Abstract
The nucleolus is an informative model structure for studying how chromatin-regulated transcription relates to nuclear organisation. In this review, we describe how chromatin controls nucleolar structure through both the modulation of rDNA activity by convergently-evolved remodelling complexes and by direct effects upon rDNA packaging. This packaging not only regulates transcription but may also be important for suppressing internal recombination between tandem rDNA repeats. The identification of nucleolar histone chaperones and novel chromatin proteins by mass spectrometry suggests that structure-specific chromatin components remain to be characterised and may regulate the nucleolus in novel ways. However, it also suggests that there is considerable overlap between nucleolar and non-nucleolar-chromatin components. We conclude that a fuller understanding of nucleolar chromatin will be essential for understanding how gene organisation is linked with nuclear architecture.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The defining characteristic of eukaryotes is the double membrane-bound nucleus. Within the nucleus, DNA is packaged into the proteinaceous ‘super-structure’, chromatin, by association with histones, RNA molecules and other proteins. Organisms can be understood in terms of their physical structure and of the genetic information that they transmit, and chromatin links these aspects by arranging DNA into a structure which both organises and regulates its transcription. So essential is this connection between form and function that chromatin has been described as ‘a, if not the, hallmark of eukaryotic life’ (Benecke 2006).
The packaging of DNA into chromatin produces a nucleus that is structured yet dynamic, and at least partially self-organising (Misteli 2001; Lamond and Sleeman 2003). This self-organisation may be regulated by covalent modification of histones (Kimura et al. 2005; Martin and Zhang 2005; Fuchs et al. 2006) or result from the interplay of chromatin regions (Espada and Esteller 2007). It also leads to the formation of many sub-nuclear domains and bodies (Spector 2003), perhaps through the aggregation of functionally related proteins (Hancock 2004). Such structures highlight the roles of chromatin in linking nuclear function to the emergence of higher-order organisation.
The nucleolus, the site of rDNA transcription, rRNA maturation and ribosome production, is the largest such nuclear domain (reviewed Lyon and Lamond 2000). It is also the site of many other RNA-processing reactions and viral maturation and function (Raska et al. 2006; Boisvert et al. 2007; Andersen et al. 2002; Hiscox 2007), and nucleolar alterations occur in many pathologies and are, for example, of diagnostic importance in cancer (Maggi and Weber 2005). The question of how such structures form has been posed as a dichotomy between ‘self-organisation’—purely due to the biochemical processes—and regulation by ‘watchmakers’—external structure-imposing elements (Kurakin 2005). As nucleoli are reformed as a result of rDNA transcription, a self-organizing model seems appropriate (Melese and Xu 1995; Hernandez-Verdun et al. 2002), but with external inputs playing a fine-tuning role. In this review, we describe the central place of chromatin in this ‘modulated self-organization’ through direct structural effects on rDNA, and subsequent control of its transcription, and discuss how these regulate the emergence of nucleolar structure. The roles chromatin may play in other nucleolar functions are also considered.
“Born at the junction of form and function”—structure and role of the nucleolus
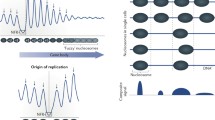
Within the nucleolus, ribosomal RNA (rRNA) genes, organised within ribosomal DNA (rDNA) as tandem repeats at nucleolar organiser regions (NORs) and separated by linker regions, are transcribed by RNA polymerase I (Pol I) to produce pre-rRNA. These transcripts are processed into three of the ribosomal RNAs and complexed with 5S RNA, probably transcribed elsewhere in the nucleus, and ribosomal proteins imported from the cytoplasm to form pre-ribosomes (Fig. 1). Nucleoli show considerable substructure in thin section transmission electron microscopy (TEM), which has traditionally been described as comprising lightly staining fibrillar centres (FCs) of about 0.1–1.0 μm in size, surrounded by or appressed to dense fibrillar component (DFC), which is usually more heavily stained than the rest of the nucleolus. The remaining parts of the nucleolus appear in the TEM to be filled with granules (called the granular component—GC); the granules are assumed to represent mainly pre-ribosomal particles in various stages of maturation (Shaw and Jordan 1995; Raska et al. 2006; Hernandez-Verdun 2005). There has been considerable debate about what the EM ultrastructure corresponds to in functional terms (Raska et al. 2006), and in particular where the transcribing genes are located. Different investigators favour either within the DFC or at the border between FC and DFC (Gonzalez-Melendi et al. 2000; Raska 2003). In fact, in spite of efforts to fit nucleolar ultrastructure into this simple tripartite model, there is considerable variation in organization between kingdoms, species, cell types and individual cells (see Fig. 1). In plants, the DFC is typically much more extensive than in animals, and often does not stain any more intensely than the GC; this distinction is also difficult to make in S. cerevisiae. Nucleoli may be associated with the nuclear membrane (as in S. cerevisiae), heterochromatic ‘knobs’, or other nuclear bodies.
Pol I transcription begins as telophase ends. Small clusters of potentiated rRNA genes become active and associate with factors from the ‘pre-nucleolar bodies’ (PNBs) which contain a subset of nucleolar proteins and RNAs that had been dispersed on entry into mitosis, a process known as nucleologenesis. As pre-rRNA processing is activated by transcription-dependent interactions between a Pol I TF and elements of the processing machinery (Kopp et al. 2007), it is possible that transcription drives nucleologenesis by stimulating processing as well as by direct control of rRNA synthesis. Inactive rDNA is visible as peripheral knobs or internal foci, or may be interspersed with active rDNA within the nucleolus (Shaw et al. 1995; Carmo-Fonseca et al. 2000; Kalmorova et al. 2007); rDNA in peripheral knobs can be recruited as active genes when required (Highett et al. 1993).
Nucleolar structure as an evolutionary adaptation of eukaryotes
There is a strong correlation between nucleolar size and rate of rDNA transcription. Formation of nucleoli also requires Pol I: when constructs containing S. cerevisiae rRNA genes connected to Pol II promoters are transferred to different locations in the genome and transcribed, full nucleoli do not form, although ribosomes are produced (Oakes et al. 1999). However, there is evidence to suggest that the form of the nucleolus is an evolutionary adaptation to increase efficiency of ribosome synthesis, rather than being purely a consequence of transcription. For example, X. laevis oocyte or mammalian nucleoli can be dissociated without stopping rDNA transcription (Gonda et al. 2003; Yu et al. 2006), suggesting that transcription-driven self-assembly cannot fully explain nucleolar structure. Similarly, transcription occurs without formation of nucleoli in Archaea and other prokaryotes (Omer et al. 2000). rRNA transcription also occurs in the absence of visible nucleoli in certain simple eukaryotes and yeast strains with few rRNA genes (Nierras et al. 1997). Archaeal transcription is controlled in a relatively simple manner by DNA-binding Hmt proteins, which are thought to have evolved into histones during the emergence of eukaryotes, when most chromatin-remodelling machinery also evolved (Ouzounis and Kyrpides 1996, Malik and Henikoff 2003). So the evolution of nucleoli may also have depended upon the evolution of more sophisticated mechanisms of chromatin regulation.
The emergence of the nucleolar structure is controlled by chromatin
During nucleologenesis not all NORs are necessarily activated. If many are present, some will typically be entirely inactive, whilst only a subset of the genes on the others will be transcribed. Only rarely are all rRNA genes active simultaneously (Sogo et al. 1984). This activation can vary during differentiation as seen in comparisons of primary and adventitious onion roots (Hasterok and Maluszynska 2000). rRNA genes are assumed to be identical in sequence but differ in their chromatin organisation, which can render them active or inactive (Grummt and Pikaard 2003). They can also exist in a poised or potentiated state in which their chromatin is open and available for transcriptional activation, but remains untranscribed (reviewed by Huang et al. 2006). Thus, there are at least three different chromatin states, with active genes in an open, accessible arrangement, inactive genes in a condensed arrangement and potentially active genes with a more dynamic nucleosomal arrangement than inactive genes (Stefanovsky and Moss 2006; Jones et al. 2007). During mitosis NORs are compacted into a condensed organisation, although they are still ∼tenfold less compact than most the chromosome (Heliot et al. 1997), and so appear as secondary constrictions in metaphase chromosomes (McClintock 1934). NORs which were active in the preceding cell cycle can be recognised by positive AgNOR staining due to the retention of proteins involved in transcriptional activity (Roussel et al. 1994).
