Abstract
Deoxyribonucleic acid (DNA) can form G-quadruplexes in the presence of certain metal ions. These play a functional role in a variety of biological process. A DNA 17-mer (PW17) was previously reported to bind hemin in the presence of excess potassium ion. The resulting stabilized G-quadruplex-hemin complex exhibits peroxidase-like activity. However, this activity is lost in the presence of one equivalent of Pb2+ ion. We exploit this property in a method for spectrophotometric detection of Pb2+. Inhibition by Pb2+ ion is reflected by a change in the Soret band of hemin and a sharp reduction in the catalytic activity towards hydrogen peroxide-mediated chromogenic oxidation of 2,2′-azino-bis(3-ethylbenzothiazoline)-6-sulfonate. The new method enables Pb2+ to be detected in the concentration range from 0.05 to 1.2 μM, with a detection limit of 27 nM. The assay shows high selectivity over other metal ions. It was successfully used to determine Pb2+ in water samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The development of detection of specific ion is linked to pressing needs for the rapid detection of toxic metals [1–4]. Of particular interest has been the detection of lead(II) (Pb2+) ion, an important pollutant with major routes of human exposure arising from lead-based paints and contaminated soils and foodstuffs [5]. Over past decades, a number of analytical methods for the detection of Pb2+ level have been reported, including those based upon atomic absorption spectrometry [6, 7], atomic emission spectrometry [8, 9], inductively coupled plasma - mass spectrometry [10–12], anodic stripping voltammetry [13–15], and high-performance liquid chromatography [16–18]. However, these methods require sophisticated equipment and/or time-consuming pretreatment steps prior to the analysis, which limit the scope of their practical applications.
To this end, some efforts have been recently directed towards the development of spectrophotometric approaches to detecting Pb2+. Spectrophotometry does not require any special equipment, and can be directly observed by the naked eyes, but it generally has a poor selectivity and sensitivity [19–25]. To circumvent these drawbacks, many new selective recognition elements and sensitive reporting probes have been introduced recently [26–32]. Among them, Pb2+-dependent deoxyribozyme (DNAzyme) as a selective recognition element has particularly attracted much attention, because it is obtained through a combinatorial biology method called in vitro selection [33]. The DNAzyme which applies Pb2+ as a cofactor consists of a substrate strand containing a single RNA base and an enzyme strand. In the presence of the cofactor (Pb2+), the DNAzyme can catalyze the cleavage of a substrate strand at its single RNA base site. Very recently, many researchers have exploited the property of the DNAzyme to specifically detection of Pb2+ with various means [27–30]. In most cases, gold nanoparticles (AuNPs) are tethered to the thio-containing DNAzyme to indicate color change [27, 28]. These labeled approaches do offer some advantages, but they are still cost- and time-consuming. Therefore, developing label-free spectrophotometric approaches to simplify the detection process are very important and attractive. Then, many researchers exploited the property of different adsorption ability of single-stranded deoxyribonucleic acid (ssDNA) and double-stranded deoxyribonucleic acid (dsDNA) on unmodified AuNPs and the phenomenon of salt-induced AuNPs aggregation to detect Pb2+ [29, 30]. In the presence of Pb2+, the DNAzyme obtained by the hybridization process of two oligonucleotides (enzyme and substrate strands) is cleaved, resulting in the release of some short ssDNA fragments which can be adsorbed onto the AuNPs and thus protect the AuNPs against aggregation under high-salt conditions (the solution remains red). In the absence of Pb2+, however, the remaining duplex can not be adsorbed onto the AuNPs and thus can not stabilize the AuNPs under the same conditions (the red-to-blue color change of the solution is observed). Because the cleavage of the DNAzyme in the presence of Pb2+ can be reflected by the color change of AuNPs, spectrophotometric approaches for detection of Pb2+ can be successfully developed. All the label-free spectrophotometry, however, involve the hybridization of two oligonucleotides by annealing of the enzyme and substrate strands. Additionally, the molar ratio between the enzyme strand and the substrate strand in the hybridization process is of great importance for detection of Pb2+ because very small numbers of unhybridized ssDNA can still stabilize AuNPs and increase the background [29].
