Abstract
During the course of life, muscle mass undergoes many changes in terms of quantity and quality. Skeletal muscle is a dynamic tissue able to hypertrophy or atrophy according to growth, ageing, physical activity, nutrition and health state. The purpose of the present review is to present the mechanisms by which exercise can induce changes in human skeletal muscle mass by modulating protein balance and regulating the fate of satellite cells. Exercise is known to exert transcriptional, translational and post-translational regulations as well as to induce epigenetic modifications and to control messenger RNA stability, which all contribute to the regulation of protein synthesis. Exercise also regulates the autophagy–lysosomal and the ubiquitin–proteasome pathways, the two main proteolytic systems in skeletal muscle, indicating that exercise participates to the regulation of the quality control mechanisms of cellular components and, therefore, to muscle health. Finally, activation, proliferation and differentiation of satellite cells can be enhanced by exercise to induce muscle remodelling and hypertrophy. Each of these mechanisms can potentially impact skeletal muscle mass, depending on the intensity, duration and frequency with which the signal appears.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During the course of life, muscle mass undergoes many changes in terms of quantity and quality. Skeletal muscle is a dynamic tissue able to hypertrophy and to atrophy according to the age, the physical activity, the nutritional state and potentially several diseases. Muscle mass accretion or loss results from net protein balance. If protein synthesis exceeds protein degradation, proteins will accumulate and muscle mass will increase. Inversely, if protein synthesis is less than degradation, loss of muscle mass will occur. In addition, satellite cell inclusion may contribute to increased muscle mass, while fibre loss results in a reduction of muscle mass. The purpose of the present review is to present the mechanisms by which exercise can exert changes in skeletal muscle mass of healthy adult humans. Undoubtedly, nutrition, and more particularly amino acids, plays an essential role in the regulation of protein balance and satellite cells by exercise. We purposely chose not to focus on this aspect here, and we refer the readers to recently published reviews [34, 97, 103, 137, 142]. Of note, the term “exercise” represents one single session of physical exercises while “exercise training” or “training” represents repeated sessions to form a whole training program. Finally, we will limit the report of animal findings to the strict minimum to favour the presentation of data acquired in human.
Effect of exercise on protein balance
Resistance exercise
Most studies on the response to exercise of human muscle protein turnover have focused on changes in the hours following resistance exercise, likely to result in muscle hypertrophy. After heavy resistance exercise, mixed muscle protein synthesis is increased for up to 48 h [119]. Similarly, after strenuous exercise, myofibrillar protein synthesis peaked at 24 h and remained elevated at 72 h [100]. Contrary to the systemic response of feeding on all skeletal muscles, physical activity only stimulates a response in the exercised muscles [99]. Over the same time period after exercise, the muscle extracellular matrix and tendon increased protein synthesis in a similar pattern [100] although probably at a slower rate [62]. The magnitude of the increase in protein synthesis in skeletal muscle after resistance exercise depends upon the intensity and the total workload. Below 40% of the one-repetition maximum (1-RM), there is no detectable increase in protein synthesis while the latter is increased by two- to threefold at intensities above 60% 1-RM [83]. Those results do not imply that intensities below 40% 1-RM cannot elicit anabolic responses. Indeed, in addition to the intensity, muscle fatigue is important to take into consideration. Increases in muscle protein synthesis at 30% 1-RM have been found to be comparable to increases at 90% 1-RM when exercise is performed to failure, not when work is matched between 30 and 90% 1-RM [20]. Practically, increasing the total workload can overcome the lack of anabolic response generally observed at a lower intensity, probably due to a higher recruitment of type II fibres following the fatiguing nature of the contractions [20]. We can therefore easily understand why repeating contractions during endurance exercise for minutes or hours, even if the intensity is rather low, can induce an anabolic response as presented in the next section. In terms of contraction mode, eccentric resistance exercise training, i.e. lengthening contractions, was found to be more potent to induce muscle hypertrophy than concentric training [127]. This effect seems to be due to the higher external loading, which is an intrinsic feature of eccentric contractions, rather than the contraction mode per se. Indeed, when total work was matched between eccentric and concentric contractions, no difference in muscle hypertrophy was observed after training [102].
In skeletal muscle, the range of the increase in protein synthesis in response to exercise (+ 80–100%) is surprising, because the rate of net accretion of muscle protein is much lower. It may take 20 weeks of intense resistance exercise to increase muscle mass by 20%. This is, of course, due to a concomitantly elevated protein degradation after acute exercise if nutrient intake is insufficient [119]. Protein ingestion after exercise increases skeletal muscle protein synthesis and net balance to a greater extent than exercise in the fasted state [149]. Measurements of protein synthesis and degradation after a 12-h fast indicate that degradation exceeds synthesis, resulting in a negative protein balance [163]. In the recovery period after exercise without nutrient provision, protein synthesis and protein degradation are increased compared with the 12-h fasted state, although net balance does not improve to a positive balance. When receiving an infusion of mixed amino acids after a fasted period, protein synthesis increases, whereas protein degradation remains the same or decreases slightly and net protein balance becomes positive. When exercise is combined with amino acids, protein synthesis increases more than after exercise or amino acid feeding alone, and protein degradation remains similar to exercise without feeding. Net protein balance is enhanced in comparison with amino acid feeding alone. For the latest perspectives regarding protein ingestion after resistance exercise training to increase muscle hypertrophy, the reader is referred to a very recent review of the literature [142]. The increase in protein synthesis after feeding is a transient storage phenomenon, whereas physical activity stimulates a longer-term adaptive response. Providing nutrition after physical activity takes advantage of the anabolic signalling pathways that physical activity has initiated by providing amino acid building blocks and energy for protein synthesis [99]. The molecular mechanisms by which exercise regulates protein synthesis and protein degradation will be presented after the section on endurance exercise.
Endurance exercise
Compared to resistance exercise, endurance exercise is characterised by a lower intensity and a longer duration from about 30 min to several hours. In contrast to resistance training for which muscle hypertrophy is the predominant adaptation, endurance training is associated with enhanced endurance capacity through the induction of shifts in substrate metabolism, mitogenesis and angiogenesis [66, 67]. Detailed comparisons between endurance and resistance exercise-induced muscle adaptations can be found in [24, 30, 31]. Compared to resistance training, fewer studies have investigated the links between endurance exercise and protein metabolism. Nonetheless, endurance exercise is also associated with a stimulation of mixed muscle protein synthesis following running, walking or swimming in both men and women [26, 59, 60, 138, 150]. Compared to resistance exercise, the response seems to be somewhat delayed as during the initial 1–1.5 h post-exercise, only minimal increases in muscle protein synthesis are observed, after which protein synthesis increases significantly [90] and can be maintained for up to 24 h, depending on the intensity of the exercise [37]. In contrast to resistance exercise, increased mixed muscle protein synthesis following endurance exercise appears to be predominantly driven by increases in sarcoplasmic and mitochondrial, rather than myofibrillar protein synthesis [38, 161]. Synthesis of mitochondrial proteins is preferentially upregulated in response to endurance exercise, while myofibrillar synthesis is preferentially upregulated in response to resistance exercise, at least in the trained state [161]. Similar to resistance exercise, muscle protein degradation responses to endurance exercise have been less studied. It has been shown that during endurance exercise, muscle protein degradation was increased [4, 26], probably for energetic purposes, by releasing free amino acids [115]. These increases in muscle protein degradation during exercise are maintained post-exercise [26, 138], however, to a lesser extent than during exercise [4]. As mentioned earlier, the next section will focus on the molecular mechanisms by which exercise regulates protein synthesis and protein degradation.
Mechanisms in exercise-induced protein balance
Protein synthesis
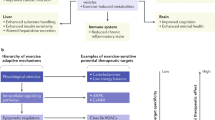
Any step of gene expression can potentially be regulated from transcriptional to translational and post-translational regulations, as well as via chemical and structural modifications of DNA or messenger RNA (mRNA) stability (Fig. 1). The ability of exercise to regulate each of those steps is detailed in the following section.
Transcriptional regulations
Most genes are regulated primarily at the level of transcription rather than translation [17]. This transcriptional regulation is mediated by transcription factors, which generally simultaneously bind DNA and RNA polymerase, as well as other factors necessary for the transcription process. Transcription factors, and their cofactors, can be regulated through reversible post-translational modifications such as phosphorylation or inactivated through mechanisms such as proteolysis. Transcription is initiated at the promoter site as an increase in the amount of an active transcription factor binds a target DNA sequence. Other proteins, known as scaffolding proteins, bind other cofactors and hold them in place. DNA sequences far from the point of initiation, known as enhancers, can aid in the assembly of this transcription machinery. While it would be impossible to establish an exhaustive list of all transcription factors, cofactors and other scaffolding proteins that are regulated by exercise, the induction of the expression of immediate early genes, such as c-Fos or c-Jun, or the expression of activating protein 1 or serum response factor is a crucial primary step in the response to contractile activity [9]. These are themselves transcription factors or components thereof, which can further influence gene expression [166].
DNA modifications
Non-genetic structural modifications of DNA and/or histones resulting in alterations in gene expression are encompassed under the term epigenetics [71]. Those modifications are tightly regulated by three major mechanisms: (1) methylation of the cytosine residues of DNA; (2) chemical modifications of specific residues of histone tails such as acetylation, methylation or phosphorylation; and (3) transcriptional regulation by microRNAs (miRNAs) [114]. The latter epigenetic modification will be developed in a specific section hereafter. The classic DNA covalent modification is methylation of cytosine, which results in the addition of a methyl group by DNA cytosine-5-methyltransferase (DNMT) enzymes [76]. The effect of DNA methylation on gene expression depends on its location within the genome. DNA methylation at the promoter and enhancer regions of genes is associated with transcriptional repression, whereas the un- or hypomethylated state is related to a transcriptionally permissive state [35]. Indeed, promoter DNA methylation changes the conformational layout of chromatin to a more condensed state, more difficult to access by the transcriptional machinery. Conversely, DNA methylation within the gene body seems to be associated with active transcription [75].
Histone post-translational modifications are the second most common epigenetic regulation. The nucleosome is formed by an octamer of histone proteins in which 147 bp of DNA is wrapped around it [74, 143]. Nucleosomes contain two copies of each one of the core histone proteins (H2A, H2B, H3 and H4). In addition, the H1 linker histone stabilises the nucleosome and the linker DNA region between nucleosomes. In the context of the present review, the tail domain of the histones is particularly susceptible to the post-translational modifications induced by resistance or endurance exercise [96] and nutrition [29]. The modifications in histone tails are controlled by histone acetyltransferases, histone deacetylases and histone demethylases, amongst others [74, 143].