The level of Pol I association with these chromatin states may depend upon the level of cytosine methylation of the rDNA, remodelling of promoter chromatin, differences in histone variants and modifications at the promoters and coding sequences (reviewed by Grummt and Pikaard 2003). rDNA can retain the resulting levels of Pol I association through mitosis, allowing epigenetic inheritance of activity state (Conconi et al. 1989; Roussel et al. 1996; Grummt 2007). The importance of chromatin state is seen particularly well in the phenomenon of nucleolar dominance in hybrid organisms in which the NORs of one genome are repressed by another (Santoro 2005; Preuss and Pikaard 2007). In a given hybrid, the same genome will be dominant whether maternal or paternal, although this can be modulated by other loci, which could have chromatin-modifying roles (Chen et al. 1998; Pontes et al. 2003; Lewis et al. 2004). Some mechanisms of nucleolar dominance are also involved in the control of rDNA transcription in non-hybrid organisms. rRNA genes can show different levels of cytosine methylation, (in Xenopus Bird et al. 1981; and wheat, Flavell et al. 1988), and this can be an essential determinant of nucleolar dominance, as inhibition of such methylation can reduce the strength of nucleolar dominance (reviewed Chen and Pikaard 1997). rDNA methylation may affect rDNA by recruiting DNMTs to methylated promoters (Majumder et al. 2006).
However, rDNA methylation is only one aspect of chromatin which can be involved in nucleolar dominance (Pikaard 1999). It is not involved in nucleolar dominance in X. laevis (La Volpe et al. 1983) and tobacco (in which rDNA is hypomethylated; Kovarik et al. 2000), and DNA is barely methylated in Drosophila. In these cases, other aspects of chromatin organisation must be involved (Pennock and Reeder 1984; Avramova 2002), especially different histone modifications. This is well demonstrated by reversion of nucleolar dominance following histone methyltransferase inhibition in onion (de la Torre et al. 1991). HMT inhibition can also increase rDNA methylation, reduce histone acetylation and cause chromatin to condense (Thompson and Flavell 1988; Eden et al. 1998; Jones et al. 1998; Lim et al. 2000) suggesting that these mechanisms operate in a concerted manner. Association of active DNA with permissive marks may also involve repackaging of transcribed regions with covalently modified replication-independent histone variants. This mechanism was first described in rDNA but is involved in wider aspects of epigenetic inheritance (Henikoff and Ahmad 2005; Schwartz and Ahmad 2005).
Chromatin controls nucleolar structure by regulating transcription, and by overall rDNA packaging
Above, it was described how active and inactive rRNA genes differ in DNA methylation, histone variant incorporation and histone modification. These processes also control Pol II transcription (reviewed by Kouzarides 2007 and Rando 2007). However, the protein complexes which establish them are not all shared between Pol I and Pol II, and this may allow rRNA genes to be regulated separately from those transcribed by other polymerases. Well-characterised examples of these include NoRC in mammals, the HDT1/HDA6 complex in Arabidopsis, and Sir2p in S. cerevisiae (Table 1). Although from different kingdoms and apparently unrelated, Pol I transcription factors do share common components such as TATA binding protein (TBP) (see, e.g. Grummt 2003). In S. cerevisiae, another interesting feature is that histone H3/H4 dimers can regulate transcription not through chromatin organisation but as components of a Pol I-associated complex, upstream activating factor (UAF; Keener et al. 1997), and reduced H3 synthesis causes dramatic reductions in Pol I transcription (Tongaonkar et al. 2005). This indicates additional direct roles for histones in control of rRNA synthesis.
The role of vertebrate UBF is particularly interesting as it demonstrates that rDNA can direct nuclear reorganisation both through transcription and independently of it. UBF binds across rDNA loci throughout the coding sequence and in the intergenic spacers. UBF-binding sequences transferred elsewhere in the genome are able to recruit UBF and other factors to form ‘pseudo-NORs’ which are transcriptionally inactive, but resemble some aspects of NORs (reviewed Moss et al. 2007). They form secondary constrictions at mitosis (O’Sullivan et al. 2002; Mais et al. 2005), and recruit factors required for rDNA transcription and processing (Prieto and McStay 2007). UBF also acts in such an ‘architectural’ manner in amplified rDNA in X. laevis oocytes (Mais and Scheer 2001) but can also function as a transcription factor (TF), possibly by facilitating promoter escape (Panov et al. 2006). Perhaps because it is able to function both as a TF and as a determinant of DNA organisation, control of UBF has emerged as a major factor in rDNA regulation. For example, it undergoes cell cycle-specific acetylation which is necessary for Pol I binding (Meraner et al. 2006).
In contrast to rDNA control in vertebrates, no protein with the DNA-binding activity of UBF has yet been found in other organisms. Instead, studies of nucleolar dominance in plants suggest that rDNA organization and activity are controlled by a combination of DNA methylation, histone modification, and chromatin remodelling. These factors feed back upon each other to produce a ‘self-reinforcing loop’, which controls rRNA expression within species as well as in hybrids (Lawrence and Pikaard 2004; McStay 2006). In a hybrid of two Arabidopsis species, nucleolar dominance was shown to involve H3K9me2 at inactive, hypermethylated NORs and H3K9ac/H3K4me/H3K14ac and histone H4 tetra-acetylation of inactive, hypomethylated NORs (Lawrence et al. 2004). This required histone deacetylation by HDA6. Point mutations in HDA6 released transgenic and endogenous repetitive elements from silencing and caused H4 hyperacetylation and H3K4 hypermethylation at the 25S rRNA gene resulting in low levels of DNA hypomethylation and chromatin decondensation. Knock-down of HDA6 caused a complete loss of nucleolar dominance, and DNA hypomethylation and increased H3K9ac, H3K14ac, H3K4me3 and H4 tetra-acetylation of previously suppressed NORs (Earley et al. 2006). Purified HDA6 counteracted HAT-mediated H3K14, H4K5 and H4K12 acetylation in vitro, suggesting a direct effect, with other changes in modification occurring downstream or through histone replacement. Aufsatz et al. (2002) proposed that this also required establishment of methylation by other proteins, as HDA6 acts at symmetrical cytosine sites rather than the asymmetric sites most prone to de novo methylation.
Whilst histone deacetylases from several species have been shown to regulate rDNA and to be located in the nucleolus, there is much less evidence for nucleolar histone acetyltransferases. Those known to act in the nucleolus also function at Pol II-transcribed genes, e.g. the H4K16-specific Sas2p/PCAF of yeast (Meijsing and Ehrenhofer-Murray 2001). Additionally, certain enzymes acetylate other proteins to control nucleolar activity. Tip60 upregulates transcription by acetylating UBF (Halkidou et al. 2004), whilst the MYST-family acetyltransferase, Esa2p, has the opposite effect in S. cerevisiae, contributing to Sir2-mediated gene silencing (Clarke et al. 2006). As many organisms have multiple unattributed HATs in their genomes it remains possible that some are rDNA-specific.
rDNA transcription underlies nucleolar-chromatin organisation, but conversely, NORs can also be regulated by chromatin at other loci. Centromeric heterochromatin is involved in yeast nucleologenesis (Pluta et al. 1995) and exogenous heterochromatic NORs alter mitotic interactions of endogenous NORs in rye (Caperta et al. 2002). Heterochromatin on other chromosomes can also affect NORs: introduction of heterochromatic B-chromosomes from rye into wheat lines caused endogenous NORs to become more compacted and less active (Morais-Cecilio et al. 2000). It is known that heterochromatin is involved in maintaining overall nuclear organisation in Drosophila (Csink and Henikoff 1996) and it is therefore possible that heterochromatin could regulate nucleolar organisation in a similar manner.
