Abstract
Rapid and accurate detection and identification of Staphylococcus aureus (S. aureus) are of great significance for food safety, environmental monitoring, early clinical diagnosis, and prevention of the spread of drug-resistant bacteria. Herein, we design a fluorometric aptasensor for ultra-sensitive, specific, and rapid detection of S. aureus. The apasensor combines the enrichment and separation of magnetic nanoparticles (MNPs), the biotin-streptavidin conjugation system, and a single S. aureus can release four signaling probes for signal amplification. Aptamer acts as a specific biorecognition element of S. aureus. Four FAM-labeled partially complementary sequences (FAM-pcDNAs) were used as signaling probes. The aptamers were sequential hybridized with the four FAM-pcDNAs to form aptamer&pcDNAs, which were then bound to MNPs via the biotin-streptavidin. When the aptamer specifically recognizes and binds to S. aureus, the FAM-pcDNAs signaling probes are replaced and released into the supernatant. The concentration of S. aureus can be quantified by measuring the fluorescence intensity (λexc/em = 492/520 nm) of the replaced signaling probe FAM-pcDNAs. The results show that the proposed fluorometric aptasensor displays good specificity, ultra-high sensitivity (1.23 cfu/mL), wide linear range (1 ~ 108 cfu/mL), and fast detection speed (~ 1.5 h). The recovery test verifies further that the proposed fluorometric aptasensor can detect S. aureus in spiked blood samples. Since aptamers are easy to customize, we believe that fluorometric aptasensors based on multiple amplification have broad prospects in the construction of practical high-performance biosensors for bacterial detection.
Key points
• Multiple amplification-based fluorometric aptasensor for S. aureus is developed
• The aptasensor displays high specificity with a LOD of 1.23 CFU/mL
• The aptasensor can directly detect S. aureus in spiked blood samples
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
S. aureus is one of the most pathogenic pathogens in the world, and was first discovered and named by Dr. Alexander Ogston in 1880 (Ogston 1882). S. aureus naturally exists in water, sewage, air, environmental surfaces, and food (Lian et al. 2015), and produces one or more staphylococcal enterotoxins (SE), such as SEA, SEB, and SEC, etc (Principato and Qian 2014), which are the main causes of food-borne poisoning (Hulme 2017). S. aureus is also the most common pathogen which can cause skin and soft tissue infections (impetigo, folliculitis, and scalded skin syndrome), pneumonia, endocarditis, osteomyelitis, pseudomembranous enteritis, and even serious systemic infections such as bacteremia, sepsis and toxic shock syndrome (Tong et al. 2015). In recent years, methicillin-resistant S. aureus (MRSA) has posed a serious threat to global public health and the infection rate of MRSA remains high in developed countries (40 ~ 60%) as well as developing countries (> 70%) (Chen and Huang 2014; Fluit et al. 2001).
S. aureus has become a pathogen that seriously endangers global public health. Therefore, rapid and accurate detection and identification of S. aureus are of great significance to food safety, environmental monitoring, early clinical diagnosis, and prevention of the spread of drug-resistant bacteria. The gold standard for detecting S. aureus is still cultivation and identification, the results are accurate and reliable, economical and simple, but it is very time-consuming, usually taking 1 ~ 2 days to form visible colonies on agar plates, and then 1 ~ 2 days for biochemical identification and serological typing. There are many newly developed rapid and automated detection methods in recent years, such as enzyme-linked immunosorbent assay (ELISA) (Nouri et al. 2018), polymerase chain reaction (PCR) (Umesha and Manukumar 2018), next-generation sequencing, and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF–MS) (Gill et al. 2019; Rajapaksha et al. 2019), etc., which require only 1 ~ 5 h. However, these methods require expensive instruments, complicated sample preparation, and extensive experience, and most experiments also require culture enrichment and isolation to improve the detection sensitivity, thereby greatly extending the detection time. Therefore, it is still challenging to develop a rapid, specific, and ultra-sensitive detection method for S. aureus.
Aptamer technology has shown its promising application in the fields of medicine and life sciences since the RNA-type oligonucleotide aptamers were screened from a library containing approximately 1015 oligonucleotide molecules by Systematic Evolution of Ligands exponential enrichment (SELEX) (Ellington and Szostak 1990; Tuerk and Gold 1990). Aptamers are a specific oligonucleotide sequences (most commonly single-stranded DNA or RNA), usually ranging in length from 25 to 90 bases, and their binding to the target is based on the diversity of single-stranded nucleic acid structure and spatial conformation (Tok et al. 2000). Aptamers can not only bind to the corresponding ligand with strong affinity (the affinity dissociation constants are generally 1 pM ~ 1 mM) and high specificity similar to antibodies, but also have many advantages compared to antibodies, such as small molecular weight, low synthesis cost, easy of modification, wide selection of ligands (small molecules, ions, proteins, and even cells), no immunogenicity, and a higher specificity, stability, and reproducibility (Labib et al. 2016; Liu et al. 2021a; Moutsiopoulou et al. 2019).
