Abstract
Chloroplasts are generally known as eukaryotic organelles whose main function is photosynthesis. They perform other functions, however, such as synthesizing isoprenoids, fatty acids, heme, iron sulphur clusters and other essential compounds. In non-photosynthetic lineages that possess plastids, the chloroplast genomes have been reduced and most (or all) photosynthetic genes have been lost. Consequently, non-photosynthetic plastids have also been reduced structurally. Some of these non-photosynthetic or “cryptic” plastids were overlooked or unrecognized for decades. The number of complete plastid genome sequences and/or transcriptomes from non-photosynthetic taxa possessing plastids is rapidly increasing, thus allowing prediction of the functions of non-photosynthetic plastids in various eukaryotic lineages. In some non-photosynthetic eukaryotes with photosynthetic ancestors, no traces of plastid genomes or of plastids have been found, suggesting that they have lost the genomes or plastids completely. This review summarizes current knowledge of non-photosynthetic plastids, their genomes, structures and potential functions in free-living and parasitic plants, algae and protists. We introduce a model for the order of plastid gene losses which combines models proposed earlier for land plants with the patterns of gene retention and loss observed in protists. The rare cases of plastid genome loss and complete plastid loss are also discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the beginning of the twentieth century, Mereschkowsky formulated the hypothesis that plastids were once free-living cyanobacteria (Mereschkowsky 1905). A bacterial origin was later suggested for mitochondria (Wallin 1927). These ideas, however, were neglected until plastid DNA (Ris and Plaut 1962) and intramitochondrial fibers with DNA characteristics (Nass and Nass 1963) were discovered. Thereafter, a modern complex theory proposing that these eukaryotic organelles originated from prokaryotic organisms was postulated (Sagan 1967). The process of the conversion of an intracellular prokaryotic (i.e., bacterial) endosymbiont into an organelle is called primary endosymbiosis. The primary endosymbiotic origin of mitochondria and primary plastids (plastids bounded by two membranes) from α-proteobacteria and cyanobacteria, respectively, is currently the generally accepted paradigm.
Unlike facultative endosymbionts, endosymbiotic organelles are dependent on their host cells and they are inherited by progeny. The conversion of an endosymbiont into an organelle is, however, a complex process accompanied by the transfer of endosymbiont genes into the host nucleus, a process known as endosymbiotic gene transfer (EGT), and consequent evolution of a mechanism for the import of the protein products of these genes back into the endosymbiont/organelle (Cavalier-Smith and Lee 1985; Giovannoni et al. 1988; Gray et al. 1999; Yang et al. 1985; for reviews see Kleine et al. 2009; Ku et al. 2015; Zimorski et al. 2014). With EGT and the evolution of import machinery, the endosymbionts gradually lose their own essential genes and become organelles, retaining miniature versions of their genomes which indicate their origin. The genomes of primary plastids and mitochondria encode approximately 100–250 (Green 2011) and a few dozen genes (Gray et al. 1999; Zíková et al. 2016), respectively. Phylogenies based on bacterial, mitochondrial and plastid genes provide direct evidence that mitochondria and primary plastids are descended from α-proteobacteria and cyanobacteria, respectively (Giovannoni et al. 1988; Gray et al. 1999; Yang et al. 1985).
As a result of the primary endosymbiosis, eukaryotes belonging to the Archaeplastida supergroup (glaucophytes, red and green algae and land plants) possess plastids bounded by two membranes (Fig. 1) which most likely arose via a single engulfment of a cyanobacterium by a heterotrophic host (Archibald 2015; Ponce-Toledo et al. 2017; Rockwell et al. 2014). The nature of the ancestor of primary plastids of Archaeplastida is currently unclear. It was proposed that it could have been a relatively complex nitrogen-fixing (heterocyst-forming) cyanobacterium (Deschamps et al. 2008; Deusch et al. 2008). However, a recent phylogenetic analysis of 97 plastid-encoded proteins and their cyanobacterial homologs suggested that it was more likely a freshwater/terrestrial cyanobacterium related to Gleomargarita litophora (Ponce-Toledo et al. 2017). It was speculated that the successful cyanobacterial endosymbiosis might have been facilitated by a chlamydial parasite of the host (Becker et al. 2008; Facchinelli et al. 2013; Moustafa et al. 2008). On the other hand, more recent investigation of the origin of the key “chlamydial effector” enzymes of Archaeplastida casts doubt on this hypothesis (Domman et al. 2015).
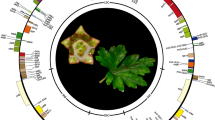
The distribution of non-photosynthetic plastids among eukaryotes. The unrooted eukaryotic tree includes five supergroups—SAR (Stramenopiles, Alveolata and Rhizaria) (violet), Amoebozoa (yellow), Opisthokonta (blue), Excavata (orange) and Archaeplastida (green). Haptophyta and Cryptophyta (both gray) are lineages of uncertain phylogenetic position. The unclear relationships between lineages are shown by dashed gray lines. The plastids gained by primary endosymbiosis/es are bounded by two membranes, while plastids gained by secondary or tertiary (or alternatively quaternary) endosymbioses are bounded either by three or by four membranes as schematically shown by the number of circular lines. Non-photosynthetic plastids are shown in white, and the number and color of circular lines corresponds to the photosynthetic plastids from which they were derived. The primary plastids of Archaeplastida including glaucophytes (bright blue), red algae (red), green algae and land plants (green) likely arose via a single primary endosymbiosis and they are bounded by two membranes (two circular lines). The presence of non-photosynthetic plastids among red and green algae and land plants is shown in white. Paulinella chromatophora (Rhizaria) represents an example of another independent primary endosymbiosis. Its plastids are also bounded by two membranes (dark blue, two circular lines). Complex plastids of red algal origin can be found in Stramenopiles (plastids bounded by four membranes), alveolates (plastids bounded by three or four membranes), and haptophytes and cryptophytes (both containing plastids bounded by four membranes). If plastids are present in apicomplexan and perkinsozoan species, they are exclusively non-photosynthetic, while both photosynthetic and non-photosynthetic complex plastids can be found among Stramenopiles, dinoflagellates, chrompodellids and cryptophytes. Chlorarachniophytes (Rhizaria) possessing plastids bounded by four membranes and euglenophytes (Excavata) possessing plastids bounded by three membranes arose via two independent secondary endosymbioses of green algae. Evidence for the loss of photosynthesis and retention of non-photosynthetic plastids exists among euglenophytes (white circle, three green circular lines). The residual nuclei of endosymbionts, nucleomorphs, are present between the second and the third plastid membranes of chlorarachniophytes (dark green dot) and cryptophytes (dark red dot), while they were lost in other complex plastids. Some apicomplexans and dinoflagellates have likely lost plastids completely (crossed circles), while genome-less non-photosynthetic plastids occur among perkinsozoans (asterisk) and likely also among chrompodellids and euglenophytes (asterisks with question marks). It is currently unknown if colpodellid plastids are bounded by four membranes like the plastids of their sister group, chromerids, or only by three membranes. Hence, the outermost fourth membrane of chrompodellids is indicated by a dashed circular line
The plastid genomes of Archaeplastida are typically up to 200 kb in size and they encode up to 250 genes (for a review see Archibald 2015; Rockwell et al. 2014). Considerable diversity of plastid genome sizes has been discovered in red algae with the largest observed in Corynoplastis japonica (1.13 Mbp) (Muñoz-Gómez et al. 2017). Many cyanobacterial genes were transferred to the host nucleus via EGT and a large number of them currently encode proteins that function in plastids, i.e., proteins involved in plastid gene expression, division, trafficking and metabolism (Kleine et al. 2009; Martin et al. 1998). These nucleus-encoded plastid proteins have to be targeted to plastids. The N-terminal targeting pre-sequence called the transit peptide serves in most cases as a signal for plastid import. The proteins are recognized and post-translationally transferred through the translocon of the outer chloroplast membrane (TOC) and the translocon of the inner chloroplast membrane (TIC). These TOC and TIC complexes consist of transit peptide receptors, protein conducting channels, regulatory elements and TOC–TIC translocon-connecting subunits (Gagat et al. 2013; Nakai 2015a; Schleiff and Becker 2011).
Another independent primary endosymbiosis of cyanobacteria occurred in a rhizarian amoeba Paulinella chromatophora (Marin et al. 2005; Nowack et al. 2008; Yoon et al. 2006). Paulinella chromatophora cells contain two kidney-shaped intracellular symbionts called chromatophores. The division of chromatophores is synchronized with the division of the P. chromatophora cell and phylogenetic studies confirmed that chromatophores are related to the Synechococcus/Prochlorococcus lineage of cyanobacteria (Marin et al. 2005; Nowack et al. 2008; Yoon et al. 2006). Although the chromatophore genome is reduced in comparison to its free-living relatives, its size is relatively large (1.02 Mb) and its coding capacity (867 protein-coding genes) is much larger than any other known plastid genome (Nowack et al. 2008). There is some evidence of a role for the Golgi apparatus in the transport of nucleus-encoded proteins through the chromatophore outer membrane (Nowack and Grossman 2012) and some homologs of cyanobacterial and plant TIC proteins are encoded by the chromatophore genome (Bodył et al. 2010; Mackiewicz et al. 2012), but the exact mechanism of protein import into chromatophores is unknown. The differences between the protein targeting machineries of the chromatophore and plastids of Archaeplastida reflect the fact that these organelles originated independently and that chromatophores arose relatively recently (~ 60–200 million years ago) (Marin et al. 2005; Nowack 2014; Nowack et al. 2008) compared to the primary plastids of Archaeplastida (~ 1–2 billion years ago) (Falkowski et al. 2004; Parfrey et al. 2011; Yoon et al. 2004).
After the primary endosymbiosis of a cyanobacterium in the ancestor of Archaeplastida, several “higher order” or “serial” endosymbiotic events (secondary, tertiary or even quarternary) occurred independently in various ancestrally heterotrophic lineages. During these endosymbioses, eukaryotic cells already possessing a plastid were engulfed by other eukaryotes. Such endosymbioses gave rise to organisms bearing complex plastids surrounded by three or four membranes. In the process of secondary endosymbiosis, various non-photosynthetic eukaryotic lineages independently engulfed red or green algae. While the endosymbiont plastids and their genomes have been retained, the nucleus of the endosymbiotic alga has been lost in most cases. Interesting exceptions include members of Cryptophyta and Chlorarachniophyta, which have retained a remnant of the endosymbiont nucleus in the form of a vestigial eukaryotic nucleus—the nucleomorph (Douglas et al. 1991; Gilson et al. 2006; for reviews see Archibald 2009; Keeling 2013; Maier et al. 2000; Moore and Archibald 2009; Vesteg et al. 2009). A single secondary endosymbiosis or more likely a series of secondary and tertiary (or even quarternary) endosymbioses of unknown number resulted in the emergence of diverse groups with complex plastids of red algal origin—stramenopiles, haptophytes, cryptophytes, apicomplexans, chromerids and dinoflagellates (Fig. 1) (Archibald 2009; Baurain et al. 2010; Cavalier-Smith 1999; Keeling 2013; Petersen et al. 2014; Zimorski et al. 2014). Two independent secondary endosymbioses with green algae led to the origin of the plastids of euglenids and chlorarachniophytes, while a plastid replacement by a secondary green plastid (“serial secondary endosymbiosis”) occurred in the dinoflagellate genus Lepidodinium (McFadden 2001; Stiller 2014; Watanabe et al. 1990). In addition, in some other dinoflagellate lineages “serial endosymbioses” occurred which led to the replacement of their ancestral peridinin-containing plastids by plastids of haptophyte, cryptophyte or diatom origin (Chesnick et al. 1997; Dodge 1969; Dorrell et al. 2017; Hackett et al. 2003; Schnepf and Elbrächter 1988; Takishita et al. 2002; Tengs et al. 2000; Yamada et al. 2017).
Since complex plastids possess typically three or four membranes, protein import into these plastids is more complex than protein import into primary plastids. Although the mechanisms of protein import into plastids differ among organisms possessing complex plastids, the first steps generally require the involvement of the endomembrane system (for reviews see Maier et al. 2015; Sheiner and Striepen 2013; Vesteg et al. 2009).
The typical functions of mitochondria and chloroplasts are respiration and photosynthesis, respectively. However, pathways providing other essential functions for the cell (e.g., synthesis of amino acids, isoprenoids, fatty acids, heme or iron sulphur clusters) are also localized in mitochondria and/or plastids (Kleffmann et al. 2004; Ralph et al. 2004; Sickmann et al. 2003; Terashima et al. 2011; Zíková et al. 2016). In some lineages, the respiratory or photosynthetic functions of these organelles have been reduced or completely lost and consequently, these organelles have also been structurally reduced. Some of these organelles were overlooked or unrecognized for decades. Since it was not possible to identify them easily and the evidence for some of them was initially based only on the presence of their nucleus-encoded hallmark proteins, they were called “cryptic organelles” (Archibald and Keeling 2002; Klinger et al. 2013; Williams and Keeling 2003). The reduced forms of mitochondria include hydrogenosomes (hydrogen-producing organelles), mitosomes and several intermediate forms, which usually do not contain the respiratory chain and the genome (Makiuchi and Nozaki 2014; Müller et al. 2012). Interestingly, a complete loss of the mitochondrion is also possible as exemplified by Monocercomonoides sp., an excavate possessing no identifiable traces of mitochondria (Karnkowska et al. 2016).
Non-photosynthetic plastids are relatively common, but in contrast to reduced mitochondria they have generally retained a reduced genome, although there are some exceptions (Molina et al. 2014; Smith and Asmail 2014; Smith and Lee 2014). Non-photosynthetic plastids have been found in many heterotrophic land plants (e.g., parasites, mycoheterotrophs) as well as in free-living (e.g., Euglena longa) and parasitic (e.g., Apicomplexa, Perkinsus sp., Helicosporidium sp.) protists (Figs. 1, 2). This review summarizes current knowledge about non-photosynthetic plastids, their genomes, structures and potential functions. We also discuss rare cases of complete plastid genome loss and plastid loss.
