Abstract
Which bacterial taxonomic groups can be used in monitoring saline water methanol-utilizing denitrification and whether nitrate is transformed into N2 in the process are unclear. Therefore, methylotrophic bacterial communities of two efficiently functioning (nitrate/nitrite reduction was 63–96 %) tropical and cool seawater reactors at a public aquarium were investigated with clone library analysis and 454 pyrosequencing of the 16S rRNA genes. Transformation of nitrate into N2 was confirmed using 15N labeling in incubation of carrier material from the tropical reactor. Combining the data with previous study results, Methylophaga and Hyphomicrobium were determined to be suitable target genera for monitoring the function of saline water methanol-fed denitrification systems. However, monitoring was not possible at the single species level. Interestingly, potential nitrate-reducing methylotrophs within Filomicrobium and closely related Fil I and Fil II clusters were detected in the reactors suggesting that they also contributed to methylotrophic denitrification in the saline environment.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Denitrification, step-wise reduction of water-soluble nitrate (NO3 −) via nitrite (NO2 −) to gaseous nitric oxide (NO), nitrous oxide (N2O) and di-nitrogen (N2) by facultative anaerobic heterotrophic bacteria, provides an important biotechnological water treatment process for nitrogen (N) removal. Denitrification is mostly a community process, as many denitrifiers perform only partial denitrification reducing NO3 − to NO2 − or to N2O, and only some bacterial species are capable of the whole denitrification chain from NO3 − to N2 gas [10]. Due to the low C:N ratio of the influent water in many N removal systems, an external carbon source, usually methanol, is added to the process. Methanol-utilizing denitrification systems are widely applied in municipal wastewater treatment plants [18]. In addition, these systems are especially important for reducing toxic inorganic N compounds in closed marine facilities, for example, in public aquaria [21] and aquaculture farms [25].
The physicochemical and technical aspects of methanol-utilizing denitrification processes have been comprehensively characterized [18, 24–26]. However, denitrification is usually measured by NO x reduction, that is, the disappearance of NO3 −/NO2 − [23]. This indirectly measured denitrification rate denotes the conversion of water-soluble NO − x into gaseous forms, but the proportions of NO, N2O and N2 in the end-product are not specified. Thus far, only a few studies have been conducted on direct measurements of gaseous end-products [17, 27]. These studies mostly focused on N2O production [17, 27], and the conversion of NO3 − to N2 has only very rarely been measured or even confirmed in water treatment facilities [6].
Optimal control and operation of wastewater treatment processes would also greatly benefit from microbiological data [27, 43], such as monitoring the presence and abundance of taxonomic groups crucial for system function [27]. The search for potential target taxonomic groups for monitoring saline water methanol-utilizing denitrification processes should be carried out in efficiently functioning systems and should focus on methylotrophic (C1-compound utilizing) organisms as they play a key role in the current processes, by utilizing methanol as an electron donor in denitrification and by transforming methanol into various extracellular organic compounds, which can then be utilized by co-occurring non-methylotrophic denitrifiers [27]. Methylotrophic bacteria of the genera Methylophaga and Hyphomicrobium have been shown to dominate the two previously studied saline water methanol-fed denitrification systems: a moving bed biofilm reactor at a seawater aquarium [3, 21, 22] and a laboratory-scale continuously stirred tank reactor (CSTR) that treats synthetic saline wastewater [32]. In addition to these two genera, Azoarcus and Paracoccus were important methylotrophs in CSTR [32]. Further analyses in the aquarium showed the genetic potential of Methylophaga to reduce NO3 − to NO2 − and Hyphomicrobium to complete the denitrification by converting NO2 − into N2 [2, 3, 42]. However, physicochemical and biological variations among bioreactors could lead to differences in the community composition. Thus, other methylotrophs could be important for the function of these systems given that methylotrophy is a quite widely dispersed trait among bacteria [19], and many can also conduct partial or complete denitrification [4, 5, 8, 12, 20, 45]. Therefore, more studies are needed to determine suitable target taxonomic groups for monitoring the function of saline water methanol-fed denitrification processes.
This study investigated NO − x reduction and the bacterial communities of two methanol-utilizing denitrifying bioreactors in a marine fish aquarium operating at two water circulation temperatures. We aimed to confirm the microbiological transformation of NO3 − to N2 using batch incubations and the 15N tracer technique. Furthermore, we aimed to find suitable target taxonomic groups for monitoring the function of saline water methanol-utilizing denitrification processes. This was achieved via a clone library and 454 pyrosequencing analysis of the 16S rRNA genes and comparison of the results for the two study reactors to those for previously studied systems [3, 21, 32]. We specifically focused on the analysis of taxonomic groups that harbor known methylotrophs.
