Abstract
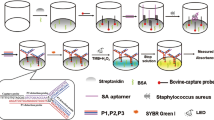
We have developed a specific method for the visual detection of Staphylococcus aureus based on aptamer recognition coupled to tyramine signal amplification technology. A biotinylated aptamer specific for S. aureus was immobilized on the surface of the wells of a microplate via biotin-avidin binding. Then, the target bacteria (S. aureus), the biotinylated-aptamer-streptavidin-HRP conjugates, biotinylated tyramine, hydrogen peroxide and streptavidin-HRP were successively placed in the wells of the microplate. After adding TMB reagent and stop solution, the intensity of the yellow reaction product can be visually inspected or measured with a plate reader. Under optimized conditions, there is a linear relationship between absorbance at 450 nm and the concentration of S. aureus in the 10 to 107 cfu mL−1 concentration range (with an R2 of 0.9976). The limit of detection is 8 cfu mL−1.

A visual detection method for Staphylococcus aureus was based on aptamer recognition coupled to tyramine signal amplification. The linear range was from 10 to 107 cfu mL-1 and the limit of detection was 8 cfu mL-1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Staphylococcus aureus is a Gram-positive bacterium; it is an important human and veterinary pathogen that causes a wide variety of infections, ranging from benign skin infections to life threatening diseases [1] such as pneumonia, endocarditis, osteomyelitis, arthritis, and sepsis [2]. Furthermore, S. aureus is the most frequent cause of bacteremia and hospital-acquired infection. In many countries, S. aureus is the second or third most common pathogen that causes outbreaks of food poisoning [3]. Therefore, it is of great importance to develop a simple, specific, and sensitive detection method for S. aureus.
Several methods have been developed to detect S. aureus, including the traditional microbiological culture method, enzyme-linked immunosorbent assay (ELISA) [4, 5], the polymerase chain reaction (PCR) technique [6, 7], Loop mediated AMPlification (LAMP) [8]. These methods have been successfully applied in various fields. However, some of the methods have deficiencies in stability, price, accuracy, detection limits and time, and the products of most of these methods cannot be visually observed. The development of a novel detection method with the advantages of simplicity, speed, sensitivity, and ease of observation has therefore attracted increased attention and remains a challenge [9, 10].
Aptamers are DNA or RNA molecules that are commonly obtained in vitro using a combinatorial chemistry technique known as systematic evolution of ligands by exponential enrichment (SELEX) [11]. Because they bind to their target molecules with high affinity and specificity, aptamers have a wide range of applications for the detection of certain pathogenic bacteria [12, 13], toxins [14–16], heavy metal ions [17, 18] and illegal food additives [19–21]. In addition, aptamers have previously been applied in the diagnosis and therapy of disease [22, 23].
Tyramide signal amplification (TSA) technology uses horseradish peroxidase (HRP) to catalyze the deposition of biotin-labeled tyramide molecules for localized signal enhancement, thus using the sensitivity of an enzyme-based assay [24]. This technology has been used in many fields, such as in situ hybridization techniques [25, 26], analysis and detection [27], and disease diagnosis [28, 29].
In this paper, aptamer recognition coupled with tyramine signal amplification was used for the direct detection of S. aureus by visual inspection. Aptamers with high affinity and specificity can specifically recognize and bind to S. aureus, which ensures the specificity of the developed method. Moreover, the TSA technology was applied in this work to effectively amplify the signals. In addition, the linear correlation, signal intensity, linear range and limit of detection of the developed method were compared with the method without the TSA technology. Compared to the existing methods, such as the microbiological culture method, ELISA and PCR, the developed method offers the merits of simplicity, rapidity, sensitivity, stability and low cost. This work is expected to launch a new field of TSA coupled with aptamer applications.