Another good example of the potential importance in the regulation of nucleolar chromatin is seen in the Sir2 (Silent information regulator2)-like family of proteins (sirtuins). These were identified in S. cerevisiae as rDNA-binding proteins which use NAD+ reduction to fuel H3/H4 deacetylation during heterochromatin formation and, thus, repress transcription (Table 1; Gotta et al. 1997; Imai et al. 2000). Sir2p has homologues with roles in nucleolar chromatin throughout eukaryotes, although the RENT complex with which it associates is not conserved. In humans, it has recently been shown that a homologue, SIRT1, instead interacts with a complex termed eNoSC (Murayama et al. 2008). This also contains SUV39H1, which mediates the repressive histone modification, H3K9 dimethylation, and is argued to allow repression of rDNA transcription in response to falling cellular energy levels (Murayama et al. 2008).
In many organisms, sirtuin activity is linked to longevity. This could reflect the fact that sirtuins can mediate cellular energy levels, possibly in the same way as caloric restriction, which is known to increase longevity. Sir2p also controls aging in S. cerevisiae, perhaps by forming inactive rDNA into heterochromatin to prevent recombination, which produces small loops of rDNA termed minicircles, which are believed to be toxic to cells (Gottlieb et al. 1989; Fritze et al. 1997; Parsons et al. 2003; Kobayashi et al. 2004). Sirtuins may have roles in genome stability in other organisms (e.g. rice; Huang et al. 2007), so this mechanism may not be restricted to fungi. However, some sirtuins seem to mediate aging pathways through down-regulation of p53 (Vaziri et al. 2001). Sirtuins also act upon other substrates entirely, including tubulin and acetyl-CoA synthetase (North et al. 2003; Hallows et al. 2006). Sirtuins have been shown to interact with nucleoli to activate Pol I transcription (Ford et al. 2006) by mechanisms which remain unclear.
The level of rRNA synthesis at active genes can also be controlled by the loading rate of Pol I complexes at the promoters and by elongation efficiency (Stefanovsky and Moss 2006). Elongation control has been argued to be particularly important for controlling rRNA synthesis in human cells as control through elongation is more significant than that of initiation; furthermore, alterations in elongation levels are sufficient to explain changes to cell growth in cancer lines (Moss et al. 2007 and references therein). Diminished capacity for transcribing to the end of the gene may also explain the reduced rRNA synthesis of Drosophila rDNA inserted into retrotransposable elements (Ye and Eickbush 2006). Histone modification and UBF phosphorylation may also control transcription via elongation, perhaps by accumulation of permissive histone modifications or replication-independent histones such as H3.3 in the 3′-parts of the genes. rDNA-associated histones may also be required for control of elongation (Tongaonkar et al. 2005).
The different systems which control rDNA transcription may act in varying combinations in different cells or species. For example, the Arabidopsis Cape Verde Island (Cvi) ecotype has 80% as many rRNA genes as Colombia ecotype (Col) but more than twice the proportion of these are hypermethylated and thus inactive (Riddle and Richards, 2002). As it is unlikely that Cvi requires less than 40% of the number of rRNA molecules that Col does, Cvi probably controls rRNA content by methylation-mediated repression of most of its rRNA genes, whilst Col has more active genes but controls their activity through other methods (Riddle and Richards 2002). These could include Pol I loading and extension rates.
The three states of nucleolar chromatin: roles for novel components?
Nucleolar chromatin is also likely to include further components of rDNA-specific enzyme complexes, nucleolar-specific histone modifications or histone variants. It is possible that such components contribute to the role of chromatin in regulating nucleolar structure as well as transcription.
Nucleolar histone variants
Histones with variant sequences are obvious candidates for nucleolus-specific chromatin components. This has already been shown in the case of a potential nucleolar H2A variant in human and possible enrichment of an H2A in carp nucleoli (Allis et al. 1982; Alvarez et al. 2006). As many organisms have multiple histone variants, it is possible that rDNA may also associate with nucleolus-specific histone variants more generally. This would parallel the association of histone variants with other nuclear domains such as centromeres, telomeres and Barr bodies (Table 2). Linker H1 histones are also enriched in the nucleolus in lily and Arabidopsis (Tanaka et al. 1999; Pendle et al. 2005). Linker histones repress inactive human rDNA and maintain epigenetic control of rDNA in tobacco, indicating roles in chromatin domain stabilisation within NORs (Slusarczyk et al. 2003). Linker histones interact with nucleolin in human cells (Erard et al. 1988; see below) and compete with Pol I for upstream binding sites in yeast (Kermekchiev et al. 1997) suggesting that they could link the maintenance of chromatin domains with transcription. In contrast with the reports of nucleolar-specific histone variants, nucleolar-specific histone modifications are conspicuous by their absence (Table 2), although various H4 acetylations are enriched at replicating NORs in the plants V. faba and A. thaliana (Houben et al. 1996; Jasencakova et al. 2000). The combinatorial complexity of the histone code suggests that other modifications could have roles in NOR replication or other nucleolar functions, but these may vary between species.
Nucleolar histone chaperones
Several nucleolar proteins also act as histone-binding proteins. These include nucleolin and nucleophosmin (also called B23, NO38 and numatrin) and novel proteins such as NO29 (Sharma 2004) and the human peptidyl prolyl cis–trans isomerase protein, FKBP (Kuzuhara and Horikoshi 2004). Nucleolin is a DNA-binding protein which is essential for Pol I transcription (Richards et al. 2007) and nucleolar integrity (Pontvianne et al. 2007). It is required for H1-related chromatin compaction (Erard et al. 1988) and the SWI/SNF2-remodelling of H2A/H2B dimers (Angelov et al. 2006) and, as an AgNOR protein, remains associated with NOR chromatin through the cell cycle (Roussel et al. 1994). Because it has many other functions (reviewed Mongelard and Bouvet 2007), nucleolin provides potential links between nucleolar-chromatin organisation and other nuclear processes.
Nucleophosmin (NPM) is part of the small nucleophosmin/nucleoplasmin histone chaperone family (Frehlick et al. 2007), and regulates rRNA transcription. NPM interacts with FRGY2, which can mediate the disassembly of Xenopus nucleoli whilst not disrupting rRNA transcription (Gonda et al. 2006), but how this links with its histone-binding activities remains unclear. Nucleophosmin associates with H3/H4 dimers via its N-terminal domain, and, via a C-terminal acidic tract, with H2A/H2B, in the assembly of nucleosomes (Okuwaki et al. 2001). It may be important for the binding and reassembly of nucleosomes disrupted during the high rates of transcription which occur at rDNA. The related NPM2 (nucleoplasmin) is also a histone chaperone, suggested by structural studies to be specific for H3/H4 tetramers (Namboodiri et al. 2004). As with nucleolin, nucleophosmin acts in many different ways—its deposition is linked to poly-adenylation, for example (Palaniswamy et al. 2006). Hence, it too could integrate histone remodelling with a range of other nucleolar processes.
Nucleolar roles of small RNAs
In many organisms, silencing of repetitive DNA often occurs through mechanisms involving small RNA species and it has recently been shown that the siRNA pathway in Drosophila plays a structural role by forming rDNA into inactive heterochromatin, which blocks recombination (Peng and Karpen 2007). This occurs via H3K9 dimethylation of rDNA and inactive microsatellites. Disruption of the pathway caused DNA reorganisation, including dissociation of the nucleolus into dispersed rDNA foci. Non-nucleolar rDNA also became excised from the chromosomes. As mutation of a ligase essential for the homologous recombination pathway rescued this, siRNA-mediated heterochromatinisation seems to be involved in preventing unwanted homologous recombination.
