Abstract
A colorimetric microplate assay for determination of Staphylococcus aureus DNA is described. Linear padlock probes were designed to recognize target sequences. After DNA binding, the linear padlock probes were circularized by ligation and then hybridize with biotin-labeled capture probes. Biotin-labeled capture probes act as primers to initiate the RCA. The biotin-labeled RCA products hybridize with digoxin-labeled signal probes fixed on streptavidin-functionalized wells of a 96-well plate. To enhance sensitivity, an AuNP-anti-digoxigenin-POx-HRP conjugate was added to the wells and then bound to digoxin-labeled signalling probes. The oxidation of tetramethylbenzidine (TMB) by H2O2 produces a color change from colorless to blue via HRP catalysis. After the reaction was terminated, absorbance is measured at 450 nm. For target sequences of Staphylococcus aureus, the detection limit is 1.2 pM. For genomic DNA, the detection limit is 7.4 pg.μL−1. The potential application of the method was verified by analyzing spiked food samples.

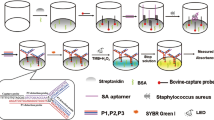
Schematic representation of rolling circle amplification and functionalized AuNP-based colorimetric determination of Staphylococcus aureus. The method uses streptavidin-functionalized 96-well plates and RCA as a molecular tool and AuNP-anti-digoxigenin-POx-HRP as signal transduction markers to increase sensitivity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Staphylococcus aureus (S. aureus) can be found in many foods and can cause food poisoning, skin infections and even sepsis, and being considered as an important food-borne pathogen [1,2,3,4,5]. The gold standard for determination of S. aureus is direct microscopic examination and culture, but it requires 4–7 days and is tedious. It is necessary to develop quickly and accurately method for determination of S. aureus, which will be helpful to control pathogens [6].
Some rapid methods have been developed, among which the molecular recognition method is the most important. Nucleic acid amplification plays a major role in the determination of bacterial pathogens [7], many methods of nucleic acid amplification have been developed such as nested-PCR [8], real time-PCR [9] and nucleic acid sequence-based amplification (NASBA) [10]. However, these methods require sophisticated thermo-cycling systems to amplify the target DNA as well as expensive instruments to report the results. Rolling circle amplification (RCA), as a signal-amplification tool, have been studied [7, 11,12,13] The RCA include three major steps: the first, target sequences are hybridized perfectly with the 5′ and 3′ terminus of linear padlock probes, it is disallowing any mismatch (even if single-nucleotide mismatch) [7]. The ends of linear padlock probes are ligated by E. coli DNA ligase. The second, Exonuclease I and Exonuclease III are added into the above-ligation resulting mixture to remove the non-circularized padlock probes. Because non-circularized padlock probes may produce self-ligation amplification in the subsequent RCA amplification [14]. The end, the capture probes as primer initiates the RCA reaction under constant temperature via phi29 DNA polymerase catalysis, the RCA products are very long reduplicated single-strand DNA sequences that linked to capture probes. RCA is not only the mismatch being restrained but also has a low incidence of false positive results compared to PCR [11]. RCA has been used successfully to the determination of many kinds of microorganism [11, 15, 16]. The RCA products can be used for on-chip detection through signal centralization and site-anchored amplification [7, 17, 18]. The RCA products also can hybridize with a series of signal probes to output various response signals [13, 18, 19].
RCA-based biosensors have been developed to detect bacterial pathogens, such as RCA based optomagnetic determination [20], RCA-based electrochemical biosensors [21], gold nanowire based electrical DNA biosensors [16], RCA-based surface plasmon resonance (SPR) biosensors [14], RCA-based quartz crystal microbalance (QCM) biosensors [22] and RCA-based terahertz (THz) spectroscopy biosensors [23], etc. These RCA-based assays are sensitive but require expensive instruments, restricting their utility. The nanomaterial-based colorimetric methods shows satisfactory results and the results can be visually observed [5, 24]. Herein, a novel RCA-based colorimetric method was established in 96-well plates for determination of S. aureus. The 96-well plates offered a high throughput platform, so this RCA-based colorimetric method was significant advantages in high throughput, ultrasensitive and rapid determination.
