Abstract
Nucleoli are formed on the basis of ribosomal genes coding for RNAs of ribosomal particles, but also include a great variety of other DNA regions. In this article, we discuss the characteristics of ribosomal DNA: the structure of the rDNA locus, complex organization and functions of the intergenic spacer, multiplicity of gene copies in one cell, selective silencing of genes and whole gene clusters, relation to components of nucleolar ultrastructure, specific problems associated with replication. We also review current data on the role of non-ribosomal DNA in the organization and function of nucleoli. Finally, we discuss probable causes preventing efficient visualization of DNA in nucleoli.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Nucleoli have been known since the works of Wagner (1835) and Valentin (1836) as the most conspicuous components of cell nucleus. A century later, it was found that these nuclear bodies are assembled around certain chromosomal loci, termed afterward “nucleolus organizer regions (NORs)” (Heitz 1931; McClintock 1934). Subsequent findings indicated that structure and the main function of nucleoli are based upon transcription of ribosomal genes. In recent studies, multiple other genomic regions have been found within and closely adjacent to the nucleoli. This review is focused on peculiarities of ribosomal DNA and on the role of non-ribosomal DNA sequences in organization and function of nucleoli.
Ribosomal DNA (rDNA)
rDNA locus
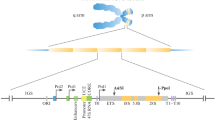
Ribosomal DNA is responsible for production of ribosomal RNAs. Thus, in mammalian cells, there are three kinds of coding regions which produce 18S RNA of the small ribosomal subunit, as well as 28S and 5.8S RNAs of the large ribosomal subunit (Gonzalez and Sylvester 1995). Accordingly, each transcription unit includes three genes, separated by internal transcribed spacers, ITS1 and ITS2, and flanked by external spacers, 5′ ETS and 3′ ETS (Fig. 1). These spacers vary significantly in composition and size in different species (reviewed in Nazar 2004).
Organization of rDNA in mammalian cell. NOR nucleolus organizer region, the cluster of rDNA loci, NTS (IGS) non-transcribed (intergenic) spacer, 5′-ETS, 3′-ETS external transcribed spacers, ITS1, 2 internal transcribed spacers, UCE/CPE promoter including upstream control element (UCE) and core promoter element (CPE), Sal Sal box, the sequence serving as transcription terminator, En enhancer
The transcription units of rDNA locus are separated from each other by non-transcribed, or intergenic, spacers, NTS or IGS (Fig. 1). In yeast and infusoria, the spacers are rather uniform (Philippsen et al. 1978; Wild and Gall 1979). But generally, structure and length of the IGS vary to a large extent not only from species to species, but also within the species, and even within a single individual (Lewin 1980; Wellauer and Dawid 1977; Reeder et al. 1976). The satellite contents of the DNA are particularly variable (Gonzalez et al. 1992a; Gonzalez and Sylvester 1995, 2001; Maden et al. 1987; Sasaki et al. 1987). Most frequently, the small repetitive arrays get increased or reduced in number as a result of slipped-strand mispairing and other errors associated with replication (Tautz et al. 1986; Levinson and Gutman 1987). Human IGS includes various kinds of DNA repeats, both tandemly and non-tandemly arranged; they include simple sequence motifs, microsatellites (2–6 bp), long repeats (cca 2 kb), as well as transposable elements. Among the latter, Alu, belonging to the short interspersed elements (SINE), are predominant (Tautz et al. 1986; Gonzalez et al. 1989; Gonzalez and Sylvester 1995). Some of rDNA repeats also contain nucleotide substitutions, supplementary microsatellite clusters, and, more seldom, extended deletions (Ryskov et al. 1993; Braga et al. 1995; Kupriyanova et al. 2015).
It has long been known that the rDNA spacers include regulatory sequences. Thus, the promoter of each ribosomal gene is partly or entirely situated in IGS, upstream of the start codon. Typically, it consists of two parts: core domain and upstream control domain (Haltiner et al. 1986; Clos et al. 1986; Paule 1994; Reeder 1992; Doelling and Pikaard 1995). Another essential part of the spacer is the terminator. It may be absent in rDNA of Xenopus and Drosophila, and in these cases, the transcription is ceased by a kind of processing (Labhart and Reeder 1986; McStay and Reeder 1986; Tautz and Dover 1986; De Winter and Moss 1986). In mammalian cells, transcription of rDNA is terminated at the 3′ end of each gene, at a sequence motif called “Sal box” with the length of 18 bp in mouse and 11 bp in human (Grummt et al. 1985; Kuhn et al. 1988). Sal box binds the TTF-1 (transcription termination factor 1) protein, which is essential for arresting RNA polymerase I (pol I) (Grummt et al. 1985, 1986; La Volpe et al. 1985; Bartsch et al. 1987; Pfleiderer et al. 1990; Diermeier et al. 2013; reviewed in Németh et al. 2013). But it seems that one such site is not sufficient, for each human or murine rDNA unit is provided with 10 terminators, T 1–10 (reviewed in Diermeier et al. 2013).
Other sites regulating the expression of rDNA have been found in the non-transcribed spacers. A usual component of IGS is the enhancer; enhancers appear as clusters of repeating sequences distanced from the regulated region (Pikaard et al. 1990; Moss et al. 1985). In mammalian cells, there are also one or more reduced transcription units situated ~2 kb upstream of the core promoter. Such units include spacer promoter, spacer terminator, and a minigene producing transcripts of ~150 bp, termed promoter-associated RNAs (pRNAs), which are involved in rDNA silencing (Sylvester et al. 2003; Mayer et al. 2006, 2008; Moss et al. 2007; McStay and Grummt 2008; Santoro et al. 2010; Anosova et al. 2015).
Until recently, extensive IGS regions of mammalian cells had been regarded as receptacles of useless, “junk” sequences. But this idea will probably follow the fate of the general “junk DNA” theory. Remarkably, RNA-seq analysis of human and murine rDNA revealed a specific pattern of low-abundance expression over the entire IGS region (Zentner et al. 2011, 2014; reviewed by Jacob et al. 2012), indicating that the spacer has a complex functional organization. The same idea is suggested by the data of Chip and Chip-seq analysis which show a regular distribution of pol I and its transcription factors throughout the intergenic region (Copenhaver et al. 1994; Hu et al. 1994; O’Sullivan et al. 2002; Zentner et al. 2011, 2014). A special role belongs to the IGS in the recent theory of “nucleolar detention,” according to which nucleoli may serve as a kind of lumber room for useless proteins and other cell components (Audas et al. 2012a). The process is triggered when RNAs produced by loci situated within the IGS recognize and “capture” proteins furnished with a “detention signal” (Audas et al. 2012a, b; Jacob et al. 2012; Diermeier et al. 2013; Padeken and Heun 2014; reviewed in Lam and Trinkle-Mulcahy 2015).
Repetitive arrays
The presence of numerous ribosomal gene copies in one cell is one of the most remarkable characteristics of rDNA. There are exceptions to this rule. For instance, some species of Protozoa and Myxomycetes have only one chromosomal gene which is amplified into a set of extrachromosomal inverted repeats in the course of somatic growth (Lewin 1980). But generally, metazoan genomes contain several hundred ribosomal gene copies (Birnstiel et al. 1971); in plants, this number often reaches several thousands (Rogers and Bendich 1987).
Usually, rDNA is arranged in clusters of tandem repeats, nucleolus organizer regions (NORs). These regions were first discovered as secondary constrictions of mitotic chromosomes (Heitz 1931; McClintock 1934). But later, it was found that some of the NORs make no secondary constrictions; such clusters are transcriptionally silent and may appear both within and without nucleoli (Sullivan et al. 2001; Strohner et al. 2001; Kalmárová et al. 2007).
In certain cases, rDNA is organized as inverted repeats (Bergold et al. 1983). In amphibia (Birnstiel et al. 1971; Bird 1978), insects (Birnstiel et al. 1971), and fungi (Butler and Metzenberg 1993), ribosomal genes are amplified into numerous extrachromosomal copies (reviewed in Moss and Stefanovsky 1995). Human diploid genome contains about 400–600 copies of a 43-kbp unit (Moss et al. 2006; Stults et al. 2009). Human NORs with an average size of 3 Mbp are situated on the short arms of the acrocentric chromosomes 13, 14, 15, 21, and 22 (Henderson et al. 1972; Long and Dawid 1980; Puvion-Dutilleul et al. 1991).
The abundance of ribosomal gene repeats not only enables the cell to regulate the production of ribosomal RNA more efficiently, but also increases the frequency of recombination. The number of repeats varies as a result of unequal homologous exchange, and this may cause damage to the cell (La Volpe et al. 1984; Mroczka et al. 1984; Erickson and Schmickel 1985; Sylvester et al. 1986, 1989; Cassidy et al. 1986; Dumenco and Wejksnora 1986; Tower et al. 1989; Stults et al. 2009). The partial silencing of rDNA seems to be an important factor in maintaining stability of the loci (Peng and Karpen 2007). Gene conversion is regarded as an additional stabilizing process, since it reduces the variability (Gonzalez and Sylvester 1995; Elder and Turner 1995). Tandem arrangement of rDNA increases the risk of inappropriate transcription; therefore, isolation of each repeat from its neighbors on the DNA strand seems necessary. Such demarcating function is ascribed to insulators (Valenzuela and Kamakaka 2006). Association of CTCF protein with human and murine rDNA at the spacer promoter region suggests the presence of an insulator element here (Torrano et al. 2006; van de Nobelen et al. 2010; Zentner et al. 2011). Remarkably, CTCF depletion leads to disorganization of nucleolar structure and overexpression of ribosomal genes (Hernández-Hernández et al. 2012).
Studies of restriction products show that the repeats within NORs vary in length and structure (Kominami et al. 1981; Gonzalez et al. 1985, 1990; Maden et al. 1987; Sasaki et al. 1987). Variants with tissue-specific expression were found among murine rDNA repeats (Tseng et al. 2008). In situ hybridization on the preparation of isolated DNA fibers, “molecular combing” (Bensimon et al. 1994; Michalet et al. 1997; Anglana et al. 2003; Caburet et al. 2005; Tseng et al. 2008) revealed high percentage of non-canonical, including palindromic, sequences (about one-third of the repeats), and great variability in the length of IGS (from 10 to 50 kb) in several types of human cells (Lebofsky and Bensimon 2005; Caburet et al. 2005). It seems that the variability in the length of NORs provides each person with a unique rDNA electrophoretic karyotype, a kind of “fingerprints” (Stults et al. 2008).
