Abstract
A plethora of studies have uncovered numerous important genes with agricultural significance in staple crops. However, when it comes to orphan crops like minor millet, genomic research lags significantly behind that of major crops. This situation has promoted a focus on exploring research opportunities in minor millets, particularly in finger millet, using cutting-edge methods. Finger millet, a coarse cereal known for its exceptional nutritional content and ability to withstand environmental stresses represents a promising climate-smart and nutritional crop in the battle against escalating environmental challenges. The existing traditional improvement programs for finger millet are insufficient to address global hunger effectively. The lack of utilization of high-throughput platforms, genome editing, haplotype breeding, and advanced breeding approaches hinders the systematic multi-omics studies on finger millet, which are essential for pinpointing crucial genes related to agronomically important and various stress responses. The growing environmental uncertainties have widened the gap between the anticipated and real progress in crop improvement. To overcome these challenges a combination of cutting-edge multi-omics techniques such as high-throughput sequencing, speed breeding, mutational breeding, haplotype-based breeding, genomic selection, high-throughput phenotyping, pangenomics, genome editing, and more along with integration of deep learning and artificial intelligence technologies are essential to accelerate research efforts in finger millet. The scarcity of multi-omics approaches in finger millet leaves breeders with limited modern tools for crop enhancement. Therefore, leveraging datasets from previous studies could prove effective in implementing the necessary multi-omics interventions to enrich the genetic resource in finger millet.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Finger millet: a versatile crop in fragile ecosystem
Finger millet is categorized into minor millets often referred as ‘Ragi’, ‘Bulo’, ‘Wimbi’ or ‘Madua’, ‘bird foot millet’. It comes fourth after other major millets in cereal crop production scenario in Africa and Asia (Rathore et al. 2019). It is self-pollinated crop, naturally allotetraploid (2n = 4x = 36, AABB, disomic inheritance), and genome size of 1593 Mb often found in two subspecies coracana and africana (Ramashia et al. 2018). The poorest populations in Africa and Asia cultivate and consume large quantities of this annual herbaceous cereal crop. Ethiopia or neighboring country (Uganda) is recognized as the primary center for finger millet but in India, records of its cultivation goes back to more than three thousand years ago in the Western Ghats (Hilu and Johnson 1992; Fuller DQ. 2024). The finger millet era is quite significant in the context of a growing population because of its high nutraceutical content and adaptability under changing climatic conditions (Kumar et al. 2018; Yang et al. 2022a). Finger millet thrives in loose soils that range from clay to sandy loam. It is a vital staple food crop in regions with a shortage of arable land because it is also highly adaptable to marginal or poor soils (Chisi and Peterson 2019; Dhanushkodi et al. 2023). Their exceptional high shelf life (more than 10 years) and resistance to insect/pests during storage make them a reliable famine crop that can be cultivated round the year (Gupta et al. 2017). In general, it tends to adapt to fragile ecosystems as well as nutritionally superior to major staple crops (Puranik et al. 2017). Finger millet stands out as a ‘super cereal’ due to its high calcium content of up to 0.45 g per 100 g of seed, which is 5 to 30 times more than other major cereals (Puranik et al. 2017). Its grain is devoid of gluten, rich in micronutrients like Zn, Cu, Cr, and Mg (Maharajan et al. 2022). With respect to negligible gluten, their seeds are a good reservoir of dietary fiber, iron, and the important source of vital amino acids (Odeny et al. 2020). Finger millet also has antioxidant nature and it tackle with oxidative stress through its high phytochemical molecules like polyphenols and flavonoids (Jideani 2012). Other human health advantages of finger millet include its hypoglycemic, hypocholesterolemia, and anti-ulcerative properties. In addition to being a nutritious food for babies, it has been used in making cakes, bread, and sweets (Ceasar and Ignacimuthu 2011).
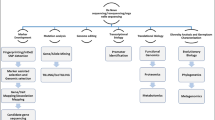
Recent status of Finger millet as an orphan crop is reflected due to its low genomic information. Finger millet’s intricate biology, self-pollinating nature, and small flowers make artificial crosses and breeding efforts challenging (Bančič et al. 2024). It is a cross of two diploid species thought to have originated in East Africa. The first of which is Eleusine indica, which is often widespread throughout Africa i.e. a sub-genome and another parent diploid pre-B sub-genome information is still hidden (possibly extinct) (Zhang et al. 2019). Despite being more valuable than other cereals it also possesses a C4 mechanism that effectively concentrates carbon (Hittalmani et al. 2017). Its adaptability in low moisture regions and high temperature could be due to genes implicated in the C4 mechanism (Li et al. 2021). It is important to compare the C4 mechanism network genes to those that have already thoroughly characterized in model crops and reveal the diversity at genomic regions (Saha et al. 2016). Considering finger millet enormous significance in the human diet very few efforts are governed to establish genetic resources and high throughput breeding strategies compared to other cereals. The development of genomic resources has been impeded due to the allotetraploid nature with an unknown B sub-genome. The fact to not discriminating between sub-genomes finger millet could lead to incorrectly calling homologous vs. homoeologous single nucleotide polymorphism (SNP) sites (Hittalmani et al. 2017; Hatakeyama et al. 2018). Establishing crop genetics by delving into molecular resources is key for unlocking desirable traits, with recent advancements in next generation sequencing accelerating genome and transcriptome sequencing of various plants like millets (Fig. 1). The whole genome sequence (WGS) is now available for model plants and numerous cereal crops, even those with more complex genomes. However, the WGS of finger millet has been delayed compared to other major cereals, making it as one of the last cereal genomes to be sequenced. Many agronomic traits of finger millets have been enriched using conventional breeding techniques, and novel candidate genes have also been discovered in the crop using genomic and transcriptomic approaches (Hittalmani et al. 2017). A decade following the draft genome sequence of rice, the first draft genome of finger millet was published (Hittalmani et al. 2017). Although genetic and genomic studies of finger millet have been constrained by the absence of whole genome sequencing (WGS), the recent release of its draft genome is anticipated to significantly advance research. This WGS of finger millet is expected to be a valuable resource for future breeding programs and for gaining a deeper understanding of the crop’s history (Wambi et al. 2020). This analysis delves into the intricate details of various biotechnological studies that have been carried out to shed light on the unrevealed portion of finger millet across different platforms, such as plant tissue culture, genomics, transcriptomics, proteomics, and metabolomics. By gathering insights from diverse finger millet sources like literatures and biological databases, we aim to pinpoint research gaps and suggest potential interventions for enhancing crops through the application of advanced omics techniques. Through the utilization of multi-omics approaches, we can create innovative frameworks that bridge the gap between the genetic blueprint and the observable traits of plants. Those effective information paving the way for a better comprehensive of finger millet genetic makeup relative to the key staple crops which is essential for global food and nutritional security.
Nutritional values of finger millet
Finger millet is acclaimed as a potential ‘super cereal’ by the United Nations Academies because of its exceptional nutrient content and diverse benefits compared to other major cereal crops (National Research Council, 1996). It is notably rich in minerals and offers a higher micronutrient density than both rice and wheat (Devi et al. 2014). Finger millet contains calcium levels that are 5 to 10 times higher than those found in commonly consumed cereals such as brown rice, wheat, or maize, and it is three times more effective than milk (Kumar et al. 2016). It contains phosphorous and iron required for bones, metabolic processes and oxygen and blood circulation in body (Talwar et al. 2020). Finger millet is also rich in minerals like iron and fiber. Production of proteins requires amino acids such as tryptophan and lysine, and these amino acids are lacking in grain endosperm, which typically comprises 1% lysine and 0.35% tryptophan, compared to the about 5% lysine and approximate 1.1% tryptophan, which are required for adequate human nutrition (Maharajan et al. 2022) In comparison with major grains, finger millet possesses a high level of tryptophan (Maharajan et al. 2022). It is also rich in essential amino acids such as methionine and lysine that are required for human health and growth but are deficient in most other cereals (Adeboye et al. 2021). Finger millet also contains the two essential polyunsaturated fatty acids, linoleic acid and linolenic acid, which are important for the human diet (Fernandez et al. 2003a). Their metabolized byproducts support the normal development of the central nervous system (Birch et al. 2007; Jacobson et al. 2008). Additionally, finger millet is rich in vitamins such as thiamin, riboflavin, niacin, and tocopherols, which are crucial for metabolic processes and overall development (Yang et al. 2022a). Broadly, finger millet is packed with antioxidant, anticarcinogenic, antimicrobial, and antidiabetic properties (Fig. 2).
In vitro studies of finger millet
Regeneration in finger millet
Finger millets genetic quality could be improved using both traditional methods and modern approaches like genetic engineering and transgenics. The principal advantages of regeneration of plant and transformation approach in Finger millet improvements includes:
-
i.
The micropropagation of cloned plantlets.
-
ii.
Regeneration of desired transgene containing plant.
Regeneration in finger and pearl millet is often achieved through Shoot Apical Meristem (SAM), which has reservoir of proliferating cells (Eapen and George 1990). The mesocotyl was also utilized as explant for finger millet (Rangan 1973; Heyser and Nabors 1982). The combinations of different plant growth regulators (PGRs) in culture medium determine the regeneration of explant. Most preferably, the use of auxin and cytokinin ratio determines the regeneration and developmental process in finger millet. Previous studies revealed the use of auxin [2,4-Dichlorophenoxyacetic acid (2,4 D) for development and maintenance of callus in finger millet (Srivastav and Kothari 2002). However, the increased level of auxin resulted in lower regeneration efficiency and leads to the abnormal growth (Nabors et al. 1983). Shoot formation was observed in the majority of cases when embryogenic calli were transferred to a medium with a low auxin to cytokinin ratio (Girgi et al. 2002). However, other numerous researches explained about successfully achieving regeneration through the providing cytokinin only in medium (Latha et al. 2005; Nethra et al. 2009).
Transformation in finger millets
Few studies have focused on optimizing genetic transformation techniques for finger millet. Latha and colleagues (2005) reported a reproducible genetic transformation protocol using the gene gun method, where callus derived from seeds was employed for the transformation process. Additionally, Ceasar and Ignacimuthu (2009) utilized shoot apex explants for Agrobacterium-mediated transformation in finger millet. For improving the nutritional value and stress adaptability including drought, saline and high temperature in finger millet (Fig. 3) by using key genes requires an appropriate genetic transformation protocol (Ceasar and Ignacimuthu, 2009). The need of crop improvement approach using genetic engineering is based on efficient genetic transformation protocols to address complex stress responses under adverse conditions. Due to the limited genetic variability among cross-compatible germplasms may restrict conventional techniques approaches. The initial efforts to transform finger millet using the direct biolistic technique were undertaken by Gupta et al. in 2001. They compared and analyzed the effectiveness of five different promoter genes on the expression of a reporter gene. The utilization of genetic transformation for sustainable breeding in finger millet is still in its infancy mostly due to limitations linked to a scarcity of sufficient genomic resources (Hittalmani et al. 2017). The optimized transformation conditions for effective transformation and regeneration were discussed in few researches, the majority of which used the Agrobacterium tumefaciens mediated transformation approach (Table 1).