Epigenetic modifications, and more particularly methylation of the promoters of peroxisome proliferator-activated receptor-gamma coactivator-1 alpha (PGC-1α), pyruvate dehydrogenase kinase 4 (PDK4) and peroxisome proliferator-activated receptor-delta (PPAR-δ) as well histone acetylation of glucose transporter 4 (GLUT4) and myocyte enhancer factor 2 (MEF2), have been shown to contribute to metabolic adaptations to endurance exercise [11, 95, 114]. In the context of the present review on the control of muscle mass, a very recent report shows that human skeletal muscle possesses an epigenetic memory of hypertrophy [136]. The number of hypomethylated loci across the genome after reloading was twice versus earlier loading. Specific genes, namely AXIN1, glutamate receptor ionotropic kainate 2 (GRIK2), calcium/calmodulin-dependent kinase 4 (CAMK4) and tumor necrosis factor receptor-associated factor 1 (TRAF1), were hypomethylated with enhanced expression after loading, and those genes maintained their hypomethylated status during unloading while muscle mass returned to control levels. Those results suggest a memory of the methylation signature of those genes following earlier hypertrophy [136]. GRIK2, TRAF1, bicaudal C homolog 1 (BICC1) and stromal antigen 1 (STAG1) were epigenetically sensitive to acute exercise as they were hypomethylated after resistance exercise, and this hypomethylation was maintained 22 weeks later with the largest increase in gene expression and muscle mass after reloading [136]. All together, those results indicate an underestimated but important epigenetic role for a large number of genes in muscle hypertrophy and memory.
mRNA stability
Two classes of short RNA molecules, small interfering RNA (siRNA) and miRNA, have been identified as sequence-specific post-transcriptional regulators of gene expression [145]. Pri-microRNA transcripts are first processed into ~ 70-nucleotide pre-miRNA with imperfect stem loop by Drosha inside the nucleus. Pre-miRNAs are transported to the cytoplasm by Exportin 5 and are processed into miRNA duplexes by dicer, a multidomain enzyme of the RNase III family. Dicer also processes long double-stranded molecules into siRNA duplexes. The nascent siRNA and miRNA are double-stranded duplexes. These duplexes need to be unwound before they can be assembled into an RNA-induced silencing complex (RISC). Only one strand (~ 21–25 nucleotides) of the miRNA duplex or the siRNA duplex is preferentially assembled into the RISC, which subsequently acts on its target by translational repression or mRNA cleavage, depending, at least in part, on the level of complementarity between the small RNA and its target [61]. Of note, each tissue can express specific miRNA, called myomiR in muscle [129].
While miRNA may be released from skeletal muscle to the systemic circulation, here, we will focus on the effect of exercise on the regulation of miRNA in skeletal muscle. If it is clear that they are required for muscle development and regeneration, the role of miRNA in muscle maintenance and adaptation during adulthood has not been well characterised up to now [80]. In 2007, it was shown for the first time that miRNA levels could be modulated by changes in mechanical demand [94]. In a mechanical overload mouse model, the plantaris muscle showed a ~ 50% decrease in miR-1 and miR-133 levels [94]. In humans, resistance exercise resulted in a decrease in miR-1 expression in skeletal muscle [39]. As miR-1 targets insulin-like growth factor 1 (IGF-1) and the IGF-1 receptor, it has been suggested that a decrease in miR-1 would potentiate activation of the IGF-1/protein kinase B (PKB) signalling cascade [42]. In addition, there is evidence to suggest that the magnitude of change in miRNA expression following resistance exercise training could predict whether a person would respond well or less well to the exercise [33]. In young adult males, changes in miRNA expression in the vastus lateralis muscle corresponded to differences in muscle hypertrophy after 12 weeks of resistance training [33]. Twenty-one miRNAs were profiled, all showing no significant change in the high-responder group, whereas an increase in miR-451 and a decrease in miR-378 as well as a tendency towards decreased miR-26a and miR-29a were found in the low-responder group. Not only resistance exercise regulates miRNA expression; several studies found that adaptations to endurance exercise were under the control of specific miRNA as well.
The targeted deletion of miR-208b or miR-499 revealed that these two miRNAs were required to establish the slow-twitch fibre phenotype as KO mice for either miRNA resulted in a muscle with significantly more fast-twitch fibres [152]. Consistent with this finding, these knockout mice exhibited reduced exercise capacity when subjected to forced running [152]. Using the same exercise paradigm, Safdar et al. [131] found that treadmill running increased the expression of miR-181, miR-1 and miR-107 and reduced miR-23 expression. These changes in miRNA expression were associated with increased expression of the miR-23 target and PGC-1α, as well as downstream targets of PGC-1α involved in mitochondrial biogenesis, namely aminolevulinic acid synthase, citrate synthase and cytochrome c [131]. In addition, PGC-1α has been found to be targeted by miR-696, another miRNA that is downregulated in response to endurance exercise [6]. Similar results have been found for miR-494, the downregulation of which corresponded to a concomitant increase in gene targets involved in mitochondrial biogenesis [167]. In addition to mitochondrial biogenesis, miRNAs may be involved in regulating adaptations involving oxygen delivery to the muscle via increased capillary density. The levels of miR-16 in the soleus muscle of rats following swim training decreased whereas the expression of vascular endothelial growth factor and its respective type 2 receptor increased [44]. In untrained human participants, acute endurance exercise increased the expression of miR-1 and miR-133 in the quadriceps muscle, while resting levels of miR-1, miR-133a, miR-133b and miR-206 were lower following 12 weeks of training than before training [108]. In addition, the changes observed following the acute exercise pre-training were abolished post-training, suggesting that miRNA levels in response to exercise are sensitive to training status [108]. Similarly, increased miR-1, miR-133a, miR-133-b and miR-181a expression and decreased miR-9, miR-23a, miR-23b and miR-31 expression were found following an acute exercise [130]. After 10 days of training, miR-1 and miR-31 expression was still increased and decreased, respectively, while miR-29b expression was increased, confirming that the training status is part of the regulation of miRNA expression.
Translational and post-translational regulations
The translation rate may be regulated by rapid changes within a few minutes. In this case, it implicates the activity or association of components of the translational machinery, which are primarily mediated by changes in the states of phosphorylation of translation factors and specific RNA-binding proteins. Over the longer term, hours to days, the control of protein synthesis involves changes in the levels of translation factors and ribosomes [121]. The process of translation is divided into three stages: initiation, elongation and termination. Each stage requires translation factors that transiently associate with the ribosome. For a detailed description of the molecular regulations of the translation phase by exercise, with a specific emphasis on the mammalian target of rapamycin (mTOR) pathway, the reader is referred to the following reviews: [2, 8, 45, 55, 70, 155].
Increased ribosomal function seems required to induce net protein synthesis and muscle hypertrophy [10]. Enhanced protein translation rates can be achieved by increased ribosomal efficiency and/or elevated ribosomal capacity via ribosome biogenesis. Both processes are regulated at least in part by mTOR complex 1 (mTORC1) activity. Recent findings suggest that increased mTORC1 activity following compensatory overload hypertrophy in a murine model has a larger impact on translational efficiency than capacity during the first few days following loading [56]. Nevertheless, mTORC1 also plays a central role in ribosome biogenesis by regulating the synthesis of both ribosomal proteins and ribosomal RNA [92]. In humans, until recently, most studies have focused on ribosomal efficiency after acute resistance exercise and long-term resistance training but several lines of evidence now suggest that ribosomal biogenesis may be a key rate-limiting factor in the regulation of resistance training-induced myofibre hypertrophy [10]. Ribosomal RNA represents 80 to 85% of total RNA but is less easy to quantify than total RNA. Therefore, the magnitude of changes in the amount of total RNA is often used as a surrogate of changes in the levels of ribosomal RNA. Over the course of resistance training, total RNA amount appears linked to the magnitude of myofibre [140] and muscle [45] hypertrophy, suggesting that ribosomal RNA amount also varies with changes in muscle growth.
Protein degradation
Studies examining human skeletal muscle protein turnover have focused predominantly on muscle protein synthesis. This is not surprising, considering protein synthetic responses to a variety of stressors in healthy muscle, including nutrition and exercise, are generally more robust and sustained than those related to protein degradation [118, 119]. In skeletal muscle, protein degradation is supported by four major proteolytic systems: the calcium-dependent calpains [169], the autophagy–lysosomal proteases/cathepsins [18], the cysteine protease caspase enzymes and the ubiquitin–proteasome system [43]. Evidence has accumulated that these systems work as partners during muscle proteolysis rather than independently. The control of the degradation rate per se is accomplished in two distinct ways: in the short term, by controlling the enzymatic activities of proteases and/or the access of substrates to these proteases, and in the long term, by controlling, at both transcriptional and translational levels, the synthesis of the proteases and their accessory enzymes [144].
Autophagy is a highly conserved degradation mechanism by which bulk cytoplasmic, long-lived proteins and organelles are degraded by the lysosomal enzymes (cathepsins) [132]. Cathepsins are also capable of degrading myofibrillar proteins such as troponin T, myosin heavy chain or tropomyosin [15]. Autophagy is particularly active in skeletal muscle, where it can be evaluated using specific molecular markers of activation such as unc-51-like kinase 1 (ULK1) phosphorylation and specific proteins, indicating increased autophagosome content, such as total microtubule-associated protein light chain 3 (LC3), LC3-II and LC3-II/LC3-I ratio, or autophagosome degradation, such as p62 [89]. LC3-II, the lipidated form of LC3-I, directly reflects the presence of autophagosomes [12]. LC3-II is recruited at both the inner and outer membranes during vesicle elongation, where it remains bound until autophagosomes fuse with lysosomes [123]. Thus, the LC3-II/LC3-I ratio is recognised as a reliable marker of autophagosome synthesis [81]. p62 binds both to aggregated proteins and to LC3-II and is degraded with the autophagosome content after fusion with lysosome and, as such, helps to determine the amount of autophagosomes discarded by the lysosomes [113].