Experimental
Reagents and apparatus
Anhydrous sodium carbonate (Na2CO3), sodium bicarbonate (NaHCO3), 30 % hydrogen peroxide (30 % H2O2), sodium chloride (NaCl), potassium chloride (KCl), disodium hydrogen phosphate (Na2HPO4 · 12H2O), dipotassium hydrogen phosphate (K2HPO4), agar powder, tryptone, yeast extract, N, N-Dimethylformamide (DMF), ethanol, triethylamine, Tween-20, and bovine serum albumin (BSA) were purchased from the Sinopharm Chemical Reagent Co., Ltd. (Shanghai, China) (http://www.reagent.com.cn). The EL-TMB Chromogenic Reagent kit, Streptavidin-HRP, N-hydroxysuccinimide biotin (NHS Bioth) and Streptavidin from Streptomyces avidinii were purchased from the Shanghai Sangon Biological Science & Technology Company (Shanghai, China) (http://www.sangon.com). Tyramine hydrochloride was purchased from the Shanghai yuanye Bio-Technology Co., Ltd. (Shanghai, China) (http://www.yuanyebio.com.cn). Staphylococcus aureus ATCC 29213 was obtained from the American Type Culture Collection (ATCC) (http://www.atcc.org). The S. aureus aptamer [30] 5′-biotin-C6-GCA ATG GTA CGG TAC TTC CTC GGC ACG TTC TCA GTA GCG CTC GCT GGT CAT CCC ACA GCT ACG TCA AAA GTG CAC GCT ACT TTG CTA A-3′ was synthesized by the Shanghai Sangon Biological Science & Technology Company (Shanghai, China) (http://www.sangon.com). The signals were scanned using a Molecular Devices SpectraMax M5 plate reader (M5, Molecular Devices, U.S.A.). The ultrapure water used in the experiments was prepared with a Millipore Direct-Q® 3 system (Merck Millipore, MA, U.S.A.) and had a resistivity of 18.2 MΩ cm.
Preparation of biotin–tyramine
The Biotin-tyramine conjugate was prepared according to Hopman’s method [31]: 10 mg Bio-NHS was dissolved in 1 mL DMF to prepare solution A; 2.8 mg tyramine hydrochloride was dissolved in 0.28 mL DMF, and 2.8 μL triethylamine was added into solution A to prepare solution B; then, solution A and solution B were mixed sufficiently to react in the dark at room temperature for 2 h. Finally, 8.72 mL ethanol was added to prepare 10 mL biotin-tyramine stock solution, which was stored at 4 °C in the dark.
Assembly of aptamer onto the microplate wells
First, the streptavidin (1 mg mL−1) was diluted 1:100 with carbonate buffer (pH 9.6), and 200 μL diluted streptavidin was added into each zone of the microplates and incubated at 4 °C overnight. After washing with wash buffer (0.01 mol L−1 PBS, 0.05 % Tween-20) 3 times (1 min per wash), each zone of the microplates was blocked with blocking buffer (BSA in 0.01 mol L−1 PBS) for 1 h at room temperature to prevent the appearance of false positive signals. Subsequently, the microtiterplates were washed 3 times with wash buffer, and 10 μL biotinylated aptamer was added and incubated at 37 °C. After 30 min, the microplates were washed 3 times with wash buffer.
Analytical procedure
A 100 μL aliquot of the sample solution containing S. aureus was added to each zone of the microplates. The biotinylated aptamer and streptavidin-HRP were mixed and then incubated at 37 °C for 30 min at the same time. The microplates were washed 3 times with wash buffer, and 10 μL aptamer and streptavidin-HRP complex was added to the microplates and incubated at 37 °C for 30 min. After washing 5 times with wash buffer, each zone of the microplates was loaded with 10 μL biotin-tyramine. A small volume of hydrogen peroxide (0.5 %, v/v) was also added to the diluted solution to act as the oxidizing agent of tyramine [32]. The microtiterplates were incubated at 37 °C for 30 min and then washed 3 times with wash buffer and air dried. Subsequently, 10 μL streptavidin-HRP was added into each zone of the microplate. The plate was incubated at 37 °C for 30 min and then washed 5 times with wash buffer and air dried. Finally, 100 μL TMB reagent was added to each well for 20 min at room temperature. After adding 50 μL stop solution, the absorbance at 450 nm was recorded using a Molecular Devices SpectraMax M5 plate reader. This procedure is illustrated in Fig. 1 and the yellow color is formed on reaction in the microplate.
Treatment of the water samples
The feasibility of applying the developed method to measure S. aureus levels in samples was validated using water samples obtained from Tai Lake, and the treatment of the water samples was as follows: the water samples were left to stand for at least 30 min to precipitate macroaggregates and seston. Then, the water samples were centrifuged for at least 35 min at 12000 rpm (5430R, Eppendorf, Germany) to remove the tiny seston, and the supernatant was filtered with a 0.45 μm filtration membrane. Finally, all samples were sterilized at 121 °C for at least 20 min to ensure that they did not contain any microorganisms that would influence the experimental results.