These important roles for siRNAs in nucleolar integrity may be specific to Drosophila, perhaps due to the lack of DNA methylation in this organism, but it remains an intriguing possibility that small RNA pathways could be involved in regulating rDNA in other species. siRNAs are encoded between non-NOR 5S rRNA genes of Arabidopsis (Llave et al. 2002), and could also regulate nucleolar transcription or structure. hda6 mutants were derepressed in transcription at centromeric repeats (Athila; May et al. 2005) through the RNA-directed DNA methylation pathway; hence the effect on NORs might also involve small RNAs (Aufsatz et al. 2002). Both nucleoli and Cajal Bodies of Arabidopsis are involved in maturation of small RNAs (Shaw et al. 1998; Pontes et al. 2006; Li et al. 2006a). Small RNAs are also matured in small ncRNA-processing bodies in which ASSYMETRIC LEAVES1 (AS1) and AS2 interact with the rRNA gene-regulating HDT proteins to regulate leaf development (Ueno et al. 2007), although HDTs have been proposed to be solely nucleolar (Lawrence et al. 2004) or found throughout the nucleus (Zhou et al. 2004). These structures may represent the perinucleolar microRNA processing bodies to which DCL1 localises (Song et al. 2007). The observation of a direct interaction between an ncRNA-processing pathway and nucleolar-chromatin-remodelling proteins suggests that chromatin could also play important roles in non-canonical nucleolar functions. Little is known about the roles of small RNAs in mammalian nucleoli, although miRNA-206 has been shown to bind ribosomal proteins in the GC in rat (Politz et al. 2006).
Further chromatin components identified by MS
Finally, mass spectrometry has been used to determine the proteomes of nucleoli from human and Arabidopsis culture cells (Andersen et al. 2002; Scherl et al. 2002; Pendle et al. 2005; Lam et al. 2007) and has identified potential nucleolar chromatin components which remain uncharacterised. MS analyses of the human nucleolus have to date identified 5 linker histones, 5 H2A isoforms, H2A.F/Z, 6 H2B isoforms, 2 H3 isoforms, a HAT-B subunit, HDAC 1 and 2, a FACT component, a CAF1 subunit, three chromobox proteins, and a splice-variant of a bromodomain-like protein: a total of 28 out of c. 700 (www.lamondlab.com/NOPdb/). The Arabidopsis nucleolar proteome so far contains two linker histones, six H2A isoforms, H2A.F/Z, five H2B isoforms, three H3.1 isoforms and one H3.2, at least one H4, a FACT component, an HDAC, three putative HAT components and the four HDAC-like HDT proteins (Pendle et al. 2005; McKeown et al. unpublished data), i.e. 27 out of c. 650. Hence, the ratios of chromatin proteins in both proteomes appear to be similar.
Such studies emphasise that many chromatin components may remain to be discovered. However, current studies of the dynamics of nuclear proteins suggest that few nuclear proteins are likely to be wholly excluded from the nucleolus, so the nucleolar proteome should be thought of as part of a continuum with the nucleoplasm. Different histone isoforms may associate with nuclear regions in different proportions, for example, and understanding nucleolar chromatin will require a consideration both of instances of ‘nucleolar isoforms’ and subtler changes to the whole histone profile. Better quantitation of mass spectrometry data may provide the information necessary to elucidate this.
Conclusions
Studies of nucleoli illustrate the way in which chromatin acts at the interface between structure and function. Different organisms use different sets of chromatin-remodelling proteins to deploy components, often shared with Pol II-transcribed genes, to selectively activate a subset of rRNA genes. The level of transcription is controlled by chromatin-mediated loading of the Pol I complex at promoters and this, in turn, modulates the structure of the nucleolus. Nucleolar structure is also maintained and modulated by heterochromatinisation of inactive rDNA by siRNAs or sirtuins or both. Chromatin at other NORs or B-chromosomes may also affect transcription.
This level of regulation of chromatin structure, and the organisation of the resultant nucleolus, may be necessary to allow transcription from repeated DNA without activating silencing mechanisms. In at least some instances, it may also be connected with the maintenance of rDNA in different epigenetic states through replication. Finally, organising rRNA transcription and processing into a defined compartment containing different sub-regions may also allow other cellular processes to take advantage of this compartmentalization, especially those requiring RNA-processing steps.
The nucleolus represents an example of the way activity can control architecture, yet also the manner in which this architecture is modulated to ensure the efficiency of overall nuclear function; this might be described as a system of ‘modulated self-organization’. In the future, it should be expected that this model will be extended to incorporate components of uncertain function, such as the HDACs and HATs identified in proteomic screens, and small RNA pathways, and to explain what role the histone-interactions of well-known proteins such as nucleolin and nucleophosmin may play. The possibility that particular histone modifications and variants may have specialised roles within the nucleolus should also be explored. In this way, the nucleolus will continue to shed light on the relationship between gene function and nuclear organisation in eukaryote cells, and hence on the relationship between structure and activity in biological systems.
Abbreviations
- DNMT:
-
DNA methyltransferase
- HAT:
-
histone acetyltransferase
- HDAC:
-
histone deacetylase
- HMT:
-
histone methyltransferase
- MS:
-
mass spectrometry
- NOR:
-
nucleolar organiser region
- rDNA:
-
ribosomal DNA
- rRNA:
-
ribosomal RNA
- UBF:
-
upstream binding factor.
References
Allis CD, Ziegler YS, Gorovsky MA, Olmsted JB (1982) A conserved histone variant enriched in nucleoli of mammalian cells. Cell 31:131–136
Alvarez M, Quezada C, Molina A, Krauskopf M, Vera MI, Thiry M (2006) Ultrastructural changes of the carp (Cyprinus carpio) hepatocyte nucleolus during seasonal acclimatization. Biol Cell 98:457–463
Andersen JS, Lyon CE, Fox AH, Leung AKL, Lam YW, Steen H, Mann M, Lamond AI (2002) Directed proteomic analysis of the human nucleolus. Curr Biol 12:1–11
Angelov D, Bondarenko VA, Almagro S, Menoni H, Mongelard F, Hans F, Mietton F, Studitsky VM, Hamiche A, Dimitrov S, Bouvet P (2006) Nucleolin is a histone chaperone with FACT-like activity and assists remodelling of nucleosomes. EMBO J 25:1669–1679
Aufsatz W, Mette MF, van der Winden J, Matzke M, Matzke AJM (2002) HDA6, a putative histone deacetylase needed to enhance DNA methylation induced by double-stranded RNA. EMBO J 21:6832–6841
Avramova ZV (2002) Update on heterochromatin. Plant Physiol 129:40–49
Benecke A (2006) Chromatin code, local non-equilibrium dynamics, and the emergence of transcription regulatory programs. Eur Phys J E 19:353–366