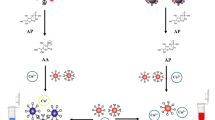
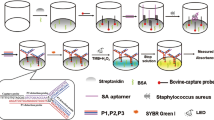
In recent years, biosensors based on aptamers as specific biorecognition elements have been widely used for efficient and rapid identification of pathogens, and have been proposed as the most attractive alternatives for the traditional detection of pathogens (Cesewski and Johnson 2020; Furst and Francis 2019). Fluorometric aptasensor is one of the most commonly used biosensors, with high sensitivity, wide linear range, and fast detection, and has been widely used for pathogen detection (Chen et al. 2022). In this research, we have fabricated a fluorometric aptasensor for ultra-sensitive, specific, and rapid detection of S. aureus. The apasensor combines the enrichment and separation of magnetic nanoparticles (MNPs), the biotin-streptavidin conjugation system, and a single S. aureus can release four signaling probes for signal amplification. As shown in the Scheme 1, biotin-aptamer acts as a specific biorecognition element of S. aureus and forms a hybrid sequence (aptamer&pcDNAs) with four partially complementary sequences (FAM-pcDNAs) through base pairing. Aptamer&pcDNAs immobilize on the streptavidin-MNPS to form aptamer&pcDNAs@MNPS as detection probe due to the strong affinity interaction between biotin-streptavidin. When S. aureus is present in the sample, the aptamer can specifically recognize and binds to S. aureus, and the FAM-pcDNAs signaling probes are replaced and released into the supernatant. The concentration of S. aureus in the sample can be quantified by measuring the fluorescence intensity (λexc/em = 492/520 nm) of the replaced signaling probe FAM-pcDNAs.
Materials and methods
Bacteria and reagents
S. aureus ATCC25923, Staphylococcus epidermidis ATC12228 (S. epidermidis), Klebsiella pneumoniae ATCC 700603 (K. pneumoniae), Acinetobacter baumannii ATCC 19606 (A. baumannii), Pseudomonas aeruginosa ATCC 27853 (P. aeruginosa), Escherichia coli ATCC 25922 (E. coli), Enterococcus faecium ATCC 29212 (E. faecium), Proteus mirabilis ATCC 12453 (P. mirabilis), Streptococcus pyogenes ATCC 19615 (S. pyogenes), and Streptococcus pneumoniae ATCC 49619 (S. pneumoniae) were obtained from the Xiangya Hospital of Central South University. From a pure culture, a single colony was selected for bacterial enrichment and different concentrations of bacteria were prepared with Tris–HCl (10 mM, pH 7.4). The standard plate count method was used to count the bacteria.
Streptavidin-MNPS (50 mg/mL), LB broth medium, Tris–HCl, PBS, agarose, 4S Red Plus nucleic acid stain, running buffer, loading buffer, and rabbit whole blood were purchased from Shanghai Sangon Biological Engineering Technology and Services. The aptamer sequence, partially complementary sequence (pcDNA), and fully complementary sequence (fcDNA) required in this article are shown in Table 1, which were synthesized by Shanghai Sangon Biological Engineering Technology and Services. Washing buffer (10 mM Tris–HCl, with 2 M NaCl, 1 mM EDTA, pH 7.4), blocking buffer (0.1 M NaCl, 0.1 M PBS, 0.05% Tween-20, pH 7.4), and hybridization buffer (10 mM Tris–HCl, with 150 mM NaCl, 5 mM KCl, 2 mM MgCl2, 1 mM CaCl2, pH 7.4) were prepared according to the literature (Shi et al. 2019).
Preparation of the aptamer&pcDNAs
Add 100 μL of different concentrations (2 μM, 4 μM, 6 μM, 8 μM, 10 μM) of FAM-pcDNA1, FAM-pcDNA2, FAM-pcDNA3, and FAM-pcDNA4 to 100 μL of biotin aptamer (5 μM). Heat at 95 °C for 5 min to denature the S. aureus aptamer (Mazzaracchio et al. 2019), and then cool down naturally at room temperature in the dark with shaking (110 r/min) for hybridization. Twenty microliters of each sample were loaded on agarose gel (4%) and electrophoresis was performed at a constant potential of 120 V for 20 min. The gel was photographed, and the gray-scale of the DNA electrophoresis bands on the gel image was analyzed by gel imaging analysis software, thereby semi-quantitative analysis of the DNA content.
Preparation of aptamer&pcDNAs@MNPS
Streptavidin-MNPS was washed three times with washing buffer and resuspended in hybridization buffer. Next, 200 μL aptamer&pcDNAs (1 μM) were added and incubated with 200 μL streptavidin-MNPS (0 mg/mL, 0.25 mg/mL, 0.5 mg/mL, 1 mg/mL, 2 mg/mL, 4 mg/mL, 8 mg/mL, 16 mg/mL) at 37℃ for 30 min in the dark under shaking (110 r/min) to generate aptamer&pcDNAs@MNPS detection probe. The aptamer&pcDNAs@MNPS was further washed twice with 400 μL washing buffer until no DNA was detected in the discarded solution. The fluorescence intensity (λexc/em = 492/520 nm) in all isolated supernatants were measured to estimate the content of unbound aptamer&pcDNAs. Aptamer&pcDNAs@MNPS were then resuspended in blocking buffer (37 ℃, 30 min) to block the exposed unbound sites of the streptavidin-MNPS. Prior to use, the aptamer&pcDNAs@MNPS were resuspended in hybridization buffer and stored at 4 °C in the dark.
