitrification filters of two municipal wastewater treatment plants: the Viikinmäki wastewater treatment plant in Helsinki, Finland (WWTPA), and the Salo wastewater treatment plant in Salo, Finland (WWTPB) (Table 1). WWTPA is a large plant with one of the largest denitrification filter systems in the world, whereas WWTPB is a small-sized plant (Table 1). Methanol-fed denitrification filters have been functioning since 2004 and 2007 in WWTPA and WWTPB, respectively. In both sites, the denitrification is preceded by an aerobic stage (activated sludge) where nitrification occurs. The samples from the denitrification filter of WWTPA were collected from the same denitrification cell at 5- to 9-day intervals during a 10-week follow-up period (27 August 2008–28 October 2008). The samples from the denitrification filter of WWTPB were collected once (2 October 2008). In addition, samples from the inflow of the denitrification systems were collected once (from WWTPA 10 November 2008 and from WWTPB 2 October 2008).
The biofilter samples were taken from the backwash water channel. Backwashing consists of air-sparging and washing, which detaches biomass from the carrier material. Samples of the backwash water (1 sample per sampling date in WWTPA, 2 replicate samples in WWTPB) and polystyrene carrier material beads escaping from the WWTPB biofilter were collected into sterile 50 ml plastic containers. Bacteria in the inflow of the systems were collected by filtering 100–200 ml water using Sarstedt Filtropur S 0.2 polyethersulfone filters. The samples were stored at −20 °C before further processing within 1–2 months.
Background data and NO −x reduction
Online monitoring data of the WWTPs were used as background data in this study. For WWTPA, water flow (Wf), methanol addition rate (Metf), inflow and outflow concentrations of NO3 − + NO2 − (henceforth NO −x in and NO −x out, respectively) in the studied denitrification cell, as well as inflow temperature (T) and inflow concentrations of O2 (O2in), suspended solids (SSin), PO4 3− (PO4 3− in), total phosphorous (TPin), and outflow concentrations of SS (SSout), PO4 3− (PO4 3− out), and TP (TPout) in the whole denitrification system were measured hourly. Daily averages (for the time period 20 August 2008–31 October 2008) were then calculated. For WWTPB, daily averages (for the time period 1 September 2008–31 October 2008) for Wf and Metf along with T, NO −x in, PO4 3− in, SSin, and O2in and NO −x out, PO4 3− out, and SSout were calculated for the whole denitrification system. The NO −x load (µmol s−1) in the inflow (LNO −x in) and outflow (LNO −x out) water was calculated from Wf and NO −x in or NO −x out. Denitrification in the filters was calculated either as relative (%) or actual (µmol s−1) NO −x reduction as follows:
Denitrification in this study refers to the conversion of water-soluble NO −x into gaseous forms, but the proportions of NO, N2O, and N2 in the end product are not separated.
Molecular microbiological analyses
DNA extraction of each sample—from 10 mg of freeze-dried backwash sample material from WWTPA and WWTPB, from five frozen carrier beads from WWTPB (sample WWTPB_Car), and from the Filtropur filters containing the inflow water samples—was carried out as previously described [32].
For the LH-PCR analysis, PCR was performed using the universal bacterial primers F8 (5′-AGA GTT TGA TCM TGG CTC AG-3′) (1:4 ird700-labelled) [41] and PRUN518r (5′-ATT ACC GCG GCT GCT GG-3′) [26], with a GeneAmp PCR system 9600 (Perkin Elmer), in previously described reaction mixtures [31]. For the PCR reaction, the following program was used: an initial denaturation step at 95 °C for 5 min, 30 cycles of amplification (94 °C for 30 s, 53 °C for 1 min, 72 °C for 3 min), and final elongation at 72 °C for 15 min. The LH-PCR analysis was done as previously described [31]. The relative area (%), that is, the relative abundance of each LH-PCR peak was defined as a ratio of the total peak area (sum of the areas of all peaks) of the sample.
PCR for the clone library analyses of 16S rRNA was performed using the universal bacterial primers 27F (5′-AGAGTTTGATCMTGGCTCAG-3′) [19] and 907R (5′-CCGTCAATTCMTTTGAGTTT-3′) [13], and cloning and sequencing (Sanger sequencing) of the PCR amplicons was done as in Rissanen et al. [32]. For the clone libraries, PCR products of the samples from WWTPA on all sampling dates (WWTPA—library), PCR products of the replicate samples of backwash water (WWTPB—library), and the carrier materials of WWTPB (WWTPB_Car—library) were pooled separately.
The bacterial communities of WWTPA were also studied via 454-pyrosequencing. Equal amounts of nucleic acid extracts from each sampling date were pooled before PCR reactions, and the PCR and sequencing was performed as previously described [32].
Sequence analysis
The analysis of the clone library and 454-pyrosequencing library sequences was done as previously described [32]. Putative methylotrophic operational taxonomic units (OTUs) (97% identity threshold) were determined based on the previous literature [1, 2, 5, 10, 18, 27, 34, 35]. Clone library OTUs assigned to the methylotrophic families found from both biofilters (Methylophilaceae and Hyphomicrobiaceae) were subjected to phylogenetic tree analyses, as described previously [32]. In addition, phylogenetic classification was linked to the LH-PCR peaks in silico using the length and taxonomical data obtained in the clone library analysis.