Improved traits in finger millet through transformation
Biotic stress tolerance
Finger millet plants were genetically engineered to enhance resistance to leaf blast disease by incorporating an antifungal protein (PIN) gene from prawns (Latha et al. 2005). Additionally, the rice chitinase 11 gene (Chi11) was introduced into the GPU45 genotype via Agrobacterium-mediated transformation, which also improved resistance to the disease (Ignacimuthu and Ceasar 2012). Upon comparison with wild plants both studies concluded that the transgenic lines with overexpressing transgenes exhibited enhanced resistance to leaf blast disease. Biotechnology is vital for protecting crop yield and ensuring sustainability in agriculture. It advances breeding techniques and deepens our understanding of plant molecular mechanisms, particularly in addressing devastating diseases such as leaf blast in finger millet. Advanced breeding strategies like Genome Wide Association Studies (GWAS) and gene editing using CRISPR/Cas have become the key to accelerating finger millet improvement programs by enhancing biotic stress tolerance (Mane et al. 2024).
Abiotic stress tolerance
Transgenic finger millet plants have been developed to enhance salt tolerance through different genetic modifications. One method involved overexpressing the sorghum vacuolar H+ pyrophosphatase (SbVPPase) gene (Anjaneyulu et al. 2014). Another approach utilized the Na+/H+ antiporter from Pennisetum glaucum (PgNHX1) and the vacuolar H+-pyrophosphatase from Arabidopsis thaliana (AVP1) to increase salt resistance (Jayasudha et al. 2014).The overexpression of Porteresia coarctata serine-rich protein (PcSrp) gene has shown enhanced tolerance against salinity in finger millet (Mahalakshmi et al. (2006). Furthermore, finger millet overexpressing a bacterial mannitol-1-phosphate dehydrogenase (mtlD) gene showed improved growth under drought and salinity stress condition compared to wild plants (Hema et al. 2014).
Genomic studies of finger millet
Available genomic data in finger millet
Finger millet generally contains fewer genomic resources than other major grains till date according to available data at biological databases (Supplementary Table 1) (https://www.ncbi.nlm.nih.gov; https://phytozome-next.jgi.doe.gov/info/Ecoracana_v1.1), which hinders the crop improvement process. Only 1,934 ESTs were linked to abiotic and biotic stresses in finger millet (Ceasar et al. 2018). The availability of major cereals are much adequate as compare to finger millet. Finger millet does not yet have a complete gene and unigene sequence. Although several genome assemblies are available for other cereals, there is currently only one genome assembly for finger millet (ASM218045v1) (Ceasar et al. 2018). As compared to other major cereal grains, fewer protein sequences have been reported till date (Kayastha et al. 2023). Comparative genomics has been widely applied to explore minor cereals and can effectively contribute to the marker-assisted selection (MAS) approach for addressing various environmental factors in finger millet (Hittalmani et al. 2017). While numerous SSR and SNP markers have been developed in grass genomes, the number reported in finger millet is completely sparse, with only 10,337 SSRs and 23,285 SNPs identified (Gimode et al. 2016). Moreover, reports of abundant genetic resources with several efficient contributions have been made through WGS in finger millet. These resources are easily accessible in the NCBI GenBank database (Table 2).
Genetic diversity analysis by marker-assisted selection (MAS)
Several studies have shown the usefulness of genetic markers in investigating diversity at genetic level and QTLs mapping in finger millet. Genetic diversity analysis is crucial for crop improvement as it provides essential information for breeding lines and linkage studies (Mohammadi and Prasanna 2003). Understanding genetic diversity is vital for assessing genome variation among different genotypes and developing populations for molecular marker analysis. For finger millet, which is cultivated across diverse climatic conditions in Asia and Africa, analyzing genetic diversity is important for evaluating the adaptability of genotypes to various biotic and abiotic stresses. Genotypes that can withstand a range of stresses often exhibit greater allele variation compared to more susceptible ones, making them valuable for breeding programs (Ceasar et al. 2018). Gupta et al. (2010) investigated three finger millet genotypes with different seed coat colors (brown, white, and golden), analyzing their morphological, physiological, and biochemical traits using 10 RAPD and 10 ISSR markers. Patil and Kale (2013) examined genetic diversity among 32 finger millet genotypes and found that RAPD markers revealed greater polymorphism than ISSR markers, utilizing 45 RAPD primers in their study. Additionally, Ramakrishnan et al. (2016) used RAPD markers to evaluate the genetic diversity and population structure of finger millet collected from various geographical regions. Further, several genetic markers were employed to examine 42 finger millet collected from several southern Indian (Rajendram et al. 2016). To create a new breeding population, genotypes containing diversity information can be crossed with elite cultivar and employed as parents of interest. For a high throughput examination of genetic diversity, haplotype-based diversity analysis has also been utilized recently for a number of plants (Varshney et al. 2021).
Identified QTLs for agronomical important traits in finger millet
The agronomical traits of finger millet, such as yield, disease resistance, drought tolerance, and nutritional enhancement, have been identified using microsatellite markers (Ceasar et al. 2018). The application of association mapping-based discovery of QTLs helped in development of biofortified finger millet (Kumar et al. 2015). The analysis conducted on 113 finger millet genotypes using 23 anchored SSR markers revealed a total of 9 QTLs linked to Ca content (Kumar et al. 2015). Hence, it is crucial to identify the QTLs controlling these traits and the candidate genes responsible for deviation in Ca accumulation to ensure their effective integration into breeding and transgenic approaches. An association mapping study using SSR markers has identified four QTLs for finger blast resistance and one QTL for neck blast resistance in finger millet. (Babu et al. 2014a). Additionally, the genomic markers RM262, FMBLEST32, and UGEP81 were found to be linked with finger blast (Babu et al. 2014). Additionally, in the same year, Babu et al. (2014b) identified two QTLs for tryptophan content and one QTL for protein content in the aforementioned finger millet genotypes using 120 SSR markers. The importance of mapping blast disease resistance along with agronomic traits was well described (Ramakrishnan et al. 2016). Employing association study, 7 QTLs associated with agronomic traits and blast disease resistance were identified and advocated their use in further MAB programme in finger millet (Ramakrishnan et al., 2016). Despite the importance of finger millet, its genomic resources still lag behind those of major cereal crops, which is why the their genetic architecture of agronomically important traits has remained elusive (Banerjee et al. 2020). The genomic regions governing important agronomic traits and blast resistance in finger millet were recently explored through genome-wide association mapping. This study utilized genotyping-by-sequencing (GBS) to generate 2977 high-quality single nucleotide polymorphism (SNP) markers, resulting in the identification of four common marker-trait association linked to neck blast resistance. Notably, three of these marker-trait associations for neck blast resistance were found to have orthologous in related genera, indicating potential involvement in plant pathogen defense (Sood et al. 2023). Till date, finger millet has low genomic information especially number of SNPs identified compare to major cereals (Bančič et al. 2024). Genotyping-by-sequencing were utilized to find 5226 SNP markers with the above 5% minimum allele frequency (MAF) without missing reads used for genetic diversity analyzes of the finger millet (Brhane et al. 2022). Similarly, the transcription factor family such a nuclear factor Y (NFY) that governs for diverse genes controlling the stress tolerance were studied (Rani et al. 2023).
The significance of identifying candidate genes in finger millet
Among all cereal grains, finger millet has tremendous potential due to its numerous qualities. The role of potential genes involved in multiple purposes such as nutritional enrichment, acclimation to adverse conditions, and other beneficial bioenergetic pathways in finger millet are predicted to be effective by current advancements in genomic research (Sood et al. 2016). The status of the initiated studies that provide the importance and role of the expression of candidate genes in finger millet is summarized in Table 3.
Candidate genes involved in calcium (ca) transport
The expression analysis of several Calcium (Ca) transporter related genes was analyzed in contrasting genotypes GP-1 (low Ca level) and GP-45 (high Ca level) (Mirza et al. 2014). Another study was located about calcium sensor genes through the transcriptome analysis in developing spikes of finger millet (Singh et al. 2014). In order key genes in Ca transport, genomic and transcriptomic sequencing were performed in the developing spikes of finger millet (Singh et al. 2015). Drawing from the expression and bioinformatics studies, EcCIPK24 play a pivotal role in high Ca accumulation in finger millet seed (Chinchole et al. 2017). A more thorough understanding of the intricate mechanisms governing Ca transport in finger millet can be attained through the application of high-resolution investigations, such as multi-omics approaches. In order to achieve this, newly popular gene editing methods such as CRISPR/Cas9 might be useful in collaboration with genome sequencing to achieve desirable effects (Ceasar et al. 2016).
Candidate genes plays role in the nitrogen (N) metabolism
Expression analysis of genes related to nitrogen uptake and assimilation was conducted in finger millet using contrasting genotypes (Gupta et al. 2013). Expression analysis of EcDof1 and EcDof, which are influenced by light and N concentration, demonstrate a complex regulation pattern in N regulation in finger millet (Gupta et al. 2014).
Candidate genes plays role in phosphate (P) transport
Phosphorus, an essential nutrient for plant growth and development, is absorbed into plant cells via phosphate transporter proteins (Pudake et al. 2017a). Along with phosphate transporter proteins, symbiosis with arbuscular mycorrhiza play vital role in the absorption and transportation of phosphorus from soil (Pudake et al. 2017a). EcPT4 is an arbuscular mycorrhiza-specific phosphate transporter gene in finger millet whose expression was found to be correlated with its colonization in the roots. In finger millet, only four EcPT genes have been characterized thus far. However, in Arabidopsis, ten phosphate transporter genes have been reported (Baker et al. 2015), indicating the potential for identifying more of these genes in finger millet. Although closely related, foxtail millet has been found to contain 12 phosphate transporter (PT) genes, which were studied for their expression patterns, phosphate transport assays in yeast, and in planta function through RNAi-induced downregulation (Ceasar et al. 2014, 2017). Among these, four PT genes were identified in finger millet using partial transcript sequences.