In this paper, a new strategy for spectrophotometric detection of Pb2+ is described. It employs a unimolecular G-quadruplex peroxidase-like DNAzyme (PW17) reported by Willner’s group [34, 35] both as a selective recognition element and a sensitive reporting probe, and thus eliminates the elaborate design of the PW17. In addition, the analytical method can avoid a laborious hybridization step in order to more facilitate Pb2+ detection. This assay is based on different binding abilities and peroxidase-like DNAzyme activities of the G-quadruplex towards K+ and Pb2+. It was found the PW17 had much higher binding ability towards Pb2+ than towards K+, but different from the excellent hemin-binding affinity and peroxidase-like DNAzyme activity exhibited by the K+-stabilized PW17, the Pb2+-stabilized PW17 exhibited neither hemin-binding affinity nor peroxidase-like DNAzyme activity. Thus, upon addition of Pb2+ to the K+-stabilized hemin-PW17 DNAzyme, the substitution of K+ by Pb2+ in the PW17 G-quadruplex results in the loss of the binding ability towards hemin and the inhibition of DNAzyme activity. These changes can be reflected easily in color change corresponding to the catalyzed hydrogen peroxide (H2O2)-mediated oxidation of ABTS, based upon which a Pb2+ spectrophotometric approach is constructed
Experimental
Reagents and chemicals
The PW17 oligonucleotide purified with polyacrylamide gel electrophoresis (PAGE), hemin, ABTS were purchased from Sangon Biotechnology Co., Ltd. (Shanghai, China, http://www.sangon.com/). Lead (II) acetate was purchased from Tianjin Huadong Chemical Reagent Company (Tianjin, China, http://www.tjhuadong.com/). H2O2 (30% (m/v)) was purchased from Sinopharm Group Chemical Reagent Co., Ltd. (Shanghai, China, http://www.reagent.com.cn/). Before use, the PW17 oligonucleotide was dissolved in 10 mM Tris(hydroxymethyl)aminomethane-acetic acid (Tris-HAc) buffer solution (pH 7.4). The as-prepared solution was heated to 95 °C for 5 min and allowed to cool naturally to room temperature in 2 h to anneal the oligonucleotide. The concentration of the oligonucleotide was determined by measuring the UV absorbance at 260 nm. Unless specified, other reagents and chemicals were of at least analytical reagent grade. The water used throughout all experimentals was purified by a Milli-Q system (Millipore, Bedford, MA, http://www.millipore.com/).
Instrumentation
Kinetic and spectral measurements were performed on a Cary 50 UV-visible spectrophotometer (Varian, USA, http://www.varianinc.com.cn/) using quartz cuvettes with an optical path length of 1 cm at room temperature.
General procedure of spectrophotometric determination of Pb2+
A typical spectrophotometric analysis was realized as following procedure: firstly, 3 μL of 70 μM PW17, an appropriate volume of 50 μM Pb2+ and 150 μL Tris-HAc buffer (50 mM Tris-Ac, pH 8.0, 1.0 mM KAc, 0.1% (w/v) Triton X-100, 2% DMSO) and an appropriate volume of water were micropipetted to give a volume of 296 μL and incubated for 2 h. Secondly, 4.2 μL of 50 μM hemin was added to the solution and allowed to stand for 1 h. Thirdly, 60 μL of water and 20 μL of 50 mM ABTS solution were added to the resulting solution, following by the addition of 20 μL of 50 mM H2O2 to initial the ABTS-H2O2 reaction. After the as-prepared solution was stirred for 30 s, the kinetic-spectral data were collected from 400 nm to 1,000 nm, every 1 nm, at 12 s intervals during 234 s. The 420 nm (the maximal absorption wavelength of ABTS•+) absorbance at 234 s was herein applied for quantitative analysis of Pb2+. And at 234 s, the photographs of the resulting mixtures were taken with Canon IXUS 110 IS digital camera.
Spectroscopic analysis of hemin-PW17 complexes
8.6 μL 70 μM PW17, an appropriate volume of 50 μM Pb2+ and/or 1 mM K+, and 200 μL Tris-HAc buffer (50 mM Tris-Ac, pH 8.0, 0.1% (w/v) Triton X-100, 2% DMSO) and an appropriate volume of water were micorpipetted to give a volume of 388 μL, and incubated for 2 h, allowing PW17 to fold into the G-quadruplex structures. Then, 12 μL of 50 μM hemin was added to the as-prepared solution. The mixture solution was kept at room temperature for 1 h, allowing the PW17 to bind hemin properly. Finally, the Soret band of hemin (centered at 396–405 nm) was recorded by using UV-visible absorption spectroscopy. The proper binding of the G-quadruplex PW17 to hemin could be reflected by the hyperchromicity of the hemin Soret band.