Procedure for S. aureus detection
First, S. aureus sample (300 μL) was added to 300 μL aptamer&pcDNAs@MNPS probe solution, and co-incubated in the dark under shaking (110 r/min). After magnetic separation, centrifuge was performed to separate unbound S. aureus from FAM-pcDNAs in the detection solution. Finally, FAM-pcDNAs in the supernatant were transferred to Hitachi F4500 matching cuvette to measure its fluorescence intensity (λexc/em = 492/520 nm).
Results
Preparation of aptamer&pcDNAs
Agarose gel electrophoresis was performed to prove the occurrence of the hybridization of biotin-aptamer with four partially complementary sequences (FAM-pcDNAs). As can be seen in Fig. 1a, lanes 1, 3, and 5 represent aptamer, FAM-pcDNAs, and aptamer&pcDNAs, respectively. The lane 5 has a light band, and the size of the light band is larger than that of the aptamer and the partially complementary sequences (FAM-pcDNAs), indicating that the aptamer&pcDNAs hybrid sequence was successfully prepared. The size of aptamer&pcDNAs in agarose gel (about 200 bp) is larger than the size of one aptamer (88 bases) plus four partially complementary sequences (22 bases), which may be because the modification of -(CH2)6-FAM on the pcDNAs affect the charge, molecular size, and shape in the aptamer&pcDNAs.
a Agarose gel electrophoresis of aptamer (20 μL, 2 μM), pcDNAs (20 μL, 2 μM), and aptamer&pcDNAs (20 μL, 10 μM aptamer + 10 μM of each of the four pcDNA); b the gray value of aptamer&pcDNAs after different concentrations of pcDNAs (2 μM, 4 μM, 6 μM, 8 μM, 10 μM) incubated with aptamer (5 μM).The concentrations ratio of pcDNAs to aptamer was 0.4, 0.8, 1.2, 1.6, 2.0; c the typical UV–Vis absorption curves of the Tris- HCl (Blank), aptamer&pcDNAs, MPNS, and aptamer&pcDNAs@MPNS; d the fluorescence intensity of all separated solution after aptamer&pcDNAs incubated (37 ℃, 30 min) with different concentrations of streptavidin-MNPS (0 mg/mL,0.25 mg/mL,0.5 mg/mL, 1 mg/mL, 2 mg/mL, 4 mg/mL, 8 mg/mL, 16 mg/mL). Error bars indicate standard deviation (n = 3)
We tried to optimize the concentration ratio (0.4, 0.8, 1.2, 1.6, 2.0) between the FAM-pcDNAs (2 μM, 4 μM, 6 μM, 8 μM, 10 μM) and biotin-aptamer (5 μM) to improve hybridization efficiency. As shown in Fig. 1b, the gray-scale of aptamer&pcDNAs increased with the increase of concentration ratio of FAM-pcDNAs to biotin-aptamer from 0.4 to 2.0. When the concentration ratio of FAM-pcDNAs to biotin-aptamer increased up to 1.2, the gray value increased insignificantly. Therefore, the optimal concentration ratio of FAM-pcDNAs to biotin-aptamer is 1.2, and it will be used for subsequent experiments.
Preparation of aptamer&pcDNAs@MNPS
UV–Vis absorption spectroscopy analysis was performed to confirm the formation of the detection probe aptamer&pcDNAs@MNPS. As shown in Fig. 1c, Tris–HCl (Blank) has no absorption peak, while streptavidin-MNPS has an absorption peak at 206 nm, and the baseline is increased, which may be attributed to streptavidin and MNPS respectively. Two absorption peaks were observed at 206 nm and 260 nm for aptamer&pcDNAs, which were characteristic absorption peaks of biotin and DNA respectively. The baseline induced by streptavidin-MNPS also increased, providing evidence for the successful preparation of aptamer&pcDNAs@MNPS.
We tried to optimize the concentration of streptavidin-MNPS (0 mg/mL, 0.25 mg/mL, 0.5 mg/mL, 1 mg/mL, 2 mg/mL, 4 mg/mL, 8 mg/mL, 16 mg/mL) to improve combining efficiency. As shown in Fig. 1d, the fluorescence intensity in the separated solution after aptamer&pcDNAs incubated with streptavidin-MNPS decreased with the increase of concentration of streptavidin-MNPS from 0.25 mg/mL to 16 mg/mL. When the concentration of streptavidin-MNPS was increased up to 8 mg/mL, the fluorescence intensity in the separated solution decreased insignificantly, indicating that almost all aptamer&pcDNAs in solution have been bound to the streptavidin-MNPS. Therefore, the optimal concentration of streptavidin-MNPS was 8 mg/mL, and it was used for subsequent experiments. According to the standard curve established by fluorescence intensity (λexc/em = 492/520 nm) and aptamer&pcDNAs concentration, it can be calculated that about 21.27 pmol (11.50 μg) of aptamer&pcDNAs bind to 1 mg of streptavidin-MNPS when the streptavidin-MNPS concentration is 8 mg/mL.