The distribution of non-photosynthetic plastids among green algae and land plants. The green circle represents a photosynthetic primary green plastid bounded by two membranes (two circular lines), which was present in the common ancestor of green algae and plants, and photosynthesis was retained in almost all green algal and plant lineages (not shown). Green algae and land plants are divided into two monophyletic lineages—(1) chlorophytes including prasinophyte green algae (basal yellow lineage) and chlorophyte green algae (yellow), and (2) streptophytes including streptophyte green algae (basal blue lineage) and land plants (blue). The light blue triangle indicates angiosperms including magnoliids, monocots and eudicots. The names of the lineages in which species with non-photosynthetic plastid(s) (white circles, two circular lines) have been described are underlined and the species or orders are noted. The asterisks (*) indicate that Polytomella sp. and Rafflesia lagascae (Malphigiales) have probably completely lost their plastid genomes
Non-photosynthetic plastids in parasitic red algae
Red algae as well as glaucophytes, green algae and land plants possess primary, two-membrane-bounded plastids of cyanobacterial origin and they all belong to the supergroup Archaeplastida (Fig. 1). The loss of photosynthesis among red algae is tightly connected with a parasitic lifestyle and it has occurred more than a hundred times in this group of organisms (Blouin and Lane 2012; Salomaki and Lane 2014). Parasitic red algae are usually assigned either as adelphoparasites (Greek “adelpho” = kin) that invade closely related species, or alloparasites (Greek “állos” = other) that can infect not only their sister taxa but also lineages that are more evolutionarily distant from them (Blouin and Lane 2012).
It has been shown that all adelphoparasites studied to date have lost their plastids and they incorporate dedifferentiated plastids from hosts into their spores (Goff and Coleman 1995; Salomaki et al. 2015). Before an adelphoparasite fuses with a host cell, it forms a conjunction with it. When cells fuse, the parasite delivers its nucleus and organelles into the host cell. In this heterokaryotic cell (containing both nuclei from the parasite and the host), the parasite nucleus and mitochondria divide alongside the division of the host nucleus, mitochondria and plastids. The host nucleus and mitochondria thereafter disappear and the host cell contains only the nucleus and mitochondria of the parasite and the plastids of the host (Goff and Coleman 1995). Most adelphoparasites are unable to reactivate photosynthesis in the stolen plastids. Nevertheless, in Janczewskia sp. or Plocamiocolax sp., mature reproductive spores are pigmented, although their plastids are colorless in vegetative tissues (Court 1980). It has been hypothesized that colorless genome-containing plastids in red algal parasites may perform pyrimidine, amino acid and fatty acid biosynthesis (Goff and Coleman 1995) or, in adelphoparasites, they might be present only as a consequence of the parasite–host cellular fusions without any other reason (Goff and Zuccarello 1994). Experimental evidence for any function of these plastids is lacking.
Photosynthetic activity has been observed in the alloparasite Choreocolax polysiphoniae, but there is a possibility that this photosynthetic tissue comes from the host (Callow et al. 1979; Goff and Zuccarello 1994; Kugrens and West 1973). Recently, the genome of a non-photosynthetic plastid in this alloparasite has been sequenced (Salomaki et al. 2015). Its 90 kb genome encodes 98 genes while having lost all genes for light harvesting and photosynthesis (Table 1; Online Resource 1) (Salomaki et al. 2015). Amino acid, fatty acid and isoprenoid biosynthesis likely occur in these plastids (Salomaki et al. 2015).
Non-photosynthetic plastids in green algae
Multiple independent losses of photosynthesis have occurred in Chloroplastida including in green algae and in land plants (Figs. 1, 2). In this chapter, we briefly describe non-photosynthetic plastids in parasitic and free-living colorless green algae and the following chapter will be devoted to non-photosynthetic plastids in land plants.
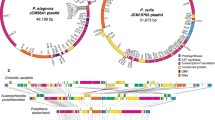
Cryptic plastids in parasitic green algae
The cryptic plastid of parasitic Helicosporidium sp. possesses a highly reduced plastid genome (~ 40 kb) which is devoid of photosynthesis-related genes (Table 1; Online Resource 1) (de Koning and Keeling 2006). Helicosporidium plastid metabolism includes tetrapyrrole (heme), isoprenoid and fatty acid synthesis, as well as several amino acid biosynthetic pathways (de Koning and Keeling 2004; Pombert et al. 2014). Although the majority of genes responsible for photosynthesis have been lost from the nuclear genome of Helicosporidium, some photosynthesis-related genes, such as those encoding the enzymes of the Calvin cycle, have been retained (Pombert et al. 2014). The predominantly free-living opportunistic pathogen Prototheca wickerhamii is closely related to mixotrophic green alga Auxenochlorella protothecoides (Yan et al. 2015). The Prototheca plastid genome encodes no genes for photosynthetic functions except for six ATP synthase subunits (Knauf and Hachtel 2002) (Table 1; Online Resource 1). The biochemical pathways localized in P. wickerhamii plastids include amino acid, carbohydrate, isoprenoid, lipid and tetrapyrrole biosynthesis as well as the thioredoxin reduction pathway (Borza et al. 2005).
Cryptic plastids in free-living green algae
Non-photosynthetic plastids with a genome and functional ribosomes have been retained in the colorless chlorophytes Polytoma obtusum and Polytoma uvella (Vernon et al. 2001). The P. uvella 230 kb plastid genome (GC content 23%) is the largest sequenced plastid genome among non-photosynthetic organisms (Figueroa-Martinez et al. 2017a, b) (Online Resource 1). It encodes 25 proteins (including clpP, ftsH and ycf1 genes), 2 plastid rRNAs and 27 tRNAs (Table 1). The function of the ycf1 gene product may be connected to TOC/TIC complexes in plants (Kikuchi et al. 2013) and thus it was suggested to be renamed tic214 (Nakai 2015a). However, there are ongoing debates as to whether YCF1 is really a key TIC component or whether it has another function (Bölter and Soll 2017; de Vries et al. 2015, 2017; Nakai 2015a, b). The ycf1 gene is present only in some plastid genomes of green algae and land plants (Table 1), suggesting that it had been acquired by the common ancestor of Chloroplastida and subsequently lost in some of their lineages (e.g., Poacae) (de Vries et al. 2017).
The large size of the P. uvella plastid genome is a consequence of the expansion of intergenic DNA repeats and the accumulation of introns (Figueroa-Martinez et al. 2017a). Many photosynthetic relatives of P. uvella also contain large plastid genomes with repeat-rich intergenic regions, which are also common in plastid genomes from other, unrelated organisms (George et al. 2015; Gogniashvili et al. 2016; Kong and Yang 2016; do Vieira et al. 2016). Such repeat regions might have mediated gene losses leading to the reduced gene content in the P. uvella plastid genome (Figueroa-Martinez et al. 2017a). On the other hand, a free-living lifestyle might have facilitated the retention of intergenic repeats and introns in the P. uvella plastid genome, in contrast to parasites possessing non-photosynthetic plastids whose genomes are generally highly compact (Figueroa-Martinez et al. (2017b).
The complete loss of plastid genomes was for a long time regarded as impossible because there were no known cases of plastids without genomes. Recently, such plastids were reported in four Polytomella species (Smith and Lee 2014) (Online Resource 1). No genes required for plastid division or for the replication and repair of plastid DNA were found in their nuclear genomes, but genes encoding plastid-targeted proteins were present and expressed (Smith and Lee 2014). Polytomella sp. and P. uvella plastids represent outstanding examples of diametrically different outcomes of the evolution of non-photosynthetic plastids in chlamydomonadalean green algae (Figueroa-Martinez et al. 2017a, b).
The loss of a plastid genome might not be a functionally simple event even in secondary heterotrophs, because some plastid-encoded genes functionally unrelated to photosynthesis are essential in many green algae. For example, the plastid trnE gene encoding plastid Glu-tRNA is highly conserved in chlamydomonadalean algae and is required for the synthesis of heme (Atteia et al. 2005; Barbrook et al. 2006; Beale 1999; Howe and Smith 1991; Smith and Asmail 2014; Smith and Lee 2014). Glu-tRNA import from mitochondria into plastids has been well described in the parasitic plant Epifagus virginiana (Wolfe et al. 1992) and in the subterrestrial orchid Rhizanthella gardneri (Delannoy et al. 2011), but Glu-tRNA is not encoded by the Polytomella mitochondrial genome (Smith et al. 2010). Hypothetically, nucleus-encoded Glu-tRNA might be imported into the plastid from the cytosol, although there is currently no evidence for this in Polytomella (Smith and Lee 2014). In some other plastid-bearing lineages, such as in apicomplexan parasites, the initial step of heme synthesis occurs in mitochondria through the Shemin pathway without any need for plastid Glu-tRNA (Oborník and Green 2005). However, the Shemin pathway seems to be absent in Polytomella. Hence, the details of heme synthesis in these algae are currently unknown (Smith and Lee 2014).
Non-photosynthetic plastids in land plants
The loss of photosynthesis occurred independently multiple times among various lineages of angiosperms and in the bryophyte Aneura mirabilis (Wickett et al. 2008) (Fig. 2). Land plants with reduced plastid genomes include parasites of plants and mycoheterotrophs which are both often non-photosynthetic. The degree of plastid genome reduction varies among species (Table 1; Online Resource 1).
Aneura mirabilis
The epiparasitic liverwort A. mirabilis is able to grow buried underground without sunlight and can obtain carbon dioxide from its association with mycorrhiza-forming fungi (Bidartondo et al. 2003). It is the only known non-photosynthetic bryophyte to date. It possesses a 108 kb circular plastid genome containing large and small single copy (LSC and SSC) regions and two IRs (inverted repeats) between them. Due to the retention of GC rich IRs, the GC content of its plastid (40.6%) is higher than in other heterotrophic plants and algae (Online Resource 1). The plastid genome encodes 116 unique genes. Seventy-seven of these are genes of unknown function. Twenty-five genes have been partially or completely lost from the A. mirabilis plastid genome, while photosynthetic psa and psb genes together with a portion of pet genes, ndh genes and the ccsA gene have become pseudogenes (Table 1) (Wickett et al. 2008). Wickett et al. (2008) infer that the A. mirabilis plastid is essential for fatty acids biosynthesis, because it contains an apparently functional accD gene encoding a subunit of acetyl-CoA carboxylase (ACC). ACC produces malonyl-CoA used for the de novo synthesis of fatty acids which occurs in plastids in plants (Ohlrogge and Browse 1995).
A non-photosynthetic magnoliid
A bizarre non-photosynthetic plant, Hydnora visseri, belongs to one of the oldest parasitic plant lineages—Hydnoraceae (Naumann et al. 2013). It contains a highly reduced ~ 27 kb plastid genome encoding only 24 genes including 4 rrn genes, 3 trn genes (trnE, trnfM, trnI-CAU), several genes for ribosomal proteins, and accD, ycf1 and ycf2 genes (Naumann et al. 2016) (Table 1). The ycf2 gene could be involved in water-usage efficiency (Ruiz-Nieto et al. 2015). Despite its extreme reduction, the H. visseri plastid genome has retained the typical plastid genome structure with IRs. However, the IRs are shortened and the plastid genome has a very low GC content (~ 24%) (Naumann et al. 2016) (Online Resource 1).
Non-photosynthetic plastids of angiosperm monocots
The mycoheterotrophic lifestyle and the loss of photosynthesis are quite common in several monocot orders—Petrosaviales, Liliales, Asparagales, Pandanales and Dioscoreales (Merckx and Freudenstein 2010) (Fig. 2). The complete plastid genome sequences are currently available for a single representative of Petrosaviales, Petrosavia stellaris (Logacheva et al. 2014), for various species of orchids belonging to the Asparagales order (Barrett and Davis 2012; Barrett et al. 2014; Delannoy et al. 2011; Funk et al. 2007; Logacheva et al. 2011; McNeal et al. 2007; Schelkunov et al. 2015), for Sciaphila densiflora (Pandanales) (Lam et al. 2015) and Thismia tentaculata (Dioscoreales) (Lim et al. 2016) (Table 1; Online Resource 1).
The P. stellaris plastid genome is ~ 104 kb in size and contains two IRs which split the genome into LSC and SSC regions (Logacheva et al. 2014); the GC content is 37.5% (Online Resource 1). It shows a wide variety of rearrangements in comparison to plastid genomes in other monocots. It has retained only a minority of photosynthetic genes (rbcL, psbZ, petG and atp) and some genes related to other plastid metabolic functions—accD and clpP encoding ATP-dependent Clp protease proteolytic subunits (Table 1). This plastid genome appears to be in an early stage of reduction and rbcL probably performs a non-photosynthetic function (Logacheva et al. 2014).
The GC content of the 92 kb plastid genome of the orchid Neottia nidus-avis is 34.4% and it appears to have typical plastid genome architecture (Logacheva et al. 2011) (Online Resource 1). It has lost all plastid genes encoding proteins involved in photosynthesis and transcription, while it has retained most genes whose products are involved in translation. The accD and clpP genes have been retained as well as two genes encoding proteins of unknown function (Logacheva et al. 2011) (Table 1).
Sequence analyses of plastid genomes from other mycoheterotrophic species of orchids, Epipogium aphyllum and Epipogium roseum, have revealed drastic reduction (31 kb encoding 27 genes and 19 kb encoding 29 genes, respectively) (Table 1; Online Resource 1) and indicated that these genomes have undergone multiple rearrangements (Schelkunov et al. 2015). The plastid genome of E. aphyllum has a tripartite structure. It contains two IRs located next to each other and only one single copy region. In contrast, the plastid genome of E. roseum has extremely reduced IRs and it contains two single copy regions. The GC contents of both genomes are very low (30 and 33%, respectively). Schelkunov et al. (2015) hypothesize that the retention of the plastid genome in Epipogium sp. is necessary for translation of accD and/or clpP proteins.
Another orchid species, Rhizanthella gardneri, possesses a 59 kb plastid genome encoding only 37 unique genes, none of which is involved in photosynthesis (Delannoy et al. 2011) (Table 1; Online Resource 1). accD and clpP are among the genes that have been retained in the R. gardneri plastid genome (Delannoy et al. 2011).
In contrast to R. gardneri, the plastid genome of the orchid Corallorhiza striata var. vreelandii with 137.5 kb in size is in a relatively early stage of reduction (Barrett and Davis 2012) (Table 1; Online Resource 1). The plastid-encoded photosynthesis-related genes (including rbcL) have been either lost or pseudogenized. The rbcL gene has been pseudogenized several times independently within this genus (Barrett and Davis 2012; Barrett and Freudenstein 2008). All plastid-encoded ATP synthase subunit genes are putatively functional in C. striata (Barrett and Davis 2012), A. mirabilis (Wickett et al. 2008) and Cuscuta sp. (Funk et al. 2007; McNeal et al. 2007), while they have been lost in R. gardneri and N. nidus-avis (Delannoy et al. 2011; Logacheva et al. 2011). C. striata has retained many housekeeping genes along with accD, clpP, matK, ycf1 and ycf2 genes (Table 1). Besides the plastid genome of C. striata, another three non-photosynthetic plastid genome sequences from the Corallorhiza genus recently became available (Barrett et al. 2014).
