Abstract
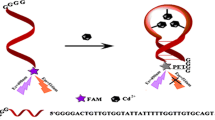
We rationally designed an ultrasensitive and label-free sensing platform for determination of cadmium (Cd). The sensing platform contains G-quadruplex-Cd(II) specific aptamer (GCDSA) constructed by incorporating G-rich sequence at the end of 5' and the critical domain of the Cd-4 aptamer. GCDSA designed act as both a special recognition sequence for Cd2+ and a signal DNAzyme. In absence of Cd2+, GCDSA may mainly exist in a random coil sequence. Upon addition of Cd2+, GCDSA could probably be induced to fold into a G-quadruplex structure. The generation of plentiful active G-quadruplex interacts with hemin to form a peroxidase-like DNAzyme, leading to increased absorbance signal of the sensing system. A4 was directly proportional to the two segments of concentrations for Cd2+, with the detection of limit of 0.15 nM. The proposed method avoids the labeled oligonucleotides and allows directly quantitative analysis of the samples by cheap instruments, with an excellent dynamic range.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Merchant Research & Consulting Ltd, “Cadimum: Global Market Trends and Prospects to 2027”, 2014, Market Publishers Ltd., London
W. Tang, W. Zhang, Y. Zhao, H. Zhang, and B. Shan, Ecol. Indic., 2017, 81, 295.
X. M. Zhao, L. A. Yao, Q. L. Ma, G. J. Zhou, L. Wang, Q. L. Fang, and Z. C. Xu, Chemosphere, 2018, 194, 107.
W. Ji, Z. Chen, D. Li, and W. Ni, Energy Procedia, 2012, 16, 27.
J. Yan, G. Quan, and C. Ding, Procedia Environ. Sci., 2013, 18, 78.
L. Shi, H. Cao, J. Luo, P. Liu, T. Wang, G. Hu, and C. Zhang, Ecotoxicol. Environ. Saf., 2017, 145, 24.
L. Barregard, G. Bergström, and B. Fagerberg, Environ. Res., 2014, 135, 311.
M. Mikowska and R. S´wiergosz-Kowalewska, Chemosphere, 2018, 199, 625.
X. Zeng, X. Xu, H. M. Boezen, J. M. Vonk, W. Wu, and X. Huo, Environ. Pollut., 2017, 230, 838.
W. M. L. Shan, Y. B. Yi, Z. Y. Zhou, and F. Le, PLoS One, 2014, 9, 1.
X. A. Yang, M. B. Chi, Q. Q. Wang, and W. B. Zhang, Anal Chim. Acta, 2015, 869, 11.
H. Yu, X. Ai, K. Xu, C. Zheng, and X. Hou, Analyst, 2016, 141, 1512.
R. Sun, G. Ma, X. Duan, and J. Sun, Spectrochim. Acta, Part B, 2018, 141, 22.
X. Yu, B. Chen, M. He, H. Wang, and B. Hu, Talanta, 2018, 179, 279.
R. S. Amais, A. Virgilio, D. Schiavo, and J. A. Nóbrega, Microchem. J., 2015, 120, 64.
G. Peng, Y. Lu, Q. He, D. Mmereki, X. Tang, Z. Zhong, and X. Zhao, Water Sci. Technol., 2016, 73, 2781.
A. T. Duarte, M. B. Dessuy, M. G. R. Vale, B. Welz, and J. B. de Andrade, Talanta, 2013, 115, 55.
H. Tinas, N. Ozbek, and S. Akman, Microchem. J., 2018, 138, 316.
V. Devesa and D. Vélez, in “Encyclopedia of Food and Health”, 2016, Academic Press, Oxford, DOI: https://doi.org/10.1016/B978-0-12-384947-2.00096-9, 543.
J. Huang, X. Su, and Z. Li, Biosens. Bioelectron., 2017, 96, 127.
L. Mo, J. Li, Q. Liu, L. Qiu, and W. Tan, Biosens. Bioelectron., 2017, 89, 20.
I. Palchetti and F. Bettazzi, in “Reference Module in Chemistry”, 2017, Molecular Sciences and Chemical Engineering, Elsevier, DOI: https://doi.org/10.1016/B978-0-12-409547-0.