Characterization of streptavidin-MNPS and aptamer&pcDNAs@MNPS
The streptavidin-MNPS and aptamer&pcDNAs@MNPS were characterized by FTIR, XPS spectra, and zeta potential to further confirm the successful assembly of aptamer&pcDNAs@MNPS. As shown in Fig. 2a, the main characteristic FTIR adsorption peaks of streptavidin-MNPS are 3229 cm−1 and 3189 cm−1 (stretching vibration of O–H and N–H), 2987 cm−1 (stretching vibration of C–H), and 2600 cm−1 (stretching vibration of S–H), 1630 cm−1, 1551 cm−1, and 1460 cm−1 (skeleton vibration of benzene ring), 1403 cm−1 (deformation vibration of -CH3), 1296 cm−1 (deformation vibration of N–H), 1135 cm−1 and 910 cm−1 (stretching vibration of C–O–C), 1035 cm−1 (stretching vibration of C–O), 664 cm−1 (deformation vibration of C–H), and 581 cm−1 and 468 cm−1 (vibration of Fe–O). After the aptamer&pcDNAs immobilized on streptavidin-MNPS, the new characteristic adsorption peaks 1180 cm−1 (stretching vibrations of C-O and P-O-C which mainly in nucleic acids) and 1730 cm−1 (stretching vibration of C = O which mainly in A/T/C/G base cyclic ketone) are observed, indicating the successful assembly of aptamer&pcDNAs@MNPS.
a FT-IR spectra of MPNS and aptamer&pcDNAs@MPNS; b the narrow-scan C1s XPS of MPNS and aptamer&pcDNAs@MPNS; c The narrow-scan N1s XPS of MPNS and aptamer&pcDNAs@MPNS; d the narrow-scan O1s XPS of MPNS and aptamer&pcDNAs@MPNS; e the narrow-scan P2p XPS of MPNS and aptamer&pcDNAs@MPNS; f the zeta potential of MPNS and aptamer&pcDNAs@MPNS
The chemical composition and states of the chemical elements of streptavidin-MNPS and aptamer&pcDNAs@MNPS were analyzed by XPS. We can clearly see the abundant C–O–C, C-O–H (Fig. 2b), and N–H (Fig. 2c) in the aptamer&pcDNAs@MNPS, which are almost absent in streptavidin-MNPS, indicating that the aptamer&pcDNAs@MNPS were assembled successfully. Similarly, we can see that the main form of O1s in aptamer&pcDNAs@MNPS are O-C, O = C, and O–H, which are different from O-Fe in streptavidin-MNPS as (Fig. 2d). What is more, there was a peak at 133.2 eV only for aptamer&pcDNAs@MNPS not for streptavidin-MNPS in the narrow-scan P2p XPS (Fig. 2e), which could be attributed to the phosphate skeleton in nucleic acids, further indicating the successful assembly of aptamer&pcDNAs@MNPS. As shown in Fig. 2f, the zeta potential of Streptavidin-MNPS is about − 26.87 mV, while the zeta potential of aptamer&pcDNAs@MNPS is − 27.57 mV under the same pH condition. This is because the nucleic acid is negatively charged and could make the zeta potential of aptamer&pcDNAs@MNPS more negative.
Optimization of experimental parameters for S. aureus detection
The incubation temperature (4 ℃, 25 °C, 37 ℃) and time (30 min, 60 min, 120 min) of aptamer&pcDNAs@MNPS with S. aureus are the critical experimental parameters that should be optimized to enhance the performance of the aptasensor. As shown in Fig. 3a, the fluorescence intensity of the detection solution increased with the increase of incubation temperature from 4 to 37 ℃ for 120 min. Therefore, 37 ℃ was the optimal temperature for the incubation of aptamer&pcDNAs@MPNS with S. aureus and used for subsequent experiments. As shown in Fig. 3b, the fluorescence intensity of the detection solution increased with the increase of incubation time from 30 to 120 min at 37 ℃. The fluorescence intensity increased insignificantly as the incubation time increased over 60 min. Therefore, the optimal incubation time between the aptamer&pcDNAs@MPNS and S. aureus was determined to be 60 min, and it was used for subsequent experiments.