The palindromic structures may cause fork stalling and/or arrest by forming hairpin structures during lagging-strand synthesis, which apparently results in significant slowing down of rDNA replication. Thus, in HeLa cells, the average speed of replication fork for the whole genome is 1.7 µm/min, but only 1 µm/min for rDNA (Lebofsky and Bensimon 2005). If the palindromes are pseudogenes (Caburet et al. 2005), they must be non-functional by definition (Mighell et al. 2000). But it is still unknown whether the length and composition of IGS have any impact on transcription of the adjacent genes.
The multiplicity and high sequence similarity of rDNA repeats greatly hinder their study. For that reason, NORs were excluded from the initial sequencing and analysis of the human genome.
Active and silent rDNA
It is typical for the clustered rDNA that its transcriptionally active genes are interspersed by transcriptionally silent repeats (Conconi et al. 1989; Santoro 2005, 2014; Zillner et al. 2015). The active genes are characterized by hypomethylation of CpG sites and histone modifications generally associated with transcriptionally active nucleoplasmic chromatin (i.e., H3K4me3 and H3K9ac), whereas transcriptionally silent rDNA is condensed, hypermethylated, and marked with repressive histone modifications (i.e., H3K27me3 and H4K20me3) (Heintzman et al. 2007; McKeown and Shaw 2009; Zentner et al. 2011; Zillner et al. 2013; reviewed in McStay and Grummt 2008; Shaw and McKeown 2011). Key role in the silencing scheme belongs to the nucleolar remodeling complex (NoRC) (Strohner et al. 2001). Targeting NoRC to rDNA leads to repositioning of a promoter-bound nucleosome, changes in histone modifications, increase in DNA methylation, and silencing of rRNA genes (Zhou and Grummt 2005; Li et al. 2006; Mayer et al. 2006, 2008; Schmitz et al. 2010; Anosova et al. 2015). On the other hand, nucleosome remodeling and deacetylation complex (NuRD) creates chromatin state in which rDNA is poised for transcription, though not yet transcribed (Xie et al. 2012).
It has been established that the active ribosomal genes form loops in which a promoter is joined to the terminator. Transcription termination factor 1 (TTF-1) and protooncogene c-Myc seem to be particularly important for this connection (Németh and Längst 2008; Pontvianne et al. 2013; Li and Hann 2013). Both proteins regulate the association of epigenetically activated rDNA genes with the nucleolar matrix (Shiue et al. 2014). TTF-1 binds to an upstream site, termed T 0, located 170 bp upstream of the transcription start site (Clos et al. 1986). This is required for efficient transcription initiation and for the recruitment of chromatin remodeling complexes that establish distinct epigenetic states of rRNA genes. Interaction of TTF-1 with CSB (Cockayne Syndrome protein B), NoRC, or NuRD leads to the establishment of active, silent, or poised state of chromatin, respectively (Strohner et al. 2001; Santoro et al. 2002; Yuan et al. 2007; Xie et al. 2012; Diermeier et al. 2013).
In steadily cycling cells, chromatin structure of ribosomal genes is maintained through multiple rounds of cell division (e.g., Li et al. 2006, reviewed in Birch and Zomerdijk 2008; Santoro and De Lucia 2005; Guetg et al. 2012). From prophase to late anaphase, the gene activity is efficiently blocked by cdc2/cyclin B-directed phosphorylation of SL-1 and other transcription factors (Heix et al. 1998; Voit et al. 2015). Nevertheless, the components of pol I transcription machinery, including the upstream binding factor (UBF) and promoter selectivity complex (SL1), can be detected on certain NORs even in metaphase (Babu and Verma 1985; Moss et al. 1985; Weisenberger and Scheer 1995; Jordan et al. 1996; Roussel and Hernandez-Verdun 1994; Roussel et al. 1996; Gebrane-Younes et al. 1997; Sirri et al. 1999, 2008; O’Sullivan et al. 2002; Leung et al. 2004; Prieto and McStay 2005). Such NORs, termed “transcriptionally competent” or just “competent” (Dousset et al. 2000; Savino et al. 2001), are transcribed, while the other, “non-competent” NORs remain silent during interphase (Weisenberger and Scheer 1995; Roussel et al. 1996; Gebrane-Younes et al. 1997). The competence, which can be revealed by UBF or silver nitrate staining, is regularly distributed among the different chromosomes (Héliot et al. 2000; Smirnov et al. 2006). After S phase, some NORs may become “asymmetrical,” when only one of the daughter chromatids acquires the competence signal. The presence of such NORs causes mitotic asymmetry (Kalmárová et al. 2008).
Organization of rDNA in the nucleolus
Structure of nucleoli is based upon transcriptionally active rDNA (Henderson et al. 1972; Long and Dawid 1980; Puvion-Dutilleul et al. 1991; Raska 2003; Raska et al. 2006a, b; Cmarko et al. 2008; Sirri et al. 2008). Crude versions of that structure appear on ectopical loci in the form of “pseudo-NORs” or “neo-NORs” produced experimentally on the basis of simple UBF binding arrays (Mais et al. 2005; Prieto and McStay 2007; Grob et al. 2014).
It has been known that nucleoli are usually formed at the end of mitosis around competent NORs, which gradually unfold and fuse into a few bodies. But the organization of rDNA in the interphase is still not understood. On the one hand, hypotonically isolated and spread ribosomal genes appear as so-called Christmas trees, in which the “tree stem” represents a single DNA fibril, from which the transcripts grow like branches (Miller and Beatty 1969; Trendelenburg et al. 1974; Scheer and Zentgraf 1982; Trendelenburg and Puvion-Dutilleul 1987; Mougey et al. 1993; Scheer et al. 1997; Albert et al. 2011). On the other hand, electron microscopical studies show that transcription of rDNA and the first steps of rRNA processing take place in the FC/DFC units, i.e., fibrillar centers (FC) surrounded by dense fibrillar components (DFC) (Fig. 2). The transcribed part of rDNA as well as the transcription signal after pulse labeling has been observed in the DFC or at the border between DFC and FC (Raška et al. 1983a, b, 1995; Ochs et al. 1985; Raska et al. 1989, 2006a, b; Scheer and Benavente 1990; Hozák et al. 1993, Cmarko et al. 2000; Melcák et al. 1996; Koberna et al. 2002; Casafont et al. 2006; Shaw and McKeown 2011). But it proved to be very difficult to find out how the elements of “Christmas trees” are accommodated among the elements of nucleolar ultrastructure.
A schematic representation of nucleolus-associated DNA. Nu nucleolus, Np nucleoplasm, RC chromosome carrying ribosomal genes (ribosomal chromosome), Cen centromere, PR proximal flanking region, DR distal flanking region, NRC non-ribosomal chromosome, FC/DFC FC/DFC unit—the center of rDNA transcription consisting of fibrillar center (FC) surrounded with dense fibrillar component (DFC). Green dots represent granular component of the nucleolus
There are reasons to believe that each FC/DFC unit typically accommodates one transcriptionally active rDNA repeat (Haaf et al. 1991; Haaf and Ward 1996; Denissov et al. 2011), which forms multiple coils passing through DFC and adjacent FC area (Reeder and Lang 1997; Cheutin et al. 2002; Puvion-Dutilleul et al. 1991; Derenzini et al. 2006; McStay and Grummt 2008). Pictures of osmium amine staining show the presence of some DNA in the FC, but the status and composition of this DNA still have not been determined (Derenzini et al. 2006, 2014). Further studies are needed to establish the position of inactive rDNA repeats, as well as the “poised genes” (reviewed in Németh and Längst 2011). The latter, together with silent genes, may be localized in the fibrillar center but, upon activation, move toward the DFC (Raska et al. 2006b).
According to the data of chromatin capture analysis, the promoter of each active gene is joined to the respective terminator and may be close topographically to several loci of the gene (Grewal et al. 2005; Arabi et al. 2005; Gomez-Roman et al. 2006; Grandori et al. 2005; Németh et al. 2008; Németh and Längst 2008; Shiue et al. 2009, 2014; Denissov et al. 2011; Lykke-Andersen et al. 2011; Xie et al. 2012; Diermeier et al. 2013; reviewed in Németh and Längst 2011). Based on such data, and considering increased binding of promoter selectivity complex SL1 over the entire region, Denissov et al. (2011) proposed a “core-helix model.” According to it, a single ribosomal gene occupying the FC/DFC unit assumes the form of rotating cylindrical solenoid. The transcribing pol I complexes driven by actin revolve around the SL1-containing core, which is situated in the FC and serves as an anchor for both the promoter and the terminator of the rDNA repeat; the nascent rRNAs exit radially into the DFC. Remarkably, this chiefly speculative model seems to be the only hypothesis describing organization of rDNA in the interphase nucleoli. So far, it is not even known whether replication of ribosomal genes occurs within FC/DFC units or in other nucleolar structures.
Reproduction of rDNA
Replication of rDNA should be viewed in connection with two circumstances: the great number of the gene copies in the cell and the ongoing transcription, which may even intensify during S phase (Gorski et al. 2008). Accordingly, each cycling cell must have means to avoid two significant dangers. On the one hand, the multiple tandemly repeated rDNA arrays may be recombination hotspots and thus present a potential source of genomic instability (Stults et al. 2009; Ide et al. 2010). This risk is probably diminished by alternation of silenced and active repeats in each array (Santoro 2005, 2014).
On the other hand, collision of the swiftly running replication and transcription machineries (reviewed in Magdalou et al. 2014) must be prevented, which requires a special spatio-temporal arrangement of replication. Thus, in yeast cells, each rDNA repeat has one potential origin of replication, and clusters of synchronously firing origins are separated by a few units with silent origins (Pasero et al. 2002). Besides, there is an efficient fork barrier situated at the 3′ end of each transcription unit; it arrests the upstream moving of the forks, which prevents collision of replication and transcription complexes (Brewer and Fangman 1988; Linskens and Huberman 1988; reviewed in Rothstein et al. 2000).