The mycoheterotroph Sciaphila densiflora (Pandanales) has a very small plastid genome (21.5 kb) with a high GC content which encodes 28 non-photosynthetic genes (Lam et al. 2015) including accD, clpP and matK genes (Table 1; Online Resource 1). Wicke et al. (2013) propose that it is on the way to complete plastid genome loss.
Thismia tentaculata, a mycoheterotrophic representative of Dioscoreales, possesses a non-photosynthetic plastid with a genome which is only 16 kb in size and thus the smallest known plastid genome among angiosperm monocots. It includes only 5 rRNA genes and 7 protein encoding genes including accD (Table 1). Interestingly, its IRs probably arose secondarily, while the original ones were lost (Lim et al. 2016) (Online Resource 1).
Non-photosynthetic plastids of angiosperm eudicots
Non-photosynthetic plants are also quite common within the eudicot lineage of angiosperms (Fig. 2). The complete sequences of plastid genomes are currently available for two Pilostyles species belonging to the Apodanthaceae family (Cucurbitales), two species from Ericales, the Cuscuta genus from the Convolvulaceae family (Solanales) and many species from the Orobanchaceae family (Lamiales) (Table 1; Online Resource 1).
Although plastid-like structures were observed in Rafflesia lagascae (Rafflesiaceae, Malphigiales), the endoparasite of Tetrastigma sp., it was impossible to detect a plastid genome using multiple approaches (Molina et al. 2014). Hence, R. lagascae is the only known plant species putatively without a plastid genome (Molina et al. 2014) (Online Resource 1).
Pilostyles aethiopica and Pilostyles hamiltonii, parasites of Fabaceae, contain 11 and 15 kb plastid genomes retaining 5 and 6 genes, respectively (Table 1; Online Resource 1) (Bellot and Renner 2016). The fatty acid biosynthesis pathway is likely the essential metabolic pathway occurring in Pilostyles plastids, since the accD gene has been retained in both plastid genomes (Bellot and Renner 2016) (Table 1).
Two non-photosynthetic plastid genomes from plants of the Ericaceae family have been sequenced only recently (Gruzdev et al. 2016; Logacheva et al. 2016). Monotropa uniflora and Hypopitys monotropa possess 46 kb (Logacheva et al. 2016) and 35 kb (Gruzdev et al. 2016; Logacheva et al. 2016) plastid genomes, respectively. Unlike in M. uniflora, the plastid genome of H. monotropa lacks IR regions and has a slightly higher GC content (Online Resource 1); it also encodes four additional tRNA genes. clpP and accD genes are encoded by the H. monotropa plastid genome (Table 1), but their sequences are highly divergent. clpP is absent from the M. uniflora plastid genome and accD is present in the form of a pseudogene (Logacheva et al. 2016).
Photosynthetic ability is variable in species of the parasitic Cuscuta genus (Convolvulaceae) (Braukmann et al. 2013). Although all Cuscuta spp. plastid genomes lack ndh, rpl32 and rpl16, they have retained many plastid genes that are generally required for photosynthesis, such as rbcL, psa, psb, pet and atp (Table 1), indicating that some species might be cryptically photosynthetic (Braukmann et al. 2013).
Lathraea squamaria (Orobanchaceae) is a holoparasite that likely exhibits the lowest degree of plastid genome reduction among non-photosynthetic land plants (Samigullin et al. 2016) (Table 1; Online Resource 1). The 150.5 kb plastid genome of L. squamaria has retained all photosynthetic genes typically present in plant plastid genomes, but some of them are pseudogenized (Table 1; Online Resource 1). Epifagus virginiana (Orobanchaceae), a parasite of American beech roots, possesses a highly reduced 70 kb plastid genome with the common quadripartite structure (Wolfe et al. 1992) (Online Resource 1). In contrast to L. squamaria, the E. virginiana plastid genome has lost all photosynthesis-related genes (Wolfe et al. 1992) (Table 1; Online Resource 1). Another orobanchid, Cistanche deserticola, has retained only a single photosynthesis-related gene, psbM, and almost a whole set of tRNA genes is encoded by its 105 kb plastid genome (Table 1; Online Resource 1). The retention of these genes might be a consequence of either a reduced level of parasitism or a recent transition to parasitism (Li et al. 2013). All three plastid genomes encode the clpP gene, while the accD gene has become a pseudogene in the C. deserticola plastid genome (Online Resource 1).
Non-photosynthetic plastids in alveolates
Alveolates include ciliates, dinoflagellates, perkinsozoans, apicomplexans, colpodellids, chromerids and other related taxa (Adl et al. 2012; Burki 2014). While ciliates are non-photosynthetic, species with photosynthetic and/or non-photosynthetic complex plastids can be found among the other lineages (Fig. 1), collectively classified as Myzozoa. Either the common myzozoan ancestor possessed photosynthetic plastids of a red algal origin (Cavalier-Smith and Chao 2004) which were replaced via secondary and tertiary (or alternatively quaternary) endosymbiosis multiple times (at least in dinoflagellates), or there were independent higher order endosymbioses in the ancestors of individual myzozoan lineages. In either case, photosynthesis was lost many times among Myzozoa (Janouškovec et al. 2015).
Apicoplasts: rudimentary plastids in apicomplexan parasites
Apicoplasts are highly reduced four-membrane-bounded non-photosynthetic plastids of apicomplexan parasites such as Toxoplasma gondii (which causes toxoplasmosis) and Plasmodium falciparum (which causes malaria) (McFadden and Yeh 2017; Waller et al. 1998; Wilson 2004). The complete apicoplast genome sequences are available for many apicomplexans parasites (Brayton et al. 2007; Cai et al. 2003; Gardner et al. 2005; Imura et al. 2014). The size of apicoplast genomes ranges from 28 to 48 kb. The first sequenced apicoplast genome was that of Plasmodium falciparum (Wilson et al. 1996). It is highly reduced (35 kb), it has a low GC content (14%) and it contains approximately 30 protein-coding genes including orfs; rRNA genes and 25 tRNA genes (Arisue et al. 2012; Wilson et al. 1996; Wilson and Williamson 1997) (Table 1; Online Resource 1). Apicoplast genome sequences are generally very similar to each other (Cinar et al. 2016), but they differ in their arrangement of the genes into clusters (Huang et al. 2015), in GC content (Cinar et al. 2016) (Online Resource 1) and in the presence of several ribosomal protein genes and genes of unknown function (Cinar et al. 2016; Huang et al. 2015) (Table 1).
Inhibition studies in P. falciparum and T. gondii have revealed that the apicoplast and its genome are necessary for the survival of apicomplexan parasites (Wilson 2004). Several biosynthetic pathways occur in the apicoplast, i.e., the non-mevalonate DOXP isoprenoid biosynthesis pathway, heme biosynthesis and the type II fatty acid biosynthetic pathway (Ralph et al. 2004; Wilson 2004, 2005; Yeh and DeRisi 2011). The treatment of P. falciparum by inhibitors of apicoplast metabolism, gene expression and replication should have only a minimal harmful effect on humans. Besides the inhibitors of apicoplast metabolic pathways, several different antibiotics have been used for malaria treatment—the inhibitors of apicoplast DNA replication (e.g., ciprofloxacin), transcription (e.g., rifampin) and translation (e.g., macrolids) (for a review see Gaillard et al. 2016; Mukherjee and Sadhukhan 2016).
Antibiotic treatments leading to the loss of apicoplast and cell death in P. falciparum can be reversed in the asexual blood stage by the addition of isopentenyl pyrophosphate (IPP), the isoprenoid precursor, suggesting that IPP is the only essential compound synthesized in apicoplasts during this life cycle stage (Yeh and DeRisi 2011). These experiments (Yeh and DeRisi 2011) have demonstrated that the loss of the apicoplast in P. falciparum is principally possible, but only under very specific conditions. Importantly, there is no evidence for the presence of apicoplasts in the apicomplexans Cryptosporidium parvum (Abrahamsen et al. 2004; Zhu et al. 2000), Gregarina niphandrodes (Toso and Omoto 2007) and Ascogregarina taiwanensis (Templeton et al. 2010). These species have likely lost the apicoplasts secondarily. The ability to synthesize fatty acids via a type I cytosolic pathway instead of the type II plastid-dependent pathway might have been a prerequisite for complete plastid loss in these species (Abrahamsen et al. 2004; Templeton et al. 2010; Toso and Omoto 2007; Zhu et al. 2000).
Cryptic plastids in Colpodellida
Colpodellids are closely related to apicomplexan parasites. Interestingly, transcripts with plastid-targeting presequences were found in the transcriptomic data of three predatory colpodellids—Voromas pontica, Alphamonas edax and Colpodella angusta—suggesting the presence of non-photosynthetic plastids (Gile and Slamovits 2014; Janouškovec et al. 2015) (Fig. 1). The loss of photosynthesis likely occurred independently several times in various colpodellids and apicomplexans (Gile and Slamovits 2014; Janouškovec et al. 2015). The biosynthetic pathways present in the plastids of different colpodellid species probably differ, but isoprenoid synthesis seems to be preserved in the plastids of all studied species (Janouškovec et al. 2015). There is no evidence for the presence of genomes in colpodellid plastids.
Cryptic plastids in Dinoflagellata
Despite the fact that about 50% of dinoflagellates are non-photosynthetic, dinoflagellates are generally thought to have had a photosynthetic ancestor which possessed complex, peridinin-containing, three-membrane-bounded plastids of red algal origin (Cavalier-Smith 2002; Janouškovec et al. 2017) (Fig. 1). This implies multiple losses of photosynthetic ability and plastids in the dinoflagellate lineage. One of the free-living non-photosynthetic dinoflagellates is an osmotrophic marine species Crypthecodinium cohnii. Analysis of C. cohnii ESTs revealed the presence and expression of numerous genes whose products possess N-terminal plastid-targeting presequences, indicating the presence of a cryptic plastid (Sánchez-Puerta et al. 2007). One of these genes is RbcL which is often encoded by plastid genomes in other non-photosynthetic lineages (Wolfe and de Pamphilis 1998). The presence of a nuclear RbcL transcript in C. cohnii ESTs is, however, not surprising, because dinoflagellates are known to use a prokaryotic type II form of RuBisCO, which was apparently acquired by horizontal transfer from α-proteobacteria to their nucleus (Delwiche and Palmer 1996; Morse et al. 1995; Rowan et al. 1996; Whitney et al. 1995). In addition, several metabolic pathways occur in C. cohnii’s putative non-photosynthetic plastid—the non-mevalonate (DOXP) pathway of isoprenoid biosynthesis, iron–sulfur cluster assembly and probably the heme biosynthesis pathway (Sánchez-Puerta et al. 2007). Analysis of EST data from another free-living dinoflagellate, Oxyrrhis marina, revealed the presence of transcripts encoding enzymes involved in similar metabolic pathways (Slamovits and Keeling 2008). Since these enzymes possess N-terminal plastid-targeting presequences, O. marina also likely contains a cryptic plastid-derived organelle (Slamovits and Keeling 2008). The transcriptomes of Noctiluca scintillans and O. marina have revealed the presence of plastid pathways for the synthesis of isoprenoids and tetrapyrroles, an additional ferredoxin redox system and the iron–sulfur cluster assembly pathway, while the pathway for fatty acid biosynthesis is likely absent (Janouškovec et al. 2017). There is currently no evidence for the presence of plastid genomes in these non-photosynthetic dinoflagellates. It has been suggested that it might be difficult to achieve a complete loss of a plastid in free-living non-photosynthetic dinoflagellates ancestrally possessing peridinin-containing plastids due to the dependency on plastid metabolic pathways (Janouškovec et al. 2017). Indeed, the only dinoflagellate in which the complete plastid loss is well-corroborated to date is the parasite Hematodinium sp. (Gornik et al. 2015). Nevertheless, various dinoflagellate lineages have replaced their peridinin-containing plastids by other secondary or tertiary (or even quaternary) plastids and this plastid replacement was accompanied by the reduction or complete loss of the ancestral peridinin-containing plastid (Hehenberger et al. 2014).
Cryptic plastid in Perkinsus sp.
The loss of photosynthesis also occurred in the oyster parasite Perkinsus sp. (Matsuzaki et al. 2008) (Fig. 1). This alveolate genus is closely related to dinoflagellates and it possesses a small, vestigial, four-membrane-bounded plastid compartment (Robledo et al. 2011; Teles-Grilo et al. 2007) which is the location of isoprenoid synthesis (Grauvogel et al. 2007; Matsuzaki et al. 2008; Robledo et al. 2011), fatty acid synthesis and iron–sulfur cluster assembly (Stelter et al. 2007). It has probably lost the plastid genome as well as the heme biosynthesis pathway (Robledo et al. 2011).
Cryptic plastids in Stramenopiles
Pteridomonas danica and Ciliophrys infusionum, both belonging to Dictyochophyceae, have independently lost photosynthesis (Fig. 1) while retaining plastid remnants. Although their plastid genomes have not been sequenced to date, there is evidence that their non-photosynthetic plastids contain at least rbcL and rbcS genes (Sekiguchi et al. 2002) (Online Resource 1).
The loss of photosynthesis has also occurred independently several times in the diatom Nitzschia spp. (Kamikawa et al. 2015b). Plastid-like structures have been observed in several isolates of the genus Nitzschia, and for many of them, plastid 16S rRNA sequences have been obtained suggesting that their plastids contain genomes (Kamikawa et al. 2015a). The plastid genome of Nitzschia sp. NIES-3851 is almost 68 kb in size with a GC content of only 23% (Online Resource 1). It lacks all photosynthesis-related genes, except for atp genes, and it encodes house-keeping genes (rrn, rps, rpl, trn); secA, secY and tatC genes (encoding thylakoid membrane translocators) and suf genes (encoding proteins involved in iron–sulfur cluster assembly) (Kamikawa et al. 2015a) (Table 1). Kamikawa et al. (2015b) propose that the ATP synthase complex located in the thylakoid membrane may allow for ATP hydrolysis to maintain a proton gradient sufficient for the translocation of proteins into thylakoids. Thus, the retention of atp genes might be important in the early stages of plastid genome reduction in other parasitic and free-living lineages (Kamikawa et al. 2015b).