Results and discussion
Effect of the concentration of BSA
To prevent the appearance of a false positive because of non-specific adsorption, the microplates were coated with BSA in 0.01 mol L−1 PBS. In this experiment, we selected different concentrations (5, 10, 15, 20, 25 mg mL−1) of BSA in PBS as the blocking solution to block the bottom of the microplates that were not coated with streptavidin. According to the experimental results (Fig. 2), the non-specific adsorption was reduced and the colors of the blank sample (sterile LB medium) were increasingly lighter with increasing BSA concentration; when the concentration of BSA was 20 mg mL−1, the bottom of the microplates that were not coated with streptavidin were largely blocked with BSA, and the effect of non-specific adsorption on the experiment was minimized. However, for the concentrations of 20 and 25 mg mL−1, there was a slight difference between the intensity of the signals that correlated with the concentration of BSA. Consequently, we selected 20 mg mL−1 as the optimized concentration of BSA.
Optimization of the concentration of aptamer
The microplate was covered with biotinylated aptamer through specific binding between biotin and streptavidin. The function of the aptamer was to capture S. aureus and attach it to the bottom of the microplates in this experiment; therefore, the concentration of aptamer was directly correlated with the amount of captured S. aureus and the intensity of the signals. To evaluate the impact of the aptamer concentrations on the experimental results, we selected different concentrations of aptamer that ranged from 10−4 mol L−1 to 10−12 mol L−1. As shown in Fig. 3, the intensity of the signals became increasingly stronger as the concentration of aptamer increased. But, for the concentrations of 10−4, 10−5 and 10−6 mol L−1, there was a slight difference between the intensity of the signals from the aptamer. Thus, we selected 10−6 mol L−1 as the optimized concentration of aptamer in this experiment.
Optimization of the dilution of Streptavidin-HRP
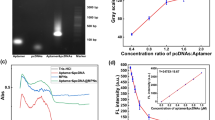
In this work, streptavidin-HRP was added to the microplate at two different steps. The role of the first addition of streptavidin-HRP was mainly to catalyze the tyramine deposition, and the signal intensity was minimally affected by the concentration of streptavidin-HRP [32], which was consistent with our experimental results; hence, this factor will not be discussed here, and 1:4500 was selected as the ideal dilution factor of the first addition of streptavidin-HRP. The second addition of streptavidin-HRP had a significant influence on the signal intensity. The streptavidin-HRP (1 mg mL−1) was diluted to 1:500, 1:1500, 1:3000, 1:4500, and 1:6000, and it was used for the detection of the sample (the concentration of S. aureus in the sample was 106 cfu mL−1) and the blank (sterile LB medium). Figure 4 indicates that the signal intensity of the samples decreased substantially as the streptavidin-HRP dilution ratio increased from 1:500 to 1:6000. For the blank, the signals decreased as the streptavidin-HRP dilution ratio increased from 1:500 to 1:1500; however, there was a slight difference between the intensity of the signals as the dilution ratio increased from 1:3000 to 1:4500 and to 1:6000. By considering the impact of the streptavidin-HRP dilution ratio on the signal intensity of the sample wells and blank wells, we selected 1:1500 as the ideal dilution ratio of the streptavidin-HRP.
Optimization of the dilution of biotin-tyramine
Tyramine is a phenolic compound, and HRP can catalyze biotin-tyramine to produce highly reactive phenolic radicals (tyramide radicals) in the presence of H2O2, which react covalently with electron-rich moieties of protein molecules that are present in the vicinity of the HRP label. Figure 5 indicates that the intensity of the signals gradually increases as the biotin–tyramine dilution ratio increases from 1:50 to 1:100, and the intensity declines sharply as the biotin–tyramine dilution ratio increases from 1:100 to 1:800. This trend may occur because HRP can catalyze the dimerization of tyramine when it is present at a high concentration, which is likely caused by the generation of free radicals [33]. However, when the tyramine is applied in lower concentrations, the probability of dimerization is reduced because the binding of highly reactive intermediates to electron-rich moieties of proteins is favored [34]. The number of highly reactive intermediates that are generated by HRP catalysis was low when the concentration of tyramine was too low. Therefore, the 1:100 dilution was suitable for the experiment.