In human cells, rDNA replication may be initiated all over the IGS and even upon the genes (Lebofsky and Bensimon 2005), though the potential origins situated upstream of the transcribed region are used more frequently (Little et al. 1993; Yoon et al. 1995; Gencheva et al. 1996; Scott et al. 1997). The replication forks may terminate and converge at variable sites throughout the rDNA repeat (Little et al. 1993). Fork barriers exist (Gerber et al. 1997; Akamatsu and Kobayashi 2015), but often fail to stop the progress of DNA polymerase complex, so that replication proceeds at far distances in both directions (Edenberg and Huberman 1975; Lebofsky and Bensimon 2005). Neighboring origins fire within 60 min of each other; the distance between them varies from tens of kilobases to a couple hundred kilobases, with an average of 80 kb (Lebofsky and Bensimon 2005). In such conditions, additional mechanisms must be engaged to protect genome stability. Separation of rRNA gene transcription and replication domains probably could make up for the relaxed timing and spacing (Pliss et al. 2005). Such separation may be achieved by regulation at the level of FC/DFC units; indeed, in vivo observations indicate that transcription is suspended in the units involved in the replication (Smirnov et al. 2014; also our new data prepared for publication).
Non-ribosomal DNA in nucleoli
Clusters of rDNA repeats, which include the transcribed and non-transcribed parts, are usually regarded as the founders of nucleoli. But this role may be shared by the regions of the same chromosomes adjacent to NORs (Gonzalez et al. 1989, 1992; Kaplan et al. 1993) (Fig. 2). In human cells, these regions have a very similar structure upon all five acrocentric chromosomes (Floutsakou et al. 2013). The proximal flanking sequences, which are positioned in the neighborhood of centromeres, consist largely of satellite DNA, frequently undergo recombination, and have numerous analogues in other parts of the genome. In contrast, the distal sequences, which are situated closer to the telomeres, exhibit low segmental duplication, but contain chromatin signature characteristic of promoters, as well as putative genes, interspersed among marks associated with heterochromatin. These sequences may regulate the activity of the NORs and participate in the structural organization of the nucleoli by anchoring rDNA to perinucleolar chromatin (Floutsakou et al. 2013).
Microscopic studies have shown that various other parts of the genome may regularly or occasionally find their way to the nucleoli. In different cell types and species, satellite DNA of centromeres is a common component of the perinucleolar shell of condensed late replicating chromatin and appears also in the interior of nucleoli (Comings 1980; Manuelidis 1984; Manuelidis and Borden 1988; Haaf and Schmid 1989, 1991; Bartholdi 1991; Billia and Deboni 1991; Ochs and Press 1992; Léger et al. 1994; Carvalho et al. 2001; Wong et al. 2007). Particularly, centromeres of chromosomes with a lower content of G-dark bands tend to be localized at the nucleolus (Carvalho et al. 2001). Telomeres, together with telomerase components (Rawlins and Shaw 1990; Vourc’h et al. 1993; Armstrong et al. 2001; Zhang et al. 2004), territories of human chromosomes 1, 9, and Y (Stahl et al. 1976; Léger et al. 1994), as well as parts of acrocentric chromosomes (Kalmárová et al. 2008; Pliss et al. 2015), are often found within or very close to nucleoli. Data of 3C analysis suggest that ribosomal genes may interact with repetitive sequences belonging to other chromosomes (O’Sullivan et al. 2009). Functional significance of this interaction is not clear.
Abundant data have been recently obtained by sequence analysis of nucleolus-associated domains (NADs), which represent the entire DNA content of isolated nucleoli (Németh et al. 2010; van Koningsbruggen et al. 2010). In the studies of human cells, it was found that NADs, not counting the ribosomal genes, constitute about 4 % of the genome and include sequences from all chromosomes. The bulk of these domains consists of AT-rich sequences, satellite repeats (mainly alpha-, beta-, GAATG/CATTC types), members of the zinc-finger, olfactory receptor defensin and immunoglobulin protein-coding gene families, transcriptionally active 5S rRNA genes, and tRNA genes (Matera et al. 1995; Thompson et al. 2003; van Koningsbruggen et al. 2010; Németh et al. 2010). Analysis of the transcriptional status and chromatin feature showed that NADs contain mainly inactive chromosomal regions (Németh and Längst 2011; van Koningsbruggen et al. 2010).
There is still an uncertainty about the composition of NADs. The often used term “nucleolar association” is somewhat ambiguous; it embraces sequences of two essentially different compartments, nucleolar interior and nucleolar periphery, since these are swept together in methods based on the isolation of nucleoli. Besides, this isolation requires breaking of NOR-bearing and perhaps some other chromosomes, which may introduce further errors. This problem has been partly solved by using in situ hybridization to confirm the data of deep sequencing (van Koningsbruggen et al. 2010).
Factors directing various DNA sequences toward the nucleolus, as well as consequences of the perinucleolar positioning, have become a focus of intensive study recently (reviewed in Padeken and Heun 2014; also reviewed in Matheson and Kaufman 2015). Localization of NADs in the perinucleolar region is correlated with heterochromatin formation and transcriptional silencing (Zhang et al. 2007; Pandey et al. 2008; Mohammad et al. 2008; Fedoriw et al. 2012; Jakociunas et al. 2013; Yang et al. 2015). Thus, the inactivating center of X chromosome (Xic) is associated with nucleoli (Zhang et al. 2007). Deletion of Xic locus reduces this association (Csankovszki et al. 2001). Remarkably, admittance to the perinucleolar region is not guaranteed by the DNA sequence, for only inactive X chromosome is found among the NADs (Zhang et al. 2007).
The proteins involved in NAD localization often also regulate rDNA transcription and/or nucleolar structure. For example, CTCF (Yusufzai et al. 2004; van de Nobelen et al. 2010; Huang et al. 2013), NCL (Roger et al. 2003; Rickards et al. 2007; Cong et al. 2012), NPM1 (Murano et al. 2008), and Ki-67 (Rahmanzadeh et al. 2007; Booth et al. 2014) all regulate transcription of ribosomal genes. Likewise, depletion of modulo (NCL) in flies disrupts nucleolar structure as demonstrated by immunofluorescence (Padeken et al. 2013), and after depletion of Ki-67 in human cells, their nucleoli become fewer and smaller (Booth et al. 2014).
Long noncoding RNAs (lncRNAs) seem to be essential for regulation of structure and function of the perinucleolar region (Mohammad et al. 2008; Jacob et al. 2013; Padeken and Heun 2014; Matheson and Kaufman 2015). One of such lncRNAs produced by a locus situated on inactive X (Xi) chromosome was named Firre (Yang et al. 2010). Firre is required for normal perinucleolar positioning of the mouse Xi (Yang et al. 2015). This protein binds CTCF, which may regulate both the silencing of Xi and its association with nucleolus (Hacisuleyman et al. 2014; Yang et al. 2015).
By attracting various segments of chromatin, constraining their mobility, and removing them from the transcriptionally active environment, nucleoli, together with nuclear periphery, play an essential part in the dynamic organization of the genome (Berger et al. 2008; Matheson and Kaufman 2015). Clustering around nucleoli might contribute to a more stable positioning of the DNA elements (Padeken and Heun 2014). Experiments with late replication labeling (Cremer and Cremer 2001) and GFP-tagged histones (van Koningsbruggen et al. 2010) indicate that after mitosis, perinucleolar chromatin partly returns to the nucleoli and partly moves to the nuclear lamina. Thus, composition of the perinucleolar region shows a degree of stability, though it may exchange components with lamina-associated domains (LADs) (van Koningsbruggen et al. 2010; Németh and Längst 2011; Kind et al. 2013).
“Invisibility” of nucleolar DNA
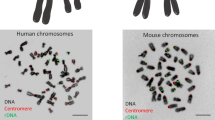
It can be seen from the aforesaid that nucleoli contain different sorts of DNA in both loose and condensed states (Fig. 2). But for some reason, we do not observe in nucleoli the alternation of dense and sparse chromatin foci which is typical for nucleoplasm. Moreover, it is well known that on preparations of cells stained with DAPI or other DNA markers, nucleoli usually appear as dark holes (Fig. 3). The intranucleolar signal is extremely weak, though exceeding the background. We find a similar pattern in distribution of various histones revealed by antibodies or as GFP constructs (e.g., Müller et al. 2007). This shows a striking contrast with results of in situ hybridization staining of transcribed and non-transcribed rDNA (e.g., Junera et al. 1995) or immunostaining for such DNA-binding proteins as UBF or TTF-1. Replication signal observed after incorporation of various nucleotides also has an amazingly low intensity within nucleoli (e.g., O’Keefe et al. 1992), especially during early S phase, when transcriptionally active ribosomal genes are replicated. Dimitrova (2011) attempted to explain this phenomenon by suggesting that certain parts of rDNA may leave nucleoli, get replicated in nucleoplasm or at nucleolar periphery, and afterward return to their former positions. But study of incorporated nucleotides in vivo (Smirnov et al. 2014) revealed no significant exchange of DNA between nucleoli and nucleoplasm or nucleolar periphery.
Even electron microscopy studies fail to clarify the matter. Thus, osmium amine reaction on ultrathin sections (Derenzini et al. 2006) reveals patches of condensed DNA in nucleoli, comparable with those in nucleoplasm. But only a pale homogenous staining appears in the area of fibrillary centers; the supposedly coiled DNA of active ribosomal genes is scarcely detected.
Bearing these data in mind, we will consider the following hypotheses:
-
1.
Special biophysical properties of nucleoli.
Since the intranucleolar space is characterized by extremely crowded condition, all processes in it are strongly influenced by short-range entropic forces which compel macromolecules to “crystallize” into nanostructures (Hancock 2014). One consequence of this may be redistribution of electric charges and low permeability of the nucleolus for DNA staining reagents, as well as certain antibodies, GFP constructs, etc. This hypothesis appears to be the easiest to verify experimentally.
-
2.
Peculiar structure of chromatin in the nucleolus.
The state of rDNA chromatin may be “unusual.” At any rate, standard ChIP protocol proves to be rather inefficient when applied to nucleolar components (Prieto and McStay 2008). This hypothesis, as well as the previous one, suggests that DNA is present in nucleoli in sufficient quantity, but for some reason eludes detection.
-
3.
Extremely low concentration of DNA in the nucleolus.