The predictions of Kamikawa et al. (2017) have recently revealed that plastids of Nitzschia sp. NIES-3581 have lost isopentenyl pyrophosphate (IPP) synthesis and Rubisco-dependent carbon fixation, while they have retained other metabolic pathways such as amino acid biosynthesis, glycolysis/gluconeogenesis and the reductive pentose phosphate pathway—the latter two being parts of the Calvin–Benson cycle. The retention of proteins for both of these metabolic pathways may be linked to plastid amino acid biosynthesis (Kamikawa et al. 2017).
Non-photosynthetic plastids bounded by four membranes are also present in the chrysophyte Spumella sp., although it is unknown if they possess a plastid genome or if they have any essential metabolic function (Yubuki et al. 2008). In addition, the heterotrophic ameba Leukarachnion sp. has been placed among photosynthetic Stramenopiles in phylogenies, suggesting the loss of photosynthesis in this genus (Ben Ali et al. 2002; Grant et al. 2009). However, PCR amplification of the rbcL gene was unsuccessful and plastid-like structures have not been observed (Grant et al. 2009). All these findings indicate that the loss of photosynthesis is common among Stramenopiles (Grant et al. 2009) and the presence of reduced plastids have been documented at least in some of them (Belcher and Swale 1976; Kamikawa et al. 2015b; Sekiguchi et al. 2002).
Cryptic plastids in Cryptophyta
In the freshwater genus Cryptomonas, at least three independent losses of photosynthesis have occurred (Hoef-Emden 2005) (Fig. 1). Interestingly, the nucleomorph, the vestigial nucleus of the secondary endosymbiont, has been retained in all three non-photosynthetic Cryptomonas species (Hoef-Emden 2005). The Cryptomonas paramecium nucleomorph genome is ~ 500 kb in size and it encodes 466 proteins (Tanifuji et al. 2011).
The C. paramecium plastid is likely essential for fatty acid biosynthesis, since its genome encodes acyl carrier protein (acpP gene) (Donaher et al. 2009). The 77.7 kb plastid genome of C. paramecium with 38% GC content is 1.5-fold smaller than the plastid genomes of its photosynthetic relatives, Guillardia theta and Rhodomonas salina (Donaher et al. 2009; Hoef-Emden 2005), and it encodes 82 protein-coding genes, 3 rRNAs and 27 tRNAs (Donaher et al. 2009) (Table 1; Online Resource 1). It has lost IRs and most photosynthesis-related genes except for rbcL, rbcS, cbbX (Rubisco activase) and atp genes. Interestingly, secA, secY and tatC genes have also been retained in the C. paramecium plastid genome, similar to Nitzschia sp. NIES-3581 (Table 1). The co-retention of genes encoding subunits of ATP-synthase and thylakoid membrane translocons in both Nitzschia sp. NIES-3581 and C. paramecium (Table 1) might indicate that thylakoids are developed, and that thylakoid membrane translocation dependent upon ATP hydrolysis is functional in both of these organisms. ATP hydrolysis seems to be crucial for the maintenance of proton motive force and Tat- and Sec-mediated import of proteins into thylakoids even in Arabidopsis thaliana (Kohzuma et al. 2017). However, the function of thylakoids in C. paramecium and Nitzschia sp. NIES-3581 remains unknown. In addition to stromal rbcS and rbcL, the chll gene involved in tetrapyrrole (heme and chlorophyll) synthesis and the petF gene encoding ferredoxin (which might be associated with thylakoids) have been also retained in the plastid genome of C. paramecium, but not in Nitzschia sp. NIES-3581 (Table 1), indicating that the function of their thylakoids might be different.
Cryptic plastids in Euglenophyta
At least five branches within the Euglenophyceae have secondarily lost photosynthesis (Marin 2004; Marin et al. 2003) (Fig. 1). Whether Euglena quartana (previously Khawkinea quartana), Phacus ocellatus (previously Hyalophacus ocellatus), Lepocinclis acus var. hyalina or Trachelomonas reticulata have lost their plastid compartments completely, or they still possess residual plastids and genomes, is currently unknown (Marin 2004). Nevertheless, the free-living E. longa, a colorless flagellate closely related to the photosynthetic Euglena gracilis (Marin 2004; Müllner et al. 2001; Triemer et al. 2006), possesses a non-photosynthetic plastid bounded by three membranes and containing a genome.
Euglena longa circular 73-kb plastid DNA is about half the size of E. gracilis chloroplast DNA (143 kb) (Hallick et al. 1993) (Online Resource 1). The genes for photosynthesis-related proteins are absent from E. longa plastid DNA, except for rbcL (Gockel and Hachtel 2000) (Table 1; Online Resource 1). There are also significant differences in gene order between E. gracilis and E. longa chloroplast genomes. The transcripts of plastid-encoded tufA, rbcL, ribosomal protein genes, 23S and 16S rRNA genes, and three unassigned ORFs as well as a 53-kDa rbcL gene product were detected in E. longa (Gockel et al. 1994; Gockel and Hachtel 2000). This indicates that transcriptional and translational machinery in E. longa plastids is functional. The large subunit of Rubisco is the only plastid-encoded protein with a function other than genome maintenance and expression; so it seems that the whole genome is retained only to express this gene. Záhonová et al. (2016) have recently confirmed that both rbcL and nucleus-encoded RbcS are translated in E. longa, but their abundance is very low in comparison with E. gracilis. Moreover, the rbcL subunit is highly divergent and the function of Rubisco in E. longa is unclear (Záhonová et al. 2016).
Interestingly, the loss of photosynthetic ability in E. gracilis can be experimentally induced using various physical (e.g., UV) and chemical (e.g., antibiotics) agents which affect bacterial as well as chloroplast DNA replication or gene expression—a process termed “bleaching” (for a review see Krajčovič et al. 2002). E. gracilis bleached mutants lack most parts (some of them probably all) of the chloroplast genome. The plastid remnants have minimal (if any) metabolic activity, making them ideal storage compartments for proteins and value-added metabolites of biotechnological interest produced by nuclear transformation of Euglena (for a review see Krajčovič et al. 2015). If there is sufficient carbon source in the medium, the viability of E. gracilis bleached mutant cells is unaffected in comparison to photosynthetically active wild type cells (Hadariová et al. 2017).
Euglena longa was once viewed as a naturally bleached form of E. gracilis (for a review see Bodył 1996). The treatment of E. longa with antibiotics also results in the loss of plastid genes (Hadariová et al. 2017). However, in contrast to E. gracilis, this is fatal for E. longa (Hadariová et al. 2017). This suggests that an intact plastid genome is obligatory for E. longa survival, although it still remains unknown which plastid-encoded gene(s) and plastid-localized metabolic pathway(s) play an essential role in E. longa (Hadariová et al. 2017).
Plastid genome reduction, functions of non-photosynthetic plastids and the rarity of complete plastid loss
Reduced non-photosynthetic plastids are present in both parasitic and free-living as well as unicelluar and multicellular eukaryotes. Parasitic organisms are often reduced, possibly as a consequence of their adaptations to a parasitic lifestyle. Since they can metabolically profit from their hosts, many of their metabolic pathways could be on their way to complete loss. The reductive evolution of parasitic eukaryotes is not restricted to plastids and their genomes, and nuclear genomes, other organelles, like mitochondria, peroxisomes, endoplasmatic reticulum, Golgi complex, and the metabolism and structure of the entire cell, are often reduced as well. On the other hand, the presence of reduced plastids or mitochondria in the free-living eukaryotic lineages suggests that organellar reductive evolution is not restricted to parasites. In some lineages, e.g., metamonads, the reduction of mitochondrial genomes and metabolism has apparently preceded the evolution of parasitism (Klinger et al. 2016; Leger et al. 2017). The facultatively parasitic P. wickerhamii is an example of an alga in which the loss of photosynthesis likely served as a pre-adaptation to the evolution of occasional parasitism. Thus, in some parasites, the reductive evolution of plastids might have been initially a consequence of adaptation to free-living heterotrophy, a likely pre-condition for an evolutionary transition to a parasitic lifestyle. Evidence for massive reduction or even complete loss of endosymbiotic organelles exists in free-living as well as in parasitic eukaryotes and the mechanisms and selective pressures for reduction might differ between different lineages.
A model for the order of plastid gene losses
The loss of autotrophy is generally connected with elevated non-synonymous changes in photosynthesis-related genes, their pseudogenization and subsequent loss. Concomitantly, essential plastid genes are transferred into the nucleus (Wolfe et al. 1992; for reviews see Barrett and Davis 2012; Krause 2008). A model for plastid genome evolution in non-photosynthetic plants has been suggested by various authors (Barrett and Davis 2012; Barrett et al. 2014; Wicke et al. 2016). According to this model, ndh genes encoding subunits of the NAD(P)H dehydrogenase complex involved in photooxidative stress response (Martin and Sabater 2010) are the first genes lost during the transition from an autotrophic to a facultatively heterotrophic and/or parasitic lifestyle. The ndh genes are, however, often absent not only from plastid genomes of non-photosynthetic eukaryotes but also from the plastid genomes of phototrophs, i.e., Euglenophyceae (Hallick et al. 1993; Hrdá et al. 2012). Subsequently, genes encoding subunits of photosystems (psa, psb and pet) are lost, and thereafter the loss of genes encoding RNA-polymerase subunits (rpo) and transcription (inf) and translation factors (tufA) occurs. The next reductive step includes the loss of atp, rbcL and non-essential house-keeping plastid genes. The final reductive stage includes the loss of other metabolic genes (clpP, accD) and housekeeping genes (remaining rpl, rps, rpo and trnE) (Wicke et al. 2016). Recently, this model has been slightly modified. It is proposed that except for ndh, psa, psb and pet genes, most plastid-encoded genes can persist in plastid genomes until the complete transition to heterotrophy occurs (Graham et al. 2017).
The distribution of plastid genes among non-photosynthetic plastids (Table 1) suggests that not all lineages have lost plastid genes in the order suggested by the model described above. The model proposed here (Fig. 3) combines models proposed earlier for land plants (Barrett and Davis 2012; Barrett et al. 2014; Graham et al. 2017) with the patterns of gene retention/loss observed in protists (Table 1; Fig. 3).
The model for the order of chloroplast gene losses in land plants and protists. The model for the order of plastid gene losses proposed earlier by Barrett and Davis (2012), Barrett et al. (2014) and Graham et al. (2017) in land plants is represented by the upper blue line. ndh genes are lost first followed by photosynthesis-related psa, psb and pet genes. The rpl and rps genes encoding ribosomal proteins are being lost from the plastid genome continuosly (interrupted line). These reductive steps are common to land plants and protists (genes in black rectangles in the middle of the figure). After the loss of photosynthetic genes from the plastid genomes of land plants, rpo genes are lost next. The RNA polymerase rpo genes can be lost from plastid genomes of land plants easily, because plants also possess nucleus-encoded phage-derived polymerase. In contrast, rpo genes in protist plastid genomes have to be maintained until the plastid genomes are completely lost, because protists do not possess a second RNA polymerase mechanism. After the loss of rpo genes from plant plastid genomes, the loss of rbcL and atp genes occurs. Housekeeping (rrn, the rest of the rpl and rps genes) and essential metabolic genes (such as accD encoding acetyl-CoA carboxylase used for de novo synthesis of fatty acids and trnE, essential for the synthesis of heme) dissapear from the plant plastid genomes last (blue rectangles, blue line). After the loss of ndh and photosynthetic genes from protist plastid genomes, the loss of rbcL and atp genes occurs (lower red line, red rectangles). The last step in the reduction of protist plastid genomes includes the loss of housekeeping genes (rrn, the remaining rpl and rps genes, rpo, and the tufA gene which is not present in plastid genomes of land plants) and of essential metabolic genes found in most reduced plant plastid genomes (red boxes). accD (a red rectangle with a question mark) has been found to be encoded only in the plastid genomes of Archaeplastida
The first difference from the previous model is the timing of the loss of rpo genes, which are lost much earlier in non-photosynthetic land plants than in protists. The probable reason for this difference is the fact that land plants possess two types of RNA polymerases—subunits of one of them are encoded by the plastid genome (cyanobacterial PEP) (Hu and Bogorad 1990; Hu et al. 1991) and a single subunit of the phage-like (NEP) polymerase is encoded by the nuclear genome (phage-derived NEP) (Allison et al. 1996), while protists possess only a single PEP complex partly encoded by the plastid genome (Smith and Purton 2002). Since the ancestor of land plants likely possessed two mechanisms for plastid transcription, they are more prone to the loss of rpo plastid genes than protists, because after their loss, they can still use the second NEP mechanism (for a review see Shiina et al. 2005). In contrast, the protists are completely dependent upon plastidial PEP RNA-polymerase, and thus they retain rpo plastid genes until their plastid genome is completely lost.
The second difference is the timing of the loss of the tufA gene encoding a translation factor. This gene is retained in all plastid genomes of non-protosynthetic protists, but absent from all plastid genomes of non-photosynthetic land plants. The reason for this difference is unknown, but it could be theoretically explained by one successful transfer of this gene into the nucleus (or its functional replacement by a nucleus-encoded protein) in the common ancestor of land plants followed by the loss of this gene from the plastid genome.
The third and last difference concerns the interesting case of accD, which encodes a subunit of acetyl-CoA carboxylase. This gene is absent from all complex non-photosyntetic plastids (i.e. apicoplasts, E. longa, etc.), but it is present in almost all studied genomes of non-photosynthetic plastids in Archaeplastida lineage with three exceptions (P. uvella, M. uniflora and C. deserticola). In M. uniflora and C. deserticola, accD is present as a pseudogene indicating that the function of this plastid gene has been lost only very recently, while in P. uvella, this gene seems to be absent from plastid genome completely (Table 1). Nevertheless, it might be possible that in these three cases, the functional version of this gene has been transferred to the nucleus and protein products are targeted back to plastid to keep the pathway functional (Graham et al. 2017). The loss of the accD gene from the plastid genome has been observed in various plant lineages, for example in Poales (Harris et al. 2013; Katayama and Ogihara 1996) and its function has been replaced by a nucleus-encoded ACC enzyme (Cai et al. 2008; Sasaki and Nagano 2004).
Similarly to the previous models, in our model rpl and rps genes are gradually being lost in both land plants and protists, likely hand in hand with their successful transfer into the nucleus, while clpP/clpC (involved in plastid protein degradation), rrn genes encoding plastid rRNAs and the trnE gene encoding Glu-rRNA are kept until the final stage of plastid reduction.
Why are plastid genomes retained in non-photosynthetic plastids?