Procedure for the determination of Pb2+ in lake water samples
Water samples were taken from South Lake (Changchun, China) and Jingyue Lake (Changchun, China). All the samples collected were spiked with Pb2+ at different concentration levels, filtered through a 0.22 μm membrane (http://www.dingguobio.com/), then centrifuged for 15 min at 14,000 rpm, and finally analyzed with the general procedure.
Results and discussion
Mechanistic basis for the system
At early stage, Willner and co-workers [34, 35] reported a peroxidase-like DNAzyme, PW17, which possessed extremely high hemin-binding affinity and peroxidase-like DNAzyme activity in the presence of excess K+. Then, this excellent property of PW17 was exploited to visually detect various analytes, such as DNA [36], small molecules [37, 38] and protein [39]. Almost all the research focus on this issue has been related to the design of PW17, with few works on the basic studies (e.g. structure-function relationship study). Very recently, the detailed basic studies on PW17 reported by Dong and co-workers showed that PW17 had typical characteristics of parallel and antiparallel G-quadruplex in the presence of K+ and Pb2+, respectively [40]. Different conformation of PW17 stabilized by both metal ions can be directly reflected in hemin-binding affinities and DNAzyme activities. It was found that K+-stabilized PW17 exhibited very high hemin-binding affinity and DNAzyme activity, while PW17 in the presence of Pb2+ had almost no ability to bind hemin and thus displayed no DNAzyme activity. On the other hand, compared with K+-stabilized PW17, Pb2+-stablized PW17 had a compact structure. This contributed to the unusual high efficiency of Pb2+ at stabilizing PW17. In general, one equiv of Pb2+ sufficed to induce and stablize a stable G-quadruplex, whereas K+ at the millimolar level was required to do so. Therefore, this gave us a inspiration to develop a Pb2+ spectrophotometric approaches based on K+-Pb2+-switched G-quadruplex of unimolecular PW17 both as a sensitive sensing platform and a selective recognition element.
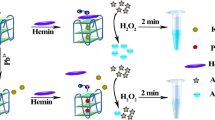
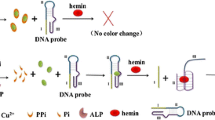
Scheme 1 illustrates the utilization of PW17 for visual detection of Pb2+ in the ABTS-H2O2 reaction system. In the presence of excess K+, PW17 folds into a parallel G-quadruplex that can bind tightly to hemin to form the hemin-G-quadruplex DNAzyme. As a result, it can catalyze the oxidation of colorless ABTS by H2O2 to produce green-colored ABTS•-. After addition of Pb2+, Pb2+ can easily substitute K+ in PW17 in order to form a antiparallel G-quadruplex conformation, and release hemin cofactor. This causes a sharp reduction in the DNAzyme activity accompanied by a less color change in the ABTS- H2O2 system, enabling the detection of aqueous Pb2+ with the naked eye.
The G-quadruplex-based DNAzyme for visual Pb2+ detection. After addition of K+ and hemin, the random conformation of PW17 is folded into the hemin-PW17 DNAzyme which can catalyze H2O2-mediated oxidation of ABTS to produce colored ABTS•-. When adding Pb2+ to such the system, it can replace K+ in hemin-PW17 DNAzyme and release the hemin cofactor, and thus no DNAzyme is formed, reflecting in a reduction in the catalytic activity towards ABTS oxidation by H2O2
Spectrophotometric detection of Pb2+
It is well recognized that the binding of DNA G-quadruplexes to hemin results in an obvious hyperchromicity of the Soret band of hemin [41, 42]. This characteristics was herein employed to investigate the hemin-PW17 interaction in the presence or absence of K+ and/or Pb2+. Fig. 1a shows the spectroscopic analysis of hemin-PW17 complex. The uncomplexed hemin had a Soret absorption band peaked at 396 nm. After incubation with PW17, there was little hyperchromicity in the hemin Soret band, indicating that almost no hemin-PW17 interaction occurred. Then, upon addition of K+, the PW17 was found to cause a large hyperchromicity in the Soret band and a red-shift from 396 to 403 nm, while a hypochromicity in the Soret band was observed upon addition of Pb2+. As previously reported [40], K+ and Pb2+ facilitated the conformation alteration of PW17 from a random coil structure to a parallel or antiparallel G-quadruplex structure, respectively. Thus, it can be assumed that a parallel G-quadruplex structure is beneficial for binding to hemin, but a antiparallel G-quadruplex structure not. Furthermore, when adding Pb2+ to K+-simulated hemin-PW17 complex, the hyperchromicity in the Soret band was clearly reduced, indicating that K+ in the PW17 was replaced by Pb2+, accompanying a conformation change from parallel G-quadruplex to antiparallel G-quadruplex.