Materials and methods
Sampling site and sampling for molecular microbiology
The public fish aquarium SEA LIFE Helsinki Finland (https://www.visitsealife.com/helsinki/) is divided into two water recirculation systems (warm tropical, AQUAR_T, and cold North Atlantic, AQUAR_C) and has a total water volume of 420 m3 with 27–34 ppt salinity. Both circulation systems have their own fluidized-bed type denitrification reactors that receive NO3 −-rich water from the nitrification stage. The volume of each reactor is ~330 L of which ~90 L is the fluidized carrier material bed (oolitic sand, 100 kg, density = 1.5 kg/L). The temperature inside the denitrification reactors of AQUAR_T and AQUAR_C was 23–24 and 18–19 °C, respectively. Two samples of oolitic sand for molecular microbiological analyses were collected twice, on 10 November 2008 and 8 September 2010, from the AQUAR_T and AQUAR_C reactors, in sterile 50 mL plastic containers and stored at −20 °C before processing within 1–2 months. The reactors utilized methanol as their carbon source except AQUAR_C in 2008 when a mixture of methanol and saccharose was used until the sampling time point after which only methanol was used.
Denitrification measurements
The NO − x concentrations inside the reactors, near the reactor outlet (NO − x out) and in the inflow water feeding the reactors (NO − x inflow) were measured using Spectroquant® nitrate and nitrite test kits (Merck Millipore, Germany) with a Spectroquant® Nova 60 photometer (Merck Millipore, Germany) from both reactors 1–2 times per month for 2.3 years (time period 24 September 2008 to 28 December 2010). The methanol addition (Metf, mmol/h) and water flow (W f, L/h) rates were adjusted by the operators and for this study reported for 1.5–2 month periods before bacterial sampling in 2008 and 2010 (24 September to 10 November 2008 and 13 July to 8 September 2010). Hourly NO − x loads (LNO − x inflow and LNO − x out, mmol N/h) were calculated using the W f and NO − x concentration values for these time periods. Denitrification was then estimated indirectly as the relative NO − x reduction (%):
and as the actual NO − x reduction:
which was converted into the NO − x reduction rate of the carrier material (µmol N/Lcar/h).
Direct denitrification measurement was performed with bottle incubation in 2010. Batches of oolitic sand (~36 mL) collected from the middle of the filter bed in AQUAR_T were put in 120 mL glass bottles (11 bottles altogether). The bottles were filled with anoxic reactor water by submerging them in the reactor, and while submerged, they were closed with caps that had butyl rubber stoppers. Each bottle was injected with ~219 µmol (~7 mg) of methanol and ~50 µmol (~0.7 mg) of NO3 −-N, which had a 2 % 15N isotope label, and then shaken briefly but vigorously. The NO3 − stock solution (0.1 M NO3 −-N) was prepared from NaNO3 and K15NO3 − (Cambridge Isotope Laboratories, Inc., MA, USA). The total NO3 −-N concentration after the addition was ~664 µM, which is approximately the same as the average NO − x inflow concentration (~688 µM) during the ~2-month period before sampling in 2010. The bottles were incubated non-shaken at 21–22 °C and sacrificed in batches of 3–4 bottles after 2 h (4 bottles), 6 h (4 bottles) and 21 h (3 bottles) of incubation. One non-incubated water sample taken from the reactor before the incubation periods served as the 0 time point control. The water subsamples were stored in 12 mL borosilicate glass Exetainer® tubes with screw-capped butyl rubber septa (Labco Ltd., High Wycombe, UK), and the microbial activity in the vials was terminated by adding 0.1 mL of ZnCl2 (1 g/mL). The concentration and 15N content of the N2 gas in the water was measured as in Tiirola et al. [41]. Denitrification was calculated as the rate of the total N2 gas accumulation and converted to the N2 production rate of the carrier material (µmol N/Lcar/h). In addition, reduction of NO3 − to N2 gas was verified by the accumulation of excess 15N-containing N2 gas. The concentration of the excess 15N-containing N2 gas, [excess 15N], was calculated for each sample as
where [N2sample] is the N2 gas concentration in the incubated sample and the at%15Nsample and the at%15Nzero are the 15N content (in %) of the N2 gas in the incubated and non-incubated (0 time point) samples, respectively.
Molecular microbiological analyses
DNA was extracted from 0.5 to 0.6 g of frozen oolitic sand from each sample using glass bead beating and phenol–chloroform extraction, which was followed by isopropanol–NaCl precipitation (pH 8) and dissolution of the DNA pellet in TE buffer.
PCR for the clone library analyses of the 16S rRNA genes was performed with primers 27F (5′-AGAGTTTGATCMTGGCTCAG-3′)/907R (5′-CCGTCAATTCMTTTGAGTTT-3′) as previously described [36], but using the following program: initial denaturation at 95 °C for 5 min and 30 cycles of amplification (94 °C for 30 s, 53 °C for 1 min, 72 °C for 3 min) and final elongation at 72 °C for 15 min. The PCR products of the replicate samples were pooled for subsequent cloning and sequencing, which was carried out as previously described [36].
To get deeper insight into the bacteria inhabiting the methanol-fed denitrification reactors, the bacterial community in AQUAR_T, which used methanol as the sole carbon source, was studied with 454 pyrosequencing. Equal amounts of nucleic acid extracts from replicate samples of AQUAR_T in 2008 were pooled before the PCR reactions. PCR amplification of the 16S rRNA genes using primers 341F (5′-CCTACGGGNGGCWGCAG-3′)/805R (5′-GACTACHVGGGTATCTAATCC-3′), subsequent purification steps and sequencing were performed as previously described [33].