It would be the simplest explanation of the phenomenon: There is just too little DNA to be detected. But does this view agree with the facts? For one thing, DNA is not homogeneously distributed in the nucleolar volume, but forms foci of variable density, which may be seen on the electronograms. On the other hand, the average concentration of DNA in the nucleoli does not seem to be particularly low. As we have seen, cca 4 % of human genome belong to NADs. Although the larger part of this may be represented by DNA sequences in the perinucleolar region (Fig. 2), NORs alone constitute roughly 1 % of human genome (about 30 Mb) (Németh and Längst 2011). To this, we must add those non-ribosomal NADs which are known to be intranucleolar, e.g., Alu, Kpn elements, pentameric arrays of chromosome 15 (Kaplan et al. 1993), centromeres (Ochs and Press 1992), together with the adjacent regions. “Core nucleoli” obtained from isolated nuclei by centrifugation and extensive nuclease treatment are deprived of ribosomal genes, yet still contain about 1 % of the total nuclear DNA (Bolla et al. 1985). Thus, nucleoli will claim about 2 % of human genome. But how much is the volume occupied by this DNA? Since mammalian cells typically contain 2–3 nucleoli with average diameter of 1–3 μm (e.g., Smetana et al. 2006), and average diameter of the nucleus in such cells is about 6 μm (Alberts et al. 2002), the ratio of nucleolar volume to the volume of the nucleus lies between 4 and 40 %. These estimates indicate that mean value of DNA concentration in nucleoplasm is generally higher than in nucleoli. Nevertheless, the differences are not so great as to account for the extraordinary low intensity of DAPI or replication labeling.
Thus, none of the three examined hypotheses seems conclusive. Perhaps, their combination will provide a solution in the future. But for the present, the invisibility of DNA in the nucleolus still remains a riddle.
References
Akamatsu Y, Kobayashi T (2015) The human RNA polymerase I transcription terminator complex acts as a replication fork barrier that coordinates the progress of replication with rRNA transcription activity. Mol Cell Biol 35:1871–1881
Albert B, Léger-Silvestre I, Normand C, Ostermaier MK, Pérez-Fernández J, Panov KI, Zomerdijk JC, Schultz P, Gadal O (2011) RNA polymerase I-specific subunits promote polymerase clustering to enhance the rRNA gene transcription cycle. J Cell Biol 192:277–293
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002) DNA and chromosomes. In: Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (eds) Molecular biology of the cell, 4th edn. Garland Science, New York, pp 191–234
Anglana M, Apiou F, Bensimon A, Debatisse M (2003) Dynamics of DNA replication in mammalian somatic cells: nucleotide pool modulates origin choice and interorigin spacing. Cell 114:385–394
Anosova I, Melnik S, Tripsianes K, Kateb F, Grummt I, Sattler M (2015) A novel RNA binding surface of the TAM domain of TIP5/BAZ2A mediates epigenetic regulation of rRNA genes. Nucl Acids Res 43:5208–5220
Arabi A, Wu S, Ridderstrale K, Bierhoff H, Shue C, Fatyol K, Fahlen S, Hydbring P, Soderberg O, Grummt I, Larsson LG, Wright A (2005) c-Myc associates with ribosomal DNA and activates RNA polymerase I transcription. Nat Cell Biol 7:303–310
Armstrong SJ, Franklin FC, Jones GH (2001) Nucleolus-associated telomere clustering and pairing precede meiotic chromosome synapsis in Arabidopsis thaliana. J Cell Sci 114:4207–4217
Audas TE, Jacob MD, Lee S (2012a) The nucleolar detention pathway: a cellular strategy for regulating molecular networks. Cell Cycle 11:2059–2062
Audas TE, Jacob MD, Lee S (2012b) Immobilization of proteins in the nucleolus by ribosomal intergenic spacer noncoding RNA. Mol Cell 45:147–157
Babu KA, Verma RS (1985) Structural and functional aspects of nucleolar organizer regions (NORs) of human chromosomes. Int Rev Cytol 94:151–176
Bartholdi MF (1991) Nuclear distribution of centromeres during the cell cycle of human diploid fibroblasts. J Cell Sci 99:255–263
Bartsch I, Schoneberg C, Grummt I (1987) Evolutionary changes of sequences and factors that direct transcription termination of human and mouse ribosomal genes. Mol Cell Biol 7:2521–2529
Bensimon A, Simon A, Chiffaudel A, Croquette V, Heslot F, Bensimon D (1994) Alignment and sensitive detection of DNA by a moving interface. Science 265:2096–2098
Berger AB, Cabal GG, Fabre E, Duong T, Buc H, Nehrbass U, Olivo-Marin JC, Gadal O, Zimmer C (2008) High-resolution statistical mapping reveals gene territories in live yeast. Nat Methods 5:1031–1037
Bergold PJ, Campbell GR, Littau VC, Johnson EM (1983) Sequence and hairpin structure of an inverted repeat series at termini of the Physarum extrachromosomal rDNA molecule. Cell 32:1287–1299
Billia F, Deboni U (1991) Localization of centromeric satellite and telomeric DNA sequences in dorsal root ganglion neurons, in vitro. J Cell Sci 100:219–226
Birch JL, Zomerdijk JC (2008) Structure and function of ribosomal RNA gene chromatin. Biochem Soc Trans 36(Pt 4):619–624. doi:10.1042/BST0360619
Bird AP (1978) A study of early events in ribosomal gene amplification. Cold Spring Harb Symp Quant Biol 42:1179–1183
Birnstiel ML, Chipchase M, Speirs J (1971) The ribosomal RNA cistrons. Prog Nucl Acid Res Mol Biol 11:351
Bolla RI, Braaten DC, Shiomi Y, Hebert MB, Schlessinger D (1985) Localization of specific rDNA spacer sequences to the mouse L-cell nucleolar matrix. Mol Cell Biol 5:1287–1294
Booth DG, Takagi M, Sanchez-Pulido L, Petfalski E, Vargiu G, Samejima K, Imamoto N, Ponting CP, Tollervey D, Earnshaw WC, Vagnarelli P (2014) Ki-67 is a PP1-interacting protein that organises the mitotic chromosome periphery. Elife 27(3):e01641. doi:10.7554/eLife.01641
Braga EA, Kapanadze B, Kupriyanova NS, Ivanova GM, Brodyanskii VM, Nechvolodov KK, Skutov GA, Ryskov AP, Nosikov NN, Yankovskii NK (1995) Distribution analysis of 7 microsatellite motifs in cosmids of human chromosome 13 library. Mol Biol 29:584–590
Brewer BJ, Fangman WL (1988) A replication fork barrier at the 3′ end of yeast ribosomal RNA genes. Cell 55:637–643
Butler DK, Metzenberg RL (1993) Amplification of the nucleolus organizer region during the sexual phase of Neurospora crassa. Chromosoma 102:519–525
Caburet S, Conti C, Schurra Lebofsky CR, Edelstein SJ, Bensimon A (2005) Human ribosomal RNA gene arrays display a broad range of palindromic structures. Genome Res 15:1079–1085
Carvalho C, Pereira HM, Ferreira J, Pina C, Mendonca D, Rosa AC, Carmo-Fonseca M (2001) Chromosomal G-dark bands determine the spatial organization of centromeric heterochromatin in the nucleus. Mol Biol Cell 12:3563–3572
Casafont I, Navascues J, Pena E, Lafarga M, Berciano MT (2006) Nuclear organization and dynamics of transcription sites in rat sensory ganglia neurons detected by incorporation of 5′-fluorouridine into nascent RNA. Neurosci 14:453–462
Cassidy BG, Yang-Yen HF, Rothblum LI (1986) Transcriptional role for the nontranscribed spacer of rat ribosomal DNA. Mol Cell Biol 6:2766–2773
Cheutin T, O’Donohue MF, Beorchia A, Vandelaer M, Kaplan H, Deféver B, Ploton D, Thiry M (2002) Three-dimensional organization of active rRNA genes within the nucleolus. J Cell Sci 115(Pt 16):3297–3307
Clos J, Normann A, Ohrlein A, Grummt I (1986) The core promoter of mouse rDNA consists of two functionally distinct domains. Nucl Acids Res 14:7581–7595
Cmarko D, Verschure PJ, Rothblum LI, Hernandez-Verdun D, Amalric F, van Driel R, Fakan S (2000) Ultrastructural analysis of nucleolar transcription in cells microinjected with 5-bromo-UTP. Histochem Cell Biol 113:181–187
Cmarko D, Smigova J, Minichova L, Popov A (2008) Nucleolus: the ribosome factory. Histol Histopathol 23:1291–1298
Comings DE (1980) Arrangement of chromatin in the nucleus. Hum Genet 53:131–143
Conconi A, Widmer RM, Koller T, Sogo JM (1989) Two different chromatin structures coexist in ribosomal RNA genes throughout the cell cycle. Cell 57:753–761
Cong R, Das S, Ugrinova I, Kumar S, Mongelard F, Wong J, Bouvet P (2012) Interaction of nucleolin with ribosomal RNA genes and its role in RNA polymerase I transcription. Nucl Acids Res 40:9441–9454. doi:10.1093/nar/gks720
Copenhaver GP, Putnam CD, Denton ML, Pikaard CS (1994) The RNA polymerase I transcription factor UBF is a sequence-tolerant HMG-box protein that can recognize structured nucleic acids. Nucl Acids Res 22:2651–2657
Cremer T, Cremer C (2001) Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat Rev Genet 2:292–301
Csankovszki G, Nagy A, Jaenisch R (2001) Synergism of Xist RNA, DNA methylation, and histone hypoacetylation in maintaining X chromosome inactivation. J Cell Biol 153:773–784
De Winter RF, Moss T (1986) The ribosomal spacer in Xenopus laevis is transcribed as part of the primary ribosomal RNA. Nucl Acids Res 14:6041–6051
Denissov S, Lessard F, Mayer C, Stefanovsky V, van Driel M, Grummt I, Moss T, Stunnenberg HG (2011) A model for the topology of active ribosomal RNA genes. EMBO Rep 12:231–237