Structures and properties of PW17 in Tris-HAc buffer solution (25 mM Tris-HAc, 0.05% (w/v) Triton X-100, 1% DMSO, pH 8.0) under different experimental conditions (a: hemin; b: hemin + PW17 + Pb2+; c: hemin + PW17; d: hemin + PW17 + K+ + Pb2+; e: hemin + PW17 + K+ ). a UV-visible absorption spectra of 1.5 mM complexes of hemin-PW17 complex in the presence or absence of K+ and/or Pb2+. b UV-visible absorption spectra (at 234 s) of the ABTS-H2O2 reaction catalyzed by 0.525 μM complexes of hemin and PW17 in the presence or absence of K+ and/or Pb2+. c Kinetic profiles at the maximal absorption wavelength of ABTS•+ (420 nm) under the experimental conditions in Fig 1B
Similar phenomena were also observed when the hemin-PW17 in the presence or absence of K+ and/or Pb2+ was under investigation in the ABTS-H2O2 system (Fig. 1b and c). The uncomplexed hemin exhibited a very low catalytic activity towards the H2O2-mediated oxidation of ABTS. After incubation with PW17, a little increase was observed in the catalytic activity, and no clear color change could be observed (Fig. S1, see Electronic Supplementary Material). Subsequently, upon addition of K+, there was a sharp increase in the catalytic activity (a pronounced color change from colorless to green could be observed, see Fig. S1), while the absorbance after adding Pb2+ was almost comparable to the background imparted by hemin catalysis (the solution color remained unchanged, see Fig. S1). All these results suggested that the hemin-PW17 DNAzyme was formed in the presence of K+, while not in the presence of Pb2+. Upon addition of Pb2+ to K+-simulated hemin-PW17 DNAzyme, the absorbance changes slowed and the solution remained pale green (Fig. S1). This gave a evidence of Pb2+ inhibition on K+-simulated hemin-PW17 DNAzyme and thus the development of a spectrophotometric approaches for detection of Pb2+ was possible.
Effect of concentrations of K+ on the Pb2+ detection
The inhibition extent of Pb2+ on hemin-PW17 DNAzyme was mainly influenced by the concentrations of K+. Thus, the effect of K+ in the concentrations range of 0.4–1,500 μM on the absorbance of hemin-PW17 DNAzyme in the absence and presence of Pb2+ was herein examined (Fig. 2a and b). The results showed that both in the absence and presence of Pb2+, the absorbance at the maximal absorption wavelength of ABTS•+ (420 nm) increased dramatically with an increased concentration of K+, and reached a plateau when the concentration of K+ exceeded 500 μM (Fig. 2a). Additionally, when the ΔA, where ΔA = Absorbance (K+)—Absorbance (Pb2+ and K+), was used for plotting, a peak value could be obtained at 500 μM (Fig. 2b). Therefore, 500 μM K+ was taken as the optimal concentration of K+.