While studying autophagy in human is not an easy task due to its very dynamic nature, the activation of autophagy in skeletal muscle has been found to occur through AMP-activated kinase (AMPK) and its downstream target ULK1. In human, the regulation of autophagy mainly relies on exercise intensity, less on the nutritional state, which differs somewhat from studies in mice [73, 135]. The autophagy-related and autophagy–regulatory genes are upregulated after an ultraendurance running race [72]. Those results contrast with the reduction in autophagosome synthesis and content, assessed by the ratio LC3-II/LC3-I and LC3-II protein levels, respectively, after endurance exercise of shorter duration (60–120 min) at moderate intensities (50–70% of VO2max) [47, 101, 135]. The effect of resistance exercise on the regulation of autophagy has been much less studied, but it has been shown that the LC3-II/LC3-I ratio and LC3-II protein levels were decreased within the first few hours during the recovery period [48, 52]. It remains to be established whether regular endurance and strength training regulate basal autophagy in humans.
Calcium-dependent μ-calpain, m-calpain and p94 contribute to muscle protein degradation [50], with p94 being the most highly expressed in skeletal muscle [79]. Calpain enzymatic activity is upregulated in response to endurance exercise [50]. The mechanism by which calpains contribute to muscle protein degradation is probably by cleaving myofibrillar proteins into smaller fragments for subsequent degradation by the ubiquitin–proteasome system [54]. Similar to the calpains, after cleaving by caspase-9, caspase-3 becomes activated and initiates muscle proteolysis by degrading myofibrillar proteins into smaller fragments [40]. In addition to cleaving myofibrils, caspase-3 is also able to activate proteasome activity [154], thereby both providing substrate to the ubiquitin–proteasome machinery and increasing ubiquitin–proteasome-mediated protein degradation. Before degradation by the proteasome, cleaved myofibril segments are ubiquitylated by muscle-specific ubiquitin ligases, amongst which atrogin-1 (MAFbx) and muscle RING finger-1 (MuRF-1). Poly-ubiquitylated proteins are subsequently degraded by amino acid hydrolysis within the 26S subunit of the proteasome [51].
Immediately following resistance exercise, MuRF-1 and MAFbx mRNA levels are rapidly augmented, together with their upstream transcription factor forkhead box 1 (FoxO1) [36, 107], with MuRF-1 reaching a peak at 1–2 h post-exercise and returning to baseline levels within 8 h [86]. This molecular regulation seems to be differentially affected by the contraction mode, with increases in MuRF-1 and FoxO1 mRNA levels only seen in response to concentric contractions, not after eccentric contractions [107]. After eccentric contractions, MAFbx mRNA levels were even decreased while the mRNA levels of the structural components of the ubiquitin–proteasome system, namely proteasome subunit α1, ubiquitin splice forms I and II as well as MuRF-2 and MuRF-3, were increased [107]. This difference in molecular regulation between the two contraction modes is likely dependent on the increased levels of damage and remodeling required for eccentric compared to concentric contraction. Following 10 weeks of resistance training, the response of the ubiquitin–proteasome system during the recovery period is attenuated [141], reflecting a similar adaptation of the muscle protein degradation rate after training [120]. This suggests that most of the muscular remodeling due to resistance training occurs in the early phases of the training period. Following moderate endurance exercise, both MuRF1 and MAFbx mRNA levels were increased, suggesting a role for the ubiquitin–proteasomal pathway in regulating post-endurance exercise protein degradation as well [116]. Similar to resistance exercise, MuRF1 mRNA levels after endurance exercise follow a temporal pattern peaking between 1 and 2h, but being maintained for up to 24 h post-exercise, unlike resistance exercise [86]. Contrary to resistance training, the upregulation of the ubiquitin–proteasome system seems to persist after 10 weeks of endurance training [141], highlighting the different regulations of protein metabolism between resistance and endurance exercise.
Effect of exercise on satellite cells
Satellite cells are a population of muscle-derived stem cells responsible for myofibre development and renewal [91]. They are located outside sarcolemma and under the basal lamina of the muscle fibre [25]. Normally, in resting skeletal muscles, satellite cells are generally in a non-proliferative, quiescent state, but they have the ability to re-enter the cell cycle to generate new muscle fibres or to provide new myonuclei during post-natal growth [134]. Activation, proliferation and fusion of this population of cells are required by a myofibre when undergoing myofibrillar protein growth to maintain a constant nuclear/cytoplasmic ratio. Each step is tightly regulated by the myogenic regulatory factors (MRFs), a family of four members (MyoD, Myf-5, myogenin and MRF4, also called Myf-6) [122].
Resistance exercise training has been shown to increase the number of satellite cells after several days [32], weeks [111] or years [77]. This increase can be maintained as long as the muscles are subjected to training. The cessation of training is associated with the termination of satellite cell activation [78]. The activation of satellite cells can be attributed to exercise per se, exercise-induced localised ultrastructural damage, exercise-induced segmental fibre damage, exercise-induced release of inflammatory substances and/or exercise-induced release of growth factors [77]. It seems that the amount of muscle fibre damage is not correlated to the changes in satellite cells following training. The trigger for satellite cell activation could rather be the exercise-induced ultrastructural muscle damage since fibres with more ultrastructural damage also contained higher proportions of active satellite cells [128]. While less studied, we understand from the latter that the activation of satellite cells is not restricted to resistance exercise but that endurance exercise can activate them as well, the critical factors being the nature of contraction, eccentric versus concentric, and the intensity of the session [1]. In contrast to resistance exercise, it seems that the activation of satellite cells in response to endurance exercise leads to muscle adaptations rather than hypertrophy. The key triggers of satellite cell activation by endurance exercise seem to be metabolic factors, such as nitric oxide (NO), nicotinamide dinucleotide (NAD) and sirtuins (SIRTs), as well as oxygen availability according to emerging findings in vitro [1].
Additional mechanisms that could be shared by endurance and resistance exercise contribute to the activation of satellite cells [14]. It has been proposed that hepatocyte growth factor (HGF) activates satellite cells and that IGF-I and fibroblast growth factor (FGF) increase the proliferation of satellite cells once they are activated [5, 146]. The discovery of two IGF-I isoforms in skeletal muscle, mechanical growth factor (MGF) and IGF-IEa, has suggested that MGF initiates satellite cell activation and proliferation, while IGF-IEa promotes differentiation of proliferating satellite cells [168].
In addition to the aforementioned factors, myostatin has direct effects on the proliferation and/or differentiation of numerous muscle cell lines [85, 126]. The effect on muscle fibre number is likely to result from the activity of myostatin on myoblast proliferation and/or differentiation during development, while the effects on fibre size appear to be mediated through the action of myostatin on muscle satellite cells [153]. Myostatin inhibits proliferation through an upregulation of p21 and decreases in the levels of both cyclin-dependent kinase 2 and phosphorylated Rb, resulting in cell cycle inhibition [147]. The effect on differentiation appears to occur through downregulation of the myogenic differentiation factors MyoD, Myf-5 and myogenin [85].
It can be hypothesised that exercise can induce activation of satellite cells without proliferation, proliferation and withdrawal from differentiation, proliferation and differentiation to provide more myonuclei and proliferation and differentiation to generate new fibres or to repair segmental fibre injuries [77]. Although the studies mentioned above tend to favour the idea that satellite cell addition participates to muscle hypertrophy following exercise, it seems that skeletal muscle is capable of hypertrophy via a mere increase in protein synthesis with no additional satellite cell incorporation [93]. In accordance with the latter view, some studies have shown that contraction-induced skeletal muscle growth occurs with no change in total DNA per muscle [164, 165] and that inhibition of satellite cell proliferation does not prevent muscle growth [87]. The requirement of satellite cell activation for skeletal muscle hypertrophy could be linked to the type of growth stimulus, the magnitude of the growth response, the age of the subjects and the time of sampling after the applied stimulus. Muscle growth consists of multiple phases, including accelerated transcriptional and translational responses followed by possible satellite cell addition during the later stages of hypertrophy. It is likely that satellite cell activation is necessary only if a certain threshold of myofibre size is reached [109].
Initiating signals leading to muscle hypertrophy
The main cellular events leading to muscle hypertrophy are summarised in Fig. 2. Via receptor binding and cellular signals, certain cytokines (others are obviously involved in muscle degradation), hormones and growth factors are sensed and activate a network of signal transduction pathways that result in the nuclear translocation or activation of transcription factors [124]. Active transcription factors change the expression of the major muscle growth regulators, IGFI-Ea, MGF and myostatin, or other muscle-specific genes. IGFI-Ea, MGF and insulin activate the phosphoinositide-3-kinase (PI3K)/PKB/mTOR pathway, which enhances protein synthesis via increased translational initiation and the synthesis of ribosomal proteins for ribosome biogenesis. Availability of amino acids will activate mTOR signalling, whereas an increased energy demand sensed by AMPK will inhibit mTOR.
Molecular mechanisms potentially leading to muscle hypertrophy. See text for details. AR, androgenic receptor; HGF, hepatocyte growth factor; IGF, insulin-like growth factor; MGF, mechanical growth factors; MAPK, mitogen-activated protein kinase; AMPK, AMP-activated protein kinase; mTOR, mammalian target of rapamycin
In addition to stimulate muscle fibre hypertrophy, IGFI-Ea, MGF, testosterone, myostatin and various other factors also regulate an increased proliferation and/or differentiation of satellite cells [64]. Hypertrophy is thus produced not only through binding of growth factors on/in the skeletal muscle fibres but also through binding to receptors on/in activated satellite cells. Although IGF-I protein is one major mediator of these hypertrophic effects, it has been suggested that the isoform from which IGF-I is produced can affect its potency. MGF has been proposed to be more potent for promoting rat skeletal muscle hypertrophy [53]. However, this proposal has been countered in another study in young growing rats, where the ability of MGF to promote muscle hypertrophy was not greater than IGF-IEa, with a barely non-existent effect in adult muscle [13]. It is thus possible that MGF can promote hypertrophy only when there is an active satellite cell pool, as observed in growing animals [13]. A decreased response of both IGF-I splice variants to resistance exercise with age has been suggested and tested in human as well, but opposite results have been found. Some studies reported a blunted response of MGF mRNA after resistance exercise in older compared to young subjects [58, 117] while a recent study found a higher response instead [3]. This discrepancy cannot be explained by the intensity of the exercise used in those different studies as both 60–65% 1-RM and 80–85% 1-RM induced similar increases in IGF-IEa and MGF mRNA levels in human skeletal muscle [160]. Of note, there is considerable variation between individuals in the response of IGF-IEa and MGF mRNA to exercise [58], which contributes to the difficulty to draw clear conclusions about their regulation and their role in muscle accretion.