Derenzini M, Pasquinelli G, O’Donohue MF, Ploton D, Thiry M (2006) Structural and functional organization of ribosomal genes within the mammalian cell nucleolus. J Histochem Cytochem 54:131–145
Derenzini M, Olins AL, Olins DE (2014) Chromatin structure in situ: the contribution of DNA ultrastructural cytochemistry. Eur J Histochem 58:2307
Diermeier SD, Németh A, Rehli M, Grummt I, Längst G (2013) Chromatin-specific regulation of mammalian rDNA transcription by clustered TTF-I binding sites. PLoS Genet 9:e1003786. doi:10.1371/journal.pgen.1003786
Dimitrova DS (2011) DNA replication initiation patterns and spatial dynamics of the human ribosomal RNA gene loci. J Cell Sci 124:2743–2752
Doelling J, Pikaard C (1995) The minimal ribosomal RNA promoter of Arabidopsis thaliana includes a critical element at the transcription initiation site. Plant J 8:683–692
Dousset T, Wang C, Verheggen C, Chen D, Hernandez-Verdun D, Huang S (2000) Initiation of nucleolar assembly is independent of RNA polymerase I transcription. Mol Biol Cell 11:2705–2717
Dumenco VM, Wejksnora PJ (1986) Characterization of the region around the start point of transcription of ribosomal RNA in the Chinese hamster. Gene 46:227–235
Edenberg HJ, Huberman JA (1975) Eukaryotic chromosome replication. Annu Rev Genet 9:245–284
Elder JF Jr, Turner BJ (1995) Concerted evolution of repetitive DNA sequences in eukaryotes. Q Rev Biol 70:297–320
Erickson JM, Schmickel RD (1985) A molecular basis for discrete size variation in human ribosomal DNA. Am J Hum Genet 37:311–325
Fedoriw AM, Calabrese JM, Mu W, Yee D, Magnuson T (2012) Differentiation-driven nucleolar association of the mouse imprinted Kcnq1 locus. G3 2(12):1521–1528. doi:10.1534/g3.112.004226
Floutsakou I, Agrawal S, Nguyen TT, Seoighe C, Ganley AR, McStay B (2013) The shared genomic architecture of human nucleolar organizer regions. Genome Res 23:2003–2012
Gebrane-Younes J, Fomproix N, Hernandez-Verdun D (1997) When rDNA transcription is arrested during mitosis, UBF is still associated with non-condensed rDNA. J Cell Sci 110:2429–2440
Gencheva M, Anachkova B, Russev G (1996) Mapping the sites of initiation of DNA replication in rat and human rRNA genes. J Biol Chem 271:2608–2614
Gerber JK, Gögel E, Berger C, Wallisch M, Müller F, Grummt I, Grummt F (1997) Termination of mammalian rDNA replication: polar arrest of replication fork movement by transcription termination factor TTF-I. Cell 90:559–567
Gomez-Roman N, Felton-Edkins ZA, Kenneth NS, Goodfellow SJ, Athineos D, Zhang J, Ramsbottom BA, Innes F, Kantidakis T, Kerr ER, Brodie J, Grandori C, White RJ (2006) Activation by c-Myc of transcription by RNA polymerases I, II and III. Biochem Soc Symp 73:141–154
Gonzalez IL, Sylvester JE (1995) Complete sequence of the 43-kb human ribosomal DNA repeat: analysis of the intergenic spacer. Genomics 27:320–328
Gonzalez IL, Sylvester JE (2001) Human rDNA: evolutionary patterns within the genes and tandem arrays derived from multiple chromosomes. Genomics 73:255–263
Gonzalez IL, Gorski JL, Campen TJ, Dorney DJ, Erickson JM, Sylvester JE, Schmickel RD (1985) Variation among human 28S ribosomal RNA genes. Proc Natl Acad Sci 82:7666–7670
Gonzalez IL, Petersen R, Sylvester JE (1989) Independent insertion of alu elements in the human ribosomal spacer and their concerted evolution. Mol Biol Evol 6:413–423
Gonzalez IL, Chambers C, Gorski JL, Stambolian D, Schmickel RD, Sylvester JE (1990) Sequence and structure correlation of human ribosomal transcribed spacers. J Mol Biol 212:27–35
Gonzalez IL, Wu S, Li W-M, Kuo BA, Sylvester JE (1992) Human ribosomal RNA intergenic spacer sequence. Nucl Acids Res 20:5846
Gorski SA, Snyder SK, John S, Grummt I, Misteli T (2008) Modulation of RNA polymerase assembly dynamics in transcriptional regulation. Mol Cell 30:486–497
Grandori C, Gomez-Roman N, Felton-Edkins ZA, Ngouenet C, Galloway DA, Eisenman RN, White RJ (2005) c-Myc binds to human ribosomal DNA and stimulates transcription of rRNA genes by RNA polymerase I. Nat Cell Biol 7:311–318
Grewal SS, Li L, Orian A, Eisenman RN, Edgar BA (2005) Myc-dependent regulation of ribosomal RNA synthesis during Drosophila development. Nat Cell Biol 7:295–302
Grob A, Colleran C, McStay B (2014) Construction of synthetic nucleoli in human cells reveals how a major functional nuclear domain is formed and propagated through cell division. Genes Dev 28:220–230
Grummt I, Maier U, Ohrlein A, Hassouna N, Bachellerie JP (1985) Transcription of mouse rDNA terminates downstream of the 3″ end of 28S RNA and involves interaction of factors with repeated sequences in the 3″ spacer. Cell 43:801–810
Grummt I, Rosenbauer H, Niedermeyer I, Maier U, Ohrlein A (1986) A repeated 18 bp sequence motif in the mouse rDNA spacer mediates binding of a nuclear factor and transcription termination. Cell 45:837–846
Guetg C, Scheifele F, Rosenthal F, Hottiger MO, Santoro R (2012) Inheritance of silent rDNA chromatin is mediated by PARP1 via noncoding RNA. Mol Cell 45:706–707
Haaf T, Schmid M (1989) Centromeric association and non-random distribution of centromeres in human tumour cells. Hum Genet 81:137–143
Haaf T, Schmid M (1991) Chromosome topology in mammalian interphase nuclei. Exp Cell Res 192:325–332
Haaf T, Ward DC (1996) Inhibition of RNA polymerase II transcription causes chromatin decondensation, loss of nucleolar structure, and dispersion of chromosomal domains. Exp Cell Res 224:163–173
Haaf T, Hayman DL, Schmid M (1991) Quantitative determination of rDNA transcription units in vertebrate cells. Exp Cell Res 193:78–86
Hacisuleyman E, Goff LA, Trapnell C et al (2014) Topological organization of multichromosomal regions by the long intergenic noncoding RNA Firre. Nat Struct Mol Biol 21:198–206
Haltiner M, Smale ST, Tjian R (1986) Two distinct promoter elements in the human rRNA gene identified by linker scanning mutagenesis. Mol Cell Biol 6:227–235
Hancock R (2014) The crowded nucleus. Int Rev Cell Mol Biol 307:15–26
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, Barrera LO, Van Calcar S, Qu C, Ching KA et al (2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39:311–318
Heitz E (1931) Die Ursache der gesetzmassigen Zahl, Lage, Form und Groesse pflanzlicher Nukleolen. Planta 12:775–844
Heix J, Vente A, Voit R, Budde A, Michaelidis TM, Grummt I (1998) Mitotic silencing of human rRNA synthesis: inactivation of the promoter selectivity factor SL1 by cdc2/cyclin B-mediated phosphorylation. EMBO J 17:7373–7381
Héliot L, Mongelard F, Klein C, O’Donohue MF, Chassery JM, Robert-Nicoud M, Usson Y (2000) Nonrandom distribution of metaphase AgNOR staining patterns on human acrocentric chromosomes. J Histochem Cytochem 48:13–20
Henderson AS, Warburton D, Atwood KC (1972) Location of ribosomal DNA in the human chromosome complement. Proc Natl Acad Sci USA 69:3394–3398
Hernández-Hernández A, Soto-Reyes E, Ortiz R, Arriaga-Canon C, Echeverría-Martinez OM, Vázquez-Nin GH, Recillas-Targa F (2012) Changes of the nucleolus architecture in absence of the nuclear factor CTCF. Cytogenet Gen Res 136:89–96
Hozák P, Schöfer C, Sylvester J, Wachtler F (1993) A study on nucleolar DNA: isolation of DNA from fibrillar components and ultrastructural localization of different DNA probes. J Cell Sci 104:1199–1205
Hu CH, McStay B, Jeong SW, Reeder RH (1994) xUBF, an RNA polymerase I transcription factor, binds crossover DNA with low sequence specificity. Mol Cell Biol 14:2871–2882
Huang K, Jia J, Wu C, Yao M, Li M, Jin J, Jiang C, Cai Y, Pei D, Pan G, Yao H (2013) Ribosomal RNA gene transcription mediated by the master genome regulator protein CCCTC-binding factor (CTCF) is negatively regulated by the condensin complex. J Biol Chem 288:26067–26077. doi:10.1074/jbc.M113.486175
Ide S, Miyazaki T, Maki H, Kobayashi T (2010) Abundance of ribosomal RNA gene copies maintains genome integrity. Science 327:693–696
Jacob MD, Audas TE, Mullineux ST, Lee S (2012) Where no RNA polymerase has gone before: novel functional transcripts derived from the ribosomal intergenic spacer. Nucleus 3:315–319
Jacob MD, Audas TE, Uniacke J, Trinkle-Mulcahy L, Lee S (2013) Environmental cues induce a long noncoding RNA-dependent remodeling of the nucleolus. Mol Biol Cell 24:2943–2953. doi:10.1091/mbc.E13-04-0223
Jakociunas T, Domange Jordö M, Aït Mebarek M, Bünner CM, Verhein-Hansen J, Oddershede LB, Thon G (2013) Subnuclear relocalization and silencing of a chromosomal region by an ectopic ribosomal DNA repeat. Proc Natl Acad Sci USA 110:E4465–E4473. doi:10.1073/pnas.1315581110
Jordan P, Mannervik M, Tora L, Carmo-Fonseca M (1996) In vivo evidence that TATA-binding protein/SL1 colocalizes with UBF and RNA polymerase I when rRNA synthesis is either active or inactive. J Cell Biol 133:225–234