Sequence analysis
In the analysis of the clone library sequences, the Mothur program package [38] was used for sequence alignment, chimera-checking (chimera.uchime executable), classification of sequences into operational taxonomic units (OTUs; 97 % identity threshold) and taxonomic classification of the OTUs (using the Ribosomal Database Project database). Variations in the community structure among the samples were analyzed with hierarchical clustering (UPGMA linkage, Bray–Curtis distances) using PAST version 3.09 [13]. Representative sequences of the OTUs assigned to taxonomic groups of known methylotrophs derived from the previous literature [8, 19, 21, 22, 32, 45], in this case Hyphomicrobiaceae and Methylophaga, were subjected to phylogenetic tree analyses [neighbor-joining (NJ) method, Kimura-2 distances, pairwise exclusion of gaps] using Mega 5.05 [40].
Tags and primer sequences, as well as low-quality sequences (containing ambiguous nucleotides and homopolymers longer than eight nucleotides) were removed from the 454 pyrosequencing library. Sequences (~200 bp) were thereafter processed as described above for the clone library analysis except phylogenetic trees were not constructed.
The 16S rRNA gene sequences of the clone libraries were deposited in the NCBI GenBank (Accession Numbers KP098736-KP098970, KP098976-KP098984 and KP098989-KP099005). The 454 pyrosequencing data were deposited in the NCBI SRA database (SRX646347).
Results
Functioning of denitrifying reactors
The operator-adjusted W f and Metf were lower in 2010 than in 2008 in both reactors during the 1.5–2 month period before bacterial sampling (Table 1). The average hourly NO − x load in the inflow decreased in AQUAR_T and increased in AQUAR_C from 2008 to 2010 (Table 1). The average molar ratio of the added methanol to the N load methanol:NO − x -Ninflow (mol:mol) decreased from 2008 to 2010 in both reactors (Table 1). The ratio was higher in AQUAR_C than in AQUAR_T in 2008 and vice versa in 2010 (Table 1).
The relative NO − x reduction, on average, was slightly higher in AQUAR_C (91 %, range 81–95 %) than in AQUAR_T (86 %, range 63–96 %) during the 2.3 year monitoring period (24 September 2008 to 28 December 2010, excluding the very low values caused by the pumping and carbon dosage problems on 4 November 2008 in both reactors and on 4 June 2009 in AQUAR_C; Fig. 1). When averaged over the 1.5–2 month period before the bacterial sampling, the average actual NO − x reduction rate expressed per carrier (oolitic sand) volume was lower in 2008 than in 2010 in both reactors (Table 1). The rate was higher in AQUAR_T than in AQUAR_C in 2008 and vice versa in 2010 (Table 1).
Concentration of NO − x in the inflow and the outflow and the relative NO − x reduction in the denitrification reactors, AQUAR_T and AQUAR_C of the sea water aquarium from 24 September 2008 to 28 December 2010. The sampling dates for the microbial studies (10 November 2008 and 8 September 2010) are indicated with arrows
Bottle incubation tests confirmed that N2 was produced and NO3 − was transformed into N2 by microbes in AQUAR_T (Fig. 2). The estimated N2 production rate of the carrier material was 56.4 µmol N/Lcar/h.
Concentrations (average ± standard deviation) and estimated production rates of N2 gas and excess 15N (in N2 gas) in incubation bottles after 2 h (n = 4), 6 h (n = 4) and 21 h (n = 3) of incubation (at room temperature) of the carrier material and water from AQUAR_T with methanol and 15N-labeled NO3 −. One non-incubated water sample taken from the reactor before the incubation served as the 0 time point control (n = 1). The standard deviations of the N2 and 15N concentrations at time point 2 h are very low and masked behind the symbol
Variation in microbial community structure
The clone library analysis of the 16S rRNA genes indicated that the variation in the bacterial community structure was higher between the reactors than between the years (Table 2; Online Resource 1). Variation between the years in the community structure was considerably higher in the cooler reactor AQUAR_C, which was first fed with a mixture of methanol and saccharose, than in the warmer reactor AQUAR_T (Table 2; Online Resource 1).
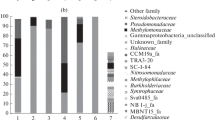
Different taxonomic groups were assigned to a putative methylotrophic function based on the previous literature. The methylotrophic groups detected in the reactors were the genera Methylophaga, Hyphomicrobium, Filomicrobium, as well as clusters Fil I and Fil II, which were more closely related to Filomicrobium than to Hyphomicrobium (Table 2; Figs. 3, 4). Betaproteobacterial methylotrophs were not detected. In the phylogenetic tree, the OTUs assigned to the genus Hyphomicrobium were positioned between the previously assigned Hyphomicrobium clusters I and II [35] (Fig. 3). Most of the OTUs and sequences assigned to Methylophaga clustered close to M. nitratireducenticrescens, whereas one of the OTUs had its closest relative in M. thiooxydans (Fig. 4).