Candidate genes in abiotic stress tolerance
Enhancing abiotic stress tolerance in crop plants is one of the key objectives of crop improvement, including for finger millet (Rakkammal et al. 2023). A plethora of reports suggest severe loss in yield of finger millet due to drought stress (Krishna et al. 2021; Mwangoe et al. 2022; Gebreyohannes et al. 2024). The growth and yield of finger millet are severely affected by biotic and abiotic stresses, including drought stress (David et al. 2021; Krishna et al. 2021; Maqsood and Ali, 2007; Parvathi et al. 2013). EcDehydrin7, a drought responsive gene was isolated from finger millet (Singh et al. 2014) and its role in drought tolerance was demonstrated by overexpressing it in tobacco (Singh et al. 2015). Expression analysis of finger millet under drought stress, employing a novel gravimetric approach, led to the identification of stress-responsive genes (Parvathi et al. 2013). Few transcriptomic profiling was performed to understand the role of drought specific genes in finger millet (Ramgowda et al. 2017; Parvathi et al. 2019). RNA-seq analysis from the leaf tissue of finger millet (GPU-28 variety) under drought stress condition at field level identified a set differentially expressed gene (Parvathi et al. 2019). This analysis also identified regulatory genes viz. PP2A, CIPK31, FPS and SRPRα associated with drought stress in finger millet (Parvathi et al. 2019). Few transcriptomic analyses were performed to understand the role of drought responsive genes in finger millet (Ramgowda et al. 2017; Parvathi et al. 2019). TBP Associated Factor 6 (EcTAF6), a regulator of drought response, was discovers by constructing cDNA library of the plant (Parvathi and Nataraja 2017). Comparative transcriptomics profiling was carried out on contrasting genotypes of finger millet against salinity stress to identify abiotic stress related genes, such as EcNAC67 TF (Rahman et al. 2016). EcNAC1 gene was discovered to be significantly upregulated in response to salt stress and was associated with resistance to other abiotic stimuli (Ramegowda et al. 2012). CBL interacting protein kinase 31 gene was discovered and characterized to confer drought resistance in finger millet (Nagarjuna et al. 2016). TBP-associated factors 6 (TAFs) gene was also identified under the drought stress condition in finger millet (Parvathi and Nataraja 2017). Finger millet Overexpression of EcbZIP17 gene from finger millet in tobacco were showed tolerance to heat and salinity stress (Ramakrishna et al. 2018) (Table 3).
Genes associated with biotic stress
Researchers have continuously discovered and characterized numerous biotic and anti-fungal proteins (AFPs) synthesized or secreted by host plant against evading pest and pathogens (Gupta et al. 2017). Prolamin is a seed storage protein in association with other proteins, tackles insects and proteases activity and acts as a barrier to their invasions and digestion of the plant (Gupta et al. 2011). The secretion of protease inhibitors by finger millet and other crop grains hinders the digestive enzymes in the insect gut, thereby controlling the population of phytophagous insects (Gatehouse and Gatehouse, 1998). α-amylase/trypsin inhibitor (RBI), a bifunctional protease inhibitor has been isolated and characterized in finger millet (Strobl al., 1995). Subsequently, RBI was cloned in E. coli, and a notable rise in RBI transcript levels was observed in the leaves of finger millet when infected with Rhizopus oryzae and Curvularia lunata (Sen and Dutta 2012). Recognizing QTLs connected to crucial physiological traits like blast resistance is advantageous for molecular breeders, looking to integrate these genes into locally well-adapted germplasm. The identification of molecular markers linked to blast resistance enables the development of blast-resistant genotypes through marker-assisted selection (Panwar et al. 2011). Babu et al. (2024) used a comparative genomics approach to identify genes/QTLs linked to the blast, a most devastating disease of finger millet. In rice, nine R genes, known to confer blast disease resistance, belong to nucleotide binding site-leucine rich repeat (NBS-LRR) family (Panwar et al. 2011). Based on homology searchers using rice NBS-LRR region as bait, finger millet NBS-LRR regions were also identified, showing similarity in their mode of action against blast disease (Panwar et al. 2011; Babu et al. 2014). Association mapping studies led to the identification of five key QTLs associated with finger blast and neck blast (Babu et al. 2014). The finger millet plant’s neck and finger blast resistance genes are located on the second and sixth linkage groups, respectively, suggesting that these chromosomes are the primary centers for blast resistance genes. In finger millet, the genes associated with neck and finger blast resistant were found to be situated on the second and sixth linkage group, respectively, suggesting that these chromosomes are the main hub for blast resistant genes (Babu et al. 2014b). Breakthroughs in recombinant DNA technology and molecular tools have greatly facilitated the establishment of finger millet as a model system for studying interactions between plants and microbes. Gupta et al. (2001) conducted the initial investigation into transformation in finger millet, assessing the effectiveness of five gene promoters for enhancing the expression of target genes in the plant. Cloning, and overexpression of PIN protein into finger millet resulted in increased resistance against leaf blast (Latha et al. 2005). With the help of rDNA technology, chitinase genes was isolated, cloned and overexpressed in crop plants including finger millet to combat fungal pathogens (Cletus et al. 2013). Phytohormones signaling play a pivotal role in understanding disease resistance mechanism in crop plants. In finger millet, the jasmonic acid signaling pathway was studied using the Jasmonate ZIM-domain (JAZ) protein family (Sen et al. 2016).
Transcriptomic studies
A hypothetical model was utilized to analyze openly accessible gene expression data, leading to a better understanding of the transport and calcium supply in embryonic seeds (Goel et al. 2012). Mirza et al. 2014 reported that the finger millet genotype, recognized for its high calcium content, exhibited significant expression levels of transporter genes during the grain filling stages. As discussed by Hittalmani et al. (2017), the draft of the ML-365 complete genome spanned 1196 Mb, representing approximately 82% of the expected genome size. Genome analysis revealed the presence f 85,243 genes, highlighting the repetitive nature of the genome. The de novo assembly of genotype GPU-28 yielded 80,777 and 90,830 transcript, which were compared between control and drought stressed areas (Parvathi et al. 2019). Differential expression of genes (DEGs) and proteins (DEPs) in response to drought stress were identified at the transcriptome and proteomic levels (Li et al. 2021). Transcriptomic sequencing data from finger millet leaf tissues yielded an average of 24 million reads. A total of 684 transcripts exhibited DEGs between heat stressed samples of both genotypes (Goel et al. 2023). Additionally, the transcriptomic sequences of finger millet are limited in biological databases and were collected under less stress effects. Only specific genotypes of finger millet underwent comprehensive transcriptomic sequencing, particularly under stressors like blast, salinity, and drought (Supplementary Table 2).
Epigenomics
Epigenetic changes such as DNA methylation and histone methylation have been extensively studied in pear millets. The significance of salicylic acid in root growth and seedling development has been revealed through DNA methylation studies. An innovative approach using methylation-sensitive amplified polymorphism analysis to investigate the cytosine DNA methylation profile in pearl millet (Ngom et al. 2017). In foxtail millet, the CHH DNA methylation context gradually increases during grain filling stages. This specific methylation pattern in CHH sites suppresses genes associated with carbohydrate and protein metabolism to enhance grain development (Wang et al. 2021). Furthermore, an epigenetic study in foxtail millet showed a notable decrease in DNA methylation levels in salinity-tolerant genotypes compared to susceptible ones. Tolerant genotypes displayed DNA methylation alterations in various genes, including ABC transporters, WARKY transcription factors, and serine threonine-protein kinases, under prolonged salinity stress conditions (Pandey et al. 2017).
Proteomic studies
The investigation focused on the expression of PBF dof TF in finger millet tissues with varying properties such as seed protein content and spike coloration. The data revealed an increase in the expression levels of all three genotypes during spike development ompared to each other. Interestingly, a strong correlation was observed between higher levels of PBF dof expression at the beginning of development and the protein content in grains (Gupta et al. 2011). Using peptide mass fingerprinting, calreticulin, a calcium-binding protein, was identified in developing finger millet spikes (Singh et al. 2015). While some studies have highlighted proteins associated with calcium saturation in finger millet, there is a lack of protein profiling specifically for nutritional proteins (Singh et al. 2023). Examination of finger millet genotypes rich in calcium focused on the embryo and aleurone layer, revealing an increase in calcium accumulation during grain filling in developing seeds. The ability of millet to withstand stress remains unexplored at proteomics level. This lack of understanding may be attributed to the scarcity of genomic data, although ongoing scientific advancements continue to raise hopes of achieving sustainable agriculture objectives.
Metabolomic and ionomic studies of finger millet
Various analytical techniques such as Nuclear magnetic resonance (NMR), electrospray ionization mass spectrometry (ESI-MS), and high-performance liquid chromatography (HPLC) have been employed to characterize the polyphenols present in finger millet. These polyphenols include cinnamic acid derivatives (such as syringic, trans-cinnamic, and p-coumaric acids), benzoic acid derivatives (including gallic, protocatechuic, p-hydroxybenzoic, vanillic, and ferulic acid), and quercetin (a flavonoid) (Chandrasekara and Shahidi 2010; Banerjee et al. 2012). It is imperative to expand this knowledge to encompass other nutritionally significant metabolites to enhance the nutritional value of plants. An investigation on Magnaporthe grisea infected ragi samples collected from leaves, necks, and fingers utilized Thin Layer Chromatography (TLC) to detect volatile compounds in the infected plant extracts. Furthermore, Gas Chromatography-Mass Spectrometry (GC-MS) analysis was employed to determine the volatile compounds in the crude metabolite extracts from the infected plant parts affected by blast (Shanmugapackiam et al. 2019).
Multi-omics approaches to improve the genetic gain of finger millet
Transgenic plant production
The success of plant transformation is contingent upon the effectiveness of regeneration. Therefore, determining the most suitable technique for each plant species and ecotype is imperative before initiating the transformation process. Several transgenic studies in finger millet have demonstrated that proper regeneration can be achieved using shoot apical meristem as an explant (Table 1). Characterization millets using standard tissue culture and transgenic protocols will enhance our understanding of their resilience to various climate conditions. In-depth investigations on finger millet, such as the cellular localization of transgenes and the construction of recombinants using promoters with GUS and GFP, have been conducted (Table 1). These specific promoters may play a crucial role in identifying important genes and tracing the signals involved in nutrient accumulation in grains, as well as biotic and abiotic tolerance (Fig. 4). The application of recombinant DNA technology and cutting-edge genome editing tools, combined with the recently unveiled draft genome, may play a crucial role in advancing the genetic progress of crops. This will allow for the assessment of findings in comparison to analogous studies in model plants for the examination of candidate gene expression.