The signalling pathways leading to transcriptional and translational changes in skeletal muscle in response to resistance exercise are still not fully understood. Four potential stimuli that may regulate these processes have been proposed: mechanical load or stress, intracellular calcium, hypoxia and redox state [9]. These are thought to be first messengers in a signalling cascade in which various transcription factors, hormones and other regulatory proteins are activated. Signal transduction pathways shown to be activated in response to various forms of contraction include those involving AMPK [162], calcineurin [98], extracellular signal-regulated kinase 1/2 (ERK1/2) and p38 [159], c-Jun N-terminal kinase (JNK) [7], nuclear factor kappa B (NF-κB) [65], PI3K/PKB/mTOR [151] and protein kinase C (PKC) [125].
Calcineurin is a calcium–calmodulin-activated protein phosphatase that dephosphorylates the transcription factor nuclear factor of activated T cells (NFAT), enabling its nuclear translocation and DNA binding. The calcineurin pathway has been linked not only to the regulation of skeletal muscle growth but also to the conversion of fast-to-slow phenotype [112] although, currently, the link with muscle growth is questioned. Various sensors of mechanical strain seem to possess the ability to translate strain into chemical signals that induce the activation of skeletal muscle gene promoters [27]. A possible candidate sensor of the increase in mechanical strain is focal adhesion kinase, a protein localised to the sarcolemma [46]. The serum response factor, which is a transcription factor, is a substrate of focal adhesion kinase, thereby providing a transcriptional link between membrane, the genome and subsequent expression of muscle protein. The putative link between focal adhesion kinase and serum response factor is the β1 integrin–RhoA signalling [156]. Different modes of exercise affect ERK1/2 and p38 MAPK likely in an intensity-dependent manner [106, 159]. However, only those stimuli likely to result in hypertrophy, such as high-frequency electrical stimulation, increased ribosomal protein S6 kinase (S6K1) and PKB phosphorylation [106]. IGFI-Ea, MGF and myostatin are not directly regulated by stretch, overload or muscle contraction, but by the signal transduction pathways that sense these stimuli and, consequently, regulate the availability of these muscle growth factors for receptor binding. The major step controlling the availability of IGFI-Ea, MGF and myostatin in response growth-inducing stimuli appears to be transcriptional regulation [124]. In summary, muscle growth stimuli lead to the activation of a signal transduction network and to a changed availability of the major muscle growth factors IGFI-Ea, MGF and myostatin. The activated signal transduction pathways and changed growth factor availability will then regulate the activity of muscle growth executors, which are the translational or protein synthesis machinery and, possibly, satellite cells.
It must be stated here that the view implying a major role of IGF-I in skeletal muscle hypertrophy induced by mechanical loading through the activation/proliferation of satellite cells and/or the increase in protein synthesis has been questioned [139]. It was found that IGF-IR was not necessary for the induction of skeletal muscle growth in response to mechanical loading, whereas components of the PKB pathway were activated. Those results were confirmed by others [157, 158]. Resistance exercise was performed either with arm muscles alone or with both arm and leg muscles to induce either low or high elevations in growth hormone, testosterone and IGF-1. Yet, despite markedly different systemic concentrations between the two groups, there was no difference in acute mTORC1 signalling or in muscle protein synthesis or in long-term adaptations to training in terms of mass and strength gains [157, 158]. However, suppression of testosterone production via the use of a gonadotropin-releasing hormone analogue in humans ablates muscle growth responses to resistance exercise [84], thus suggesting that testosterone remains an integral part of protein metabolic responses to exercise. These results suggest that IGF-I and growth hormone are not the key factors necessary for initiation of growth response to mechanical load and that other factors are probably responsible for the activation of the PKB/mTORC1 pathway, such as, for example, intrinsic mechanosensors.
Mechanotransduction is the process of converting mechanical signals that are sensed in response to cellular movement into molecular signals, and numerous candidate “mechanosensors” have been suggested in skeletal muscle. One target recently found was phospholipase D, which increases the production of the lipid second messenger phosphatidic acid in a mechanosensitive manner. Phosphatidic acid signalling was found to be upstream of contraction-induced activation of mTOR. Indeed, pharmacological inhibition of phospholipase D impaired activation of mTOR in response to muscle contractions [110]. The focal adhesion complexes are other possible mechanosensitive sensors that link the extracellular matrix to the cytoplasmic cytoskeleton. They consist of a variety of extracellular matrix receptors/integrins, intracellular cytoskeletal and signalling molecules [82]. Interactions of extracellular matrix proteins with integrin receptors stimulate intracellular signalling pathways important in cell growth and migration in adult skeletal muscle [133]. Activation of integrin receptors appears to be a common feature of muscle remodeling in response to, amongst others, endurance exercise [148]. Focal adhesion kinase (FAK), which localises to focal adhesion complexes, is a non-receptor tyrosine kinase, which can be phosphorylated at Tyr397, and thereby activated, upon engagement of integrin receptors [22, 28]. A growing body of evidence has associated FAK activation with responses to mechanical stress in skeletal muscle [41]. Indeed, FAK phosphorylation has been shown to be increased in overload models in mice [46, 57] and resistance exercise in humans [161]. In addition, local overexpression of FAK in rodent skeletal muscle stimulated muscle hypertrophy [82]. All together, those results indicate that FAK is a legitimate mechanosensitive component of muscle hypertrophy.
Concurrent exercise
Endurance and resistance exercise induce different physiological and molecular adaptations that could potentially interfere with each other. Performing endurance and resistance exercise concurrently could be detrimental for some adaptations, from which the interference hypothesis was raised [16]. At the molecular level, it was originally thought that the activation of AMPK by endurance exercise could inhibit the mTOR pathway, which is activated by resistance exercise [30]. In the context of the present review, consequent to the reduction in mTOR signalling, the typical anabolic response after resistance exercise would be reduced as well [105]. However, this potential mechanism was nuanced depending on whether a single exercise or a training process was considered [49]. Without questioning, the inhibitory effect of AMPK on mTOR, which appears during exercise, seems to be lower in humans than in rodents [69]. Moreover, during a training process of a few weeks, the hypertrophic response was not altered by the addition of endurance training to a resistance training [88]. It seems that a rest period of at least 6–24 h is needed to avoid any interference between both modalities of exercise, at least when looking at muscle hypertrophy [104]. All together, the recent findings seem to indicate that the interference phenomenon originally described by Hickson in 1980 [63] could be reduced or totally abrogated if training parameters are planned appropriately [49].
Interindividual variability
While it is now well established that resistance exercise stimulates muscle protein synthesis and promotes muscle mass and strength gains, substantial variability exists following standardised resistance training programs in the magnitude of those gains from one individual to another. Changes in muscle size ranging from 3% up to almost 60% have been measured following 12 weeks of resistance training in healthy young adults [68]. In addition to alterations in satellite cell population, myogenic gene expression, miRNA and gene polymorphisms, it has recently been postulated that the circadian rhythms and underlying molecular clock signals could contribute to this variability as well [23]. For example, taking care of performing resistance exercise at a time when cortisol levels are low may allow for increased plasma IGF-1 levels to, in turn, allow the subsequent activation of S6K1. It has been shown that the performance of resistance exercise in the evening compared to the morning was associated with reduced plasma cortisol levels, indicating the potential for evening over morning to reduce the catabolic environment and to promote muscle hypertrophy adaptations [19, 21].
Perspectives
While the physiological adaptations to endurance and resistance training seem relatively different and specific to the modality of training, this could only be apparent. Indeed, a priori, an increase in the mitochondrial pool and efficiency is a hallmark of endurance training. Considering the huge amount of energy protein synthesis and muscle accretion required, the role of mitochondria to provide ATP during muscle hypertrophy has maybe been neglected so far. If this reveals true, any training strategy aiming at increasing mitochondrial content and function could help to speed up the accretion of muscle mass, assuming the availability of amino acids is optimal. For sure, this would ask a fine balance between endurance and resistance exercise to avoid any interference as mentioned above but this hypothesis is probably worth to be tested.
Conclusion
In conclusion, both resistance and endurance exercise regulate protein balance and satellite cell inclusion. However, resistance training has a higher potential than endurance training to increase muscle mass. While several mechanisms leading to muscle hypertrophy have been discovered, further investigation should focus on the specific regulation of those mechanisms according to the training status, the age and the gender.