Junera HR, Masson C, Geraud G, Hernandez-Verdun D (1995) The three-dimensional organization of ribosomal genes and the architecture of the nucleoli vary with G1, S and G2 phases. J Cell Sci 108:3427–3441
Kalmárová M, Smirnov E, Mašata M, Koberna K, Ligasová A, Popov A, Raška I (2007) Positioning of NORs and NOR-bearing chromosomes in relation to nucleoli. J Struct Biol 160:49–56
Kalmárová M, Kovácik L, Popov A, Testillano SP, Smirnov E (2008) Asymmetrical distribution of the transcriptionally competent NORs in mitosis. J Struct Biol 163:40–44. doi:10.1016/j.jsb.2008.04.002
Kaplan FS, Murray J, Sylvester JE, Gonzalez IL, O’Connor JP, Doering JL, Muenke M, Emanuel BS, Zasloff MA (1993) The topographic organization of repetitive DNA in the human nucleolus. Genomics 15:123–132
Kind J, Pagie L, Ortabozkoyun H, Boyle S, de Vries SS, Janssen H, Amendola M, Nolen LD, Bickmore WA, van Steensel B (2013) Single-cell dynamics of genome–nuclear lamina interactions. Cell 153:178–192
Koberna K, Malínský J, Pliss A, Mašata M, Večeřová J, Fialová M, Bednár J, Raška I (2002) Ribosomal genes in focus: new transcripts label the dense fibrillar components and form clusters indicative of “Christmas trees” in situ. J Cell Biol 157:743–748
Kominami R, Urano Y, Mishima Y, Muramatsu M (1981) Organization of ribosomal RNA gene repeats of the mouse. Nucl Acids Res 14:3219–3233
Kuhn A, Normann A, Bartsch I, Grummt I (1988) The mouse ribosomal gene terminator consists of three functionally separable sequence elements. EMBO J 7:1497–1502
Kupriyanova NS, Netchvolodov KK, Sadova AA, Cherepanova MD, Ryskov AP (2015) Non-canonical ribosomal DNA segments in the human genome, and nucleoli functioning. Gene 572:237–242
La Volpe A, La Mantia G, Gargiulo G, Malva C (1984) Regulation of rRNA gene number in Drosophila melanogaster: new aspects resulting from the use of free duplications. Mol Gen Genet 194:485–488
La Volpe A, Simeone A, Simeone A, D’Esposito M, Scotto L, Fidanza V, de Falco A, Boncinelli E (1985) Molecular analysis of the heterogeneity region of the human ribosomal spacer. J Mol Biol 183:213–223
Labhart P, Reeder RH (1986) Characterization of three sites of RNA 3′ end formation in the Xenopus ribosomal gene spacer. Cell 45:431–443
Lam YW, Trinkle-Mulcahy L (2015) New insights into nucleolar structure and function. F1000Prime Rep 7:48
Lebofsky R, Bensimon A (2005) DNA replication origin plasticity and perturbed fork progression in human inverted repeats. Mol Cell Biol 25:6789–6797
Léger I, Guillaud M, Krief B, Brugal G (1994) Interactive computer-assisted analysis of chromosome 1 colocalization with nucleoli. Cytometry 16:313–323
Leung AK, Gerlich D, Miller G, Lyon C, Lam YW, Lleres D, Daigle N, Zomerdijk J, Ellenberg J, Lamond AI (2004) Quantitative kinetic analysis of nucleolar breakdown and reassembly during mitosis in live human cells. J Cell Biol 166:787–800
Levinson G, Gutman GA (1987) High frequencies of short frameshifts in poly-CA/TG tandem repeats borne by bacteriophage M13 in Escherichia coli K-12. Nucl Acids Res 15:5323–5338
Lewin B (1980) Gene expression, vol 2. Wiley, New York
Li Z, Hann SR (2013) Nucleophosmin is essential for c-Myc nucleolar localization and c-Myc-mediated rDNA transcription. Oncogene 32:1988–1994
Li J, Langst G, Grummt I (2006) NoRC-dependent nucleosome positioning silences rRNA genes. EMBO J 25:5735–5741
Linskens MHK, Huberman JA (1988) Organization of replication of ribosomal DNA in Saccharomyces cerevisiae. Mol Cell Biol 8:4927–4935
Little RD, Platt TH, Schildkraut CL (1993) Initiation and termination of DNA replication in human rRNA genes. Mol Cell Biol 13:6600–6613
Long EO, Dawid IB (1980) Repeated genes in eukaryotes. Annu Rev Biochem 49:727–764
Lykke-Andersen S, Mapendano CK, Jensen TH (2011) An ending is a new beginning: transcription termination supports re-initiation. Cell Cycle 10:863–865
Maden BE, Dent CL, Farrell TE, Garde J, McCallum FS, Wakeman JA (1987) Clones of human ribosomal DNA containing the complete 18 S-rRNA and 28 S-rRNA genes. Characterization, a detailed map of the human ribosomal transcription unit and diversity among clones. Biochem J 246:519–527
Magdalou I, Lopez BS, Pasero P, Lambert SAE (2014) The causes of replication stress and their consequences on genome stability and cell fate. Semin Cell Dev Biol 30:154–164
Mais C, Wright JE, Prieto JL, Raggett SL, McStay B (2005) UBF-binding site arrays form pseudo-NORs and sequester the RNA polymerase I transcription machinery. Genes Dev 19:50–64
Manuelidis L (1984) Different central nervous system cell types display distinct and nonrandom arrangements of satellite DNA sequences. Proc Natl Acad Sci 81:3123–3127
Manuelidis L, Borden J (1988) Reproducible compartmentalization of individual chromosome domains in human CNS cells revealed by in situ hybridization and three-dimensional reconstruction. Chromosoma 96:397–410
Matera AG, Frey MR, Margelot K, Wolin SL (1995) A perinucleolar compartment contains several RNA polymerase III transcripts as well as the polypyrimidine tract-binding protein, hnRNP I. J Cell Biol 129:1181–1193
Matheson TD, Kaufman PD (2015) Grabbing the genome by the NADs. Chromosoma. doi:10.1007/s00412-015-0527-8
Mayer C, Schmitz KM, Li J, Grummt I, Santoro R (2006) Intergenic transcripts regulate the epigenetic state of rRNA genes. Mol Cell 22:351–361
Mayer C, Neubert M, Grummt I (2008) The structure of NoRC-associated RNA is crucial for targeting the chromatin remodelling complex NoRC to the nucleolus. EMBO Rep 9:774–780
McClintock B (1934) The relationship of a particular chromosomal element to the development of the nucleoli in Zea mays. Zeit Zellforsch Mik Anat 21:294–328
McKeown PC, Shaw PJ (2009) Chromatin: linking structure and function in the nucleolus. Chromosoma 118:11–23. doi:10.1007/s00412-008-0184-2
McStay B, Grummt I (2008) The epigenetics of rRNA genes: from molecular to chromosome biology. Annu Rev Cell Dev Biol 24:131–157
McStay B, Reeder RH (1986) A termination site for Xenopus RNA polymerase I also acts as an element of an adjacent promoter. Cell 47:913–920
Melčák I, Risueño MC, Raška I (1996) Ultrastructural non-isotopic mapping of nucleolar transcription sites in onion protoplasts. J Struct Biol 116:253–263
Michalet X, Ekong R, Fougerousse F, Rousseaux S, Schurra C, Hornigold N, van Slegtenhorst M, Wolfe J, Povey S, Beckmann JS, Bensimon A (1997) Dynamic molecular combing: stretching the whole human genome for high-resolution studies. Science 277:1518–1523
Mighell AJ, Smith NR, Robinson PA, Markham AF (2000) Vertebrate pseudogenes. FEBS Lett 468:109–114
Miller OL Jr, Beatty BR (1969) Visualization of nucleolar genes. Science 164:955–957
Mohammad F, Pandey RR, Nagano T, Chakalova L, Mondal T, Fraser P, Kanduri C (2008) Kcnq1ot1/Lit1 noncoding RNA mediates transcriptional silencing by targeting to the perinucleolar region. Mol Cell Biol 28:3713–3728. doi:10.1128/MCB.02263-07
Moss T, Stefanovsky VY (1995) Promotion and regulation of ribosomal transcription in eukaryotes by RNA polymerase I. In: Cohn WE, Moldave K (eds) Progress in nucleic acids and molecular biology. Academic Press, San Diego, pp 25–66
Moss T, Mitchelson K, De Winter RFJ (1985) The promotion of ribosomal transcription in eukaryotes. Oxf Surv Eukaryot Genes 2:207–250
Moss T, Stefanovsky V, Langlois F, Gagnon-Kugler T (2006) A new paradigm for the regulation of the mammalian ribosomal RNA genes. Biochem Soc Trans 34:1079–1081
Moss T, Langlois F, Gagnon-Kugler T, Stefanovsky V (2007) A housekeeper with power of attorney: the rRNA genes in ribosome biogenesis. Cell Mol Life Sci 64:29–49
Mougey EB, O’Reilly M, Osheim Y, Miller OL Jr, Beyer A, Sollner-Webb B (1993) The terminal balls characteristic of eukaryotic rRNA transcription units in chromatin spreads are rRNA processing complexes. Genes Dev 7:1609–1619
Mroczka DL, Cassidy B, Busch H, Rothblum LI (1984) Characterization of rat ribosomal DNA: the highly repetitive sequences that flank the ribosomal RNA transcription unit are homologous and contain RNA polymerase III transcription initiation sites. J Mol Biol 174:141
Müller WG, Rieder D, Karpova TS, John S, Trajanoski Z, McNally JG (2007) Organization of chromatin and histone modifications at a transcription site. J Cell Biol 177:957–967
Murano K, Okuwaki M, Hisaoka M, Nagata K (2008) Transcription regulation of the rRNA gene by a multifunctional nucleolar protein, B23/nucleophosmin, through its histone chaperone activity. Mol Cell Biol 28:3114–3126
Nazar RN (2004) Ribosomal processing and ribosome biogenesis in eukaryotes. IUBMB Life 56:457–465
Németh A, Längst G (2008) Chromatin organization of active ribosomal RNA genes. Epigenetics 3:243–245
Németh A, Längst G (2011) Genome organization in and around the nucleolus. Trends Genet 27:149–156
Németh A, Guibert S, Tiwari VK, Ohlsson R, Langst G (2008) Epigenetic regulation of TTF-I-mediated promoter–terminator interactions of rRNA genes. EMBO J 27:1255–1265