Plant system biology approach for multiplexing effect at genomic level
Enriching the genomic resources for genetic diversity
The journey of multi-omics techniques through comparative genomics proposes orthologous GWAS for candidate gene exploration in finger millet. The preliminary genome information of model cereal crops will be beneficial for relative genomics, which might assist in discovering alleles regulating a variety of crucial characteristics in finger millet. Nevertheless, ongoing exploration through multi-omics approaches is useful in intricate genetic structure analysis and unleashes full potential for superior cultivar development in finger millet (Sharma et al. 2022). Advanced and focused investigation is crucial to completely comprehend the array of molecular markers and significant candidate genes associated with desired traits in finger millet (Fig. 5), even though these studies provide enlightening information about the genetics of the crop. The manners in which millet get unveiled in case of nutritional security and climate resilience such methods are implementing such methods in future farming systems may better equip them to withstand adverse climatic conditions and ensure food security (Rani et al. 2023).
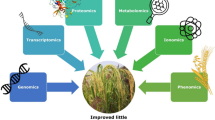
Scientists are increasingly turning to multi-omics approaches, such as genomics, transcriptomics, and proteomics, to explore the extensive genetic diversity in finger millet beyond conventional understanding. These methodologies provide novel insights into molecular diversity within breeding populations by bridging the gap between genotype and phenotype. They enable more precise marker identification and enhance phenotypic predictions (Weckwerth et al. 2020; Pazhamala et al. 2021). Associative transcriptomics investigates how DNA sequence variation correlates with both phenotypic variation and transcript abundance in crops like finger millet (Harper et al. 2012). Studies on cis-expression quantitative trait loci (eQTLs) support the understanding of phenotypic variability in maize, including aspects related to domestication and adaptability (Lemmon et al. 2014). Transcriptome-wide association studies (TWAS) and expression read depth genome-wide association studies explore the relationship between mRNA expression levels and phenotypic traits (Lin et al. 2017); Kremling et al. 2019). Through GWAS and comparative genomics with rice, conserved genetic regions controlling nutrient accumulation in finger millet have been identified (Puranik et al. 2020). Unlike genetic variations affected by random allelic effects, transcript levels provide deeper insights into the fundamental regulatory mechanisms governing quantitative traits and aid in identifying causative genes (Kremling et al. 2019). Furthermore, whole transcriptome analysis offers opportunities to elucidate potential pathways influencing nutritional quality in finger millet (Mirza et al. 2014; Hittalmani et al. 2017). Conducting global transcriptome analysis is crucial for comprehending mechanisms related to other biological variables impacting dietary quality. Epigenetic studies and advanced proteomics techniques are instrumental in enhancing the precision of protein discovery in finger millet (Fernandez et al. 2003a). Recent proteomics innovations, such as Matrix-Assisted Laser Desorption/Ionization Time-of-Flight (MALDI-Tof), HPLC, GLC, coupled with mutagenesis applications, offer effective means to uncover complex traits associated with specific genes or markers. These technologies play a crucial role in the breeding process, where high-throughput phenotyping and sequencing approaches can effectively screen numerous lines (Weckwerth et al. 2020). Metabolomic technologies, which are often more cost-effective than NGS methods, are particularly valuable in finger millet for comprehensive metabolome analyses. This approach allows the identification of biomarkers associated with nutritional traits and provides critical insights for targeted breeding and improvement strategies (Singh et al. 2022; Dhanushkodi et al. 2023).
Sequencing technologies for understanding genomic variation
Until recently, advanced molecular biology tools like NGS were primarily utilized in major commercial crops. However, these high-throughput technologies are now applicable to orphan or neglected crops such as finger millet. This enables the analysis of molecular markers, comparative studies using transcript sequences, genetic diversity deciphering, and WGS of crops. High-confidence variant calling and assembly of extended contigs are facilitated by high-quality reference-genomes, significantly enhancing genetic investigations like gene identification and modification. Polyploid crops like finger millet, techniques such as Hi-C sequencing and Bio-Nano genomics optical mapping have revolutionized genome assembly and improving SNP calling, facilitating haplotype breeding with high fidelity. The integration of trait-linked SNP genotyping and molecular assessment of genetic distance during parent line creation has proven beneficial in maximizing yield through inbreeding and cross-breeding strategies across various plant species (Baggett et al. 2021). Furthermore, the application of NGS and GBS in millet breeding enhances breeding efficiency by elucidating the impact of changing environmental conditions on crop performance. While SSRs have historically been the preferred markers for assessing genetic diversity, their sequencing process has been labor-intensive. Advances now allow for the validation of causative regions influencing heterotic groups more efficiently (Baggett et al. 2021).
High throughput phenotyping
Recent advancements in plant phenotyping have greatly benefited from breakthroughs in sensor technology and image capturing techniques, leading to significant improvements in cost-effectiveness and efficiency (Mir et al. 2012). Although automated platforms have yet to be fully utilized in studying crops like finger millet, these technologies facilitate sensor-to-plant and plant-to-sensor interactions, enabling dynamic monitoring of plant responses at various levels-from organs to entire fields. Innovative applications such as 3D modeling with X-ray computed tomography are reducing delays in subsurface phenotyping, allowing for in situ assessment of root morphology. The evolution towards digitalized and AI-driven field-based phenotyping (see Fig. 5) reflects the growing need for precise monitoring of plant performance under real-world conditions. Aerial platforms, including manned and unmanned aircraft and satellites, play a pivotal role in phenotyping by offering variable payload capacities and high-resolution imaging capabilities (Jin et al. 2020). These platforms enhance the ability to capture comprehensive data on crop growth and environmental interactions across large spatial scales.
Utilization of Germplasm resources for Mining Superior haplotypes
The untapped evolutionary data within millet germplasm collections present a significant contrast to the comprehensive studies conducted on other major grains. The genetic diversity and taxonomic relationships within the germplasm pool of finger millet remain largely unexplored, hindering efforts to identify useful traits for crop improvement. Addressing these gaps is crucial for securing future agricultural sustainability by uncovering important key genes and alleles in plant genetic resources, including germplasm lines and landraces stored in repositories. In the upcoming era, there is a pressing need to employ morphological, cytogenetic, and numerical taxonomy techniques to systematically study diversity and relationship in finger millet. Current enrichment programs for finger millet have not fully capitalized on the potential benefit of integration of diverse alleles into breeding efforts through advanced genetic and genomic research. Establishment core collections of germplasm, encompassing the broad spectrum of new combinations found in both wild and cultivated varieties facilitates the identification of key alleles for desirable traits. Successful quantitative trait studies (Fig. 5) have been achieved through various genetic populations, including MAS, GWAS, and multi-parental populations such as Multi-parent Advanced Generation Inter-Cross (MAGIC) and Nested Association Mapping (NAM). Recent advancements in sequencing and genotyping technologies enable the evaluation of Plant Genetic Resources (PGRs) and genetic populations across environmental conditions and growing seasons, assessing their agronomic and nutritional properties (Varshney et al. 2020). The application of pangenomics and GWAS methodologies holds promise for identifying superior alleles or genes associated with desirable attributes. Genomic breeding approaches must be embraced to accelerate the enhancement of agriculturally significant crops like finger millet.
Improving genetic gain by genomic selection approach
Genomic selection (GS) represents a groundbreaking breeding strategy that identifies superior genotypes based genomic estimated breeding values (GEBVs), derived from genotypic data distributed uniformly across the genome of a training population. This approach is particularly effective in a characterizing genotype within inbred populations and is essential for accelerating the genetic improvement of neglected crops. To optimize the selection process and hybrid prediction capabilities, three GS model were compared using different grain genotyping platforms: traditional GBS, RAD-seq, and tunable GBS. In sorghum, multi-trait analysis has demonstrated improved GS for enhancing yield and tolerance to abiotic stresses. In the context of finger millet, GS has emerged as a crucial tool for enriching genetic gain and facilitating germplasm improvement. Breeding programs now utilize genotypic information and GEBVs to identify candidate offspring with the potential to enhance line improvement. The calculation of GEBVs integrates advanced genome-wide sequencing with precise phenotypic data, expediting genetic progress in breeding efforts. This approach enhances our ability to assess genetic significance and accelerate the genetic gain in targeted crops. The advent of sequencing technologies has generated vast amounts of marker data, challenging their inclusion in prediction models. GS mitigates this challenge by determining GEBVs through comprehensive genome-wide marker profiling, enabling line selection before field phenotyping and thereby shortening the breeding cycle. A key advantage of GS lies in its ability to expedite parental selection, which is crucial for determining the genetic merit of tested individuals. GS model incorporates additional environmental variables and their correlation with genotype to enhance the accuracy of GEBV predictions across diverse contexts. This comprehensive approach underscores the efficacy of modern breeding strategies aimed at enhancing crop genetic improvement.
Pangenomics
The concept of the pangenome has gained popularity with the sequencing of numerous plant genomes. A pangenome consists of core genes, which are present in all selected individuals, and dispensable genes, which are absent in at least one individual. Understanding the pangenome allows us to tailor the ideal gene content for crop varieties, optimizing breeding outcomes. This approach holds immense potential for finger millet by providing insights into and identifying crucial genes. Super-pangenomes encompass core genes and haplotypic variations, are essential for advancing pangenomics to breed the next generation of crops. Plant breeders can transition from traditional progeny selection to utilizing causative allelic information derived from pangenomics (Fig. 5). These methodologies provide valuable haplotype information to create superior combinations of genes and haplotypes. Researchers can use this approach to facilitate the identification of QTLs and genes associated with significant agronomic traits. These gene data may enhance our ability to harness genetic diversity and accelerate the development of improved finger millet varieties tailored to meet agricultural and nutritional demands.