References
Abreu P, Mendes SV, Ceccatto VM, Hirabara SM (2017) Satellite cell activation induced by aerobic muscle adaptation in response to endurance exercise in humans and rodents. Life Sci 170:33–40. https://doi.org/10.1016/j.lfs.2016.11.016
Adams GR, Bamman MM (2012) Characterization and regulation of mechanical loading-induced compensatory muscle hypertrophy. Compr Physiol 2:2829–2870. https://doi.org/10.1002/cphy.c110066
Ahtiainen JP, Hulmi JJ, Lehti M, Kraemer WJ, Nyman K, Selanne H, Alen M, Komulainen J, Kovanen V, Mero AA, Philippou A, Laakkonen EK, Hakkinen K (2016) Effects of resistance training on expression of IGF-I splice variants in younger and older men. Eur J Sport Sci 16:1055–1063. https://doi.org/10.1080/17461391.2016.1185164
Andersen G, Orngreen MC, Preisler N, Jeppesen TD, Krag TO, Hauerslev S, van Hall G, Vissing J (2015) Protein-carbohydrate supplements improve muscle protein balance in muscular dystrophy patients after endurance exercise: a placebo-controlled crossover study. Am J Physiol Regul Integr Comp Physiol 308:R123–R130. https://doi.org/10.1152/ajpregu.00321.2014
Anderson J, Pilipowicz O (2002) Activation of muscle satellite cells in single-fiber cultures. Nitric Oxide 7:36–41
Aoi W, Naito Y, Mizushima K, Takanami Y, Kawai Y, Ichikawa H, Yoshikawa T (2010) The microRNA miR-696 regulates PGC-1{alpha} in mouse skeletal muscle in response to physical activity. Am J Physiol Endocrinol Metab 298:E799–E806. https://doi.org/10.1152/ajpendo.00448.2009
Aronson D, Boppart MD, Dufresne SD, Fielding RA, Goodyear LJ (1998) Exercise stimulates c-Jun NH2 kinase activity and c-Jun transcriptional activity in human skeletal muscle. Biochem Biophys Res Commun 251:106–110. https://doi.org/10.1006/bbrc.1998.9435
Atherton PJ, Phillips BE, Wilkinson DJ (2015) Exercise and regulation of protein metabolism. Prog Mol Biol Transl Sci 135:75–98. https://doi.org/10.1016/bs.pmbts.2015.06.015
Baar K, Blough E, Dineen B, Esser K (1999) Transcriptional regulation in response to exercise. Exerc Sport Sci Rev 27:333–379
Bamman MM, Roberts BM, Adams GR (2018) Molecular regulation of exercise-induced muscle fiber hypertrophy. Cold Spring Harb Perspect Med 8. https://doi.org/10.1101/cshperspect.a029751
Barres R, Yan J, Egan B, Treebak JT, Rasmussen M, Fritz T, Caidahl K, Krook A, O’Gorman DJ, Zierath JR (2012) Acute exercise remodels promoter methylation in human skeletal muscle. Cell Metab 15:405–411. https://doi.org/10.1016/j.cmet.2012.01.001
Barth S, Glick D, Macleod KF (2010) Autophagy: assays and artifacts. J Pathol 221:117–124. https://doi.org/10.1002/path.2694
Barton ER (2006) Viral expression of insulin-like growth factor-I isoforms promotes different responses in skeletal muscle. J Appl Physiol (1985) 100:1778–1784. https://doi.org/10.1152/japplphysiol.01405.2005
Bazgir B, Fathi R, Rezazadeh Valojerdi M, Mozdziak P, Asgari A (2017) Satellite cells contribution to exercise mediated muscle hypertrophy and repair. Cell J 18:473–484
Bechet D, Tassa A, Taillandier D, Combaret L, Attaix D (2005) Lysosomal proteolysis in skeletal muscle. Int J Biochem Cell Biol 37:2098–2114. https://doi.org/10.1016/j.biocel.2005.02.029
Berryman N, Mujika I, Bosquet L (2018) Concurrent training for sports performance: the two sides of the medal. Int J Sports Physiol Perform:1–22. https://doi.org/10.1123/ijspp.2018-0103
Beyersmann D (2000) Regulation of mammalian gene expression. EXS 89:11–28
Bird JW, Carter JH, Triemer RE, Brooks RM, Spanier AM (1980) Proteinases in cardiac and skeletal muscle. Fed Proc 39:20–25
Bird SP, Tarpenning KM (2004) Influence of circadian time structure on acute hormonal responses to a single bout of heavy-resistance exercise in weight-trained men. Chronobiol Int 21:131–146
Burd NA, West DW, Staples AW, Atherton PJ, Baker JM, Moore DR, Holwerda AM, Parise G, Rennie MJ, Baker SK, Phillips SM (2010) Low-load high volume resistance exercise stimulates muscle protein synthesis more than high-load low volume resistance exercise in young men. PLoS One 5:e12033. https://doi.org/10.1371/journal.pone.0012033
Burley SD, Whittingham-Dowd J, Allen J, Grosset JF, Onambele-Pearson GL (2016) The differential hormonal milieu of morning versus evening may have an impact on muscle hypertrophic potential. PLoS One 11:e0161500. https://doi.org/10.1371/journal.pone.0161500
Calalb MB, Polte TR, Hanks SK (1995) Tyrosine phosphorylation of focal adhesion kinase at sites in the catalytic domain regulates kinase activity: a role for Src family kinases. Mol Cell Biol 15:954–963
Camera DM (2018) Anabolic heterogeneity following resistance training: a role for circadian rhythm? Front Physiol 9:569. https://doi.org/10.3389/fphys.2018.00569
Camera DM, Smiles WJ, Hawley JA (2016) Exercise-induced skeletal muscle signaling pathways and human athletic performance. Free Radic Biol Med 98:131–143. https://doi.org/10.1016/j.freeradbiomed.2016.02.007
Campion DR (1984) The muscle satellite cell: a review. Int Rev Cytol 87:225–251
Carraro F, Stuart CA, Hartl WH, Rosenblatt J, Wolfe RR (1990) Effect of exercise and recovery on muscle protein synthesis in human subjects. Am J Phys 259:E470–E476. https://doi.org/10.1152/ajpendo.1990.259.4.E470
Carson JA, Wei L (2000) Integrin signaling’s potential for mediating gene expression in hypertrophying skeletal muscle. J Appl Physiol (1985) 88:337–343. https://doi.org/10.1152/jappl.2000.88.1.337
Cary LA, Guan JL (1999) Focal adhesion kinase in integrin-mediated signaling. Front Biosci 4:D102–D113
Cheng Z, Zheng L, Almeida FA (2018) Epigenetic reprogramming in metabolic disorders: nutritional factors and beyond. J Nutr Biochem 54:1–10. https://doi.org/10.1016/j.jnutbio.2017.10.004
Coffey VG, Hawley JA (2007) The molecular bases of training adaptation. Sports Med 37:737–763
Coffey VG, Hawley JA (2017) Concurrent exercise training: do opposites distract? J Physiol 595:2883–2896. https://doi.org/10.1113/JP272270
Crameri RM, Langberg H, Magnusson P, Jensen CH, Schroder HD, Olesen JL, Suetta C, Teisner B, Kjaer M (2004) Changes in satellite cells in human skeletal muscle after a single bout of high intensity exercise. J Physiol 558:333–340. https://doi.org/10.1113/jphysiol.2004.061846
Davidsen PK, Gallagher IJ, Hartman JW, Tarnopolsky MA, Dela F, Helge JW, Timmons JA, Phillips SM (2011) High responders to resistance exercise training demonstrate differential regulation of skeletal muscle microRNA expression. J Appl Physiol (1985) 110:309–317. https://doi.org/10.1152/japplphysiol.00901.2010
Deane CS, Wilkinson DJ, Phillips BE, Smith K, Etheridge T, Atherton PJ (2017) “Nutraceuticals” in relation to human skeletal muscle and exercise. Am J Physiol Endocrinol Metab 312:E282–E299. https://doi.org/10.1152/ajpendo.00230.2016
Deaton AM, Bird A (2011) CpG islands and the regulation of transcription. Genes Dev 25:1010–1022. https://doi.org/10.1101/gad.2037511
Deldicque L, Atherton P, Patel R, Theisen D, Nielens H, Rennie MJ, Francaux M (2008) Effects of resistance exercise with and without creatine supplementation on gene expression and cell signaling in human skeletal muscle. J Appl Physiol (1985) 104:371–378. https://doi.org/10.1152/japplphysiol.00873.2007
Di Donato DM, West DW, Churchward-Venne TA, Breen L, Baker SK, Phillips SM (2014) Influence of aerobic exercise intensity on myofibrillar and mitochondrial protein synthesis in young men during early and late postexercise recovery. Am J Physiol Endocrinol Metab 306:E1025–E1032. https://doi.org/10.1152/ajpendo.00487.2013
Donges CE, Burd NA, Duffield R, Smith GC, West DW, Short MJ, Mackenzie R, Plank LD, Shepherd PR, Phillips SM, Edge JA (2012) Concurrent resistance and aerobic exercise stimulates both myofibrillar and mitochondrial protein synthesis in sedentary middle-aged men. J Appl Physiol (1985) 112:1992–2001. https://doi.org/10.1152/japplphysiol.00166.2012
Drummond MJ, McCarthy JJ, Fry CS, Esser KA, Rasmussen BB (2008) Aging differentially affects human skeletal muscle microRNA expression at rest and after an anabolic stimulus of resistance exercise and essential amino acids. Am J Physiol Endocrinol Metab 295:E1333–E1340. https://doi.org/10.1152/ajpendo.90562.2008
Du J, Wang X, Miereles C, Bailey JL, Debigare R, Zheng B, Price SR, Mitch WE (2004) Activation of caspase-3 is an initial step triggering accelerated muscle proteolysis in catabolic conditions. J Clin Invest 113:115–123. https://doi.org/10.1172/JCI18330
Durieux AC, Desplanches D, Freyssenet D, Fluck M (2007) Mechanotransduction in striated muscle via focal adhesion kinase. Biochem Soc Trans 35:1312–1313. https://doi.org/10.1042/BST0351312
Elia L, Contu R, Quintavalle M, Varrone F, Chimenti C, Russo MA, Cimino V, De Marinis L, Frustaci A, Catalucci D, Condorelli G (2009) Reciprocal regulation of microRNA-1 and insulin-like growth factor-1 signal transduction cascade in cardiac and skeletal muscle in physiological and pathological conditions. Circulation 120:2377–2385. https://doi.org/10.1161/CIRCULATIONAHA.109.879429
Fagan JM, Waxman L, Goldberg AL (1987) Skeletal muscle and liver contain a soluble ATP + ubiquitin-dependent proteolytic system. Biochem J 243:335–343
Fernandes T, Magalhaes FC, Roque FR, Phillips MI, Oliveira EM (2012) Exercise training prevents the microvascular rarefaction in hypertension balancing angiogenic and apoptotic factors: role of microRNAs-16, -21, and -126. Hypertension 59:513–520. https://doi.org/10.1161/HYPERTENSIONAHA.111.185801