Phylogenetic tree (neighbor-joining method) of Hyphomicrobiaceae based on a comparative analysis of 16S rRNA gene sequences in clone libraries showing the phylogenetic position of the operational taxonomic units (OTUs; at 97 % sequence similarity) in AQUAR_C and AQUAR_T in 2008 and 2010 (symbols). Hyphomicrobium clusters I and II were previously defined by Rainey et al. [35]. The numbers in the parentheses after the OTU number indicate the number of sequences within that OTU. The numbers at the nodes indicate the percentage of occurrence in 1000 bootstrapped trees (bootstrap values >50 % are shown)
Phylogenetic tree (neighbor-joining method) of Methylophaga based on a comparative analysis of 16S rRNA gene sequences in clone libraries showing the phylogenetic position of the operational taxonomic units (OTUs; at 97 % sequence similarity) in AQUAR_C and AQUAR_T at 2008 and 2010 (symbols). The numbers in the parentheses after the OTU number indicate the number of sequences within that OTU. The numbers at the nodes indicate the percentage of occurrence in 1000 bootstrapped trees (bootstrap values >50 % are shown)
The relative abundance of the putative methylotrophs was considerably higher in the warmer reactor AQUAR_T than in the cooler reactor AQUAR_C in both years (Table 2). In addition, methylotrophs were more abundant in 2010 than in 2008 in AQUAR_C, whereas the opposite took place in AQUAR_T (Table 2). The relative abundance of Methylophaga was considerably higher in the warmer reactor AQUAR_T than in the cooler reactor AQUAR_C (Table 2). In AQUAR_C, Methylophaga were absent in 2008 and present at very low numbers in 2010. Correspondingly, there was a higher abundance of Methylophaga in 2010 than in 2008 in AQUAR_T (Table 2). Methylophaga in AQUAR_T had their closest relative in M. nitratireducenticrescens, whereas those in AQUAR_C were most closely related to M. thiooxydans (Fig. 4). Of the family Hyphomicrobiaceae, Hyphomicrobium were much more abundant in the warmer reactor AQUAR_T than in the cooler reactor AQUAR_C. There was a higher abundance of Hyphomicrobium in 2010 than in 2008 in AQUAR_C and vice versa in AQUAR_T (Table 2). In contrast, the bacteria of the Fil I cluster were much more abundant in AQUAR_C than in AQUAR_T (Table 2). In AQUAR_T, Fil I bacteria were absent in 2008 and present at very low abundance in 2010. Correspondingly, there was a higher abundance of Fil I bacteria in 2010 than in 2008 in AQUAR_C. Fil II cluster bacteria were present in low abundance in both reactors and were absent in AQUAR_T in 2010 (Table 2). Filomicrobium cluster bacteria were present in low abundance and only in AQUAR_T (Table 2; Fig. 3).
The relative abundance of other bacteria also varied between reactors and years and was generally higher in AQUAR_C than in AQUAR_T (Table 2). Within Gammaproteobacteria, there was a considerably large group of unclassified 16S rRNA gene sequences that probably represented clades without cultured representatives (Table 2). Of Alphaproteobacteria (other than methylotrophs), Rhodobacteraceae and Phyllobacteriaceae as well as non-methylotrophic Hyphomicrobiaceae within Maritalea and Mar I clusters were detected (Table 2; Fig. 3). According to BLAST searches [1] of the NCBI nr-database, the detected Rhodobacteraceae were most closely related to the non-methylotrophic genus Roseovarius. Of the Proteobacteria, also Deltaproteobacteria were present. Other detected phyla were Acidobacteria, Actinobacteria, Bacteroidetes, Chloroflexi, Deferribacteres, Planctomycetes and Spirochaetes (Table 2).
There were differences in the results of the clone library analysis and 454 pyrosequencing (Table 2). Most importantly, 454 pyrosequencing had lower resolution to detect Alphaproteobacteria or resolve different genera within Hyphomicrobiaceae and resulted in a higher proportion of unclassified bacterial sequences than the clone library analysis (Table 2). Since high taxonomic resolution was necessary to identify clusters especially within Hyphomicrobiaceae, the focus in this study was on the clone library results.
Discussion
Both study reactors removed N very efficiently. The relative NO − x reduction was even slightly higher than measured at other methanol-fed sea water denitrification bioreactors that reduced 65–90 % [22] and up to 88 % [24] of NO − x . In contrast, the average actual NO − x reduction rates were within the lower end of the range (300–9000 µmol N/Lcar/h) measured at other systems [21, 23, 24]. The differences in the actual NO − x reduction rates are mostly due to 3–10 times higher N loads (as expressed per carrier volume) in the previously studied reactors [23, 24]. However, the differences can be also partially attributed to the use of deaeration systems to remove O2 in the inflow and the amendment of the trace metal solution in the previously studied reactors, which enhanced denitrification [21, 23]. To our knowledge, this is the first study of saline water methanol-utilizing denitrification systems that confirmed NO3 − is reduced to N2. However, the N2 production measured in the batch tests was only 6–7 % of the actual NO − x reduction rates in the AQUAR_T reactor. This difference is very likely due to the lack of shaking during incubation, which reduced the contact between NO3 − and the bacteria on the carrier material. However, the possible formation of other gases, N2O [17] and NO, during the incubation periods can also explain a small part of this discrepancy.