A strategy of haplotype-based breeding technique
Utilizing pangenomics, GWAS and system biological approaches offer the potential to identify superior haplotype related to agronomically important trait like stress resilience and complex trait such as yield in finger millet. Integrating these methods with breeding and multi-omics approaches could accelerate the development of superior cultivars. In genomics-assisted breeding (GAB), successful strategies such as backcrossing and MAS have been pivotal in generating improved cultivars (Varshney et al. 2021). Most of these efforts have employed biparental population to pinpoint DNA markers associated with quantitative traits. Advances in whole genome sequencing coupled with extensive phenotypic data now enable the identification and validation of effective alleles through breeding programs. This process involves accurately assembling recombinant haplotype that exhibit desired phenotypes. Studying the natural diversity within finger millet, a particularly vulnerable cereal crop, provides insights into its genetic architecture and enhances our understanding of genotype-to-phenotype interactions. This approach is poised to revolutionize the identification of key traits crucial for crop improvement. NGS approaches have proven effective in analysing genetic diversity and establishing marker-trait associations (MTA)s in various crops, particularly through the accumulation of genomic resources such as SNPs (Sehgal et al. 2012). These resources form a substantial repository for enhancing our understanding of genetic variability and trait inheritance in major grains. Overall, leveraging these advanced genomic and computational tools holds immense promise for accelerating the development and unrevealing of superior finger millet varieties tailored to meet future agriculture demands.
Mutation breeding approach
The mutational breeding approach facilitates the recombination of new genetic lines to uncover valuable genetic information. In pearl millet, mutational breeding has been explored for enhancing agronomic traits through treatment like ethyl-methane sulfonate (EMS) on seeds. The Target Induced Local Lesions in Genomes (TILLING) approach has similarly been employed across various crop plants to identify unique allelic variants associated with nutritional quality and stress tolerance (Kayastha et al. 2023). Additionally, NGS opens avenues for integrating advanced biotechnological methods with mutant breeding techniques such as MutMap, MutMap+, SHOREmap, Mutmap-Gap, and QTLseq (Varshney et al. 2020). These approaches have the potential to expedite improvement programs (Fig. 5) in neglected crops like finger millet. The diversity of these methodologies is crucial for gaining insights into genetic information and understanding agronomically significant traits. Overall, combining mutational breeding with sophisticated biotechnological tools holds promise for accelerating the enhancement of finger millet and other underutilized crops. This approach may enable targeted improvements in traits essential for agricultural sustainability (Kayastha et al. 2023).
Genome editing based fast forward breeding
Genome editing (GE), particularly utilizing CRISPR/Cas9 technology, has revolutionized plant genome manipulation. Technological advancements in GE have expanded its application to various crops, including the development of point mutations in field-grown crops through cytosine and adenine base editors (Yan et al. 2018). Traditional CRISPR/Cas9 delivery methods primarily rely on DNA-free GE techniques, which present a promising alternative for creating modified crops with reduced risk of off-target effects. Recent innovations in GE include the integration of search-and-replace editing, an enhancement over single base substitutions, allowing for small insertions, deletions, point mutations, and combinatorial edits (Anzalone et al. 2019). While this technique has only been tested in human cells, it is expected to be effective in plant cells as well. This advancement brings researchers closer to achieving precise gene content management in crops. GE plays a crucial role in regulating gene expression in plants and is a promising approach for developing improved varieties. Techniques involving Cas9, Cas12a, and Cas12b are used to modify transcript function, holding significant potential for advancing agricultural practices. To date, GE in millets has been limited, with only two successful cases reported in foxtail millet. Arabidopsis and rice are well-established model plants for C3 photosynthesis, yet a comparable model for C4 plants was previously lacking. However, Yang and colleagues have recently established foxtail millet as a model for C4 plants, a development that, combined with advancements in CRISPR/Cas technology, is set to transform plant research and agricultural improvement (Peng et al. 2021). The application of GE in millets has faced challenges due to the complexity of millet genomes, limited regeneration protocols, and delayed genome sequencing. However, ongoing research on finger millet to enhance resistance to fungi and other biotic stressors is crucial for targeting specific genes for genetic modifications and improving grain yields. This research aims to develop robust resistance against various pests and pathogens by elucidating the molecular mechanisms of resistance in finger millet grains. Although single antifungal genes have provided modest resistance, creating more robust resistance to infections remains a challenge. Future efforts should focus on gene pyramiding or advanced CRISPR/Cas approaches to enhance resistance to a range of infections. The continued refinement of editing capabilities and transformation techniques may significantly advance genetic improvement strategies for finger millet.
Single cell omics approach
Single-cell omics techniques have revolutionized biological studies by providing unique insights into cellular complexities. Unlike traditional bulk sequencing, single-cell sequencing can identify individual cell characteristics, unveiling various cell types, dynamic states, and rare cell groups hidden in bulk data. Driven by high-throughput technologies, this method offers a detailed view of genomes, transcriptomes, proteomes, and epigenetics, revealing hidden intricacies in cellular landscapes. Single cell genomics sheds new light on biological functions by examining genetic variants at the cellular level, despite facing challenges due to limited DNA from single cell. Advances in genomic research have led to the development of Whole-genome amplification (WGA) technologies like degenerate oligonucleotide primed (DOP)-PCR and multiple displacement amplification (MDA). Although these methods are prone to artifacts, they have paved the way for innovations such as primary template-directed amplifications (PTA) and Multiplexed end-tagging amplification of complementary strands (META-CS) for more accurate single-cell genome analysis. Microfluidic-based WGA techniques are emerging as promising alternatives offering automation, integration and simplification of workflows for single-cell genomics. This is exemplified by single-stranded sequencing using microfluidic reactor (SISSOR), which hallows for precise DNA isolation and amplification in nanoscale compartments. The fusion of diverse omics data in one cell offers a comprehensive look at cellular processes, aiding in uncovering intricate interactions and networks. This ultimately advances our knowledge of cellular system and the plant-disease interactive nature (Lim et al. 2024).
Deep Learning and Artificial intelligence
The recent application of AI, convolutional neural networks, and support vector system to identify and predict plant biotic stressors using high-throughput imaging and hyperspectral data represents a sophisticated approach that can enhance our understanding of the genetic nature of finger millet under adverse conditions. Recent improvements in machine learning (ML) have been driven by advancement in high-throughput image recognition (Van Dijk et al. 2021). Artificial intelligence (AI) and deep learning utilize statistical methods and machine learning (ML) enabling computer systems to learn from previous data and predict the outcomes of input data (Samuel et al. 1959) (Fig. 5). Systems biology, a multidisciplinary method that integrates computer science, mathematics, and biological sciences aims to quantitatively analyze the abundance of transcriptomics, proteomics and metabolomics datasets to anticipate informative pathway. ML is crucial for correlating cellular processes with molecular mechanisms involving causal genes and functional pathways, connecting integrational information across various omics networks. In this context, systems-level elucidation of the genes and their intricate functional interactions will benefit from data-driven network analysis techniques like multiplex network and interconnected network (Hass et al. 2017). High-throughput phenotyping data provide a big data challenge, necessitating enhanced computational ecosystems supported by deep ML tools for data storage, processing, and analysis. These tools are essential for drawing insightful conclusions from multidimensional datasets. The effectiveness of ML tools has been evaluated for genome-wide predictions, including their involvement in genotype-phenotype relationships (Liu et al. 2016). Applications of ML is in advanced plant science range from characterizing genomic areas and genome function to understanding the complexity of plant responses to stressors (Van Dijk et al. 2021). However, the main obstacle to utilizing ML for faster crop breeding is the existing dearth of high high-quality labeled data on large populations. This bottleneck might be alleviated with the aid of automated ML techniques and synthetic data synthesis. Cross scale meta-analyses must be developed through these multiscale AI investigations. Accelerating breeding requires both the application of cutting-edge methods like GS and ML, and a better understanding of the genomic basis of agronomically important plant trait.
The optimal contribution selection method for highlighting finger millet agricultural values
In fragile ecosystems, employing advanced improvement programs in major cereals to enrich the genetic base with exotic alleles is crucial. Efficient approaches, such as optimal contributions selection (OCS), can accelerate the harnessing of exotic alleles from the allelic diversity pool. OCS can strike a balance between revealed allelic information and genetic relatedness in exotic crosses with elite populations. Various OCS approaches can maximize contributions from individuals across multiple breeding cycles based on different types of machine learning (ML) (Woolliams et al. 2015). However, this strategy has not yet been applied to the development of finger millet. Genetic algorithms used to create evolutionary algorithms (EA) are significant applications for ML. Several ML and deep learning (DL) algorithms can be combined with EA-based search strategies to aid in optimization. One such advancement is differential evolution (DE), which can optimize the selection process to address the challenging issue of mate selection in animal and plant breeding (Kinghorn et al. 2011). Using pedigree and genomic data, DE-based OCS was used to simulate genetic improvement of complex traits in self-pollinating grain crops (Gorjanc et al. 2018). This methodology was applied to simulate genetic improvement in wheat for heat stress tolerance, grain yield, disease resistance, and other complex traits, aiming to safeguard wheat yields over the next 60 years of anthropogenic climate change (Cowling et al. 2019). This method has an evolutionary impact and can improve and clarify the impact of agronomically important traits in finger millet. A simulated stacking of advantageous wheat haplotypes demonstrated the importance of genetic algorithms in crossing designs for long-term genetic gain. ML can shorten cycle times and speed up breeding by optimizing mating strategies using non-inbred lines (Cobb et al. 2019). Although non-inbred lines often have less accurate prediction than pure lines, this can be improved by using pedigree or genomic data, as seen in animal breeding. To predict long-term genetic development in selfing crops based on F1 recurrent selection, Cowling et al. used pedigree information, and Gorjanc et al. (2018) used genomic information in conjunction with OCS. When considering an additional year each cycle for selfing the F3, the time required to double the economic index using F1 and F3-derived recurrent selection in combination with OCS was comparable. These investigations reaffirm the importance of non-inbred selection for ‘fast-forward breeding’ that combines OCS with abiotic stress tolerance to increase grain yield. This approach, integrating advanced genetic algorithms and ML with traditional breeding strategies, holds great promise for accelerating the genetic improvement of finger millet and other underutilized crops.
Speed breeding
Advanced omics breeding techniques discussed in intervention strategies can be leveraged to improve traits that shorten the generation cycle time, thereby increasing the rate of genomic information (Watson et al. 2018). Advanced platforms, including the fast generation cycling system (Yan et al. 2017) and speed breeding, have been promoted to address issues with in vitro genotype dependence, the necessity for specialized facilities, and the desire to lower costs per plant for large-scale breeding. To enhance breeding programs, the speed breeding strategy should be applied to finger millet. Speed breeding conditions can be achieved in all genetic improvement programs, such as by extending the photoperiod to increase generation turnover by one or two generations annually. Recently, more sophisticated single seed descent (SSD) platforms have emerged, utilizing high-quality controlled environments and photoperiod control with light quality manipulation to exploit prehistoric plant light responses, like shade avoidance, to achieve synchronized flowering across various phenotypes. The ideal conditions for prompt elicitation of blooming time differ by species, yet there are opportunities within related species to develop universally applicable growth conditions. Accelerated SSD techniques are more effective than traditional in-season generation turnover and are particularly useful when multiple traits with varying heritability must be selected simultaneously. Breeding strategies like optimal contributions selection (OCS), haplotype-based breeding (HHB), genomic selection (GS), and genome editing (GE) can be integrated with speed breeding. For instance, simulated studies indicate that using GS in conjunction with rapid gene fixation can shorten the breeding cycle and increase genetic gain per unit time (Jighly et al. 2019). This strategy represents an evolutionary step in genetic improvement and a potent new integrative method for plant development, particularly in underutilized species like finger millet and other millets.