The existence of EGT—the transfer of genes from organellar genomes to the nucleus—raises the question of why organellar genomes are retained at all. Several hypotheses have been proposed to explain this phenomenon. They assume that some membrane bound components of electron transport chain complexes either have biochemical properties which would complicate their transport across membranes (Popot and de Vitry 1990), or they would cause mis-targeting to the ER (Bjorkholm et al. 2015). The “CoRR” (Co-location for Redox Regulation) hypothesis assumes that the expression of individual organelle-encoded subunits enables fine-tuned regulation of the electron transport chain in the organelles (Allen 1993, 2003). None of these hypotheses is, however, directly applicable to non-photosynthetic plastids, because they generally do not retain electron transport chains.
The “Limited transfer window” hypothesis (Barbrook et al. 2006) relates the number of plastids in a cell to the retention of plastid genes. Since gene transfers from the plastid to the nucleus require plastid lysis, they are unlikely to occur in protists containing only a single plastid which is essential for their survival. This hypothesis could be linked to the evolution of coordination between host and endosymbiont division in the early stages of endosymbiosis. In the beginning of the conversion of an endosymbiont into an organelle, there could have been many endosymbiont cells present in the host cell and a massive EGT could have occurred prior to the host nucleus taking control over endosymbiont division (Doolittle 1998). One of the outcomes of this control is the existence of protist lineages with single plastids. In these cases, the EGT window has been closed (Barbrook et al. 2006). This hypothesis is compellingly supported by experiments involving Chlamydomonas reinhardtii, a unicellular green alga possessing a single chloroplast. It was shown that this species is much less prone to plastid to nucleus gene transfers than tobacco containing hundreds of chloroplasts per cell (Huang et al. 2003; Lister et al. 2003; Stegemann et al. 2003). Smith et al. (2011) analyzed the traces of EGT from plastid to nucleus in 30 genome sequences from mono- and polyplastidial species including monoplastidial apicomplexans, and the extreme rarity of EGT in monoplastidial species was confirmed.
Another hypothesis explaining why non-photosynthetic plastids are still present and why they have retained their genomes suggests that it is due to the presence of some essential tRNA genes, i.e., Glu-tRNA (trnE), a precursor of tetrapyrrole (heme and chlorophyll) biosynthesis (Barbrook et al. 2006). If this is the only pathway for tetrapyrrole synthesis available for an organism, the plastid genome encoding trnE has to be retained. Another example of an essential tRNA encoded by the P. falciparum apicoplast genome is fMet-tRNA, which is utilized by the mitochondrion. The P. falciparum mitochondrial genome is also highly reduced and does not contain any tRNA genes (Wilson and Williamson 1997). All tRNA gene products have to be imported into mitochondria from the cytosol. Since the nucleus-encoded Met-tRNA cannot be processed to prokaryotic fMet-tRNA, this tRNA species has to be imported into mitochondria from the apicoplast (Barbrook et al. 2006).
Various unrelated non-photosynthetic organisms (A. mirabilis, P. stellaris, C. gronovii, C. paramecium, E. longa, P. danica and C. infusionum) have also retained the plastid gene encoding the large subunit of Rubisco (rbcL) (Table 1). The function of the Rubisco protein in these non-photosynthetic plastids is unclear, but it could still be involved in carbon fixation. On the other hand, several alternative functions have been also proposed—it might act as oxygenase, it could be involved in glycine and serine biosynthesis (Miziorko and Lorimer 1983), it could be required for an alternative lipid biosynthesis pathway (Schwender et al. 2004) or it might perform another as-yet unknown function (Sánchez-Puerta et al. 2007; Sekiguchi et al. 2002; Wolfe and de Pamphilis 1998; Záhonová et al. 2016). Hence, the essential nature of rbcL might also prevent chloroplast genome loss.
Another example of essential genes might be clp genes—the clpP gene encoded in the plastid genomes of green algae and the clpC gene encoded in the plastid genomes of red algae and plastids of red algal origin (Table 1). clp genes encode proteases that may function in transit peptide cleavage during protein import into plastids as well as in degradation of aberrant proteins (Adam 2000; Clarke 1999). Despite their important role in many plastids, Cahoon et al. (2003) argued that the clpP gene does not necessarily have to be essential for plant survival due to its dispensability from the plastid genomes of non-photosynthetic mutants or albino plants lacking plastid ribosomes. Therefore, clpP genes might have an essential function only in plastids with active gene expression (Cahoon et al. 2003). However, clp genes are absent from at least some non-photosynthetic plastid genomes (Table 1). In these cases, clp genes might either be present in the nucleus, or they might not be essential at all.
Finally, several relatively well-documented cases (Polytomella, Rafflesia) of complete plastid genome loss exist (Molina et al. 2014; Smith and Lee 2014) (Online Resource 1). Other organisms such as Perkinsus and colpodellids also likely lack plastid genomes while retaining a plastid compartment, although the data are currently not sufficient to be certain about this conclusion (Gile and Slamovits 2014; Grauvogel et al. 2007; Janouškovec et al. 2015; Matsuzaki et al. 2008; Robledo et al. 2011; Teles-Grilo et al. 2007). Nevertheless, it seems that complete loss of chloroplast DNA is indeed possible under certain circumstances, while the genome-less plastid compartment remains functional, as in the case of genome-less derivates of mitochondria—hydrogenosomes and mitosomes (Klinger et al. 2016).
What prevents plastid loss after the loss of photosynthesis?
It is commonly assumed that plastids have been retained in non-photosynthetic organisms because they perform essential function(s) unrelated to photosynthesis. Several metabolic pathways frequently operate in non-photosynthetic plastids, but the situation varies among organismal groups. The most common biochemical pathways operating in non-photosynthetic plastids are: (1) tetrapyrrole (heme) synthesis, which requires plastid-encoded Glu-tRNA (encoded by the trnE gene), (2) the non-mevalonate (DOXP) pathway for the synthesis of IPP (precursors for the isoprenoid biosynthesis), and (3) fatty acid biosynthesis (Ralph et al. 2004; for a review see; Seeber and Soldati-Favre 2010). None of the final products of these pathways have to be strictly synthesized in the plastid compartment. There are several examples of organisms in which alternative pathways for the synthesis of the same final products exist in the cytosol or which are able to obtain these products from the environment (the host cell in the case of parasites). In some organisms with photosynthetic ancestors (or with ancestors possessing non-photosynthetic plastids) such as the dinoflagellate Hematodinium sp. or the apicomplexan C. parvum, plastids are undetectable using currently available, highly sophisticated techniques, and the most likely interpretation is their complete loss. The plastid independence of the pathways mentioned above in the ancestors of these organisms likely allowed the complete loss of the plastid compartment. In Hematodinium sp., fatty acids and heme are synthesized in the cytosol and by a combination of cytosolic and mitochondrial pathways, respectively (Gornik et al. 2015). The requirements for IPP essential for isoprenoid synthesis are apparently satisfied by uptake from the host (Gornik et al. 2015). Interestingly, complete plastid loss has been experimentally induced in the bloodstream stages of P. falciparum. After the treatment of this parasite with antibiotics which completely destroy the apicoplast and concomitant supplementation with IPP in the media for growth of the blood stream form, cells lacking apicoplasts remained viable and divided (Yeh and DeRisi 2011). C. parvum represents an example of an apicomplexan which naturally lacks apicoplasts (Abrahamsen et al. 2004; Zhu et al. 2000). Plastid independence of the FAS pathway was a likely precondition for apicoplast loss in this case.
Taken together, examples from nature as well as laboratory experiments have shown that there is no universal reason preventing the loss of non-photosynthetic plastids. Once evolution “blindly finds a solution” for a cell to compensate for essential plastid-localized biochemical pathways, the plastid compartment can be lost. Since the maintenance of a useless organelle is costly (the cost of retaining mitochondria has been discussed, e.g., by Lynch and Marinov 2017), the retention of non-photosynthetic plastids for performing just one or a few essential metabolic pathways may be viewed also as an evolutionary trap from which only a few species have successfully escaped.
Conclusion
In this review, we have summarized current knowledge of non-photosynthetic plastids which have arisen independently by reductive evolutionary processes among several unrelated phototrophic eukaryotic lineages. The vast majority of non-photosynthetic plastids contain genomes lacking any genes whose products are directly involved in photosynthesis in their photosynthetic relatives. By the comparison of coding capacity of plastid genomes present in non-photosynthetic plastids, we have proposed an updated model for the order of plastid gene losses after (or concomitant with) the transition to an obligatory heterotrophic lifestyle (Fig. 3). Although the scenarios for plastid gene loss are quite similar among evolutionarily distant groups, i.e., losing ndh, psa, psb and most pet genes as the first ones and clp, trnE and rrn genes as the last ones, some differences between certain lineages should be stressed: (1) relatively early loss of rpo genes in most land plants and their retention in all unicellular eukaryotes, (2) the retention of the accD gene in Archaeplastida but none of the other phylogenetic groups, and (3) the retention of tufA by all non-photosynthetic plastid genomes except for those of non-photosynthetic land plants (Table 1; Fig. 3).
In contrast to quite frequent massive reduction of gene content in non-photosynthetic plastid genomes, complete plastid genome loss is extremely rare but possible. Some essential biochemical pathways are restricted exclusively to non-photosynthetic genome-less plastid compartments and they cannot be easily replaced by pathways localized to other compartments. This might be a likely reason for the retention of a genome-less plastid. Loss of the plastid genome combined with retention of the plastid compartment has been documented in a parasitic plant, R. lagascae, in four free-living algal species of the Polytomella genus and an alveolate parasite Perkinsus sp. (Figs. 1, 2), but plastid genome loss is potentially more frequent among alveolates (e.g., candidate genome-less plastids in colpodellids).
Curiously, the complete loss of the plastid genome (and likely the plastid itself) may be induced in the laboratory in E. gracilis during bleaching by antibiotics without any addition of compounds which would compensate for any plastid-localized pathway (Krajčovič et al. 2002), as well as in P. falciparum after antibiotics-induced destruction of apicoplasts and IPP-donation-mediated rescue in the bloodstream stage (Yeh and DeRisi 2011). Although the complete loss of the plastid compartment seems to be even less frequent than plastid genome loss, these experiments suggest that it is principally possible even in experimental models under certain conditions. Hence, it is likely that complete plastid loss, though extremely rare, is possible also in nature and it has been indeed credibly documented for four protists—apicomplexans Cryptosporidium parvum (Abrahamsen et al. 2004; Zhu et al. 2000), Gregarina niphandrodes (Toso and Omoto 2007) and Ascogregarina taiwanensis (Templeton et al. 2010), and the marine parasitic dinoflagellate Hematodinium sp. (Gornik et al. 2015) (Fig. 1).
We propose that the distantly related non-photosynthetic lineages with non-photosynthetic plastids are caught in an evolutionary trap that does not permit the loss of the chloroplast compartment in which at least one essential metabolic pathway is performed. In only a few cases, evolution has found a way to compensate for the essential plastid-localized pathways and thereafter, natural selection favored the loss of the compartment—a way out of the trap. Such solutions might include the relocalization of pathway to other compartment(s), horizontal acquisition of a pathway compensating for the plastid-localized pathway or evolution of a mechanism for the uptake of the essential plastid metabolite from the external environment or host. Interestingly, most organisms for which complete plastid loss has been proposed belong to Myzozoa, one of the major eukaryotic lineages whose ancestor likely possessed a complex plastid(s) of red algal origin, and they are parasites. This suggests that Myzozoa might be more prone to plastid loss than any other eukaryotic lineage and that parasitism is in most cases an important prerequisite for plastid loss.