Combining the results from the two study reactors and the two previously studied systems [3, 21, 32] shows that putative methylotrophic bacteria belonging to Methylophaga and Hyphomicrobium are very common in efficiently functioning saline water methanol-fed denitrification systems. This result indicates that these bacteria are crucial for the function of the process. Thus, these two genera can serve as targets when monitoring the function of saline-water methanol-utilizing denitrification systems. However, there were considerable species-level differences between the systems. Hyphomicrobium nitrativorans, which dominated in a previously studied system [3, 21], were not found at all in the AQUAR reactors, and Methylophaga nitratireducenticrescens, which inhabited a previously studied reactor [3] and AQUAR_T, were not found in AQUAR_C. Thus, no single Methylophaga or Hyphomicrobium species can be determined for use as a general target species for monitoring the function of saline water methanol-fed denitrification systems.
Interestingly, this study is also the first to show that members of Filomicrobium sp. and Filomicrobium-related Fil I and Fil II clusters can be abundant in efficiently functioning saline water methanol-fed denitrification systems. In addition to results from cultivation-based [45] and genomic [14] studies of Filomicrobium strains, further confirmation of the bacteria’s methylotrophic metabolism was provided by the concurrent increase in the relative abundance of Hyphomicrobium, Methylophaga and Fil I bacteria in AQUAR_C from 2008 to 2010 when the C source changed from saccharose + methanol to solely methanol. In 2008, the methylotrophs were probably over-competed by more efficiently growing non-methylotrophs, for example, in Bacteroidetes, Rhodobacteraceae and Maritalea. Furthermore, the concurrent decrease in Hyphomicrobium and Fil II bacteria in AQUAR_T from 2008 to 2010 could be due to the decrease in the availability of methanol (a decrease in Metf and methanol:NO x -Ninflow before sampling).
Previous studies also suggest that Filomicrobium participate in denitrification by dissimilatory reduction of NO3 − to NO2 − [14, 45]. Thus, in addition to Methylophaga [2, 3], Filomicrobium, Fil I and Fil II bacterial clusters could couple methylotrophy with the reduction of NO3 − to NO2 − and supply it for denitrifiers capable of further denitrification steps [3]. However, previously studied strains of Methylophaga thiooxydans [7] and M. nitratireducenticrescens [42] also had the NO − 2 reductase gene, although it was truncated in M. nitratireducenticrescens [42]. In addition, M. nitratireducenticrescens had genes for NO and N2O− reduction [42]. This suggests that Methylophaga species coupling methylotrophy to NO2 −, NO and N2O reduction could exist in nature and in saline water methanol-fed denitrification systems. Since only a few Filomicrobium strains have been characterized for their metabolic potential [14, 45], it is possible that some species within Filomicrobium, Fil I and Fil II clusters are also capable of this.
Thorough comparative analyses of the factors that affect the growth and activity of Filomicrobium and Hyphomicrobium species do not exist. Therefore, many possible factors might have affected the genera- and species-level variation of Hyphomicrobiaceae between the AQUAR reactors and previously studied systems [3, 21]. In a comparison of denitrifying Hyphomicrobium species, NO3 − significantly affected the growth and activity of Hyphomicrobium species with H. nitrativorans growing and denitrifying at higher NO3 − concentrations than the other studied species, H. zavarzinii and H. denitrificans [30]. Thus, the considerably lower N loads could explain the presence and dominance of Hyphomicrobium species other than H. nitrativorans in the AQUAR reactors. Similarly, Filomicrobium, Fil I and Fil II bacteria could have been favored in the lower N load conditions that prevail in the AQUAR system. Furthermore, O2 constantly enters AQUAR reactors due to the lack of a preceding deaeration step but is rapidly consumed there based on the anoxic conditions [(O2) < 0.1 mg/L, below the detection limit of the O2 probe] that prevail inside the reactors [9]. Thus, variation in the O2 availability between the systems can also explain the differences in the bacterial communities. Part of the Hyphomicrobium and Filomicrobium OTUs in the AQUAR reactors may have been favored by O2. Analogously, as in aerobic methane oxidation coupled with denitrification (AME-D) [46], these bacteria could have contributed to the overall denitrification performance by consuming O2 and by aerobically converting methanol to organic substrates utilizable by non-methylotrophic denitrifiers. However, in addition to variating NO3 − and O2, differences in the inocula (the original bacterial community that colonized the reactors) and in the carrier materials between the reactors as well as the amendment of the trace metal solution and the higher addition of methanol in the previously studied systems [3, 21, 22] could have also affected the differences in the bacterial communities.