Materials and methods
Materials and apparatus
The single-stranded oligonucleotides were synthesized by Genewiz Biotechnology Co., Ltd. (Suzhou, China, www.genewiz.com). These sequences were showed in Table S1. E. coli DNA ligase, Exonuclease I, Exonuclease III, and Phi29 DNA polymerase were purchased from New England Biolabs (USA, www.neb-online.de). QuickCut™ EcoRI, QuickCut™ HindIII and QuickCut™ BamHI were purchased from Takara (Dalian, China, www.takarabio.com). TIANamp Bacteria DNA Kit was obtained from TIANGEN (Beijing, China, www.tiangen.com). Anhydrous citric acid trisodium salt (98%) was obtained from Acros Organics (USA, www.acros.com). The HAuCl4, streptavidin (SA), anti-digoxigenin-POx, horseradish peroxidase (HRP), bovine serum albumin (BSA), N-ethyl-N′-(3-di-methylaminopropyl) carbodiimide (EDC), sodium acetate, dimethyl sulfoxide (DMSO), β-cyclodextrin, urea hydrogen peroxide, and tetramethylbenzidine (TMB) were purchased from Sigma-Aldrich (USA, www.sigmaaldrich.com). Sodium carbonate and sodium bicarbonate were purchased from Sinopharm Chemical Reagent Co., Ltd. (Shanghai, China, www.sinoreagent.com). Biotin-secondary antibody was obtained from Dingguo Changsheng Biotechnology Co., Ltd. (Beijing, China, www.dingguo.com). The F96 Maxisorp nunc-immuno plate (96-well plates) was purchased from Thermo scientific (Denmark, www.thermofisher.com). All other chemicals were analytical grade. The substrate solution A consisted of 100 mM sodium acetate-citric acid buffer (pH 5.0), 4.56 mM urea hydrogen peroxide, and 2.2 mM β-cyclodextrin. The substrate solution B was prepared by adding 42 mM TMB into DMSO. The working solution contained 100 mM Tris-HCl (pH 7.5), 5% (w/v) saccharose, 0.5% (w/v) BSA, and 0.5% (v/v) PEG 200. 1 L washing buffer (PBST pH 7.4) consisted of 1.79 g NaH2PO4·2H2O, 13.76 g Na2HPO4·12H2O, 9 g NaCl and 5 mL 10% (w/w) Tween-20. The solutions were prepared with deionized water from a Milli-Q system (Millipore, Billerica, MA, USA, www.merckmillipore.com). UV–vis absorption was measured on a microplate reader (Thermo fisher, USA, www.thermofisher.com).
Preparation of streptavidin-functionalized 96-well plates
Initially, biotin-secondary antibody (0.25 μg well−1) was placed in the 96-well plates and then incubated at 37 °C for 3 h. Excess biotin-secondary antibody was removed and the 96-well plates were washed with washing buffer. 200 μL well−1 of 0.5% skim milk (w/v) was added into the 96-well plates to prevent nonspecific adsorption. After incubating at 37 °C for 1 h, the 96-well plates were washed with washing buffer. Subsequently, 1 μg well−1 streptavidin were added into the 96-well plates and then incubated at 37 °C for 1 h. Excess reagent was decanted and then the 96-well plates were washed with washing buffer, being stored at 4 °C.
Synthesis of AuNP-anti-digoxigenin-POx-HRP
AuNPs is a colloidal solution forming electron dense regions between negatively charged particles. The AuNPs is able to associate with proteins via Van der Waals forces, the proteins forming a shell around each AuNPs particle [25].To increase the loading amount of HRP toward one reaction event, HRP was conjugated to anti-digoxigenin-POx via condensation reaction between the amidogen and the carboxyl in the presence of EDC [26], and then was conjugated with AuNPs [27] (Fig. 1b).
Schematic diagram of RCA colorimetric method for determination of S. aureus DNA. a Diagram of the linear padlock probes. The linear padlock probes included: the test region which complemented with target sequences (T1 and T2); the general region which offer tandem reduplicated sequences on the RCA products for signal probe hybridizing (G); the special part which can be recognized by capture probe (S). b Principle of the synthetic of AuNP-anti-digoxigenin-POx-HRP conjugates. c Schematic illustration of the RCA colorimetric method.
AuNPs (~15 nm diameter) were prepared as following [28]. All glassware was immersed in chromic acid for 24 h and rinsed with deionized water, then dried by oven. Before being used, trisodium citrate solution and HAuCl4 were filtered through a 0.22-μm filter. 2.25 mL of 1% (w/v) trisodium citrate was added into 100 mL boiling 0.01% (w/v) HAuCl4 with vigorous stirring. The color of the mixture turned from pale yellow to deep blue, the final color changed to wine red. The wine-red mixture solution kept boiling for 15 min with continuous stirring, then was moved away heating source. The mixture solution was stirred continuously until cooled to room temperature. The product was stored in dark place at 4 °C and used to synthesize AuNP-anti-digoxigenin-POx-HRP conjugates.