16S rRNA gene sequences of the clone libraries were deposited into the EMBL database (accession numbers KP098594—KP098735, KP098971—KP098975, and KP098985—KP098988). The 454-pyrosequencing data were deposited into the NCBI SRA database (SRX646346).
Statistical analyses
Bray–Curtis dissimilarities among the samples were calculated from the relative abundances of the LH-PCR peaks. Temporal variations in the structure of the bacterial communities of WWTPA were then analyzed by non-metric multidimensional scaling (NMS) of the LH-PCR peak data. Changes in the WWTPA community structures were correlated with variations in the background parameters using Mantel’s test. In addition, temporal variations in the relative abundances of the LH-PCR peaks affiliated with methylotrophs and non-methylotrophs were correlated with variations in the background parameters using either Pearson correlation analysis (for normally distributed variables, normality tested using the Shapiro–Wilk test) or Spearman’s correlation analysis (for non-normally distributed variables). For background parameters, the average daily values for the time period between the two samplings were used in the correlation analyses. Temporal and inter-system variations in the community structures were also analyzed by hierarchical clustering (UPGMA linkage) using the LH-PCR data. The NMS analysis and Mantel’s test were performed in PC-ORD 6.0 [24], and cluster analysis was done using PAST version 3.09 [11]. The correlation analyses were performed in PASW 18.0 (PASW Statistics 18, Release Version 18.0.0, SPSS, Inc., 2009, Chicago).
Results
Performance of the denitrification biofilters
As is typical for WWTPs in Northern countries in autumn, Wf increased and T decreased during the study period in both filter systems (Fig. 1, Online Resource 1). NO −x in and O2in were generally higher and more variable in WWTPB (NO −x in: 700–2900 µmol/L; O2in: 1–215 µmol/L) than in WWTPA (NO −x in: 500–1000 µmol/L; O2in: 40–110 µmol/L). In addition, NO −x in decreased in WWTPB and O2in in WWTPA during the study period (Fig. 1, Online Resource 1). The higher NO −x in in WWTPB compared to WWTPA could be due to possible differences in the total N concentrations feeding the WWTPs, the nitrification efficiency between WWTPA and WWTPB, or the lack of a pre-denitrification system in WWTPB (Table 1). In the filters, Metf is controlled by a feedback loop that controls the NO3-N concentration inside the filter cells [7]. As a result, Metf followed LNO −x in tightly, and they both controlled the actual NO −x reduction rate (µmol/s) in the systems (Fig. 1, Online Resource 1). This kept the C:N ratio in the inflow (Metf:LNO −x inflow ratio), as well as the relative NO −x reduction and the NO −x out concentration, relatively stable in both systems. However, the relative NO −x reduction and NO −x out concentration were higher and lower, respectively, and temporally more stable, and Metf:LNO −x inflow was lower in WWTPA (Metf:LNO −x inflow ratio: 0.90–1.13; relative NO −x reduction: 82–93%; NO −x out: 66–99 µmol/L) than in WWTPB (Metf:LNO −x inflow ratio: 0.98–1.18; relative NO −x reduction: 64–90%; NO −x out: 128–870 µmol/L, when the exceptional values of 25 October were excluded) (Fig. 1, Online Resource 1). When estimated per carrier volume, the load of NO −x , O2 and methanol feeding as well as the actual NO −x reduction rate were on average lower in WWTPA (NO −x : 570 µmol/m3/s; O2: 50 µmol/m3/s; methanol: 590 µmol/m3/s; actual NO −x reduction: 510 µmol/m3/s) than in WWTPB (NO −x : 890 µmol/m3/s; O2: 60 µmol/m3/s; methanol: 930 µmol/m3/s; actual NO −x reduction: 730 µmol/m3/s). The higher O2 load increases the requirement for electron donors for O2 reduction (to allow anaerobic conditions for denitrification), which explains the higher Metf:LNO −x inflow ratio in WWTPB than in WWTPA. Furthermore, the average surface load was higher and the average hydraulic retention time (HRT) lower in the biofilter of WWTPA (Table 1).
NO −x reduction, operating conditions, and microbial community dynamics in the denitrification filter of WWTPA (the 10-week follow-up period of microbial communities [27 August 2008–28 October 2008] is framed). a Temperature and the concentration of NO −x and O2 in the inflow, concentration of NO −x in the outflow, and the relative NO −x reduction. b NO −x load in the inflow and outflow, actual NO −x reduction rate, water flow, methanol addition rate, and methanol:NO −x ratio in the inflow. c Results of non-metric multidimensional scaling analysis of LH-PCR peak abundance data (1. axis shown, explaining 90% of the variability in community structure) and relative abundance of methylotrophs, Hyphomicrobium (peak 466 bp) and Methylophilaceae (peak 521 bp), as well as their sum as a biomarker of methylotrophs and the relative abundance of non-methylotrophs (sum of all peaks except 466, 521 and 524 bp) based on the LH-PCR peak data
Differences in the bacterial community structures between the biofilters
Based on the UPGMA clustering of the LH-PCR data, conditions within the biofilters shaped the original bacterial communities (communities of the inflow water) in both WWTPA and WWTPB (Online Resource 2, 3). The bacterial communities of the WWTPA and WWTPB samples clustered separately (Table 2; Online Resource 2, 3), except for the carrier material of WWTPB, which more resembled the backwash water of WWTPA than that of WWTPB (Table 2, Online Resource 2).