Abbreviations
- EGT:
-
Endosymbiotic gene transfer
- TOC:
-
Translocon of outer chloroplast membrane
- TIC:
-
Translocon of inner chloroplast membrane
- Glu-tRNA:
-
Glutamyl-tRNA
- acetyl-CoA:
-
Acetyl co-enzyme A
- ACC:
-
Acetyl-CoA carboxylase
- IR:
-
Inverted repeat
- LSC:
-
Large single copy
- SSC:
-
Small single copy
- fMet-tRNA:
-
Formylmetionyl-tRNA
- Met-tRNA:
-
Metionyl-tRNA
- DOXP:
-
Non-mevalonate isoprenoid biosynthesis pathway
- IPP:
-
Isopentenyl pyrophosphate
- FAS:
-
Fatty acid synthesis
- CoRR:
-
Co-location for redox regulation
References
Abrahamsen MS, Templeton TJ, Enomoto S et al (2004) Complete genome sequence of the apicomplexan, Cryptosporidium parvum. Science 16:441–445
Adam Z (2000) Chloroplast proteases: possible regulators of gene expression? Biochimie 82(6):647–654
Adl SM, Simpson AG, Lane CE et al (2012) The revised classification of eukaryotes. J Eukaryot Microbiol 59(5):429–514
Allen JF (1993) Control of gene expression by redox potential and the requirement for chloroplast and mitochondrial genomes. J Theor Biol 165:609–631
Allen JF (2003) The function of genomes in bioenergetic organelles. Philos Trans R Soc Lond B Biol Sci 358(1429):19–38
Allison LA, Simon LD, Maliga P (1996) Deletion of rpoB reveals a second distinct transcription system in plastids of higher plants. EMBO J 15(11):2802
Archibald JM (2009) The puzzle of plastid evolution. Curr Biol 19(2):R81–R88
Archibald JM (2015) Genomic perspectives on the birth and spread of plastids. PNAS 112(33):10147–10153
Archibald JM, Keeling PJ (2002) Recycled plastids: a ‘green movement’ in eukaryotic evolution. Trends Genet 18(11):577–584
Arisue N, Hashimoto T, Mitsui H et al (2012) The Plasmodium apicoplast genome: conserved structure and close relationship of P. ovale to rodent malaria parasites. Mol Biol Evol 29(9):2095–2099
Atteia A, van Lis R, Beale SI (2005) Enzymes of the heme biosynthetic pathway in the nonphotosynthetic alga Polytomella sp. Eukaryot Cell 4:2087–2097
Barbrook AC, Howe CJ, Purton S (2006) Why are plastid genomes retained in nonphotosynthetic organisms? Trends Plant Sci 11:101–108
Barrett CF, Davis JI (2012) The plastid genome of the mycoheterotrophic Corallorhiza striata (Orchidaceae) is in the relatively early stages of degradation. Am J Bot 99:1513–1523
Barrett CF, Freudenstein JV (2008) Molecular evolution of rbcL in the mycoheterotrophic coralroot orchids (Corallorhiza Gagnebin, Orchidaceae). Mol Phylogen Evol 47:665–679
Barrett CF, Freudenstein JV, Li J et al (2014) Investigating the path of plastid genome degradation in an early-transitional clade of heterotrophic orchids, and implications for heterotrophic angiosperms. Mol Biol Evol 31:3095–3112
Baurain D, Brinkmann H, Petersen J et al (2010) Phylogenomic evidence for separate acquisition of plastids in cryptophytes, haptophytes, and stramenopiles. Mol Biol Evol 27(7):1698–1709
Beale SI (1999) Enzymes of chlorophyll biosynthesis. Photosynth Res 60:43–73
Becker B, Hoef-Emden K, Melkonian M (2008) Chlamydial genes shed light on the evolution of photoautotrophic eukaryotes. BMC Evol Biol 8:203
Belcher JH, Swale EMF (1976) Spumella elongata nov. comb.—a colourless flagellate from soil. Arch Protistenkd 118:215–220
Bellot S, Renner SS (2016) The plastomes of two species in the endoparasite genus Pilostyles (Apodanthaceae) each retain just five or six possibly functional genes. Genome Biol Evol 8(1):189–201
Ben Ali A, De Baerea R, De Wachtera R, Van de Peer Y (2002) Evolutionary relationships among heterokont algae (the autotrophic stramenopiles) based on combined analyses of small and large subunit ribosomal RNA. Protist 153:123–132
Bidartondo MI, Bruns TD, Weiss M, Sérgio C, Read DJ (2003) Specialized cheating of the ectomycorrhizal symbiosis by an epiparasitic liverwort. Proc R Soc Lond B Biol Sci 270(1517):835–842
Bjorkholm P, Harish A, Hagstrom E, Ernst AM, Andersson SG (2015) Mitochondrial genomes are retained by selective constraints on protein targeting. Proc Natl Acad Sci USA 112:10154–10161
Blouin NA, Lane CE (2012) Red algal parasites: models for a life history evolution that leaves photosynthesis behind again and again. Bioessays 34(3):226–235
Bodył A (1996) Is the origin of Astasia longa an example of the inheritance of acquired characteristics? Acta Protozool 35:87–94
Bodył A, Mackiewicz P, Stiller JW (2010) Comparative genomic studies suggest that the cyanobacterial endosymbionts of the amoeba Paulinella chromatophora possess an import apparatus for nuclear-encoded proteins. Plant Biol 12(4):639–649
Bölter B, Soll J (2017) Ycf1/Tic214 is not essential for the accumulation of plastid proteins. Mol Plant 10(1):219–221
Borza T, Popescu CE, Lee RW (2005) Multiple metabolic roles for the nonphotosynthetic plastid of the green alga Prototheca wickerhamii. Eukaryot Cell 4:253–261
Braukmann T, Kuzmina M, Stefanović S (2013) Plastid genome evolution across the genus Cuscuta (Convolvulaceae): two clades within subgenus Grammica exhibit extensive gene loss. J Exp Bot 64:977–989
Brayton KA, Lau AO, Herndon DR et al (2007) Genome sequence of Babesia bovis and comparative analysis of apicomplexan hemoprotozoa. PLoS Pathog 3(10):e148
Burki F (2014) The eukaryotic tree of life from a global phylogenomic perspective. Cold Spring Harb Perspect Biol 6(5):a016147
Cahoon AB, Cunningham KA, Stern DB (2003) The plastid clpP gene may not be essential for plant cell viability. Plant Cell Physiol 44(1):93–95
Cai X, Fuller AL, McDougald LR, Zhu G (2003) Apicoplast genome of the coccidian Eimeria tenella. Gene 321:39–46
Cai Z, Guisinger M, Kim HG et al (2008) Extensive reorganization of the plastid genome of Trifolium subterraneum (Fabaceae) is associated with numerous repeated sequences and novel DNA insertions. J Mol Evol 67(6):696–704
Callow JA, Callow ME, Evans LV (1979) Nutritional studies of the parasitic red alga Choreocolax polysiphoniae. New Phytol 83:451–462
Cavalier-Smith T (1999) Principles of protein and lipid targeting in secondary symbiogenesis: euglenoid, dinoflagellate, and sporozoan plastid origins and the eukaryote family tree. J Eukaryot Microbiol 46(4):347–366
Cavalier-Smith T (2002) The phagotrophic origin of eukaryotes and the phylogenetic classification of Protozoa. Int J Syst Evol Microbiol 52:297–354
Cavalier-Smith T, Chao EE (2004) Protalveolate phylogeny and systematics and the origins of Sporozoa and dinoflagellates (phylum Myzozoa nom. nov.). Eur J Protistol 40(3):185–212
Cavalier-Smith T, Lee JJ (1985) Protozoa as hosts for endosymbioses and the conversion of symbionts into organelles. J Eukaryot Microbiol 32(3):376–379
Chesnick JM, Hooistra WH, Wellbrock U, Medlin LK (1997) Ribosomal RNA analysis indicates a benthic pennate diatom ancestry for the endosymbionts of the dinoflagellates Peridinium foliaceum and Peridinium balticum (Pyrrhophyta). J Eukaryot Microbiol 44:314–320
Cinar HN, Qvarnstrom Y, Wei-Pridgeon Y et al (2016) Comparative sequence analysis of Cyclospora cayetanensis apicoplast genomes originating from diverse geographical regions. Parasit Vectors 9(1):611
Clarke AK (1999) ATP-dependent Clp proteases in photosynthetic organisms—a cut above the rest! Ann Bot 83(6):593–599
Court GJ (1980) Photosynthesis and translocation studies of Laurencia spectabilis and its symbiont Janczewskia gardneri (Rhodophyceae). J Phycol 16:270–279
de Koning AP, Keeling PJ (2004) Nucleus-encoded genes for plastid-targeted proteins in Helicosporidium: functional diversity of a cryptic plastid in a parasitic alga. Eukaryot Cell 3:1198–1205
de Koning AP, Keeling PJ (2006) The complete plastid genome sequence of the parasitic green alga Helicosporidium sp. is highly reduced and structured. BMC Biol 4:12
de Vries J, Sousa FL, Bölter B, Soll J, Gould SB (2015) YCF1: a green TIC? Plant Cell 27(7):1827–1833
de Vries J, Archibald JM, Gould SB (2017) The carboxy terminus of YCF1 contains a motif conserved throughout > 500 Myr of streptophyte evolution. Genome Biol Evol 9(2):473–479
Delannoy E, Fijii S, Colas des Francs C, Brundett M, Small I (2011) Rampant gene loss in the underground orchid Rhizanthella gardneri highlights evolutionary constraints on plastid genomes. Mol Biol Evol 28:2077–2086
Delwiche CF, Palmer JD (1996) Rampant horizontal transfer and duplication of Rubisco genes in eubacteria and plastids. Mol Biol Evol 13(6):873–882
Deschamps P, Colleonic C, Nakamura Y et al (2008) Metabolic symbiosis and the birth of the plant kingdom. Mol Biol Evol 25:536–548
Deusch O, Landan G, Roettger M et al (2008) Genes of cyanobacterial origin in plant nuclear genomes point to heterocyst-forming plastid ancestor. Mol Biol Evol 25:748–761
do Vieira LN,·dos Anjos KG, Faoro H, Fraga HPdeF, Greco TM, Pedrosa FdeO, de Souza EM, Rogalski M, de Souza RF, Guerra MP (2016) Phylogenetic inference and SSR characterization of tropical woody bamboos tribe Bambuseae (Poaceae: Bambusoideae) based on complete plastid genome sequences. Curr Genet 62:443–453. doi:10.1007/s00294-015-0549-z
Dodge JD (1969) Observations on the fine structure of the eyespot and associated organelles in the dinoflagellate Glenodinium foliaceum. J Cell Sci 5:479–93
Domman D, Horn M, Embley TM, Williams TA (2015) Plastid establishment did not require a chlamydial partner. Nat Commun 6:6421. doi:10.1038/ncomms7421
Donaher N, Tanifuji G, Onodera NT et al (2009) The complete plastid genome sequence of the secondarily nonphotosynthetic alga Cryptomonas paramecium: reduction, compaction, and accelerated evolutionary rate. Genome Biol Evol 1:439–448
Doolittle WF (1998) You are what you eat: a gene transfer ratchet could account for bacterial genes in eukaryotic nuclear genomes. Trends Genet 14(8):307–311
Dorrell RG, Gile G, McCallum Méheust R, Bapteste EP, Klinger CM, Brillet-Guéguen L, Freeman KD, Richter DJ, Bowler C (2017) Chimeric origins of ochrophytes and haptophytes revealed through an ancient plastid proteome. eLife 2017 6:e23717. doi:10.7554/eLife.23717
Douglas SE, Murphy CA, Spencer DF, Gray MW (1991) Cryptomonad algae are evolutionary chimaeras of two phylogenetically distinct unicellular eukaryotes. Nature 350(6314):148–151
Facchinelli F, Colleoni C, Ball SG, Weber AP (2013) Chlamydia, cyanobiont, or host: Who was on top in the ménage à trois? Trends Plant Sci 18(12):673–679
Falkowski PG, Katz ME, Knoll AH et al (2004) The evolution of modern eukaryotic phytoplankton. Science 305(5682):354–360
Figueroa-Martinez F, Nedelcu AM, Smith DR, Reyes-Prieto A (2017a) The plastid genome of Polytoma uvella is the largest known among colorless algae and plants and reflects contrasting evolutionary paths to nonphotosynthetic lifestyles. Plant Physiol 173(2):932–943
Figueroa-Martinez F, Nedelcu AM, Reyes-Prieto A, Smith DR (2017b) The plastid genomes of nonphotosynthetic algae are not so small after all. Commun Integ Biol 10(1):e1283080
Funk HT, Berg S, Krupinska K, Maier UG, Krause K (2007) Complete DNA sequences of the plastid genomes of two parasitic flowering plant species, Cuscuta reflexa and Cuscuta gronovii. BMC Plant Biol 7:45
Gagat P, Bodyl A, Mackiewicz P (2013) How protein targeting to primary plastids via the endomembrane system could have evolved? A new hypothesis based on phylogenetic studies. Biol Direct 8(1):18
Gaillard T, Madamet M, Tsombeng FF, Dormoi J, Pradines B (2016) Antibiotics in malaria therapy: which antibiotics except tetracyclines and macrolides may be used against malaria? Malar J 15(1):556
Gardner MJ, Bishop R, Shah T et al (2005) Genome sequence of Theileria parva, a bovine pathogen that transforms lymphocytes. Science 309(5731):134–137
George B, Bhatt BS, Awasthi M, George B, Singh AK (2015) Comparative analysis of microsatellites in chloroplast genomes of lower and higher plants. Curr Genet 61:665–677
Gile GH, Slamovits CH (2014) Transcriptomic analysis reveals evidence for a cryptic plastid in the colpodellid Voromonas pontica, a close relative of chromerids and apicomplexan parasites. PloS one 9(5):e96258
Gilson PR, Su V, Slamovits CH et al (2006) Complete nucleotide sequence of the chlorarachniophyte nucleomorph: nature’s smallest nucleus. Proc Natl Acad Sci 103(25):9566–9571
Giovannoni S, Turner S, Olsen G et al (1988) Evolutionary relationships among cyanobacteria and green chloroplasts. J Bacteriol 170:3584–3592
Gockel G, Hachtel W (2000) Complete gene map of the plastid genome of the nonphotosynthetic euglenoid flagellate Astasia longa. Protist 151:347–351
Gockel G, Hachtel W, Baier S et al (1994) Genes for chloroplast apparatus are conserved in the reduced 73-kb plastid DNA of the nonphotosynthetic euglenoid flagellate Astasia longa. Curr Genet 26:256–262
Goff LJ, Coleman AW (1995) Fate of parasite and host organelle DNA during cellular transformation of red algae by their parasites. Plant Cell 7(11):1899–1911
Goff LJ, Zuccarello G (1994) The evolution of parasitism in red algae: Cellular interactions of adelphoparasites and their hosts. J Phycol 30:695–720
Gogniashvili M, Jinjikhadze T, Maisaia I, Akhalkatsi M, Adam Kotorashvili A, Kotaria N, Beridze T, Dudnikov AJ (2016) Complete chloroplast genomes of Aegilops tauschii Coss. and Ae. cylindrica Host sheds light on plasmon D evolution. Curr Genet 62:791–798. doi:10.1007/s00294-016-0583-5
Gornik SG, Cassin AM, MacRae JI et al (2015) Endosymbiosis undone by stepwise elimination of the plastid in a parasitic dinoflagellate. Proc Natl Acad Sci 112(18):5767–5772