Figueiredo VC, Markworth JF (2015) Mechanisms of protein synthesis activation following exercise: new pieces to the increasingly complex puzzle. J Physiol 593:4693–4695. https://doi.org/10.1113/JP271431
Fluck M, Carson JA, Gordon SE, Ziemiecki A, Booth FW (1999) Focal adhesion proteins FAK and paxillin increase in hypertrophied skeletal muscle. Am J Phys 277:C152–C162
Fritzen AM, Madsen AB, Kleinert M, Treebak JT, Lundsgaard AM, Jensen TE, Richter EA, Wojtaszewski J, Kiens B, Frosig C (2016) Regulation of autophagy in human skeletal muscle: effects of exercise, exercise training and insulin stimulation. J Physiol 594:745–761. https://doi.org/10.1113/JP271405
Fry CS, Drummond MJ, Glynn EL, Dickinson JM, Gundermann DM, Timmerman KL, Walker DK, Volpi E, Rasmussen BB (2013) Skeletal muscle autophagy and protein breakdown following resistance exercise are similar in younger and older adults. J Gerontol A Biol Sci Med Sci 68:599–607. https://doi.org/10.1093/gerona/gls209
Fyfe JJ, Bishop DJ, Stepto NK (2014) Interference between concurrent resistance and endurance exercise: molecular bases and the role of individual training variables. Sports Med 44:743–762. https://doi.org/10.1007/s40279-014-0162-1
Gissel H (2005) The role of Ca2+ in muscle cell damage. Ann N Y Acad Sci 1066:166–180. https://doi.org/10.1196/annals.1363.013
Glickman MH, Ciechanover A (2002) The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev 82:373–428. https://doi.org/10.1152/physrev.00027.2001
Glynn EL, Fry CS, Drummond MJ, Dreyer HC, Dhanani S, Volpi E, Rasmussen BB (2010) Muscle protein breakdown has a minor role in the protein anabolic response to essential amino acid and carbohydrate intake following resistance exercise. Am J Physiol Regul Integr Comp Physiol 299:R533–R540. https://doi.org/10.1152/ajpregu.00077.2010
Goldspink G (2005) Mechanical signals, IGF-I gene splicing, and muscle adaptation. Physiology (Bethesda) 20:232–238. https://doi.org/10.1152/physiol.00004.2005
Goll DE, Thompson VF, Li H, Wei W, Cong J (2003) The calpain system. Physiol Rev 83:731–801. https://doi.org/10.1152/physrev.00029.2002
Goodman CA (2014) The role of mTORC1 in regulating protein synthesis and skeletal muscle mass in response to various mechanical stimuli. Rev Physiol Biochem Pharmacol 166:43–95. https://doi.org/10.1007/112_2013_17
Gordon BS, Liu C, Steiner JL, Nader GA, Jefferson LS, Kimball SR (2016) Loss of REDD1 augments the rate of the overload-induced increase in muscle mass. Am J Physiol Regul Integr Comp Physiol 311:R545–R557. https://doi.org/10.1152/ajpregu.00159.2016
Gordon SE, Fluck M, Booth FW (2001) Selected contribution: skeletal muscle focal adhesion kinase, paxillin, and serum response factor are loading dependent. J Appl Physiol (1985) 90:1174–1183; discussion 1165. https://doi.org/10.1152/jappl.2001.90.3.1174
Hameed M, Orrell RW, Cobbold M, Goldspink G, Harridge SD (2003) Expression of IGF-I splice variants in young and old human skeletal muscle after high resistance exercise. J Physiol 547:247–254. https://doi.org/10.1113/jphysiol.2002.032136
Harber MP, Crane JD, Dickinson JM, Jemiolo B, Raue U, Trappe TA, Trappe SW (2009) Protein synthesis and the expression of growth-related genes are altered by running in human vastus lateralis and soleus muscles. Am J Physiol Regul Integr Comp Physiol 296:R708–R714. https://doi.org/10.1152/ajpregu.90906.2008
Harber MP, Konopka AR, Jemiolo B, Trappe SW, Trappe TA, Reidy PT (2010) Muscle protein synthesis and gene expression during recovery from aerobic exercise in the fasted and fed states. Am J Physiol Regul Integr Comp Physiol 299:R1254–R1262. https://doi.org/10.1152/ajpregu.00348.2010
He L, Hannon GJ (2004) MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5:522–531. https://doi.org/10.1038/nrg1379
Heinemeier KM, Schjerling P, Heinemeier J, Magnusson SP, Kjaer M (2013) Lack of tissue renewal in human adult Achilles tendon is revealed by nuclear bomb (14)C. FASEB J 27:2074–2079. https://doi.org/10.1096/fj.12-225599
Hickson RC (1980) Interference of strength development by simultaneously training for strength and endurance. Eur J Appl Physiol Occup Physiol 45:255–263
Hill M, Goldspink G (2003) Expression and splicing of the insulin-like growth factor gene in rodent muscle is associated with muscle satellite (stem) cell activation following local tissue damage. J Physiol 549:409–418. https://doi.org/10.1113/jphysiol.2002.035832
Hollander J, Fiebig R, Gore M, Ookawara T, Ohno H, Ji LL (2001) Superoxide dismutase gene expression is activated by a single bout of exercise in rat skeletal muscle. Pflugers Arch 442:426–434
Holloszy JO (1967) Biochemical adaptations in muscle. Effects of exercise on mitochondrial oxygen uptake and respiratory enzyme activity in skeletal muscle. J Biol Chem 242:2278–2282
Holloszy JO, Coyle EF (1984) Adaptations of skeletal muscle to endurance exercise and their metabolic consequences. J Appl Physiol Respir Environ Exerc Physiol 56:831–838. https://doi.org/10.1152/jappl.1984.56.4.831
Hubal MJ, Gordish-Dressman H, Thompson PD, Price TB, Hoffman EP, Angelopoulos TJ, Gordon PM, Moyna NM, Pescatello LS, Visich PS, Zoeller RF, Seip RL, Clarkson PM (2005) Variability in muscle size and strength gain after unilateral resistance training. Med Sci Sports Exerc 37:964–972
Hughes DC, Ellefsen S, Baar K (2018) Adaptations to endurance and strength training. Cold Spring Harb Perspect Med 8. https://doi.org/10.1101/cshperspect.a029769
Jacobs BL, Goodman CA, Hornberger TA (2014) The mechanical activation of mTOR signaling: an emerging role for late endosome/lysosomal targeting. J Muscle Res Cell Motil 35:11–21. https://doi.org/10.1007/s10974-013-9367-4
Jaenisch R, Bird A (2003) Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet 33(Suppl):245–254. https://doi.org/10.1038/ng1089
Jamart C, Benoit N, Raymackers JM, Kim HJ, Kim CK, Francaux M (2012) Autophagy-related and autophagy-regulatory genes are induced in human muscle after ultraendurance exercise. Eur J Appl Physiol 112:3173–3177. https://doi.org/10.1007/s00421-011-2287-3
Jamart C, Naslain D, Gilson H, Francaux M (2013) Higher activation of autophagy in skeletal muscle of mice during endurance exercise in the fasted state. Am J Physiol Endocrinol Metab 305:E964–E974. https://doi.org/10.1152/ajpendo.00270.2013
Jenuwein T, Allis CD (2001) Translating the histone code. Science 293:1074–1080. https://doi.org/10.1126/science.1063127
Jones PA (2012) Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet 13:484–492. https://doi.org/10.1038/nrg3230
Jones PA, Baylin SB (2007) The epigenomics of cancer. Cell 128:683–692. https://doi.org/10.1016/j.cell.2007.01.029
Kadi F, Charifi N, Denis C, Lexell J, Andersen JL, Schjerling P, Olsen S, Kjaer M (2005) The behaviour of satellite cells in response to exercise: what have we learned from human studies? Pflugers Arch 451:319–327. https://doi.org/10.1007/s00424-005-1406-6
Kadi F, Schjerling P, Andersen LL, Charifi N, Madsen JL, Christensen LR, Andersen JL (2004) The effects of heavy resistance training and detraining on satellite cells in human skeletal muscles. J Physiol 558:1005–1012. https://doi.org/10.1113/jphysiol.2004.065904
Kinbara K, Sorimachi H, Ishiura S, Suzuki K (1997) Muscle-specific calpain, p94, interacts with the extreme C-terminal region of connectin, a unique region flanked by two immunoglobulin C2 motifs. Arch Biochem Biophys 342:99–107. https://doi.org/10.1006/abbi.1997.0108
Kirby TJ, McCarthy JJ, Peterson CA, Fry CS (2016) Synergist ablation as a rodent model to study satellite cell dynamics in adult skeletal muscle. Methods Mol Biol 1460:43–52. https://doi.org/10.1007/978-1-4939-3810-0_4
Klionsky DJ (2007) Autophagy: from phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol 8:931–937. https://doi.org/10.1038/nrm2245
Klossner S, Durieux AC, Freyssenet D, Flueck M (2009) Mechano-transduction to muscle protein synthesis is modulated by FAK. Eur J Appl Physiol 106:389–398. https://doi.org/10.1007/s00421-009-1032-7
Kumar V, Selby A, Rankin D, Patel R, Atherton P, Hildebrandt W, Williams J, Smith K, Seynnes O, Hiscock N, Rennie MJ (2009) Age-related differences in the dose-response relationship of muscle protein synthesis to resistance exercise in young and old men. J Physiol 587:211–217. https://doi.org/10.1113/jphysiol.2008.164483
Kvorning T, Andersen M, Brixen K, Madsen K (2006) Suppression of endogenous testosterone production attenuates the response to strength training: a randomized, placebo-controlled, and blinded intervention study. Am J Physiol Endocrinol Metab 291:E1325–E1332. https://doi.org/10.1152/ajpendo.00143.2006
Langley B, Thomas M, Bishop A, Sharma M, Gilmour S, Kambadur R (2002) Myostatin inhibits myoblast differentiation by down-regulating MyoD expression. J Biol Chem 277:49831–49840. https://doi.org/10.1074/jbc.M204291200
Louis E, Raue U, Yang Y, Jemiolo B, Trappe S (2007) Time course of proteolytic, cytokine, and myostatin gene expression after acute exercise in human skeletal muscle. J Appl Physiol (1985) 103:1744–1751. https://doi.org/10.1152/japplphysiol.00679.2007
Lowe DA, Alway SE (1999) Stretch-induced myogenin, MyoD, and MRF4 expression and acute hypertrophy in quail slow-tonic muscle are not dependent upon satellite cell proliferation. Cell Tissue Res 296:531–539