a Optimization of incubation temperature (4 ℃, 25 ℃, 37 ℃) between the aptamer&pcDNAs@MPNS and S. aureus for 120 min; b optimization of incubation time (30 min, 60 min, 120 min) between the aptamer&pcDNAs@MPNS and S. aureus at 37 ℃; c, d, e, f the fluorescence emission spectra of the detection supernatant contained signaling probe after aptamer&cDNA@MPNS incubation (37 ℃, 60 min) with different concentrations of S. aureus (101 cfu/mL, 104 cfu/mL, 107 cfu/mL): c only one FAM-fcDNA signaling probe, d two FAM-pcDNAs signaling probes, e three FAM-pcDNAs signaling probes, f four FAM-pcDNAs signaling probes
Then, we compared the performance of four FAM-pcDNAs signaling probes with one FAM-fcDNA signaling probes, two FAM-pcDNAs signaling probes, and three FAM-pcDNAs signaling probes. As shown in Fig. 3c, when one FAM-fcDNA signaling probes were used to react with low (101 cfu/mL), medium (104 cfu/mL), and high (107 cfu/mL) concentrations of S. aureus, the relatively low fluorescence intensities were obtained (51.19, 183.52, 747.85, respectively). The fluorescence intensity enhanced with the increase of FAM-pcDNAs bound on the signaling probe. When using the four FAM-pcDNAS signaling probes, the corresponding fluorescence intensities were enhanced about twofold, reaching to 148.55, 596.81, and 2342.73, respectively, indicating that this strategy can be used to improve the detection sensitivity.
The morphology of aptamer&S. aureus@MNPS was observed by scanning electron microscopy (SEM) to verify that the aptamer@MNPS could specifically recognizes and binds to S. aureus, thereby releasing the FAM-pcDNAs signaling probe for detection. The morphology of streptavidin-MNPS (Fig. 4a, b) has no significant difference from that of aptamer&pcDNAs@MNPS (Fig. 4c, d). As shown in Fig. 4e, f, the aptamer@MNPs are obviously bound and aggregated on the surface of approximately spherical S. aureus with a diameter of about 0.8 μm by aptamer-specific biorecognition element after aptamer&pcDNAs@MNPS incubated with S. aureus (108 cfu/mL) at 37 ℃ for 60 min. The results indicate that the FAM-pcDNAs signaling probe could be replaced and released from aptamer&pcDNAs@MNPS by S. aureus, and the content of the replaced FAM-pcDNAs signaling probe is related to the concentration of S. aureus.
a, b SEM image of Streptavidin-MNPS at different magnifications: a × 10,000, b × 50,000; c, d SEM image of aptamer&pcDNAs@MPNS at different magnifications: c × 10,000, d × 50,000; e, f SEM image of aptamer&S.aureus@MPNS at different magnifications: e × 10,000, f × 50,000. The arrow points to the S. aureus
The linearity, sensitivity, and selectivity of fluorometric aptasensor
Figure 5a shows the fluorescence emission spectra of the detection supernatant after incubation of aptamer&pcDNAs@MNPS with different concentrations of S. aureus (1 ~ 108 cfu/mL) at 37 ℃ for 60 min. The fluorescence intensity of the separated supernatant increased with the increase of S. aureus concentration from 1 to 108 cfu/mL. It displays a good linear relationship between the logarithm of the fluorescence intensity and the logarithm of the S. aureus concentration from 1 to 108 cfu/mL as shown in Fig. 5b. The linear regression equation could be expressed as logF (a.u.) = 0.22logC (cfu/mL) + 1.87 (R2 = 0.99). The limit of detection (LOD) and limit of quantification (LOQ) were calculated according to the following equations (Zhou et al. 2017): LOD = 3σ/k and LOQ = 10σ/k, where σ = 0.0899 (standard deviation of Tris–HCl solution), k = 0.22 (slope of the calibration curve). Our LOD and LOQ are calculated to be 1.23 cfu/mL and 4.09 cfu/mL, respectively.
a The fluorescence emission spectra of the detection supernatant contained FAM-pcDNAs after aptamer&pcDNAs@MPNS incubation (37 ℃, 60 min) with different concentrations of S. aureus (1 ~ 108 cfu/mL); b the relationship curve between the logarithmic values of fluorescence intensity with the logarithmic values of S. aureus concentration (1 ~ 108 cfu/mL); c selectivity of the aptasensor toward different bacteria. The concentration of S. aureus is 104 cfu/mL and the concentration of other bacteria is 108 cfu/mL; error bars indicate standard deviation (n = 3)
The aptamer&pcDNAs@MPNS were incubated with Tri-HCl (blank) at 37 ℃ for 60 min to determine whether the FAM-pcDNAs signaling probes would spontaneously detach from the aptamer. The results in Fig. 5c show that the probe FAM-pcDNAs detached hardly from the aptamer spontaneously as the fluorescence intensity of the separated supernatant changed little before and after the reaction between aptamer&pcDNAs@MPNS with Tri-HCl. In order to explore the specificity of the aptasensor, the prepared aptamer&pcDNAs@MPNS were incubated with E. coli, A. baumannii, E. faecium, P. aeruginosa, P. mirabilis, S. epidermidis, S. pyogenes, S. pneumoniae, and K. pneumoniae at 37 ℃ for 60 min. As shown in Fig. 5c, the fluorescence intensity of S. aureus (104 cfu/mL) is much stronger than that of other bacteria (108 cfu/mL), indicating that there is no non-specific binding between aptamer&pcDNAs@MPNS and other bacteria. The above results showed that the proposed aptasensor owns excellent specificity due to the high specificity of aptamer.