The lower relative abundance of methylotrophs in the cooler reactor AQUAR_C than in AQUAR_T can be first explained by the saccharose addition in 2008 leading to over-competition of non-methylotrophic organisms in AQUAR_C. In 2010, the difference can be attributed to the lower availability of methanol (lower Metf and methanol:NO − x inflow) in AQUAR_C. Variation in the availability of methanol can also explain the lower abundance of Hyphomicrobium and Methylophaga and the higher abundance of Fil I bacteria in AQUAR_C than in AQUAR_T. However, differences in temperature could also play a role here since the lowest limits of the growth temperature ranges of Methylophaga (M. nitratireducenticrescens, 15–37 °C) [42] and Hyphomicrobium (H. nitrativorans, 15–35 °C) [29] are higher than that of Filomicrobium (e.g., F. insigne, 4–45 °C) [45], which indicates that Filomicrobium-related Fil I bacteria grew better in the cooler AQUAR_C reactor. The growth temperature range of M. thiooxydans, the closest cultured relative of Methylophaga in AQUAR_C, is not known. However, the absence of M. thiooxydans in the previously studied sea water reactor with a slightly lower temperature, 16–18 °C [3], than in AQUAR_C, 18–19 °C, suggests that temperature variations do not explain the differences in the Methylophaga species between AQUAR_C and AQUAR_T. M. thiooxydans can grow on a wider range of carbon substrates (methanol, fructose, monomethylamine) than M. nitratireducenticrescens (methanol) [42]. Thus, differences in the Methylophaga species between the reactors could be explained by M. thiooxydans surviving better with lower methanol availability.
Many other detected phyla, that is, Acidobacteria, Actinobacteria, Bacteroidetes, Chloroflexi, Deferribacteres, Planctomycetes and Proteobacteria (other than the detected methylotrophs), contain species capable of either partial or complete denitrification [11, 27, 37, 39, 44] suggesting that they contribute to the overall denitrification performance of the AQUAR reactors and utilize the organic compounds produced by methylotrophs. However, recent cultivation studies have shown that methylotrophy is present within Bacteroidetes [28] and Actinobacteria [15]. In addition, DNA stable isotopic probing (SIP) analyses, although not able to distinguish between direct C1 utilization and cross-feeding, raise speculation about the existence of unclassified methylotrophic Gammaproteobacteria [31], as well as methylotrophic members within Bacteroidetes, Deferribacteres [16] and Acidobacteria [34]. Further studies are needed to show whether members of these groups couple methylotrophy with denitrification in saline water systems and could also be considered potential target taxonomic groups for monitoring purposes.
Conclusions
This study confirmed that NO3 was transformed into N2 in a functioning saline water methanol-fed denitrification system. Combining the results of this study with those of previous studies show that Methylophaga and Hyphomicrobium can serve as target genera in monitoring the function of saline water methanol-utilizing denitrification systems. However, species-level differences among the systems, stemming from differences in the physicochemical and biological conditions, prevent the determination of individual Methylophaga or Hyphomicrobium species to be used as general target species in process monitoring. Interestingly, other bacteria with potential methylotrophic and NO3 −-reducing metabolism, within Filomicrobium and Fil I and Fil II clusters, were found, suggesting they also contributed to saline water methylotrophic denitrification. Whether bacteria within Methylophaga, Filomicrobium, Fil I and Fil II clusters can only reduce NO3 − to NO2 − or reduce NO2 −, NO and N2O, and whether other detected bacteria are able to couple methylotrophy with denitrification in the saline water denitrification bioreactors, is still to be resolved. These studies should employ high-throughput culturing methods to isolate target organisms, as well as metagenomics and metatranscriptomics.
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Auclair J, Lépine F, Parent S, Villemur R (2010) Dissimilatory reduction of nitrate in seawater by a Methylophaga strain containing two highly divergent narG sequences. ISME J 4:1302–1313
Auclair J, Parent S, Villemur R (2012) Functional diversity in the denitrifying biofilm of the methanol-fed marine denitrification system at the Montreal Biodome. Microb Ecol 63:726–735
Baytshtok V, Lu H, Park H, Kim S, Yu R, Khandran K (2009) Impact of varying electron donors on the molecular microbial ecology and biokinetics of methylotrophic denitrifying bacteria. Biotechnol Bioeng 102:1527–1536