Samples of the sheared biomass in the backwash water were used in comparing the methylotrophic communities between WWTPA and WWTPB. The relative abundance of putative methylotrophs was much higher in WWTPB than in WWTPA (Table 2). Methylophilaceae and Hyphomicrobiaceae were the dominant methylotrophic families that were found in both biofilters, whereas Paracoccus (Rhodobacteraceae) and Methyloversatilis (Rhodocyclaceae) were found only in WWTPB (Table 2; Figs. 2, 3). According to the clone library analyses, Hyphomicrobiaceae had a much higher relative abundance in WWTPA than in WWTPB, whereas the opposite was observed for Methylophilaceae (Table 2). In contrast to the backwash sample, the carrier material of WWTPB did not harbor Paracoccus or Methyloversatilis, but rather Bradyrhizobium. The carrier material of WWTPB also had a higher and lower relative abundance of Hyphomicrobiacea and Methylophilacea, respectively, than the backwash material of WWTPB (Table 2).
Phylogenetic tree (neighbor joining method) of the 16S rRNA gene clone libraries of the Hyphomicrobiaceae assigned operational taxonomic units (OTUs) (at 97% sequence similarity) in the studied denitrification filters. Hyphomicrobium clusters were previously defined by Rainey et al. [30]. The numbers in brackets after the OTU number indicate the number of sequences within that OTU. The numbers at the nodes indicate the percentages of occurrence in 1000 bootstrapped trees (bootstrap values >50% are shown)
Phylogenetic tree (neighbor joining method) of the 16S rRNA gene clone libraries of the Methylophilaceae assigned OTUs. Methylophilaceae clusters were defined in this study (see tree details in the legend of Fig. 2)
Hyphomicrobiaceae was represented by only 2 OTUs in the clone libraries. These OTUs belonged to Hyphomicrobium cluster II [30] (Table 2; Fig. 2). OTU 16 was shared between WWTPA and WWTPB. The other OTU, OTU 22, likely representing a different Hyphomicrobium species, was only found in the carrier material of WWTPB (Fig. 2), where it was more abundant than OTU 16. 454-pyrosequencing had a lower resolution for detecting Hyphomicrobiaceae than the clone library analysis (Table 2), but it showed 7 Hyphomicrobiaceae OTUs in WWTPA, of which the dominant one, harboring almost all (91%) of the Hyphomicrobiaceae sequences in the 454-pyrosequencing library, was identical to OTU 16 in the clone library (Fig. 2).
Bacteria within Methylophilaceae, consisting of ten OTUs, were divided into four groups (Table 2; Fig. 3). Three of the groups, that is, clusters Met I, Methylotenera I, and Methylotenera II (clustering according to this study), included eight OTUs covering the majority of the observed Methylophilaceae sequences (Table 2; Fig. 3). Methylotenera I and Methylotenera II were closely related to the cultured members of the genus Methylotenera (Fig. 3), while the Met I cluster probably represented a novel species of Methylotenera with no cultured representatives so far. The fourth group included two rare OTUs that were not closely affiliated to known Methylophilaceae genera (Table 2; Fig. 3). Strikingly, despite the high relative abundance of Methylophilaceae, the backwash material of WWTPB had only one Methylophilaceae OTU, and it belonged to cluster Met I (Fig. 3). Cluster Met I was also the most abundant group of Methylophilacea in the carrier material of WWTPB, whereas it was absent in WWTPA (Table 2; Fig. 3). In contrast, clusters Methylotenera I and II were found in the backwash material of WWTPA and also in the carrier material of WWTPB (Table 2; Fig. 3). Methylotenera I was much more abundant than Methylotenera II in WWTPA, but it was only slightly less abundant than Methylotenera II in the carrier material of WWTPB (Table 2). 454-pyrosequencing found 6 Methylophilaceae OTUs in WWTPA, of which the dominant OTU, harboring almost all (99%) of the Methylophilaceae sequences in the 454-pyrosequencing library, was identical to Methylotenera OTU 6 (within cluster Methylotenera I) in the clone library analyses (Fig. 3). Furthermore, 454-pyrosequencing of 16S rRNA gene amplicons revealed a marginal abundance (≤1% of 16S rRNA sequences) of the following putative methylotrophs: Methylocystaceae, Methylococcaceae, Acinetobacter, and Flavobacterium in WWTPA (Table 2). 454-pyrosequencing also resulted in a higher proportion of unclassified bacterial sequences than the clone library analysis (Table 2).