Németh A, Conesa A, Santoyo-Lopez J, Medina I, Montaner D, Péterfia B, Solovei I, Cremer T, Dopazo J, Längst G (2010) Initial genomics of the human nucleolus. PLoS Genet 6:e1000889. doi:10.1371/journal.pgen.1000889
Németh A, Perez-Fernandez J, Merkl P, Hamperl S, Gerber J, Griesenbeck J, Tscochner H (2013) RNA polymerase I termination: Where is the end? Biochem Biophys Acta 1829:306–317
O’Sullivan AC, Sullivan GJ, McStay B (2002) UBF binding in vivo is not restricted to regulatory sequences within the vertebrate ribosomal DNA repeat. Mol Cell Biol 22:657–668
Ochs RL, Press RI (1992) Centromere autoantigens are associated with the nucleolus. Exp Cell Res 200:339–350
Ochs RL, Lischwe MA, Spohn WH, Busch H (1985) Fibrillarin: a new protein of the nucleolus identified by autoimmune sera. Biol Cell 54:123–133
O’Keefe RT, Henderson SC, Spector DL (1992) Dynamic organization of DNA replication in mammalian cell nuclei: spatially and temporally defined replication of chromosome-specific α-satellite DNA sequences. J Cell Biol 116:1095–1110
O’Sullivan JM, Sontam DM, Grierson R, Jones B (2009) Repeated elements coordinate the spatial organization of the yeast genome. Yeast 26:125–138
Padeken J, Heun P (2014) Nucleolus and nuclear periphery: velcro for heterochromatin. Curr Opin Cell Biol 28:54–60
Padeken J, Mendiburo MJ, Chlamydas S, Schwarz HJ, Kremmer E, Heun P (2013) The nucleoplasmin homolog NLP mediates centromere clustering and anchoring to the nucleolus. Mol Cell 25(50):236–249. doi:10.1016/j.molcel.2013.03.002
Pandey RR, Mondal T, Mohammad F, Enroth S, Redrup L, Komorowski J, Nagano T, Mancini-Dinardo D, Kanduri C (2008) Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell 32:232–246
Pasero P, Bensimon A, Schwob E (2002) Single-molecule analysis reveals clustering and epigenetic regulation of replication origins at the yeast rDNA locus. Genes Dev 16:2479–2484
Paule M (1994) Transcription of ribosomal RNA by eukaryotic RNA polymerase I. In: Conaway RC, Conaway JW (eds) Transcription: mechanisms and regulation. Raven Press, New York, pp 83–106
Peng JC, Karpen GH (2007) H3K9 methylation and RNA interference regulate nucleolar organization and repeated DNA stability. Nat Cell Biol 9:25–35
Pfleiderer C, Smid A, Bartsch I, Grummt I (1990) An undecamer DNA sequence directs termination of human ribosomal gene transcription. Nucl Acids Res 18:4727–4736
Philippsen P, Kramer RA, Davis RW (1978) Cloning of the yeast ribosomal DNA repeat unit in SstI and HindIII lambda vectors using genetic and physical size selections. J Mol Biol 123:371–386
Pikaard CS, Pape LK, Henderson SL, Ryan K, Paalman MH, Lopata MA, Reeder RH, Sollner-Webb B (1990) Enhancers for RNA polymerase I in mouse ribosomal DNA. Mol Cell Biol 10:4816–4825
Pliss A, Koberna K, Vecerova J, Malinsky J, Masata M, Fialova M, Raska I, Berezney R (2005) Spatio-temporal dynamics at rDNA foci: global switching between DNA replication and transcription. J Cell Biochem 94:554–565
Pliss A, Fritz AJ, Stojkovic B, Ding H, Mukherjee L, Bhattacharya S, Xu J, Berezney R (2015) Non-random patterns in the distribution of NOR-bearing chromosome territories in human fibroblasts: a network model of interactions. J Cell Physiol 230:427–439
Pontvianne F, Blevins T, Chandrasekhara C, Mozgova I, Hassel C, Pontes OM, Tucker S, Mokros P, Muchova V, Fajkus J et al (2013) Subnuclear partitioning of rRNA genes between the nucleolus and nucleoplasm reflects alternative epiallelic states. Genes Dev 27:1545–1550
Prieto JL, McStay B (2005) Nucleolar biogenesis: the first small steps. Biochem Soc Trans 33:1441–1443
Prieto JL, McStay B (2007) Recruitment of factors linking transcription and processing of pre-rRNA to NOR chromatin is UBF-dependent and occurs independent of transcription in human cells. Genes Dev 21:2041–2054
Prieto JL, McStay B (2008) Pseudo-NORs: a novel model for studying nucleoli. Biochim Biophys Acta 1783:2116–2123
Puvion-Dutilleul F, Bachellerie JP, Puvion E (1991) Nucleolar organization of HeLa cells as studied by in situ hybridization. Chromosoma 100:395–409
Rahmanzadeh R, Hüttmann G, Gerdes J, Scholzen T (2007) Chromophore-assisted light inactivation of pKi-67 leads to inhibition of ribosomal RNA synthesis. Cell Prolif 40:422–430
Raska I (2003) Oldies but goldies: searching for Christmas trees within the nucleolar architecture. Trends Cell Biol 13:517–525
Raska I, Reimer G, Jarnik M, Kostrouch Z, Raska K Jr (1989) Does the synthesis of ribosomal RNA take place within nucleolar fibrillar centers or dense fibrillar components. Biol Cell 65(1):79–82
Raska I, Shaw PJ, Cmarko D (2006a) New insights into nucleolar architecture and activity. Int Rev Cytol 255:177–235
Raska I, Shaw PJ, Cmarko D (2006b) Structure and function of the nucleolus in the spotlight. Curr Opin Cell Biol 18:325–334
Raška I, Rychter Z, Smetana K (1983a) Fibrillar centers and condensed nucleolar chromatin in resting and stimulated human lymphocytes. Zeitschrift fur mikroskopischanatomische Forschung 97(1):15–32
Raška I, Armbruster BL, Frey JR, Smetana K (1983b) Analysis of ring-shaped nucleoli in serially sectioned human lymphocytes. Cell Tissue Res 234(3):707–711
Raška I, Dundr M, Koberna K, Melčák I, Risueño MM, Török I (1995) Does the synthesis of ribosomal RNA take place within nucleolar fibrillar centres or dense fibrillar components? A critical appraisal. J Struct Biol 114:1–22
Rawlins DJ, Shaw PJ (1990) Localization of ribosomal and telomeric DNA sequences in intact plant nuclei by in situ hybridization and three-dimensional optical microscopy. J Microsc 157:83–89
Reeder R (1992) Regulation of transcription by RNA polymerase I. In: McKnight SL, Yamamoto KR (eds) Transcriptional regulation. Cold Spring Harbour Laboratory Press, Cold Spring Harbour, pp 315–347
Reeder RH, Lang WH (1997) Terminating transcription in eukaryotes: lessons learned from RNA polymerase I. Trends Biochem Sci 22:473–477
Reeder RH, Brown DD, Wellauer PK, Dawid IB (1976) Patterns of ribosomal DNA spacer lengths are inherited. J Mol Biol 105:507–516
Rickards B, Flint SJ, Cole MD, LeRoy G (2007) Nucleolin is required for RNA polymerase I transcription in vivo. Mol Cell Biol 27:937–948
Roger B, Moisand A, Amalric F, Bouvet P (2003) Nucleolin provides a link between RNA polymerase I transcription and pre-ribosome assembly. Chromosoma 111:399–407
Rogers SO, Bendich AJ (1987) Ribosomal RNA genes in plants: variability in copy number and in the intergenic spacer. Plant Mol Biol 9:509
Rothstein R, Michel B, Gangloff S (2000) Replication fork pausing and recombination or “gimme a break”. Genes Dev 14:1–10
Roussel P, Hernandez-Verdun D (1994) Identification of Ag-NOR proteins, markers of proliferation related to ribosomal gene activity. Exp Cell Res 214:465–472
Roussel P, Andre C, Comai L, Hernandez-Verdun D (1996) The rDNA transcription machinery is assembled during mitosis in active NORs and absent in inactive NORs. J Cell Biol 133:235–246
Ryskov AP, Kupriianova NS, Kapanadze BI, Nechvolodov KK, Pozmogova GE, Prosniak MI, Iankovskiĭ NK (1993) Frequency of various mini- and micro-satellite sequences in DNA of human chromosome 13. Genetika 29:1750–1754
Santoro R (2005) The silence of the ribosomal RNA genes. Cell Mol Life Sci 62:2067–2079
Santoro R (2014) Analysis of chromatin composition of repetitive sequences: the ChIP-Chop assay. Methods Mol Biol 1094:319–328
Santoro R, De Lucia F (2005) Many players, one goal: how chromatin states are inherited during cell division. Biochem Cell Biol 83:332–343
Santoro R, Li J, Grummt I (2002) The nucleolar remodeling complex NoRC mediates heterochromatin formation and silencing of ribosomal gene transcription. Nat Genet 32:393–396
Santoro R, Schmitz KM, Sandoval J, Grummt I (2010) Intergenic transcripts originating from a subclass of ribosomal DNA repeats silence ribosomal RNA genes in trans. EMBO Rep 11:52–58
Sasaki T, Okazaki T, Muramatsu M, Kominami R (1987) Variation among mouse ribosomal RNA genes within and between chromosomes. Mol Biol Evol 4:594–601
Savino TM, Gebrane-Younes J, De Mey J, Sibarita JB, Hernandez-Verdun D (2001) Nucleolar assembly of the rRNA processing machinery in living cells. J Cell Biol 153:1097–1110
Scheer U, Benavente R (1990) Functional and dynamic aspects of the mammalian nucleolus. Review. Bioessays 12:14–21
Scheer U, Zentgraf H (1982) Morphology of nucleolar chromatin in electron microscopic spread preparations. In: Busch Rothblum (ed) The cell nucleus. Academic Press, New York, p 143
Scheer U, Xia B, Merkert H, Weisenberger D (1997) Looking at Christmas trees in the nucleolus. Chromosoma 105:470–480
Schmitz KM, Mayer C, Postepska A, Grummt I (2010) Interaction of noncoding RNA with the rDNA promoter mediates recruitment of DNMT3b and silencing of rRNA genes. Genes Dev 24:2264–2269
Scott RS, Truong KY, Vos JM (1997) Replication initiation and elongation fork rates within a differentially expressed human multicopy locus in early S phase. Nucl Acids Res 25:4505–4512