AuNP-anti-digoxigenin-POx-HRP were prepared by following the steps. Firstly, the AuNPs solution was adjusted to pH 6.0 using 0.1 M K2CO3, and then the anti-digoxigenin-POx, HRP (the enzyme activity ratio of anti-digoxigenin-POx: HRP = 1:375) and 20 μM EDC were added. The mixture was stirred for 3 h and then centrifuged at 11603 rcf for 20 min at 4 °C. Secondly, the precipitate was re-suspended in the mixed solution of final concentration of 0.8% BSA (w/v) and 0.4% PEG20000 (w/v), keeping for 30 min, being centrifuged at 11603 rcf for 20 min at 4 °C. Finally, the precipitate was re-suspended in the working solution and stored at 4 °C for subsequent experiment.
The RCA-based colorimetric method
Rolling circle amplification reaction
The linear padlock probes and the target sequence were denatured at 95 °C for 5 min and then immediately put into ice bath for 10 min. Next, the mixture was incubated at 50 °C for 60 min, and then 10 U of E. coli DNA ligase was added. The ligation mixture incubated at 30 °C for 60 min. To remove non-circularized padlock probes and linear oligonucleotides in the ligation mixture, 10 μL of Exonuclease mixture (1 × buffer I, 10 U Exonuclease I, and 10 U Exonuclease III) was added into each reaction. The mixture solution was incubated at 37 °C for 1 h and then was terminated by inactivated at 95 °C for 15 min. The end, the above-products was added into 25 μL RCA reaction system (1× phi 29 DNA polymerase buffer, 1 μM biotin labeled capture probes), and incubated at 50 °C for 30 min. Then, added 5 U phi29 DNA polymerase, 1 mM dNTPs and 5 μg BSA into this RCA reaction system, which was incubated at 35 °C for 90 min and inactivated at 65 °C for 10 min. The resulting products, which were 5′ terminus biotin labeled long reduplicated single-strand DNA sequences, were used for further analysis.
Microbiological determination on the streptavidin functionalized 96-well plates
First, the 5′ terminus biotin labeled RCA products hybridized with signal probes (1 μL of the RCA products were hybridized with 2 μM signal probes at 60 °C for 30 min) were added into the streptavidin-functionalized 96-well plates and incubated at 37 °C for 30 min, then the plates were washed by washing buffer. Second, after adding 100 μL AuNP-anti-digoxigenin-POx-HRP, the plates were incubated at 37 °C for 30 min, and then washed with washing buffer. Final, the substrate mixture (contain substrate solution A and substrate solution B) was added into the plates, then the plates were placed at room temperature for 20 min, the stop solution being added into the plates to stop the reaction. The result was measured by microplate reader at 450 nm.
Preparation of bacterial genomic DNA
E. coli O157: H7 (ATCC 35150), E. coli (CICC 10305), S. aureus (ATCC 25923), S. epidermidis (ATCC 35984), S. saprophyticus (ATCC 15305), Shigella soonei (CTCC 21535), L. monocytogenes (ATCC 7644), and P. aeruginosa (ATCC 27853) were used in this experiment. They were cultured overnight at 37 °C in LB medium. Genomic DNA was isolated according to the instructions of TIANamp Bacteria DNA Kit. The extracted DNA was eluted with TE buffer and stored at −20 °C. The concentration of microbial genomic DNA was determined according to the manufacturer’s instructions. Before the RCA reaction, the extracted DNA was fragmented through enzymolysis using Quick Cut™ Bam HI, Quick Cut™ Hind III, and Quick Cut™ Eco RI [29] for 30 min.
Preparation of artificial contaminated food samples
Milk and chicken breast were purchased from local markets and washed with 0.85% NaCl (sterile). The chicken breast sample was homogenized (samples/sterile 0.85%NaCl 1:10) by homogenizer with 8000 r min−1 for 2 min. The milk sample was 10-fold diluted with sterile 0.85% NaCl. Two groups of artificial contaminated milk samples and chicken breast samples were prepared. The S. aureus of exponential growth phase was added into the milk samples and the chicken breast samples. The above sample diluting solution was put into 7.5%NaCl broth medium, and incubated at 37 °C for overnight. The bacterial genomic DNA was isolated according to the instructions of TIANamp Bacteria DNA Kit. The extracted DNA was eluted with TE buffer and stored at −20 °C. Before being analyzed by the proposed RCA-based colorimetric method, the extracted DNA was fragmented through enzymolysis using Quick Cut™ Bam HI, Quick Cut™ Hind III, and Quick Cut™ Eco RI for 30 min.
Results and discussion
Principle of the RCA-based colorimetric method
Principle of probes design
Specific sequences in the 16S rDNA of the S. aureus were selected as the target sequences. For accurate and rapid identification of S. aureus, three kinds of probes were designed to the established RCA-based colorimetric method, which included linear padlock probes, biotin-labeled capture probes, and digoxin-labeled signal probes (Table S1).