The abundant non-methylotrophic bacterial groups (≥5% of 16S rRNA sequences in any of the libraries) included Acidobacteria, Actinobacteria, Bacteroidetes (other than Flavobacterium), Chloroflexi, Comamonadaceae, Deltaproteobacteria, Planctomycetes, and Rhodocyclaceae (other than Methyloversatilis) (Table 2).
Temporal variation in the bacterial community in the WWTPA biofilter
The bacterial community structure changed over time (non-metric multidimensional scaling analysis, Fig. 1), along with a temporal change in several operational parameters (Fig. 1). The fluctuations in the community structure were correlated with variations in Wf (Mantel’s test, r = 0.36, p < 0.05, n = 10), LNO −x in (r = 0.61, p < 0.05, n = 10), Metf (r = 0.55, p < 0.05, n = 10), and T (r = 0.59, p < 0.05, n = 10). In addition, the community structure correlated with the actual NO −x reduction rate (r = 0.62, p < 0.05).
To study the variation of the methylotrophic taxa in WWTPA, the phylogenetic classification was linked to the LH-PCR peaks in silico using the length and taxonomic data obtained from the clone library analyses (Online Resource 3). All the clone library sequences with a size of 466 bp in the area amplifiable by LH-PCR primers belonged to OTU 16 within the Hyphomicrobium II cluster, and all the sequences of genus Hyphomicrobium had the size of this peak (see Fig. 2). The sequences assigned to Methylophilaceae were found only within peaks 521 bp and 524 bp, and they dominated only within peak 521 bp (73%), which was also the largest peak in the LH-PCR profiles of WWTPA (Online Resource 3). Peak 521 bp consisted mostly of OTU 6 within the Methylotenera I cluster (67%) and for the smaller part of the unclassified Methylophilaceae OTU 137 (6%) (see Fig. 3), Burkholderiales (13%), Rhodocyclales (7%, not Methyloversatilis), and Bacteroidetes (7%, not Flavobacterium). Thus, LH-PCR peaks 466 and 521 bp were chosen as biomarkers of Hyphomicrobium and Methylophilaceae, respectively. Furthermore, the sum of LH-PCR peaks 466 and 521 bp was used as a general biomarker for methylotrophs, whereas the sum of all peaks excluding methylotrophic peaks 466, 521, and 524 bp (see above) was used as a biomarker for non-methylotrophs.
During the study period, there was a negative correlation between the relative abundances of Hyphomicrobium and Methylophilaceae (r = −0.91, p < 0.001) (Fig. 4). The relative abundance of Hyphomicrobium increased as Metf, Wf, and LNO −x in increased (Metf: r = 0.74, p < 0.05; Wf, ρ = 0.67, p < 0.05; LNO −x in, r = 0.80, p < 0.05, n = 10) (Figs. 1, 4), while the opposite took place with Methylophilaceae (Metf: r = −0.74, p < 0.05; Wf, ρ = −0.66, p < 0.05; LNO −x in, r = −0.77, p < 0.05, n = 10). The relative abundance of Methylophilaceae also increased as T increased (r = 0.67, p < 0.05, n = 10), while there was no correlation between T and Hyphomicrobium (r = −0.62, p = 0.06, n = 10) (Fig. 4). The relative abundance of total methylotrophs decreased as Metf and LNO −x in increased (Metf: r = −0.73, p < 0.05; LNO −x in, r = −0.77, p < 0.05, n = 10) and T decreased (r = 0.67, p < 0.05), while the opposite took place with non-methylotrophs (Metf: r = 0.79, p < 0.05; LNO −x in: r = 0.80, p < 0.05; T: r = −0.72, p < 0.05, n = 10) (Fig. 4). An increase in the relative abundance of Hyphomicrobium (r = 0.77, p < 0.05, n = 10) and non-methylotrophs (r = 0.80, p < 0.05, n = 10) and a decrease in Methylophilaceae (r = −0.77, p < 0.05, n = 10) and total methylotrophs (r = −0.76, p < 0.05, n = 10) also occurred with the increase in the actual NO −x reduction rate (Figs. 1, 4).
Correlation between the relative abundance of the peaks assigned to a Hyphomicrobium (peak 466 bp) and Methylophilaceae (peak 521 bp) and b methylotrophs (sum of 466 and 521 bp) and non-methylotrophs (sum of all peaks except 466, 521, and 524 bp) in the length heterogeneity-PCR (LH-PCR) analysis of WWTPA samples during the 10-week monitoring period. Physicochemical and process variables correlating (p < 0.05) with the relative abundance of both groups in either a or b; the sign of the correlations are shown with black-colored text and dashed-line arrow, whereas those correlating only with one of the groups are shown as gray-colored text and dashed-line arrow
Discussion
Bacteria belonging to genus Hyphomicrobium inhabited both WWTP biofilters. This agrees with the results from many previous studies (e.g. [2, 27, 29]) indicating that bacteria in Hyphomicrobium are crucial for the function of methanol-utilizing denitrification processes. Moreover, this further confirms that Hyphomicrobium is a suitable target genus for monitoring denitrification in full-scale methanol-fed WWTP biofilters [23].