Conclusion
Recent studies emphasize that despite finger millet’s resilience to climate change, it still encounters challenges like salinity, nutrient deficiency, drought, and fungal infections in harsh conditions. Innovative approaches such as multi-omics methods and molecular breeding are unveiling how genetic and environmental factors influence finger millet traits. Previously, the exploration of finger millet’s functional genes was limited without whole genome sequencing. However, with multi-omics strategies, including epigenomics, transcriptomics, proteomics, and metabolomics, whole genome sequencing can now reveal genetic components and pathways related to nutrient accumulation and stress tolerance. Exploring these genetic inquiries can be effectively carried out on model plants such as Arabidopsis thaliana, rice, and wheat, laying the foundation for upcoming studies on finger millet. Utilizing innovative genomics instruments, this analysis outlines a path for enhancing finger millet varieties by pinpointing proteins associated with crucial agricultural traits, conducting haplotype examinations, and expediting breeding procedures.
Data availability
No datasets were generated or analysed during the current study.
References
Adeboye AS, Babarinde G, Oyedepo J et al (2021) Smart crops for climate change and food security in Africa. Food Secur Safety: Afr Perspect 415–449
Anjaneyulu E, Reddy PS, Sunita MS et al (2014) Salt tolerance and activity of antioxidative enzymes of transgenic fingermillet overexpressing a vacuolar H+-pyrophosphatase gene (SbVPPase) fromSorghum bicolor. J Plant Physiol 171:789–798
Anzalone AV et al (2019) Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576:149–157
Babitha KC, Vemanna R, Nataraja K et al (2015) Overexpression of EcbHLH57 transcription factor from Eleusine Coracana L. In Tobacco confers tolerance to Salt, oxidative and Drought stress. PLoS ONE 10:1–21
Babu BK, Agrawal P, Pandey D et al (2014a) Association mapping of agro-morphological characters among the global collection of finger millet genotypes using genomic SSR markers. Mol Biol Rep 41:5287–5297
Babu BK, Agrawal P, Pandey D et al (2014b) Comparative genomics and association mapping approaches for opaque2 modifier genes in finger millet accessions using genic, genomic and candidate gene-based simple sequence repeat markers. Mol Breed 34:1261–1279
Babu BK, Dinesh P, Agrawal P et al (2014c) Comparative genomics and association mapping approaches for blast resistant genes in finger millet using SSRs. PLoS ONE 9:e99182
Babu BK, Pandey D, Agrawal P et al (2014d) Molecular analysis of world collection of finger millet accessions for blast disease resistance using functional SSR markers. SABRAO J Breed Genet 46:202–216
Babu BK, Pandey D, Agrawal P et al (2014e) In-silico mining, type and frequency analysis of genic microsatellites of finger millet (Eleusine coracana (L.) Gaertn.): a comparative genomic analysis of NBS-LRR regions of finger millet with rice. Mol Biol Rep 41:3081–3090
Babu B, Kumar A, Yadav S, Sharma A, Singh S, Patel V, Choudhury S (2024) Comparative genomics approach to identify genes/QTLs linked to blast in finger millet (Eleusine coracana (L.) Gaertn). Front Plant Sci 15:102345
Baggett C, Koffi K, Sineshchekov O, et al (2021). Integration of SNP genotyping and molecular markers in crop breeding: A case study in millet improvement. Front Plant Sci 12:678945
Baker A, Ceasar S, Palmer A et al (2015) Replace, reuse, recycle: improving the sustainable use of phosphorus by plants. J Exp Bot 66:3523–3540
Bančič J, Damaris A, Odeny H et al (2024) Genomic and phenotypic characterization of finger millet indicates a complex diversification history. Plant Geno 17:e20392
Banerjee S, Sanjay K, Chethan S, Malleshi N (2012) Finger millet (Eleusine coracana) polyphenols: investigation of their antioxidant capacity and antimicrobial activity. Afr J Food Sci 6:362–374
Banerjee P, Maitra S, Banerjee P (2020) The role of small millets as functional food to combat malnutrition in developing countries. Indian J Nat Sci 10(60):20412–20417
Bhatt D, Saxena S, Jain S et al (2013) Cloning, expression and functional validation of drought inducible ascorbate peroxidase (Ec-apx1) from Eleusine coracana. Mol Bio Rep 40:1155–1165
Birch EE, Garfield S, Casteneda Y et al (2007) Visual activity and cognitive outcomes at 4 years of age in a double-blind, randomized trial of long-chain polyunsaturated fatty acid-supplemented infant formula. Early Hum Dev 83:279–284
Brhane A, Kassahun B, Assefa K, Fikre A, Tesfaye K, Spillane C, Jones H (2022) Genome-wide association study of grain yield and related traits in Ethiopian finger millet (Eleusine coracana (L.) Gaertn) landraces. BMC Genomics 23:5226
Ceasar SA (2018) Feeding world population amidst depleting phosphate reserves: the role of biotechnological interventions. Open Biotechnol J 12:51–55
Ceasar SA, Ignacimuthu S (2011) Agrobacterium-mediated transformation of finger millet (Eleusine coracana (L.) Gaertn.) Using shoot apex explants. Plant Cell Rep 30:1759–1770
Ceasar SA, Hodge A, Baker A, Baldwin S (2014) Phosphate concentration and arbuscular mycorrhizal colonisation influence the growth, yield and expression of twelve PHT1 family phosphate transporters in foxtail millet (Setaria italica). PLoS ONE 9:e108459
Ceasar SA, Rajan V, Prykhozhij S et al (2016) Insert, remove or replace: a highly advanced genome editing system using CRISPR/Cas9. Biochim. Biophys Acta 1863:2333–2344
Ceasar SA, Baker A, Ignacimuthu S (2017) Functional characterization of the PHT1 family transporters of foxtail millet with development of a novel Agrobacterium-mediated transformation procedure. Sci Rep 7:14064
Chandrasekara A, Shahidi F (2010) Content of insoluble bound phenolics in millets and their contribution to antioxidant capacity. J Agric Food Chem 58:6706–6714
Chinchole M, Pathak R, Singh U, Kumar A (2017) Molecular characterization of EcCIPK24 gene of finger millet (Eleusine coracana) for investigating its regulatory role in calcium transport. 3 Biotech 7:267
Chisi M, Peterson G (2019) Breeding and agronomy in Sorghum and millets. AACC Inter Pre-23-50.
Cletus J, Balasubramanian V, Vashisht D, Sakthivel N (2013) Transgenic expression of plant chitinases to enhance disease resistance. Biotechnol Lett 35:1719–1732
Cobb JN, Anderson JA, Boddupalli PM, et al. (2019) Enhancing the rate of genetic gain in public-sector plant breeding programs: lessons from the breeder’s equation. Theor Appl Genet 132:627–645
Cowling WA et al (2019) Modeling crop breeding for global food security during climate change. Food Energy Secur 8:e00157
Dagnachew L, Villiers S, Sewalem D et al (2014) Genetic diversity and eco-geographical distribution of Eleusine Species Collected from Ethiopia. Afr Crop Sci J 22:45–58
David RHA, Ramakrishnan M, Maharajan T et al (2021) Mining QTL and genes for root traits and biochemical parameters under vegetative drought in south Indian genotypes of finger millet (Eleusine coracana (L.) Gaertn) by association mapping and in silico comparative genomics. Biocatal Agric Biotechnol 32:101935
Devi PB, Vijayabharathi R, Sathyabama S et al (2014) Health benefits of Finger millet (Eleusine coracana L.) polyphenols and dietary fiber: a review. J food Sci tech 51:1021–1040
Dhanushkodi V, Hemavathy T, Shenbagavalli S et al (2023) A review on Nutritional properties and Health benefits of Finger millet. Inter J Pla Soi Sci 35(18):753–761
Dida MM, Oduori C, Manthi S et al (2021) 440 novel sources of resistance to blast disease in Finger millet. Crop Sci 61(1):250–262
Eapen S, George L (1990) Influence of phytohormones, carbohydrates, aminoacids, growth supplements and antibiotics on somatic embryogenesis and plant differentiation in finger millet. Plant Cell Tissue Organ Cult 22:87–93
Fernandez D, Vanderjagt D, Millson M et al (2003) Fatty acid, amino acid and trace mineral composition of Eleusine coracana (Pwana) seeds from northern Nigeria. Plant Foods Hum Nutr 58:1–10
Fuller DQ (2014) Finger Millet: Origins and Development. Encyclopedia of Global Archaeology, 2783–2785
Gebreyohannes A, Shimelis H, Mashilo J et al (2024) Finger millet (Eleusine coracana) improvement: challenges andprospects—A review. Plant Breed 143:350–374
Gimode D, Odeny D, De Villiers E et al (2016) Identification of SNP and SSR markers in Finger Millet using next generation sequencing technologies. PLoS ONE 11:1–23
Girgi M, O’Kennedy M, Morgenstern A et al (2002) Transgenic and herbicide resistant pearl millet (Pennisetum glaucum L.) R.Br. Via microprojectile bombardment of scutellar tissue. Mol Breed 10:243–252
Goel A, Gaur V, Arora S et al (2012) In silico analysis of expression data for identification of genes involved in spatial accumulation of calcium in developing seeds of rice. OMICS 16:402–413
Goel S, Singh AK, Yadav N, Tripathi A, Pandey D, Kumar A (2023). Transcriptomic insights into heat stress tolerance in finger millet: Differential gene expression analysis. J Plant Res 136:1021–1034
Gorjanc G et al (2018) Optimal cross selection for long-term genetic gain in two-part programs with rapid recurrent genomic selection. Theor Appl Genet 131:1953–1966
Gupta P, Raghuvanshi S, Tyagi A (2001) Assessment of the efficiency of various gene promoters via biolistics in leaf and regenerating seed callus of millets, Eleucine coracana and Echinochloa crusgalli. Plant Biotechnol 18:275–282
Gupta R, Verma K, Joshi D et al (2010) Assessment of genetic relatedness among three varieties of finger millet with variable seed coat color using RAPD and ISSR markers. Genet Eng Biotechnol J 2:1–9
Gupta N, Gupta A, Singh N, Kumar A (2011) Differential expression of PBF Dof transcription factor in different tissues of three finger millet genotypes differing in seed protein content and color. Plant Mol Biol Rep 29:69–76
Gupta A, Gaur V, Gupta S, Kumar A (2013) Nitrate signals determine the sensing of nitrogen through differential expression of genes involved in nitrogen uptake and assimilation in finger millet. Funct Integr Genomics 13:179–190
Gupta SM, Arora S, Mirza N et al (2017) Finger millet: a certain crop for an uncertain future and a solution to food insecurity and hidden hunger under stressful environments. Fron Plant sci 8:643