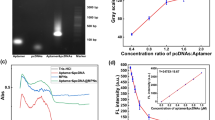
As shown in Fig. 1a, the linear padlock probes include three parts: T1 and T2 represent the test region locating at the 5′ and 3′terminus, this region is complementary to target sequences; G indicate a general region which provides the reduplicated sequences on the RCA products for signal probe hybridizing; S represent a special part which can be recognized by the 5′terminus biotin-labeled capture probe. For making sure the linear padlock probes effectively recognizing the target sequence, asymmetrical design was applied to the linear padlock probes, which was lengthening the 5′terminus sequence and shortening the 3′terminus sequence [14]. The linear padlock probes can discriminate the single-nucleotide mismatch [7, 14]. This design improved the recognition ability of the linear padlock probes to the target sequence effectively. In order to substantiate the above designs, the products of the ligation reactions and the RCA were analyzed by 2% agarose gel electrophoresis. As shown in Fig. S1A, the migration velocity, which the products of linear padlock ligation treated with target sequences, is slower than only the linear padlock probes. This confirm that the linear padlock probes are ligated into a ring only when the target sequences being present. Fig. S1B display that the RCA products only from target sequences has a band near the sample hole, but the RCA products of single-nucleotide mismatch and negative control not have it. It is verified that the probes of the RCA had the ability to identify single-nucleotide mismatch.
Each capture probes contain an additional (C)10 space linker at 5′terminus, which can improve the efficiency of circularized padlock probes hybridization and enhance signal fixation on the 96-well plate. The signal probes have the same (C)10 space linker as the capture probes.
Principle of the RCA-based colorimetric method
As shown in Fig. 1c, the principle of the RCA-based colorimetric method is presented schematically. The key of the RCA-based colorimetric method is the RCA reaction. The RCA include three major steps: the first, target sequence hybridize perfectly with the 5′ and 3′ terminus of linear padlock probes, and then them are ligated and circularized by E. coli DNA ligase, disallowing any mismatch (even if single-nucleotide mismatch). The second, Exonuclease I and Exonuclease III are added into the above-ligation mixture to remove the unreacted padlock probes, which is able to produce self-ligation amplification in the subsequent RCA amplification. The self-ligation amplification can cause strong background signals [14]. The end, the biotin-labeled capture probes act as primers to initiate the RCA reaction. The RCA products are 5′ terminus biotin-labeled long single-stranded DNAs, which have tandem reduplicated sequences [7].
5′ terminus digoxigenin-labeled signal probes hybridize with the general region of RCA products. This hybridized mixture is fixed on streptavidin-functionalized wells of a 96-well plate. To enhance sensitivity, AuNP-anti-digoxigenin-POx-HRP is added to the wells and then bound to digoxigenin-labeled signal probes. Finally, the oxidation of tetramethylbenzidine (TMB) by H2O2 produces a color change from colorless to blue via HRP catalysis. After the reaction is terminated, which is measured by microplate reader at 450 nm.
Preparation of AuNP-anti-digoxigenin-POx-HRP
Optimizing the synthetic of AuNP-anti-digoxigenin-POx-HRP
The following parameters were optimized to get AuNP-anti-digoxigenin-POx-HRP conjugated: (a) the pH of reaction mixture; (b) concentration of EDC; (c) the ratio of anti-digoxigenin-POx: HRP. Respective data and Figures are given in the Electronic Supporting Material.
In short, the following experimental conditions were found to give best results: (a) Optimal pH of reaction mixture: pH 6.0; (b) Optimal concentration of EDC: 20 μM EDC; (c) Best ratio of anti-digoxigenin-POx and HRP: 1 U:375 U.
Characterization of AuNP-anti-digoxigenin-POx-HRP
The AuNP-anti-digoxigenin-POx-HRP conjugates were rinsed three times, and then were characterized by transmission electron micrographs (TEM), UV–vis absorption spectrum, and FTIR spectra. The TEM of AuNP-anti-digoxigenin-POx-HRP conjugates and AuNPs are shown in Fig.S3. The TEM displays that it is not significant difference between the AuNPs and the AuNP-anti-digoxigenin-POx-HRP conjugates.
As Fig. 2a showing, the UV–vis absorption spectrum of the functionalized and non-functionalized AuNPs all have a plasmon band near 520 nm, which is due to the surface plasmon resonance of the AuNPs [26]. It illustrate that the functionalized AuNPs had dispersity as good as the AuNPs, the result is consistent with the TEM. It is difference between the UV–vis absorption spectrum of functionalized and non-functionalized AuNPs. The functionalized AuNPs have an absorption at 280 nm which is characteristic protein peak, proving the anti-digoxigenin-POx-HRP conjugates being attached to the AuNPs.