Bird A, Taggart M, Macleod D (1981) Loss of rDNA methylation accompanies the onset of ribosomal gene activity in early development of X. laevis. Cell 26:381–390
Boggs BA, Connors B, Sobel RE, Chinault AC, Allis CD (1996) Reduced levels of histone H3 acetylation on the inactive X chromosome in human females. Chromosoma 105:303–309
Boisvert FM, van Koningsbruggen S, Navascues J, Lamond AI (2007) The multifunctional nucleolus. Nature Rev Mol Cell Biol 8:574–585
Brown SE, Szyf M (2007) Epigenetic programming of the rRNA promoter by MBD3. Mol Cell Biol 27:4938–4952
Caperta AD, Neves N, Morais-Cecilio L, Malho R, Viegas W (2002) Genome restructuring in rye affects the expression, organization and disposition of homologous rDNA loci. J Cell Sci 115:2839–2846
Carmo-Fonseca M, Mendes-Soares L, Campos I (2000) To be or not to be in the nucleolus. Nature Cell Biol 2:E107–E112
Cervantes MD, Xi XH, Vermaak D, Yao MC, Malik HS (2006) The CNA1 histone of the ciliate Tetrahymena thermophila is essential for chromosome segregation in the germline micronucleus. Mol Biol Cell 17:485–497
Chen ZJ, Pikaard CS (1997) Epigenetic silencing of RNA polymerase I transcription: a role for DNA methylation and histone modification in nucleolar dominance. Genes Dev 11:2124–2136
Chen ZJ, Comai L, Pikaard CS (1998) Gene dosage and stochastic effects determine the severity and direction of uniparental ribosomal RNA gene silencing (nucleolar dominance) in Arabidopsis allopolyploids. Proc Natl Acad Sci USA 95:14891–14896
Chen DY, Belmont AS, Huang S (2004) Upstream binding factor association induces large scale chromatin decondensation. Proc Natl Acad Sci USA 101:15106–15111
Clarke AS, Samal E, Pillus L (2006) Distinct roles for the essential MYST family HAT Esa1p in transcriptional silencing. Mol Biol Cell 17:1744–1757
Conconi A, Widmer RM, Koller T, Sogo JM (1989) Two different chromatin structures coexist in ribosomal RNA genes throughout the cell cycle. Cell 57:753–761
Costanzi C, Stein P, Worrad DM, Schultz RM, Pehrson JR (2000) Histone macroH2A1 is concentrated in the inactive X chromosome of female preimplantation mouse embryos. Development 127:2283–2289
Csink AK, Henikoff S (1996) Genetic modification of heterochromatic association and nuclear organization in Drosophila. Nature 381:529–531
de la Torre C, Giminez-Abian JF, Gonzalez-Fernandez A (1991) Dominance of a NOR (nucleolar organizer region) over its allele and over its sister NOR after asymmetric 5-azacytidine substitution in plant chromosomes. J Cell Sci 100:667–674
Earley K, Lawrence RJ, Pontes O, Reuther R, Encisco AJ, Silva M, Neves N, Gross M, Viegas W, Pikaard CS (2006) Erasure of histone acetylation by Arabidopsis HDA6 mediates large-scale gene silencing in nucleolar dominance. Genes Dev 20:1283–1293
Eden S, Hashimshony T, Keshet I, Cedar H, Thorne AW (1998) DNA methylation models histone acetylation. Nature 394:842
Erard MS, Belenguer P, Caizerguesferrer M, Pantaloni A, Amalric F (1988) A major nucleolar protein, nucleolin, induces chromatin decondensation by binding to histone H1. Eur J Biochem 175:525–530
Espada J, Esteller M (2007) Epigenetic control of nuclear architecture. Cell Mol Life Sci 64:449–457
Fernandez-Capetillo O, Lee A, Nussenzweig M, Nussenzweig A (2004a) H2AX: the histone guardian of the genome. DNA Repair 3:959–967
Fernandez-Capetillo O, Allis CD, Nussenzweig A (2004b) Phosphorylation of histone H2B at DNA double-strand breaks. J Exp Med 199:1671–1677
Flavell RB, O’Dell M, Thompson WF (1988) Regulation of cytosine methylation in ribosomal DNA and nucleolus organizer expression in wheat. J Mol Biol 204:523–534
Ford E, Voit R, Liszt G, Magin C, Grummt I, Guarente L (2006) Mammalian Sir2 homolog SIRT7 is an activator of RNA polymerase I transcription. Genes Dev 20:1075–1080
Frehlick LJ, Eirin-Lopez JM, Ausio J (2007) New insights into the nucleophosmin/nucleoplasmin family of nuclear chaperones. Bioessays 29:49–59
Fritze CE, Verschueren K, Strich R, Esposito RE (1997) Direct evidence for SIR2 modulation of chromatin structure in yeast rDNA. EMBO J 16:6495–6509
Fuchs J, Demidov D, Houben A, Schubert I (2006) Chromosomal histone modification patterns—from conservation to diversity. Trends Plant Sci 11:199–208
Galy V, Olivo-Marin JC, Scherthan H, Dove V, Rasalou N, Nehrbass U (2000) Nuclear pore complexes in the organisation of silent telomeric chromatin. Nature 403:108–112
Gonda K, Fowler J, Katoku-Kikyo N, Haroldsen J, Wudel J, Kikyo N (2003) Reversible disassembly of somatic nucleoli by the germ cell proteins FRGY2a and FRGY2b. Nature Cell Biol 5:205–210
Gonda K, Wudel J, Nelson D, Katoku-Kikyo N, Reed P, Tamada H, Kikyo N (2006) Requirement of the protein B23 for nucleolar disassembly induced by the FRGY2a family proteins. J Biol Chem 281:8153–8160
Gonzalez-Melendi P, Beven A, Boudonck K, Abranches R, Wells B, Dolan L, Shaw P (2000) The nucleus: a highly organized but dynamic structure. J Microsc 198:199–207
Gotta M, Strahl-Bolsinger S, Renauld H, Laroche T, Kennedy BK, Grunstein M, Gasser SM (1997) Localization of Sir2p: the nucleolus as a compartment for silent information regulators. EMBO J 16:3243–3255
Gottlieb S, Esposito RE (1989) A new role for a yeast transcriptional silencer gene, Sir2, in regulation of recombination in ribosomal DNA. Cell 56:771–776
Grummt I (2003) Life on a planet of its own: regulation of RNA polymerase I in the nucleolus. Genes Dev 17:1691–1702
Grummt I (2007) Different epigenetic layers engage in complex crosstalk to define the epigenetic state of mammalian rRNA genes. Hum Mol Gen 16:R21–R27
Grummt I, Pikaard CS (2003) Epigenetic silencing of RNA polymerase I transcription. Nature Rev Mol Cell Biol 4:641–649
Halkidou K, Logan IR, Cook S, Neal DE, Robson CN (2004) Putative involvement of the histone acetyltransferase Tip60 in ribosomal gene transcription. Nucl Acids Res 32:1654–1665
Hallows WC, Lee S, Denu JM (2006) Sirtuins deacetylate and activate mammalian acetyl-CoA synthetases. Proc Natl Acad Sci USA 103:10230–10235
Hancock R (2004) Internal organisation of the nucleus: assembly of compartments by macromolecular crowding and the nuclear matrix model. Biol Cell 96:595–601
Hasterok R, Maluszynska J (2000) Different rRNA gene expression in primary and adventitious roots of Allium cepa. Folia Hist Cytobiol 38:181–184
Heliot L, Kaplan H, Lucas L, Klein C, Beorchia A, Doco-Fenzy M, Menager M, Thiry M, O’Donohue M-F, Ploton D (1997) Electron tomography of metaphase nucleolar organizer regions: evidence for a twisted-loop organization. Mol Biol Cell 8:2199–2216
Henikoff S, Ahmad K (2005) Assembly of variant histones into chromatin. Ann Rev Cell Dev Biol 21:133–153
Hernandez-Verdun D (2005) Tracking the interactions of rRNA processing proteins during nucleolar assembly in living cells. Medec Sci 21:1025–1027
Hernandez-Verdun D, Roussel P, Gebrane-Younes J (2002) Emerging concepts of nucleolar assembly. J Cell Sci 115:2265–2270
Highett MI, Rawlins DJ, Shaw PJ (1993) Different patterns of rDNA distribution in Pisum sativum nucleoli correlate with different levels of nucleolar activity. J Cell Sc 104:843–852