Graham SW, Lam VK, Merckx VS (2017) Plastomes on the edge: the evolutionary breakdown of mycoheterotroph plastid genomes. New Phytol 214(1):48–55
Grant J, Tekle YI, Anderson OR, Patterson DJ, Katz LA (2009) Multigene evidence for the placement of a heterotrophic amoeboid lineage Leukarachnion sp. among photosynthetic stramenopiles. Protist 160(3):376–385
Grauvogel C, Reece KS, Brinkmann H, Petersen J (2007) Plastid isoprenoid metabolism in the oyster parasite Perkinsus marinus connects dinoflagellates and malaria pathogens-new impetus for studying alveolates. J Mol Evol 65:725–729
Gray MW, Burger G, Lang BF (1999) Mitochondrial evolution. Science 283(5407):1476–1481
Green BR (2011) Chloroplast genomes of photosynthetic eukaryotes. Plant J 66(1):34–44
Gruzdev EV, Mardanov AV, Beletsky AV et al (2016) The complete chloroplast genome of parasitic flowering plant Monotropa hypopitys: extensive gene losses and size reduction. Mitochondrial DNA Part B Resour 1(1):212–213
Hackett JD, Maranda L, Yoon HS, Bhattacharya D (2003) Phylogenetic evidence for the cryptophyte origin of the plastid of Dinophysis (Dinophysiales, Dinophyceae). J Phycol 39:440–448
Hadariová L, Vesteg M, Birčák E, Schwartzbach SD, Krajčovič J (2017) An intact plastid genome is essential for the survival of colorless Euglena longa but not Euglena gracilis. Curr Genet 63(2):331–334
Hallick RB, Hong L, Drager RG et al (1993) Complete sequence of Euglena gracilis chloroplast DNA. Nucleic Acids Res 21:3537–3544
Harris ME, Meyer G, Vandergon T, Vandergon VO (2013) Loss of the acetyl-CoA carboxylase (accD) gene in Poales. Plant Mol Biol Report 31(1):21–31
Hehenberger E, Imanian B, Burki F, Keeling PJ (2014) Evidence for the retention of two evolutionary distinct plastids in dinoflagellates with diatom endosymbionts. Gen Biol Evol 6(9):2321–2334
Hoef-Emden K (2005) Multiply independent losses of photosynthesis and differing evolutionary in the genus Cryptomonas (Cryptophyceae): Combined phylogenetic analyses of DNA sequences of the nuclear and the nucleomorph ribosomal operons. J Mol Evol 60:183–195
Howe CJ, Smith AG (1991) Plants without chlorophyll. Nature 349:109
Hrdá Š, Fousek J, Szabová J, Hampl V, Vlček Č (2012) The plastid genome of Eutreptiella provides a window into the process of secondary endosymbiosis of plastid in euglenids. PLoS One 7(3):e33746
Hu J, Bogorad L (1990) Maize chloroplast RNA polymerase: the 180-, 120-, and 38-kilodalton polypeptides are encoded in chloroplast genes. Proc Natl Acad Sci 87(4):1531–1535
Hu J, Troxler RF, Bogorad L (1991) Maize chloroplast RNA polymerase: the 78-kilodalton polypeptide is encoded by the plastid rpoC1 gene. Nucleic Acids Res 19(12):3431–3434
Huang CY, Ayliffe MA, Timmis JN (2003) Direct measurement of the transfer rate of chloroplast DNA into the nucleus. Nature 422(6927):72
Huang Y, He L, Hu J et al (2015) Characterization and annotation of Babesia orientalis apicoplast genome. Parasites vectors 8(1):543
Imura T, Sato S, Sato Y et al (2014) The apicoplast genome of Leucocytozoon caulleryi, a pathogenic apicomplexan parasite of the chicken. Parasitol Res 113(3):823–828
Janouškovec J, Tikhonenkov DV, Burki F et al (2015) Factors mediating plastid dependency and the origins of parasitism in apicomplexans and their close relatives. Proc Natl Acad Sci 112(33):10200–10207
Janouškovec J, Gavelis GS, Burki F et al (2017) Major transitions in dinoflagellate evolution unveiled by phylotranscriptomics. Proc Natl Acad Sci 114(2):E171–E180
Kamikawa R, Tanifuji G, Ishikawa SA et al (2015a) Proposal of a twin-arginine translocator system-mediated constraint against loss of ATP synthase genes from nonphotosynthetic plastid genomes. Mol Biol Evol 32:2598–2604
Kamikawa R, Yubuki N, Yoshida M et al (2015b) Multiple losses of photosynthesis in Nitzschia (Bacillariophyceae). Phycol Res 63(1):19–28
Kamikawa R, Moog D, Zauner S et al (2017) A non-photosynthetic diatom reveals early steps of reductive evolution in plastids. Mol Biol Evol 34(9):2355–2366
Karnkowska A, Vacek V, Zubáčová Z et al (2016) A eukaryote without a mitochondrial organelle. Curr Biol 26(10):1274–1284
Katayama H, Ogihara Y (1996) Phylogenetic affinities of the grasses to other monocots as revealed by molecular analysis of chloroplast DNA. Curr Genet 29(6):572–581
Keeling PJ (2013) The number, speed, and impact of plastid endosymbioses in eukaryotic evolution. Annu Rev Plant Biol 64:583–607
Kikuchi S, Bédard J, Hirano M et al (2013) Uncovering the protein translocon at the chloroplast inner envelope membrane. Science 339(6119):571–574
Kleffmann T, Russenberger D, von Zychlinski A et al (2004) The Arabidopsis thaliana chloroplast proteome reveals pathway abundance and novel protein functions. Curr Biol 14(5):354–362
Kleine T, Maier UG, Leister D (2009) DNA transfer from organelles to the nucleus: the idiosyncratic genetics of endosymbiosis. Annu Rev Plant Biol 60:115–138
Klinger CM, Nisbet RE, Ouologuem DT, Roos DS, Dacks JB (2013) Cryptic organelle homology in apicomplexan parasites: insights from evolutionary cell biology. Curr Opin Microbiol 16(4):424–431
Klinger CM, Karnkowska A, Herman EK, Hampl V, Dacks JB (2016) Phylogeny and evolution. In: Molecular parasitology. Springer, Vienna, pp 383–408
Knauf U, Hachtel W (2002) The genes encoding subunits of ATP synthase are conserved in the reduced plastid genome of the heterotrophic alga Prototheca wickerhamii. Mol Genet Genom 267(4):492–497
Kohzuma K, Froehlich JE, Davis GA, Temple JA, Minhas D, Dhingra A, Cruz JA, Kramer DM (2017) The role of light–dark regulation of the chloroplast ATP synthase. Front Plant Sci 8:1248. doi:10.3389/fpls.2017.01248
Kong W, Yang J (2016) The complete chloroplast genome sequence of Morus mongolica and a comparative analysis within the Fabidae clade. Curr Genet 62:165–172. doi:10.1007/s00294-015-0507-9
Krajčovič J, Ebringer L, Schwartzbach SD (2002) Reversion of endosymbiosis? The case of bleaching in Euglena. In: Systems J, Seckbach (eds) Symbiosis: mechanism and model. Kluwer Academic Publishers, Dordrecht, pp 185–206
Krajčovič J, Vesteg M, Schwartzbach SD (2015) Euglenoid flagellates: a multifaceted biotechnology platform. J Biotechnol 202:135–145
Krause K (2008) From chloroplasts to “cryptic” plastids: evolution of plastid genomes in parasitic plants. Curr Genet 54:111–121
Ku C, Nelson-Sathi S, Roettger M et al (2015) Endosymbiotic gene transfer from prokaryotic pangenomes: Inherited chimerism in eukaryotes. Proc Natl Acad Sci 112(33):10139–10146
Kugrens P, West JA (1973) The ultrastructure of an alloparasitic red alga Choreocolax polysiphoniae. Phycol 12(3–4):175–186
Lam VKY, Gomez MS, Graham SW (2015) The highly reduced plastome of mycoheterotrophis Sciaphila (Triuridaceae) is colinear with its green relatives and is under strong purifying selection. Genome Biol Evol 7:2220–2236
Leger MM, Kolisko M, Kamikawa R et al (2017) Organelles that illuminate the origins of Trichomonas hydrogenosomes and Giardia mitosomes. Nat Ecol Evol 1:0092
Li X, Zhang TC, Qiao Q et al (2013) Complete chloroplast genome sequence of holoparasite Cistanche deserticola (Orobanchaceae) reveals gene loss and horizontal gene transfer from its host Haloxylon ammodendron (Chenopodiaceae). PloS one 8(3):e58747
Lim GS, Barrett CF, Pang CC, Davis JI (2016) Drastic reduction of plastome size in the mycoheterotrophic Thismia tentaculata relative to that of its autotrophic relative Tacca chantrieri. Am J Bot 103(6):1129–1137
Lister DL, Bateman JM, Purton S, Howe CJ (2003) DNA transfer from chloroplast to nucleus is much rarer in Chlamydomonas than in tobacco. Gene 316:33–38
Logacheva MD, Schelkunov MI, Penin AA (2011) Sequencing and analysis of plastid genome in mycoheterotrophic orchid Neottia nidus-avis. Genome Biol Evol 3:1296–1303
Logacheva MD, Schelkunov MI, Nuraliev MS, Samigullin TH, Penin AA (2014) The plastid genome of mycoheterotrophic monocot Petrosavia stellaris exhibits both gene losses and multiple rearrangements. Genome Biol Evol 6(1):238–246
Logacheva MD, Schelkunov MI, Shtratnikova VY, Matveeva MV, Penin AA (2016) Comparative analysis of plastid genomes of non-photosynthetic Ericaceae and their photosynthetic relatives. Sci Rep 6:30042
Lynch M, Marinov GK (2017) Membranes, energetics, and evolution across the prokaryote-eukaryote divide. eLife 6:e20437
Mackiewicz P, Bodył A, Gagat P (2012) Possible import routes of proteins into the cyanobacterial endosymbionts/plastids of Paulinella chromatophora. Theory Biosci 131(1):1–18
Maier UG, Douglas SE, Cavalier-Smith T (2000) The nucleomorph genomes of cryptophytes and chlorarachniophytes. Protist 151(2):103–109
Maier UG, Zauner S, Hempel F (2015) Protein import into complex plastids: cellular organization of higher complexity. Eur J Cell Biol 94(7):340–348
Makiuchi T, Nozaki T (2014) Highly divergent mitochondrion-related organelles in anaerobic parasitic protozoa. Biochimie 100:3–17
Marin B (2004) Origin and fate of chloroplasts in the euglenoida. Protist 155:13–14
Marin B, Palm A, Klingberg M, Melkonian M (2003) Phylogeny and taxonomic revision of plastid-containing euglenophytes based on SSU rDNA sequence comparison and synapomorphic signatures in the SSU rRNA secondary structure. Protist 154:99–145
Marin B, Nowack CM, Melkonian M (2005) A plastid in the making: evidence for a second primary endosymbiosis. Protist 156:425–432
Martin M, Sabater B (2010) Plastid ndh genes in plant evolution. Plant Physiol Biochem 48:636–645
Martin W, Stoebe B, Goremykin V et al (1998) Gene transfer to the nucleus and the evolution of chloroplasts. Nature 393:162–165
Matsuzaki M, Kuroiwa H, Kuroiwa T, Kita K, Nozaki H (2008) A cryptic algal group unveiled: a plastid biosynthesis pathway in the oyster parasite Perkinsus marinus. Mol Biol Evol 25:1167–1179
McFadden GI (2001) Primary and secondary endosymbiosis and the origin of plastids. J Phycol 37(6):951–959
McFadden GI, Yeh E (2017) The apicoplast: now you see it, now you don’t. Int J Parasitol 47(2):137–144
McNeal JR, Arumugunathan K, Kuehl JV, Boore JL, dePamphilis CV (2007) Systematics and plastid genome evolution of the cryptically photosynthetic parasitic plant genus Cuscuta (Convolvulaceae). BMC Biol 5:55
Merckx V, Freudenstein JV (2010) Evolution of mycoheterotrophy in plants: a phylogenetic perspective. New Phytol 185(3):605–609
Mereschkowsky C (1905) Über Natur und Ursprung der Chromatophoren im Pflanzenreiche. Biol Centralbl 25:593–604 (English translation In: Martin W, Kowallik KV (1999) Eur J Phycol 34:287–295)
Miziorko HM, Lorimer GH (1983) Ribulose-1, 5-bisphosphate carboxylase-oxygenase. Annu Rev Biochem 52(1):507–535
Molina J, Hazzouri KM, Nickrent D et al (2014) Possible loss of the chloroplast genome in the parasitic flowering plant Rafflesia lagascae (Rafflesiaceae). Mol Biol Evol 31:793–803
Moore CE, Archibald JM (2009) Nucleomorph genomes. Annu Rev Genet 43:251–264
Morse D, Salois P, Markovic P, Hastings JW (1995) A nuclear-encoded form II RuBisCO in dinoflagellates. Science 268(5217):1622
Moustafa A, Reyes-Prieto A, Bhattacharya D (2008) Chlamydiae have contributed at least 55 genes to Plantae with predominantly plastid functions. PloS One 3:e2205
Mukherjee A, Sadhukhan GC (2016) Anti-malarial drug design by targeting Apicoplasts: new perspectives. J Pharmacopunct 19(1):7
Müller M, Mentel M, van Hellemond JJ et al (2012) Biochemistry and evolution of anaerobic energy metabolism in eukaryotes. Microbiol Mol Biol Rev 76(2):444–495
Müllner AN, Angeler DG, Samuel R, Linton EW, Triemer RE (2001) Phylogenetic analysis of phagotrophic, phototrophic and osmotrophic euglenoids by using the nuclear 18S rDNA sequence. Int J Syst Evol Microbiol 51:783–791
Muñoz-Gómez SA, Mejía-Franco FG, Durnin K et al (2017) The new red algal subphylum Proteorhodophytina comprises the largest and most divergent plastid genomes known. Curr Biol 27(11):1677–1684
Nakai M (2015a) The TIC complex uncovered: the alternative view on the molecular mechanism of protein translocation across the inner envelope membrane of chloroplasts. Biochim Biophys Acta Bioenerget 1847(9):957–967
Nakai M (2015b) YCF1: a green TIC: response to the de Vries et al. Commentary. Plant Cell 27:1834–1838
Nass MMK, Nass S (1963) Intramitochondrial fibers with DNA characteristics. II. Enzymatic and other hydrolytic treatments. J Cell Biol 19:613–629
Naumann J, Salomo K, Der JP et al (2013) Single-copy nuclear genes place haustorial Hydnoraceae within Piperales and reveal a cretaceous origin of multiple parasitic angiosperm lineages. PLoS One 8(11):e79204
Naumann J, Der JP, Wafula EK et al (2016) Detecting and characterizing the highly divergent plastid genome of the nonphotosynthetic parasitic plant Hydnora visseri (Hydnoraceae). Genome Biol Evol 8(2):345–363
Nowack EC (2014) Paulinella chromatophora-rethinking the transition from endosymbiont to organelle. Acta Societ Botanicorum Poloniae 83(4)
Nowack EC, Grossman AR (2012) Trafficking of protein into the recently established photosynthetic organelles of Paulinella chromatophora. Proc Natl Acad Sci 109(14):5340–5345
Nowack EC, Melkonian M, Glöckner G (2008) Chromatophore genome sequence of Paulinella sheds light on acquisition of photosynthesis by eukaryotes. Curr Biol 18(6):410–418
Oborník M, Green BR (2005) Mosaic origin of the heme biosynthesis pathway in photosynthetic eukaryotes. Mol Biol Evol 22:2343–2353