Gupta S, Pathak R, Gupta S et al (2018) Identification and molecular characterization of Dof Transcription Factor Gene Family preferentially expressed in developing spikes of Eleusine Coracana L. 3 Biotech 8:1–13
Harper AL et al (2012) Associative transcriptomics of traits in the polyploid crop species Brassica napus. Nat Biotechnol 30:798–802
Hass H et al (2017) Mathematical model of early reelin-induced src family kinase-mediated signaling. PLoS ONE 12:1–16
Hatakeyama M, Aluri S, Balachadran M et al (2018) Multiple hybrid de novo genome assembly of finger millet, an orphan allotetraploid crop. DNA Res 25:39–47
Hema R, Vemanna R, Sreeramulu S et al (2014) Stable expression of mtlD gene imparts multiple stress tolerance in finger millet. PLoS ONE 9:e99110
Heyser JW, Nabors MW (1982) Regeneration of proso millet from embryogenic calli derived from various plant parts. Crop Sci 22:1070–1074
Hilu KW, Johnson JL (1992) Ribosomal DNA variation in Finger millet and wild species of Eleusine (Poaceae). Theo App Gen 83:895–902
Hittalmani S, Mahesh H, Shirke M et al (2017) Genome and transcriptome sequence of finger millet (Eleusine coracana (L.) Gaertn.) Provides insights into drought tolerance and nutraceutical properties. BMC Genomics 18:465
Ignacimuthu S, Ceasar S (2012) Development of transgenic finger millet (Eleusine coracana (L.) Gaertn.) Resistant to leaf blast disease. J Biosci 37:135–147
Jacobson JL, Jacobson SW, Muckle C et al (2008) Beneficial effects of a polyunsaturated fatty acid on infant development: evidence from the Inuit of arctic Quebec. J Pediatr 152:356–364
Jayasudha BG, Sushma A, Prashantkumar S, Sashidhar V (2014) An efficient in-vitro agrobacterium mediated transformation protocol for raising salinity tolerant transgenic finger millet (Eleusine coracana (L.) Gaertn). Plant Arch 14:823–829
Jideani IA (2012) Digitaria exilis (acha/fonio), Digitaria iburua (iburu/fonio) and eluesine coracana (tamba/Finger millet)–Non-conventional cereal grains with potentials. Sci Res Ess 7(45):3834–3843
Jighly A et al (2019) Boosting genetic gain in allogamous crops via speed breeding and genomic selection. Front Plant Sci 10:1364
Jin X et al (2020) High-throughput estimation of crop traits: a review of ground and aerial phenotyping platforms. IEEE Trans Geosci Remote Sens 9:200–231
Kanwal P, Gupta S, Arora S, Kumar A (2014) Identification of genes involved in Carbon Metabolism from Eleusine Coracana (L.) Gaertn for understanding their light-mediated Entrainment and Regulation. Plant Cell Rep 33:1403–1411
Kayastha S, Sahoo JP, Mahapatra M, Panda N (2023) Understanding the molecular breeding and omics approaches for finger millet (Eleusine coracana L.) improvement towards a global sustainable nutritional security. Crop Des 3(1):100049
Kinghorn BP (2011) An algorithm for efficient constrained mate selection. Genet Sel Evol 43:4
Kirti D, Patel S, Jha S (2020) Optimization of regeneration and transformation system in finger millet (Eleucine coracana L). Int J Chem Stud 8(3):1678–1683
Kremling KA et al (2019) Transcriptome-wide association supplements genome-wide association in Zea mays. G3(Bethesda):3023–3033
Krishna TP, Maharajan T, Roch G et al (2020) Hybridization and hybrid detection through molecular markers in Finger Millet [Eleusine Coracana (L.) Gaertn]. J Crop Improv 34(3):335–355
Krishna SS, Reddy YAN, Kumar RLR (2021) Assessment of traits for grain yield under drought in finger millet. Plant Physiol Rep 26(1):84–94
Kumar A, Yadav S, Panwar P et al (2015) Identification of anchored simple sequence repeat markers Associated with Calcium Content in Finger Millet (Eleusine Coracana (L.) Gaertn). Proc Natl Acad Sci India Sect B: Biol Sci 85(1):311–317
Kumar A, Metwal M, Kaur S et al (2016) Nutraceutical value of Finger millet [Eleusine coracana (L.) Gaertn.], and their improvement using omics approaches. Fro Plant sci 7:934
Kumar A, Tomer V, Kaur A et al (2018) Millets: a solution to agrarian and nutritional challenges. Agri food Sect. 7(1):1–15
Latha AM, Rao K, Reddy V (2005) Production of transgenic plants resistant to leaf blast disease in finger millet (Eleusine coracana (L.) Gaertn). Plant Sci 169:657–667
Lemmon ZH et al (2014) The role of cis regulatory evolution in maize domestication. PLoS Genet 10:e1004745
Li J, Wang Y, Wang L et al (2021) Integration of transcriptomic and 513 proteomic analyses for Finger millet [Eleusine coracana (L.) Gaertn.] In response to drought stress. Plos 514 One 16(2):e0247181
Lim J, Park C, Kim M et al (2024) Advances in single-cell omics and multi-omics for high resolution molecular profilin. Exp Mol Med 56:515–526
Lin H et al (2017) Substantial contribution of genetic variation in the expression of transcription factors to phenotypic variation revealed by eRD-GWAS. Genome Biol 18:192
Liu JX, Howell S (2010) Endoplasmic reticulum protein Quality Control and its relationship to environmental stress responses in plants. Plant Cell 22:2930–2942
Liu F, Li D, Yu H, Kan Z, Liu Z, Sun D, Ding X (2016) PEDLA: predicting enhancers with a deep learning-based algorithmic framework. Sci Rep 6:28517.
Maharajan T, Ceasar SA, Krishna TPA (2022) Finger Millet (Eleusine coracana (L.) Gaertn): nutritional importance and nutrient transporters. CRC Crit Rev Plant Sci 41(1):1–31
Mane RS, Prasad BD, Sahni S, Quaiyum Z (2024) Biotechnological approaches toward Blast Disease Resistance in Finger Millet. Int J Plant Soil Sci 36(7):422–428
Manyasa EO, Tongoona P, Shanahan P, Mgonja M (2015) Genetic Diversity in East African Finger Millet (Eleusine Coracana (L.) Gaertn) Landraces Based on SSR Markers and Some Qualitative Traits. Plant Genetic Resources: Characterization and Utilization 13: 45–55
Mir RR et al (2012) Integrated genomics, physiology and breeding approaches for improving drought tolerance in crops. Theor Appl Genet 125:625–645
Mirza N, Taj G, Arora S, Kumar A (2014) Transcriptional expression analysis of genes involved in regulation of calcium translocation and storage in finger millet (Eleusine coracana L. Gartn). Gene 550:171–179
Mohammadi S, Prasanna B (2003) Analysis of genetic diversity in crop plants-salient statistical tools and considerations. Crop Sci 43:1235–1248
Mwangoe J, Kimurto PK, Ojwang PPO (2022) Identification of drought tolerant finger millet (Eleusine coracana) lines based on morpho-physiological characteristics and grain yield. Afr J Plant Sci 16(4):47–60
Nabors MW, Heyser J, Dykes T, De Mott K (1983) Long duration high frequency plant regeneration from cereal tissue cultures. Planta 157:385–391
Nagarjuna K, Parvathi M, Sajeevan R et al (2016) Full-length cloning and characterization of abiotic stress responsive CIPK31-like gene from finger millet, a drought-tolerant crop. Curr Sci 111:890–894
Nethra N, Gowda R, Gowda P (2009) Influence of culture medium on callus proliferation and morphogenesis in finger millet. Univ Bern 167–178
Ngom B, Sarr I, Kimatu J et al (2017) Genome-wide analysis of cytosine DNA methylation revealed salicylic acid promotes defense pathways over seedling development in pearl millet. Plant Signal Behav 12
Obidiegwu ON, Parzies H, Obidiegwu J (2014) Development and genotyping potentials of EST-SSRs in Finger Millet (E. Coracana (L.) Gaertn). Int J Genet Genomics 2:42–46
Odeny DA, Niazi A, Tesfaye K et al (2020) Genomic designing for climate smart Finger millet. Genomic Des of climate-smart cereal crops 287–307
Pandey G, Yadav CB, Sahu PP et al (2017) Salinity induced differential methylation patterns in contrasting cultivars of foxtail millet (Setaria italica L). Plant Cell Rep 36:759–772
Panwar P, Nath M, Yadav V, Kumar A (2010) Comparative evaluation of genetic diversity using RAPD, SSR and cytochrome P450 gene based markers with respect to calcium content in Finger millet (Eleusine coracana L. Gaertn). J Gen 89:121–133
Panwar P, Jha A, Pandey P et al (2011) Functional markers based molecular characterization and cloning of resistance gene analogs encoding NBS-LRR disease resistance proteins in finger millet (Eleusine coracana). Mol Biol Rep 38:3427–3436
Parvathi MS, Nataraja K (2017) Discovery of stress responsive TATAbox binding protein associated Factor6 (TAF6) from finger millet (Eleusine coracana (L.) Gaertn). J Plant Biol 60:335–342
Parvathi M, Nataraja K, Yashoda B et al (2013) Expression analysis of stress responsive pathway genes linked to drought hardiness in an adapted crop, finger millet (Eleusine coracana). J Plant Biochem Biotechnol 22:193–201
Parvathi MS, Nataraja K, Reddy Y et al (2019) Transcriptome analysis of Finger Millet (Eleusine Coracana (L.) Gaertn.) Reveals Unique Drought responsive genes. J Genet 98:1–12
Patil J, Kale A (2013) Study of genetic diversity in finger millet (Elesuine Coracana L.) genotypes using RAPD markers. Int J Integr Sci Innov Technol 2:31–36
Pazhamala LT et al (2021) Systems biology for crop improvement: prospects and challenges. Plant Genome 14:e2009
Peng R, Zhang B (2021) Foxtail Millet: a New Model for C4 plants. Trends Plant Sci 26(3):199–201
Pudake RN, Mehta C, Mohanta T et al (2017) Expression of four phosphate transporter genes from Finger millet (Eleusine coracana L.) in response to mycorrhizal colonization and pi stress. 3 Biotech 7(1):1–13
Puranik S, Kam J, Sahu P et al (2017) Harnessing finger millet to combat calcium deficiency in humans: challenges and prospects. Front Plant Sci 8:1311
Puranik S, Sahu P, Beynon S et al (2020) Genome-wide association mapping and comparative genomics identifies genomic regions governing grain nutritional traits in Finger millet (Eleusine coracana L. Gaertn). Plants People Planet 2(6):649–662
Rahman H, Jagadeeshselvam N, Valarmathi R et al (2014) Transcriptome analysis of salinity responsiveness in contrasting genotypes of finger millet (Eleusine coracana L.) through RNA-sequencing. Plant Mol Biol 85:485–503
Rahman H, Ramanathan V, Nallathambi J et al (2016) Over-expression of a NAC 67 transcription factor from finger millet (Eleusine coracana L.) confers tolerance against salinity and drought stress in rice. BMC Biotechnol 16:35
Rajendram R, Nirmalakumari A, Senthil N, Muthiah AR (2016) Genetic diversity analysis in finger millet (Eleusine coracana L. Gaertn) using SSR markers. J Food Legumes 29:123–128.