a Effect of K+ concentrations in the absence (a) and presence (b) of Pb2+ on the hemin-PW17 DNAzyme activity. Experimental conditions: cPW17 = 0.525 μM, chemin = 0.525 μM, cABTS = 2.5 mM, cH2O2 = 2.5 mM, cPb2+ = 1.2 μM. The working buffer buffer (pH 8.0) contains 25 mM Tris-HAc, 0.05% (w/v) Triton X-100, and 1% DMSO. b ΔA versus concentrations of K+ curve for Pb2+ detection where ΔA = Absorbance (K+, 420 nm)—Absorbance (Pb2+ and K+, 420 nm)
Linearity and sensitivity for the Pb2+ detection
To quantitatively detect Pb2+ using the method, the absorbance of the system in the presence of various concentration of Pb2+ was measured at 420 nm as a function of time (Fig. 3a). The 420 nm absorbance at 234 s versus the concentration of Pb2+ (0–3.0 μM) was plotted in Fig. 3b. It was clearly observed that the absorbance increase was highly sensitive to the concentration of Pb2+ over the range of 0.05–1.2 μM and the absorbance increased linearly with the concentration (Fig. 3 Inset). The absorbance could be fitted as the equation of A = 0.68–0.23 c (μM) with the correlation coefficient of 0.998, and the detection limit was determined to be 27 nM (3σ/slope), which compared well with those achieved by the sensitive spectrophotometric analytical methods based on AuNPs [27–30].
a Kinetic absorbance-time curves for Pb2+ with different concentrations (a to h: 0, 0.05, 0.5, 0.7, 1.0, 1.2, 1.5, 3.0 μM). The concentration of K+ is 0.5 mM. Other experimental conditions are as in Fig. 2. b Absorbance at 420 nm versus Pb2+ concentration curve. Inset: derived calibration curve
Interference study
The selectivity of the system was also investigated by testing the response of the spectrophotometric assay to other metal ions, including Pb2+, Hg2+, Zn2+, Cd2+, Mn2+, Fe3+, Fe2+, Co2+, Ni2+, Cu2+, Cr3+, Al3+, Mg2+, Ca2+, Li+, Na+, Ba2+ and some common anions, such as F−, SO 2−4 , NO −3 (Fig. 4). The results demonstrated excellent selectivity over various metal ions and some common anions. There was almost no decrease of the K+-stabilized PW17 catalytic activity in the presence of various metal ions except Pb2+, demonstrating that the assay had good selectivity towards Pb2+.
Selectivity study corresponding to the analysis of Pb2+. Absorbance ratio = Absorbance (metal ion, 420 nm) / Absorbance (Blank, 420 nm).The concentration of all the metal ions was 1.2 μM. Other experimental conditions are as in Fig. 2
Application: determination of Pb2+ in lake water samples
The applications of the method were evaluated for determination of Pb2+ in two lake water samples. The results are listed in Table 1. As can be seen in Table 1, the mean recoveries of such samples were around 100%. The results revealed that the method developed here could be applicable to natural systems.
Conclusion
A label-free spectrophotometric method for detection of Pb2+ with unimolecular G-quadruplex peroxidase-like DNAzyme, PW17, both as a selective recognition element and a sensitive reporter is introduced. It is based upon the inhibitory effect of Pb2+ on K+-stabilized peroxidase-like DNAzyme and the phenomenon of the catalytic H2O2-mediated oxidation of ABTS by the DNAzyme to green-colored product, ABTS•-. This assay could enable Pb2+ to be detected at 27 nM with spectrophotometry in a facile way, with high selectivity against other metal ions. In addition, another great significance of the work lies in providing a convenient means to visually detect conformation change of G-quadruplex.