Ohlrogge J, Browse J (1995) Lipid biosynthesis. Plant Cell 7(7):957
Parfrey LW, Lahr DJ, Knoll AH, Katz LA (2011) Estimating the timing of early eukaryotic diversification with multigene molecular clocks. Proc Natl Acad Sci 108(33):13624–13629
Petersen J, Ludewig AK, Michael V et al (2014) Chromera velia, endosymbioses and the rhodoplex hypothesis—plastid evolution in cryptophytes, alveolates, stramenopiles, and haptophytes (CASH lineages). Genome Biol Evol 6(3):666–684
Pombert JF, Blouin NA, Lane C, Boucias D, Keeling PJ (2014) A lack of parasitic reduction in the obligate parasitic green alga Helicosporidium. PLoS Genet 10(5):e1004355
Ponce-Toledo RI, Deschamps P, López-García P et al (2017) An early-branching freshwater cyanobacterium at the origin of plastids. Curr Biol 27(3):386–391
Popot JL, de Vitry C (1990) On the microassembly of integral membrane proteins. Annu Rev Biophys Biophys Chem 19:369–403
Ralph SA, van Dooren G, Waller R et al (2004) Metabolic maps and functions of the Plasmodium falciparum apicoplast. Nat Rev Microbiol 2:203–216
Ris H, Plaut W (1962) Ultrastructure of DNA-containing areas in the chloroplast of Chlamydomonas. J Cell Biol 13:383–391
Robledo JAF, Caler E, Matsuzaki M et al (2011) The search for the missing link: a relic plastid in Perkinsus? Int J Parasitol 41(12):1217–1229
Rockwell NC, Lagarias JC, Bhattacharya D (2014) Primary endosymbiosis and the evolution of light and oxygen sensing in photosynthetic eukaryotes. Front Ecol Evol 2(66). doi:10.3389/fevo.2014.00066
Rowan R, Whitney SM, Fowler A, Yellowlees D (1996) Rubisco in marine symbiotic dinoflagellates: form II enzymes in eukaryotic oxygenic phototrophs encoded by a nuclear multigene family. Plant Cell (3):539–553
Ruiz-Nieto JE, Aguirre-Mancilla CL, Acosta-Gallegos JA et al (2015) Photosynthesis and chloroplast genes are involved in water-use efficiency in common bean. Plant Physiol Biochem 86:166–173
Sagan L (1967) On the origin of mitosing cells. J Theor Biol 14(3):225IN1–274IN6
Salomaki ED, Lane CE (2014) Are all red algal parasites cut from the same cloth? Acta Societ Botanicorum Poloniae 83(4):369–375
Salomaki ED, Nickles KR, Lane CE (2015) The ghost plastid of Choreocolax polysiphoniae. J Phycol 51(2):217–221
Samigullin TH, Logacheva MD, Penin AA, Vallejo-Roman CM (2016) Complete plastid genome of the recent holoparasite Lathraea squamaria reveals earliest stages of plastome reduction in Orobanchaceae. PLoS One 11:e0150718
Sánchez-Puerta MV, Lippmeier JC, Apt KE, Delwiche CF (2007) Plastid genes in a non-photosynthetic dinoflagellate. Protist 158:105–117
Sasaki Y, Nagano Y (2004) Plant acetyl-CoA carboxylase: structure, biosynthesis, regulation, and gene manipulation for plant breeding. Biosci Biotech Biochem 68(6):1175–1184
Schelkunov MI, Shtratnikova VY, Nuraliev MS et al (2015) Exploring the limits for reduction of plastid genomes: a case study of the mycoheterotrophic orchids Epipogium aphyllum and Epipogium roseum. Genome Biol Evol 7:1179–1191
Schleiff E, Becker T (2011) Common ground for protein translocation: access control for mitochondria and chloroplasts. Nat Rev Mol Cell Biol 12(1):48–59
Schnepf E, Elbrächter M (1988) Cryptophycean-like double membrane-bound chloroplast in the dinoflagellate, Dinophysis Ehrenb.: evolutionary, phylogenetic and toxicological implications. Plant Biol 101(2):96–203
Schwender J, Goffman F, Ohlrogge JB, Shachar-Hill Y (2004) Rubisco without the Calvin cycle improves the carbon efficiency of developing green seeds. Nature 432:779–782
Seeber F, Soldati-Favre D (2010) Metabolic pathways in the apicoplast of apicomplexa. Int Rev Cell Mol Biol 281:161–228
Sekiguchi H, Moriya M, Nakayama T, Inouye I (2002) Vestigial chloroplasts in heterotrophic stramenopiles Pteridomonas danica and Ciliophrys infusionum (Dictyophyceae). Protist 153:157–167
Sheiner L, Striepen B (2013) Protein sorting in complex plastids. Biochim Biophys Acta (BBA) Mol Cell Res 1833(2):352–359
Shiina T, Tsunoyama Y, Nakahira Y, Khan MS (2005) Plastid RNA polymerases, promoters, and transcription regulators in higher plants. Int Rev Cytol 244:1–68
Sickmann A, Reinders J, Wagner Y et al (2003) The proteome of Saccharomyces cerevisiae mitochondria. Proc Natl Acad Sci 100(23):13207–13212
Slamovits CH, Keeling PJ (2008) Plastid-derived genes in the nonphotosynthetic alveolate Oxyrrhis marina. Mol Biol Evol 25(7):1297–1306
Smith DR, Asmail SR (2014) Next-generation sequencing data suggest that certain nonphotosynthetic green plants have lost their plastid genomes. New Phytol 204:7–11
Smith DR, Lee RW (2014) A plastid without a genome: evidence from the nonphotosynthetic green algal genus Polytomella. Plant Physiol 164:1812–1819
Smith AC, Purton S (2002) The transcriptional apparatus of plastids. Eur J Phycol 37:301–311
Smith DR, Hua J, Lee RW (2010) Evolution of linear mitochondrial DNA in three known lineages of Polytomella. Curr Genet 56:427–438
Smith DR, Crosby K, Lee RW (2011) Correlation between nuclear plastid DNA abundance and plastid number supports the limited transfer window hypothesis. Genome Biol Evol 3:365–371
Stegemann S, Hartmann S, Ruf S, Bock R (2003) High-frequency gene transfer from the chloroplast genome to the nucleus. Proc Natl Acad Sci 100(15):8828–8833
Stelter K, El-Sayed NM, Seeber F (2007) The expression of a plant-type ferredoxin redox system provides molecular evidence for a plastid in the early dinoflagellate Perkinsus marinus. Protist 158:119–130
Stiller JW (2014) Toward an empirical framework for interpreting plastid evolution. J Phycol 50(3):462–471
Takishita K, Koike K, Maruyama T, Ogata T (2002) Molecular evidence for plastid robbery (Kleptoplastidy) in Dinophysis, a dinoflagellate causing diarrhetic shellfish poisoning. Protist 153:293–302
Tanifuji G, Onodera NT, Wheeler TJ et al (2011) Complete nucleomorph genome sequence of the nonphotosynthetic alga Cryptomonas paramecium reveals a core nucleomorph gene set. Genome Biol Evol 3:44–54
Teles-Grilo ML, Tato-Costa J, Duarte SM et al (2007) Is there a plastid in Perkinsus atlanticus (Phylum Perkinsozoa)? Eur J Protistol 43(2):163–167
Templeton TJ, Enomoto S, Chen W-J et al (2010) A genome-sequence survey for Ascogregarina taiwanensis supports evolutionary affiliation but metabolic diversity between a gregarine and Cryptosporidium. Mol Biol Evol 27:235–248
Tengs T, Dahlberg OJ, Shalchian-Tabrizi K et al (2000) Phylogenetic analyses indicate that the 19′ hexanoyloxy-fucoxanthin-containing dinoflagellates have tertiary plastids of haptophyte origin. Mol Biol Evol 17(5):718–729
Terashima M, Specht M, Hippler M (2011) The chloroplast proteome: a survey from the Chlamydomonas reinhardtii perspective with a focus on distinctive features. Curr Genet 57(3):151–168
Toso MA, Omoto CK (2007) Gregarina niphandrodes may lack both a plastid genome and organelle. J Eukaryot Microbiol 54(1):66–72
Triemer RE, Linton E, Shin W et al (2006) Phylogeny of the Euglenales based upon combined SSU and LSU rDNA sequence comparisons and description of Discoplastis gen. nov. (Euglenophyta). J Phycol 42:731–740
Vernon D, Gutell RR, Cannone JJ, Rumpf RW, Birky CW (2001) Accelerated evolution of functional plastid rRNA and elongational factor genes due to reduced protein synthetic load after the loss of photosynthesis in the chlorophyte alga Polytoma. Mol Biol Evol 18:1810–1822
Vesteg M, Vacula R, Krajčovič J (2009) On the origin of chloroplasts, import mechanisms of chloroplast-targeted proteins, and loss of photosynthetic ability. Folia Microbiol 54:303–321
Waller RF, Keeling PJ, Donald RGK et al (1998) Nuclear encoded proteins target to the plastid in Toxoplasma gondii and Plasmodium falciparum. Cell Biol 95:12352–12357
Wallin IE (1927) Symbionticism and the origin of species. Bailliere, Tindall and Cox 171, London
Watanabe MM, Suda S, Inouya I, Sawaguchi T, Chihara M (1990) Lepidodinium viride gen. et sp. nov. (Gymnodinaiales, Dinophyta), a green dinoflagellate with a chlorophyll a- and b-containing endosymbiont. J Phycol 26(4):741–751
Whitney SM, Shaw DC, Yellowlees D (1995) Evidence that Some dinoflagellates contain a ribulose-1,5-bisphosphate carboxylase/oxygenase related to that of the $\alpha $-proteobacteria. Proc R Soc Lond B Biol Sci 259(1356):271–275
Wicke S, Müller KF, dePamphilis CW, Quandt D, Wickett NJ, Zhang Y, Renner SS, Schneeweiss GM (2013) Mechanisms of functional and physical genome reduction in photosynthetic and nonphotosynthetic parasitic plants of the broomrape family. Plant Cell 25:3711–3725
Wicke S, Müller KF, Quandt D, Bellot S, Schneeweiss GM (2016) Mechanistic model of evolutionary rate variation en route to a nonphotosynthetic lifestyle in plants. Proc Natl Acad Sci USA 113(32):9045–9050
Wickett NJ, Zhang Y, Hansen SK et al (2008) Functional gene losses occur with minimal size reduction in the plastid genome of the parasitic liverwort Aneura mirabilis. Mol Biol Evol 25:393–401
Williams BA, Keeling PJ (2003) Cryptic organelles in parasitic protists and fungi. Adv Parasitol 54:9–68
Wilson RJM (2004) Plastid functions in the Apicomplexa. Protist 155:11–12
Wilson RJM (2005) Parasite plastids: approaching the endgame. Biol Rev Camb Philos Soc 80:129–153
Wilson RJM, Williamson DH (1997) Extrachromosomal DNA in the Apicomplexa. Microbiol Mol Biol Rev 61:1–16
Wilson RJM, Denny PW, Preiser PR et al (1996) Complete gene map of the plastid-like DNA of the malaria parasite Plasmodium falciparum. J Mol Biol 261:155–172
Wolfe A, de Pamphilis CW (1998) The effect of relaxed functional constraints on the photosynthetic gene rbcL in photosynthetic and nonphotosynthetic parasitic plants. Mol Biol Evol 15:1243–1258
Wolfe KH, Morden CW, Palmer JD (1992) Function and evolution of a minimal plastid genome from a nonphotosynthetic parasitic plant. Proc Natl Acad Sci USA 89:10648–10652
Yamada N, Sym SD, Horiguchi T (2017) Identification of highly divergent diatom-derived chloroplasts in dinoflagellates, including a description of Durinskia kwazulunatalensis sp. nov. (Peridiniales, Dinophyceae). Mol Biol Evol 34:1335–1351
Yan D, Wang Y, Murakami T et al (2015) Auxenochlorella protothecoides and Prototheca wickerhamii plastid genome sequences give insight into the origins of non-photosynthetic algae. Sci Rep 5:14465
Yang D, Oyaizu Y, Oyaizu H, Olsen GJ, Woese CR (1985) Mitochondrial origins. Proc Natl Acad Sci USA 82:4443–4447
Yeh E, DeRisi JL (2011) Chemical rescue of malaria parasites lacking an apicoplast defines organelle function in blood-stage Plasmodium falciparum. PLoS Biol 9(8):e1001138
Yoon HS, Hackett JD, Ciniglia C, Pinto G, Bhattacharya D (2004) A molecular timeline for the origin of photosynthetic eukaryotes. Mol Biol Evol 21(5):809–818
Yoon HS, Reyes-Prieto A, Melkonian M, Bhattacharya D (2006) Minimal plastid evolution in the Paulinella endosymbiont. Curr Biol 16:R670-R672
Yubuki N, Nakayama T, Inouye I (2008) A unique life cycle and perennation in a colorless chrysophyte Spumella sp. J Phycol 44(1):164–172
Záhonová K, Füssy Z, Oborník M, Eliáš M, Yurchenko V (2016) RuBisCO in non-photosynthetic alga Euglena longa: divergent features, transcriptomic analysis and regulation of complex formation. PLoS One 11(7):e0158790
Zhu G, Marchewka MJ, Keithly JS (2000) Cryptosporidium parvum appears to lack a plastid genome. Microbiol 146:315–321
Zíková A, Hampl V, Paris Z, Týč J, Lukeš J (2016) Aerobic mitochondria of parasitic protists: diverse genomes and complex functions. Mol Biochem Parasitol 209(1):46–57
Zimorski V, Ku C, Martin WF, Gould SB (2014) Endosymbiotic theory for organelle origins. Curr Opin 22C:38–48
Acknowledgements
This work was supported by the Scientific Grant Agency of the Slovak Ministry of Education and the Academy of Sciences (Grant VEGA 1/0535/17), by project ITMS 26210120024 supported by the Research & Development Operational Programme funded by the ERDF, by the Czech Science foundation Project nr. 16-25280S, by the Ministry of Education, Youth and Sports of CR within the National Sustainability Program II (Project BIOCEV-FAR) LQ1604 and by the project ‘‘BIOCEV’’ (CZ.1.05/1.1.00/02.0109). We thank Dr. Heather Esson (Laboratory of Evolutionary Protistology, Institute of Parasitology, Biology Centre CAS, České Budějovice, Czech Republic) for language revision of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by M. Kupiec.
Electronic supplementary material
Below is the link to the electronic supplementary material.
294_2017_761_MOESM1_ESM.pdf
Online Resource 1 Characteristics of plastid genomes in organisms with non-photosynthetic plastids. Plastid genome sizes – "0 bp" indicates that there is probably no plastid genome present. Gene number includes only protein-coding genes. rRNA genes, tRNA genes, pseudogenes and orfs are not included. Number of photosynthetic genes comprises psa, psb, pet and rbcL genes. IRs (inverted repeats) –"Yes" indicates presence, "No" indicates absence, "Cryptic" indicates the possibility of IRs existence, "Secondary" indicates that original IRs have been lost and new ones have arisen secondarily. A question mark indicates the lack of data (PDF 334 KB)
Rights and permissions
About this article
Cite this article
Hadariová, L., Vesteg, M., Hampl, V. et al. Reductive evolution of chloroplasts in non-photosynthetic plants, algae and protists. Curr Genet 64, 365–387 (2018). https://doi.org/10.1007/s00294-017-0761-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-017-0761-0