Hiscox JA (2007) RNA viruses: hijacking the dynamic nucleolus. Nature Rev Microbiol 5:119–127
Houben A, Belyaev ND, Turner BM, Schubert I (1996) Differential immunostaining of plant chromosomes by antibodies recognizing acetylated histone H4 variants. Chromosome Res 4:191–194
Huang J, Moazed D (2003) Association of the RENT complex with nontranscribed and coding regions of rDNA and a regional requirement for the replication fork block protein Fob1 in rDNA silencing. Genes Dev 17:2162–2176
Huang S, Rothblum LI, Chen DY (2006) Ribosomal chromatin organization. Biochem Cell Biol 84:444–449
Huang LM, Sun QW, Qin FJ, Li C, Zhao Y, Zhou DX (2007) Down-regulation of a SILENT INFORMATION REGULATOR2-related histone deacetylase gene, OsSRT1, induces DNA fragmentation and cell death in rice. Plant Physiol 144:1508–1519
Idei S, Kondo K, Turner BM, Fukui K (1996) Tomographic distribution of acetylated histone H4 in plant chromosomes, nuclei and nucleoli. Chromosoma 105:293–302
Imai S-I, Armstrong CM, Keaberlein M, Guarente L (2000) Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature 403:795–800
Jasencakova Z, Meister A, Walter J, Turner BM, Schubert I (2000) Histone H4 acetylation of euchromatin and heterochromatin is cell cycle dependent and correlated with replication rather than with transcription. Plant Cell 12:2087–2100
Jones PL, Veenstra GJC, Wade PA, Vermaak D, Kass SU, Landsberger N, Strouboulis J, Wolffe AP (1998) Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nature Gen 19:187–191
Jones HS, Kawauchi J, Braglia P, Alen CM, Kent NA, Proudfoot NJ (2007) RNA polymerase I in yeast transcribes dynamic nucleosomal rDNA. Nature Struct Mol Biol 14:123–130
Kalmarova M, Smirnov E, Masata M, Koberna K, Ligasova A, Popov A, Raska I (2007) Positioning of NORs and NOR-bearing chromosomes in relation to nucleoli. J Struct Biol 160:49–56
Keener J, Dodd JA, Lalo D, Nomura M (1997) Histones H3 and H4 are components of upstream activation factor required for the high-level transcription of yeast rDNA by RNA polymerase I. Proc Natl Acad Sci USA 94:13458–13462
Kermekchiev M, Workman JL, Pikaard CS (1997) Nucleosome binding by the polymerase I transactivator upstream binding factor displaces linker histone H1. Mol Cell Biol 17:5833–5842
Kimura A, Matsubara K, Horikoshi M (2005) A decade of histone acetylation: marking eukaryotic chromosomes with specific codes. J Biochem 138:647–662
Kobayashi T, Horiuchi T, Tongaonkat P, Vu L, Nomura M (2004) SIR2 regulates recombination between different rDNA repeats, but not recombination within individual rRNA genes. Cell 117:441–453
Kohlmaier A, Savarese F, Lachner M, Martens J, Jenuwein T, Wutz A (2004) A chromosomal memory triggered by Xist regulates histone methylation in X inactivation. PLoS Biol 2:991–1003
Kopp K, Gasiorowski JZ, Chen D, Gilmore R, Norton JT, Wang C, Leary DJ, Chan EKL, Dean DA, Huang S (2007) Pol I transcription and pre-rRNA processing are coordinated in a transcription-dependent manner in mammalian cells. Mol Biol Cell 18:394–403
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705
Kovarik A, Koukalova B, Lim KY, Matyasek R, Lichtenstein CP, Leitch AR, Bezdek M (2000) Comparative analysis of DNA methylation in tobacco heterochromatic sequences. Chrom Res 8:527–541
Kurakin A (2005) Self-organization versus Watchmaker: stochastic dynamics of cellular organization. Biol Chem 386:247–254
Kuzuhara T, Horikoshi M (2004) A nuclear FK506-binding protein is a histone chaperone regulating rDNA silencing. Nature Struct Mol Biol 11:275–283
La Volpe A, Taggart M, McStay B, Bird A (1983) DNaseI-hypersensitive sites at promoter-like sequences in the space of Xenopus laevis and Xenopus borealis ribosomal DNA. Nucl Acids Res 11:5361–5380
Lam YW, Lamond AI, Mann M, Andersen JS (2007) Analysis of nucleolar protein dynamics reveals the nuclear degradation of ribosomal proteins. Curr Biol 17:749–760
Lamond AI, Sleeman JE (2003) Nuclear substructure and dynamics. Curr Biol 13:R825–R828
Lawrence RJ, Pikaard CS (2004) Chromatin turn ons and turn offs of ribosomal RNA genes. Cell Cycle 3:880–883
Lawrence RJ, Earley K, Pontes O, Silva M, Chen ZJ, Neves N, Viegas W, Pikaard CS (2004) A concerted DNA methylation/histone methylation switch regulates rRNA gene dosage control and nucleolar dominance. Mol Cell 113:599–609
Lewis MS, Cheverud JM, Pikaard CS (2004) Evidence for nucleolus organizer regions as the units of regulation in nucleolar dominance in Arabidopsis thaliana interecotype hybrids. Genetics 167:931–939
Li CF, Pontes O, El-Shami M, Henderson IR, Bernatavichute YV, Chan SWL, Lagrange T, Pikaard CS, Jacobsen SE (2006a) An ARGONAUTE4-containing nuclear processing center colocalized with Cajal bodies in Arabidopsis thaliana. Cell 126:93–106
Li CH, Mueller JE, Bryk M (2006b) Sir2 represses endogenous polymerase II transcription units in the ribosomal DNA nontranscribed spacer. Mol Biol Cell 17:3848–3859
Li JW, Santoro R, Koberna K, Grummt I (2005) The chromatin remodelling complex NoRC controls replication timing of rRNA genes. EMBO J 24:120–127
Lim KY, Kovarik A, Matyasek R, Bezdek M, Lichtenstein CP, Leitch AR (2000) Gene conversion of ribosomal DNA in Nicotiana tabacum is associated with undermethylated, decondensed and probably active gene units. Chromosoma 109:161–172
Llave C, Kasschau KD, Rector MA, Carrington JC (2002) Endogenous and silencing-associated small RNAs in plants. Plant Cell 14:1605–1619
Lyon CE, Lamond AI (2000) The nucleolus. Curr Biol 10:R323–R323
Maggi LB, Weber JD (2005) Nucleolar adaptation in human cancer. Cancer Invest 23:599–608
Mais C, Scheer U (2001) Molecular architecture of the amplified nucleoli of Xenopus oocytes. J Cell Sci 114:709–718
Mais C, Wright JE, Prieto JL, Raggett SL, McStay B (2005) UBF-binding site arrays form pseudo-NORs and sequester the RNA polymerase I transcription machinery. Genes Dev 19:50–64
Majumder S, Ghoshal K, Datta J, Smith DS, Bai SM, Jacob ST (2006) Role of DNA methyltransferases in regulation of human ribosomal RNA gene transcription. J Biol Chem 281:22062–22072
Malik HS, Henikoff S (2003) Phylogenomics of the nucleosome. Nature Struct Biol 10:882–891
Marian CO, Bordoli SJ, Goltz M, Santarella RA, Jackson LP, Danilevskaya O, Beckstette M, Meeley R, Bass HW (2003) The maize Single myb histone 1 gene, Smh1, belongs to a novel gene family and encodes a protein that binds telomere DNA repeats in vitro. Plant Physiol 133:1336–1350
Martin C, Zhang Y (2005) The diverse functions of histone lysine methylation. Nature Rev Mol Cell Biol 6:838–849
May BP, Lippman ZB, Fang YD, Spector DL, Martienssen RA (2005) Differential regulation of strand-specific transcripts from Arabidopsis centromeric satellite repeats. PLoS Gen 1:705–714
McClintock B (1934) The relation of a particular chromosomal element to the development of the nucleoli in Zea mays. Z Zellforsch Mikrol Anat 21:294–328
McStay B (2006) Nucleolar dominance: a model for rRNA gene silencing. Genes Dev 20:1207–1214