Blood sample analysis
The proposed fluorometric aptasensor was used for the recovery test of blood samples spiked with different concentrations of S. aureus from 103 to 106 cfu/ mL to validate the feasibility of the method. As shown in Table 2, compared with the plate counting method, the spiked recovery rate between 90.6% and 116.2% demonstrated the applicability of fluorometric aptasensor for the detection of S. aureus in blood samples.
Discussions
The aptamer has higher specificity and affinity, can specifically capture S. aureus and distinguish it from other bacteria, endowing the apatsensor good selectivity. The high sensitivity of aptasensors is usually achieved by using one or more signaling amplification methods (enzyme-catalyzed reactions and/or nanomaterials) (Table 3). However, the wide application of enzyme-catalyzed reactions such as rolling cycle amplification (RCA) (Xu et al. 2018), strand displacement reaction (SDA) (Cai et al. 2020), hybridization Chain reaction (HCR) (Chai et al. 2021), and hairpin DNA cascade hybridization reaction (HD-CHR) (Borum and Jokerst 2021) are limited due to their complex reactions, harsh conditions, and susceptibility to environmental factors. Nanomaterials, as efficient transducers and amplifiers, play an increasingly important role in electrochemical-based aptasensors (such as differential pulse voltammetry (DPV) and electrochemical impedance spectroscopy (EIS)) (Abbaspour et al. 2015; Ranjbar and Shahrokhian 2018), but their stability and reproducibility still need to be improved. Novel optical methods such as surface enhanced Raman scattering (SERS) were also limited by the tedious sample preparation and the expensive equipment required (Liu et al. 2021b). In our research, we design a fluorometric aptasensor with comparable high sensitivity (1.23 cfu/mL) to other conjugated enzymes or/and nanomaterials studies by simply combining the enrichment and separation of MNPs, the biotin-streptavidin conjugation system, and a single S. aureus can release four signaling probes. The proposed aptasensor also showed a wide linear range from 1 to 108 cfu/mL and fast detection speed (~ 1.5 h). In the follow-up study, we will explore the use of different fluorescently labeled detection probes to achieve simultaneous detection of multiple pathogens, which is of great significance in practical applications.
To be acknowledged, there are still some limitations need to be addressed when applying the proposed fluorometric aptasensor for real samples analysis. First, the fluorescent molecules (FAM) used in our proposed fluorometric aptasensor are unstable, have low fluorescence intensity, and are prone to photobleaching, so the entire experiment needs to be performed in the dark and is susceptible to interference by autofluorescence or background fluorescence, which is not conducive to practical applications. Compared with traditional organic fluorescent dyes (FAM, fluorescein isothiocyanate, and acridine orange), fluorescent nanomaterials such as quantum dots (QDs) (Fu et al. 2020; Tao et al. 2021), upconversion nanoparticles (UCNPs) (Ouyang et al. 2021a, 2021b) and carbon dots (Cui et al. 2019; Pebdeni et al. 2020) that have emerged in recent years have many advantages, such as wider absorption spectra, size-tunable narrower emission spectra, and better stability, which can be subsequently used to improve the proposed fluorometric aptasensor. Second, although the proposed fluorometric aptasensor achieved good results in buffers (simple components) and good recoveries in blood samples (103 ~ 106 cfu/mL) by optimizing the experimental conditions, the interferences from multiple substances (ions, small molecules, proteins or carbohydrates, etc.) in blood sample requires further optimization to ensure their compatibility in real samples. Third, simultaneous detection of multiple pathogens with the proposed fluorometric aptasensor in real samples still needs to be explored.
In summary, although there are still some challenges to be addressed, the proposed multiple amplification-based fluorometric aptasensor have broad prospects in the construction of practical high-sensitive biosensors for early detection of bacteria due to the easy replacement of aptamers.
Data availability
All data generated or analyzed during this study are included in this published article.