Methylophilaceae were also important components of the bacterial communities in both biofilters, which is in accordance with results from laboratory-scale methanol-fed denitrification systems [10, 29, 36]. In addition, Methylophilaceae were abundant in pilot-scale activated sludge reactors during a period of high nitrate and methanol concentration [12] and in a full-scale, methanol-fed, activated sludge plant [33]. Since the first indication of the methylotrophic denitrification capability of Methylophilaceae was shown in 2004 [10], Methylophilaceae were not even targeted [Methylophilaceae-specific fluorescence in situ hybridized (FISH) probes were not used] in a previous study of a full-scale WWTP biofilter (a sand filter) [21, 27]. However, the addition of methanol led to enrichment of Betaproteobacteria in the biofilter [27], and it can be suggested that this was at least partially due to the growth of Methylophilaceae. Together, these results suggest that, besides Hyphomicrobium, bacteria belonging to Methylophilaceae are crucial for the function of methanol-utilizing denitrification processes. Furthermore, the results from the WWTPA and WWTPB biofilters and methanol-affected activated sludge systems [12, 33] indicate that, of the family Methylophilaceae, the bacteria belonging to genus Methylotenera, which includes species that couple methylotrophy to denitrification [16], can be important components of methanol-fed denitrification systems. In addition, many yet uncultivated species of Methylotenera probably also exist, as exemplified by the abundant Cluster Met I detected in WWTPB. However, Methylobacillus [29, 36] and Methylophilus [29] as well as another, thus far uncultivated Methylophilaceae genus [10] (Fig. 3) were determined to be the primary methanol-consuming Methylophilaceae in previous laboratory-scale studies of methanol-utilizing denitrification. Thus, Methylophilaceae can be used as a target family for monitoring denitrification in full-scale methanol-fed WWTP biofilters, although there can be variation in the genera and species mediating the process between different systems.
The considerable differences between the bacterial communities within the biofilters and in the water feeding the biofilters indicate that prevailing physicochemical conditions are very strong determinants of the bacterial community structure inside the biofilters. A change in the primary C source from multicarbon sources (present in the feed water) to methanol can exert an especially strong structuring force on the bacterial communities [36]. We suggest that differences in the biofilter communities between WWTPA and WWTPB are mostly due to variations in physicochemical conditions, but the effect of variations in the original inocula (bacteria from preceding activated sludge stage) cannot be completely ruled out.
Many possible physicochemical factors might have affected the differences between the filters. The higher abundance of methylotrophs in WWTPB than in WWTPA could be explained by the higher availability of methanol (higher Metf:LNO −x inflow and higher Metf estimated per carrier volume). As a higher O2 load caused the higher Metf:LNO −x inflow in WWTPB, the higher abundance of methylotrophs could be due to a higher contribution of aerobic methylotrophs and methylotrophs performing aerobic denitrification in WWTPB. Analogous to aerobic methane oxidation coupled with denitrification (AME-D) [43], these methylotrophs could have contributed to the overall denitrification performance by consuming O2 and by converting methanol to substrates utilizable by non-methylotrophic denitrifiers. However, higher HRT and lower surface load, which act through decreasing the input of bacteria (mostly non-methylotrophic) from the preceding activated sludge stage and through lowering the physical force exerted on the carrier material, might have also favored the growth and development of methylotrophs over non-methylotrophs in WWTPB.
Capable of aerobic denitrification, Paracoccus tolerates O2 better than Hyphomicrobium, which thrive in anoxic conditions, and thus Paracoccus were favored in the surface zones of the biofilm in a previously studied full-scale biofilter (a sand filter) [21]. This is in accordance with our results on the higher and lower relative abundance of Paracoccus and Hyphomicrobium, respectively, in the sheared biomass of the backwash water (representing more aerobic surface biofilm) than in the carrier material (representing deeper anoxic biofilm) in WWTPB. Similarly, the lower O2 load (as expressed per carrier volume) could explain the higher abundance of Hyphomicrobium and the absence of Paracoccus in WWTPA. Since some Methylotenera strains are aerobic [3, 14] or perform aerobic denitrification [25], the higher abundance of Methylophilaceae in the sheared biomass than in the carrier material could also be due to differences in O2 availability. However, it could also be due to differences in NO −x and methanol availability, which is expected to be higher in the biofilm surface. The results indicate that Cluster Met I, which was the sole Methylophilaceae group in the sheared biomass of WWTPB, was especially favored by the higher availability of O2, NO −x , and/or methanol. Therefore, the lower O2, NO −x , and methanol load (as expressed per carrier volume) could both explain the lower abundance of Methylophilacea and the absence of Cluster Met I in WWTPA. However, as discussed below for the temporal variation in the bacterial community in WWTPA, the lower abundance of Methylophilacea and higher abundance of Hyphomicrobium in WWTPA could also be due to a lower HRT and higher surface load, which could favor Hyphomicrobium over Methylophilacea. In addition, as there are variations in the response of different Hyphomicrobium species to varying NO3 − [23], the differential distribution of the two Hyphomicrobium species (OTUs) between the sheared biomass and carrier material in WWTPB was probably due to the decreased availability of NO3 − deeper in the biofilm. Finally, Methyloversatilis and Paracoccus gain an ecological advantage by shifting between using C1-carbon and multicarbon substrates [2, 4, 34]. Their presence in WWTPB, but not in WWTPA, might also reflect higher temporal variation in the availability of methanol or higher and temporally more variable availability of other C sources (present in feed water or produced from methanol) in WWTPB.