The FTIR spectra of AuNPs and AuNP-anti-digoxigenin-POx-HRP conjugates are shown in Fig. 2b. Comparing with the FTIR spectrum of AuNPs, new characteristic peaks of the functionalized AuNPs appear at 1578 cm−1 (corresponding to the bending vibration of N-H), 1415 cm−1, 1337 cm−1 and 1146 cm−1 (stretching vibrations of C-N), and 843 cm−1 (out-of-plane bending vibration). The broad band at near 3476 cm−1 became stronger, sharper and appeared a slight drift, which is assigned to the stretching vibration of N-H. In addition, the peak at near 1643 cm−1 became stronger is corresponded to the stretching vibration of C=O. These changes suggested that the anti-digoxigenin-POx-HRP conjugates are combined with AuNPs successfully.
Signal amplification effect of AuNP-anti-digoxigenin-POx-HRP
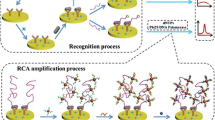
AuNPs as a nanocarrier to synthesize AuNP-anti-digoxigenin-POx-HRP was researched. AuNP-anti-digoxigenin-POx-HRP and anti-digoxigenin-POx were utilized to detect target sequences of concentration from 10−9 to 10−5 M. The Fig. 3a is the diagram of the signal amplification using AuNP-anti-digoxigenin-POx-HRP. As shown in Fig. 3b, when anti-digoxigenin-POx is used, the UV–vis absorption decrease with the target concentration decreasing. The concentration of target sequences is below 10−8 M, the UV absorption value do not change. When AuNP-anti-digoxigenin-POx-HRP is used, the UV–vis absorption reduce significantly with the target concentration from 10−5 to 10−9 M. When the RCA products initiated by 10−5 M target sequences are detected using anti-digoxigenin-POx or AuNP-anti-digoxigenin-POx-HRP, the UV–vis absorption of anti-digoxigenin-POx is 1.890 ± 0.077 (n = 3); while, the UV–vis absorption of AuNP-anti-digoxigenin-POx-HRP is 3.981 ± 0.093 (n = 3), being 2.11 times as much as anti-digoxigenin-POx. Therefore, the AuNP-anti-digoxigenin-POx-HRP improve the amplification signal significantly.
Effects of signal amplification effect of AuNP-anti-digoxigenin-POx-HRP. a The schematic diagram of the signal amplification using AuNP-anti-digoxigenin-POx-HRP. b The UV–vis absorption value of using anti-digoxigenin-POx and AuNP-anti-digoxigenin-POx-HRP to detect different concentrations of target sequences.(n = 3)
Optimization of method
The following parameters were optimized: (a) amount of linear padlock probes; (b) hybridization temperature between linear padlock probes and target sequences; (c) the time of the RCA; (d) the time of RCA products bonding with streptavidin functional 96-well plate. Respective text, data and Figures are given in the Electronic Supporting Material.
So, the following experimental conditions were found to give best results: (a) amount of linear padlock probes: 1 μM linear padlock probes; (b) hybridization temperature between linear padlock probes and target sequences: 50 °C; (c) the time of the RCA: 90 min; (d) the time of RCA products bonding with streptavidin functional 96-well plate: 30 min.
Sensitivity of the RCA-based colorimetric method
The sensitivity of the RCA-based colorimetric method was evaluated using optimization conditions. Figure 4 shows that there is a good linear relationship between UV–vis absorption value and the logarithm of the target sequences concentration from 10−9 to 10−6 M with a correlation coefficient of 0.99344. The limits of detection (3S/N) of the RCA-based colorimetric method is 1.2 × 10−12 M of target sequences. These products have been analyzed by 2% agarose gel electrophoresis, the result of agarose gel electrophoresis is agreed with that of these findings (Fig. S7).
The relationship between the fluorescence intensity (recorded at 450 nm) and target sequences concentrations.Inset:The standard curve of this method upon addition of 10−9 to 10−6 M target sequences (n = 3). Photographs of the 96-well ELISA plate corresponding to different concentration of target sequences
To evaluate the sensitivity of the method in bacterial samples, the different concentrations of bacterial genomic DNA was analyzed using the proposed RCA-based colorimetric method. Microbial genomic DNA was isolated by TIANamp Bacteria DNA Kit. The concentration of microbial genomic DNA was determined according to the manufacturer’s instructions. The UV–vis absorption values of different concentrations bacterial genomic DNA are shown in Fig. 5. There is a good linear relationship between the UV–vis absorption value and the logarithm of the microbe genomic DNA concentration from 1 × 10−2 to 1 ng μL−1 with a correlation coefficient of 0.99394. The limits of detection (3S/N) of this assay is 7.36 × 10−3 ng μL−1 for the microbe genomic DNA. For verify these results, these products were analyzed by 2% agarose gel electrophoresis. As showed in Fig. S8, the electrophoretic results correspond to that of the UV–vis absorption.