Lundberg TR, Fernandez-Gonzalo R, Gustafsson T, Tesch PA (2013) Aerobic exercise does not compromise muscle hypertrophy response to short-term resistance training. J Appl Physiol (1985) 114:81–89. https://doi.org/10.1152/japplphysiol.01013.2012
Martin-Rincon M, Morales-Alamo D, Calbet JAL (2018) Exercise-mediated modulation of autophagy in skeletal muscle. Scand J Med Sci Sports 28:772–781. https://doi.org/10.1111/sms.12945
Mascher H, Ekblom B, Rooyackers O, Blomstrand E (2011) Enhanced rates of muscle protein synthesis and elevated mTOR signalling following endurance exercise in human subjects. Acta Physiol (Oxf) 202:175–184. https://doi.org/10.1111/j.1748-1716.2011.02274.x
Mauro A (1961) Satellite cell of skeletal muscle fibers. J Biophys Biochem Cytol 9:493–495
Mayer C, Grummt I (2006) Ribosome biogenesis and cell growth: mTOR coordinates transcription by all three classes of nuclear RNA polymerases. Oncogene 25:6384–6391. https://doi.org/10.1038/sj.onc.1209883
McCarthy JJ, Esser KA (2007) Counterpoint: satellite cell addition is not obligatory for skeletal muscle hypertrophy. J Appl Physiol (1985) 103:1100–1102; discussion 1102–1103. https://doi.org/10.1152/japplphysiol.00101.2007a
McCarthy JJ, Esser KA (2007) MicroRNA-1 and microRNA-133a expression are decreased during skeletal muscle hypertrophy. J Appl Physiol (1985) 102:306–313. doi:https://doi.org/10.1152/japplphysiol.00932.2006
McGee SL, Hargreaves M (2011) Histone modifications and exercise adaptations. J Appl Physiol (1985) 110:258–263. https://doi.org/10.1152/japplphysiol.00979.2010
McGee SL, Walder KR (2017) Exercise and the skeletal muscle epigenome. Cold Spring Harb Perspect Med 7. https://doi.org/10.1101/cshperspect.a029876
McGlory C, van Vliet S, Stokes T, Mittendorfer B, Phillips SM (2018) The impact of exercise and nutrition in the regulation of skeletal muscle mass. J Physiol. https://doi.org/10.1113/JP275443
Meissner JD, Kubis HP, Scheibe RJ, Gros G (2000) Reversible Ca2+-induced fast-to-slow transition in primary skeletal muscle culture cells at the mRNA level. J Physiol 523(Pt 1):19–28
Miller BF, Hansen M, Olesen JL, Schwarz P, Babraj JA, Smith K, Rennie MJ, Kjaer M (2007) Tendon collagen synthesis at rest and after exercise in women. J Appl Physiol (1985) 102:541–546. https://doi.org/10.1152/japplphysiol.00797.2006
Miller BF, Olesen JL, Hansen M, Dossing S, Crameri RM, Welling RJ, Langberg H, Flyvbjerg A, Kjaer M, Babraj JA, Smith K, Rennie MJ (2005) Coordinated collagen and muscle protein synthesis in human patella tendon and quadriceps muscle after exercise. J Physiol 567:1021–1033. https://doi.org/10.1113/jphysiol.2005.093690
Moller AB, Vendelbo MH, Christensen B, Clasen BF, Bak AM, Jorgensen JO, Moller N, Jessen N (2015) Physical exercise increases autophagic signaling through ULK1 in human skeletal muscle. J Appl Physiol (1985) 118:971–979. https://doi.org/10.1152/japplphysiol.01116.2014
Moore DR, Young M, Phillips SM (2012) Similar increases in muscle size and strength in young men after training with maximal shortening or lengthening contractions when matched for total work. Eur J Appl Physiol 112:1587–1592. https://doi.org/10.1007/s00421-011-2078-x
Morton RW, Murphy KT, McKellar SR, Schoenfeld BJ, Henselmans M, Helms E, Aragon AA, Devries MC, Banfield L, Krieger JW, Phillips SM (2018) A systematic review, meta-analysis and meta-regression of the effect of protein supplementation on resistance training-induced gains in muscle mass and strength in healthy adults. Br J Sports Med 52:376–384. https://doi.org/10.1136/bjsports-2017-097608
Murach KA, Bagley JR (2016) Skeletal muscle hypertrophy with concurrent exercise training: contrary evidence for an interference effect. Sports Med 46:1029–1039. https://doi.org/10.1007/s40279-016-0496-y
Nader GA (2006) Concurrent strength and endurance training: from molecules to man. Med Sci Sports Exerc 38:1965–1970. https://doi.org/10.1249/01.mss.0000233795.39282.33
Nader GA, Esser KA (2001) Intracellular signaling specificity in skeletal muscle in response to different modes of exercise. J Appl Physiol (1985) 90:1936–1942. https://doi.org/10.1152/jappl.2001.90.5.1936
Nedergaard A, Vissing K, Overgaard K, Kjaer M, Schjerling P (2007) Expression patterns of atrogenic and ubiquitin proteasome component genes with exercise: effect of different loading patterns and repeated exercise bouts. J Appl Physiol (1985) 103:1513–1522. https://doi.org/10.1152/japplphysiol.01445.2006
Nielsen S, Scheele C, Yfanti C, Akerstrom T, Nielsen AR, Pedersen BK, Laye MJ (2010) Muscle specific microRNAs are regulated by endurance exercise in human skeletal muscle. J Physiol 588:4029–4037. https://doi.org/10.1113/jphysiol.2010.189860
O’Connor RS, Pavlath GK, McCarthy JJ, Esser KA (2007) Last word on point:counterpoint: satellite cell addition is/is not obligatory for skeletal muscle hypertrophy. J Appl Physiol (1985) 103:1107. https://doi.org/10.1152/japplphysiol.00502.2007
O’Neil TK, Duffy LR, Frey JW, Hornberger TA (2009) The role of phosphoinositide 3-kinase and phosphatidic acid in the regulation of mammalian target of rapamycin following eccentric contractions. J Physiol 587:3691–3701. https://doi.org/10.1113/jphysiol.2009.173609
Olsen S, Aagaard P, Kadi F, Tufekovic G, Verney J, Olesen JL, Suetta C, Kjaer M (2006) Creatine supplementation augments the increase in satellite cell and myonuclei number in human skeletal muscle induced by strength training. J Physiol 573:525–534. https://doi.org/10.1113/jphysiol.2006.107359
Olson EN, Williams RS (2000) Calcineurin signaling and muscle remodeling. Cell 101:689–692
Pankiv S, Clausen TH, Lamark T, Brech A, Bruun JA, Outzen H, Overvatn A, Bjorkoy G, Johansen T (2007) p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J Biol Chem 282:24131–24145. https://doi.org/10.1074/jbc.M702824200
Pareja-Galeano H, Sanchis-Gomar F, Garcia-Gimenez JL (2014) Physical exercise and epigenetic modulation: elucidating intricate mechanisms. Sports Med 44:429–436. https://doi.org/10.1007/s40279-013-0138-6
Pasiakos SM, Carbone JW (2014) Assessment of skeletal muscle proteolysis and the regulatory response to nutrition and exercise. IUBMB Life 66:478–484. https://doi.org/10.1002/iub.1291
Pasiakos SM, McClung HL, McClung JP, Urso ML, Pikosky MA, Cloutier GJ, Fielding RA, Young AJ (2010) Molecular responses to moderate endurance exercise in skeletal muscle. Int J Sport Nutr Exerc Metab 20:282–290
Petrella JK, Kim JS, Cross JM, Kosek DJ, Bamman MM (2006) Efficacy of myonuclear addition may explain differential myofiber growth among resistance-trained young and older men and women. Am J Physiol Endocrinol Metab 291:E937–E946. https://doi.org/10.1152/ajpendo.00190.2006
Phillips SM, Hartman JW, Wilkinson SB (2005) Dietary protein to support anabolism with resistance exercise in young men. J Am Coll Nutr 24:134S–139S
Phillips SM, Tipton KD, Aarsland A, Wolf SE, Wolfe RR (1997) Mixed muscle protein synthesis and breakdown after resistance exercise in humans. Am J Phys 273:E99–E107. https://doi.org/10.1152/ajpendo.1997.273.1.E99
Phillips SM, Tipton KD, Ferrando AA, Wolfe RR (1999) Resistance training reduces the acute exercise-induced increase in muscle protein turnover. Am J Phys 276:E118–E124
Proud CG (2007) Signalling to translation: how signal transduction pathways control the protein synthetic machinery. Biochem J 403:217–234. https://doi.org/10.1042/BJ20070024
Puri PL, Sartorelli V (2000) Regulation of muscle regulatory factors by DNA-binding, interacting proteins, and post-transcriptional modifications. J Cell Physiol 185:155–173. https://doi.org/10.1002/1097-4652(200011)185:2<155::AID-JCP1>3.0.CO;2-Z
Ravikumar B, Sarkar S, Davies JE, Futter M, Garcia-Arencibia M, Green-Thompson ZW, Jimenez-Sanchez M, Korolchuk VI, Lichtenberg M, Luo S, Massey DC, Menzies FM, Moreau K, Narayanan U, Renna M, Siddiqi FH, Underwood BR, Winslow AR, Rubinsztein DC (2010) Regulation of mammalian autophagy in physiology and pathophysiology. Physiol Rev 90:1383–1435. https://doi.org/10.1152/physrev.00030.2009
Rennie MJ, Wackerhage H, Spangenburg EE, Booth FW (2004) Control of the size of the human muscle mass. Annu Rev Physiol 66:799–828. https://doi.org/10.1146/annurev.physiol.66.052102.134444
Richter EA, Derave W, Wojtaszewski JF (2001) Glucose, exercise and insulin: emerging concepts. J Physiol 535:313–322
Rios R, Carneiro I, Arce VM, Devesa J (2001) Myostatin regulates cell survival during C2C12 myogenesis. Biochem Biophys Res Commun 280:561–566. https://doi.org/10.1006/bbrc.2000.4159
Roig M, O’Brien K, Kirk G, Murray R, McKinnon P, Shadgan B, Reid WD (2009) The effects of eccentric versus concentric resistance training on muscle strength and mass in healthy adults: a systematic review with meta-analysis. Br J Sports Med 43:556–568. https://doi.org/10.1136/bjsm.2008.051417
Roth SM, Martel GF, Ivey FM, Lemmer JT, Tracy BL, Metter EJ, Hurley BF, Rogers MA (2001) Skeletal muscle satellite cell characteristics in young and older men and women after heavy resistance strength training. J Gerontol A Biol Sci Med Sci 56:B240–B247
Russell AP, Lamon S (2015) Exercise, skeletal muscle and circulating microRNAs. Prog Mol Biol Transl Sci 135:471–496. https://doi.org/10.1016/bs.pmbts.2015.07.018