In accordance with the results from the comparison of the biofilters, many possible physicochemical factors might have affected the temporal variation in the bacterial community structure within the WWTPA biofilter. The overall bacterial community structure changed due to variations in the availability of electron acceptors (NO −x ) and donors (methanol) as well as in temperature, which have also previously been shown to affect denitrifying communities [9, 40]. In addition, changes in the water flow, which act through changing the HRT and surface load, possibly affected the community structure. However, due to the covariation among these factors (Fig. 1) and the relatively small sample size, it is impossible to specify the effects of each variable. In contrast to explaining differences between the biofilters, the availability of O2 [the O2 concentration and the O2 flow (µmol s−1) (data not shown)] did not affect the temporal variation in the community structure in WWTPA.
Assigning taxonomies to the LH-PCR peaks allowed for analysis of the relationship between the physicochemical factors and bacterial communities at the level of major functional and methylotrophic groups. Methylotrophs and non-methylotrophs as well as the key methylotrophic groups, Methylophilaceae and Hyphomicrobium, responded differently to variations in the physicochemical factors. Since the bulk of methylotrophs consisted of Methylophilacea in every sampling occasion, the variation in the relative abundance of methylotrophs tightly followed that of Methylophilaceae.
The decrease in Methylophilaceae (and total methylotrophs) and increase in Hyphomicrobium and non-methylotrophs with increasing NO −x and methanol loads contrast with the above comparison between WWTPA and WWTPB. This discrepancy could be due to the dominant Methylophilaceae group in WWTPA, Methylotenera I, having a slower growth rate and a lesser response to increases in NO −x and methanol than the dominant group in WWTPB, Cluster Met I. However, differences in the water flow acting through changes in the HRT and surface load provide a more unifying explanation for the community variations both between the biofilters and within WWTPA. With an increased water flow (lowered HRT and increased surface load), the input of non-methylotrophic bacteria from the preceding activated sludge stage was increased, which could have lowered the relative abundance of Methylophilaceae (and total methylotrophs). Furthermore, increased physical disturbance due to increased water flow could have caused the selective removal of Methylophilaceae, which would further contribute to the decrease in methylotrophs as well as to the increase in Hyphomicrobium. Prosthecae and buds of Hyphomicrobium [42] might have provided firmer attachment to the carrier material than the flagellum and ‘prostheca-like’ structures of Methylotenera [15]. In addition, decreased temperature could have decreased the growth rate of Methylophilaceae (and total methylotrophs), which could have also contributed to the observed community variations.
Physicochemical factors can control microbial process rates both directly by affecting the short-term cell function and indirectly by affecting the microbial community structure in the longer term [40]. The correlation between the community structure and function (actual NO −x reduction rate) in the WWTPA biofilter suggests that physicochemical factors controlled the denitrification rate of the biofilter indirectly by modifying the community composition. However, this study cannot rule out the importance of direct control of physicochemical factors on cell function. The decrease in Methylophilaceae and total methylotrophs and increase in Hyphomicrobium and non-methylotrophs with an increasing actual NO −x reduction rate are surprising and contrast with the results from a laboratory reactor in which the relative abundance of Methylophilaceae increased and that of Hyphomicrobium did not change with increasing denitrification rate [10]. However, this discrepancy is probably due to differing expressions of the process rate, expressed as per biofilter or per volume of carrier material in our study and as per mass of biomass (mixed liquor volatile suspended solids [MLVSS]) in Ginige et al. [10]. Unfortunately, MLVSS was not analyzed in this study. However, the higher actual NO −x reduction rate with an increasing relative abundance of non-methylotrophs suggests that non-methylotrophs can efficiently support the N removal of methanol-fed denitrification systems, especially during periods of high N load. In those conditions, methylotrophs might have increasingly allocated more of the methanol C into extracellular substances than into biomass and thus supported the activity of non-methylotrophs.
Conclusions
Combining the results of the two WWTP biofilters with those of previous studies confirms that bacteria in genus Hyphomicrobium and family Methylophilaceae are crucial components of methanol-utilizing denitrification. Thus, Hyphomicrobium and Methylophilaceae can be used as target taxonomic groups to monitor the function of full-scale methanol-fed denitrification biofilters of WWTPs. Although Methylotenera was the major Methylophilaceae genus in the studied WWTP biofilters, other genera (Methylophilus and Methylobacillus) may be more important in other systems. There were differences in the bacterial communities between the biofilters. In addition, 10-week monitoring of one of the biofilters showed temporal variation in the bacterial community. Variation in the loads of NO −x and O2 as well as in the methanol addition rate, water flow rate (acting through changing HRT and surface load), and temperature were all potential candidates affecting the structure of the bacterial communities. Methylotrophs and non-methylotrophs as well as Hyphomicrobium and Methylophilaceae responded differently to these variations. Furthermore, the correlation of the bacterial community structure with the process function (actual NO −x reduction rate) in the temporally monitored biofilter indicates that fluctuating physicochemical conditions affected the denitrification rate indirectly by affecting the community composition. Further temporal monitoring and/or experimental studies combined with modern sophisticated culture-independent (stable isotope probing of DNA/RNA, metatranscriptomics, metagenomics) as well as culture-dependent (high-throughput culturing) techniques are needed to resolve the exact mechanisms underlying the observed relationship among the physicochemical factors, bacterial communities (methylotrophs, non-methylotrophs, Hyphomicrobium, and Methylophilaceae), and process function.