The relationship between the fluorescence intensity (recorded at 450 nm) and genomic DNA concentrations.Inset:The standard curve of this method upon addition of 1 × 10−2 to 1 ng μL−1 M target sequences (n = 3). Photographs of the 96-well ELISA plate corresponding to different concentration of genomic DNA
The sensitivity of the method was compared with that of other similar methods (Table 1). The sensitivity of this method is more or equal to the documents, and the method has obvious advantages in terms of high throughput, sensitivity, costs and speed.
Specificity
Bacterial genomic DNA, including E. coli O157:H7, E. coli, S. aureus, S. epidermidis, S. saprophyticus, Shigella soonei, L. monocytogenes, P. aeruginosa, were used to evaluate the specificity of this method. Each microbial genomic DNA was tested by this method, the results were reported through the UV–vis absorption. Subsequently, the results were validated by 2% agarose gel electrophoresis. As Fig. 6 shows that the UV–vis absorption produced by S. aureus and the mixed are 1.439 ± 0.056, 1.213 ± 0.062 (n = 3) respectively, which is significantly higher than that of other bacterial DNA samples (p < 0.01). These results are in accord with that of agarose gel electrophoresis analysis (Fig.S9). These can be concluded that the RCA-based colorimetric method provided a specificity and effective means to detect S. aureus.
Analysis artificial contaminated food samples
To confirm the practicability of this method for the detection of bacterial genomic DNA in food samples, artificial contaminated milk samples and chicken breast samples were analyzed using this colorimetric method. As shown in Table S2, the differences among the contaminated samples and the negative controls can be distinguished by the UV–vis absorption values. The UV–vis absorption values of artificial contaminated milk and chicken breast samples represent 1.563 ± 0.068 and 1.442 ± 0.044 (n = 3), separately. The above results can be significantly distinguished from the negative controls (0.529 ± 0.041 and 0.460 ± 0.029 (n = 3), p < 0.01). These results are verified by agarose gel electrophoresis (Fig.S9).
Conclusions
A colorimetric microplate assay for detecting Staphylococcus aureus DNA was established on streptavidin-functionalized 96-well plates using rolling circle amplification (RCA) as a molecular detection tool and AuNP-anti-digoxigenin-POx-HRP as an efficient signal transduction markers. In the method, RCA offered high sensitivity and specific recognition for the target DNA of Staphylococcus aureus. The RCA products were very long reduplicated single-strand DNA sequences, which can hybridize with AuNP-anti-digoxigenin-POx-HRP to output amplified colorimetric signals. Under the optimum parameters, the detection limit of the colorimetric microplate assay for detecting target sequences was 1.2 pM and detecting bacterial genomic DNA was 7.36 pg μL−1. Using the method to detect artificial contaminated food samples, the potential application of the proposed RCA-based colorimetric method was verified. The method is a user-friendly diagnostic tool for bacterial pathogens. Future research can design more much individual species-specific padlock probes to improve the multiplex detection capabilities of the RCA-based colorimetric method. This method may be also expanded for other bacteria and viruses.
References
Kadariya J, Smith TC, Thapaliya D (2014) Staphylococcus aureus and staphylococcal food-borne disease: an ongoing challenge in public health. Biomed Res Int 2014:9. https://doi.org/10.1155/2014/827965
Wei C, Zhong J, Hu T, Zhao X (2018) Simultaneous detection of Escherichia coli O157:H7, Staphylococcus aureus and Salmonella by multiplex PCR in milk. 3. Biotech 8(1):76. https://doi.org/10.1007/s13205-018-1086-5
Zhao X, Zhao F, Wang J, Zhong N (2017) Biofilm formation and control strategies of foodborne pathogens: food safety perspectives. RSC Adv 7(58):36670–36683. https://doi.org/10.1039/C7RA02497E
Zhao X, Wei C, Zhong J, Jin S (2016) Research advance in rapid detection of foodborne Staphylococcus aureus. Biotechnol Biotechnol Equip 30(5):827–833. https://doi.org/10.1080/13102818.2016.1209433
Gill AAS, Singh S, Thapliyal N, Karpoormath R (2019) Nanomaterial-based optical and electrochemical techniques for detection of methicillin-resistant Staphylococcus aureus: a review. Microchim Acta 186(2):114. https://doi.org/10.1007/s00604-018-3186-7