Specificity
To evaluate the specificity of this method, we measured six other samples that contained Vibrio parahemolyticus, Salmonella Typhimurium, Streptococcus, E. sakazakii, E. coli and Listeria monocytogenes and one blank sample. The analysis of all samples was performed under the same conditions, and the concentrations of all bacteria were 106 cfu mL-1. Figure 6 clearly shows that the intensity of the signals of S. aureus were much stronger than the other six bacterial samples and the blank samples, mainly because of the binding of the aptamers to their target molecules with high affinity and specificity. Thus, the signal intensities of Vibrio parahemolyticus, Streptococcus, Salmonella typhimurium, E. sakazakii, E. coli, Listeria monocytogenes and the blank were weak. This result (Fig. 6) indicated the good specificity of the developed method for the detection of S. aureus.
Analytical performance
A series of concentrations of S. aureus (from 10 cfu mL−1 to 107 cfu mL−1) were investigated under optimal conditions. The results show that the intensity of the signals is linear with and the concentration of S. aureus ranged from 10 cfu mL−1 to 107 cfu mL−1 (y = 0.1750x−0.0061, R2 = 0.9976) (Fig. 7). Statistical analysis revealed that the limit of detection of S. aureus was 8 cfu mL−1.
A series of concentrations of S. aureus (from 10 cfu mL−1 to 107 cfu mL−1) were analyzed by the method without the tyramide signal amplification (TSA) technology. As the results shown, the linear correlation between the intensity of the signals and the concentration of S. aureus was 0.9921 and the linear fit that was obtained was y = 0.1191x−0.0575. The linear range (from 50 cfu mL−1 to 5 × 104 cfu mL−1) was very narrow, and the limit of detection was 40 cfu mL−1. By considering the results from the comparison between these two methods, we determined that the linear correlations between the intensity of the signals and the concentration of S. aureus and between the linear range and the limit of detection were improved significantly by the TSA technology.
The developed method were further compared with the presented some previous methods about detection of S. aureus (Table 1). As shown in Table 1, the developed method is more sensitive than others. However, the analysis time of the developed method should be improved and it would be taken into account in the future studies.
Analytical application
A series of concentrations of S. aureus were added to the treated water samples, and 100 μL dilutions were added to the 900 μL water samples. Afterward, 100 μL samples were detected by the developed method, and additional 100 μL samples were tested by the classical counting methods.
The results obtained by the visual detection method were similar to those obtained from the plate counting method. There was no significant difference between the counting method and the developed method (R2 = 0.9979, P < 0.0001), and the linear fit that was obtained was y = 0.9994x + 1.3014. It confirms that the visual detection method using aptamers coupled with tyramine signal amplification can be used for the detection of samples.
Conclusions
In conclusion, we successfully developed a specific, stable and visual detection method for S. aureus that is based on the recognition of aptamers coupled with tyramine signal amplification (TSA). With the help of the TSA technology, the detection sensitivity of the developed method reached 8 cfu mL−1, and the linear range was wide (from 10 cfu mL−1 to 107 cfu mL−1), as determined by visual inspection. Furthermore, aptamers are specific, stable, and inexpensive; therefore, the developed method is stable, inexpensive and specific for the detection for S. aureus. The developed method has the potential for wide use in the visual detection of other foodborne pathogenic bacteria in food samples.