References
Bamforth CW, Quayle JR (1978) Aerobic and anaerobic growth of Paracoccus denitrificans on methanol. Arch Microbiol 119:91–97
Baytshtok V, Lu H, Park H, Kim S, Yu R, Khandran K (2009) Impact of varying electron donors on the molecular microbial ecology and biokinetics of methylotrophic denitrifying bacteria. Biotechnol Bioeng 102:1527–1536
Beck DAC, McTaggart TL, Setboonsarng U, Vorobev A, Kalyuzhnaya MG, Ivanova N, Goodwin L, Woyke T, Lidstrom ME, Chistoserdova L (2014) The expanded diversity of Methylophilaceae from Lake Washington through cultivation and genomic sequencing of novel ecotypes. PLoS One 9:e102458
Blaszczyk M (1993) Effect of medium composition on the denitrification of nitrate by Paracoccus denitrificans. Appl Environ Microbiol 59:3951–3953
Chistoserdova L, Kalyuzhnaya MG, Lidstrom ME (2009) The expanding world of methylotrophic organisms. Annu Rev Microbiol 63:477–499
Claus G, Kutzner HJ (1985) Denitrification of nitrate and nitric acid with methanol as carbon source. Appl Microbiol Biotechnol 22:378–381
Corona F, Mulas M, Haimi H, Sundell L, Heinonen M, Vahala R (2013) Monitoring nitrate concentrations in the denitrifying post-filtration unit of a municipal wastewater treatment plant. J Process Contr 23:158–170
Gentile ME, Nyman JL, Criddle CS (2007) Correlation of patterns of denitrification instability in replicated bioreactor communities with shifts in the relative abundance and the denitrification patterns of specific populations. ISME J 1:714–728
Gentile M, Yan T, Tiquia SM, Fields MW, Nyman J, Zhou J, Criddle CS (2006) Stability in a denitrifying fluidized bed reactor. Microb Ecol 52:311–321
Ginige MP, Hugenholtz P, Daims H, Wagner M, Keller J, Blackall LL (2004) Use of stable isotope-probing, full-cycle rRNA analysis, and fluorescence in situ hybridization-microautoradiography to study a methanol-fed denitrifying microbial community. Appl Environ Microbiol 70:588–596
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4(1):9
Isazadeh S, Ozcer PO, Frigon D (2014) Microbial community structure of wastewater treatment subjected to high mortality rate due to ozonation of return activated sludge. J Appl Microbiol 117:587–596
Johnson JL (1994) Similarity analysis of DNAs. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 655–682
Kalyuzhnaya MG, Beck DA, Vorobev A, Smalley N, Kunkel DD, Lidstrom ME, Chistoserdova L (2012) Novel methylotrophic isolates from lake sediment, description of Methylotenera versatilis sp. nov. and emended description of the genus Methylotenera. Int J Syst Evol Micr 62:106–111
Kalyuzhnaya MG, Boverman S, Lara JC, Lidstrom ME, Chistoserdova L (2006) Methylotenera mobilis gen. nov., sp. nov., an obligately methylamine-utilizing bacterium within the family Methylophilaceae. Int J Syst Evol Micr 56:2819–2823
Kalyuzhnaya MG, Martens-Habbena W, Wang T, Hackett M, Stolyar SM, Stahl DA, Lidstrom ME, Chistoserdova L (2009) Methylophilaceae link methanol oxidation to denitrification in freshwater lake sediment as suggested by stable isotope probing and pure culture analysis. Environ Microbiol Reports 1:385–392
Koch G, Siegrist H (1997) Denitrification with methanol in tertiary filtration. Water Res 31:3029–3038
Kolb S (2009) Aerobic methanol-oxidizing Bacteria in soil. FEMS Microbiol Lett 300:1–10
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–175
Lemmer H, Zaglauer A, Metzner G (1997) Denitrification in a methanol-fed fixed-bed reactor. Part 1: Physico-chemical and biological characterization. Water Res 31:1897–1902
Lemmer H, Zaglauer A, Neef A, Meier H, Amann R (1997) Denitrification in a methanol-fed fixed-bed reactor. Part 2: Composition and ecology of the bacterial community in the biofilms. Water Res 31:1903–1908
Lu H, Chandran K, Stensel D (2014) Microbial ecology of denitrification in biological wastewater treatment. Water Res 64:237–254
Martineau C, Mauffrey F, Villemur R (2015) Comparative analysis of denitrifying activities of Hyphomicrobium nitrativorans, Hyphomicrobium denitrificans and Hyphomicrobium zavarzinii. Appl Environ Microbiol 81:5003–5014
McCune B, Mefford MJ (2011) PC-ORD. Multivariate analysis of ecological data. Version 6. MjM Software, Gleneden Beach, Oregon, USA
Mustakhimov I, Kalyuzhnaya MG, Lidstrom ME, Chistoserdova L (2013) Insights into denitrification in Methylotenera mobilis from denitrification pathway and methanol metabolism mutants. J Bacteriol 195:2207–2211