References
Abbaspour A, Norouz-Sarvestani F, Noon A, Soltani N (2015) Aptamer-conjugated silver nanoparticles for electrochemical dual-aptamer-based sandwich detection of staphylococcus aureus. Biosens Bioelectron 68:149–155. https://doi.org/10.1016/j.bios.2014.12.040
Borum RM, Jokerst JV (2021) Hybridizing clinical translatability with enzyme-free DNA signal amplifiers: recent advances in nucleic acid detection and imaging. Biomater Sci 9(2):347–366. https://doi.org/10.1039/d0bm00931h
Cai R, Yin F, Chen H, Tian Y, Zhou N (2020) A fluorescent aptasensor for Staphylococcus aureus based on strand displacement amplification and self-assembled DNA hexagonal structure. Mikrochim Acta 187(5):304. https://doi.org/10.1007/s00604-020-04293-9
Cao X, Li S, Chen L, Ding H, Xu H, Huang Y, Li J, Liu N, Cao W, Zhu Y, Shen B, Shao N (2009) Combining use of a panel of ssDNA aptamers in the detection of Staphylococcus aureus. Nucleic Acids Res 37(14):4621–4628. https://doi.org/10.1093/nar/gkp489
Cesewski E, Johnson BN (2020) Electrochemical biosensors for pathogen detection. Biosens Bioelectron 159:112214. https://doi.org/10.1016/j.bios.2020.112214
Chai H, Cheng W, Jin D, Miao P (2021) Recent progress in DNA hybridization chain reaction strategies for amplified biosensing. ACS Appl Mater Interfaces 13(33):38931–38946. https://doi.org/10.1021/acsami.1c09000
Chen CJ, Huang YC (2014) New epidemiology of Staphylococcus aureus infection in Asia. Clin Microbiol Infect 20(7):605–623. https://doi.org/10.1111/1469-0691.12705
Chen W, Lai Q, Zhang Y, Liu Z (2022) Recent advances in aptasensors for rapid and sensitive detection of Staphylococcus aureus. Front Bioeng Biotechnol 10:889431. https://doi.org/10.3389/fbioe.2022.889431
Cui F, Sun J, Habimana JdD, Yang X, Ji J, Zhang Y, Lei H, Li Z, Zheng J, Fan M, Sun X (2019) Ultrasensitive fluorometric angling determination of Staphylococcus aureus in vitro and fluorescence imaging in vivo using carbon dots with full-color emission. Anal Chem 91(22):14681–14690. https://doi.org/10.1021/acs.analchem.9b03916
Ellington AD, Szostak JW (1990) In vitro selection of RNA molecules that bind specific ligands. Nature 346(6287):818–822. https://doi.org/10.1038/346818a0
Fluit AC, Verhoef J, Schmitz FJ (2001) Frequency of isolation and antimicrobial resistance of gram-negative and gram-positive bacteria from patients in intensive care units of 25 European university hospitals participating in the European arm of the SENTRY Antimicrobial Surveillance Program 1997–1998. Eur J Clin Microbiol Infect Dis 20(9):617–625. https://doi.org/10.1007/s100960100564
Fu F, Zhang Y, Li L, Wang H, Li Q, Tao X, Song Y, Song E (2020) Intracellular pathogen detection based on dual-recognition units constructed fluorescence resonance energy transfer nanoprobe. Anal Chem 92(16):11462–11468. https://doi.org/10.1021/acs.analchem.0c02695
Furst AL, Francis MB (2019) Impedance-based detection of bacteria. Chem Rev 119(1):700–726. https://doi.org/10.1021/acs.chemrev.8b00381
Gill AAS, Singh S, Thapliyal N, Karpoormath R (2019) Nanomaterial-based optical and electrochemical techniques for detection of methicillin-resistant Staphylococcus aureus: a review. Mikrochim Acta 186(2):114. https://doi.org/10.1007/s00604-018-3186-7
Hulme J (2017) Recent Advances in the detection of methicillin resistant Staphylococcus aureus (MRSA). BioChip J 11(2):89–100. https://doi.org/10.1007/s13206-016-1201-9
Labib M, Sargent EH, Kelley SO (2016) Electrochemical methods for the analysis of clinically relevant biomolecules. Chem Rev 116(16):9001–9090. https://doi.org/10.1021/acs.chemrev.6b00220
Lian Y, He F, Wang H, Tong F (2015) A new aptamer/graphene interdigitated gold electrode piezoelectric sensor for rapid and specific detection of Staphylococcus aureus. Biosens Bioelectron 65:314–319. https://doi.org/10.1016/j.bios.2014.10.017
Liu LS, Wang F, Ge Y, Lo PK (2021a) Recent developments in aptasensors for diagnostic applications. ACS Appl Mater Interfaces 13(8):9329–9358. https://doi.org/10.1021/acsami.0c14788
Liu S, Li H, Hassan MM, Ali S, Chen Q (2021b) SERS based artificial peroxidase enzyme regulated multiple signal amplified system for quantitative detection of foodborne pathogens. Food Control 123. https://doi.org/10.1016/j.foodcont.2020.107733
Mazzaracchio V, Neagu D, Porchetta A, Marcoccio E, Pomponi A, Faggioni G, D’Amore N, Notargiacomo A, Pea M, Moscone D, Palleschi G, Lista F, Arduini F (2019) A label-free impedimetric aptasensor for the detection of Bacillus anthracis spore simulant. Biosens Bioelectron 126:640–646. https://doi.org/10.1016/j.bios.2018.11.017