Russell AP, Lamon S, Boon H, Wada S, Guller I, Brown EL, Chibalin AV, Zierath JR, Snow RJ, Stepto N, Wadley GD, Akimoto T (2013) Regulation of miRNAs in human skeletal muscle following acute endurance exercise and short-term endurance training. J Physiol 591:4637–4653. https://doi.org/10.1113/jphysiol.2013.255695
Safdar A, Abadi A, Akhtar M, Hettinga BP, Tarnopolsky MA (2009) miRNA in the regulation of skeletal muscle adaptation to acute endurance exercise in C57Bl/6J male mice. PLoS One 4:e5610. https://doi.org/10.1371/journal.pone.0005610
Sandri M (2013) Protein breakdown in muscle wasting: role of autophagy-lysosome and ubiquitin-proteasome. Int J Biochem Cell Biol 45:2121–2129. https://doi.org/10.1016/j.biocel.2013.04.023
Schlaepfer DD, Hauck CR, Sieg DJ (1999) Signaling through focal adhesion kinase. Prog Biophys Mol Biol 71:435–478
Schultz E, McCormick KM (1994) Skeletal muscle satellite cells. Rev Physiol Biochem Pharmacol 123:213–257
Schwalm C, Jamart C, Benoit N, Naslain D, Premont C, Prevet J, Van Thienen R, Deldicque L, Francaux M (2015) Activation of autophagy in human skeletal muscle is dependent on exercise intensity and AMPK activation. FASEB J 29:3515–3526. https://doi.org/10.1096/fj.14-267187
Seaborne RA, Strauss J, Cocks M, Shepherd S, O’Brien TD, van Someren KA, Bell PG, Murgatroyd C, Morton JP, Stewart CE, Sharples AP (2018) Human skeletal muscle possesses an epigenetic memory of hypertrophy. Sci Rep 8:1898. https://doi.org/10.1038/s41598-018-20287-3
Shamim B, Hawley JA, Camera DM (2018) Protein availability and satellite cell dynamics in skeletal muscle. Sports Med 48:1329–1343. https://doi.org/10.1007/s40279-018-0883-7
Sheffield-Moore M, Yeckel CW, Volpi E, Wolf SE, Morio B, Chinkes DL, Paddon-Jones D, Wolfe RR (2004) Postexercise protein metabolism in older and younger men following moderate-intensity aerobic exercise. Am J Physiol Endocrinol Metab 287:E513–E522. https://doi.org/10.1152/ajpendo.00334.2003
Spangenburg EE, Le Roith D, Ward CW, Bodine SC (2008) A functional insulin-like growth factor receptor is not necessary for load-induced skeletal muscle hypertrophy. J Physiol 586:283–291. https://doi.org/10.1113/jphysiol.2007.141507
Stec MJ, Kelly NA, Many GM, Windham ST, Tuggle SC, Bamman MM (2016) Ribosome biogenesis may augment resistance training-induced myofiber hypertrophy and is required for myotube growth in vitro. Am J Physiol Endocrinol Metab 310:E652–E661. https://doi.org/10.1152/ajpendo.00486.2015
Stefanetti RJ, Lamon S, Wallace M, Vendelbo MH, Russell AP, Vissing K (2015) Regulation of ubiquitin proteasome pathway molecular markers in response to endurance and resistance exercise and training. Pflugers Arch 467:1523–1537. https://doi.org/10.1007/s00424-014-1587-y
Stokes T, Hector AJ, Morton RW, McGlory C, Phillips SM (2018) Recent perspectives regarding the role of dietary protein for the promotion of muscle hypertrophy with resistance exercise training. Nutrients 10. https://doi.org/10.3390/nu10020180
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403:41–45. https://doi.org/10.1038/47412
Szewczyk NJ, Jacobson LA (2005) Signal-transduction networks and the regulation of muscle protein degradation. Int J Biochem Cell Biol 37:1997–2011. https://doi.org/10.1016/j.biocel.2005.02.020
Tang G (2005) siRNA and miRNA: an insight into RISCs. Trends Biochem Sci 30:106–114. https://doi.org/10.1016/j.tibs.2004.12.007
Ten Broek RW, Grefte S, Von den Hoff JW (2010) Regulatory factors and cell populations involved in skeletal muscle regeneration. J Cell Physiol 224:7–16. doi:https://doi.org/10.1002/jcp.22127
Thomas M, Langley B, Berry C, Sharma M, Kirk S, Bass J, Kambadur R (2000) Myostatin, a negative regulator of muscle growth, functions by inhibiting myoblast proliferation. J Biol Chem 275:40235–40243. https://doi.org/10.1074/jbc.M004356200
Timmons JA, Larsson O, Jansson E, Fischer H, Gustafsson T, Greenhaff PL, Ridden J, Rachman J, Peyrard-Janvid M, Wahlestedt C, Sundberg CJ (2005) Human muscle gene expression responses to endurance training provide a novel perspective on Duchenne muscular dystrophy. FASEB J 19:750–760. https://doi.org/10.1096/fj.04-1980com
Tipton KD, Ferrando AA, Phillips SM, Doyle D Jr, Wolfe RR (1999) Postexercise net protein synthesis in human muscle from orally administered amino acids. Am J Phys 276:E628–E634
Tipton KD, Ferrando AA, Williams BD, Wolfe RR (1996) Muscle protein metabolism in female swimmers after a combination of resistance and endurance exercise. J Appl Physiol (1985) 81:2034–2038. https://doi.org/10.1152/jappl.1996.81.5.2034
Turinsky J, Damrau-Abney A (1999) Akt kinases and 2-deoxyglucose uptake in rat skeletal muscles in vivo: study with insulin and exercise. Am J Phys 276:R277–R282
van Rooij E, Quiat D, Johnson BA, Sutherland LB, Qi X, Richardson JA, Kelm RJ Jr, Olson EN (2009) A family of microRNAs encoded by myosin genes governs myosin expression and muscle performance. Dev Cell 17:662–673. https://doi.org/10.1016/j.devcel.2009.10.013
Walsh FS, Celeste AJ (2005) Myostatin: a modulator of skeletal-muscle stem cells. Biochem Soc Trans 33:1513–1517. https://doi.org/10.1042/BST20051513
Wang XH, Zhang L, Mitch WE, LeDoux JM, Hu J, Du J (2010) Caspase-3 cleaves specific 19 S proteasome subunits in skeletal muscle stimulating proteasome activity. J Biol Chem 285:21249–21257. https://doi.org/10.1074/jbc.M109.041707
Watson K, Baar K (2014) mTOR and the health benefits of exercise. Semin Cell Dev Biol 36:130–139. https://doi.org/10.1016/j.semcdb.2014.08.013
Wei L, Wang L, Carson JA, Agan JE, Imanaka-Yoshida K, Schwartz RJ (2001) beta1 integrin and organized actin filaments facilitate cardiomyocyte-specific RhoA-dependent activation of the skeletal alpha-actin promoter. FASEB J 15:785–796. https://doi.org/10.1096/fj.00-026com
West DW, Burd NA, Tang JE, Moore DR, Staples AW, Holwerda AM, Baker SK, Phillips SM (2010) Elevations in ostensibly anabolic hormones with resistance exercise enhance neither training-induced muscle hypertrophy nor strength of the elbow flexors. J Appl Physiol (1985) 108:60–67. https://doi.org/10.1152/japplphysiol.01147.2009
West DW, Kujbida GW, Moore DR, Atherton P, Burd NA, Padzik JP, De Lisio M, Tang JE, Parise G, Rennie MJ, Baker SK, Phillips SM (2009) Resistance exercise-induced increases in putative anabolic hormones do not enhance muscle protein synthesis or intracellular signalling in young men. J Physiol 587:5239–5247. https://doi.org/10.1113/jphysiol.2009.177220
Widegren U, Wretman C, Lionikas A, Hedin G, Henriksson J (2000) Influence of exercise intensity on ERK/MAP kinase signalling in human skeletal muscle. Pflugers Arch 441:317–322
Wilborn CD, Taylor LW, Greenwood M, Kreider RB, Willoughby DS (2009) Effects of different intensities of resistance exercise on regulators of myogenesis. J Strength Cond Res 23:2179–2187. https://doi.org/10.1519/JSC.0b013e3181bab493
Wilkinson SB, Phillips SM, Atherton PJ, Patel R, Yarasheski KE, Tarnopolsky MA, Rennie MJ (2008) Differential effects of resistance and endurance exercise in the fed state on signalling molecule phosphorylation and protein synthesis in human muscle. J Physiol 586:3701–3717. https://doi.org/10.1113/jphysiol.2008.153916
Winder WW, Hardie DG (1999) AMP-activated protein kinase, a metabolic master switch: possible roles in type 2 diabetes. Am J Phys 277:E1–E10
Wolfe RR (2000) Protein supplements and exercise. Am J Clin Nutr 72:551S–557S. https://doi.org/10.1093/ajcn/72.2.551S
Wong TS, Booth FW (1990) Protein metabolism in rat gastrocnemius muscle after stimulated chronic concentric exercise. J Appl Physiol (1985) 69:1709–1717. https://doi.org/10.1152/jappl.1990.69.5.1709
Wong TS, Booth FW (1990) Protein metabolism in rat tibialis anterior muscle after stimulated chronic eccentric exercise. J Appl Physiol (1985) 69:1718–1724. https://doi.org/10.1152/jappl.1990.69.5.1718
Woodgett JR (1989) Early gene induction by growth factors. Br Med Bull 45:529–540
Yamamoto H, Morino K, Nishio Y, Ugi S, Yoshizaki T, Kashiwagi A, Maegawa H (2012) MicroRNA-494 regulates mitochondrial biogenesis in skeletal muscle through mitochondrial transcription factor A and Forkhead box j3. Am J Physiol Endocrinol Metab 303:E1419–E1427. https://doi.org/10.1152/ajpendo.00097.2012
Yang SY, Goldspink G (2002) Different roles of the IGF-I Ec peptide (MGF) and mature IGF-I in myoblast proliferation and differentiation. FEBS Lett 522:156–160
Zeman RJ, Kameyama T, Matsumoto K, Bernstein P, Etlinger JD (1985) Regulation of protein degradation in muscle by calcium. Evidence for enhanced nonlysosomal proteolysis associated with elevated cytosolic calcium. J Biol Chem 260:13619–13624
Funding
M.F. was supported by the Sports Ministry of the Brussels-Wallonia Federation. This work was supported by the Fonds Scientifique de la Recherche (FSR) from the Université catholique de Louvain and by the Fonds National de la Recherche Scientifique (FNRS, F.4504.17).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
This article is part of the special issue on Exercise Physiology: Future Opportunities and Challenges in Pflügers Archiv – European Journal of Physiology
Rights and permissions
About this article
Cite this article
Francaux, M., Deldicque, L. Exercise and the control of muscle mass in human. Pflugers Arch - Eur J Physiol 471, 397–411 (2019). https://doi.org/10.1007/s00424-018-2217-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00424-018-2217-x