Shaw PJ, McKeown PC (2011) The structure of rDNA chromatin. In: Olson MOJ (ed) The nucleolus, protein reviews, vol 15. Springer, New York, pp 43–55
Shiue CN, Berkson RG, Wright AP (2009) c-Myc induces changes in higher order rDNA structure on stimulation of quiescent cells. Oncogene 28:1833–1842
Shiue CN, Nematollahi-Mahani A, Wright AP (2014) Myc-induced anchorage of the rDNA IGS region to nucleolar matrix modulates growth-stimulated changes in higher-order rDNA architecture. Nucl Acids Res 42:5505–5517
Sirri V, Roussel P, Hernandez-Verdun D (1999) The mitotically phosphorylated form of the transcription termination factor TTF-1 is associated with the repressed rDNA transcription machinery. J Cell Sci 112:3259–3268
Sirri V, Urcuqui-Inchima S, Roussel P, Hernandez-Verdun D (2008) Nucleolus: the fascinating nuclear body. Histochem Cell Biol 129:13–31
Smetana K, Klamová H, Mikulenková D, Pluskalová M, Hrkal Z (2006) On the nucleolar size and density in human early granulocytic progenitors, myeloblasts. Eur J Histochem 50:119–124
Smirnov E, Kalmárová M, Koberna K, Zemanová Z, Malínský J, Masata M, Cvacková Z, Michalová K, Raska I (2006) NORs and their transcription competence during the cell cycle. Folia Biol (Praha) 52(3):59–70
Smirnov E, Borkovec J, Kováčik L, Svidenská S, Schröfel A, Skalníková M, Švindrych Z, Křížek P, Ovesný M, Hagen GM, Juda P, Michalová K, Cardoso MC, Cmarko D, Raška I (2014) Separation of replication and transcription domains in nucleoli. J Struct Biol 188:259–266. doi:10.1016/j.jsb.2014.10.001
Stahl A, Hartung M, Vagner-Capodano AM, Fouet C (1976) Chromosomal constitution of nucleolus-associated chromatin in man. Hum Genet 35:27–34
Strohner R, Nemeth A, Nemeth A, Jansa P et al (2001) NoRC—a novel member of mammalian ISWI-containing chromatin remodeling machines. EMBO J 20:4892–4900
Stults DM, Killen MW, Pierce HH, Pierce AJ (2008) Genomic architecture and inheritance of human ribosomal RNA gene clusters. Genome Res 18:13–18
Stults DM, Killen MW, Williamson EP, Hourigan JS, Vargas HD, Arnold SM, Moscow JA, Pierce AJ (2009) Human rRNA gene clusters are recombinational hotspots in cancer. Cancer Res 69:9096–9104
Sullivan GJ, Bridger JM, Cuthbert AP, Newbold RF, Bickmore WA, McStay B (2001) Human acrocentric chromosomes with transcriptionally silent nucleolar organizer regions associate with nucleoli. EMBO J 20:2867–2874
Sylvester JE, Whiteman DA, Podolsky R, Pozsgay JM, Respess J, Schmickel RD (1986) The human ribosomal RNA genes: structure and organization of the complete repeating unit. Hum Genet 73:193–198
Sylvester JE, Petersen R, Schmickel RD (1989) Human ribosomal DNA: novel sequence organization in a 4.5-kb region upstream from the promoter. Gene 84:193–196
Sylvester JE, Gonzalez IL, Mougey EB (2003) In: Olson M (ed) The nucleolus. Landes Bioscience, Georgetown, pp 58–73
Tautz D, Dover GA (1986) Transcription of the tandem array of ribosomal DNA in Drosophila melanogaster does not terminate at any fixed point. EMBO J 6:1267–1273
Tautz D, Trick M, Dover GA (1986) Cryptic simplicity in DNA is a major source of genetic variation. Nature 322:652–656
Thompson M, Haeusler RA, Good PD, Engelke DR (2003) Nucleolar clustering of dispersed tRNA genes. Science 302:1399–1401
Torrano V, Navascues J, Docquier F, Zhang R, Burke LJ, Chernukhin I, Farrar D, Leon J, Berciano MT, Renkawitz R et al (2006) Targeting of CTCF to the nucleolus inhibits nucleolar transcription through a poly (ADP-ribosyl)ation-dependent mechanism. J Cell Sci 119:1746–1759
Tower J, Henderson SL, Dougherty KM, Wejksnora PJ, Sollner-Webb B (1989) An RNA polymerase I promoter located in the CHO and mouse ribosomal DNA spacers: functional analysis and factor and sequence requirements. Mol Cell Biol 9:1513
Trendelenburg MF, Puvion-Dutilleul F (1987) Visualizing active genes. In: Sommerville A, Scheer U (eds) Electron microscopy in molecular biology: a practical approach. IRL Press, Oxford, pp 101–146
Trendelenburg MF, Spring H, Scheer U, Franke WW (1974) Morphology of nucleolar cistrons in a plant cell, Acetabularia mediterranea. Proc Natl Acad Sci USA 71:3626–3630
Tseng H, Chou W, Wang J, Zhang X, Zhang S, Schultz RM (2008) Mouse ribosomal RNA genes contain multiple differentially regulated variants. PLoS One 3:e1843. doi:10.1371/journal.pone.0001843
Valentin G (1836) Repertorium für anatomie und physiologie. Verlag von Veit und Comp Berlin 1:1–293
Valenzuela L, Kamakaka RT (2006) Chromatin insulators. Annu Rev Genet 40:107–138
van de Nobelen S, Rosa-Garrido M, Leers J, Heath H, Soochit W, Joosen L, Jonkers I, Demmers J, van der Reijden M, Torrano V et al (2010) CTCF regulates the local epigenetic state of ribosomal DNA repeats. Epigenet Chromat 3:17
van Koningsbruggen S, Gierlinski M, Schofield P, Martin D, Barton GJ, Ariyurek Y, den Dunnen JT, Lamond AI (2010) High-resolution whole-genome sequencing reveals that specific chromatin domains from most human chromosomes associate with nucleoli. Mol Biol Cell 21:3735–3748
Voit R, Seiler J, Grummt I (2015) Cooperative action of Cdk1/cyclin B and SIRT1 is required for mitotic repression of rRNA synthesis. PLoS Genet 11:e1005246. doi:10.1371/journal.pgen.1005246
Vourc’h C, Taruscio D, Boyle AL, Ward DC (1993) Cell-cycle-dependent distribution of telomeres, centromeres and chromosome-specific subsatellite domains in the interphase nucleus of mouse lymphocytes. Exp Cell Res 205:142–151
Wagner R (1835) Einige Bemerkungen und Fragen über das keimbläschen (vesicular germinativa). Müller’s Archiv Anat Physiol Wissenschaft Med 373–377
Weisenberger D, Scheer UA (1995) A possible mechanism for the inhibition of ribosomal RNA gene transcription during mitosis. J Cell Biol 129:561–575
Wellauer PK, Dawid IB (1977) The structural organization of rDNA in Drosophila melanogaster. Cell 10:193–212
Wild MA, Gall JG (1979) An intervening sequence in the gene coding for 25S ribosomal RNA of Tetrahymena pigmentosa. Cell 16:565–573
Wong LH, Brettingham-Moore KH, Chan L, Quach JM, Anderson MA, Northrop EL, Hannan R, Saffery R, Shaw ML, Williams E, Choo KHA (2007) Centromere RNA is a key component for the assembly of nucleoproteins at the nucleolus and centromere. Genome Res 17:1146–1160
Xie W, Ling T, Zhou Y, Feng W, Zhu Q et al (2012) The chromatin remodeling complex NuRD establishes the poised state of rRNA genes characterized by bivalent histone modifications and altered nucleosome positions. Proc Natl Acad Sci 109:8161–8166
Yang F, Babak T, Shendure J, Disteche CM (2010) Global survey of escape from X inactivation by RNA-sequencing in mouse. Genome Res 20:614–622
Yang F, Deng X, Ma W et al (2015) The lncRNA Firre anchors the inactive X chromosome to the nucleolus by binding CTCF and maintains H3K27me3 methylation. Genome Biol 16:52
Yoon Y, Sanches A, Brun C, Huberman JA (1995) Mapping of replication initiation sites in human ribosomal DNA by nascent-strand abundance analysis. Mol Cell Biol 15:2482–2489
Yuan X, Feng W, Imhof A, Grummt I, Zhou Y (2007) Activation of RNA polymerase I transcription by cockayne syndrome group B protein and histone methyltransferase G9a. Mol Cell 27:585–595
Yusufzai TM, Tagami H, Nakatani Y, Felsenfeld G (2004) CTCF tethers an insulator to subnuclear sites, suggesting shared insulator mechanisms across species. Mol Cell 13:291–298
Zentner GE, Saiakhova A, Manaenkov P, Adams MD, Scacheri PC (2011) Integrative genomic analysis of human ribosomal DNA. Nucl Acids Res 39:4949–4960
Zentner GE, Balow SA, Scacheri PC (2014) Genomic characterization of the mouse ribosomal DNA locus. G3 4:243–254. doi:10.1534/g3.113.009290
Zhang S, Hemmerich P, Grosse F (2004) Nucleolar localization of the human telomeric repeat binding factor 2 (TRF2). J Cell Sci 117:3935–3945
Zhang LF, Huynh KD, Lee JT (2007) Perinucleolar targeting of the inactive X during S phase: evidence for a role in the maintenance of silencing. Cell 129:693–706
Zhou Y, Grummt I (2005) The PHD finger/bromodomain of NoRC interacts with acetylated histone H4K16 and is sufficient for rDNA silencing. Curr Biol 15:1434–1438
Zillner K, Filarsky M, Rachow K, Weinberger M, Längst G, Németh A (2013) Large-scale organization of ribosomal DNA chromatin is regulated by Tip5. Nucl Acids Res 41:5251–5262. doi:10.1093/nar/gkt218
Zillner K, Komatsu J, Filarsky K, Kalepu R, Bensimon A, Németh A (2015) Active human nucleolar organizer regions are interspersed with inactive rDNA repeats in normal and tumor cells. Epigenomics 7:363–378
Acknowledgments
This work was supported by the Grant Agency of Czech Republic (P302/12/1885, P302/12/G157, and 13-12317J), by Charles University in Prague (PRVOUK P27/LF1/1 and UNCE 204022), and by OPPK (CZ.2.16/3.1.00/24010).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Smirnov, E., Cmarko, D., Mazel, T. et al. Nucleolar DNA: the host and the guests. Histochem Cell Biol 145, 359–372 (2016). https://doi.org/10.1007/s00418-016-1407-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-016-1407-x