References
Miyawaki A, Liopis J, Heim R, McCaffery JM, Adams JA, Ikura M, Tsien RY (1997) Nature 388:882–887
Nolan EM, Lippard SJ (2003) J Am Chem Soc 125:14270–14271
Turner APF (2000) Science 290:1315–1317
Walkup GK, Imperiali B (1996) J Am Chem Soc 118:3053–3054
Rodriguez BB, Bolbot JA, Tothill IE (2004) Biosens Bioelectron 19:1157–1167
Baralkiewicz D, Kozka M, Gramowska H, Barbara TB, Wasinkiewicz K (2004) Int J Environ Anal Chem 84:901–908
Berkkan A, Ertas N (2004) Talanta 64:423–427
Ganjali MR, Babaei LH, Badaei LH, Ziarani GM, Tarlani A (2004) Anal Sci 20:725–729
Zhang JS, Li LH, Zhang JP, Jin QH (2004) Chem J Chinese U 25:1248–1250
Hsieh HF, Chang WS, Hsieh YK, Wang CF (2009) Talanta 79:183–188
Petrov PK, Wibetoe G, Tsalev DL (2006) Spectrochim Acta Part B 61:50–57
Resano M, Marzo P, Perez-Arantegui J, Aramendia M, Cloquet C, Vanhaecke F (2008) J Anal At Spectrom 23:1182–1191
Goldcamp MJ, Underwood MN, Cloud JL, Harshman S, Ashley K (2008) J Chem Educ 85:976–979
Rievaj M, Tomcik P, Cernanska M, Janosikova Z, Bustin D (2008) Chem Anal 53:717–723
Rodriguez JA, Ibarra IS, Galan-Vidal CA, Vega M, Barrado E (2009) Electroanalysis 21:452–458
Ebdon L, Hill SJ, Rivas C (1998) Spectrochim Acta Part B 53:289–297
Pan YH, Liu XS, He XQ, Wang CH (2005) Chin J Anal Chem 33:1560–1564
Szpunar J, Pellerin P, Makarov A, Doco T, Williams P, Medina B, Lobinski R (1998) J Anal At Spectrom 13:749–754
Dai SJ, Zhang XS, Yu LY, Yang YJ (2010) Spectrochim Acta Part B 75:330–333
Aracama NZ, Araujo AN, Perez-Olmos R (2004) Anal Sci 20:679–682
Chen YY, Chang HT, Shiang YC, Hung YL, Chiang CK, Huang CC (2009) Anal Chem 81:9433–9439
Fatoki OS (1987) Environ Int 13:369–373
Lau KT, McHugh E, Baldwin S, Diamond D (2006) Anal Chim Acta 569:221–226
Ruengsitagoon W, Chisvert A, Liawruanggrath S (2004) Talanta 62:709–713
Xue H, Tang XJ, Wu LZ, Zhang LP, Tung CH (2005) J Org Chem 70:9727–9734
Huang KW, Yu CJ, Tseng WL (2010) Biosens Bioelectron 25:984–989
Liu JW, Lu Y (2004) J Am Chem Soc 126:12298–122305
Liu JW, Lu Y (2004) Chem Mater 16:3231–3238
Wang ZD, Lee JH, Lu Y (2008) Adv Mater 20:3263–3267
Wei H, Li BL, Li J, Dong SJ, Wang EK (2008) Nanotechnology 19:095501
Asano T, Yabusaki K, Wang PC, Lwasaki A (2010) Spectrochim Acta Part A 75:819–824
Ranyuk E, Douaihy CM, Bessmertnykh A, Denat F, Averin A, Beletskaya I, Guilard R (2009) Org Lett 11:987–990
Breaker RR, Joyce GF (1994) Chem Biol 1:223–229
Niazov T, Pavlov V, Xiao Y, Gill R, Willner I (2004) Nano Lett 4:1683–1687
Pavlov V, Xiao Y, Gill R, Dishon A, Kotler M, Willner I (2004) Anal Chem 76:2152–2156
Cheglakov Z, Weizmann Y, Beissenhirtz MK, Willner I (2006) Chem Commun 3205-3207
Elbaz J, Shlyahovsky B, Li D, Willner I (2008) ChemBiochem 9:232–239
Li D, Shlyahovsky B, Elbaz J, Willner I (2007) J Am Chem Soc 129:5804–5805
Shlyahovsky B, Li D, Katz E, Willner I (2007) Biosens Bioelectron 22:2570–2576
Li T, Wang EK, Dong SJ (2009) J Am Chem Soc 131:15082–15083
Travascio P, Li YF, Sen D (1998) Chem Biol 5:505–517
Travascio P, Bennet AJ, Wang DY, Sen D (1999) Chem Biol 6:779–787
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 20890022), the National Key Basic Research Development Project of China (No. 2010CB933602) and the Project of Chinese Academy of Sciences (No.KJCX2-YW-H09).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary materials
Below is the link to the electronic supplementary material.
Fig. S1
(DOC 21861 kb)
Rights and permissions
About this article
Cite this article
Wang, Y., Wang, J., Yang, F. et al. Spectrophotometric detection of lead(II) ion using unimolecular peroxidase-like deoxyribozyme. Microchim Acta 171, 195–201 (2010). https://doi.org/10.1007/s00604-010-0418-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-010-0418-x