Meijsing SH, Ehrenhofer-Murray AE (2001) The silencing complex SAS-I links histone acetylation to the assembly of repressed chromatin by CAF-I and Asf1 in S. cerevisiae. Genes Dev 15:3169–3182
Melese T, Xue Z (1995) The nucleolus—an organelle formed by the act of building a ribosome. Curr Op Cell Biol 7:319–324
Meraner J, Lechner M, Loidl A, Goralik-Schramel M, Voit R, Grummt I, Loidl P (2006) Acetylation of UBF changes during the cell cycle and regulates the interaction of UBF with RNA polymerase I. Nucl Acids Res 34:1798–1806
Mermoud JE, Popova B, Peters AHFM, Jenuwein T, Brockdorff N (2002) Histone H3 lysine 9 methylation occurs rapidly at the onset of random X chromosome inactivation. Curr Biol 12:247–251
Misteli T (2001) The concept of self-organization in cellular architecture. J Cell Biol 155:181–185
Mongelard F, Bouvet P (2007) Nucleolin: a multiFACeTed protein. Trends Cell Biol 17:80–86
Morais-Cecilio L, Delgado M, Jones RN, Viegas W (2000) Modification of wheat rDNA loci by rye B chromosomes: a chromatin organization model. Chrom Res 8:341–351
Moss T, Langlois F, Gagnon-Kugler T, Stefanovsky V (2007) A housekeeper with power of attorney: the rRNA genes in ribosome biogenesis. Cell Mol Life Sci 64:29–49
Murayama A, Ohmori K, Fujimara A, Minami H, Yasuzawa-Tanaka K, Kuroda T, Oie S, Daitoku H, Okuwaki M, Nagata K, Fukamizu A, Kimura K, Shimizu T, Yanagisawa J (2008) Epigenetic control or rDNA loci in response to intracellular energy status. Cell 133:627–639
Namboodiri VMH, Akey IV, Schmidt-Zachmann MS, Head JF, Akey CW (2004) The structure and function of Xenopus NO38-core, a histone chaperone in the nucleolus. Structure 12:2149–2160
Ng HH, Ciccone DN, Morshead KB, Oettinger MA, Struhl K (2003) Lysine-79 of histone H3 is hypomethylated at silenced loci in yeast and mammalian cells: a potential mechanism for position-effect variegation. Proc Natl Acad Sci USA 100:1820–1825
Nierras CR, Liebman SW, Warner JR (1997) Does S. cerevisiae need an organized nucleolus? Chromosoma 105:444–451
North BJ, Marshall BL, Borra MT, Denu JM, Verdin E (2003) The human Sir2 ortholog, SIRT2, is an NAD(+)-dependent tubulin deacetylase. Mol Cell 11:437–444
Oakes ML, Siddiqi I, Vu L, Aris J, Nomura M (1999) Transcription factor UAF, expansion and contraction of ribosomal DNA (rDNA) repeats, and RNA polymerase switch in transcription of yeast rDNA. Mol Cell Biol 19:8559–8569
Oakes ML, Siddiqi I, French SL, Vu L, Sato M, Aris JP, Beyer AL, Nomura M (2006) Role of histone deacetylase Rpd3 in regulating rRNA gene transcription and nucleolar structure in yeast. Mol Cell Biol 26:3889–3901
Okuwaki M, Matsumoto K, Tsujimoto M, Nagata K (2001) Function of nucleophosmin/B23, a nucleolar acidic protein, as a histone chaperone. FEBS Lett 506:272–276
Omer AD, Lowe TM, Russell AG, Ebhardt H, Eddy SR, Dennis PP (2000) Homologs of small nucleolar RNAs in archaea. Science 288:517–522
O’Sullivan AC, Sullivan GJ, McStay B (2002) UBF binding in vivo is not restricted to regulatory sequences within the vertebrate ribosomal DNA repeat. Mol Cell Biol 22:657–668
Ouzounis CA, Kyrpides NC (1996) Parallel origins of the nucleosome core and eukaryotic transcription from archeae. J Mol Evol 42:234–239
Palaniswamy V, Moraes KCM, Wilusz CJ, Wilusz J (2006) Nucleophosmin is selectively deposited on mRNA during polyadenylation. Nature Struct Mol Biol 13:429–435
Panov KI, Friedrich JK, Russell J, Zomerdijk J (2006) UBF activates RNA polymerase I transcription by stimulating promoter escape. EMBO J 25:3310–3322
Parsons XH, Garcia SN, Pillus L, Kadonaga JT (2003) Histone deacetylation by Sir2 generates a transcriptionally repressed nucleoprotein complex. Proc Natl Acad Sci USA 100:1609–1614
Payne C, Braun RE (2006) Histone lysine trimethylation exhibits a distinct perinuclear distribution in Plzf-expressing spermatogonia. Dev Biol 293:461–472
Pendle AF, Clark GP, Boon R, Lewandowska D, Lam YW, Andersen J, Mann M, Lamond AI, Brown JW, Shaw PJ (2005) Proteomic analysis of the Arabidopsis nucleolus suggests novel nucleolar functions. Mol Biol Cell 16:260–269
Peng JC, Karpen GH (2007) H3K9 methylation and RNA interference regulate nucleolar organization and repeated DNA stability. Nature Cell Biol 9:25–35
Pennock DG, Reeder RH (1984) In vitro methylation of HpaII sites in Xenopus laevis rDNA does not affect its transcription in oocytes. Nucl Acids Res 12:2225–2232
Percipalle P, Fomproix N, Cavellan E, Voit R, Reimer G, Kruger T, Thyberg J, Scheer U, Grummt I, Farrants AKO (2006) The chromatin remodelling complex WSTF-SNF2h interacts with nuclear myosin 1 and has a role in RNA polymerase I transcription. EMBO Reports 7:525–530
Pikaard CS (1999) Nucleolar dominance and silencing of transcription. Trends Plant Sci 4:478–483
Pluta AF, Mackay AM, Ainsztein AM, Goldberg IG, Earnshaw WC (1995) Centromere—hub of chromosomal activities. Science 270:1591–1594
Politz JCR, Zhang F, Pederson T (2006) MicroRNA-206 colocalizes with ribosome-rich regions in both the nucleolus and cytoplasm of rat myogenic cells. Proc Natl Acad Sci USA 103:18957–18962
Pontes O, Lawrence RJ, Neves N, Silva M, Lee JH, Chen ZJ, Viegas W, Pikaard CS (2003) Natural variation in nucleolar dominance reveals the relationship between nucleolus organizer chromatin topology and rRNA gene transcription in Arabidopsis. Proc Natl Acad Sci USA 100:11418–11423
Pontes O, Li CF, Nunes PC, Haag J, Ream T, Vitins A, Jacobsen SE, Pikaard CS (2006) The Arabidopsis chromatin-modifying nuclear siRNA pathway involves a nucleolar RNA processing center. Cell 126:79–92
Pontvianne F, Matia I, Douet J, Tourmente S, Medina FJ, Echeverria M, Saez-Vasquez J (2007) Characterization of AtNUC-L1 reveals a central role of nucleolin in nucleolus organization and silencing of AtNUC-L2 gene in Arabidopsis. Mol Biol Cell 18:369–379
Preuss S, Pikaard CS (2007) rRNA gene silencing and nucleolar dominance: insights into a chromosome-scale epigenetic on/off switch. Biochim Biophys Acta 1769:383–392
Prieto JL, McStay B (2007) Recruitment of factors linking transcription and processing of pre-rRNA to NOR chromatin is UBF-dependent and occurs independent of transcription in human cells. Genes Dev 21:2041–2054
Probst AV, Fagard M, Proux F, Mourrain P, Boutet S, Earley K, Lawrence RJ, Pikaard CS, Murfett J, Furner I, Vaucheret H, Sheid OM (2004) Arabidopsis histone deacetylase HDA6 is required for maintenance of transcriptional gene silencing and determines nuclear organization of rDNA repeats. Plant Cell 16:1021–1034
Rando O (2007) Global patterns of histone modifications. Curr Op Genet Devel 17:94–99
Raska I (2003) Oldies but goldies: searching for Christmas trees within the nucleolar architecture. Trends Cell Biol 13:517–525
Raska I, Shaw PJ, Cmarko D (2006) Structure and function of the nucleolus in the spotlight. Curr Op Cell Biol 18:325–334
Richards B, Flint SJ, Cole MD, LeRoy G (2007) Nucleolin is required for RNA polymerase I transcription in vivo. Mol Cell Biol 27:937–948