Beck DAC, McTaggart TL, Setboonsarng U, Vorobev A, Kalyuzhnaya MG, Ivanova N, Goodwin L, Woyke T, Lidstrom ME, Chistoserdova L (2014) The expanded diversity of Methylophilaceae from Lake Washington through cultivation and genomic sequencing of novel ecotypes. PLoS One 9:e102458
Bernet N, Delgenes N, Akunna JC, Delgenes JP, Moletta R (2000) Combined anaerobic–aerobic SBR for the treatment of piggery wastewater. Water Res 34:611–619
Boden R, Ferriera S, Johnson J, Kelly DP, Murrell JC, Schäfer H (2011) Draft genome sequence of the chemolithoheterotrophic, halophilic methylotroph Methylophaga thiooxydans DMS010. J Bacteriol 193:3154–3155
Chistoserdova L, Kalyuzhnaya MG, Lidstrom ME (2009) The expanding world of methylotrophic organisms. Annu Rev Microbiol 63:477–499
Dernjatin M (2008) Leijuva-alustaisen denitrifikaatioprosessin optimointi suljetussa kiertovesijärjestelmässä. MsC thesis, University of Helsinki, Helsinki, Finland (in Finnish)
Gentile ME, Nyman JL, Criddle CS (2007) Correlation of patterns of denitrification instability in replicated bioreactor communities with shifts in the relative abundance and the denitrification patterns of specific populations. ISME J 1:714–728
Gittel A, Kofoed MVW, Sørensen KB, Ingvorsen K, Schramm A (2012) Succession of Deferribacteres and Epsilonproteobacteria through a nitrate-treated high-temperature oil production facility. Syst Appl Microbiol 35:165–174
Hagman M, Nielsen JL, Nielsen PH, la Jansen J C (2008) Mixed carbon sources for nitrate reduction in activated sludge-identification of bacteria and process activity studies. Water Res 42:1539–1546
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4(1):9
Henriques AC, De Marco P (2015) Complete genome sequences of two strains of “Candidatus Filomicrobium marinum”, a methanesulfonate-degrading species. Genome Announc 3:e00160-15
Hung WL, Wade WG, Boden R, Kelly DP, Wood AP (2011) Facultative methylotrophs from the human oral cavity and methylotrophy in strains of Gordonia, Leifsonia, and Microbacterium. Arch Microbiol 193:407–417
Jensen S, Neufeld JD, Birkeland NK, Hovland M, Murrell JC (2008) Methane assimilation and trophic interactions with marine Methylomicrobium in deep-water coral reef sediment off the coast of Norway. FEMS Microbiol Ecol 66:320–330
Kampschreur MJ, Temmink H, Kleerebezem R, Jetten MSM, van Loosdrecht MCM (2009) Nitrous oxide emission during wastewater treatment. Water Res 43:4093–4103
Koch G, Siegrist H (1997) Denitrification with methanol in tertiary filtration. Water Res 31:3029–3038
Kolb S (2009) Aerobic methanol- oxidizing Bacteria in soil. FEMS Microbiol Lett 300:1–10
Kumaresan D, Wischer D, Hillebrand-Voiculescu AM, Murrell JC (2015) Draft genome sequences of facultative methylotrophs, Gemmobacter sp. strain LW1 and Mesorhizobium sp. strain 1M-11, isolated from Movile Cave, Romania. Genome Announc 3:e01266-15
Labbé N, Juteau P, Parent S, Villemur R (2003) Bacterial diversity in a marine methanol-fed denitrification reactor at the Montreal Biodome, Canada. Microb Ecol 46:12–21
Labbé N, Laurin V, Juteau P, Parent S, Villemur R (2007) Microbiological community structure of the biofilm of a methanol-fed, marine denitrification system, and identification of the methanol-utilizing micro-organisms. Microb Ecol 53:621–630
Labbé N, Parent S, Villemur R (2003) Addition of trace metals increases denitrification rate in closed marine systems. Water Res 37:914–920
Labelle M-A, Juteau P, Jolicoeur M, Villemur R, Parent S, Comeau Y (2005) Seawater denitrification in a closed mesocosm by a submerged moving bed biofilm reactor. Water Res 39:3409–3417
Lee PG, Lea RN, Dohmann E, Prebilsky W, Turk PE, Ying H, Whitson JL (2000) Denitrification in aquaculture systems: an example of a fuzzy logic control problem. Aquac Eng 23:37–59
Lemmer H, Zaglauer A, Metzner G (1997) Denitrification in a methanol-fed fixed-bed reactor. Part 1: physico-chemical and biological characterization. Water Res 31:1897–1902
Lu H, Chandran K, Stensel D (2014) Microbial ecology of denitrification in biological wastewater treatment. Water Res 64:237–254
Madhaiyan M, Poonguzhali S, Lee J-S, Lee KC, Sundaram S (2010) Flavobacterium glycines sp. nov., a facultative methylotroph isolated from the rhizosphere of soybean. Int J Syst Evol Microbiol 60:2187–2192
Martineau C, Villeneuve C, Mauffrey F, Villemur R (2013) Hyphomicrobium nitrativorans sp. nov., isolated from the biofilm of a methanol-fed denitrification system treating seawater at the Montreal Biodome. Int J Syst Evol Microbiol 63:3777–3781
Martineau C, Mauffrey F, Villemur R (2015) Comparative analysis of denitrifying activities of Hyphomicrobium nitrativorans, Hyphomicrobium denitrificans and Hyphomicrobium zavarzinii. Appl Environ Microbiol 81:5003–5014