Moutsiopoulou A, Broyles D, Dikici E, Daunert S, Deo SK (2019) Molecular aptamer beacons and their applications in sensing, imaging, and diagnostics. Small 15(35):e1902248. https://doi.org/10.1002/smll.201902248
Nouri A, Ahari H, Shahbazzadeh D (2018) Designing a direct ELISA kit for the detection of Staphylococcus aureus enterotoxin A in raw milk samples. Int J Biol Macromol 107(Pt B):1732–1737. https://doi.org/10.1016/j.ijbiomac.2017.10.052
Ogston A (1882) Micrococcus poisoning. J Anat Phys 17(Pt 1):24–58
Ouyang Q, Wang L, Ahmad W, Yang Y, Chen Q (2021a) Upconversion nanoprobes based on a horseradish peroxidase-regulated dual-mode strategy for the ultrasensitive detection of Staphylococcus aureus in meat. J Agric Food Chem 69(34):9947–9956. https://doi.org/10.1021/acs.jafc.1c03625
Ouyang Q, Yang Y, Ali S, Wang L, Li H, Chen Q (2021b) Upconversion nanoparticles-based FRET system for sensitive detection of Staphylococcus aureus. Spectrochim Acta, Part A 255. https://doi.org/10.1016/j.saa.2021.119734
Pebdeni AB, Hosseini M, Ganjali MR (2020) Fluorescent turn-on aptasensor of Staphylococcus aureus based on the FRET between green carbon quantum dot and gold nanoparticle. Food Anal Method 13(11):2070–2079. https://doi.org/10.1007/s12161-020-01821-4
Principato M, Qian BF (2014) Staphylococcal enterotoxins in the etiopathogenesis of mucosal autoimmunity within the gastrointestinal tract. Toxins 6(5):1471–1489. https://doi.org/10.3390/toxins6051471
Rajapaksha P, Elbourne A, Gangadoo S, Brown R, Cozzolino D, Chapman J (2019) A review of methods for the detection of pathogenic microorganisms. Analyst 144(2):396–411. https://doi.org/10.1039/c8an01488d
Ranjbar S, Shahrokhian S (2018) Design and fabrication of an electrochemical aptasensor using Au nanoparticles/carbon nanoparticles/cellulose nanofibers nanocomposite for rapid and sensitive detection of Staphylococcus aureus. Bioelectrochemistry 123:70–76. https://doi.org/10.1016/j.bioelechem.2018.04.018
Shi X, Zhang J, He F (2019) A new aptamer/polyadenylated DNA interdigitated gold electrode piezoelectric sensor for rapid detection of Pseudomonas aeruginosa. Biosens Bioelectron 132:224–229. https://doi.org/10.1016/j.bios.2019.02.053
Tao X, Liao Z, Zhang Y, Fu F, Hao M, Song Y, Song E (2021) Aptamer-quantum dots and teicoplanin-gold nanoparticles constructed FRET sensor for sensitive detection of Staphylococcus aureus. Chin Chem Lett 32(2):791–795. https://doi.org/10.1016/j.cclet.2020.07.020
Tok JB, Cho J, Rando RR (2000) RNA aptamers that specifically bind to a 16S ribosomal RNA decoding region construct. Nucleic Acids Res 28(15):2902–2910. https://doi.org/10.1093/nar/28.15.2902
Tong SY, Davis JS, Eichenberger E, Holland TL, Fowler VG Jr (2015) Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev 28(3):603–661. https://doi.org/10.1128/cmr.00134-14
Tuerk C, Gold L (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science (new York, NY) 249(4968):505–510. https://doi.org/10.1126/science.2200121
Umesha S, Manukumar HM (2018) Advanced molecular diagnostic techniques for detection of food-borne pathogens: current applications and future challenges. Crit Rev Food Sci Nutr 58(1):84–104. https://doi.org/10.1080/10408398.2015.1126701
Xu J, Guo J, Maina SW, Yang Y, Hu Y, Li X, Qiu J, Xin Z (2018) An aptasensor for staphylococcus aureus based on nicking enzyme amplification reaction and rolling circle amplification. Anal Biochem 549:136–142. https://doi.org/10.1016/j.ab.2018.03.013
Zhou Y, Marar A, Kner P, Ramasamy RP (2017) Charge-directed immobilization of bacteriophage on nanostructured electrode for whole-cell electrochemical biosensors. Anal Chem 89(11):5734–5741. https://doi.org/10.1021/acs.analchem.6b03751
Funding
This research work was supported financially by the National Natural Science Foundation of China (Grant no. 61771493) and the Natural Science Foundation of Hunan province (Grant No. 2018JJ2532 and 2022JJ40771).
Author information
Authors and Affiliations
Contributions
WC and ZCL conceived and designed research. WC, QTL, and YZL conducted experiments. WC and YKZ analyzed data and wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This work does not involve any human participation nor live animals performed by any of the listed authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, W., Zhang, Y., Lai, Q. et al. Multiple amplification-based fluorometric aptasensor for highly sensitive detection of Staphylococcus aureus. Appl Microbiol Biotechnol 106, 6733–6743 (2022). https://doi.org/10.1007/s00253-022-12057-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12057-z