Gomez A, Miller NS, Smolina I (2014) Visual detection of bacterial pathogens via PNA-based padlock probe assembly and isothermal amplification of DNAzymes. Anal Chem 86(24):11992–11998. https://doi.org/10.1021/ac5018748
Mohsen MG, Kool ET (2016) The discovery of rolling circle amplification and rolling circle transcription. Acc Chem Res 49(11):2540–2550. https://doi.org/10.1021/acs.accounts.6b00417
Dar SA, Kuenen JG, Muyzer G (2005) Nested PCR-denaturing gradient gel electrophoresis approach to determine the diversity of sulfate-reducing bacteria in complex microbial communities. Appl Environ Microbiol 71(5):2325–2330. https://doi.org/10.1128/AEM.71.5.2325-2330.2005
Fang TH, Ramalingam N, Xian-Dui D, Ngin TS, Xianting Z, Lai Kuan AT, Peng Huat EY, Hai-Qing G (2009) Real-time PCR microfluidic devices with concurrent electrochemical detection. Biosens Bioelectron 24(7):2131–2136. https://doi.org/10.1016/j.bios.2008.11.009
Cook N (2003) The use of NASBA for the detection of microbial pathogens in food and environmental samples. J Microbiol Methods 53(2):165–174. https://doi.org/10.1016/S0167-7012(03)00022-8
Sun J, Najafzadeh M, Zhang J, Vicente V, Xi L, Hoog S (2011) Molecular identification of Penicillium marneffei using rolling circle amplification. Mycoses 54:e751–e759. https://doi.org/10.1111/j.1439-0507.2011.02017.x
Xu J, Guo J, Maina SW, Yang Y, Hu Y, Li X, Qiu J, Xin Z (2018) An aptasensor for staphylococcus aureus based on nicking enzyme amplification reaction and rolling circle amplification. AnBio 549:136–142. https://doi.org/10.1016/j.ab.2018.03.013
Zhan Z, Li H, Liu J, Xie G, Xiao F, Wu X, Aguilar ZP, Xu H (2020) A competitive enzyme linked aptasensor with rolling circle amplification (ELARCA) assay for colorimetric detection of listeria monocytogenes. Food Control 107:106806. https://doi.org/10.1016/j.foodcont.2019.106806
Shi D, Huang J, Chuai Z, Chen D, Zhu X, Wang H, Peng J, Wu H, Huang Q, Fu W (2014) Isothermal and rapid detection of pathogenic microorganisms using a nano-rolling circle amplification-surface plasmon resonance biosensor. Biosens Bioelectron 62:280–287. https://doi.org/10.1016/j.bios.2014.06.066
Najafzadeh MJ, Sun J, Vicente VA, Hoog GSD (2011) Rapid identification of fungal pathogens by rolling circle amplification using Fonsecaea as a model. Mycoses 54(5):e577. https://doi.org/10.1111/j.1439-0507.2010.01995.x
Russell C, Welch K, Jarvius J, Cai Y, Brucas R, Nikolajeff F, Svedlindh P, Nilsson M (2014) Gold nanowire based electrical DNA detection using rolling circle amplification. ACS Nano 8(2):1147–1153. https://doi.org/10.1021/nn4058825
Huang R, He L, Xia Y, Xu H, Liu C, Xie H, Wang S, Peng L, Liu Y, Liu Y, He N, Li Z (2019) A sensitive Aptasensor based on a Hemin/G-Quadruplex-assisted signal amplification strategy for electrochemical detection of gastric Cancer Exosomes. Small 15(19):1900735. https://doi.org/10.1002/smll.201900735
Teng J, Ye Y, Yao L, Yan C, Cheng K, Xue F, Pan D, Li B, Chen W (2017) Rolling circle amplification based amperometric aptamer/immuno hybrid biosensor for ultrasensitive detection of Vibrio parahaemolyticus. Microchim Acta 184(9):3477–3485. https://doi.org/10.1007/s00604-017-2383-0
Yao L, Ye Y, Teng J, Xue F, Pan D, Li B, Chen W (2017) In vitro isothermal nucleic acid amplification assisted surface-enhanced Raman spectroscopic for ultrasensitive detection of Vibrio parahaemolyticus. Anal Chem 89(18):9775–9780. https://doi.org/10.1021/acs.analchem.7b01717
Tian B, Han Y, Fock J, Strömberg M, Leifer K, Hansen MF (2019) Self-assembled magnetic nanoparticle–Graphene oxide Nanotag for Optomagnetic detection of DNA. ACS Appl Nano Mater 2(3):1683–1690. https://doi.org/10.1021/acsanm.9b00127
Gou D, Xie G, Li Y, Zhang X, Chen H (2018) Voltammetric immunoassay for Mycobacterium tuberculosis secretory protein MPT64 based on a synergistic amplification strategy using rolling circle amplification and a gold electrode modified with graphene oxide, Fe3O4 and Pt nanoparticles. Microchim Acta 185(9):436. https://doi.org/10.1007/s00604-018-2972-6
Yao C, Xiang Y, Deng K, Xia H, Fu W (2013) Sensitive and specific HBV genomic DNA detection using RCA-based QCM biosensor. Sensors Actuators B Chem 181:382–387. https://doi.org/10.1016/j.snb.2013.01.063