L. Mo, J. Li, Q. Liu, L. Qiu, and W. Tan, Biosens. Bioelectron., 2017, 89, 201.
J. Huang, X. Su, and Z. Li, Biosens. Bioelectron., 2017, 96, 127.
Y. Wu, S. Zhan, L. Wang, and P. Zhou, Analyst, 2014, 139, 1550.
H. R. Lotfi Zadeh Zhad, Y. M. Rodríguez Torres, and R. Y. Lai, J. Electroanal. Chem., 2017, 803, 89.
C. K. Kwok and C. J. Merrick, Trends Biotechnol, 2017, 35, 997.
T. Fujii, P. Podbevšek, J. Plavec, and N. Sugimoto, J. Inorg Biochem., 2017, 166, 190.
D. L. Ma, H. Z. He, K. H. Leung, H. J. Zhong, D. S. Chan, and C. H. Leung, Chem. Soc. Rev., 2013, 42, 3427.
J. Kosman, A. Stanislawska, A. Gluszynska, and B. Juskowiak, Int. J. Biol. Macromol., 2017, 101, 799.
Y. Wang, Y. Wu, W. Liu, L. Chu, Z. Liao, W. Guo, G.-Q. Liu, X. He, and K. Wang, Talanta, 2018, 178, 491.
H. Sun, L. Yu, H. Chen, J. Xiang, X. Zhang, Y. Shi, Q. Yang, A. Guan, Q. Li, and Y. Tang, Talanta, 2015, 136, 210.
J. Ge, X. P. Li, J. H. Jiang, and R. Q. Yu, Talanta, 2014, 122, 85.
M. Liu, Z. Wang, L. Pan, Y. Cui, and Y. Liu, Biosens. Bioelectron., 2015, 69, 142.
Y. Y. Yan, J. Lin, T. M. Ou, J. H. Tan, D. Li, L. Q. Gu, and Z. S. Huang, Biochem. Biophys. Res. Commum., 2010, 402, 614.
A. C. Fyfe, P. W. Dunten, M. M. Martick, and W. G. Scott, J. Mol. Biol., 2015, 427, 2205.
I. M. Pedroso, L. F. Duarte, G. Yanez, A. M. Baker, and T. M. Fletcher, Biochem Biophys. Res, Commun., 2007, 358, 298.
X. Tang, Y. S. Wang, J. H. Xue, B. Zhou, J. X. Cao, S. H. Chen, M. H. Li, X. F. Wang, Y. F. Zhu, and Y. Q. Huang, J. Pharm. Biomed. Anal., 2015, 107, 258.
Y. Ding, S. Wang, J. Li, and L. Chen, TrAC, Trends Anal. Chem., 2016, 82, 175.
Z. Zhang, H. Wang, Z. Chen, X. Wang, J. Choo, and L. Chen, Biosens. Bioelectron., 2018, 114, 52.
L. Chen and J. Li, ACS Appl. Mater. Interfaces, 2014, 6, 15897.
L. Chen, X. Fu, W. Lu, and L. Chen, ACS Appl. Mater. Interfaces, 2013, 5, 284.
T. Lou, Z. Chen, Y. Wang, and L. Chen, ACS Appl. Mater. Interfaces, 2011, 3, 1568.
Y. F. Zhu, Y. S. Wang, B. Zhou, J. H. Yu, L. L. Peng, Y. Q. Huang, X. J. Li, S. H. Chen, X. Tang and X. F. Wang, Anal. Bioanal. Chem., 2017, 409, 4951.
K. K. Chun and J. M. Catherine, Trends Biotechnol., 2017, 35, 997.
K. Nucleic, A. Lawrence, and S. B. Paramjeet, Nucleic Acids Res., 2006, 34, W676.
M. I. Zarudnaya, I. M. Kolomiets, A. L. Potyahaylo, and D. M. Hovorun, Nucleic Acids Res., 2003, 31, 1375.
B. I. Kankia, G. Barany, and K. Musier-Forsyth, Nucleic Acids Res., 2005, 33, 4395.
H. Sun, X. Li, Y. Li, L. Fan, and H. B. Kraatz, Analyst, 2013, 138, 856.
X. H. Zhou, D. M. Kong, and H. X. Shen, Anal. Chim. Acta, 2010, 678, 124.
Acknowledgments
This work is subsidized by the National Natural Science Foundation of China (No. 81502850), the Natural Science Foundation of Hunan Province in China (No. 2015JJ2122) and Undergraduate inquiry learning and innovative experimental project in Hunan province (No. 336).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhou, B., Chen, YT., Yang, XY. et al. An Ultrasensitive Colorimetric Strategy for Detection of Cadmium Based on the Peroxidase-like Activity of G-Quadruplex-Cd(II) Specific Aptamer. ANAL. SCI. 35, 277–282 (2019). https://doi.org/10.2116/analsci.18P248
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2116/analsci.18P248