References
Francois O, Scherl A, Hochstrasserb D, Schrenzel J (2010) Proteomic approaches to study Staphylococcus aureus pathogenesis. J Proteomics 73:701–708
Chakraborty SP, Mahapatra SK, Sahu SK, Chattopadhyay S, Pramanik P, Roy S (2011) Nitric oxide mediated Staphylococcus aureus pathogenesis and protective role of nanoconjugated vancomycin. Asian Pac J Trop Biomed 1:102–109
Atanassova V, Meindl A, Ring C (2011) Prevalence of Staphylococcus aureus and staphylococcal enterotoxins in raw pork and uncooked smoked ham-a comparison of classical culturing detection and RFLP-PCR. Int J Food Microbial 68:105–113
Belkessam Y, Benali M, Moulessehoul S, Harrach D (2010) Polyclonal antibodies production against Staphylococcus aureus protein A: ELISA technique optimization for milk quality control. Afr J Biotechnol 9:764–769
Yazdankhah SP, Solverod L, Simonsen S (1999) Development and evaluation of an immunomagnetic separation-ELISA for the detection of Staphylococcus aureus thermostable nuclease in composite milk. Vet Microbiol 67:113–125
Pinto B, Chenoll E, Aznar R (2005) Identification and typing of food-borne Staphylococcus aureus by PCR-based techniques. Syst Appl Microbiol 28:340–352
Sabet NS, Subramaniam G, Navaratnam P, Sekaran SD (2007) Detection of methicillin- and aminoglycoside-resistant genes and simultaneous identification of S.aureus using triplex real-time PCR Taqman assay. J Microbiol Methods 68:157–162
Sowmya N, Thakur MS, Manonmani HK (2012) Rapid and simple DNA extraction method for the detection of enterotoxigenic Staphylococcus aureus directly from food samples: comparison of PCR and LAMP methods. J Appl Microbiol 133:106–113
Xu S (2012) Electromechanical biosensors for pathogen detection. Microchim Acta 178:245–260
Roda A, Mirasoli M, Roda B, Bonvicini F, Colliva C, Reschiglian P (2012) Recent developments in rapid multiplexed bioanalytical methods for foodborne pathogenic bacteria detection. Microchim Acta 178:7–28
Hamula CLA, Guthrie JW, Zhang HQ, Li XF, Le CX, Li XF (2006) Selection and analytical applications of aptamers. Trac-trend Anal Chem 25:681–691
Duan N, Sj W, Zhu CQ, Ma XY, Wang ZP, Yu Y, Jiang Y (2012) Dual-color upconversion fluorescence and aptamer-functionalized magnetic nanoparticles-based bioassay for the simultaneous detection of Salmonella Typhimurium and Staphylococcus aureus. Anal Chim Acta 723:1–6
Liu GQ, Yu XF, Xue F, Chen W, Ye YK, Yang XJ, Lian YQ, Yan Y, Zong K (2012) Screening and preliminary application of a DNA aptamer for rapid detection of Salmonella O8. Microchim Acta 178:237–244
Wu SJ, Duan N, Ma XY, Xia Y, Wang HX, Wang ZP (2013) A highly sensitive fluorescence resonance energy transfer aptasensor for staphylococcal enterotoxin B detection based on exonuclease-catalyzed target recycling strategy. Anal Chim Acta 782:59–66
Girolamo AD, McKeague M, Miller JD, DeRosa MC, Visconti A (2011) Determination of ochratoxin A in wheat after clean-up through a DNA aptamer-based solid phase extraction column. Food Chem 127:1378–1384
Wu SJ, Duan N, Wang ZP, Wang HX (2011) Aptamer-functionalized magnetic nanoparticle-based bioassay for the detection of ochratoxin a using upconversion nanoparticles as labels. Analyst 136:2306–2314
Chung CH, Kim JH, Jung J, Chung BH (2013) Nuclease-resistant DNA aptamer on gold nanoparticles for the simulantaneous detection of Pb2+ and Hg2+ inhuman serum. Biosens Bioelectron 41:827–832
Liang AH, Zhang J, Wen GQ, Liu QY, Li TS, Jiang ZL (2010) Aptamer-modified AuRu nanoparticle catalytic resonance scattering spectral assay for the determination of trace Pb2+. Chem J Chinese U 32:2366–2371
Liang AH, Zhou LP, Qin HM, Zhang Y, Ouyang HX, Jiang ZL (2011) A highly sensitive aptamer-nanogold catalytic resonance scattering spectral assay for melamine. J Fluoresc 21:1907–1921
Mei Z, Chu H, Chen W, Xue F, Liu J, Xu H, Zhang R, Zheng L (2013) Ultrasensitive one-step rapid visual detection of bisphenol A in water samples by label-free aptasensor. Biosens Bioelectron 39:26–30
Xue F, Wu JJ, Chu YQ, Mei ZL, Ye YK, Liu J, Zhang R, Peng CF, Zheng L, Chen W (2013) Electrochemical aptasensor for the determination of bisphenol A in drinking water. Microchim Acta 180:109–115