Riddle NC, Richards EJ (2002) The control of natural variation in cytosine methylation in Arabidopsis. Genetics 162:355–363
Riddle NC, Richards EJ (2005) Genetic variation in epigenetic inheritance of ribosomal RNA gene methylation in Arabidopsis. Plant J 41:524–532
Roussel P, Sirri V, Hernandez-Verdun D (1994) Quantification of Ag-NOR proteins using Ag-NOR staining on Western blots. J Histochem Cytochem 42:1513–1517
Roussel P, Andre C, Comai L, Hernandez-Verdun D (1996) The rDNA transcription machinery is assembled during mitosis in active NORs and absent in inactive NORs. J Cell Biol 133:235–346
Santoro R (2005) The silence of the ribosomal RNA genes. Cell Mol Life Sci 62:2067–2079
Santoro R, Grummt I (2001) Molecular mechanisms mediating methylation-dependent silencing of ribosomal gene transcription. Mol Cell 8:719–725
Santoro R, Grummt I (2005) Epigenetic mechanism of rRNA gene silencing: temporal order of NoRC-mediated histone modification, chromatin remodeling, and DNA methylation. Mol Cell Biol 25:2539–2546
Scherl A, Coute Y, Deon C, Calle A, Kindbeiter K, Sanchez JC, Greco A, Hochstrasser D, Diaz JJ (2002) Functional proteomic analysis of human nucleolus. Mol Biol Cell 13:4100–4109
Schwartz BE, Ahmad K (2005) Transcriptional activation triggers deposition and removal of the histone variant H3.3. Genes Dev 19:804–814
Sharma M (2004) NO29, a histone chaperone in the nucleolus. Prot Sci 13:115–116
Shaw PJ, Jordan EG (1995) The nucleolus. Ann Rev Cell Dev Biol 11:93–121
Shaw PJ, Highett MI, Beven AF, Jordan EG (1995) The nucleolar architecture of polymerase I transcription and processing. EMBO J 14:2896–2906
Shaw PJ, Beven AF, Leader DJ, Brown JWS (1998) Localization and processing from a polycistronic precursor of novel snoRNAs in maize. J Cell Science 111:2121–2128
Shogren-Knaak M, Ishii H, Sun J-M, Pazin MJ, Davie JR, Peterson CL (2006) Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311:844–847
Shou WY, Sakamoto KM, Keener J, Morimoto KW, Traverso EE, Azzam R, Hoppe GJ, Feldman RMR, DeModena J, Moazed D, Charbonneau H, Nomura M, Deshaies RJ (2001) Net1 stimulates RNA polymerase I transcription and regulates nucleolar structure independently of controlling mitotic exit. Mol Cell 8:45–55
Slusarczyk J, Wierzbicki A, Przewloka M, Tykarska T, Jerzmanowski A, Kuras M (2003) Influence of change in the proportion of H1 histone variants on microsporogenesis and development of male gametophyte in transgenic plants of tobacco (Nicotiana tabacum L.). Acta Soc Botan Pol 71:25–35
Sogo JM, Ness PJ, Widmer RM, Parish RW, Koller T (1984) Psoralen-crosslinking of DNA as a probe for the structure of active nucleolar chromatin. J Mol Biol 178:897–919
Song L, Han MH, Lesicka J, Federoff N (2007) Arabidopsis primary microRNA processing proteins HYL1 and DCL1 define a nuclear body distinct from the Cajal body. Proc Natl Acad Sci USA 104:5437–5442
Spector DL (2003) The dynamics of chromosome organization and gene regulation. Ann Rev Biochem 72:573–608
Stargell LA, Bowen J, Dadd CA, Dedon PC, Davis M, Cook RG, Allis CD, Gorovsky MA (1993) Temporal and spatial association of histone H2A variant hv1 with transcriptionally competent chromatin during nuclear development in Tetrahymena thermophila. Genes Dev 7:2641–2651
Stefanovsky V, Moss T (2006) Regulation of rRNA synthesis in human and mouse cells is not determined by changes in active gene count. Cell Cycle 5:735–739
Straight AF, Shou WY, Dowd GJ, Turck CW, Deshaies RJ, Johnson AD, Moazed D (1999) Net1, a Sir2-associated nucleolar protein required for rDNA silencing and nucleolar integrity. Cell 97:245–256
Taddei A, Hediger F, Neumann FR, Bauer C, Gasser SM (2004) Separation of silencing from perinuclear acnhoring functions in yeast Ku80, Sir4 and Esc1 proteins. EMBO J 23:1301–1312
Tanaka I, Akahori Y, Gomi K, Suzuki T, Ueda K (1999) A novel histone variant localized in nucleoli of higher plant cells. Chromosoma 108:190–199
Thompson WF, Flavell RB (1988) DNase I sensitivity of ribosomal RNA genes in chromatin and nucleolar dominance in wheat. J Mol Biol 204:535–548
Thorstensen T, Fischer A, Sandvik SV, Johnsen SS, Grini PE, Reuter G, Aalen RB (2006) The Arabidopsis SUVR4 protein is a nucleolar histone methyltransferase with preference for monomethylated H3K9. Nucl Acids Res 34:5461–5470
Tongaonkar P, French SL, Oakes ML, Vu L, Schneider DA, Beyer AL, Nomura M (2005) Histones are required for transcription of yeast rRNA genes by RNA polyrnerase I. Proc Natl Acad Sci USA 102:10129–10134
Tsang CK, Bertram PG, Ai WD, Drenan R, Zheng XFS (2003) Chromatin-mediated regulation of nucleolar structure and RNA Pol I localization by TOR. EMBO J 22:6045–6056
Ueno Y, Ishikawa T, Watanabe K, Terakura S, Iwakawa H, Okada K, Machida C, Machida Y (2007) Histone deacetylases and ASYMMETRIC LEAVES2 are involved in the establishment of polarity in leaves of Arabidopsis. Plant Cell 19:445–457
Vaziri H, Dessain SK, Eaton EN, Imai S-I, Frye RA, Pandita TK, Guarente L, Weinberg RA (2001) hSIR2(SIRT1) functions as an NAD-dependent p53 deacetylase. Cell 107:149–159
Wako T, Houben A, Furushima-Shimogawarana R, Belyaev ND, Fukui K (2003) Centromere-specific acetylation of histone H4 in barley detected through three-dimensional microscopy. Plant Mol Biol 51:533–541
Wierzbicki AT, Jerzmanowski A (2005) Suppression of histone H1 genes in Arabidopsis results in heritable developmental defects and stochastic changes in DNA methylation. Genetics 169:997–1008
Wu K, Tian L, Zhao C, Brown D, Miki B (2003) Repression of gene expression by Arabidopsis HD2 histone deacetylases. Plant J 34:241–247
Ye JQ, Eickbush TH (2006) Chromatin structure and transcription of the R1- and R2-inserted rRNA genes of Drosophila melanogaster. Mol Cell Biol 26:8781–8790
Young DW, Hassan MQ, Pratap J, Galindo M, Zaidi SK, Lee SH, Yang XQ, Xie R, Javed A, Underwood JM, Furcinitti P, Imbalzano AN, Penman S, Nickerson JA, Montecino MA, Lian JB, Stein JL, van Wijnen AJ, Stein GS (2007) Mitotic occupancy and lineage-specific transcriptional control of rRNA genes by Runx2. Nature 445:442–446
Yu Y, Maggi LB, Brady SN, Apicelli AJ, Dai MS, Lu H, Weber JD (2006) Nucleophosmin is essential for ribosomal protein L5 nuclear export. Mol Cell Biol 26:3798–3809
Zhou YG, Santoro R, Grummt I (2002) The chromatin remodeling complex NoRC targets HDAC1 to the ribosomal gene promoter and represses RNA polymerase I transcription. EMBO J 21:4632–4640
Zhou C, Labbe H, Sridha S, Wang L, Tian L, Latoszek-Green M, Yang Z, Brown D, Miki B, Wu K (2004) Repression and function of HD2-type histone deacetylases in Arabidopsis development. Plant J 38:715–724
Acknowledgement
This work was funded by the Biotechnology and Biological Sciences Research Council of the UK (BBSRC).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by E. Nigg
Rights and permissions
About this article
Cite this article
McKeown, P.C., Shaw, P.J. Chromatin: linking structure and function in the nucleolus. Chromosoma 118, 11–23 (2009). https://doi.org/10.1007/s00412-008-0184-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-008-0184-2