Yang X, Yang K, Zhao X, Lin Z, Liu Z, Luo S, Zhang Y, Wang Y, Fu W (2017) Terahertz spectroscopy for the isothermal detection of bacterial DNA by magnetic bead-based rolling circle amplification. Analyst 142:4661–4669. https://doi.org/10.1039/C7AN01438D
Wang J, Li H, Li T, Ling L (2018) Determination of bacterial DNA based on catalytic oxidation of cysteine by G-quadruplex DNAzyme generated from asymmetric PCR: application to the colorimetric detection of Staphylococcus aureus. Microchim Acta 185(9):410. https://doi.org/10.1007/s00604-018-2935-y
Bendayan M (2000) A review of the potential and versatility of colloidal gold cytochemical labeling for molecular morphology. Biotech Histochem 75(5):203–242. https://doi.org/10.3109/10520290009068433
Liu L, Xia N, Liu H, Kang X, Liu X, Xue C, He X (2014) Highly sensitive and label-free electrochemical detection of microRNAs based on triple signal amplification of multifunctional gold nanoparticles, enzymes and redox-cycling reaction. Biosens Bioelectron 53:399–405. https://doi.org/10.1016/j.bios.2013.10.026
Lai W, Zeng Q, Tang J, Zhang M, Tang D (2018) A conventional chemical reaction for use in an unconventional assay: a colorimetric immunoassay for aflatoxin B1 by using enzyme-responsive just-in-time generation of a MnO2 based nanocatalyst. Microchim Acta 185(2):92. https://doi.org/10.1007/s00604-017-2651-z
Ambrosi A, Castañeda MT, Killard AJ, Smyth MR, Alegret S, Merkoçi A (2007) Double-codified gold Nanolabels for enhanced Immunoanalysis. Anal Chem 79(14):5232–5240. https://doi.org/10.1021/ac070357m
Szemes M, Bonants P, de Weerdt M, Baner J, Landegren U, Schoen CD (2005) Diagnostic application of padlock probes—multiplex detection of plant pathogens using universal microarrays. NAR 33(8):e70–e70. https://doi.org/10.1093/nar/gni069
Li J, Chu X, Liu Y, Jiang J-H, He Z, Zhang Z, Shen G, Yu R-Q (2005) A colorimetric method for point mutation detection using high-fidelity DNA ligase. NAR 33:e168. https://doi.org/10.1093/nar/gni163
Hua D, Xu Z, Anil K, Guozang Z, Xiaoning Z, Xing-Jie L (2013) Long genomic DNA amplicons adsorption onto unmodified gold nanoparticles for colorimetric detection of bacillus anthracis. ChCom 49(1):51–53. https://doi.org/10.1039/c2cc37037a
Ning Y, Gao Q, Zhang X, Wei K, Chen L (2016) A Graphene oxide-based sensing platform for the determination of methicillin-resistant Staphylococcus aureus based on Strand-displacement polymerization recycling and synchronous fluorescent signal amplification. J Biomol Screen 21(8):851–857. https://doi.org/10.1177/1087057116653564
Zhang S, Wu Z, Shen G, Yu R (2009) A label-free strategy for SNP detection with high fidelity and sensitivity based on ligation-rolling circle amplification and intercalating of methylene blue. Biosens Bioelectron 24(11):3201–3207. https://doi.org/10.1016/j.bios.2009.03.012
Xiang Y, Zhu X, Huang Q, Zheng J, Fu W (2015) Real-time monitoring of mycobacterium genomic DNA with target-primed rolling circle amplification by a Au nanoparticle-embedded SPR biosensor. Biosens Bioelectron 66:512–519. https://doi.org/10.1016/j.bios.2014.11.021
Xiang Y, Deng K, Xia H, Yao C, Chen Q, Zhang L, Liu Z, Fu W (2013) Isothermal detection of multiple point mutations by a surface plasmon resonance biosensor with Au nanoparticles enhanced surface-anchored rolling circle amplification. Biosens Bioelectron 49:442–449. https://doi.org/10.1016/j.bios.2013.04.044
Acknowledgements
The authors are grateful for providing language help from the American Journal Experts.
Funding
This work was supported by “The National Key R&D Program of China” (No.2016YFD0401202), Special Project of Tianjin Innovation Platform (No.17PTGCCX00230), Tianjin science and technology planning project(No. 18PTSYJC00130).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The author(s) declare that they have no conflict of interest.
Human and Animal Rights
This article does not contain any studies with laboratory animals or human participants performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 1890 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Wang, J., Wang, S. et al. Rolling circle amplification based colorimetric determination of Staphylococcus aureus. Microchim Acta 187, 119 (2020). https://doi.org/10.1007/s00604-019-4082-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-019-4082-5