Rakkammal K, Maharajan T, Shriram RN et al (2023) Physiological, biochemical and molecular responses of finger millet (Eleusine coracana) genotypes exposed to short-term drought stress induced by PEG-6000. South Afr J Bot 155:45–59
Ramakrishna C, Singh S, Raghavendrarao S et al (2018) The membrane tethered transcription factor EcbZIP17 from Finger millet promotes plant growth and enhances tolerance to abiotic stresses. Sci Rep 8(1):2148
Ramakrishnan M, Ceasar S, Duraipandiyan V et al (2016) Using molecular markers to assess the genetic diversity and population structure of finger millet (Eleusine coracana (L.) Gaertn.) From various geographical regions. Genet Resour Crop Evol 63:361–376
Ramashia SE, Gwata E, Meddows-taylor S et al (2018) Some Physical and Functional properties of Finger Millet (Eleusine Coracana) obtained in sub-saharan Africa. Food Rese Inter 104:110–118
Ramegowda V, Senthil-Kumar M, Nataraja K et al (2012) Expression of a finger millet transcription factor. EcNAC1, in tobacco confers abiotic stress-tolerance. PLoS ONE 7:e40397
Ramegowda V, Gill U, Sivalingam P et al (2017) GBF3 transcription factor imparts drought tolerance in Arabidopsis thaliana. Sci Rep 7:9148
Rangan TS (1973) Morphogenic investigations on tissue cultures of Panicum miliaceum. Z Pflanzenpyhsiol Bd 72:456–459
Rani V, Joshi D, Joshi P et al (2023) Millet models for harnessing nuclear factor-Y transcription factors to engineer stress tolerance in plants: current knowledge and emerging paradigms. Planta 258(2):29
Rathore T, Singh R, Kamble D, Upadhyay A, Thangalakshmi S (2019) Review on Finger millet: Processing and value addition. Phar Inno J 8(4):283–291
Saha D, Gowda M, Arya L et al (2016) Genetic and genomic resources of small millets. Crit Rev Plant Sci 35(1):56–79
Saha D, Rana R, Arya L et al (2019) Genetic polymorphisms among and between Blast Disease resistant and susceptible Finger Millet (Eleusine Coracana (L) Gaertn). Plant Genetic Resources: Charact Utilization 15:355–365
Samuel AL (1959) Some studies in machine learning using the game of checkers. IBM J Res Dev 3:210–229
Satish L, Ceasar S, Ramesh M (2017) Improved agrobacteriummediated transformation and direct plant regeneration in four cultivars of finger millet (Eleusine coracana (L.) Gaertn). Plant Cell Tissue Organ Cult 131:547–565
Sehgal D, Rajaram V, Armstead I et al (2012) Integration of gene-based markers in a pearl millet genetic map for identifcation of candidate genes underlying drought tolerance quantitative trait loci. BMC Plant Biol 12:1–3
Sen S, Dutta S (2012) Cloning, expression and characterization of biotic stress inducible ragi bifunctional inhibitor (RBI) gene from Eleusine coracana Gaertn. Plant Biochem Biotechnol 21:66–76
Sen S, Kundu S, Dutta S (2016) Proteomic analysis of JAZ interacting proteins under methyl jasmonate treatment in finger millet. Plant Physiol Biochem 108:79–89
Shanmugapackiam S, Sankar PM, Irulandi S, Raguchander T (2019) Identification of secondary metabolites from finger millet parts infected with Magnaporthe grisea by GC-MS analysis. Chem Sci Rev Lett 6(21):88–93
Sharma M, Kothari-Chajer A, Jagga-Chugh S, Kothari S (2011) Factors influencing Agrobacterium tumefasciens-mediated genetic transformation of Eleucine coracana (L.) Gaertn, Plant Cell. Tiss Organ Cult 105:93–104
Singh UM, Chandra M, Shankhdhar S, Kumar A (2014) Transcriptome wide identification and validation of calcium sensor gene family in the developing spikes of finger millet genotypes for elucidating its role in grain calcium accumulation. PLoS ONE 9:e103963
Singh R, Singh V, Raghavendrarao S et al (2015) Expression of finger millet EcDehydrin7 in transgenic tobacco confers tolerance to drought stress. Appl Biochem Biotechnol 177:207–216
Singh S, Mahalle M, Mukhtar M, Jiwani G, Sevanthi A, Solanke A (2022) Advances in Omics 639 for enhancing abiotic stress tolerance in Finger millets. In omics of climate resilient small millets Singapore. Springer Nat Singap 257:235–640
Singh P, Singh R, Bhadauria V, Singh H (2023) The functionality and extraction of protein from Sorghum, Finger millet, and Kodo Millet: a review. International Journal of Food Science & Technology
Sood S, Kumar A, Babu B et al (2016) Gene discovery and advances in finger millet (Eleusine coracana (L.) Gaertn.) Genomics-an important nutri-cereal of future. Front Plant Sci 7:1634
Sood S, Joshi DC, Rajashekara H et al (2023) Deciphering the genomic regions governing major agronomic traits and blast resistance using genome wide association mapping in Finger millet. Gene 854:147115
Srivastav S, Kothari SL (2002) Embryogenic callus induction and efficient plant regeneration in pearl millet. Cer Res Comm 30:69–74
Strobl S, Muhlhahn P, Bernstein R et al (1995) Determination of the three-dimensional structure of the bifunctional alpha-amylase/trypsin inhibitor from ragi seeds by NMR spectroscopy. Biochemistry 34:8281–8293
Sudan J, Negi B, Arora S (2015) Oxidative stress induced expression of monodehydroascorbate reductase gene in Eleusine coracana. Physio Mol Bio Plants 21:551–558
Talwar HS, Kumar S, Madhusudhana R et al (2020) Variations in drought tolerance components and their association with yield components in Finger millet (Eleusine coracana). Funct Plant Biol 47(7):659–674
van Dijk AD et al (2021) Machine learning in plant science and plant breeding. iScience 24:101890
Varshney RK et al (2020) 5Gs for crop genetic improvement. Curr Opin Plant Biol 56:190–196
Varshney RK et al (2021) Designing future crops: genomicsassisted breeding comes of age. Trends Plant Sci 26:631–664
Wambi W, Tumwesigye W, Otienno G, Mulumba j (2020) Genetic and genomic resources for finger millet improvement: opportunities for advancing climate-smart agriculture. J Crop Improv 35(2):204–233
Wang T, Lu Q, Song H et al (2021) DNA methylation and RNA-Sequencing analysis show epigenetic function during Grain Filling in Foxtail Millet (Setaria italica L). Front Plant Sci 12
Watson A et al (2018) Speed breeding is a powerful tool to accelerate crop research and breeding. Nat Plants 4:23–29
Weckwerth W et al (2020) PANOMICS meets germplasm. Plant Biotechnol J 18:1507–1525
Woolliams JA, Berg P, Dagnachew BS, Meuwissen THE (2015) Genetic contributions and their optimization. J Anim Breed Genet 132:89–99.
Yadav S, Kumar A, Thakur A (2017) Marker-Trait Association Analysis for Grain Calcium Content in Global Collection of Finger Millet genotypes. J Food Process Technol 8:75
Yan G et al (2017) Accelerated generation of selfed pure line plants for gene identification and crop breeding. Front Plant Sci 8:1786
Yan F et al (2018) High-efficient A-T to G-C base editing by Cas9n-guided tRNA adenosine deaminase in rice. Mol Plant 11:631–634
Yang Y, Wang J, Li G et al (2022a) Shift in subsistence crop dominance from broomcorn millet to foxtail millet around 5500 BP in the western Loess Plateau. Fron Pla Sci 13:939340
Yemets AI, Radchuk V, Bayer O et al (2008) Development of transformation vectors based upon a modified plant α-tubulin gene as the selectable marker. Cell Biol Int 32:566–570
Zhang H, Hall N, Goertzen L et al (2019) Transcriptome Analysis Reveals Unique Relationships among Eleusine species and Heritage of Eleusine Coracana. G3/Genes/Genomes/Genetics 9:2029–2036
Acknowledgements
Authors acknowledge the funding received from Dr. Rajendra Prasad Central Agricultural University, Pusa, Samastipur-848125, Bihar, India.
Author information
Authors and Affiliations
Contributions
RSM and BDP contributed to the literature collection and preparing the draft of the manuscript. RSM, BDP, SS, ZQ, and VKS have read, edited, and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mane, R.S., Prasad, B.D., Sahni, S. et al. Biotechnological studies towards improvement of finger millet using multi-omics approaches. Funct Integr Genomics 24, 148 (2024). https://doi.org/10.1007/s10142-024-01438-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-024-01438-4