Wang P, Yang Y, Hong H, Zhang Y, Cai W, Fang D (2011) Aptamers as therapeutics in cardiovascular diseases. Curr Med Chem 18:4169–4174
Keefe AD, Pai S, Ellington A (2010) Aptamers as therapeutics. Nat Rev Drug Discov 9:537–550
Zhang Z, Kitching P (2000) A sensitive method for the detection of foot and mouth disease virus by in situ hybridisation using biotin-labelled oligodeoxynucleotides and tyramide signal amplification. J Virol Methods 88:187–192
Not F, Simon N, Biegala IC, Vaulot D (2002) Application of fluorescent in situ hybridization coupled with tyramide signal amplification (FISH-TSA) to assess eukaryotic picoplankton composition. Aquat Microb Ecol 28:157–166
Kubota K, Ohashi A, Imachi H, Harada H (2006) Visualization of mcr mRNA in a methanogen by fluorescence in situ hybridization with an oligonucleotide probeand two-pass tyramide signal amplification (two-pass TSA-FISH). J Microbiol Methods 66:521–528
Saha D, Acharya D, Roy D, Dhar TK (2007) Filtration-based tyramide amplification technique-a new simple approach for rapid detection of aflatoxin B1. Anal Chim Acta 387:1121–1130
Koda T, Aosasa MH, Nakaba H, Matsuda H (2003) Application of tyramide signal amplification for detection of N-glycolylneuraminic acid in human hepatocellular carcinoma. Int J Oncol 8:317–321
Strappe PM, Wang TH, McKenzie CA, Lowrie S, Simmonds P, Bell JE (1997) Enhancement of immunohistochemical detection of HIV-1 p24 antigen in brain by tyramide signal amplification. J Virol Methods 67:103–112
Cao XX, Li SH, Chen LC, Ding HM, Xu H, Huang YP, Li J, Liu N, Cao W, Zhu Y, Shen B, Shao N (2009) Combining use of a panel of ssDNA aptamers in the detection of Staphylococcus aureus. Nucleic Acids Res 37:4621–4628
Hopman AH, Ramaekers FC, Speel EJ (1998) Rapid synthesis of biotin-, digoxigenin-, trinitrophenyl-, and fluorochrome-labeled tyramides and their application for in situ hybridization using CARD amplification. J Histochem Cytochem 46:771–777
Qi HJ, Chen SH, Zhang ML, Shi H, Wang SQ (2010) DNA microarrays for visual detection of human pathogenic microorganisms based on tyramine signal amplification coupled with gold label silver stain. Anal Bioanal Chem 398:2745–2750
Zaitsu K, Ohkura Y (1980) New fluorogenic substrates for horseradish peroxidase: rapid and sensitive assays for hydrogen peroxide and the peroxidase. Anal Biochem 109:109–113
Speel EJ, Anton HN (1999) Amplification methods to increase the sensitivity of in situ hybridization: play CARD(S). J Histochem Cytochem 47:281–288
Miao TT, Wang ZP, Li S, Wang X (2011) Sensitive fluorescent detection of Staphylococcus aureus using nanogold linked CdTe nanocrystals as signal amplification labels. Microchim Acta 172:431–437
Ruan M, Niu CG, Zeng GM (2011) Rapid detection of Staphylococcus aureus via a sensitive DNA hybridization assay based on a long-time luminescent europium maker. Microchim Acta 178:105–112
Boujday S, Briandet R, Salmain M, Herry JM, Marnet PG, Gautier M, Pradier CM (2008) Detection of pathogenic Staphylococcus aureus bacteria by gold based immunosensors. Microchim Acta 163:203–209
Sung YJ, Suk HJ, Sung HY, Li T, Poo H, Kim MG (2013) Novel antibody/gold nanoparticle/magnetic nanoparticle nanocomposites for immunomagnetic separation and rapid colorimetric detection of Staphylococcus aureus in milk. Biosens Bioelectron 43:432–439
He Y, Liu HL, Xian MJ (2010) Detection and identification of Staphylococcus aureus in raw milk by hybridization to oligonucleotide microarray. Afr J Biotechnol 9:2284–2289
Acknowledgments
This work was partly supported by the S&T Supporting Project of Jiangsu (BE2011621, BE2010679, BE2012614), the National S&T Support Program of China (2012BAK08B01, 2011YQ170067), NSFC (21375049) NCET-11-0663 and JUSRP51309A.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Yuan, J., Yu, Y., Li, C. et al. Visual detection and microplate assay for Staphylococcus aureus based on aptamer recognition coupled to tyramine signal amplification. Microchim Acta 181, 321–327 (2014). https://doi.org/10.1007/s00604-013-1120-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-013-1120-6