Neufeld JD, Schäfer H, Cox MJ, Boden R, McDonald IR, Murrell JC (2007) Stable-isotope probing implicates Methylophaga spp and novel Gammaproteobacteria in marine methanol and methylamine metabolism. ISME J 1:480–491
Osaka T, Shirotani K, Yoshie S, Tsuneda S (2008) Effects of carbon source on denitrification efficiency and microbial community structure in a saline wastewater treatment process. Water Res 42:3709–3718
Peura S, Eiler A, Bertilsson S, Nykänen H, Tiirola M, Jones RI (2012) Distinct and diverse anaerobic bacterial communities in boreal lakes dominated by candidate division OD1. ISME J 6:1640–1652
Radajewski S, Webster G, Reay DS, Morris SA, Ineson P, Nedwell DB, Prosser JI, Murrell JC (2002) Identification of active methylotroph populations in an acidic forest soil by stable-isotope probing. Microbiology 148:2331–2342
Rainey FA, Ward-Rainey N, Gliesche CG, Stackebrandt E (1998) Phylogenetic analysis and intrageneric structure of the genus and the related genus Filomicrobium. Int J Syst Bacteriol 48:635–639
Rissanen AJ, Kurhela E, Aho T, Oittinen T, Tiirola M (2010) Storage of environmental samples for guaranteeing nucleic acid yields for molecular microbiological studies. Appl Microbiol Biotechnol 88:977–984
Scheuner C, Tindall BJ, Lu M, Nolan M, Lapidus A, Cheng JF, Goodwin L, Pitluck S, Huntemann M, Liolios K, Pagani I, Mavromatis K, Ivanova N, Pati A, Chen A, Palaniappan K, Jeffries CD, Hauser L, Land M, Mwrichia R, Rohde M, Abt B, Detter JC, Woyke T, Eisen JA, Markowitz V, Hugenholtz P, Göker M, Kyrpides NC, Klenk HP (2014) Complete genome sequence of Planctomyces brasiliensis type strain (DSM 5305T), phylogenomic analysis and reclassification of Planctomycetes including the descriptions of Gimesia gen. nov., Planctopirus gen. nov. and Rubinisphaera gen. nov. and emended descriptions of the order Planctomycetales and the family Planctomycetaceae. Stand Genomic Sci 9:10
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Shoun H, Kano M, Baba I, Takaya N, Matsuo M (1998) Denitrification by Actinomycetes and purification of dissimilatory nitrite reductase and azurin from Streptomyces thioluteus. J Bacteriol 180:4413–4415
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tiirola MA, Rissanen AJ, Sarpakunnas M, Arvola L, Nykänen H (2011) Stable isotope profiles of nitrogen gas indicate denitrification in oxygen-stratified humic lakes. Rapid Commun Mass Spectrom 25:1497–1502
Villeneuve C, Martineau C, Mauffrey F, Villemur R (2013) Methylophaga nitratireducenticrescens sp. nov. and Methylophaga frappieri sp. nov., isolated from the biofilm of the methanol-fed denitrification system treating the seawater at the Montreal Biodome. Int J Syst Evol Microbiol 63:2216–2222
Wagner M, Loy A, Nogueira R, Purkhold U, Lee N, Daims H (2002) Microbial community composition and function in wastewater treatment plants. Antonie van Leeuwenhoek 81:665–680
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Countinho PM, Wu M, Xie G, Haft DH, Sait M, Badger J, Barabote RD, Bradley B, Brettin TS, Brinkac LM, Bruce D, Creasy T, Daugherty SC, Davidsen TM, DeBoy RT, Detter JC, Dodson RJ, Durkin AS, Ganapathy A, Gwinn-Giglio M, Han CS, Khouri H, Kiss H, Kothari SP, Madupu R, Nelson KE, Nelson WC, Paulsen I, Penn K, Ren Q, Rosovitz MJ, Selengut JD, Shrivastava S, Sullivan SA, Tapia R, Thompson LS, Watkins KL, Yang Q, Yu C, Zafar N, Zhou L, Kuske CR (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056
Wu X-L, Yu S-L, Gu J, Zhao G-F, Chi C-Q (2009) Filomicrobium insigne sp. nov., isolated from an oil-polluted saline soil. Int J Syst Evol Microbiol 59:300–305
Zhu J, Wang Q, Yuan M, Tan GYA, Sun F, Wang C, Wu W, Lee PH (2016) Microbiology and potential applications of aerobic methane oxidation coupled to denitrification (AME-D) process: a review. Water Res 90:203–215
Acknowledgments
We thank H. Devlin, B. Thamdrup and S. Hallin for comments on an earlier version of this manuscript. This study was funded by Maa-ja Vesitekniikan Tuki ry (A.J.R.) and the Academy of Finland (projects 286642 to A.J.R. and 120089 to M.T.), as well as the European Research Council (ERC) Consolidator project 615146 (M.T.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10295_2016_1839_MOESM1_ESM.pdf
Online Resource 1 Hierarchical clustering analysis (UPGMA) of relative abundance of OTUs (using Bray-Curtis distances) in 16S rRNA gene clone library analyses of AQUAR_T and AQUAR_C in 2008 and 2010 (PDF 13 kb)
Rights and permissions
About this article
Cite this article
Rissanen, A.J., Ojala, A., Dernjatin, M. et al. Methylophaga and Hyphomicrobium can be used as target genera in monitoring saline water methanol-utilizing denitrification. J Ind Microbiol Biotechnol 43, 1647–1657 (2016). https://doi.org/10.1007/s10295-016-1839-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-016-1839-2