Muyzer G, Dewaal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturating gel electrophoresis of polymerase chain reaction amplified genes coding for 16S ribosomal RNA. Appl Environ Microbiol 59:695–700
Neef A, Zaglauer A, Meier H, Amann R, Lemmer H, Schleifer K-H (1996) Population analysis in a denitrifying sand filer: Conventional and in situ identification of Paracoccus spp. in methanol-fed biofilms. Appl Environ Microbiol 62:4329–4339
Nurse GR (1980) Denitrification with methanol: microbiology and biochemistry. Water Res 14:531–537
Osaka T, Yoshie S, Tsuneda S, Hirata A, Iwami N, Inamori Y (2006) Identification of acetate- or methanol-assimilating bacteria under nitrate-reducing conditions by stable-isotope probing. Microb Ecol 52:253–266
Rainey FA, Ward-Rainey N, Gliesche CG, Stackebrandt E (1998) Phylogenetic analysis and intrageneric structure of the genus and the related genus Filomicrobium. Int J Syst Bacteriol 48:635–639
Rissanen AJ, Kurhela E, Aho T, Oittinen T, Tiirola M (2010) Storage of environmental samples for guaranteeing nucleic acid yields for molecular microbiological studies. Appl Microbiol Biotechnol 88:977–984
Rissanen AJ, Ojala A, Dernjatin M, Jaakkola J, Tiirola M (2016) Methylophaga and Hyphomicrobium can be used as target genera in monitoring saline water methanol-utilizing denitrification. J Ind Microbiol Biot. doi:10.1007/s10295-016-1839-2
Saunders AM, Albertsen M, Vollertsen J, Nielsen PH (2016) The activated sludge ecosystem contains a core community of abundant organisms. ISME J 10:11–20
Smalley NE, Taipale S, De Marco P, Doronina NV, Kyrpides N, Shapiro N, Woyke T, Kalyuzhnaya MG (2015) Functional and genomic diversity of methylotrophic Rhodocyclaceae: description of Methyloversatilis discipulorum sp. nov. Int J Syst Evol Microbiol 65:2227–2233
Sperl GT, Hoare DS (1971) Denitrification with methanol: a selective enrichment for Hyphomicrobium species. J Bacteriol 108:733–736
Srinandan CS, D´souza G, Srivastava N, Nayak BB, Nerurkar AS (2012) Carbon sources influence the nitrate removal activity, community structure and biofilm architecture. Bioresour Technol 117:292–299
Suzuki M, Rappe MS, Giovannoni SJ (1998) Kinetic bias in estimates of coastal picoplankton community structure obtained by measurements of small-subunit rRNA gene PCR amplicon length heterogeneity. Appl Environ Microbiol 64:4522–4529
Timmermans P, Van Heute A (1983) Denitrification with methanol: Fundamental study of the growth and denitrification capacity of Hyphomicrobium sp. Water Res 17:1249–1255
Wagner M, Loy A, Nogueira R, Purkhold U, Lee N, Daims H (2002) Microbial community composition and function in wastewater treatment plants. Antonie Van Leeuwenhoek 81:665–680
Wallenstein MD, Myrold DD, Firestone M, Voytek M (2006) Environmental controls on denitrifying communities and denitrification rates: insights from molecular methods. Ecol Appl 16:2143–2152
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Wu X-L, Yu S-L, Gu J, Zhao G-F, Chi C-Q (2009) Filomicrobium insigne sp. nov., isolated from an oil-polluted saline soil. Int J Syst Evol Micr 59:300–305
Zhu J, Wang Q, Yuan M, Tan GYA, Sun F, Wang C, Wu W, Lee PH (2016) Microbiology and potential applications of aerobic methane oxidation coupled to denitrification (AME-D) process: a review. Water Res 90:203–215
Acknowledgements
We thank P. Lindholm, P. Lindell, L. Sundell, K. Murtonen, and M. Heinonen for technical assistance. We thank R. Kettunen for valuable comments on this manuscript. We also thank H. Devlin, B. Thamdrup, and S. Hallin for comments on the earlier version of this manuscript. This study was funded by Maa-ja Vesitekniikan Tuki ry for A.J.R and Academy of Finland (Projects 286642 and 140964 to A.J.R and 260797 to M.T.) as well as European Research Council (ERC) Consolidator Project 615146 to M.T.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rissanen, A.J., Ojala, A., Fred, T. et al. Methylophilaceae and Hyphomicrobium as target taxonomic groups in monitoring the function of methanol-fed denitrification biofilters in municipal wastewater treatment plants. J Ind Microbiol Biotechnol 44, 35–47 (2017). https://doi.org/10.1007/s10295-016-1860-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-016-1860-5